Submitted:
25 April 2026
Posted:
28 April 2026
You are already at the latest version
Abstract
Kelch-like ECH-associated protein 1 (Keap1) acts as a repressor of nuclear factor-erythroid 2-related factor 2 (Nrf2), a major transcription factor regulating cellular antioxidant response. Keap1 is the substrate adaptor subunit of the cullin 3-RING E3 ubiquitin ligase complex that specifically facilitates Nrf2 ubiquitination and its proteasomal degradation. Keap1 is rich in cysteine residues and several of them undergo various modifications, such as sulfhydration, nitrosylation and glutathionylation under cellular stress conditions. Some of these modifications alter the conformation of Keap1, preventing Nrf2 from ubiquitination and subsequent proteasome-mediated degradation. As a result, newly synthesised Nrf2 translocates to the nucleus to induce the expression of diverse genes involved in protecting cells against oxidative stress. Protein CoAlation is a reversible redox-dependent post-translational modification (PTM) in which coenzyme A (CoA) forms disulphide bonds with oxidised cysteine residues under oxidative or metabolic stress. In this study, we demonstrate for the first time that disulphide Keap1 dimer undergoes CoAlation in cellular response to oxidative stress induced by various oxidising compounds. Furthermore, glucose deprivation also induces CoAlation of disulphide Keap1 dimer in HEK293/Pank1β cells. We also demonstrate that Keap111 Cys-less mutant is not CoAlated in response to diamide treatment or glucose deprivation. In summary, this study uncovers a novel PTM of Keap1 by the key metabolic integrator CoA, which provides new insights into the regulation of the Keap1-Nrf2 antioxidant pathway under oxidative and metabolic stress.
Keywords:
1. Introduction
2. Materials and Methods
2.1. Reagents and Chemicals
2.2. Mammalian Cell Culture and Transient Transfection
2.3. Cell Treatment with Oxidising Agents and Metabolic Stress
2.4. Cell Lysis and Immunoprecipitation
2.5. Western Blot Analysis
3. Results
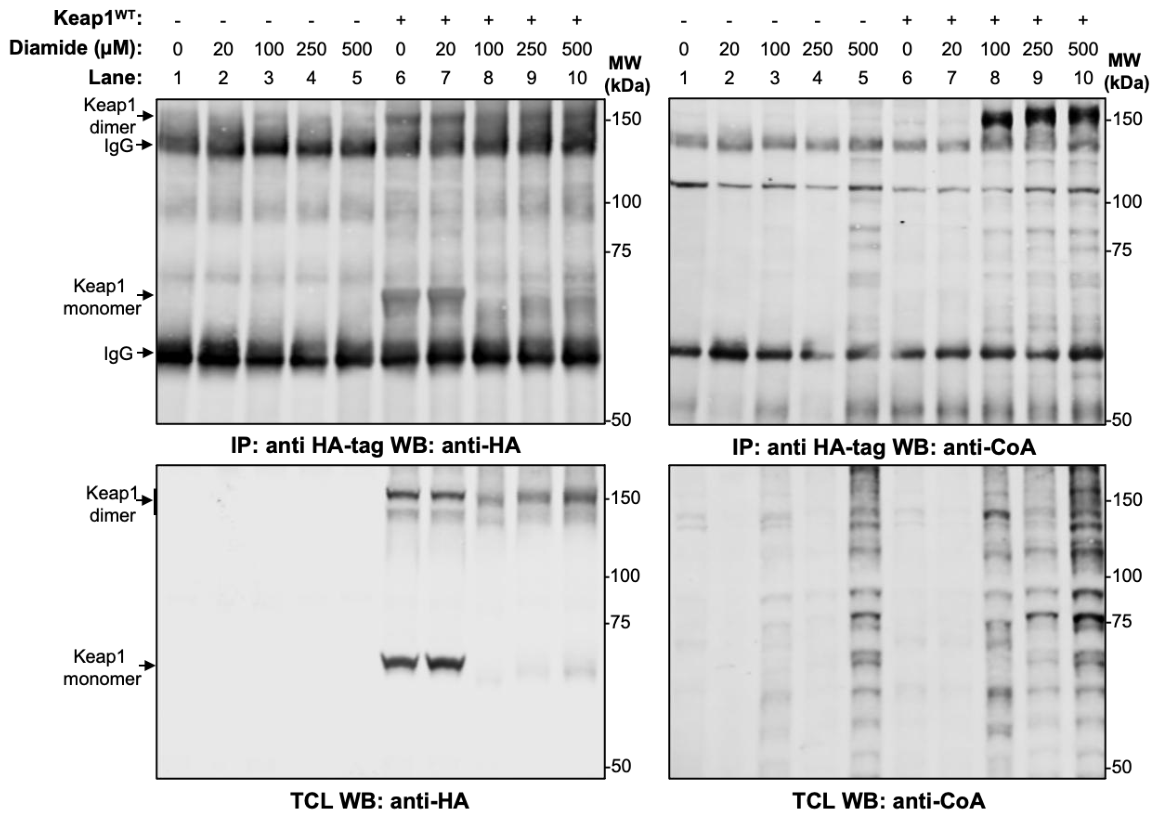
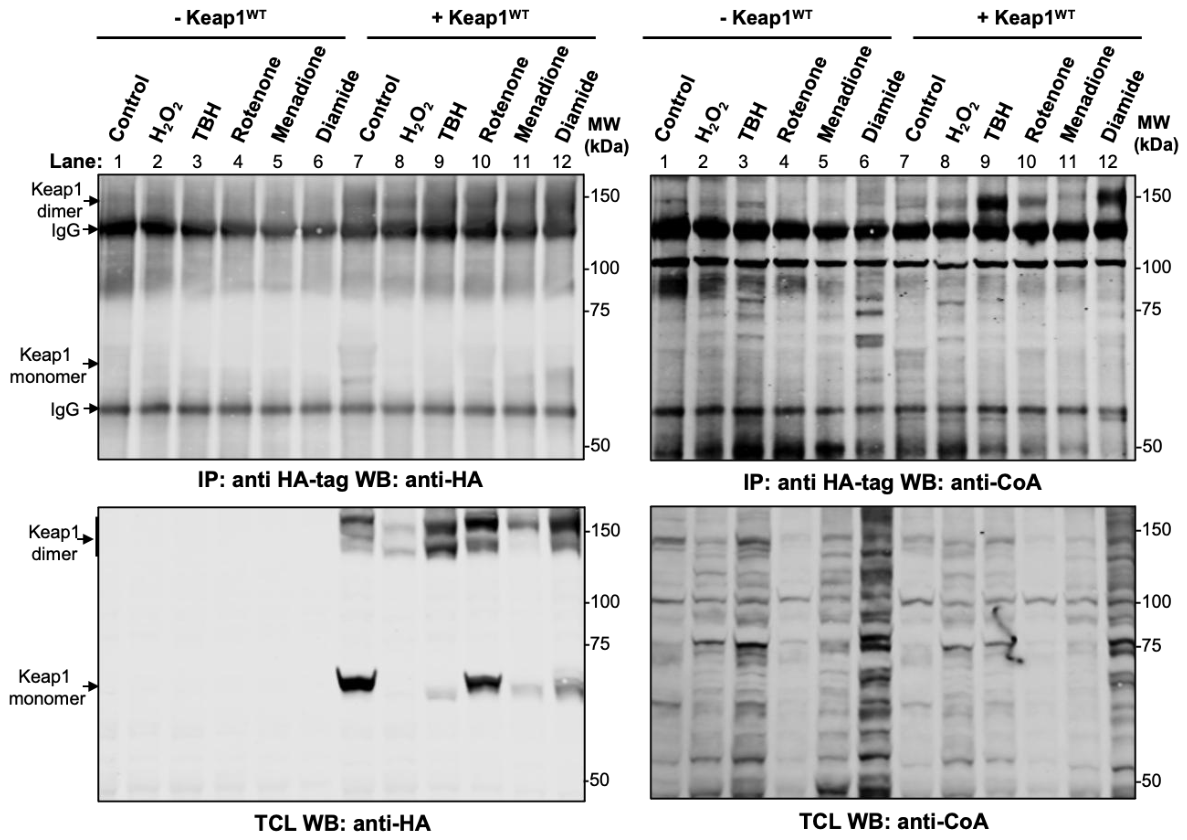
3.1. Oxidative Stress Induces CoAlation of HA-mKeap1 in HEK293/Pank1β Cells
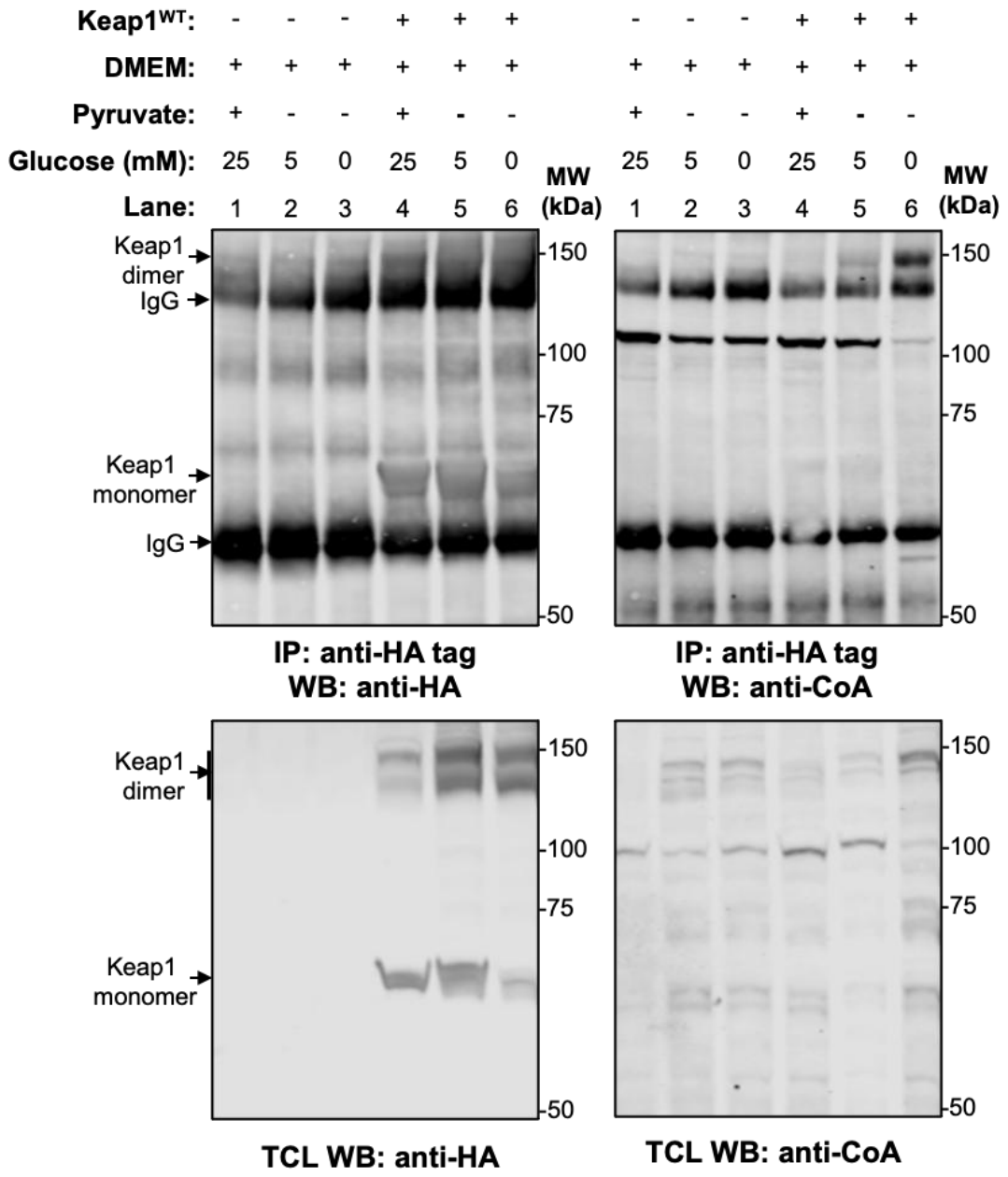
3.2. Metabolic Stress Induces CoAlation of HA-mKeap1WT in HEK293/Pank1β Cells
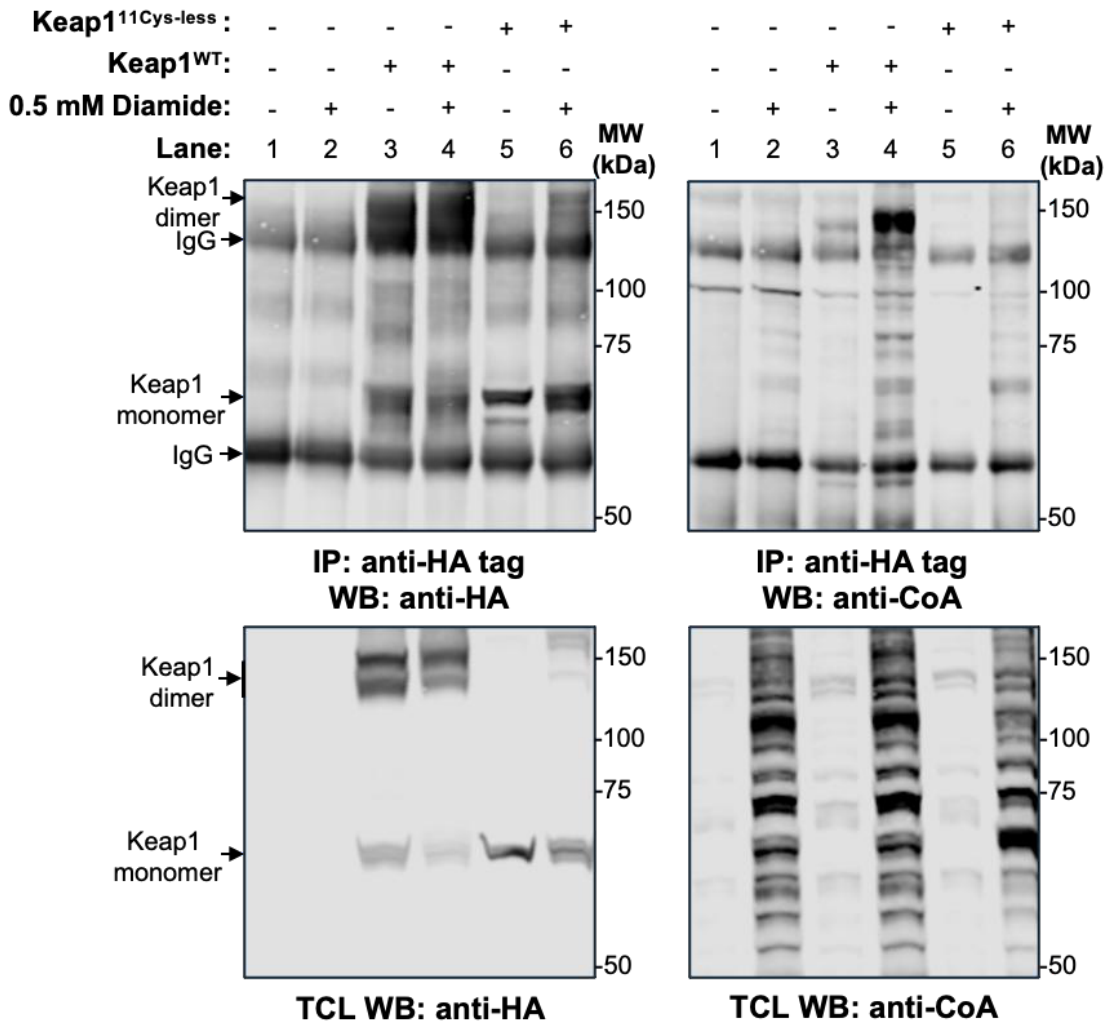
3.3. Oxidative Stress Does Not Induce CoAlation of the HA-mKeap111Cys-less Mutant
4. Discussion
Supplementary Materials
Author Contributions
Funding
Conflicts of Interest
Abbreviations
| Keap1 | Kelch-like ECH-associated protein 1 |
| Nrf2 | Nuclear factor-erythroid 2-related factor 2 |
| PTM | Post-translational modification |
| CoA | Coenzyme A |
| Redox | Reduction-oxidation |
| ROS | Reactive oxygen species |
| H2O2 | Hydrogen peroxide |
| LMW | Low-molecular-weight |
| GSH | Glutathione |
| BSH | Bacillithiol |
| MSH | Mycothiol |
| TCA | Tricarboxylic acid |
| sMAF | Small musculoaponeurotic fibrosarcoma |
| CNC | Cap’n’collar |
| CsMBE | CNC-sMaf binding element |
| Cul3 RBX1 |
Cullin3 RING-box protein 1 |
| NTD | N-terminal domain |
| BTB | Broad complex-Tramtrack-Bric-a-brac |
| IVR | Intervening region |
| CTD | C-terminal domain |
| TBH | tert-Butyl hydroperoxide |
| NEM | N-ethylmaleimide |
| SDS | Sodium dodecyl sulfate |
| DTT | Dithiothreitol |
| Tris-HCl | Tris hydrochloride |
| PIC | Protease Inhibitor Cocktail |
| mAb | Monoclonal antibody |
| HEK293 | Human embryonic kidney 293 |
| Pank1β | Pantothenate kinase 1β |
| DMEM | Dulbecco’s Modified Eagle Medium |
| FBS | Fetal bovine serum |
| RT | Room temperature |
| TCL | Total cell lysate |
| WB | Western blot |
| HA-mKeap1WT | HA-tagged wild-type mouse Keap1 |
| HA-mKeap111Cys-less | HA-tagged mouse Keap1 mutant containing 11 substituted cysteine residues |
| SDS-PAGE | SDS polyacrylamide gel electrophoresis |
| TBSt | Tris-buffer saline containing 0.1% Tween 20 |
References
- Sies, H.; Berndt, C.; Jones, D.P. Oxidative Stress. Annu Rev. Biochem. 2017, 86, 715–48. [Google Scholar] [CrossRef] [PubMed]
- Sies, H.; Belousov, V.V.; Chandel, N.S.; Davies, M.J.; Jones, D.P.; Mann, G.E.; et al. Defining roles of specific reactive oxygen species (ROS) in cell biology and physiology. Nat. Rev. Mol. Cell. Biol. 2022, 23(7), 499–515. [Google Scholar] [CrossRef]
- Tossounian, M.-A.; Zhao, Y.; Yu, B.Y.K.; Markey, S.A.; Malanchuk, O.; Zhu, Y.; et al. Low-molecular-weight thiol transferases in redox regulation and antioxidant defence. Redox Biol. 2024, 71, 103094. [Google Scholar] [CrossRef]
- Van Laer, K.; Hamilton, C.J.; Messens, J. Low-molecular-weight thiols in thiol-disulfide exchange. Antioxid. Redox Signal. 2013, 18(13), 1642–53. [Google Scholar] [CrossRef]
- Tsuchiya, Y.; Peak-Chew, S.Y.; Newell, C.; Miller-Aidoo, S.; Mangal, S.; Zhyvoloup, A.; et al. Protein CoAlation: a redox-regulated protein modification by coenzyme A in mammalian cells. Biochem J. 2017, 474(14), 2489–508. [Google Scholar] [CrossRef]
- Brett, C.; Gout, I. The two faces of coenzyme A in cellular biology. Free Radic. Biol. Med. 2025, 233, 162–73. [Google Scholar] [CrossRef]
- Tossounian, M.A.; Baczynska, M.; Dalton, W.; Newell, C.; Ma, Y.; Das, S.; et al. Profiling the Site of Protein CoAlation and Coenzyme A Stabilization Interactions. Antioxidants 2022, 11(7). [Google Scholar] [CrossRef] [PubMed]
- Tsuchiya, Y.; Zhyvoloup, A.; Baković, J.; Thomas, N.; Yu, B.Y.K.; Das, S.; et al. Protein CoAlation and antioxidant function of coenzyme A in prokaryotic cells. Biochem J. 2018, 475(11), 1909–37. [Google Scholar] [CrossRef] [PubMed]
- Baković, J.; Yu, B.Y.K.; Silva, D.; Chew, S.P.; Kim, S.; Ahn, S.H.; et al. A key metabolic integrator, coenzyme A, modulates the activity of peroxiredoxin 5 via covalent modification. Mol. Cell. Biochem. 2019, 461(1-2), 91–102. [Google Scholar] [CrossRef] [PubMed]
- Lin, C.C.; Lin, Y.T.; Chen, S.Y.; Setayeshpour, Y.; Chen, Y.; Dunn, D.E.; et al. Coenzyme A protects against ferroptosis via CoAlation of mitochondrial thioredoxin reductase. J. Clin. Invest. 2025, 135(19). [Google Scholar] [CrossRef] [PubMed]
- Malanchuk, O.; Bdzhola, A.; Palchevskyi, S.; Bdzhola, V.; Chai, P.; Pardo, O.E.; et al. Investigating the Regulation of Ribosomal Protein S6 Kinase 1 by CoAlation. Int. J. Mol. Sci. 2024, 25(16). [Google Scholar] [CrossRef]
- Tsuchiya, Y.; Byrne, D.P.; Burgess, S.G.; Bormann, J.; Baković, J.; Huang, Y.; et al. Covalent Aurora A regulation by the metabolic integrator coenzyme A. Redox Biol. 2020, 28, 101318. [Google Scholar] [CrossRef]
- Yu, B.Y.K.; Tossounian, M.A.; Hristov, S.D.; Lawrence, R.; Arora, P.; Tsuchiya, Y.; et al. Regulation of metastasis suppressor NME1 by a key metabolic cofactor coenzyme A. Redox Biol. 2021, 44, 101978. [Google Scholar] [CrossRef]
- Suzuki, T.; Motohashi, H.; Yamamoto, M. Toward clinical application of the Keap1-Nrf2 pathway. Trends Pharmacol. Sci. 2013, 34(6), 340–6. [Google Scholar] [CrossRef] [PubMed]
- Yamamoto, M.; Kensler, T.W.; Motohashi, H. The KEAP1-NRF2 System: a Thiol-Based Sensor-Effector Apparatus for Maintaining Redox Homeostasis. Physiol. Rev. 2018, 98(3), 1169–203. [Google Scholar] [CrossRef] [PubMed]
- Tossounian, M.-A.; Zhyvoloup, A.; Chatterjee, R.; Gouge, J. Dissecting the Interplay Between NRF2 and BACH1 at CsMBEs. Antioxidants 2025, 14, 1203. [Google Scholar] [CrossRef]
- Zhang, D.D.; Lo, S.C.; Cross, J.V.; Templeton, D.J.; Hannink, M. Keap1 is a redox-regulated substrate adaptor protein for a Cul3-dependent ubiquitin ligase complex. Mol. Cell. Biol. 2004, 24(24), 10941–53. [Google Scholar]
- Kobayashi, A.; Kang, M.I.; Okawa, H.; Ohtsuji, M.; Zenke, Y.; Chiba, T.; et al. Oxidative stress sensor Keap1 functions as an adaptor for Cul3-based E3 ligase to regulate proteasomal degradation of Nrf2. Mol. Cell. Biol. 2004, 24(16), 7130–9. [Google Scholar] [CrossRef]
- Furukawa, M.; Xiong, Y. BTB protein Keap1 targets antioxidant transcription factor Nrf2 for ubiquitination by the Cullin 3-Roc1 ligase. Mol. Cell. Biol. 2005, 25(1), 162–71. [Google Scholar] [CrossRef] [PubMed]
- Cullinan, S.B.; Gordan, J.D.; Jin, J.; Harper, J.W.; Diehl, J.A. The Keap1-BTB protein is an adaptor that bridges Nrf2 to a Cul3-based E3 ligase: oxidative stress sensing by a Cul3-Keap1 ligase. Mol. Cell. Biol. 2004, 24(19), 8477–86. [Google Scholar] [PubMed]
- Fourquet, S.; Guerois, R.; Biard, D.; Toledano, M.B. Activation of NRF2 by nitrosative agents and H2O2 involves KEAP1 disulfide formation. J. Biol. Chem. 2010, 285(11), 8463–71. [Google Scholar] [CrossRef]
- Yamamoto, T.; Suzuki, T.; Kobayashi, A.; Wakabayashi, J.; Maher, J.; Motohashi, H.; et al. Physiological significance of reactive cysteine residues of Keap1 in determining Nrf2 activity. Mol. Cell. Biol. 2008, 28(8), 2758–70. [Google Scholar] [CrossRef]
- Saito, R.; Suzuki, T.; Hiramoto, K.; Asami, S.; Naganuma, E.; Suda, H.; et al. Characterizations of Three Major Cysteine Sensors of Keap1 in Stress Response. Mol. Cell. Biol. 2016, 36(2), 271–84. [Google Scholar] [CrossRef] [PubMed]
- Zhang, D.D.; Hannink, M. Distinct cysteine residues in Keap1 are required for Keap1-dependent ubiquitination of Nrf2 and for stabilization of Nrf2 by chemopreventive agents and oxidative stress. Mol. Cell. Biol. 2003, 23(22), 8137–51. [Google Scholar] [CrossRef]
- Takaya, K.; Suzuki, T.; Motohashi, H.; Onodera, K.; Satomi, S.; Kensler, T.W.; et al. Validation of the multiple sensor mechanism of the Keap1-Nrf2 system. Free Radic. Biol. Med. 2012, 53(4), 817–27. [Google Scholar] [PubMed]
- McMahon, M.; Lamont, D.J.; Beattie, K.A.; Hayes, J.D. Keap1 perceives stress via three sensors for the endogenous signaling molecules nitric oxide, zinc, and alkenals. Proc. Natl. Acad. Sci. U S A 2010, 107(44), 18838–43. [Google Scholar] [CrossRef]
- Kobayashi, M.; Li, L.; Iwamoto, N.; Nakajima-Takagi, Y.; Kaneko, H.; Nakayama, Y.; et al. The antioxidant defense system Keap1-Nrf2 comprises a multiple sensing mechanism for responding to a wide range of chemical compounds. Mol. Cell. Biol. 2009, 29(2), 493–502. [Google Scholar] [CrossRef] [PubMed]
- Suzuki, T.; Muramatsu, A.; Saito, R.; Iso, T.; Shibata, T.; Kuwata, K.; et al. Molecular Mechanism of Cellular Oxidative Stress Sensing by Keap1. Cell. Rep. 2019, 28(3), 746–58.e4. [Google Scholar] [CrossRef]
- Ogura, T.; Tong, K.I.; Mio, K.; Maruyama, Y.; Kurokawa, H.; Sato, C.; et al. Keap1 is a forked-stem dimer structure with two large spheres enclosing the intervening, double glycine repeat, and C-terminal domains. Proc. Natl. Acad. Sci. U S A. 2010, 107(7), 2842–7. [Google Scholar] [CrossRef]
- Carvalho, A.N.; Marques, C.; Guedes, R.C.; Castro-Caldas, M.; Rodrigues, E.; van Horssen, J.; et al. S-Glutathionylation of Keap1: a new role for glutathione S-transferase pi in neuronal protection. FEBS Lett. 2016, 590(10), 1455–66. [Google Scholar] [CrossRef]
- Luo, Y.; Eggler, A.L.; Liu, D.; Liu, G.; Mesecar, A.D.; van Breemen, R.B. Sites of alkylation of human Keap1 by natural chemoprevention agents. J. Am. Soc. Mass. Spectrom. 2007, 18(12), 2226–32. [Google Scholar] [PubMed]
- Mills, E.L.; Ryan, D.G.; Prag, H.A.; Dikovskaya, D.; Menon, D.; Zaslona, Z.; et al. Itaconate is an anti-inflammatory metabolite that activates Nrf2 via alkylation of KEAP1. Nature. 2018, 556(7699), 113–7. [Google Scholar] [PubMed]
- Adam, J.; Hatipoglu, E.; O’Flaherty, L.; Ternette, N.; Sahgal, N.; Lockstone, H.; et al. Renal cyst formation in Fh1-deficient mice is independent of the Hif/Phd pathway: roles for fumarate in KEAP1 succination and Nrf2 signaling. Cancer Cell. 2011, 20(4), 524–37. [Google Scholar]
- Bollong, M.J.; Lee, G.; Coukos, J.S.; Yun, H.; Zambaldo, C.; Chang, J.W.; et al. A metabolite-derived protein modification integrates glycolysis with KEAP1-NRF2 signalling. Nature 2018, 562(7728), 600–4. [Google Scholar]
- Ko, Y.; Hong, M.; Lee, S.; Kumar, M.; Ibrahim, L.; Nutsch, K.; et al. S-lactoyl modification of KEAP1 by a reactive glycolytic metabolite activates NRF2 signaling. Proc. Natl. Acad. Sci. U S A. 2023, 120(20), e2300763120. [Google Scholar]
- Um, H.C.; Jang, J.H.; Kim, D.H.; Lee, C.; Surh, Y.J. Nitric oxide activates Nrf2 through S-nitrosylation of Keap1 in PC12 cells. Nitric Oxide. 2011, 25(2), 161–8. [Google Scholar]
- Xie, L.; Gu, Y.; Wen, M.; Zhao, S.; Wang, W.; Ma, Y.; et al. Hydrogen Sulfide Induces Keap1 S-sulfhydration and Suppresses Diabetes-Accelerated Atherosclerosis via Nrf2 Activation. Diabetes 2016, 65(10), 3171–84. [Google Scholar] [CrossRef] [PubMed]
- Li, N.; Wang, J.; Zang, X.; Wang, Z.; Zhang, T.; Zhao, B.; et al. H(2)S probe CPC inhibits autophagy and promotes apoptosis by inhibiting glutathionylation of Keap1 at Cys434. Apoptosis. 2021, 26(1-2), 111–31. [Google Scholar] [CrossRef]
- Gambhir, L.; Checker, R.; Thoh, M.; Patwardhan, R.S.; Sharma, D.; Kumar, M.; et al. 1,4-Naphthoquinone, a pro-oxidant, suppresses immune responses via KEAP-1 glutathionylation. Biochem Pharmacol. 2014, 88(1), 95–105. [Google Scholar]
- Wang, L.; Qu, G.; Gao, Y.; Su, L.; Ye, Q.; Jiang, F.; et al. A small molecule targeting glutathione activates Nrf2 and inhibits cancer cell growth through promoting Keap-1 S-glutathionylation and inducing apoptosis. RSC Adv. 2018, 8(2), 792–804. [Google Scholar]
- Chen, X.; Zhou, Q.; Chen, H.; Bai, J.; An, R.; Zhang, K.; et al. Glutathione Induces Keap1 S-Glutathionylation and Mitigates Oscillating Glucose-Induced β-Cell Dysfunction by Activating Nrf2. Antioxidants . 2024, 13(4). [Google Scholar] [CrossRef]
- Sun, X.; Guo, C.; Huang, C.; Lv, N.; Chen, H.; Huang, H.; et al. GSTP alleviates acute lung injury by S-glutathionylation of KEAP1 and subsequent activation of NRF2 pathway. Redox Biol. 2024, 71, 103116. [Google Scholar] [CrossRef]
- Malanchuk, O.; Panasyuk, G.; Serbin, N.; Gout, I.; Filonenko, V. Generation and characterization of monoclonal antibodies specific to Coenzyme A. Biopolym. Cell. 2015, 31, 187–92. [Google Scholar] [CrossRef]
- Wakabayashi, N.; Dinkova-Kostova, A.T.; Holtzclaw, W.D.; Kang, M.I.; Kobayashi, A.; Yamamoto, M.; et al. Protection against electrophile and oxidant stress by induction of the phase 2 response: fate of cysteines of the Keap1 sensor modified by inducers. Proc. Natl. Acad. Sci. U S A 2004, 101(7), 2040–5. [Google Scholar] [CrossRef]
- Criddle, D.N.; Gillies, S.; Baumgartner-Wilson, H.K.; Jaffar, M.; Chinje, E.C.; Passmore, S.; et al. Menadione-induced reactive oxygen species generation via redox cycling promotes apoptosis of murine pancreatic acinar cells. J. Biol. Chem. 2006, 281(52), 40485–92. [Google Scholar] [CrossRef]
- Li, N.; Ragheb, K.; Lawler, G.; Sturgis, J.; Rajwa, B.; Melendez, J.A.; et al. Mitochondrial complex I inhibitor rotenone induces apoptosis through enhancing mitochondrial reactive oxygen species production. J. Biol. Chem. 2003, 278(10), 8516–25. [Google Scholar] [CrossRef]
- Kosower, N.S.; Kosower, E.M. Diamide: an oxidant probe for thiols. Methods Enzymol. 1995, 251, 123–33. [Google Scholar]
- Zipper, L.M.; Mulcahy, R.T. The Keap1 BTB/POZ dimerization function is required to sequester Nrf2 in cytoplasm. J. Biol. Chem. 2002, 277(39), 36544–52. [Google Scholar] [CrossRef]
- Sato, M.; Yaguchi, N.; Iijima, T.; Muramatsu, A.; Baird, L.; Suzuki, T.; et al. Sensor systems of KEAP1 uniquely detecting oxidative and electrophilic stresses separately In vivo. Redox Biol. 2024, 77, 103355. [Google Scholar] [CrossRef]
- Holland, R.; Hawkins, A.E.; Eggler, A.L.; Mesecar, A.D.; Fabris, D.; Fishbein, J.C. Prospective type 1 and type 2 disulfides of Keap1 protein. Chem. Res. Toxicol. 2008, 21(10), 2051–60. [Google Scholar] [CrossRef]
- Dunlap, T.; Piyankarage, S.C.; Wijewickrama, G.T.; Abdul-Hay, S.; Vanni, M.; Litosh, V.; et al. Quinone-induced activation of Keap1/Nrf2 signaling by aspirin prodrugs masquerading as nitric oxide. Chem. Res. Toxicol. 2012, 25(12), 2725–36. [Google Scholar] [CrossRef]
- Hong, F.; Sekhar, K.R.; Freeman, M.L.; Liebler, D.C. Specific patterns of electrophile adduction trigger Keap1 ubiquitination and Nrf2 activation. J. Biol. Chem. 2005, 280(36), 31768–75. [Google Scholar] [CrossRef]
- Eggler, A.L.; Liu, G.; Pezzuto, J.M.; van Breemen, R.B.; Mesecar, A.D. Modifying specific cysteines of the electrophile-sensing human Keap1 protein is insufficient to disrupt binding to the Nrf2 domain Neh2. Proc. Natl. Acad. Sci. U S A. 2005, 102(29), 10070–5. [Google Scholar] [CrossRef]
- Dietz, B.M.; Liu, D.; Hagos, G.K.; Yao, P.; Schinkovitz, A.; Pro, S.M.; et al. Angelica sinensis and its alkylphthalides induce the detoxification enzyme NAD(P)H: quinone oxidoreductase 1 by alkylating Keap1. Chem. Res. Toxicol. 2008, 21(10), 1939–48. [Google Scholar] [CrossRef]
- Desideri, E.; Filomeni, G.; Ciriolo, M.R. Glutathione participates in the modulation of starvation-induced autophagy in carcinoma cells. Autophagy. 2012, 8(12), 1769–81. [Google Scholar] [CrossRef]
- Lettieri-Barbato, D.; Minopoli, G.; Caggiano, R.; Izzo, R.; Santillo, M.; Aquilano, K.; et al. Fasting Drives Nrf2-Related Antioxidant Response in Skeletal Muscle. Int. J. Mol. Sci. 2020, 21(20). [Google Scholar] [CrossRef]
- Tossounian, M.A.; Baczynska, M.; Dalton, W.; Peak-Chew, S.Y.; Undzenas, K.; et al. Bacillus subtilis YtpP and Thioredoxin A Are New Players in the Coenzyme-A-Mediated Defense Mechanism against Cellular Stress. Antioxidants 2023, 12(4), 938. [Google Scholar] [CrossRef]
- Ahmed, S.M.; Luo, L.; Namani, A.; Wang, X.J.; Tang, X. Nrf2 signaling pathway: Pivotal roles in inflammation. Biochim Biophys. Acta Mol. Basis Dis. 2017, 1863(2), 585–97. [Google Scholar] [CrossRef]
- Adinolfi, S.; Patinen, T.; Deen, J.A.; Pitkänen, S.; Härkönen, J.; Kansanen, E.; et al. The KEAP1-NRF2 pathway: Targets for therapy and role in cancer. Redox Biol. 2023, 63, 102726. [Google Scholar] [CrossRef]
- Yamazaki, H.; Tanji, K.; Wakabayashi, K.; Matsuura, S.; Itoh, K. Role of the Keap1/Nrf2 pathway in neurodegenerative diseases. Pathol. Int. 2015, 65(5), 210–9. [Google Scholar] [CrossRef]
- Lashley, T.; Tossounian, M.-A.; Heaven, C.N.; Wallworth, S.; Peak-Chew, S.; Bradshaw, A.; et al. Extensive Anti-CoA Immunostaining in Alzheimer’s Disease and Covalent Modification of Tau by a Key Cellular Metabolite Coenzyme A. Front Cell. Neurosci. 2021, 15, 739425. [Google Scholar] [CrossRef] [PubMed]
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2026 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).



