Submitted:
26 April 2026
Posted:
27 April 2026
You are already at the latest version
Abstract
Keywords:
Introduction
The Bayesian Framework and Its Hidden Assumptions
Admixture as a Source of Model Misspecification
Borderline VUS Instability as a Structural Phenomenon
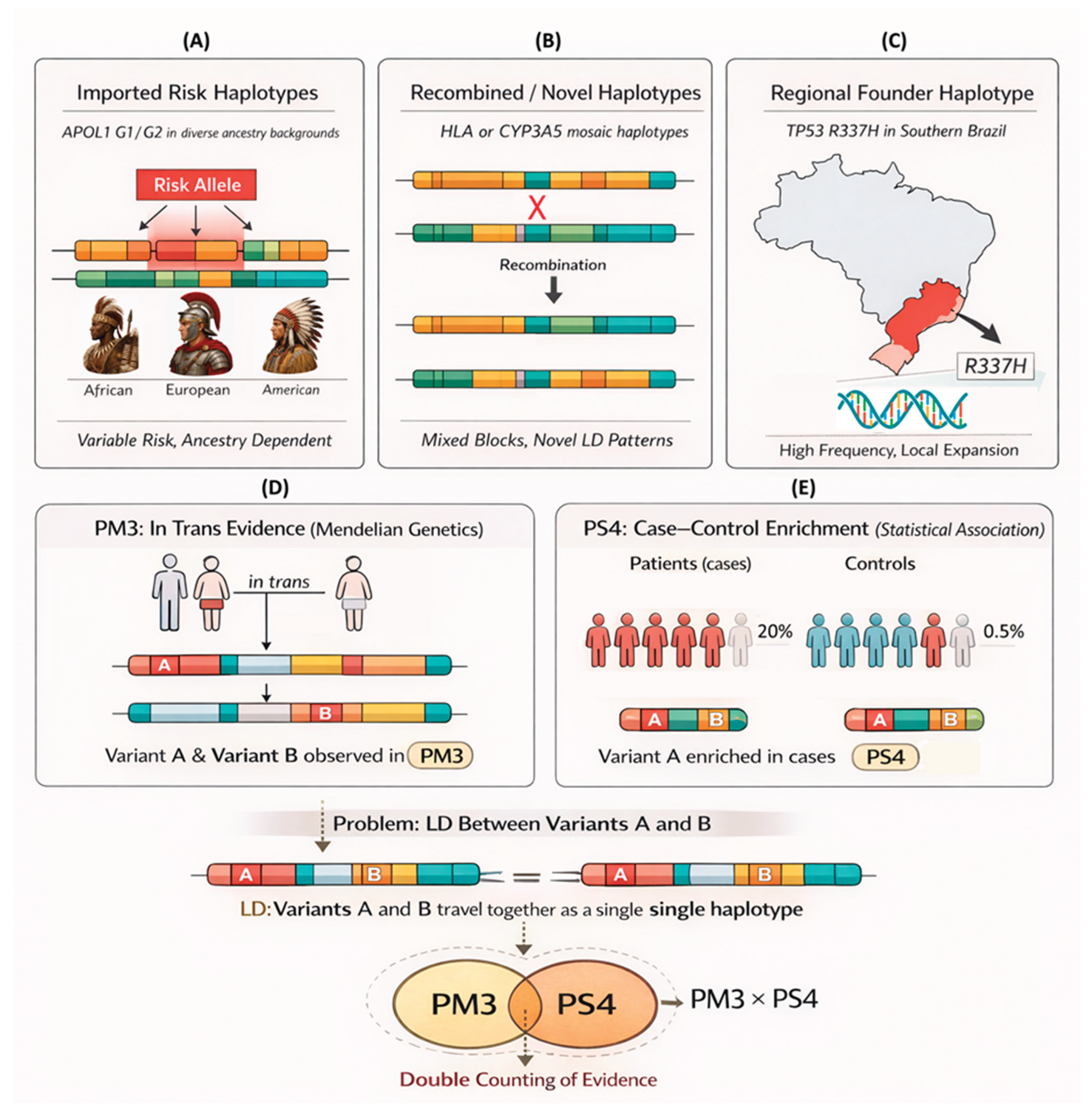
Haplotype Classes in Brazilian Populations
Toward a Haplotype-Aware Bayesian Framework
How LD Breaks the Non-Haplotype Aware Model
Correct Formulation
Three Practical Ways to Incorporate LD
Joint Likelihood
Correlation Penalty
Hierarchical Model
Translation to ACMG/AMP Criteria
Conclusions
Author Contributions
Funding
Data Availability Statement
Acknowledgments
Conflicts of Interest
References
- Achatz, M.I.W.; Olivier, M.; Calvez, F.L.; Martel-Planche, G.; Lopes, A.; Rossi, B.M.; et al. The TP53 mutation, R337H, is associated with Li-Fraumeni and Li-Fraumeni-like syndromes in Brazilian families. Cancer Letters 2007, 245, 96–102. [Google Scholar] [CrossRef] [PubMed]
- Daly, M.J.; Rioux, J.D.; Schaffner, S.F.; Hudson, T.J.; Lander, E.S. High-resolution haplotype structure in the human genome. Nat Genet 2001, 29, 229–232. [Google Scholar] [CrossRef] [PubMed]
- Gaudet, M.M.; Kirchhoff, T.; Green, T.; Vijai, J.; Korn, J.M.; Guiducci, C.; et al. Common Genetic Variants and Modification of Penetrance of BRCA2-Associated Breast Cancer. PLOS Genetics 2010, 6, e1001183. [Google Scholar] [CrossRef]
- Genovese, G.; Friedman, D.J.; Ross, M.D.; Lecordier, L.; Uzureau, P.; Freedman, B.I.; et al. Association of Trypanolytic ApoL1 Variants with Kidney Disease in African Americans. Science 2010, 329, 841–845. [Google Scholar] [CrossRef] [PubMed]
- Gravel, S. Population Genetics Models of Local Ancestry. Genetics 2012, 191, 607–619. [Google Scholar] [CrossRef] [PubMed]
- Kuehl, P.; Zhang, J.; Lin, Y.; Lamba, J.; Assem, M.; Schuetz, J.; et al. Sequence diversity in CYP3A promoters and characterization of the genetic basis of polymorphic CYP3A5 expression. Nat Genet 2001, 27, 383–391. [Google Scholar] [CrossRef] [PubMed]
- Lam, T.H.; Shen, M.; Chia, J.-M.; Chan, S.H.; Ren, E.C. Population-specific recombination sites within the human MHC region. Heredity 2013, 111, 131–138. [Google Scholar] [CrossRef] [PubMed]
- Lamason, R.L.; Mohideen, M.-A. P. K.; Mest, J.R.; Wong, A.C.; Norton, H.L.; Aros, M.C.; et al. SLC24A5, a Putative Cation Exchanger, Affects Pigmentation in Zebrafish and Humans. Science 2005, 310, 1782–1786. [Google Scholar] [CrossRef] [PubMed]
- Martin, A.R.; Kanai, M.; Kamatani, Y.; Okada, Y.; Neale, B.M.; Daly, M.J. Clinical use of current polygenic risk scores may exacerbate health disparities. Nat Genet 2019, 51, 584–591. [Google Scholar] [CrossRef] [PubMed]
- Nunes, K.; Araújo Castro e Silva, M.; Rodrigues, M.R.; Lemes, R.B.; Pezo-Valderrama, P.; Kimura, L.; et al. Admixture’s impact on Brazilian population evolution and health. Science 2025, 388, eadl3564. [Google Scholar] [CrossRef] [PubMed]
- Pena, S.D.J.; Pietro, G.D.; Fuchshuber-Moraes, M.; Genro, J.P.; Hutz, M.H.; Kehdy, F. de S. G.; et al. The Genomic Ancestry of Individuals from Different Geographical Regions of Brazil Is More Uniform Than Expected. PLOS ONE 2011, 6, e17063. [Google Scholar] [CrossRef] [PubMed]
- Reich, D.; Thangaraj, K.; Patterson, N.; Price, A.L.; Singh, L. Reconstructing Indian population history. Nature 2009, 461, 489–494. [Google Scholar] [CrossRef] [PubMed]
- Richards, S.; Aziz, N.; Bale, S.; Bick, D.; Das, S.; Gastier-Foster, J.; et al. Standards and guidelines for the interpretation of sequence variants: a joint consensus recommendation of the American College of Medical Genetics and Genomics and the Association for Molecular Pathology. Genet Med 2015, 17, 405–423. [Google Scholar] [CrossRef] [PubMed]
- Shen, J.; Oza, A.M.; del Castillo, I.; Duzkale, H.; Matsunaga, T.; Pandya, A.; et al. Consensus interpretation of the p.Met34Thr and p.Val37Ile variants in GJB2 by the ClinGen Hearing Loss Expert Panel. Genetics in Medicine 2019, 21, 2442–2452. [Google Scholar] [CrossRef] [PubMed]
- Sirugo, G.; Williams, S.M.; Tishkoff, S.A. The Missing Diversity in Human Genetic Studies. Cell 2019, 177, 26–31. [Google Scholar] [CrossRef] [PubMed]
- Tavtigian, S.V.; Greenblatt, M.S.; Harrison, S.M.; Nussbaum, R.L.; Prabhu, S.A.; Boucher, K.M.; et al. Modeling the ACMG/AMP variant classification guidelines as a Bayesian classification framework. Genetics in Medicine 2018, 20, 1054–1060. [Google Scholar] [CrossRef] [PubMed]
- Thauvin-Robinet, C.; Munck, A.; Huet, F.; Génin, E.; Bellis, G.; Gautier, E.; et al. The very low penetrance of cystic fibrosis for the R117H mutation: a reappraisal for genetic counselling and newborn screening. Journal of Medical Genetics 2009, 46, 752–758. [Google Scholar] [CrossRef] [PubMed]
| Haplotype Class | Gene | Variant / Structure | Origin | Key Property | Bayesian Impact |
|---|---|---|---|---|---|
| Imported risk | APOL1 | G1 / G2 | African | Context-dependent risk | Prior distortion |
| Recombined | HLA | Multi-locus haplotypes | Mixed | Novel LD | Likelihood error |
| Recombined | CYP3A5 | *1 / *3 | African+ European | Expression variability | Prior+ likelihood |
| Mixed functional | SLC24A5 | A111T context | European+ mixed | Functional recombination | Prior miscalibration |
| Founder | TP53 | R337H | Brazilian | Local expansion | Prior shift |
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2026 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).




