Submitted:
20 April 2026
Posted:
27 April 2026
You are already at the latest version
Abstract
Keywords:
1. Introduction
2. Materials and Methods
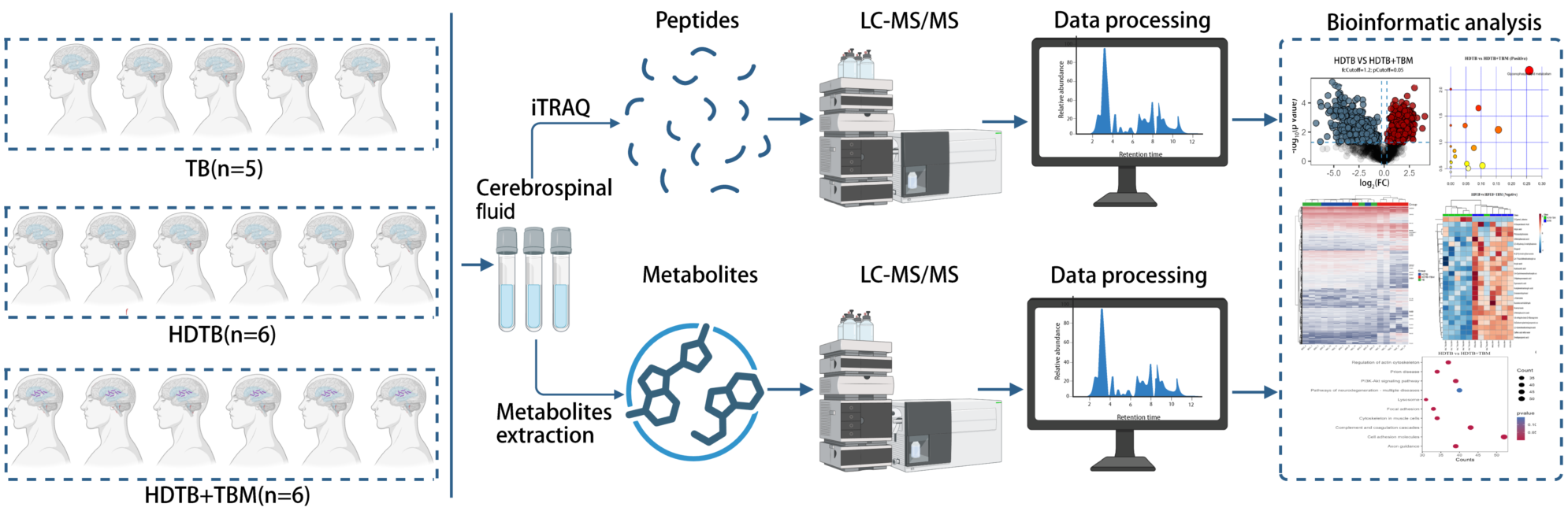
2.1. Participants and Cerebrospinal Fluid Collection
2.2. Proteomics Analysis
2.3. Non-Targeted Metabolomics Analysis
2.4. Integrated Analysis
2.4. Statistical Analysis
3. Results
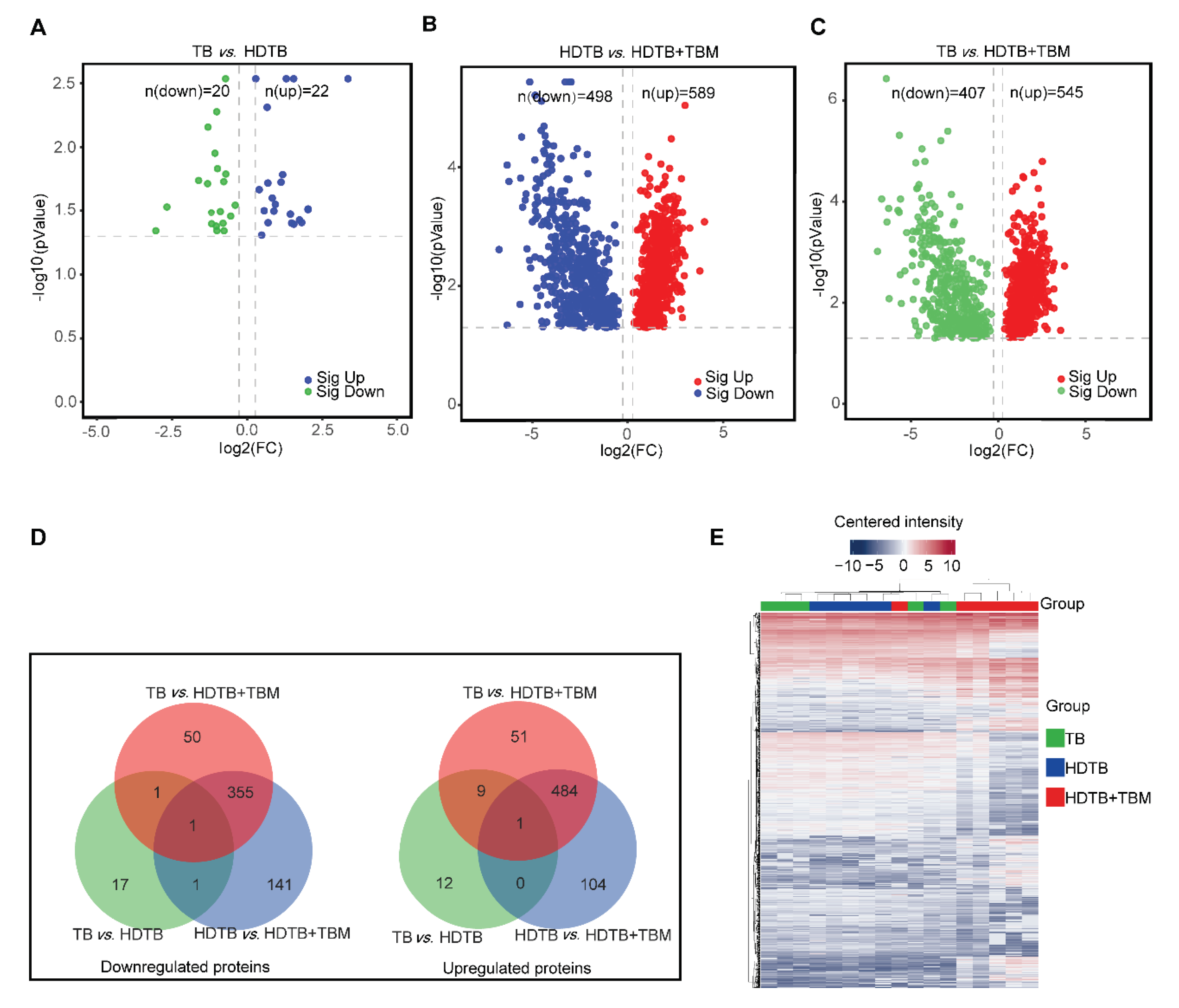
3.1. SubsectionCSF Proteomic Profiling Reveals Stage-Dependent Molecular Remodeling During Tuberculosis Progression
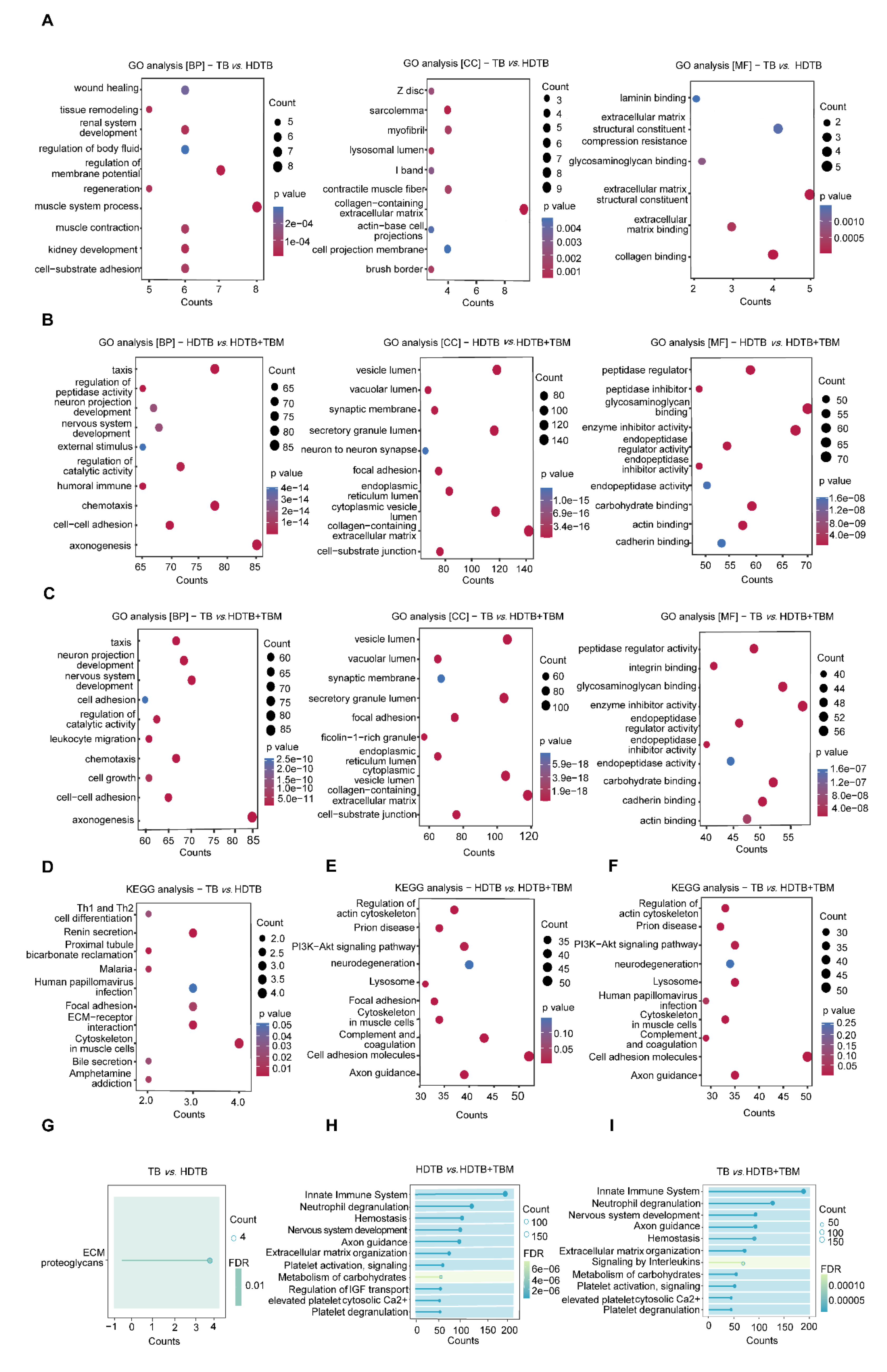
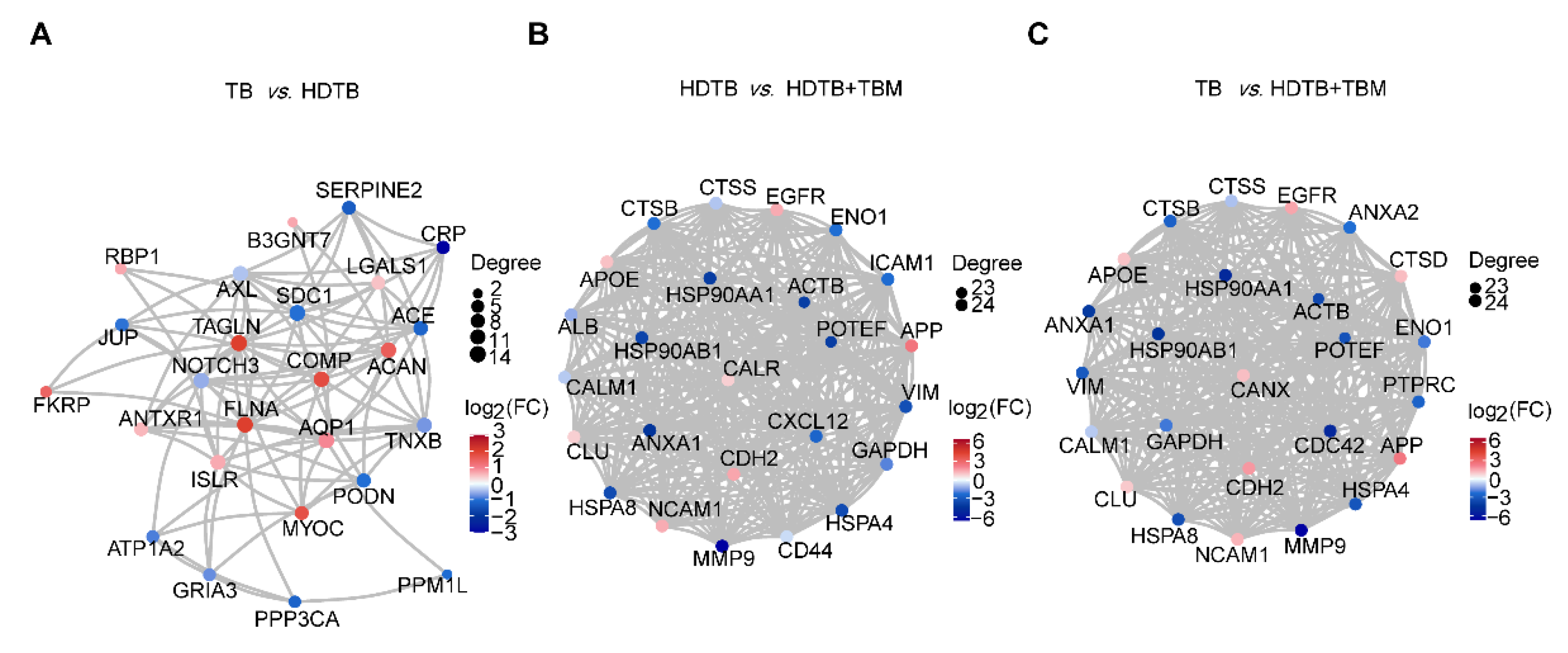
3.2. Functional Enrichment and Network Analyses Link TBM-Associated Proteomic Changes to Neuro-Immune and Vascular Pathways
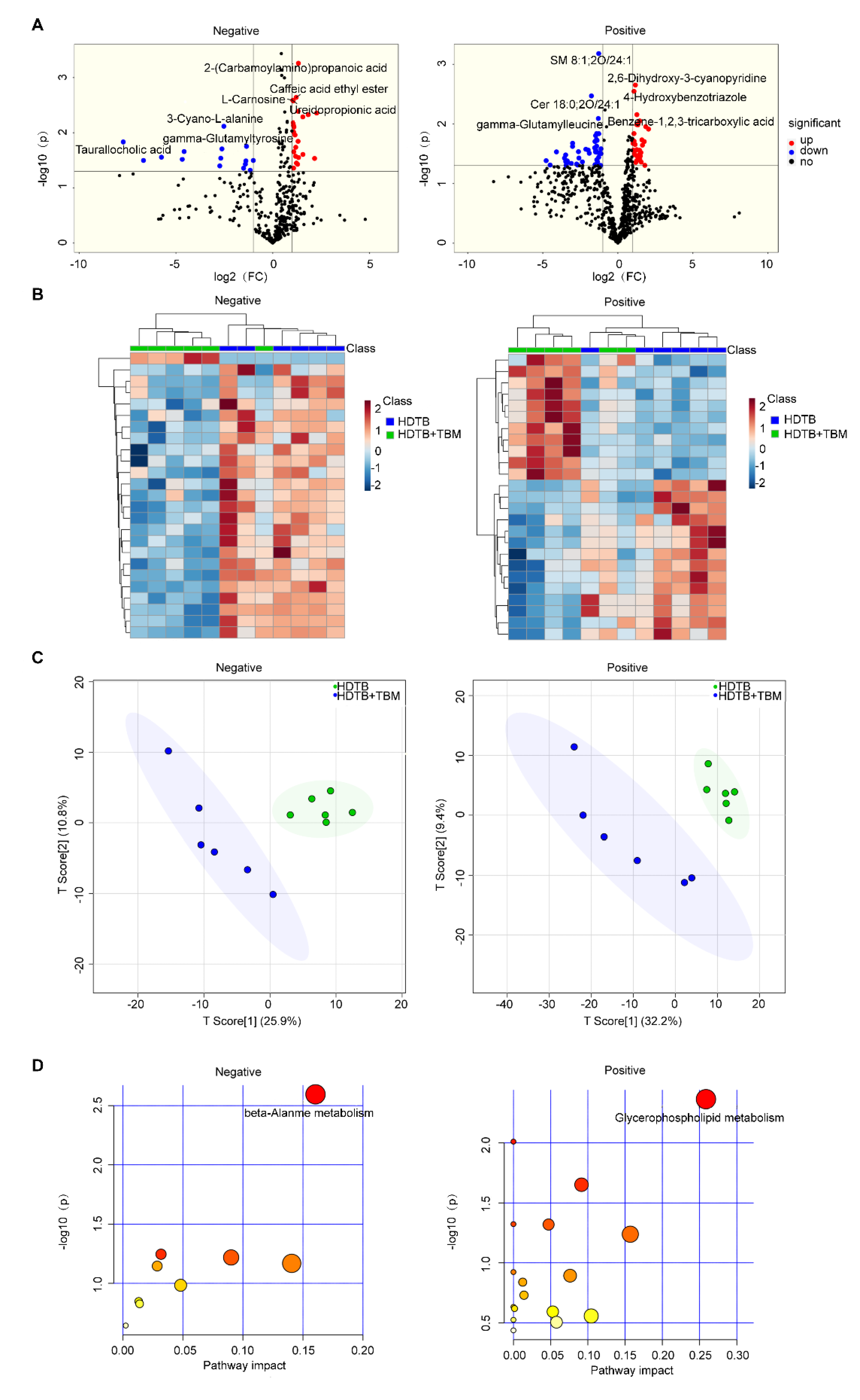
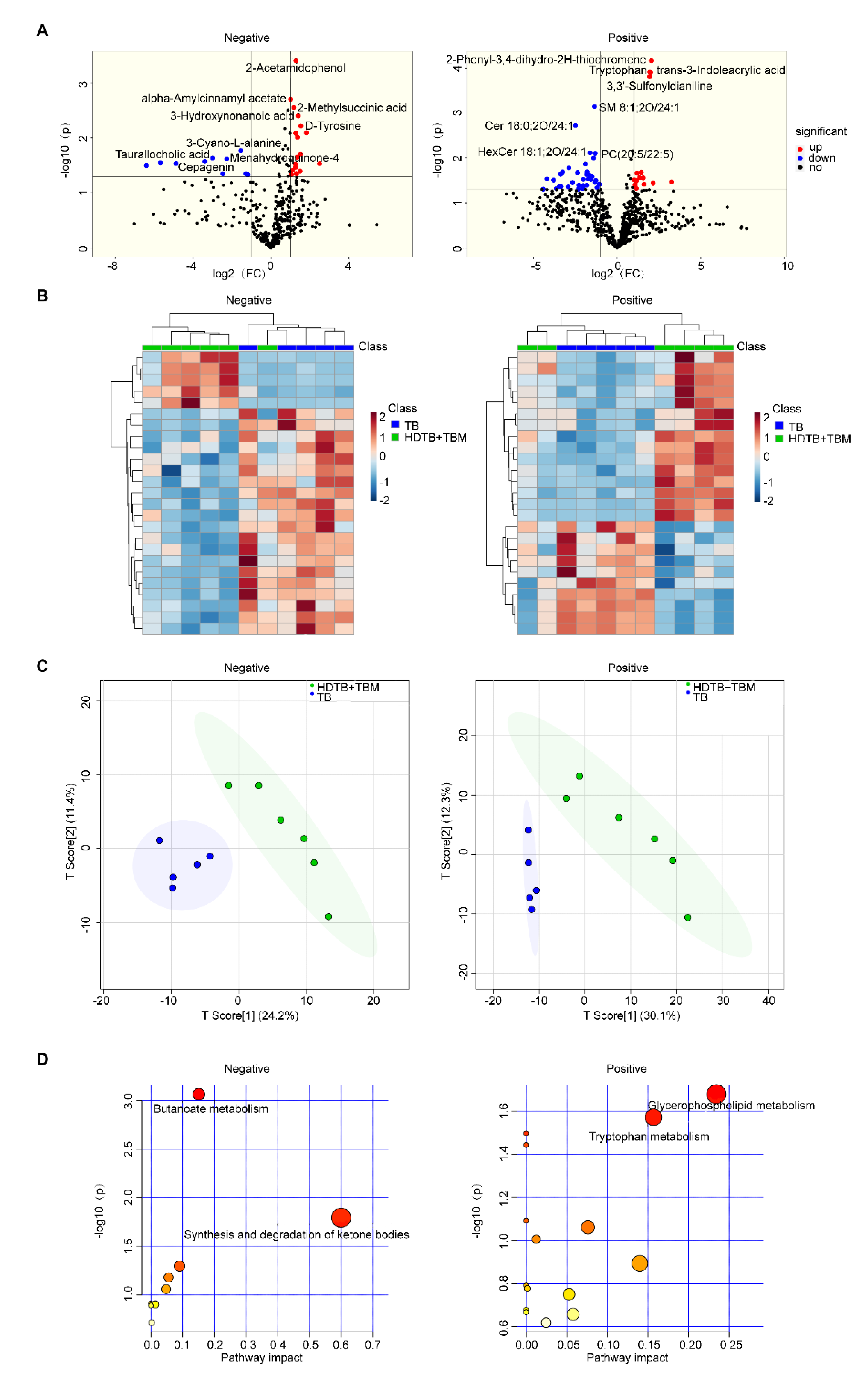
3.3. CSF Metabolomic Profiling Highlights Metabolic Reprogramming During Transition to TBM
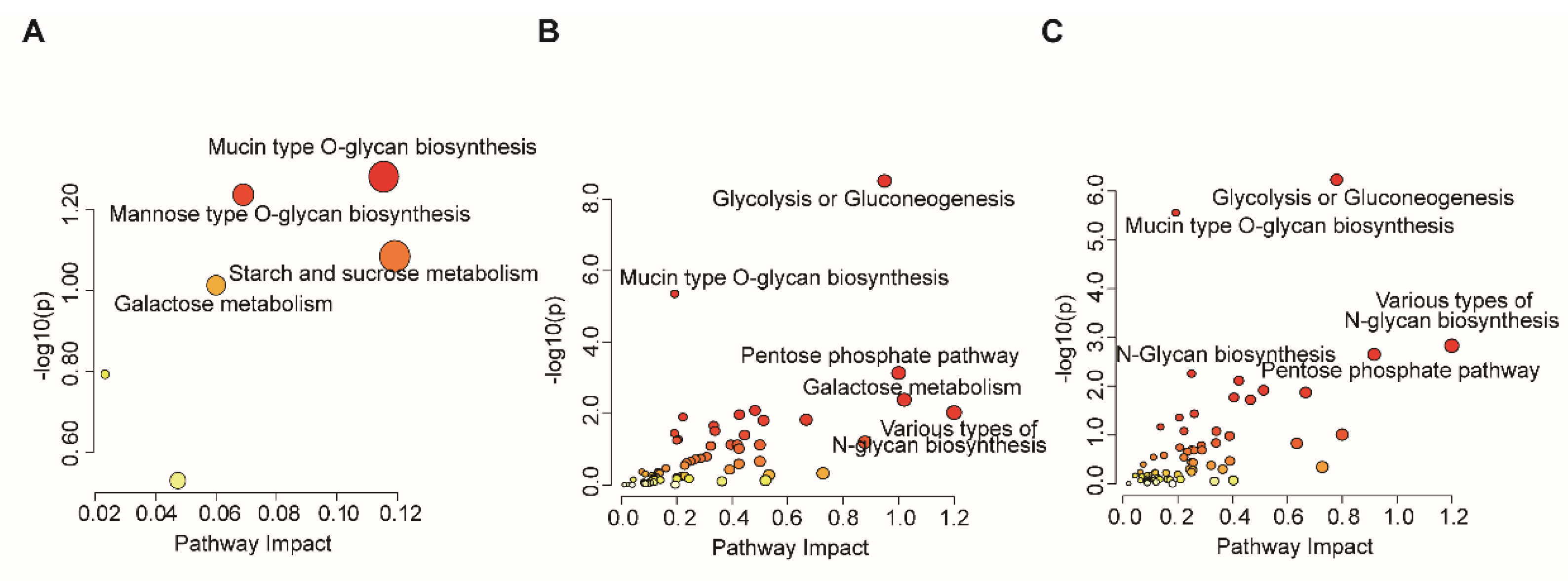
3.4. Integrated Multi-Omics Analysis Identifies Coordinated Metabolic and Glycosylation Rewiring During Meningeal Involvement
4. Discussion
5. Conclusions
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Conflicts of Interest
Abbreviations
| CSF | Cerebrospinal fluid |
| TBM | Tuberculous meningitis |
| TB | Tuberculosis |
| HDTB | Hematogenous disseminated tuberculosis |
| iTRAQ | Isotope-labeling relative and absolute quantification techniques |
| ApoB | Apolipoprotein B |
| FASP | Filter-aided sample preparation |
| LC-MS/MS | Liquid chromatography-tandem mass spectrometry |
| FC | Fold changes |
| GO | Gene Ontology |
| KEGG | Kyoto Encyclopedia of Genes and Genomes |
| HESI | Heated Electrospray Ionization |
| SEM | Standard error of the mean |
| DEPs | Differentially expressed proteins |
| DEMs | Differential metabolites |
| PLS-DA | Partial least squares discriminant analysis |
| BP | Biological process |
| CC | Cellular component |
| MF | Molecular function |
| PPI | Protein-protein interaction |
References
- Goletti, D.; Meintjes, G.; Andrade, B.B.; Zumla, A.; Shan Lee, S. Insights from the 2024 WHO Global Tuberculosis Report - More Comprehensive Action, Innovation, and Investments required for achieving WHO End TB goals. Int. J. Infect. Dis. 2025, 150, 107325. [Google Scholar] [CrossRef]
- Mehta, K.; Balazki, P.; van der Graaf, P.H.; Guo, T.; van Hasselt, J.G.C. Predictions of Bedaquiline Central Nervous System Exposure in Patients with Tuberculosis Meningitis Using Physiologically based Pharmacokinetic Modeling. Clin. Pharmacokinet. 2024, 63, 657–68. [Google Scholar] [CrossRef] [PubMed]
- Wilkinson, R.J.; Rohlwink, U.; Misra, U.K.; van Crevel, R.; Mai, N.T.H.; Dooley, K.E.; et al. Tuberculous meningitis. Nat. Rev. Neurol. 2017, 13, 581–98. [Google Scholar] [CrossRef]
- Mason, S.; Reinecke, C.J.; Solomons, R. Cerebrospinal Fluid Amino Acid Profiling of Pediatric Cases with Tuberculous Meningitis. Front Neurosci. 2017, 11, 534. [Google Scholar] [CrossRef] [PubMed]
- Niu, M.; Bai, Z.; Dong, L.; Zheng, W.; Wang, X.; Dong, N.; et al. A Novel Diagnostic Prediction Model for Distinguishing Between Tuberculous and Cryptococcal Meningitis. Clin. Med. Res. 2024, 22, 197–205. [Google Scholar] [CrossRef]
- Barreras, P.; Stern, B.J. Clinical features and diagnosis of neurosarcoidosis - review article. J. Neuroimmunol. 2022, 368, 577871. [Google Scholar] [CrossRef]
- Basu, S.; Chakraborty, S. A Comprehensive Review of the Diagnostics for Pediatric Tuberculosis Based on Assay Time, Ease of Operation, and Performance. Microorganisms 2025, 13. [Google Scholar] [CrossRef]
- Boudry, F.; Durand, F.; Goossens, C. Metabolomics Signatures of a Respiratory Tract Infection During an Altitude Training Camp in Elite Rowers. Metabolites 2025, 15. [Google Scholar] [CrossRef]
- Ganna, A.; Salihovic, S.; Sundstrom, J.; Broeckling, C.D.; Hedman, A.K.; Magnusson, P.K.; et al. Large-scale metabolomic profiling identifies novel biomarkers for incident coronary heart disease. PLoS Genet 2014, 10, e1004801. [Google Scholar] [CrossRef] [PubMed]
- Liesenfeld, D.B.; Habermann, N.; Owen, R.W.; Scalbert, A.; Ulrich, C.M. Review of mass spectrometry-based metabolomics in cancer research. Cancer Epidemiol. Biomark. Prev. 2013, 22, 2182–201. [Google Scholar] [CrossRef]
- Dai, Y.N.; Huang, H.J.; Song, W.Y.; Tong, Y.X.; Yang, D.H.; Wang, M.S.; et al. Identification of potential metabolic biomarkers of cerebrospinal fluids that differentiate tuberculous meningitis from other types of meningitis by a metabolomics study. Oncotarget 2017, 8, 100095–112. [Google Scholar] [CrossRef]
- Mu, J.; Yang, Y.; Chen, J.; Cheng, K.; Li, Q.; Wei, Y.; et al. Elevated host lipid metabolism revealed by iTRAQ-based quantitative proteomic analysis of cerebrospinal fluid of tuberculous meningitis patients. Biochem Biophys. Res. Commun. 2015, 466, 689–95. [Google Scholar] [CrossRef]
- Samuel, V.; Solomons, R.; Mason, S. Targeted metabolomics investigation of metabolic markers of Mycobacterium tuberculosis in the cerebrospinal fluid of paediatric patients with tuberculous meningitis. PLoS ONE 2024, 19, e0314854. [Google Scholar] [CrossRef] [PubMed]
- Yang, Y.; Mu, J.; Chen, G.; Zhan, Y.; Zhong, J.; Wei, Y.; et al. iTRAQ-based quantitative proteomic analysis of cerebrospinal fluid reveals NELL2 as a potential diagnostic biomarker of tuberculous meningitis. Int. J. Mol. Med. 2015, 35, 1323–32. [Google Scholar] [CrossRef] [PubMed]
- Huang, M.; Ding, Z.; Li, W.; Chen, W.; Du, Y.; Jia, H.; et al. Identification of protein biomarkers in host cerebrospinal fluid for differential diagnosis of tuberculous meningitis and other meningitis. Front Neurol. 2022, 13, 886040. [Google Scholar] [CrossRef]
- Ou, Q.; Liu, X.; Cheng, X. An iTRAQ approach to quantitative proteome analysis of cerebrospinal fluid from patients with tuberculous meningitis. Biosci. Trends 2013, 7, 186–92. [Google Scholar] [CrossRef] [PubMed]
- Mason, S.; Solomons, R. CSF Metabolomics of Tuberculous Meningitis: A Review. Metabolites 2021, 11. [Google Scholar] [CrossRef]
- Enche Ady, C.N.A.; Lim, S.M.; Teh, L.K.; Salleh, M.Z.; Chin, A.V.; Tan, M.P.; et al. Metabolomic-guided discovery of Alzheimer’s disease biomarkers from body fluid. J. Neurosci. Res. 2017, 95, 2005–24. [Google Scholar] [CrossRef]
- Kim, S.G.; Hwang, J.S.; George, N.P.; Jang, Y.E.; Kwon, M.; Lee, S.S.; et al. Integrative Metabolome and Proteome Analysis of Cerebrospinal Fluid in Parkinson’s Disease. Int. J. Mol. Sci. 2024, 25. [Google Scholar] [CrossRef]
- Niu, D.; Sun, P.; Zhang, F.; Song, F. Metabonomic analysis of cerebrospinal fluid in epilepsy. Ann. Transl. Med. 2022, 10, 449. [Google Scholar] [CrossRef]
- Ardiansyah, E.; Avila-Pacheco, J.; Nhat, L.T.H.; Dian, S.; Vinh, D.N.; Hai, H.T.; et al. Tryptophan metabolism determines outcome in tuberculous meningitis: a targeted metabolomic analysis. Elife 2023, 12. [Google Scholar] [CrossRef]
- Shi, Y.; Zhang, C.; Pan, S.; Chen, Y.; Miao, X.; He, G.; et al. The diagnosis of tuberculous meningitis: advancements in new technologies and machine learning algorithms. Front Microbiol. 2023, 14, 1290746. [Google Scholar] [CrossRef]
- Zhang, P.; Zhang, W.; Lang, Y.; Qu, Y.; Chu, F.; Chen, J.; et al. Mass spectrometry-based metabolomics for tuberculosis meningitis. Clin. Chim. Acta 2018, 483, 57–63. [Google Scholar] [CrossRef] [PubMed]
- Cresswell, F.V.; Davis, A.G.; Sharma, K.; Basu Roy, R.; Ganiem, A.R.; Kagimu, E.; et al. Recent Developments in Tuberculous Meningitis Pathogenesis and Diagnostics. Wellcome Open. Res. 2019, 4, 164. [Google Scholar] [CrossRef] [PubMed]
- Machida, N.; Arai, T.; Noguchi, K.; Oki, Y. [Change of incidence of antinuclear antibodies in serum of NOD mouse with aging]. Jikken Dobutsu 1989, 38, 345–7. [Google Scholar] [PubMed]
- Ratinam, J.; Mishra, A.K.; Muthuram, A.J.; Miraclin, A.; Chandy, G.M.; Vanjare, H.A.; et al. Role of cerebrospinal fluid C-reactive protein in tuberculous meningitis. Int. J. Mycobacteriol 2020, 9, 422–8. [Google Scholar]
- Nacarapa, E.; Munyangaju, I.; Osorio, D.; Ramos-Rincon, J.M. Predictors of Tuberculous Meningitis Mortality Among Persons with HIV in Mozambique. Trop. Med. Infect. Dis. 2025, 10. [Google Scholar] [CrossRef]
- Nasiri, M.J.; Lutfy, K.; Venketaraman, V. Challenges of Multidrug-Resistant Tuberculosis Meningitis: Current Treatments and the Role of Glutathione as an Adjunct Therapy. Vaccines 2024, 12. [Google Scholar] [CrossRef]
- Galimi, R. Extrapulmonary tuberculosis: tuberculous meningitis new developments. Eur. Rev. Med. Pharmacol. Sci. 2011, 15, 365–86. [Google Scholar]
- Smith, S.; Anderson, S.; Ballermann, B.J.; Brenner, B.M. Role of atrial natriuretic peptide in adaptation of sodium excretion with reduced renal mass. J. Clin. Invest 1986, 77, 1395–8. [Google Scholar] [CrossRef]
- Hu, X.; Liao, S.; Bai, H.; Wu, L.; Wang, M.; Wu, Q.; et al. Integrating exosomal microRNAs and electronic health data improved tuberculosis diagnosis. EBioMedicine 2019, 40, 564–73. [Google Scholar] [CrossRef] [PubMed]
- Nuwagira, E.; Huppler Hullsiek, K.; Jjunju, S.; Rutakingirwa, M.; Kasibante, J.; Tadeo, K.K.; et al. Diagnostic and Prognostic Value of Cerebrospinal Fluid Lactate and Glucose in HIV-Associated Tuberculosis Meningitis. Microbiol. Spectr. 2022, 10, e0161822. [Google Scholar] [CrossRef]
- Pan, L.; Liu, F.; Zhang, J.; Li, J.; Jia, H.; Huang, M.; et al. Genome-Wide miRNA Analysis Identifies Potential Biomarkers in Distinguishing Tuberculous and Viral Meningitis. Front Cell. Infect. Microbiol. 2019, 9, 323. [Google Scholar] [CrossRef] [PubMed]
- Wei, J.; Zhu, L.; Jian, B.; Yu, Y.; Hu, B.; Guo, L.; et al. Proteomic profiling of cerebrospinal fluid uncovers distinctive pathophysiological changes and potential biomarkers in pediatric tubercular meningitis. Front Cell. Infect. Microbiol. 2025, 15, 1662783. [Google Scholar] [CrossRef] [PubMed]
- Lin, F. Tuberculous meningitis diagnosis and treatment: classic approaches and high-throughput pathways. Front Immunol. 2024, 15, 1543009. [Google Scholar] [CrossRef]
- Manyelo, C.M.; Solomons, R.S.; Walzl, G.; Chegou, N.N. Tuberculous Meningitis: Pathogenesis, Immune Responses, Diagnostic Challenges, and the Potential of Biomarker-Based Approaches. J. Clin. Microbiol. 2021, 59. [Google Scholar] [CrossRef]
- Chen, H.L.; Lu, C.H.; Chang, C.D.; Chen, P.C.; Chen, M.H.; Hsu, N.W.; et al. Structural deficits and cognitive impairment in tuberculous meningitis. BMC Infect. Dis. 2015, 15, 279. [Google Scholar] [CrossRef]
- Jansson, D.; Wang, M.; Thomas, R.G.; Erickson, M.A.; Peskind, E.R.; Li, G.; et al. Markers of Cerebrovascular Injury, Inflammation, and Plasma Lipids Are Associated with Alzheimer’s Disease Cerebrospinal Fluid Biomarkers in Cognitively Normal Persons. J. Alzheimers Dis. 2022, 86, 813–26. [Google Scholar] [CrossRef]
- Kalita, J.; Misra, U.K.; Ranjan, P. Predictors of long-term neurological sequelae of tuberculous meningitis: a multivariate analysis. Eur. J. Neurol. 2007, 14, 33–7. [Google Scholar] [CrossRef]
- Davis, A.G.; Nightingale, S.; Springer, P.E.; Solomons, R.; Arenivas, A.; Wilkinson, R.J.; et al. Neurocognitive and functional impairment in adult and paediatric tuberculous meningitis. Wellcome Open. Res. 2019, 4, 178. [Google Scholar] [CrossRef]
- Kim, J.; Erice, C.; Rohlwink, U.K.; Tucker, E.W. Infections in the Developing Brain: The Role of the Neuro-Immune Axis. Front Neurol. 2022, 13, 805786. [Google Scholar] [CrossRef]
- Ma, Q.; Chen, J.; Kong, X.; Zeng, Y.; Chen, Z.; Liu, H.; et al. Interactions between CNS and immune cells in tuberculous meningitis. Front Immunol. 2024, 15, 1326859. [Google Scholar] [CrossRef] [PubMed]
- Yang, Z.R.; Zhang, L.F.; Zhou, B.T.; Shi, X.C.; Cao, W.; Fan, H.W.; et al. Clinical features and influencing factors of long-term prognosis in patients with tuberculous meningitis. Zhonghua Nei Ke Za Zhi 2022, 61, 764–70. [Google Scholar]
- Hachem, M.; Nacir, H. Emerging Role of Phospholipids and Lysophospholipids for Improving Brain Docosahexaenoic Acid as Potential Preventive and Therapeutic Strategies for Neurological Diseases. Int. J. Mol. Sci. 2022, 23. [Google Scholar] [CrossRef] [PubMed]
- Sarkar, C.; Lipinski, M.M. Glycerophospholipid dysregulation after traumatic brain injury. Neurochem Int. 2024, 175, 105701. [Google Scholar] [CrossRef] [PubMed]
- Yu, R.K.; Ariga, T. The role of glycosphingolipids in neurological disorders. Mechanisms of immune action. Ann. N Y Acad. Sci. 1998, 845, 285–306. [Google Scholar] [CrossRef]
- Baranano, K.W.; Hartman, A.L. The ketogenic diet: uses in epilepsy and other neurologic illnesses. Curr. Treat. Options Neurol. 2008, 10, 410–9. [Google Scholar] [CrossRef]
- Mattson, M.P. Lifelong brain health is a lifelong challenge: from evolutionary principles to empirical evidence. Ageing Res. Rev. 2015, 20, 37–45. [Google Scholar] [CrossRef]
- Murugaiyah, V.; Mattson, M.P. Neurohormetic phytochemicals: An evolutionary-bioenergetic perspective. Neurochem Int. 2015, 89, 271–80. [Google Scholar] [CrossRef]
- Zawislak, A.; Jakimowicz, P.; McCubrey, J.A.; Rakus, D. Neuron-derived transthyretin modulates astrocytic glycolysis in hormone-independent manner. Oncotarget 2017, 8, 106625–38. [Google Scholar] [CrossRef]
- Riaz, M.; Ali, Q.; Yan, L. Calcium-L-aspartate nanoparticles mitigate Boron toxicity in rice seedlings by modulating physiological, antioxidant, and cell wall mechanisms. Sci. Rep. 2025, 15, 42439. [Google Scholar] [CrossRef]
- Wu, X.; Zhang, G.; Zhang, W.; Zhou, J.; Cong, H.; Yang, G.; et al. Rumen microbiota helps Tibetan sheep obtain energy more efficiently to survive in the extreme environment of the Qinghai-Tibet Plateau. Front Microbiol. 2024, 15, 1431063. [Google Scholar] [CrossRef]
- Borsig, L.; Wolf, M.J.; Roblek, M.; Lorentzen, A.; Heikenwalder, M. Inflammatory chemokines and metastasis--tracing the accessory. Oncogene 2014, 33, 3217–24. [Google Scholar] [CrossRef]
- Davis, A.G.; Rohlwink, U.K.; Proust, A.; Figaji, A.A.; Wilkinson, R.J. The pathogenesis of tuberculous meningitis. J. Leukoc. Biol. 2019, 105, 267–80. [Google Scholar] [CrossRef]
- Isaiah, S.; Loots, D.T.; Solomons, R.; van der Kuip, M.; Tutu Van Furth, A.M.; Mason, S. Overview of Brain-to-Gut Axis Exposed to Chronic CNS Bacterial Infection(s) and a Predictive Urinary Metabolic Profile of a Brain Infected by Mycobacterium tuberculosis. Front Neurosci. 2020, 14, 296. [Google Scholar] [CrossRef]
- Deng, Q.; Ji, Y.; Liu, J.; Wen, T. Lipid reprogramming and ferroptosis crosstalk in clear cell renal cell carcinoma: metabolic vulnerabilities and therapeutic targeting. Mol. Cancer 2025, 24, 236. [Google Scholar] [CrossRef]
- Billatos, E.; Vick, J.L.; Lenburg, M.E.; Spira, A.E. The Airway Transcriptome as a Biomarker for Early Lung Cancer Detection. Clin. Cancer Res. 2018, 24, 2984–92. [Google Scholar] [CrossRef] [PubMed]
- Zhao, L.; Deng, H.; Zhang, J.; Zamboni, N.; Yang, H.; Gao, Y.; et al. Lactate dehydrogenase B noncanonically promotes ferroptosis defense in KRAS-driven lung cancer. Cell. Death Differ. 2025, 32, 632–45. [Google Scholar] [CrossRef] [PubMed]
- Dalal, K.; Yang, W.; Tian, E.; Chernish, A.; McCluggage, P.; Lara, A.J.; et al. In vivo mapping of the mouse Galnt3-specific O-glycoproteome. J. Biol. Chem. 2024, 300, 107628. [Google Scholar] [CrossRef] [PubMed]
- Sharapov, S.Z.; Timoshchuk, A.N.; Aulchenko, Y.S. Genetic control of N-glycosylation of human blood plasma proteins. Vavilovskii Zhurnal Genet Sel. 2023, 27, 224–39. [Google Scholar] [CrossRef]
- Vojta, A.; Samarzija, I.; Bockor, L.; Zoldos, V. Glyco-genes change expression in cancer through aberrant methylation. Biochim Biophys. Acta 2016, 1860, 1776–85. [Google Scholar] [CrossRef]
- Al-Habash, I.H.; Alshaeb, A.M.; Srpanova, V.B.; Alempijevic, D.; Keckarevic-Markovic, M.; Concato, M.; et al. Integrating Forensic Autopsies with Proteomic Profiling for Suicide Risk Assessment: A Comprehensive Review of Literature. Curr. Neuropharmacol. 2025, 23, 1393–404. [Google Scholar] [CrossRef]
- Wang, M.; Li, W.; Hao, J.; Gonzales, A., 3rd; Zhao, Z.; Flores, R.S.; et al. Molecularly cleavable bioinks facilitate high-performance digital light processing-based bioprinting of functional volumetric soft tissues. Nat. Commun. 2022, 13, 3317. [Google Scholar] [CrossRef]
| Variable | TB (n=5) | HDTB (n=6) | HDTB+TBM (n=6) | P Valuea | P Valueb |
|---|---|---|---|---|---|
| Age, y, mean ± SEM | 45.40±4.82 | 33.33±5.60 | 36.20±7.57 | 0.336c | 0.145c |
| BMI | 20.90±1.56 | 16.04±0.6 | 23.08±2.63 | 0.527c | 0.270c |
| Sex, female, No. (%) | 1 (20.00) | 2 (33.33) | 1 (16.67) | 0.887d | 0.621d |
| Blood routine | |||||
| WBC (109 /L), mean ± SEM | 6.76±0.84 | 6.22±0.84 | 7.14±1.11 | 0.814c | 0.666c |
| CRP ( mg/L), mean ± SEM | 18.63±12.59 | 61.79±20.07 | 21.85±6.46 | 0.805c | 0.117c |
| PCT (%),mean±SEM | 0.26±0.03 | 0.29±0.04 | 0.26±0.02 | 0.970c | 0.479c |
| CEA ( ng/mL), mean ± SEM | 3.02±0.93 | 1.73±0.41 | 1.36±0.22 | 0.039c | 0.249c |
| ESR (mm/h) , mean ± SEM | 32.2±9.68 | 43±26.26 | 55.5±11.79 | 0.195c | 0.685c |
| Cerebrospinal fluid biochemistry | |||||
| MTP-C(g/L), mean ± SEM | 0.29±0.02 | 0.26±0.08 | 0.74±0.19 | 0.047c | 0.730c |
| GLU-C (mmol/L), mean ± SEM | 4.10±0.90 | 2.90±0.10 | 2.15±0.30 | 0.039c | 0.175c |
| CL-C(mmol/L), mean ± SEM | 124.30±1.59 | 120.00±2.42 | 121.2±2.01 | 0.288c | 0.187c |
| ADA-C(u/L), mean ± SEM | 0.36±0.12 | 0.57±0.12 | 3.09±0.57 | 0.003c | 0.243c |
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2026 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license.




