Submitted:
26 April 2026
Posted:
27 April 2026
You are already at the latest version
Abstract
Keywords:
1. Introduction
2. Lysosomes and the Tumor Microenvironment
2.1. Lysosomal Degradation and Signaling Networks in Cellular and Tumor Homeostasis
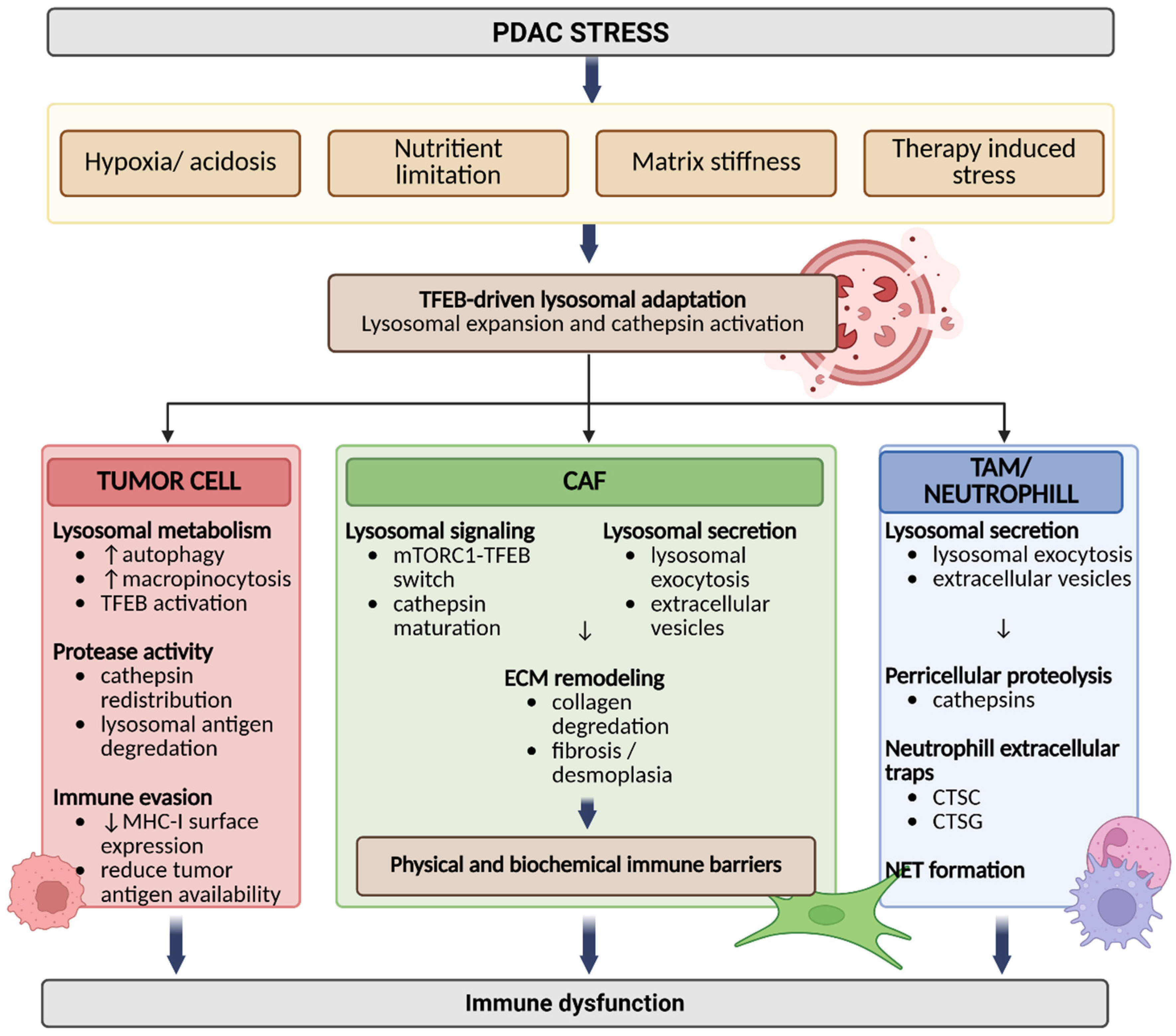
2.2. Unique Features of PDAC TME That Impact Lysosomal Function
2.2.1. Hypoxia and Low pH (Turning Lysosomal Programs Outward)
2.2.2. Nutrient Limitation (Autophagy and Scavenging Through Lysosomes)
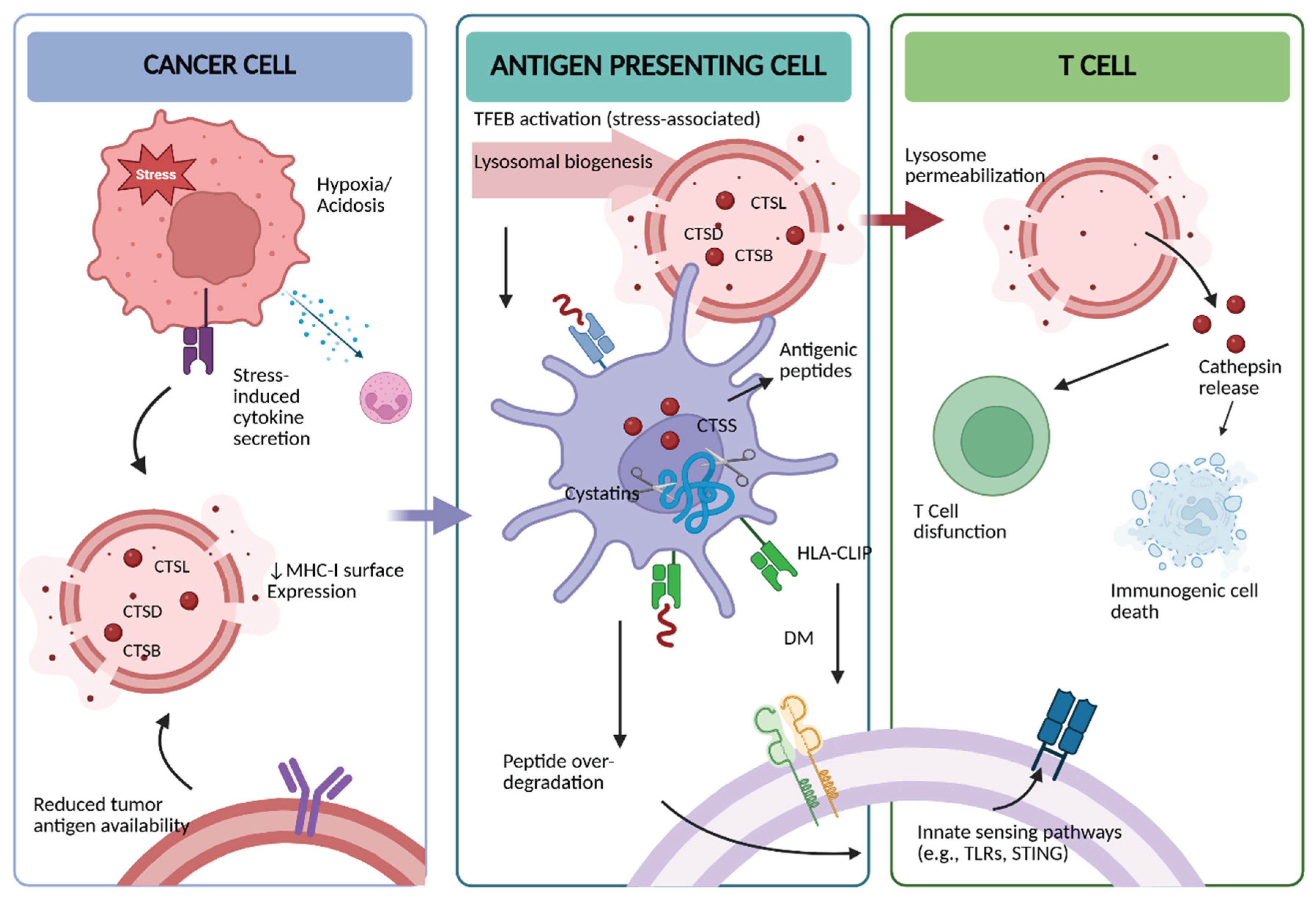
2.3. Lysosomal–Immune System Interface (Antigen Presentation, Cytokine Release, Cell Death)
3. Cathepsins as Lysosomal Effectors in Pancreatic Cancer
3.1. Overview of Cathepsins
3.2. Lysosomal Localization, Trafficking, and Activation Control of Cathepsin Activity
3.3. Non-Canonical Release into the Extracellular Space and Microenvironmental Effects
3.4. Clinical Associations and Compartmental Expression
4. Lysosome–Cathepsin Axis in Anti-Tumor Immunity in Pancreatic Cancer
4.1. Antigen Presentation: Role of Cathepsins in MHC Class II Loading in Dendritic Cells and Macrophages
4.2. Implications for T-Cell Priming in PDAC
4.3. Lysosomal Cathepsins in Stromal Remodeling and Immune Exclusion
4.4. Lysosomal Membrane Permeabilization and Cathepsin-Mediated Cell Death
5. Therapeutic Targeting of the Lysosome-Cathepsin Axis
5.1. Lysosome-Targeting Drugs
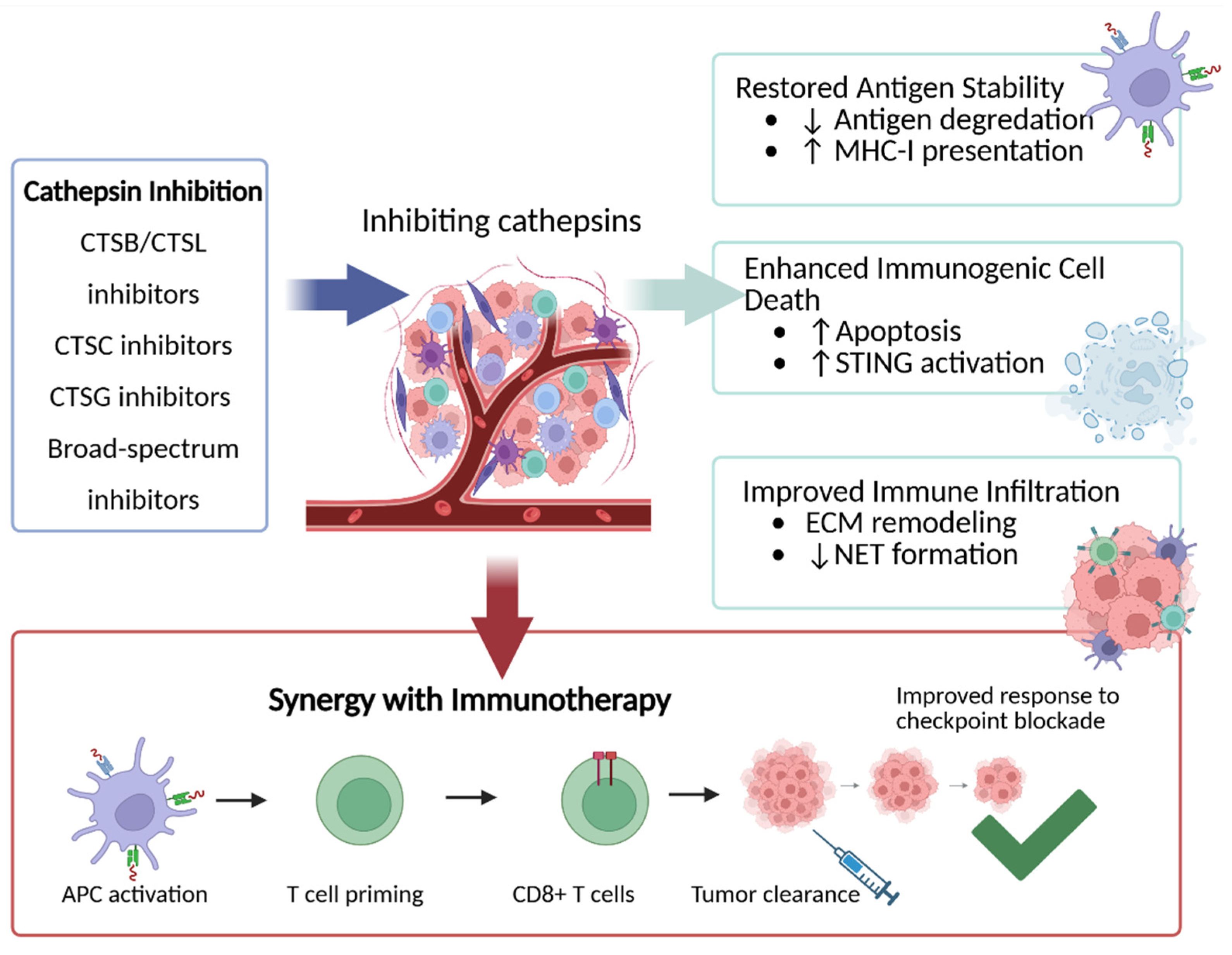
5.2. Cathepsin Inhibitors
5.3. Cathepsin-Responsive Targeting Strategies, Theranostics, and Prodrug Systems
5.4. Biomarkers and Monitoring
6. Conclusions and Future Perspectives
Author Contributions
Funding
Acknowledgments
Conflicts of Interest
Abbreviations
| APC | Antigen presenting cell |
| ATG | Autophagy related protein |
| CAF | Cancer-associated fibroblast |
| CQ | Chloroquine |
| ECM | Extracellular matrix |
| HCQ | Hydrochloroquine |
| MHC | Major histocompatibility complex |
| MMP | Matrix metalloproteinase |
| NET | Neutrophil extracellular trap |
| NK | Natural killer cell |
| PDAC | Pancreatic ductal adenocarcinoma |
| PSC | Pancreatic stellate cell |
| TAM | Tumor-associated macrophage |
| TFEB | Transcription factor EB |
| TME | Tumor microenvironment |
| uPA | Urokinase-type plasminogen activator |
References
- Lencioni, G.; Gregori, A.; Toledo, B.; Rebelo, R.; Immordino, B.; Amrutkar, M.; Xavier, C.P.R.; Kocijančič, A.; Pandey, D.P.; Perán, M.; et al. Unravelling the Complexities of Resistance Mechanism in Pancreatic Cancer: Insights from in Vitro and Ex-Vivo Model Systems. Semin. Cancer Biol. 2024, 106–107, 217–233. [Google Scholar] [CrossRef]
- Mukherji, R.; Debnath, D.; Hartley, M.L.; Noel, M.S. The Role of Immunotherapy in Pancreatic Cancer. Curr. Oncol. Tor. Ont. 2022, 29, 6864–6892. [Google Scholar] [CrossRef]
- Partyka, O.; Pajewska, M.; Kwaśniewska, D.; Czerw, A.; Deptała, A.; Budzik, M.; Cipora, E.; Gąska, I.; Gazdowicz, L.; Mielnik, A.; et al. Overview of Pancreatic Cancer Epidemiology in Europe and Recommendations for Screening in High-Risk Populations. Cancers 2023, 15, 3634. [Google Scholar] [CrossRef]
- Di Costanzo, F.; Di Costanzo, F.; Antonuzzo, L.; Mazza, E.; Giommoni, E. Optimizing First-Line Chemotherapy in Metastatic Pancreatic Cancer: Efficacy of FOLFIRINOX versus Nab-Paclitaxel Plus Gemcitabine. Cancers 2023, 15, 416. [Google Scholar] [CrossRef]
- Galindo-Vega, A.; Maldonado-Lagunas, V.; Mitre-Aguilar, I.B.; Melendez-Zajgla, J. Tumor Microenvironment Role in Pancreatic Cancer Stem Cells. Cells 2023, 12, 1560. [Google Scholar] [CrossRef] [PubMed]
- Sherman, M.H.; Beatty, G.L. Tumor Microenvironment in Pancreatic Cancer Pathogenesis and Therapeutic Resistance. Annu. Rev. Pathol. 2023, 18, 123–148. [Google Scholar] [CrossRef]
- Glapiński, F.; Zając, W.; Fudalej, M.; Deptała, A.; Czerw, A.; Sygit, K.; Kozłowski, R.; Badowska-Kozakiewicz, A. The Role of the Tumor Microenvironment in Pancreatic Ductal Adenocarcinoma: Recent Advancements and Emerging Therapeutic Strategies. Cancers 2025, 17, 1599. [Google Scholar] [CrossRef] [PubMed]
- Hartupee, C.; Nagalo, B.M.; Chabu, C.Y.; Tesfay, M.Z.; Coleman-Barnett, J.; West, J.T.; Moaven, O. Pancreatic Cancer Tumor Microenvironment Is a Major Therapeutic Barrier and Target. Front. Immunol. 2024, 15. [Google Scholar] [CrossRef] [PubMed]
- Olaoba, O.T.; Yang, M.; Adelusi, T.I.; Maidens, T.; Kimchi, E.T.; Staveley-O’Carroll, K.F.; Li, G. Targeted Therapy for Highly Desmoplastic and Immunosuppressive Tumor Microenvironment of Pancreatic Ductal Adenocarcinoma. Cancers 2024, 16. [Google Scholar] [CrossRef]
- Bräutigam, K.; Skok, K.; Szymonski, K.; Rift, C.V.; Karamitopoulou, E. Tumor Immune Microenvironment in Pancreatic Ductal Adenocarcinoma Revisited – Exploring the “Space.”. Cancer Lett. 2025, 622, 217699. [Google Scholar] [CrossRef]
- Masuda, H. Cancer-Associated Fibroblasts in Cancer Drug Resistance and Cancer Progression: A Review. Cell. Death Discov. 2025, 11, 341. [Google Scholar] [CrossRef]
- Deipenbrock, A.; Wilmes, B.E.; Sommermann, T.; Abdo, N.; Moustakas, K.; Raasch, M.; Rennert, K.; Teusch, N.E. Modelling of the Multicellular Tumor Microenvironment of Pancreatic Ductal Adenocarcinoma (PDAC) on a Fit-for-Purpose Biochip for Preclinical Drug Discovery. Lab. Chip 2025, 25, 2168–2181. [Google Scholar] [CrossRef]
- Bear, A.S.; Vonderheide, R.H.; O’Hara, M.H. Challenges and Opportunities for Pancreatic Cancer Immunotherapy. Cancer Cell. 2020, 38, 788–802. [Google Scholar] [CrossRef] [PubMed]
- Bowers, J.S.; Bailey, S.R.; Rubinstein, M.P.; Paulos, C.M.; Camp, E.R. Genomics Meets Immunity in Pancreatic Cancer: Current Research and Future Directions for Pancreatic Adenocarcinoma Immunotherapy. Oncol. Rev. 2019, 13, 430. [Google Scholar] [CrossRef]
- Settembre, C.; Perera, R.M. Lysosomes as Coordinators of Cellular Catabolism, Metabolic Signalling and Organ Physiology. Nat. Rev. Mol. Cell. Biol. 2024, 25, 223–245. [Google Scholar] [CrossRef]
- Huang, F.-F.; Cui, W.-H.; Ma, L.-Y.; Chen, Q.; Liu, Y. Crosstalk of Nervous and Immune Systems in Pancreatic Cancer. Front. Cell. Dev. Biol. 2023, 11. [Google Scholar] [CrossRef]
- Savini, M.; Zhao, Q.; Wang, M.C. Lysosomes: Signaling Hubs for Metabolic Sensing and Longevity. Trends Cell. Biol. 2019, 29, 876–887. [Google Scholar] [CrossRef]
- Pišlar, A.; Perišić Nanut, M.; Kos, J. Lysosomal Cysteine Peptidases - Molecules Signaling Tumor Cell Death and Survival. Semin. Cancer Biol. 2015, 35, 168–179. [Google Scholar] [CrossRef]
- Senjor, E.; Kos, J.; Nanut, M.P. Cysteine Cathepsins as Therapeutic Targets in Immune Regulation and Immune Disorders. Biomedicines 2023, 11, 476. [Google Scholar] [CrossRef] [PubMed]
- Zhao, K.; Sun, Y.; Zhong, S.; Luo, J.-L. The Multifaceted Roles of Cathepsins in Immune and Inflammatory Responses: Implications for Cancer Therapy, Autoimmune Diseases, and Infectious Diseases. Biomark. Res. 2024, 12, 165. [Google Scholar] [CrossRef] [PubMed]
- Cao, M.; Luo, X.; Wu, K.; He, X. Targeting Lysosomes in Human Disease: From Basic Research to Clinical Applications. Signal Transduct. Target. Ther. 2021, 6, 379. [Google Scholar] [CrossRef]
- Ren, H.; Wang, G. Autophagy and Lysosome Storage Disorders. Adv. Exp. Med. Biol. 2020, 1207, 87–102. [Google Scholar] [CrossRef]
- Tang, T.; Yang, Z.; Wang, D.; Yang, X.; Wang, J.; Li, L.; Wen, Q.; Gao, L.; Bian, X.; Yu, S. The Role of Lysosomes in Cancer Development and Progression. Cell. Biosci. 2020, 10, 131. [Google Scholar] [CrossRef]
- Ma, Y.; Galluzzi, L.; Zitvogel, L.; Kroemer, G. Autophagy and Cellular Immune Responses. Immun. Camb. Mass. 2013, 39, 211–227. [Google Scholar] [CrossRef]
- Eriksson, I.; Öllinger, K. Lysosomes in Cancer-At the Crossroad of Good and Evil. Cells 2024, 13, 459. [Google Scholar] [CrossRef]
- Bloomfield, G.; Kay, R.R. Uses and Abuses of Macropinocytosis. J. Cell. Sci. 2016, 129, 2697–2705. [Google Scholar] [CrossRef] [PubMed]
- Lim, J.P.; Gleeson, P.A. Macropinocytosis: An Endocytic Pathway for Internalising Large Gulps. Immunol. Cell. Biol. 2011, 89, 836–843. [Google Scholar] [CrossRef] [PubMed]
- Levin-Konigsberg, R.; Montaño-Rendón, F.; Keren-Kaplan, T.; Li, R.; Ego, B.; Mylvaganam, S.; DiCiccio, J.E.; Trimble, W.S.; Bassik, M.C.; Bonifacino, J.S.; et al. Phagolysosome Resolution Requires Contacts with the Endoplasmic Reticulum and Phosphatidylinositol-4-Phosphate Signalling. Nat. Cell. Biol. 2019, 21, 1234–1247. [Google Scholar] [CrossRef]
- Roche, P.A.; Furuta, K. The Ins and Outs of MHC Class II-Mediated Antigen Processing and Presentation. Nat. Rev. Immunol. 2015, 15, 203–216. [Google Scholar] [CrossRef]
- Gros, M.; Amigorena, S. Regulation of Antigen Export to the Cytosol During Cross-Presentation. Front. Immunol. 2019, 10, 41. [Google Scholar] [CrossRef] [PubMed]
- Canton, J.; Khezri, R.; Glogauer, M.; Grinstein, S. Contrasting Phagosome pH Regulation and Maturation in Human M1 and M2 Macrophages. Mol. Biol. Cell. 2014, 25, 3330–3341. [Google Scholar] [CrossRef] [PubMed]
- Commisso, C.; Davidson, S.M.; Soydaner-Azeloglu, R.G.; Parker, S.J.; Kamphorst, J.J.; Hackett, S.; Grabocka, E.; Nofal, M.; Drebin, J.A.; Thompson, C.B.; et al. Macropinocytosis of Protein Is an Amino Acid Supply Route in Ras-Transformed Cells. Nature 2013, 497, 633–637. [Google Scholar] [CrossRef]
- Lysosomes and Plasma Membrane Repair. In Current Topics in Membranes; Academic Press, 2019; Vol. 84, pp. 1–16.
- Trojani, M.-C.; Santucci-Darmanin, S.; Breuil, V.; Carle, G.F.; Pierrefite-Carle, V. Lysosomal Exocytosis: From Cell Protection to Protumoral Functions. Cancer Lett. 2024, 597, 217024. [Google Scholar] [CrossRef] [PubMed]
- Bulle, A.; Lim, K.-H. Beyond Just a Tight Fortress: Contribution of Stroma to Epithelial-Mesenchymal Transition in Pancreatic Cancer. Signal Transduct. Target. Ther. 2020, 5, 249. [Google Scholar] [CrossRef]
- Liot, S.; Balas, J.; Aubert, A.; Prigent, L.; Mercier-Gouy, P.; Verrier, B.; Bertolino, P.; Hennino, A.; Valcourt, U.; Lambert, E. Stroma Involvement in Pancreatic Ductal Adenocarcinoma: An Overview Focusing on Extracellular Matrix Proteins. Front. Immunol. 2021, 12, 612271. [Google Scholar] [CrossRef]
- Edwards, P.; Kang, B.W.; Chau, I. Targeting the Stroma in the Management of Pancreatic Cancer. Front. Oncol. 2021, 11, 691185. [Google Scholar] [CrossRef]
- Yu, M.; Tannock, I.F. Targeting Tumor Architecture to Favor Drug Penetration: A New Weapon to Combat Chemoresistance in Pancreatic Cancer? Cancer Cell. 2012, 21, 327–329. [Google Scholar] [CrossRef]
- Thomas, D.; Radhakrishnan, P. Tumor-Stromal Crosstalk in Pancreatic Cancer and Tissue Fibrosis. Mol. Cancer 2019, 18, 14. [Google Scholar] [CrossRef]
- M, S.; S, W. Cancer as an Overhealing Wound: An Old Hypothesis Revisited. Nat. Rev. Mol. Cell. Biol. 2008, 9. [Google Scholar] [CrossRef]
- Cirri, P.; Chiarugi, P. Cancer Associated Fibroblasts: The Dark Side of the Coin. Am. J. Cancer Res. 2011, 1, 482–497. [Google Scholar] [PubMed]
- Eyden, B.; Banerjee, S.S.; Shenjere, P.; Fisher, C. The Myofibroblast and Its Tumours. J. Clin. Pathol. 2009, 62, 236–249. [Google Scholar] [CrossRef]
- Klingberg, F.; Hinz, B.; White, E.S. The Myofibroblast Matrix: Implications for Tissue Repair and Fibrosis. J. Pathol. 2013, 229, 298–309. [Google Scholar] [CrossRef]
- Xiang, X.; Niu, Y.-R.; Wang, Z.-H.; Ye, L.-L.; Peng, W.-B.; Zhou, Q. Cancer-Associated Fibroblasts: Vital Suppressors of the Immune Response in the Tumor Microenvironment. Cytokine Growth Factor Rev. 2022, 67, 35–48. [Google Scholar] [CrossRef]
- Hou, W. Role of TGFβ-Activated Cancer-Associated Fibroblasts in the Resistance to Checkpoint Blockade Immunotherapy. Front. Oncol. 2025, 15, 1602452. [Google Scholar] [CrossRef]
- Liao, T.; Chen, X.; Qiu, F.; Zhang, X.; Wu, F.; Zhao, Z.; Xu, M.; Chen, M.; Shen, J.-W.; Shen, Q.; et al. Regulation of Cancer-Associated Fibroblasts for Enhanced Cancer Immunotherapy Using Advanced Functional Nanomedicines: An Updated Review. J. Nanobiotechnology 2025, 23, 166. [Google Scholar] [CrossRef]
- Wang, H.; Inoue, A.; Lei, Y.; Wu, H.; Hong, L.; Cheng, X.W. Cathepsins in the Extracellular Space: Focusing on Non-Lysosomal Proteolytic Functions with Clinical Implications. Cell. Signal. 2023, 103, 110531. [Google Scholar] [CrossRef] [PubMed]
- Sadozai, H.; Acharjee, A.; Kayani, H.Z.; Gruber, T.; Gorczynski, R.M.; Burke, B. High Hypoxia Status in Pancreatic Cancer Is Associated with Multiple Hallmarks of an Immunosuppressive Tumor Microenvironment. Front. Immunol. 2024, 15, 1360629. [Google Scholar] [CrossRef] [PubMed]
- Rahman, M.A.; Yadab, M.K.; Ali, M.M. Emerging Role of Extracellular pH in Tumor Microenvironment as a Therapeutic Target for Cancer Immunotherapy. Cells 2024, 13, 1924. [Google Scholar] [CrossRef] [PubMed]
- Vidak, E.; Javoršek, U.; Vizovišek, M.; Turk, B. Cysteine Cathepsins and Their Extracellular Roles: Shaping the Microenvironment. Cells 2019, 8, 264. [Google Scholar] [CrossRef]
- Peppicelli, S.; Calorini, L.; Bianchini, F.; Papucci, L.; Magnelli, L.; Andreucci, E. Acidity and Hypoxia of Tumor Microenvironment, a Positive Interplay in Extracellular Vesicle Release by Tumor Cells. Cell. Oncol. 2025, 48, 27–41. [Google Scholar] [CrossRef] [PubMed]
- Bogdanov, A.; Bogdanov, A.; Chubenko, V.; Volkov, N.; Moiseenko, F.; Moiseyenko, V. Tumor Acidity: From Hallmark of Cancer to Target of Treatment. Front. Oncol. 2022, 12, 979154. [Google Scholar] [CrossRef]
- Sheehan, C.; Muir, A. What’s on the Menu?: Metabolic Constraints in the Pancreatic Tumor Microenvironment. J. Clin. Invest. 135, e191940. [CrossRef]
- Mondal, G.; Debnath, J. NRF2 Activates Macropinocytosis upon Autophagy Inhibition. Cancer Cell. 2021, 39, 596–598. [Google Scholar] [CrossRef]
- Qiu, Z.; Liu, W.; Zhu, Q.; Ke, K.; Zhu, Q.; Jin, W.; Yu, S.; Yang, Z.; Li, L.; Sun, X.; et al. The Role and Therapeutic Potential of Macropinocytosis in Cancer. Front. Pharmacol. 2022, 13, 919819. [Google Scholar] [CrossRef]
- Yamamoto, K.; Venida, A.; Yano, J.; Biancur, D.E.; Kakiuchi, M.; Gupta, S.; Sohn, A.S.W.; Mukhopadhyay, S.; Lin, E.Y.; Parker, S.J.; et al. Autophagy Promotes Immune Evasion of Pancreatic Cancer by Degrading MHC-I. Nature 2020, 581, 100–105. [Google Scholar] [CrossRef]
- Piersma, B.; Hayward, M.-K.; Weaver, V.M. Fibrosis and Cancer: A Strained Relationship. Biochim. Biophys. Acta BBA-Rev. Cancer 2020, 1873, 188356. [Google Scholar] [CrossRef]
- Manoukian, P.; Damhofer, H.; Zhao, L.; van Laarhoven, H.W.M.; Bijlsma, M.F. Stromal Hedgehog Signaling Is Associated with Favorable Outcomes in Pancreatic Cancer. Int. J. Mol. Sci. 2025, 26, 5200. [Google Scholar] [CrossRef] [PubMed]
- Yamamoto, K.; Venida, A.; Perera, R.M.; Kimmelman, A.C. Selective Autophagy of MHC-I Promotes Immune Evasion of Pancreatic Cancer. Autophagy 2020, 16, 1524–1525. [Google Scholar] [CrossRef] [PubMed]
- Saftig, P.; Puertollano, R. How Lysosomes Sense, Integrate, and Cope with Stress. Trends Biochem. Sci. 2021, 46, 97–112. [Google Scholar] [CrossRef]
- Ballabio, A.; Bonifacino, J.S. Lysosomes as Dynamic Regulators of Cell and Organismal Homeostasis. Nat. Rev. Mol. Cell. Biol. 2020, 21, 101–118. [Google Scholar] [CrossRef] [PubMed]
- Evavold, C.L.; Kagan, J.C. Inflammasomes: Threat-Assessment Organelles of the Innate Immune System. Immunity 2019, 51, 609–624. [Google Scholar] [CrossRef]
- Cullen, S.P.; Kearney, C.J.; Clancy, D.M.; Martin, S.J. Diverse Activators of the NLRP3 Inflammasome Promote IL-1β Secretion by Triggering Necrosis. Cell. Rep. 2015, 11, 1535–1548. [Google Scholar] [CrossRef]
- Morawiec, M.-L.; Kubina, R.; Jabłońska, E.; Ratajczak-Wrona, W.; Stępień, S.; Gołębski, M.; Mielczarek-Palacz, A. NETs - as Predictors and Targets of Supportive Therapy for Cancer Treatment. Front. Immunol. 2025, 16. [Google Scholar] [CrossRef]
- He, X.-Y.; Ng, D.; Egeblad, M. Caught in a Web: Emerging Roles of Neutrophil Extracellular Traps in Cancer. Annu. Rev. Cancer Biol. 2022, 6, 223–243. [Google Scholar] [CrossRef]
- Repnik, U.; Hafner Česen, M.; Turk, B. Lysosomal Membrane Permeabilization in Cell Death: Concepts and Challenges. Mitochondrion 2014, 19 Pt A, 49–57. [Google Scholar] [CrossRef] [PubMed]
- Aits, S.; Jäättelä, M. Lysosomal Cell Death at a Glance. J. Cell. Sci. 2013, 126, 1905–1912. [Google Scholar] [CrossRef]
- Fujii, Y.; Asadi, Z.; Mehla, K. Cathepsins: Emerging Targets in the Tumor Ecosystem to Overcome Cancers. Semin. Cancer Biol. 2025, 112, 150–166. [Google Scholar] [CrossRef]
- Turk, V.; Stoka, V.; Vasiljeva, O.; Renko, M.; Sun, T.; Turk, B.; Turk, D. Cysteine Cathepsins: From Structure, Function and Regulation to New Frontiers. Biochim. Biophys. Acta 2012, 1824, 68–88. [Google Scholar] [CrossRef] [PubMed]
- Mohamed, M.M.; Sloane, B.F. Cysteine Cathepsins: Multifunctional Enzymes in Cancer. Nat. Rev. Cancer 2006, 6, 764–775. [Google Scholar] [CrossRef] [PubMed]
- Nägler, D.K.; Zhang, R.; Tam, W.; Sulea, T.; Purisima, E.O.; Ménard, R. Human Cathepsin X: A Cysteine Protease with Unique Carboxypeptidase Activity. Biochemistry 1999, 38, 12648–12654. [Google Scholar] [CrossRef]
- Mort, J.S.; Buttle, D.J. Cathepsin B. Int. J. Biochem. Cell. Biol. 1997, 29, 715–720. [Google Scholar] [CrossRef] [PubMed]
- Löser, R.; Pietzsch, J. Cysteine Cathepsins: Their Role in Tumor Progression and Recent Trends in the Development of Imaging Probes. Front. Chem. 2015, 3. [Google Scholar] [CrossRef]
- Bakst, R.L.; Xiong, H.; Chen, C.-H.; Deborde, S.; Lyubchik, A.; Zhou, Y.; He, S.; McNamara, W.; Lee, S.-Y.; Olson, O.C.; et al. Inflammatory Monocytes Promote Perineural Invasion via CCL2-Mediated Recruitment and Cathepsin B Expression. Cancer Res. 2017, 77, 6400–6414. [Google Scholar] [CrossRef]
- Lines, K.E.; Chelala, C.; Dmitrovic, B.; Wijesuriya, N.; Kocher, H.M.; Marshall, J.F.; Crnogorac-Jurcevic, T. S100P-Binding Protein, S100PBP, Mediates Adhesion through Regulation of Cathepsin Z in Pancreatic Cancer Cells. Am. J. Pathol. 2012, 180, 1485–1494. [Google Scholar] [CrossRef] [PubMed]
- Gocheva, V.; Chen, X.; Peters, C.; Reinheckel, T.; Joyce, J.A. Deletion of Cathepsin H Perturbs Angiogenic Switching, Vascularization and Growth of Tumors in a Mouse Model of Pancreatic Islet Cell Cancer. Biol. Chem. 2010, 391, 937–945. [Google Scholar] [CrossRef]
- Khojasteh-Leylakoohi, F.; Mohit, R.; Khalili-Tanha, N.; Asadnia, A.; Naderi, H.; Pourali, G.; Yousefli, Z.; Khalili-Tanha, G.; Khazaei, M.; Maftooh, M.; et al. Down Regulation of Cathepsin W Is Associated with Poor Prognosis in Pancreatic Cancer. Sci. Rep. 2023, 13, 16678. [Google Scholar] [CrossRef]
- Park, H.-D.; Kang, E.-S.; Kim, J.-W.; Lee, K.-T.; Lee, K.H.; Park, Y.S.; Park, J.-O.; Lee, J.; Heo, J.S.; Choi, S.H.; et al. Serum CA19-9, Cathepsin D, and Matrix Metalloproteinase-7 as a Diagnostic Panel for Pancreatic Ductal Adenocarcinoma. Proteomics 2012, 12, 3590–3597. [Google Scholar] [CrossRef] [PubMed]
- Vetvicka, V.; Fusek, M. Procathepsin D as a Tumor Marker, Anti-Cancer Drug or Screening Agent. Anticancer Agents Med. Chem. 12, 172–175. [CrossRef]
- Dumartin, L.; Whiteman, H.J.; Weeks, M.E.; Hariharan, D.; Dmitrovic, B.; Iacobuzio-Donahue, C.A.; Brentnall, T.A.; Bronner, M.P.; Feakins, R.M.; Timms, J.F.; et al. AGR2 Is a Novel Surface Antigen That Promotes the Dissemination of Pancreatic Cancer Cells through Regulation of Cathepsins B and D. Cancer Res. 2011, 71, 7091–7102. [Google Scholar] [CrossRef]
- Cruz-Monserrate, Z.; Abd-Elgaliel, W.R.; Grote, T.; Deng, D.; Ji, B.; Arumugam, T.; Wang, H.; Tung, C.-H.; Logsdon, C.D. Detection of Pancreatic Cancer Tumours and Precursor Lesions by Cathepsin E Activity in Mouse Models. Gut 2012, 61, 1315–1322. [Google Scholar] [CrossRef]
- Azuma, T.; Hirai, M.; Ito, S.; Yamamoto, K.; Taggart, R.T.; Matsuba, T.; Yasukawa, K.; Uno, K.; Hayakumo, T.; Nakajima, M. Expression of Cathepsin E in Pancreas: A Possible Tumor Marker for Pancreas, a Preliminary Report. Int. J. Cancer 1996, 67, 492–497. [Google Scholar] [CrossRef]
- Dupont, S.; Morsut, L.; Aragona, M.; Enzo, E.; Giulitti, S.; Cordenonsi, M.; Zanconato, F.; Le Digabel, J.; Forcato, M.; Bicciato, S.; et al. Role of YAP/TAZ in Mechanotransduction. Nature 2011, 474, 179–183. [Google Scholar] [CrossRef] [PubMed]
- Xiao, Y.; Zhang, H.; Ma, Q.; Huang, R.; Lu, J.; Liang, X.; Liu, X.; Zhang, Z.; Yu, L.; Pang, J.; et al. YAP1-Mediated Pancreatic Stellate Cell Activation Inhibits Pancreatic Cancer Cell Proliferation. Cancer Lett. 2019, 462, 51–60. [Google Scholar] [CrossRef] [PubMed]
- Turk, V.; Turk, B.; Turk, D. Lysosomal Cysteine Proteases: Facts and Opportunities. EMBO J. 2001, 20, 4629–4633. [Google Scholar] [CrossRef]
- Saftig, P.; Klumperman, J. Lysosome Biogenesis and Lysosomal Membrane Proteins: Trafficking Meets Function. Nat. Rev. Mol. Cell. Biol. 2009, 10, 623–635. [Google Scholar] [CrossRef]
- Komura, T.; Takabatake, H.; Harada, K.; Yamato, M.; Miyazawa, M.; Yoshida, K.; Honda, M.; Wada, T.; Kitagawa, H.; Ohta, T.; et al. Clinical Features of Cystatin A Expression in Patients with Pancreatic Ductal Adenocarcinoma. Cancer Sci. 2017, 108, 2122–2129. [Google Scholar] [CrossRef]
- Görgülü, K.; Diakopoulos, K.N.; Ai, J.; Schoeps, B.; Kabacaoglu, D.; Karpathaki, A.-F.; Ciecielski, K.J.; Kaya-Aksoy, E.; Ruess, D.A.; Berninger, A.; et al. Levels of the Autophagy-Related 5 Protein Affect Progression and Metastasis of Pancreatic Tumors in Mice. Gastroenterology 2019, 156, 203–217.e20. [Google Scholar] [CrossRef]
- B, M.; Ma, P.; C, L.; Lh, T.; S, J.; Mj, B. Gemcitabine Promotes Autophagy and Lysosomal Function through ERK- and TFEB-Dependent Mechanisms. Cell. Death Discov. 2023, 9. [Google Scholar] [CrossRef]
- Conesa-Bakkali, R.; Morillo-Huesca, M.; Martínez-Fábregas, J. Non-Canonical, Extralysosomal Activities of Lysosomal Peptidases in Physiological and Pathological Conditions: New Clinical Opportunities for Cancer Therapy. Cells 2025, 14. [Google Scholar] [CrossRef] [PubMed]
- Droga-Mazovec, G.; Bojic, L.; Petelin, A.; Ivanova, S.; Romih, R.; Repnik, U.; Salvesen, G.S.; Stoka, V.; Turk, V.; Turk, B. Cysteine Cathepsins Trigger Caspase-Dependent Cell Death through Cleavage of Bid and Antiapoptotic Bcl-2 Homologues. J. Biol. Chem. 2008, 283, 19140–19150. [Google Scholar] [CrossRef]
- Enneking, A.E.; Khorey, M.M.; Edgington-Mitchell, L.E. Nuclear Roles for Canonically Lysosomal Proteases. Mol. Cell. Biol. 2025, 45, 343–352. [Google Scholar] [CrossRef]
- Singh, N.; Das, P.; Gupta, S.; Sachdev, V.; Srivasatava, S.; Datta Gupta, S.; Pandey, R.M.; Sahni, P.; Chauhan, S.S.; Saraya, A. Plasma Cathepsin L: A Prognostic Marker for Pancreatic Cancer. World J. Gastroenterol. 2014, 20, 17532–17540. [Google Scholar] [CrossRef]
- Fujimoto, T.; Tsunedomi, R.; Matsukuma, S.; Yoshimura, K.; Oga, A.; Fujiwara, N.; Fujiwara, Y.; Matsui, H.; Shindo, Y.; Tokumitsu, Y.; et al. Cathepsin B Is Highly Expressed in Pancreatic Cancer Stem-like Cells and Is Associated with Patients’ Surgical Outcomes. Oncol. Lett. 2021, 21, 1–1. [Google Scholar] [CrossRef]
- Singh, N.; Das, P.; Datta Gupta, S.; Sahni, P.; Pandey, R.M.; Gupta, S.; Chauhan, S.S.; Saraya, A. Prognostic Significance of Extracellular Matrix Degrading Enzymes-Cathepsin L and Matrix Metalloproteases-2 [MMP-2] in Human Pancreatic Cancer. Cancer Invest. 2013, 31, 461–471. [Google Scholar] [CrossRef] [PubMed]
- Niedergethmann, M.; Wostbrock, B.; Sturm, J.W.; Willeke, F.; Post, S.; Hildenbrand, R. Prognostic Impact of Cysteine Proteases Cathepsin B and Cathepsin L in Pancreatic Adenocarcinoma. Pancreas 2004, 29, 204–211. [Google Scholar] [CrossRef]
- Kalubowilage, M.; Covarrubias-Zambrano, O.; Malalasekera, A.P.; Wendel, S.O.; Wang, H.; Yapa, A.S.; Chlebanowski, L.; Toledo, Y.; Ortega, R.; Janik, K.E.; et al. Early Detection of Pancreatic Cancers in Liquid Biopsies by Ultrasensitive Fluorescence Nanobiosensors. Nanomed. Nanotechnol. Biol. Med. 2018, 14, 1823–1832. [Google Scholar] [CrossRef]
- Olson, O.C.; Joyce, J.A. Cysteine Cathepsin Proteases: Regulators of Cancer Progression and Therapeutic Response. Nat. Rev. Cancer 2015, 15, 712–729. [Google Scholar] [CrossRef]
- Gutierrez-Ruiz, O.L.; Johnson, K.M.; Krueger, E.W.; Nooren, R.E.; Cruz-Reyes, N.; Heppelmann, C.J.; Hogenson, T.L.; Fernandez-Zapico, M.E.; McNiven, M.A.; Razidlo, G.L. Ectopic Expression of DOCK8 Regulates Lysosome-Mediated Pancreatic Tumor Cell Invasion. Cell. Rep. 2023, 42, 113042. [Google Scholar] [CrossRef] [PubMed]
- Hwang, J.-H.; Lee, S.H.; Lee, K.H.; Lee, K.Y.; Kim, H.; Ryu, J.K.; Yoon, Y.B.; Kim, Y.-T. Cathepsin B Is a Target of Hedgehog Signaling in Pancreatic Cancer. Cancer Lett. 2009, 273, 266–272. [Google Scholar] [CrossRef]
- Yuan, J.; Ofengeim, D. A Guide to Cell Death Pathways. Nat. Rev. Mol. Cell. Biol. 2024, 25, 379–395. [Google Scholar] [CrossRef] [PubMed]
- Jiang, Y.; Han, L.; Xue, M.; Wang, T.; Zhu, Y.; Xiong, C.; Shi, M.; Li, H.; Hai, W.; Huo, Y.; et al. Cystatin B Increases Autophagic Flux by Sustaining Proteolytic Activity of Cathepsin B and Fuels Glycolysis in Pancreatic Cancer: CSTB Orchestrates Autophagy and Glycolysis in PDAC. Clin. Transl. Med. 2022, 12, e1126. [Google Scholar] [CrossRef] [PubMed]
- Jakoš, T.; Pišlar, A.; Jewett, A.; Kos, J. Cysteine Cathepsins in Tumor-Associated Immune Cells. Front. Immunol. 2019, 10, 2037. [Google Scholar] [CrossRef]
- Turk, D.; Janjić, V.; Štern, I.; Podobnik, M.; Lamba, D.; Weis Dahl, S.; Lauritzen, C.; Pedersen, J.; Turk, V.; Turk, B. Structure of Human Dipeptidyl Peptidase I (Cathepsin C): Exclusion Domain Added to an Endopeptidase Framework Creates the Machine for Activation of Granular Serine Proteases. EMBO J. 2001, 20, 6570–6582. [Google Scholar] [CrossRef] [PubMed]
- Sulpizio, S.; Franceschini, N.; Piattelli, A.; Di Sebastiano, P.; Innocenti, P.; Selvaggi, F. Cathepsins and Pancreatic Cancer: The 2012 Update. Pancreatology 2012, 12, 395–401. [Google Scholar] [CrossRef]
- Gopinathan, A.; Denicola, G.M.; Frese, K.K.; Cook, N.; Karreth, F.A.; Mayerle, J.; Lerch, M.M.; Reinheckel, T.; Tuveson, D.A. Cathepsin B Promotes the Progression of Pancreatic Ductal Adenocarcinoma in Mice. Gut 2012, 61. [Google Scholar] [CrossRef]
- Brömme, D.; Li, Z.; Barnes, M.; Mehler, E. Human Cathepsin V Functional Expression, Tissue Distribution, Electrostatic Surface Potential, Enzymatic Characterization, and Chromosomal Localization. Biochemistry 1999, 38, 2377–2385. [Google Scholar] [CrossRef]
- Canè, S.; Barouni, R.M.; Fabbi, M.; Cuozzo, J.; Fracasso, G.; Adamo, A.; Ugel, S.; Trovato, R.; De Sanctis, F.; Giacca, M.; et al. Neutralization of NET-Associated Human ARG1 Enhances Cancer Immunotherapy. Sci. Transl. Med. 2023, 15, eabq6221. [Google Scholar] [CrossRef]
- Karimi, E.; Kalaki, N.S.; Akrami, S.M. Identifying Hub Genes and Pathways in Pancreatic Ductal Adenocarcinoma (PAAD): A Comprehensive in Silico Study. Biochem. Biophys. Rep. 2025, 41, 101921. [Google Scholar] [CrossRef]
- Giriyappagoudar, M.; Vastrad, B.; Horakeri, R.; Vastrad, C. Identification and Interaction Analysis of Molecular Markers in Pancreatic Ductal Adenocarcinoma by Bioinformatics and Next-Generation Sequencing Data Analysis. Bioinforma. Biol. Insights 2023, 17, 11779322231186719. [Google Scholar] [CrossRef]
- Aghdassi, A.A.; Pham, C.; Zierke, L.; Mariaule, V.; Korkmaz, B.; Rhimi, M. Cathepsin C Role in Inflammatory Gastroenterological, Renal, Rheumatic, and Pulmonary Disorders. Biochimie 2024, 216, 175–180. [Google Scholar] [CrossRef] [PubMed]
- Zheng, L.; Liu, X.; Xi, Y.; Gong, D.; Ge, B.; Wei, X.; Cai, J.; Chen, P. A Pan-Cancer Analysis of CTSC as a Candidate Prognostic and Immune-Related Biomarker. Discov. Oncol. 2025, 16, 2258. [Google Scholar] [CrossRef] [PubMed]
- Akkari, L.; Gocheva, V.; Kester, J.C.; Hunter, K.E.; Quick, M.L.; Sevenich, L.; Wang, H.-W.; Peters, C.; Tang, L.H.; Klimstra, D.S.; et al. Distinct Functions of Macrophage-Derived and Cancer Cell-Derived Cathepsin Z Combine to Promote Tumor Malignancy via Interactions with the Extracellular Matrix. Genes. Dev. 2014, 28, 2134–2150. [Google Scholar] [CrossRef]
- Wex, T.; Bühling, F.; Wex, H.; Günther, D.; Malfertheiner, P.; Weber, E.; Brömme, D. Human Cathepsin W, a Cysteine Protease Predominantly Expressed in NK Cells, Is Mainly Localized in the Endoplasmic Reticulum. J. Immunol. 2001, 167, 2172–2178. [Google Scholar] [CrossRef]
- Ding, Y.; Li, Z.; Wang, H.; Wang, Q.; Jiang, H.; Yu, Z.; Xu, M. CTSK and PLAU as Prognostic Biomarker and Related to Immune Infiltration in Pancreatic Cancer: Evidence from Bioinformatics Analysis and qPCR. Int. J. Genom. 2023, 2023, 3914687. [Google Scholar] [CrossRef]
- Zhang, T.; Chen, J.; Yang, H.; Sun, X.; Ou, Y.; Wang, Q.; Edderkaoui, M.; Zheng, S.; Ren, F.; Tong, Y.; et al. Stromal Softness Confines Pancreatic Cancer Growth through Lysosomal-Cathepsin Mediated YAP1 Degradation. Cell. Mol. Life Sci. CMLS 2024, 81, 442. [Google Scholar] [CrossRef] [PubMed]
- Huang, X.; Deng, H.; Zhang, B.; Wang, K.; Qu, Y.; Li, T.; Liu, T. The Causal Relationship between Cathepsins and Digestive System Tumors: A Mendelian Randomization Study. Front. Oncol. 2024, 14. [Google Scholar] [CrossRef]
- Mahajan, U.M.; Goni, E.; Langhoff, E.; Li, Q.; Costello, E.; Greenhalf, W.; Kruger, S.; Ormanns, S.; Halloran, C.; Ganeh, P.; et al. Cathepsin D Expression and Gemcitabine Resistance in Pancreatic Cancer. JNCI Cancer Spectr. 2019, 4, pkz060. [Google Scholar] [CrossRef] [PubMed]
- Uno, K.; Azuma, T.; Nakajima, M.; Yasuda, K.; Hayakumo, T.; Mukai, H.; Sakai, T.; Kawai, K. Clinical Significance of Cathepsin E in Pancreatic Juice in the Diagnosis of Pancreatic Ductal Adenocarcinoma. J. Gastroenterol. Hepatol. 2000, 15, 1333–1338. [Google Scholar] [CrossRef]
- Watts, C. Lysosomes and Lysosome-Related Organelles in Immune Responses. FEBS Open. Bio 2022, 12, 678–693. [Google Scholar] [CrossRef]
- DeVorkin, L.; Pavey, N.; Carleton, G.; Comber, A.; Ho, C.; Lim, J.; McNamara, E.; Huang, H.; Kim, P.; Zacharias, L.G.; et al. Autophagy Regulation of Metabolism Is Required for CD8+ T Cell Anti-Tumor Immunity. Cell. Rep. 2019, 27, 502–513.e5. [Google Scholar] [CrossRef]
- Riese, R.J.; Wolf, P.R.; Brömme, D.; Natkin, L.R.; Villadangos, J.A.; Ploegh, H.L.; Chapman, H.A. Essential Role for Cathepsin S in MHC Class II-Associated Invariant Chain Processing and Peptide Loading. Immunity 1996, 4, 357–366. [Google Scholar] [CrossRef] [PubMed]
- Gao, H.; Zhang, Z.; Deng, J.; Song, Y. Cathepsin S: Molecular Mechanisms in Inflammatory and Immunological Processes. Front. Immunol. 2025, 16, 1600206. [Google Scholar] [CrossRef]
- Berquez, M.; Li, A.L.; Luy, M.A.; Venida, A.C.; O’Loughlin, T.; Rademaker, G.; Barpanda, A.; Hu, J.; Yano, J.; Wiita, A.; et al. A Multi-Subunit Autophagic Capture Complex Facilitates Degradation of ER Stalled MHC-I in Pancreatic Cancer. BioRxiv Prepr. Serv. Biol. 2024, 10.27.620516. [Google Scholar] [CrossRef]
- Baranov, M.V.; Bianchi, F.; Schirmacher, A.; van Aart, M.A.C.; Maassen, S.; Muntjewerff, E.M.; Dingjan, I.; Ter Beest, M.; Verdoes, M.; Keyser, S.G.L.; et al. The Phosphoinositide Kinase PIKfyve Promotes Cathepsin-S-Mediated Major Histocompatibility Complex Class II Antigen Presentation. iScience 2019, 11, 160–177. [Google Scholar] [CrossRef]
- Roh, K.; Noh, J.; Kim, Y.; Jang, Y.; Kim, J.; Choi, H.; Lee, Y.; Ji, M.; Kang, D.; Kim, M.-S.; et al. Lysosomal Control of Senescence and Inflammation through Cholesterol Partitioning. Nat. Metab. 2023, 5, 398–413. [Google Scholar] [CrossRef] [PubMed]
- Oelschlaegel, D.; Weiss Sadan, T.; Salpeter, S.; Krug, S.; Blum, G.; Schmitz, W.; Schulze, A.; Michl, P. Cathepsin Inhibition Modulates Metabolism and Polarization of Tumor-Associated Macrophages. Cancers 2020, 12, 2579. [Google Scholar] [CrossRef]
- Man, S.M.; Kanneganti, T.-D. Regulation of Lysosomal Dynamics and Autophagy by CTSB/Cathepsin B. Autophagy 2016, 12, 2504–2505. [Google Scholar] [CrossRef]
- Iwama, H.; Mehanna, S.; Imasaka, M.; Hashidume, S.; Nishiura, H.; Yamamura, K.-I.; Suzuki, C.; Uchiyama, Y.; Hatano, E.; Ohmuraya, M. Cathepsin B and D Deficiency in the Mouse Pancreas Induces Impaired Autophagy and Chronic Pancreatitis. Sci. Rep. 2021, 11, 6596. [Google Scholar] [CrossRef]
- Yeung, B.H.Y.; Huang, D.-C.; Sinicrope, F.A. PS-341 (Bortezomib) Induces Lysosomal Cathepsin B Release and a Caspase-2-Dependent Mitochondrial Permeabilization and Apoptosis in Human Pancreatic Cancer Cells. J. Biol. Chem. 2006, 281, 11923–11932. [Google Scholar] [CrossRef] [PubMed]
- Parreño, M.; Casanova, I.; Céspedes, M.V.; Vaqué, J.P.; Pavón, M.A.; Leon, J.; Mangues, R. Bobel-24 and Derivatives Induce Caspase-Independent Death in Pancreatic Cancer Regardless of Apoptotic Resistance. Cancer Res. 2008, 68, 6313–6323. [Google Scholar] [CrossRef]
- Qu, M.; Zhu, C.; Sun, C.; Zhu, S.; Zhang, H.; Miao, C.; Zhou, D. Neutrophil Extracellular Traps Promote Pancreatic Cancer Progression via the STING Pathway. Gastroenterol. Res. Pract. 2025, 2025, 4950214. [Google Scholar] [CrossRef]
- Kuang, F.; Liu, J.; Li, C.; Kang, R.; Tang, D. Cathepsin B Is a Mediator of Organelle-Specific Initiation of Ferroptosis. Biochem. Biophys. Res. Commun. 2020, 533, 1464–1469. [Google Scholar] [CrossRef]
- Piffoux, M.; Eriau, E.; Cassier, P.A. Autophagy as a Therapeutic Target in Pancreatic Cancer. Br. J. Cancer 2021, 124, 333–344. [Google Scholar] [CrossRef]
- Elliott, I.A.; Dann, A.M.; Xu, S.; Kim, S.S.; Abt, E.R.; Kim, W.; Poddar, S.; Moore, A.; Zhou, L.; Williams, J.L.; et al. Lysosome Inhibition Sensitizes Pancreatic Cancer to Replication Stress by Aspartate Depletion. Proc. Natl. Acad. Sci. U. S. A 2019, 116, 6842–6847. [Google Scholar] [CrossRef]
- Fu, Z.; Cheng, X.; Kuang, J.; Feng, H.; Chen, L.; Liang, J.; Shen, X.; Yuen, S.; Peng, C.; Shen, B.; et al. CQ Sensitizes Human Pancreatic Cancer Cells to Gemcitabine through the Lysosomal Apoptotic Pathway via Reactive Oxygen Species. Mol. Oncol. 2018, 12, 529–544. [Google Scholar] [CrossRef] [PubMed]
- Klein, K.; Werner, K.; Teske, C.; Schenk, M.; Giese, T.; Weitz, J.; Welsch, T. Role of TFEB-Driven Autophagy Regulation in Pancreatic Cancer Treatment. Int. J. Oncol. 2016, 49, 164–172. [Google Scholar] [CrossRef]
- Kim, J.H.; Lee, J.; Cho, Y.-R.; Lee, S.-Y.; Sung, G.-J.; Shin, D.-M.; Choi, K.-C.; Son, J. TFEB Supports Pancreatic Cancer Growth through the Transcriptional Regulation of Glutaminase. Cancers 2021, 13. [Google Scholar] [CrossRef]
- Jain, V.; Harper, S.L.; Versace, A.M.; Fingerman, D.; Brown, G.S.; Bhardwaj, M.; Crissey, M.A.S.; Goldman, A.R.; Ruthel, G.; Liu, Q.; et al. Targeting UGCG Overcomes Resistance to Lysosomal Autophagy Inhibition. Cancer Discov. 2023, 13, 454–473. [Google Scholar] [CrossRef]
- Phadatare, P.; Debnath, J. Lysosomal Lipid Peroxidation Mediates Immunogenic Cell Death. J. Clin. Invest. 2023, 133. [Google Scholar] [CrossRef] [PubMed]
- Cheng, C.; Hu, J.; Mannan, R.; He, T.; Bhattacharyya, R.; Magnuson, B.; Wisniewski, J.P.; Peters, S.; Karim, S.A.; MacLean, D.J.; et al. Targeting PIKfyve-Driven Lipid Metabolism in Pancreatic Cancer. Nature 2025, 642, 776–784. [Google Scholar] [CrossRef] [PubMed]
- Bryant, K.L.; Der, C.J. Blocking Autophagy to Starve Pancreatic Cancer. Nat. Rev. Mol. Cell. Biol. 2019, 20, 265–265. [Google Scholar] [CrossRef] [PubMed]
- Zhao, B.; Dierichs, L.; Gu, J.-N.; Trajkovic-Arsic, M.; Axel Hilger, R.; Savvatakis, K.; Vega-Rubin-de-Celis, S.; Liffers, S.-T.; Peña-Llopis, S.; Behrens, D.; et al. TFEB-Mediated Lysosomal Biogenesis and Lysosomal Drug Sequestration Confer Resistance to MEK Inhibition in Pancreatic Cancer. Cell. Death Discov. 2020, 6, 12. [Google Scholar] [CrossRef]
- Karasic, T.B.; O’Hara, M.H.; Loaiza-Bonilla, A.; Reiss, K.A.; Teitelbaum, U.R.; Borazanci, E.; De Jesus-Acosta, A.; Redlinger, C.; Burrell, J.A.; Laheru, D.A.; et al. Effect of Gemcitabine and Nab-Paclitaxel With or Without Hydroxychloroquine on Patients With Advanced Pancreatic Cancer: A Phase 2 Randomized Clinical Trial. JAMA Oncol. 2019, 5, 993–998. [Google Scholar] [CrossRef]
- Zeh, H.J.; Bahary, N.; Boone, B.A.; Singhi, A.D.; Miller-Ocuin, J.L.; Normolle, D.P.; Zureikat, A.H.; Hogg, M.E.; Bartlett, D.L.; Lee, K.K.; et al. A Randomized Phase II Preoperative Study of Autophagy Inhibition with High-Dose Hydroxychloroquine and Gemcitabine/Nab-Paclitaxel in Pancreatic Cancer Patients. Clin. Cancer Res. Off. J. Am. Assoc. Cancer Res. 2020, 26, 3126–3134. [Google Scholar] [CrossRef]
- Cash, T.P.; Alcalá, S.; Rico-Ferreira, M.D.R.; Hernández-Encinas, E.; García, J.; Albarrán, M.I.; Valle, S.; Muñoz, J.; Martínez-González, S.; Blanco-Aparicio, C.; et al. Induction of Lysosome Membrane Permeabilization as a Therapeutic Strategy to Target Pancreatic Cancer Stem Cells. Cancers 2020, 12, 1790. [Google Scholar] [CrossRef]
- Stylianopoulos, T.; Munn, L.; Jain, R. Reengineering the Tumor Vasculature: Improving Drug Delivery and Efficacy. Trends Cancer 2018, 4. [Google Scholar] [CrossRef]
- Reinheckel, T.; Peters, C.; Krüger, A.; Turk, B.; Vasiljeva, O. Differential Impact of Cysteine Cathepsins on Genetic Mouse Models of De Novo Carcinogenesis: Cathepsin B as Emerging Therapeutic Target. Front. Pharmacol. 2012, 3. [Google Scholar] [CrossRef]
- Zhang, C.; Sun, J.; Liu, D.; Cui, Y.; Liu, S.; Sun, H. Preliminary Research on the Pathological Role of Cathepsin-B in Subcutaneous Heteroplastic Pancreatic Carcinoma in Nude Mice. Chin. Med. J. (Engl.) 2009, 122, 2489–2496. [Google Scholar] [PubMed]
- Huang, H.; Zhang, Y.; Xu, X.; Liu, Y.; Zhao, J.; Ma, L.; Lei, J.; Ge, W.; Li, N.; Ma, E.; et al. Design and Synthesis of Dual Cathepsin L and S Inhibitors and Antimetastatic Activity Evaluation in Pancreatic Cancer Cells. Bioorg. Med. Chem. Lett. 2023, 80, 129087. [Google Scholar] [CrossRef] [PubMed]
- Citarella, A.; Petrella, S.; Moi, D.; Dimasi, A.; Braga, T.; Ruberto, L.; Pieraccini, S.; Sironi, M.; Micale, N.; Schirmeister, T.; et al. Synthesis of α-Fluorocinnamate Derivatives as Novel Cathepsin S Inhibitors with in Vitro Antiproliferative Activity against Pancreatic Cancer Cells. Bioorg. Med. Chem. 2024, 115, 117987. [Google Scholar] [CrossRef]
- Elie, B.T.; Gocheva, V.; Shree, T.; Dalrymple, S.A.; Holsinger, L.J.; Joyce, J.A. Identification and Pre-Clinical Testing of a Reversible Cathepsin Protease Inhibitor Reveals Anti-Tumor Efficacy in a Pancreatic Cancer Model. Biochimie 2010, 92, 1618–1624. [Google Scholar] [CrossRef] [PubMed]
- Cho, Y.; Lee, J.H.; Kim, J.H.; Lee, S.; Yoo, S.; Jung, M.; Kim, S.J.; Yoo, H.J.; Pack, C.; Rho, J.K.; et al. Matrine Suppresses KRAS-driven Pancreatic Cancer Growth by Inhibiting Autophagy-mediated Energy Metabolism. Mol. Oncol. 2018, 12, 1203–1215. [Google Scholar] [CrossRef]
- Mirković, B.; Markelc, B.; Butinar, M.; et al. Nitroxoline impairs tumor progression in vitro and in vivo by regulating cathepsin B activity. Oncotarget 2015, 6(22), 19027–19042. [Google Scholar] [CrossRef]
- Fonović, U.P.; Mitrović, A.; Knez, D.; et al. Identification and characterization of the novel reversible and selective cathepsin X inhibitors. Sci. Rep. 2017, 7, 11459. [Google Scholar] [CrossRef]
- Mitrović, A.; Završnik, J.; Mikhaylov, G.; et al. Evaluation of novel cathepsin-X inhibitors in vitro and in vivo and their ability to improve cathepsin-B-directed antitumor therapy. Cell. Mol. Life Sci. Published. 2022, 79(1), 34. [Google Scholar] [CrossRef]
- Hadi, M.M.; Farrell, S.; Nesbitt, H.; Thomas, K.; Kubajewska, I.; Ng, A.; Masood, H.; Patel, S.; Sciscione, F.; Davidson, B.; et al. Nanotechnology-Augmented Sonodynamic Therapy and Associated Immune-Mediated Effects for the Treatment of Pancreatic Ductal Adenocarcinoma. J. Cancer Res. Clin. Oncol. 2023, 149, 5007–5023. [Google Scholar] [CrossRef]
- Han, H.; Jin, Q.; Wang, Y.; Chen, Y.; Ji, J. The Rational Design of a Gemcitabine Prodrug with AIE-Based Intracellular Light-up Characteristics for Selective Suppression of Pancreatic Cancer Cells. Chem. Commun. 2015, 51, 17435–17438. [Google Scholar] [CrossRef]
- Fang, Y.; Du, F.; Xu, Y.; Meng, H.; Huang, J.; Zhang, X.; Lu, W.; Liu, S.; Yu, J. Enhanced Cellular Uptake and Intracellular Drug Controlled Release of VESylated Gemcitabine Prodrug Nanocapsules. Colloids Surf. B Biointerfaces 2015, 128, 357–362. [Google Scholar] [CrossRef]
- Koga, Y.; Manabe, S.; Aihara, Y.; Sato, R.; Tsumura, R.; Iwafuji, H.; Furuya, F.; Fuchigami, H.; Fujiwara, Y.; Hisada, Y.; et al. Antitumor Effect of Antitissue Factor Antibody-MMAE Conjugate in Human Pancreatic Tumor Xenografts. Int. J. Cancer 2015, 137, 1457–1466. [Google Scholar] [CrossRef] [PubMed]
- Wang, Y.; Fan, W.; Dai, X.; Katragadda, U.; Mckinley, D.; Teng, Q.; Tan, C. Enhanced Tumor Delivery of Gemcitabine via PEG-DSPE/TPGS Mixed Micelles. Mol. Pharm. 2014, 11, 1140–1150. [Google Scholar] [CrossRef] [PubMed]
- Chitkara, D.; Mittal, A.; Behrman, S.W.; Kumar, N.; Mahato, R.I. Self-Assembling, Amphiphilic Polymer-Gemcitabine Conjugate Shows Enhanced Antitumor Efficacy against Human Pancreatic Adenocarcinoma. Bioconjug. Chem. 2013, 24, 1161–1173. [Google Scholar] [CrossRef] [PubMed]
- Ogbomo, S.M.; Shi, W.; Wagh, N.K.; Zhou, Z.; Brusnahan, S.K.; Garrison, J.C. 177Lu-Labeled HPMA Copolymers Utilizing Cathepsin B and S Cleavable Linkers: Synthesis, Characterization and Preliminary in Vivo Investigation in a Pancreatic Cancer Model. Nucl. Med. Biol. 2013, 40, 606–617. [Google Scholar] [CrossRef]
- von Burstin, J.; Eser, S.; Seidler, B.; Meining, A.; Bajbouj, M.; Mages, J.; Lang, R.; Kind, A.J.; Schnieke, A.E.; Schmid, R.M.; et al. Highly Sensitive Detection of Early-Stage Pancreatic Cancer by Multimodal near-Infrared Molecular Imaging in Living Mice. Int. J. Cancer 2008, 123, 2138–2147. [Google Scholar] [CrossRef] [PubMed]
- Han, H.; Valdepérez, D.; Jin, Q.; Yang, B.; Li, Z.; Wu, Y.; Pelaz, B.; Parak, W.J.; Ji, J. Dual Enzymatic Reaction-Assisted Gemcitabine Delivery Systems for Programmed Pancreatic Cancer Therapy. ACS Nano 2017, 11, 1281–1291. [Google Scholar] [CrossRef] [PubMed]
- Fan, W.; Zhang, W.; Jia, Y.; Brusnahan, S.K.; Garrison, J.C. Investigation into the Biological Impact of Block Size on Cathepsin S-Degradable HPMA Copolymers. Mol. Pharm. 2017, 14, 1405–1417. [Google Scholar] [CrossRef]
- Joubert, F.; Martin, L.; Perrier, S.; Pasparakis, G. Development of a Gemcitabine-Polymer Conjugate with Prolonged Cytotoxicity against a Pancreatic Cancer Cell Line. ACS Macro Lett. 2017, 6, 535–540. [Google Scholar] [CrossRef]
- Han, H.; Teng, W.; Chen, T.; Zhao, J.; Jin, Q.; Qin, Z.; Ji, J. A Cascade Enzymatic Reaction Activatable Gemcitabine Prodrug with an AIE-Based Intracellular Light-up Apoptotic Probe for in Situ Self-Therapeutic Monitoring. Chem. Commun. 2017, 53, 9214–9217. [Google Scholar] [CrossRef]
- Han, H.; Wang, J.; Chen, T.; Yin, L.; Jin, Q.; Ji, J. Enzyme-Sensitive Gemcitabine Conjugated Albumin Nanoparticles as a Versatile Theranostic Nanoplatform for Pancreatic Cancer Treatment. J. Colloid Interface Sci. 2017, 507, 217–224. [Google Scholar] [CrossRef]
- S, W.; D, C.; C, Z. A Cathepsin B-Sensitive Gemcitabine Prodrug for Enhanced Pancreatic Cancer Therapy. J. Pharm. Sci. 2024, 113. [Google Scholar] [CrossRef]
- Lee, G.Y.; Qian, W.P.; Wang, L.; Wang, Y.A.; Staley, C.A.; Satpathy, M.; Nie, S.; Mao, H.; Yang, L. Theranostic Nanoparticles with Controlled Release of Gemcitabine for Targeted Therapy and MRI of Pancreatic Cancer. ACS Nano 2013, 7, 2078–2089. [Google Scholar] [CrossRef]
- Pontious, C.; Kaul, S.; Hong, M.; Hart, P.A.; Krishna, S.G.; Lara, L.; Conwell, D.L.; Cruz-Monserrate, Z. Cathepsin E Expression and Activity: Role in the Detection and Treatment of Pancreatic Cancer. Pancreatol. Off. J. Int. Assoc. Pancreatol. IAP Al 2019, 19, 951–956. [Google Scholar] [CrossRef]
- Li, H.; Li, Y.; Cui, L.; Wang, B.; Cui, W.; Li, M.; Cheng, Y. Monitoring Pancreatic Carcinogenesis by the Molecular Imaging of Cathepsin E In Vivo Using Confocal Laser Endomicroscopy. PLoS ONE 2014, 9, e106566. [Google Scholar] [CrossRef]
- Abd-Elgaliel, W.R.; Cruz-Monserrate, Z.; Wang, H.; Logsdon, C.D.; Tung, C.-H. Pancreatic Cancer-Associated Cathepsin E as a Drug Activator. J. Control. Release Off. J. Control. Release Soc. 2013, 167, 221–227. [Google Scholar] [CrossRef]
- Lopez, S.; Hallali, N.; Lalatonne, Y.; Hillion, A.; Antunes, J.C.; Serhan, N.; Clerc, P.; Fourmy, D.; Motte, L.; Carrey, J.; et al. Magneto-Mechanical Destruction of Cancer-Associated Fibroblasts Using Ultra-Small Iron Oxide Nanoparticles and Low Frequency Rotating Magnetic Fields. Nanoscale Adv. 4, 421–436. [CrossRef]
- Li, T.; Chen, D.; Liu, H.; Tao, Y.; He, X.; Zang, S.; Li, J.; Zhang, L.; Li, M.; Liu, J.; et al. Spatially Targeting and Regulating Tumor-Associated Macrophages Using a Raspberry-like Micellar System Sensitizes Pancreatic Cancer Chemoimmunotherapy. Nanoscale 2022, 14, 13098–13112. [Google Scholar] [CrossRef] [PubMed]
- Kwok, H.F.; Buick, R.J.; Kuehn, D.; Gormley, J.A.; Doherty, D.; Jaquin, T.J.; McClurg, A.; Ward, C.; Byrne, T.; Jaworski, J.; et al. Antibody Targeting of Cathepsin S Induces Antibody-Dependent Cellular Cytotoxicity. Mol. Cancer 2011, 10, 147. [Google Scholar] [CrossRef]
- Deng, Y.; Yan, K.; Xu, L.; Liu, X.; Zhao, F.; Wang, S.; Zhang, F.; Liang, G.; Wang, R. “Turn-on” NIR-II Fluorescence of a Dually Quenched Probe for Sensitive Imaging of Cathepsin B in Vivo. Chem. Commun. 2025, 61, 544–547. [Google Scholar] [CrossRef] [PubMed]
- Husi, H.; Fernandes, M.; Skipworth, R.J.; Miller, J.; Cronshaw, A.D.; Fearon, K.C.H.; Ross, J.A. Identification of Diagnostic Upper Gastrointestinal Cancer Tissue Type-Specific Urinary Biomarkers. Biomed. Rep. 2019, 10, 165–174. [Google Scholar] [CrossRef]
- Tumminello, F.M.; Leto, G.; Pizzolanti, G.; Candiloro, V.; Crescimanno, M.; Crosta, L.; Flandina, C.; Montalto, G.; Soresi, M.; Carroccio, A.; et al. Cathepsin D, B and L Circulating Levels as Prognostic Markers of Malignant Progression. Anticancer Res. 1996, 16, 2315–2319. [Google Scholar]
- Roger, M.; Martínez, J.; Aparicio, J.-R.; Peiró, G.; Ruiz, F.; Compañy, L.; Casellas, J.-A.; Roger, M.; Martínez, J.; Aparicio, J.-R.; et al. EUS-FNA Cytological Material from Pancreatic Lesions: The Expression of Cathepsins and Its Predictive Value of Malignancy. Rev. Esp. Enfermedades Dig. 2018, 110, 446–450. [Google Scholar] [CrossRef]
- Cruz-Monserrate, Z.; Abd-Elgaliel, W.R.; Ji, B.; Deng, D.; Arumugan, T.; Tung, C.H.; Logsdon, C.D. Abstract 5324: Cathepsin E Expression: A New Potential Target for Pancreatic Tumor Imaging. Cancer Res. 2011, 71. [Google Scholar] [CrossRef]
- Ivry, S.L.; Sharib, J.M.; Dominguez, D.A.; Roy, N.; Hatcher, S.E.; Yip-Schneider, M.T.; Schmidt, C.M.; Brand, R.E.; Park, W.G.; Hebrok, M.; et al. Global Protease Activity Profiling Provides Differential Diagnosis of Pancreatic Cysts. Clin. Cancer Res. 2017, 23, 4865–4874. [Google Scholar] [CrossRef]
- Pescatore, R.; Milliken, N.; King, T.; Josey, D.; Palma, N.A.; Ford, L. Analytical Validation of a Serum Biomarker Signature for Detection of Early-Stage Pancreatic Ductal Adenocarcinoma. Diagnostics 2025, 15, 3177. [Google Scholar] [CrossRef] [PubMed]
- Yitzhak, Y.; Gaikwad, H.; Weiss-Sadan, T.; Merquiol, E.; Turk, B.; Blum, G. Improved Cathepsin Probes for Sensitive Molecular Imaging. Molecules 2022, 27, 842. [Google Scholar] [CrossRef] [PubMed]
| Sample type | Analyte(s) | Intended application | Key finding | Ref. |
|---|---|---|---|---|
| Urine | Cathepsin B (among candidate urinary biomarkers) | Non-invasive biomarker discovery | Urinary biomarker profiling identified tissue–type–specific candidates for upper GI cancers, including markers relevant to pancreatic cancer. | [179] |
| Serum / liquid biopsy | Enzymatic activity panel (arginase, MMP-1/3/9, cathepsins B & E, uPA, neutrophil elastase) | Early detection/ screening concept | Multiplex enzymatic “signature” proposed for potential early detection of pancreatic cancers in liquid biopsies. | [97] |
| Serum (circulating levels) | Cathepsins D, B, L | Prognosis / malignant progression | Circulating cathepsins are reported as markers associated with malignant progression. | [180] |
| Plasma | Cathepsin L | Prognosis | Plasma CTSL evaluated as a potential prognostic marker in pancreatic cancer. | [93] |
| Pancreatic juice | Cathepsin E | Diagnostics | CTSE levels are significantly elevated in pancreatic juice from PDAC patients compared to benign pancreatic disease; high diagnostic specificity | [119] |
| Serum (circulating levels) | Cathepsin B | Prognosis | Increased CTSB levels correlate with tumor burden and invasive phenotype; associated with aggressive disease biology | [94] |
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2026 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).




