Submitted:
23 April 2026
Posted:
23 April 2026
You are already at the latest version
Abstract
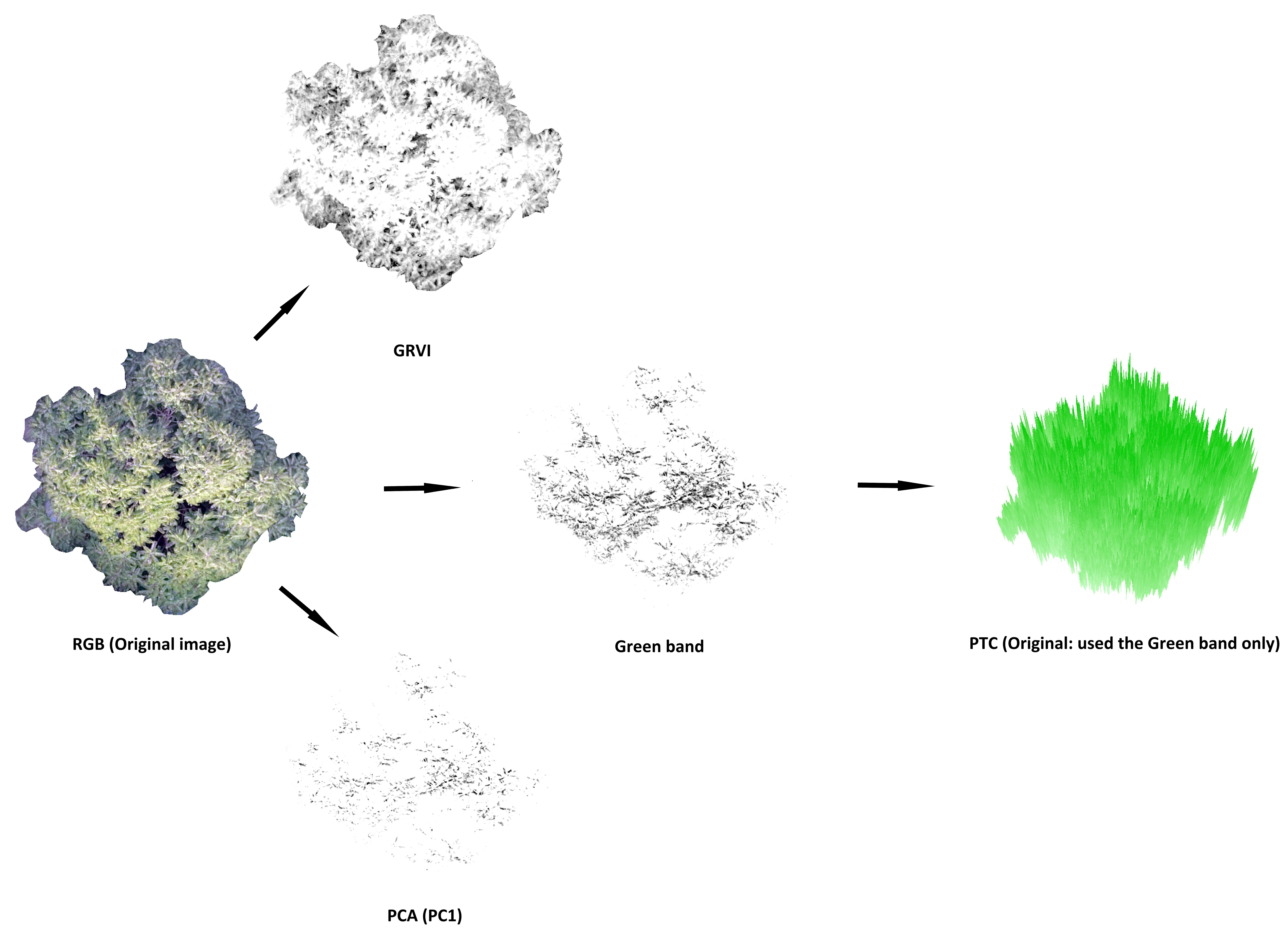
Keywords:
1. Introduction
2. Material and methods
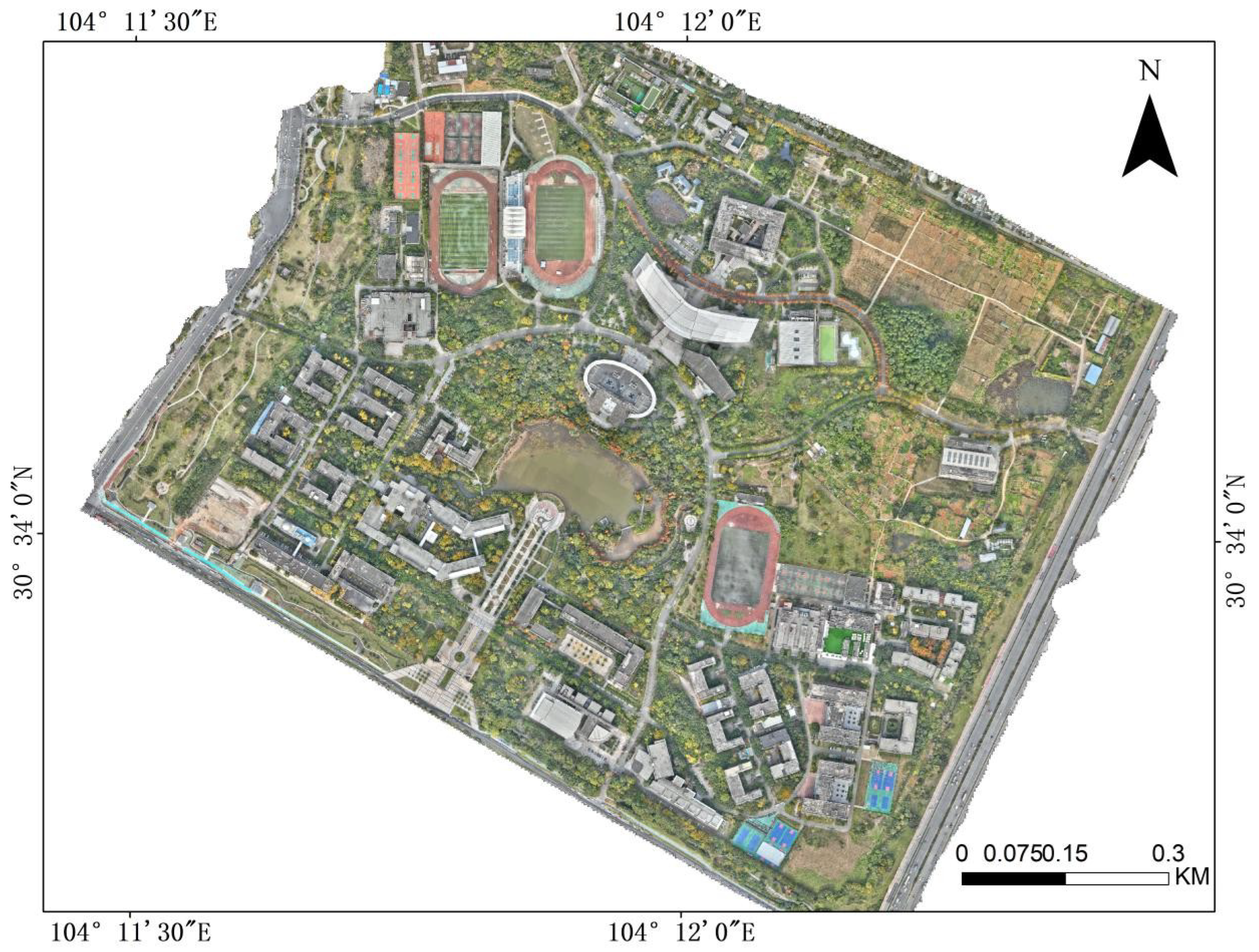
2.1. The study area and data description
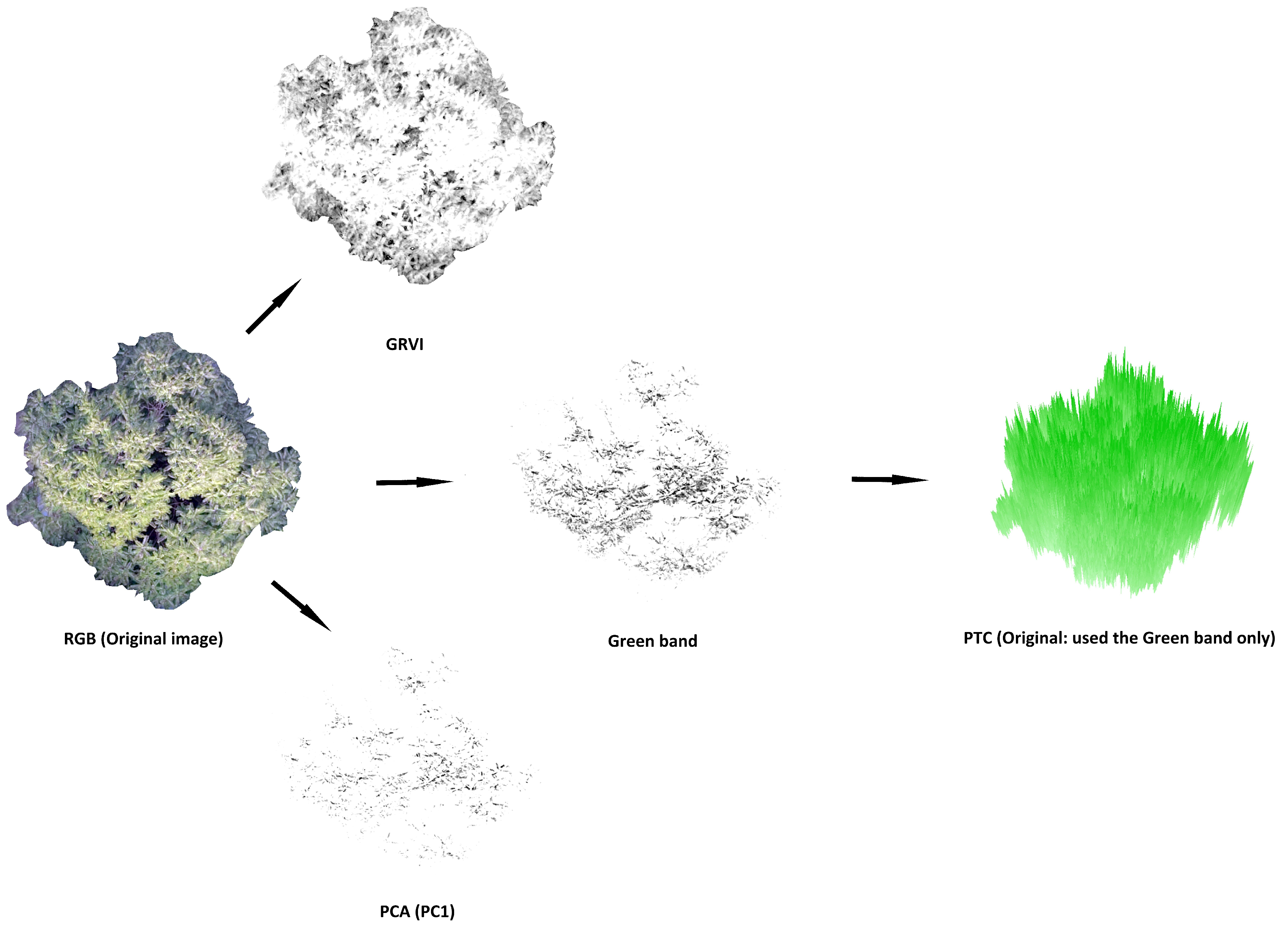
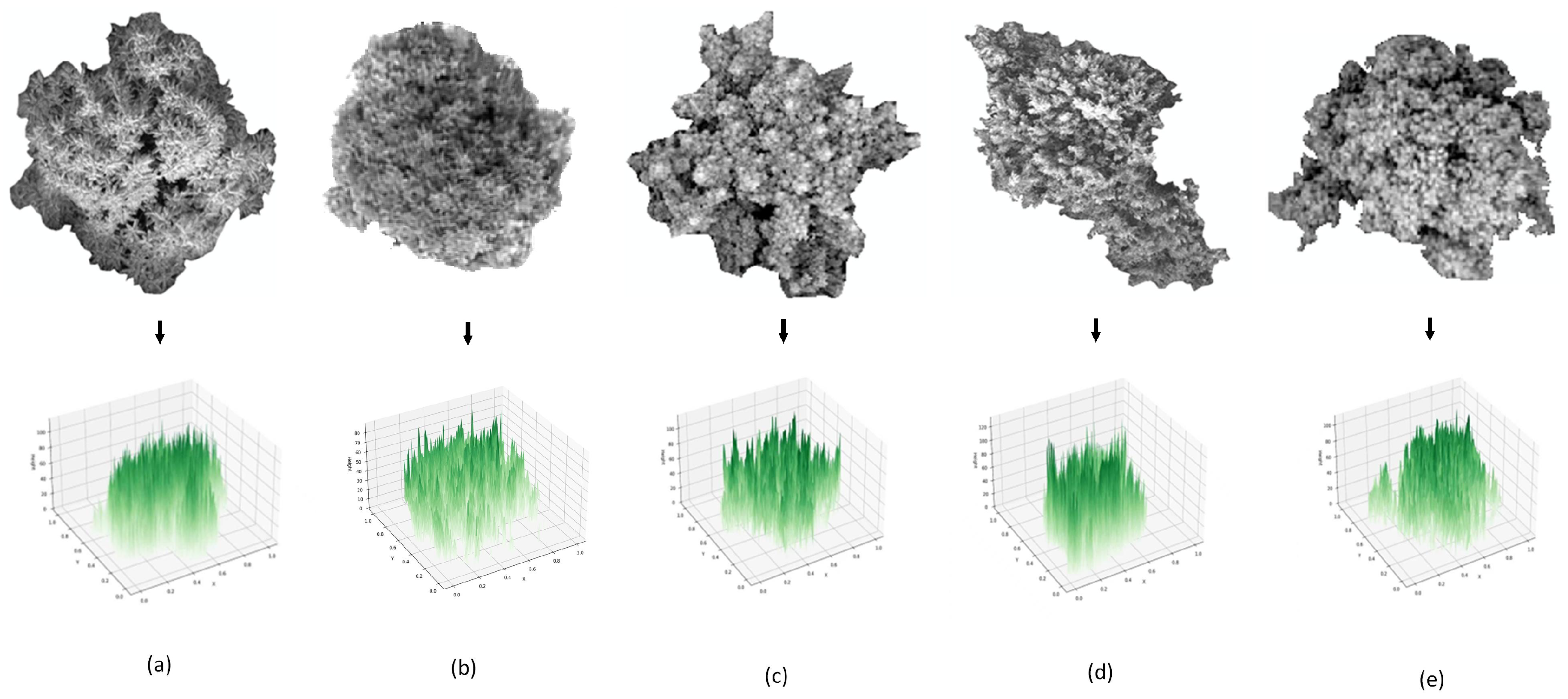
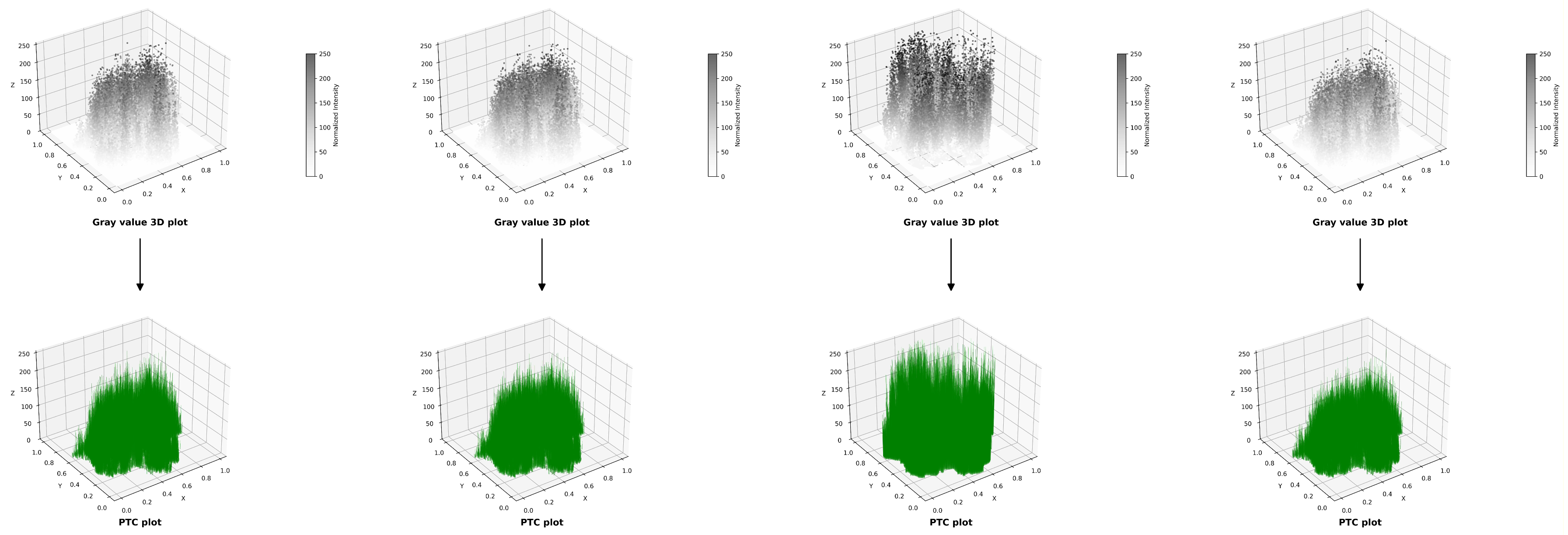
2.2. Methods: PTC, GRVI and PCA
3. Results
| Model | Data | Tree Species | Precision | Recall | F1-score | IoU |
|---|---|---|---|---|---|---|
| RF | RGB | Tree of Heaven | 0.5700 | 0.5938 | 0.5819 | 0.4199 |
| Osmanthus Tree | 0.6013 | 0.6636 | 0.6234 | 0.4646 | ||
| Chinese Banyan | 0.7211 | 0.7000 | 0.7100 | 0.5111 | ||
| Big-leaved Fig | 0.5824 | 0.6667 | 0.6243 | 0.4332 | ||
| Campphor Tree | 0.5882 | 0.4909 | 0.5441 | 0.3636 | ||
| Green | Tree of Heaven | 0.6296 | 0.6084 | 0.6169 | 0.4498 | |
| Osmanthus Tree | 0.6340 | 0.6501 | 0.6404 | 0.4749 | ||
| Chinese Banyan | 0.6907 | 0.6834 | 0.6854 | 0.5283 | ||
| Big-leaved Fig | 0.6239 | 0.6739 | 0.6459 | 0.4810 | ||
| Campphor Tree | 0.5387 | 0.5341 | 0.5347 | 0.3739 | ||
| GRVI | Tree of Heaven | 0.8283 | 0.8179 | 0.8176 | 0.7127 | |
| Osmanthus Tree | 0.7279 | 0.7595 | 0.7342 | 0.5896 | ||
| Chinese Banyan | 0.7271 | 0.6928 | 0.7041 | 0.5528 | ||
| Big-leaved Fig | 0.6292 | 0.6928 | 0.6509 | 0.4905 | ||
| Campphor Tree | 0.6699 | 0.5552 | 0.5961 | 0.4448 | ||
| PCA1 | Tree of Heaven | 0.6014 | 0.6000 | 0.6003 | 0.4312 | |
| Osmanthus Tree | 0.6510 | 0.6375 | 0.6438 | 0.4803 | ||
| Chinese Banyan | 0.6374 | 0.6708 | 0.6529 | 0.4915 | ||
| Big-leaved Fig | 0.5844 | 0.5970 | 0.5902 | 0.4206 | ||
| Campphor Tree | 0.5889 | 0.5483 | 0.5669 | 0.3975 | ||
| ResNet50 | RGB | Tree of Heaven | 0.9091 | 0.8333 | 0.8696 | 0.7692 |
| Osmanthus Tree | 0.9231 | 0.8000 | 0.8571 | 0.7500 | ||
| Chinese Banyan | 0.9333 | 0.9333 | 0.9333 | 0.8750 | ||
| Big-leaved Fig | 0.7917 | 0.9048 | 0.8444 | 0.7308 | ||
| Campphor Tree | 0.7692 | 0.8696 | 0.8163 | 0.6897 | ||
| Green | Tree of Heaven | 0.9130 | 0.8750 | 0.8936 | 0.8077 | |
| Osmanthus Tree | 0.8846 | 0.7667 | 0.8214 | 0.6970 | ||
| Chinese Banyan | 0.9000 | 0.9000 | 0.9000 | 0.8182 | ||
| Big-leaved Fig | 0.8000 | 0.9524 | 0.8696 | 0.7692 | ||
| Campphor Tree | 0.7500 | 0.7826 | 0.7660 | 0.6207 | ||
| GRVI | Tree of Heaven | 0.9565 | 0.9167 | 0.9362 | 0.8800 | |
| Osmanthus Tree | 0.6970 | 0.7667 | 0.7302 | 0.5750 | ||
| Chinese Banyan | 0.9231 | 0.8000 | 0.8471 | 0.7500 | ||
| Big-leaved Fig | 0.7619 | 0.7619 | 0.7619 | 0.6154 | ||
| Campphor Tree | 0.4800 | 0.5217 | 0.5000 | 0.3333 | ||
| PCA1 | Tree of Heaven | 0.8947 | 0.7083 | 0.7907 | 0.6538 | |
| Osmanthus Tree | 0.9524 | 0.6667 | 0.7843 | 0.6452 | ||
| Chinese Banyan | 0.8486 | 0.9333 | 0.8889 | 0.8000 | ||
| Big-leaved Fig | 0.6774 | 1.0000 | 0.8077 | 0.6207 | ||
| Campphor Tree | 0.7500 | 0.7826 | 0.7660 | 0.6774 | ||
| YOLOv10 | RGB | Tree of Heaven | 0.8119 | 0.8056 | 0.7936 | 0.6631 |
| Osmanthus Tree | 0.7971 | 0.6968 | 0.7068 | 0.5833 | ||
| Chinese Banyan | 0.7941 | 0.6896 | 0.7381 | 0.5966 | ||
| Big-leaved Fig | 0.7046 | 0.7190 | 0.7049 | 0.6107 | ||
| Campphor Tree | 0.6657 | 0.5000 | 0.5542 | 0.3923 | ||
| Green | Tree of Heaven | 0.8360 | 0.7986 | 0.7962 | 0.6675 | |
| Osmanthus Tree | 0.8656 | 0.6783 | 0.6816 | 0.5736 | ||
| Chinese Banyan | 0.8444 | 0.7673 | 0.8031 | 0.6781 | ||
| Big-leaved Fig | 0.7366 | 0.6440 | 0.6711 | 0.5536 | ||
| Campphor Tree | 0.7258 | 0.6556 | 0.6610 | 0.5276 | ||
| GRVI | Tree of Heaven | 0.8537 | 0.7461 | 0.7927 | 0.6681 | |
| Osmanthus Tree | 0.8644 | 0.7821 | 0.8171 | 0.6968 | ||
| Chinese Banyan | 0.7576 | 0.7408 | 0.7475 | 0.6033 | ||
| Big-leaved Fig | 0.7404 | 0.6765 | 0.6966 | 0.5641 | ||
| Campphor Tree | 0.6820 | 0.7582 | 0.7114 | 0.5827 | ||
| PCA1 | Tree of Heaven | 0.8731 | 0.8334 | 0.8469 | 0.7346 | |
| Osmanthus Tree | 0.6243 | 0.6482 | 0.6340 | 0.5493 | ||
| Chinese Banyan | 0.7972 | 0.7069 | 0.7491 | 0.6245 | ||
| Big-leaved Fig | 0.7365 | 0.7965 | 0.7618 | 0.6318 | ||
| Campphor Tree | 0.6304 | 0.6130 | 0.6161 | 0.4638 |
| Input Data | Random Forest | ResNet50 | YOLOv10 |
|---|---|---|---|
| RGB-PTC | |||
| Green-PTC | |||
| GRVI-PTC | |||
| PCA1-PTC |
| Model | Input | Data Variation | time (seconds) |
|---|---|---|---|
| RF | Directly | RGB | 76.90 |
| Green | 78.07 | ||
| GRVI | 79.79 | ||
| PCA1 | 77.29 | ||
| PTC | 75.28 | ||
| PTCs | RGB | 81.15 | |
| Green | 75.28 | ||
| GRVI | 83.58 | ||
| PCA1 | 77.14 | ||
| ResNet50 | Directly | RGB | 45.79 |
| Green | 23.55 | ||
| GRVI | 29.49 | ||
| PCA1 | 43.05 | ||
| PTC | 45.49 | ||
| PTCs | RGB | 29.48 | |
| Green | 45.49 | ||
| GRVI | 27.31 | ||
| PCA1 | 36.75 | ||
| YOLOv10 | Directly | RGB | 409.53 |
| Green | 412.97 | ||
| GRVI | 416.11 | ||
| PCA1 | 406.36 | ||
| PTC | 413.26 | ||
| PTCs | RGB | 416.58 | |
| Green | 413.26 | ||
| GRVI | 412.83 | ||
| PCA1 | 422.48 |
4. Discussion
5. Conclusion
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Conflicts of Interest
Abbreviations
| PTC | Pseudo tree crown |
| ITS | Individual tree species |
| GRVI | Green Red Vegetation Index |
| PCA | Principal Component Analysis |
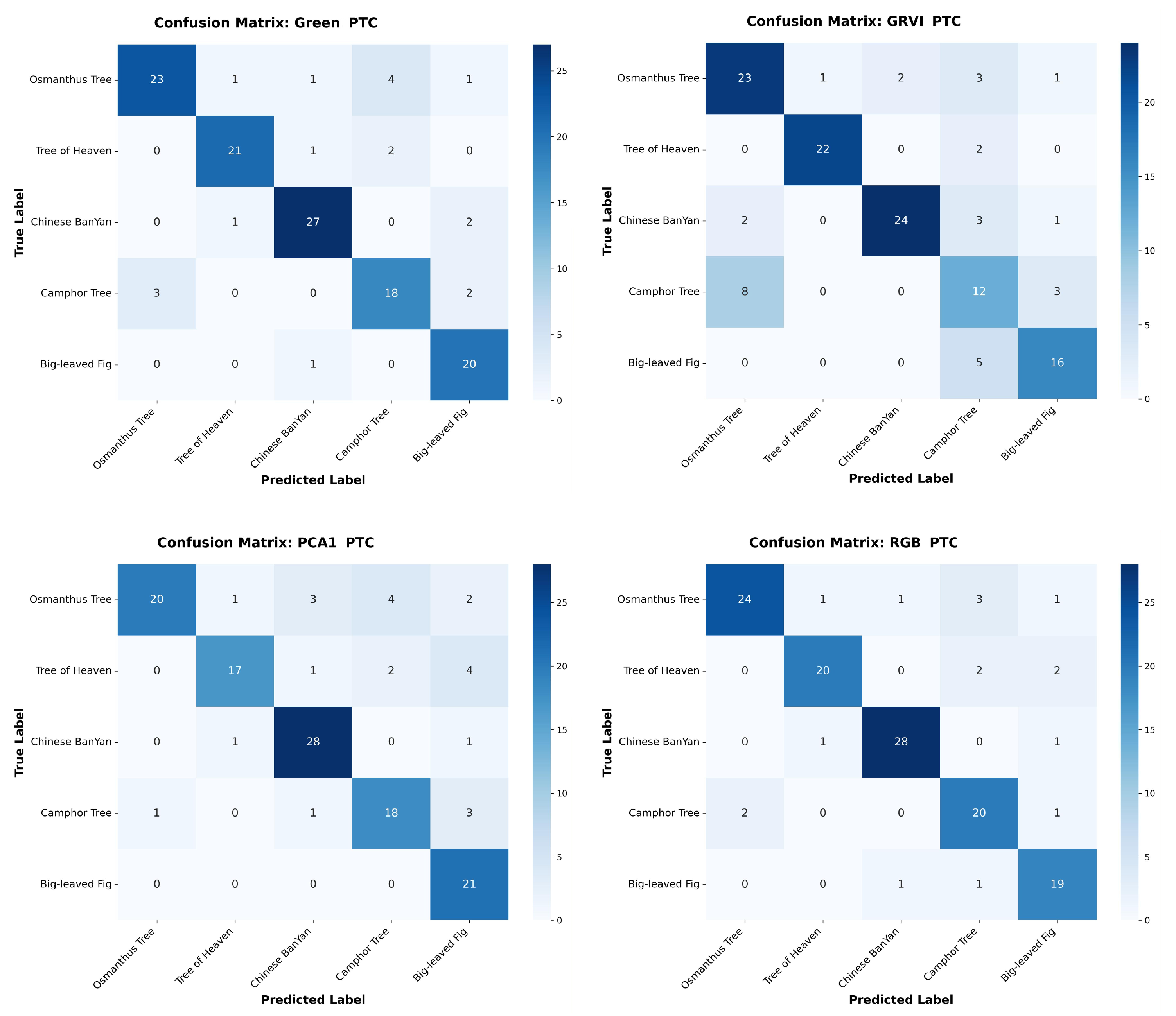
Appendix A The confusion matrices of species classification results of different PTC transformations using RF and YOLOv10
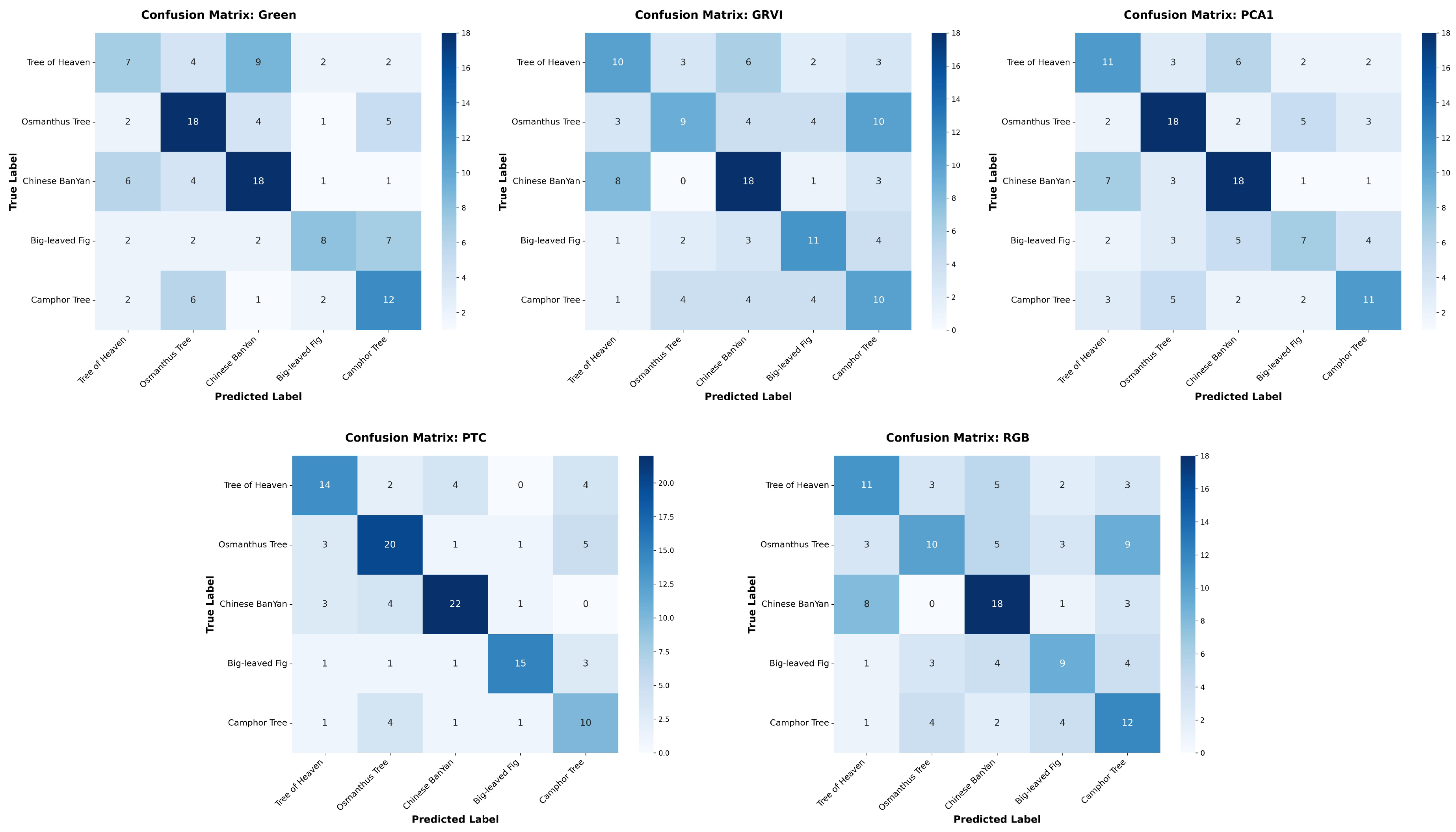
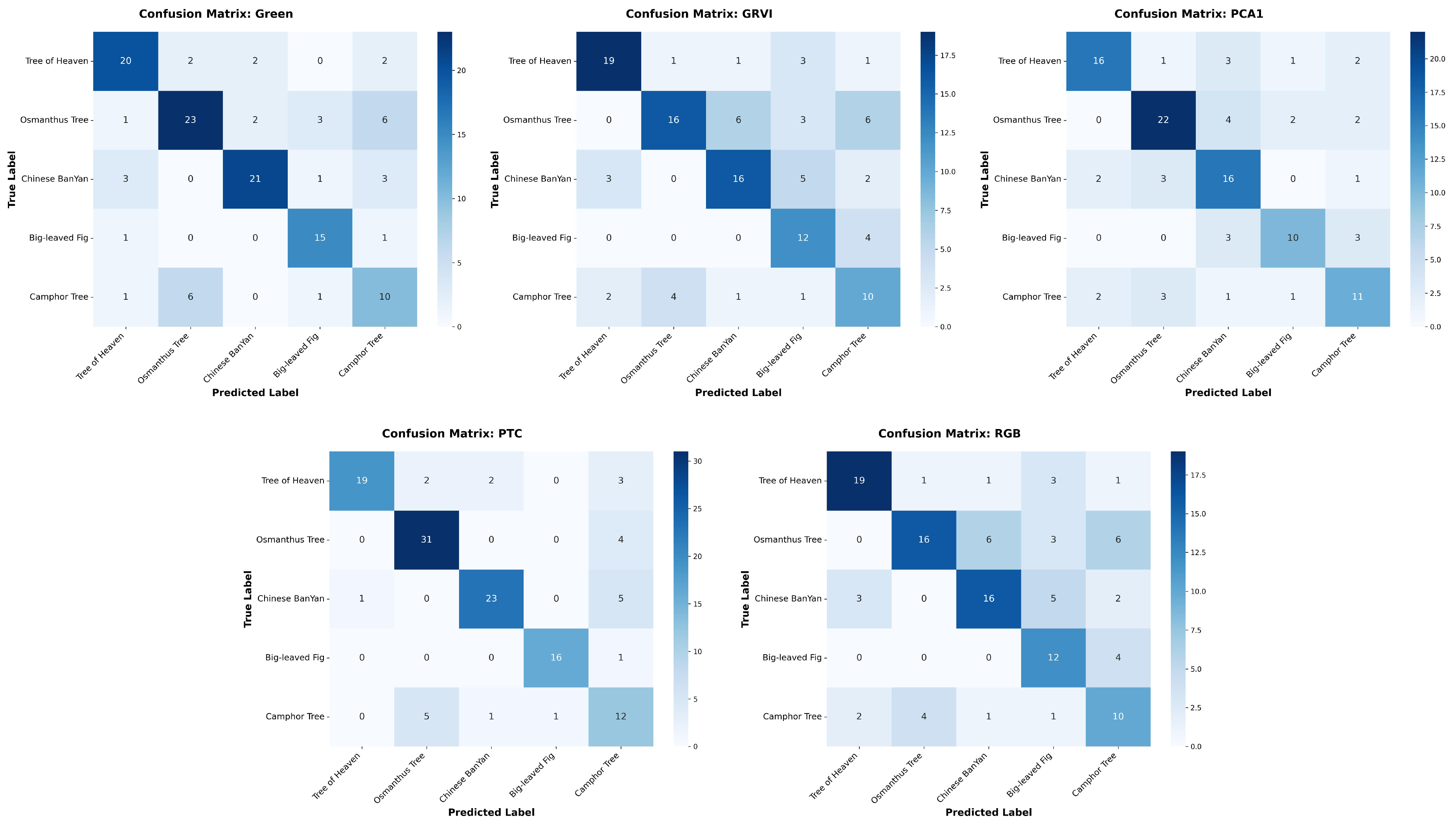
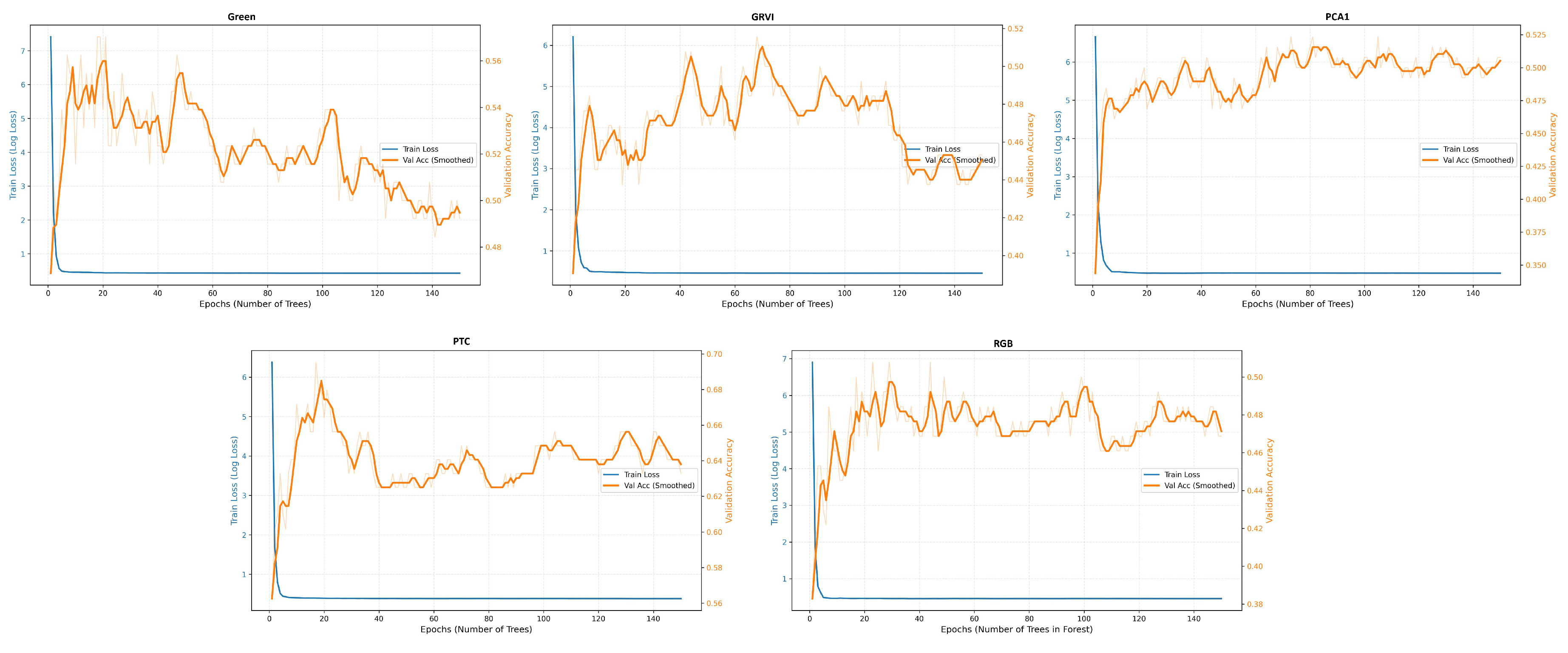
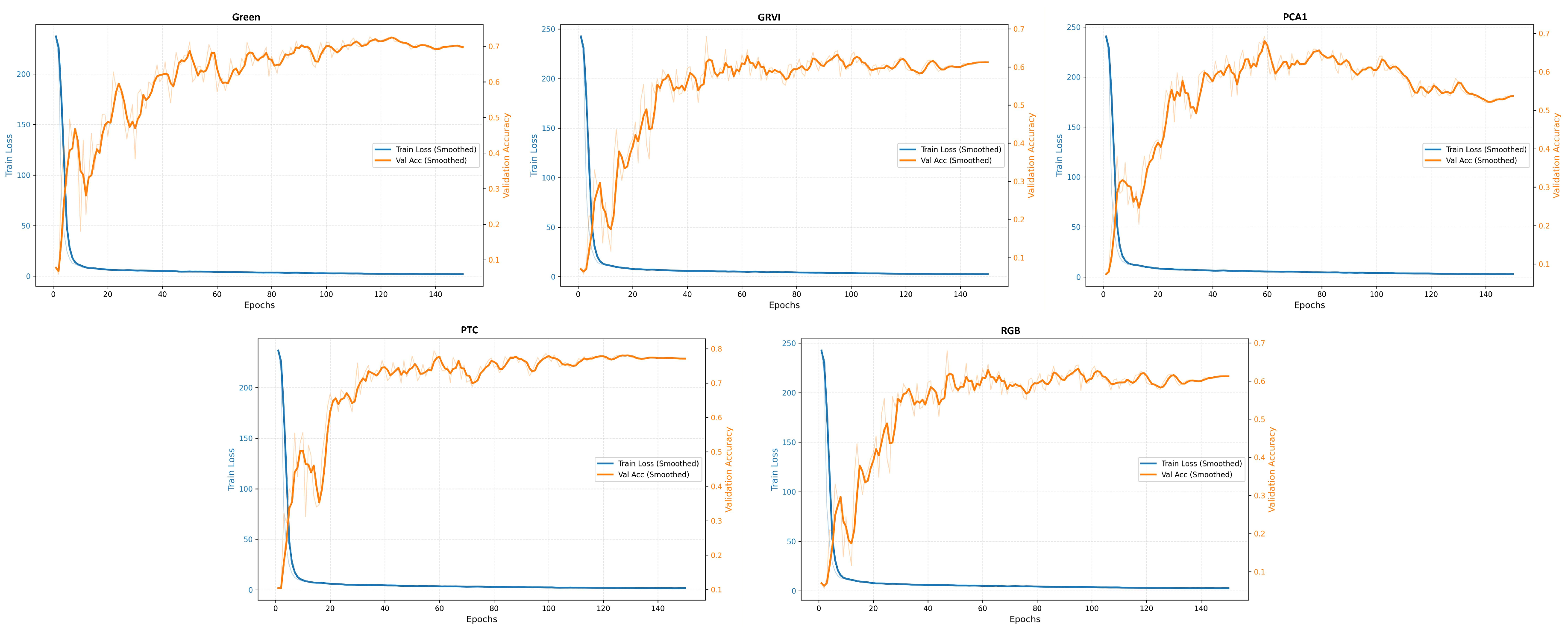
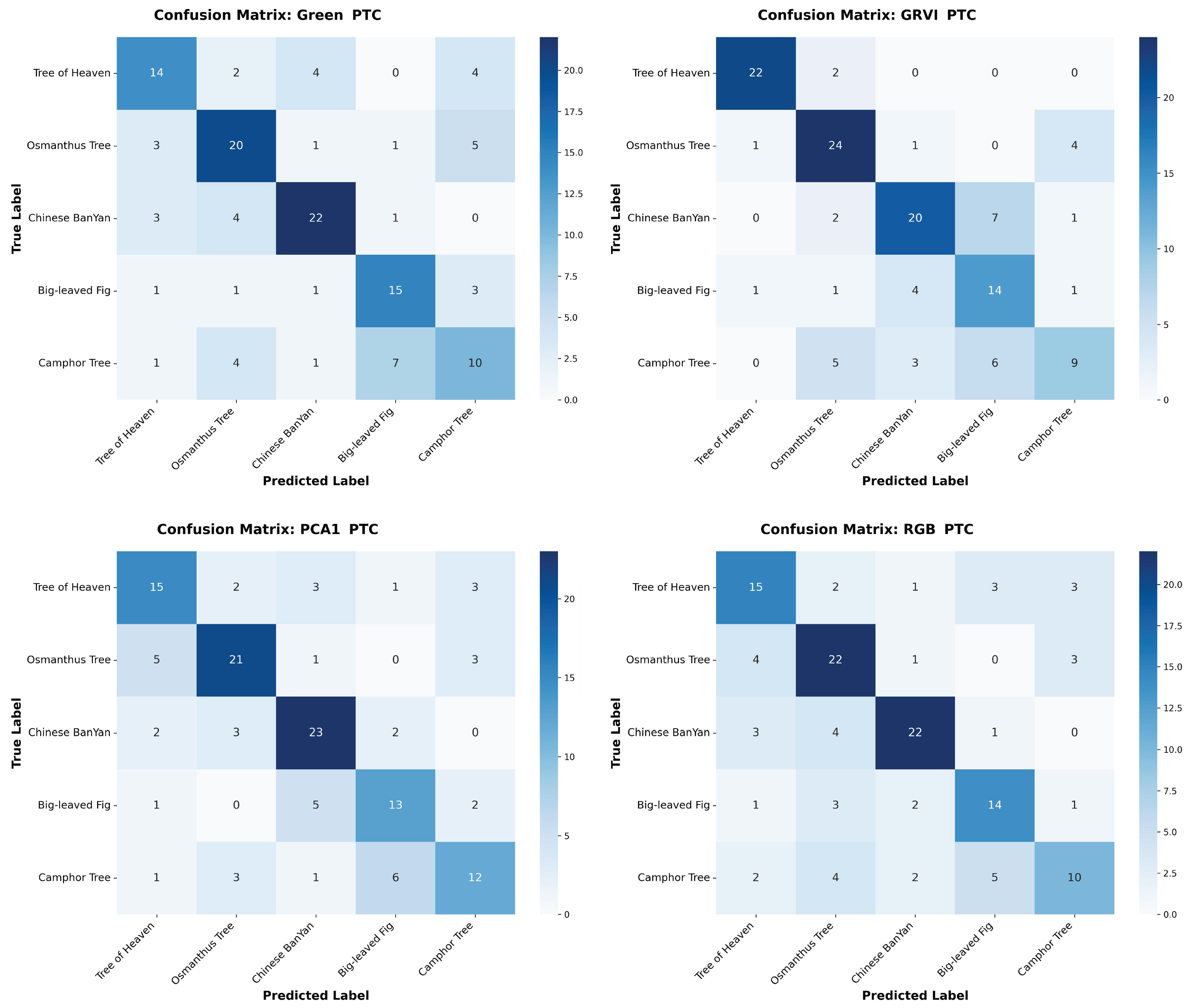
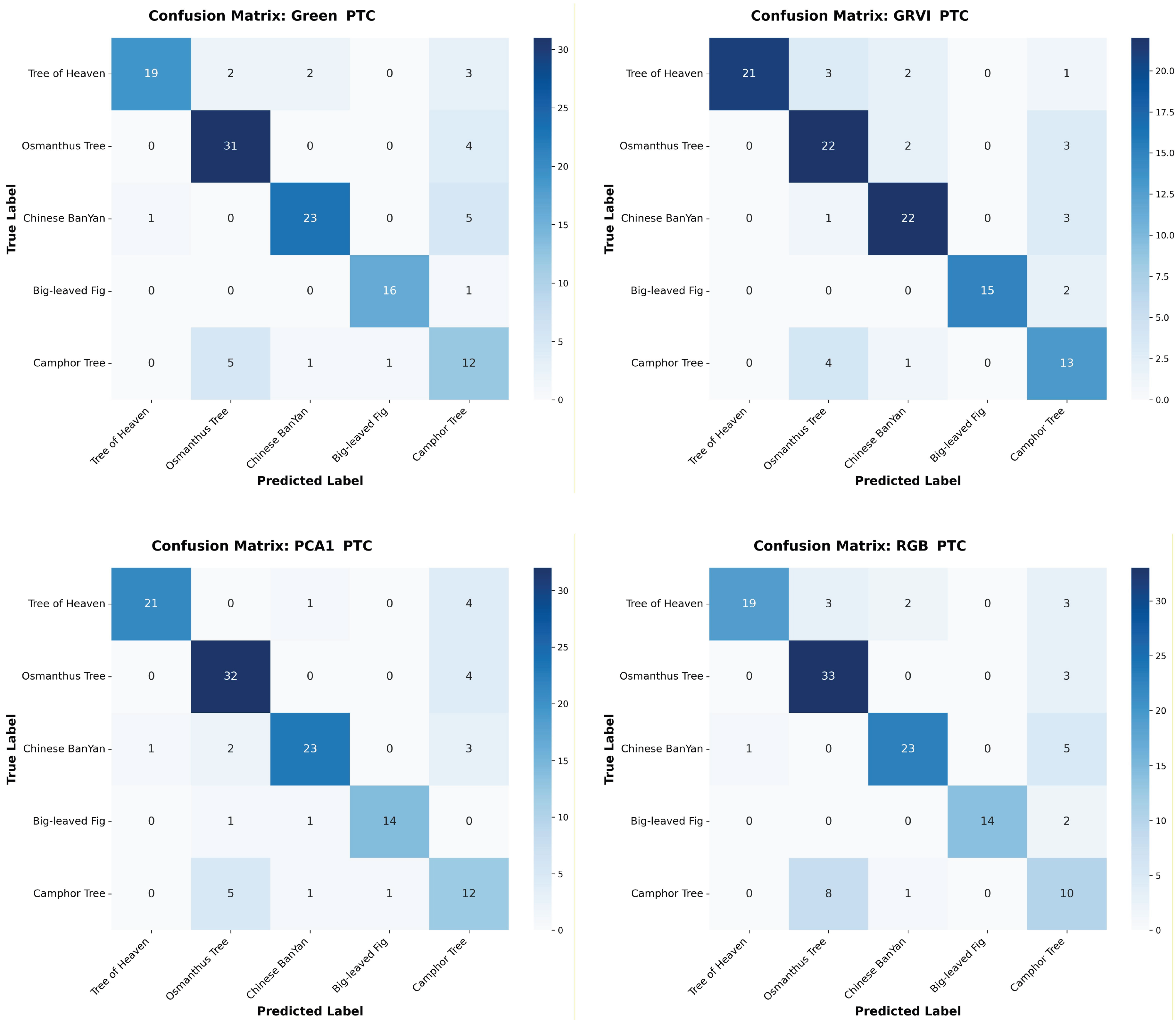
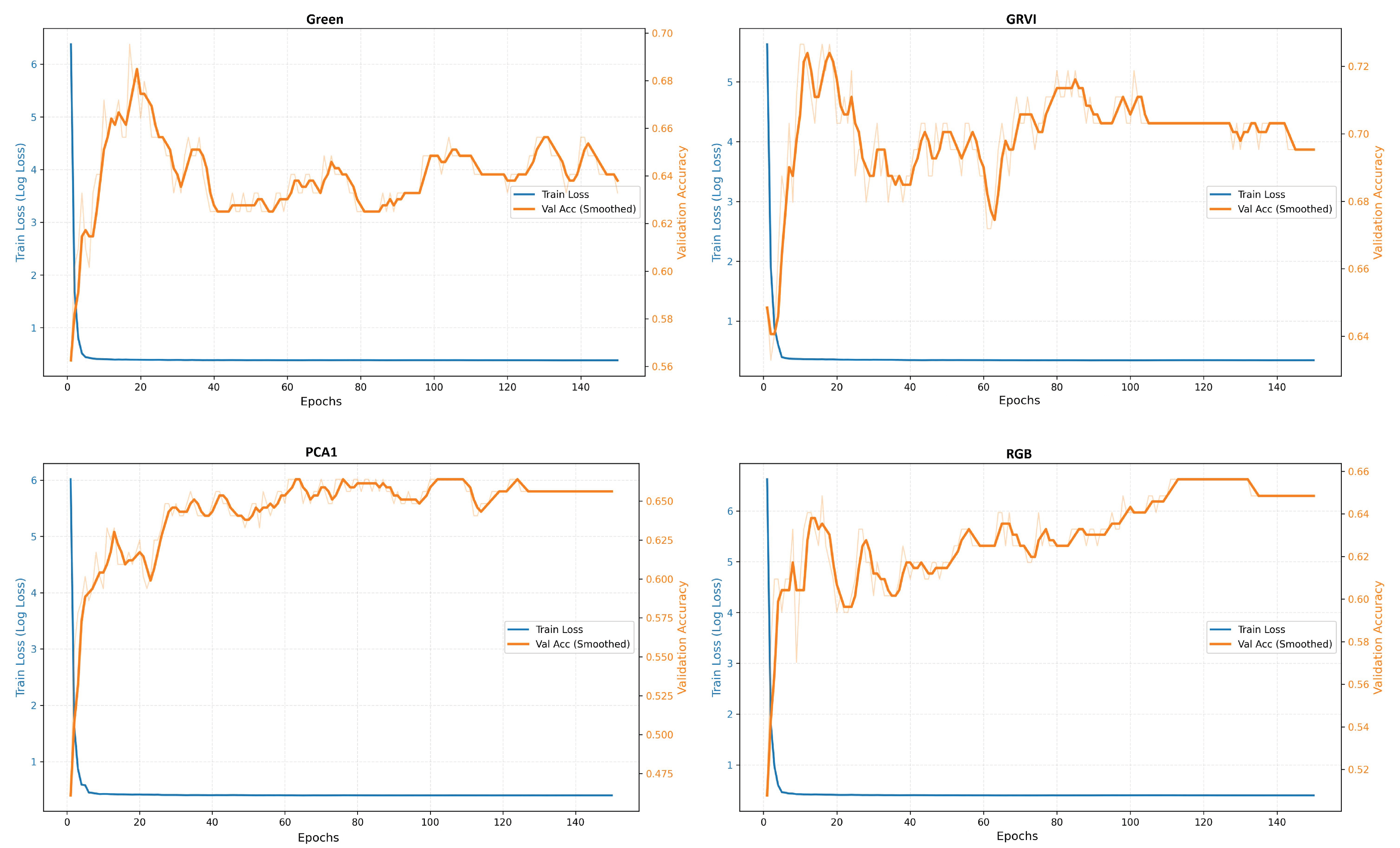
References
- Wang, K.; Wang, T.; Liu, X. A Review: Individual Tree Species Classification Using Integrated Airborne LiDAR and Optical Imagery with a Focus on the Urban Environment. Forests 2018, 10, 1. [Google Scholar] [CrossRef]
- Guo, X.; Li, H.; Jing, L.; Wang, P. Individual Tree Species Classification Based on Convolutional Neural Networks and Multitemporal High-Resolution Remote Sensing Images. Sensors 2022, 22. [Google Scholar] [CrossRef] [PubMed]
- Wang, L.; Lu, D.; Xu, L.; Robinson, D.T.; Tan, W.; Xie, Q.; Guan, H.; Chapman, M.A.; Li, J. Individual tree species classification using low-density airborne multispectral LiDAR data via attribute-aware cross-branch transformer. Remote Sensing of Environment 2024, 315, 114456. [Google Scholar] [CrossRef]
- Lei, Z.; Li, H.; Zhao, J.; Jing, L.; Tang, Y.; Wang, H. Individual Tree Species Classification Based on a Hierarchical Convolutional Neural Network and Multitemporal Google Earth Images. Remote Sensing 2022, 14. [Google Scholar] [CrossRef]
- Natesan, S.; Armenakis, C.; Vepakomma, U. Individual tree species identification using Dense Convolutional Network (DenseNet) on multitemporal RGB images from UAV. Journal of Unmanned Vehicle Systems 2020, 8, 310–333. [Google Scholar] [CrossRef]
- Cetin, Z.; Yastikli, N. The Use of Machine Learning Algorithms in Urban Tree Species Classification. ISPRS International Journal of Geo-Information 2022, 11, 226. [Google Scholar] [CrossRef]
- Bensi, M.E.; Esquivel, R.A. Unraveling the Significance of the Classification Tree Algorithm in Machine Learning: A Literature Review. Journal of Theoretical and Applied Sciences 2023, 1, 604–611. [Google Scholar] [CrossRef]
- He, X.; Han, X.; Chen, Y.; Huang, L. A Light-Weighted Fusion Vision Mamba for Multimodal Remote Sensing Data Classification. IEEE Journal of Selected Topics in Applied Earth Observations and Remote Sensing 2025, 18, 21532–21548. [Google Scholar] [CrossRef]
- Zhong, L.; Dai, Z.; Fang, P.; Cao, Y.; Wang, L. A Review: Tree Species Classification Based on Remote Sensing Data and Classic Deep Learning-Based Methods. Forest 2024, 15, 852. [Google Scholar] [CrossRef]
- Southworth, J.; Smith, A.C.; Safaei, M.; Rahaman, M.; Alruzuq, A.; Tefera, B.B.; Muir, C.S.; Herrero, H.V. Machine learning versus deep learning in land system science: a decision-making framework for effective land classification. Frontiers in Remote Sensing 2024, 5–2024. [Google Scholar] [CrossRef]
- Abreu-Dias, R.; Juan, M.S.G.; Fernando, M.R.; Luis, M.S. Advances in the Automated Identification of Individual Tree Species: A Systematic Review of Drone- and AI-Based Methods in Forest Environments. Technologies 2025, 13, 187. [Google Scholar] [CrossRef]
- Wang, Y.; Huang, H.; State, R. Cross Domain Early Crop Mapping Using CropSTGAN. IEEE Access 2024, 12, 130800–130815. [Google Scholar] [CrossRef]
- Wang, Y.; Huang, H.; State, R. Cross Domain Early Crop Mapping with Label Spaces Discrepancies using MultiCropGAN. ISPRS Annals of the Photogrammetry, Remote Sensing and Spatial Information Sciences 2024, X-1-2024, 241–248. [Google Scholar] [CrossRef]
- Xu, Z.; Wang, T.; Skidmore, A.K.; Lamprey, R. A review of deep learning techniques for detecting animals in aerial and satellite images. International Journal of Applied Earth Observation and Geoinformation 2024, 128, 103732. [Google Scholar] [CrossRef]
- Wang, W.; Zhao, C.; Wu, Y. Spatial weighting — An effective incorporation of geological expertise into deep learning models. Geochemistry 2024, 84, 126212. [Google Scholar] [CrossRef]
- Xiong, Q.; Zhang, X.; Shen, J. A prior embedding-driven architecture for long distance blind iris recognition. Biomedical Signal Processing and Control 2025, 109, 108048. [Google Scholar] [CrossRef]
- Ben Khalifa, M.A.; El Koundi, M.; Farah, I.R. Pushing boundaries in remote sensing: A comprehensive review of deep learning for spatial super-resolution. Remote Sensing Applications: Society and Environment 2025, 40, 101809. [Google Scholar] [CrossRef]
- Fan, W.; Tian, J.; Troles, J.; Döllerer, M.; Kindu, M.; Knoke, T. Comparing Deep Learning and MCWST Approaches for Individual Tree Crown Segmentation. ISPRS Ann. Photogramm. Remote Sens. Spatial Inf. Sci. X-1-2024 2024, 67–73. [CrossRef]
- Burmeister, J.M.; Richter, R.; Reder, S.; Mund, J.P.; Döllner, J. Tree Instance Segmentation in Urban 3D Point Clouds Using a Coarse-to-Fine Algorithm Based on Semantic Segmentation. ISPRS Ann. Photogramm. Remote Sens. Spatial Inf. Sci. X-4/W5-2024 2024, 79–86. [CrossRef]
- Fu, Y.; Niu, Y.; Wang, L.; Li, W. Individual-Tree Segmentation from UAV–LiDAR Data Using a Region-Growing Segmentation and Supervoxel-Weighted Fuzzy Clustering Approach. Remote Sensing 2024, 16, 608. [Google Scholar] [CrossRef]
- D’Amico, G.; Nilsson, M.; Axelsson, A.; Chirici, G. Data homogeneity impact in tree species classification based on Sentinel-2 multitemporal data case study in central Sweden. International Journal of Remote Sensing 2024, 45, 5050–5075. [Google Scholar] [CrossRef]
- Zhang, H.; Liu, B.; Yang, B.; Guo, J.; Hu, Z.; Zhang, M.; Yang, Z.; Zhang, J. Efficient tree species classification using machine and deep learning algorithms based on UAV-LiDAR data in North China. Frontiers in Forests and Global Change 2025, 8–2025. [Google Scholar] [CrossRef]
- Marconi, S.; Weinstein, B.G.; Zou, S.; Bohlman, S.A.; Zare, A.; Singh, A.; Stewart, D.; Harmon, I.; Steinkraus, A.; White, E.P. Continental-scale hyperspectral tree species classification in the United States National Ecological Observatory Network. Remote Sensing of Environment 2022, 282, 113264. [Google Scholar] [CrossRef]
- Zakrzewska, A.; Kopeć, D.; Krajewski, K.; Charyton, J. Canopy temperatures of selected tree species growing in the forest and outside the forest using aerial thermal infrared (3.6–4.9 µm) data. European Journal of Remote Sensing 2022, 55, 313–325. [Google Scholar] [CrossRef]
- Blickensdörfer, L.; Oehmichen, K.; Pflugmacher, D.; Kleinschmit, B.; Hostert, P. National tree species mapping using Sentinel-1/2 time series and German National Forest Inventory data. Remote Sensing of Environment 2024, 304, 114069. [Google Scholar] [CrossRef]
- Qin, T.; Zhao, Q. Multi-branch and multi-label tree species classification using deep learning for UAV aerial photography and Sentinel remote sensing images. Sci Rep 2025, 15. [Google Scholar] [CrossRef]
- Heinzel, J.; Koch, B. Investigating multiple data sources for tree species classification in temperate forest and use for single tree delineation. International Journal of Applied Earth Observation and Geoinformation 2012, 18, 101–110. [Google Scholar] [CrossRef]
- Wan, H.; Tang, Y.; Jing, L.; Li, H.; Qiu, F.; Wu, W. Tree Species Classification of Forest Stands Using Multisource Remote Sensing Data. Remote Sensing 2021, 13. [Google Scholar] [CrossRef]
- Cormier, K.; Zhang, K.F.; Padron-Uy, J.; Wong, A.; Gagnier, K.; Parihar, A. Data Warehouse Design for Multiple Source Forest Inventory Management and Image Processing. In Proceedings of the 2025 IEEE Canadian Conference on Electrical and Computer Engineering (CCECE); IEEE, 2025; pp. 475–480. [Google Scholar] [CrossRef]
- Liu, B.; Hao, Y.; Huang, H.; Chen, S.; Li, Z.; Chen, E.; Tian, X.; Ren, M. TSCMDL: Multimodal Deep Learning Framework for Classifying Tree Species Using Fusion of 2-D and 3-D Features. IEEE Transactions on Geoscience and Remote Sensing 2023, 61, 1–11. [Google Scholar] [CrossRef]
- Cao, Y.; Coops, N.C.; Murray, B.A.; Sinclair, I.; Geordie, R.M. M3FNet: Multi-modal multi-temporal multi-scale data fusion network for tree species composition mapping. ISPRS Journal of Photogrammetry and Remote Sensing 2026, 231, 797–814. [Google Scholar] [CrossRef]
- He, X.; Han, X.; Zhao, Y.; Chen, Y.; Zou, L. Uncertainty-Based Dendritic Model for Multimodal Remote Sensing Data Classification. IEEE Transactions on Geoscience and Remote Sensing 2026, 64, 1–18. [Google Scholar] [CrossRef]
- Hu, B.; Zhang, F.; Wang, J. On the retrival of vegetation parameters from multi-angular hyperspectral remote sensing data. In Proceedings of the 2009 IEEE Toronto International Conference Science and Technology for Humanity (TIC-STH); IEEE, 2009. [Google Scholar] [CrossRef]
- Zhang, K.; Hu, B. Individual Urban Tree Species Classification Using Very High Spatial Resolution Airborne Multi-Spectral Imagery Using Longitudinal Profiles. Remote Sensing 2012, 4, 1741–1764. [Google Scholar] [CrossRef]
- Zhang, K.; Robinson, J.; Jing, L. Canopy vertical parameters estimation using unmanned aerial vehicle (UAV) imagery. In Proceedings of the 2016 IEEE International Geoscience and Remote Sensing Symposium (IGARSS); IEEE, 2016. [Google Scholar] [CrossRef]
- Zhang, K.; Jing, L.; Robinson, J. Douglas fir productivity estimation using very high spatial resolution imagery - a case study on ground treatment impact in west Kootenay, British Columbia, Canada. In Proceedings of the 2016 IEEE International Geoscience and Remote Sensing Symposium (IGARSS); IEEE, 2016. [Google Scholar] [CrossRef]
- Balkenhol, L.; Zhang, K.F. Identifying Individual Tree Species Structure with High-Resolution Hyperspectral Imagery Using a Linear Interpretation of the Spectral Signature. In Proceedings of the 38th Canadian Symposium on Remote Sensing, 2017. [Google Scholar]
- Miao, S.; Zhang, K.F.; Zeng, H.; Liu, J. Improving artificial-intelligence-based individual tree species classification using pseudo tree crown derived from unmanned aerial vehicle imagery. Remote Sensing 2024, 16, 1849. [Google Scholar] [CrossRef]
- Zhang, K.; Zhang, T.; Liu, J. Individual tree species classification using Pseudo Tree Crown (PTC) on coniferous forests. Remote Sensing 2025, 17, 3102. [Google Scholar] [CrossRef]
- Tucker, C.J. Red and photographic infrared linear combinations for monitoring vegetation. Remote Sensing of Environment 1979, 8, 127–150. [Google Scholar] [CrossRef]
- Breiman, L. Random Forests. Machine Learning 2001, 45, 5–32. [Google Scholar] [CrossRef]
- He, K.; Zhang, X.; Ren, S.; Sun, J. Deep Residual Learning for Image Recognition. In Proceedings of the Proceedings of the IEEE Conference on Computer Vision and Pattern Recognition (CVPR), 2016; pp. 770–778. [Google Scholar] [CrossRef]
- Wang, A.; Chen, H.; Liu, L.; Chen, K.; Lin, Z.; Han, J.; Ding, G. YOLOv10: Real-Time End-to-End Object Detection. arXiv 2024, arXiv:2405.14458. [Google Scholar] [CrossRef]
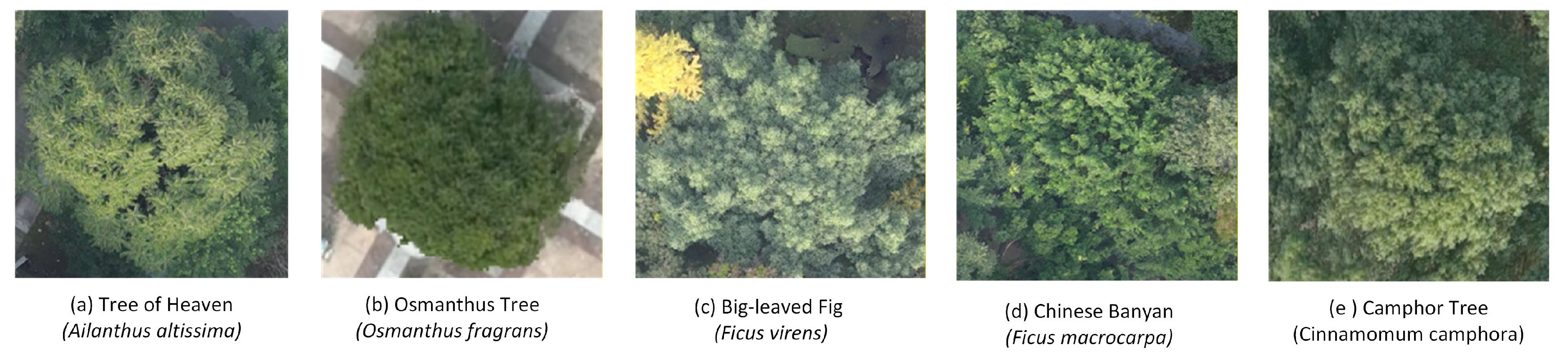

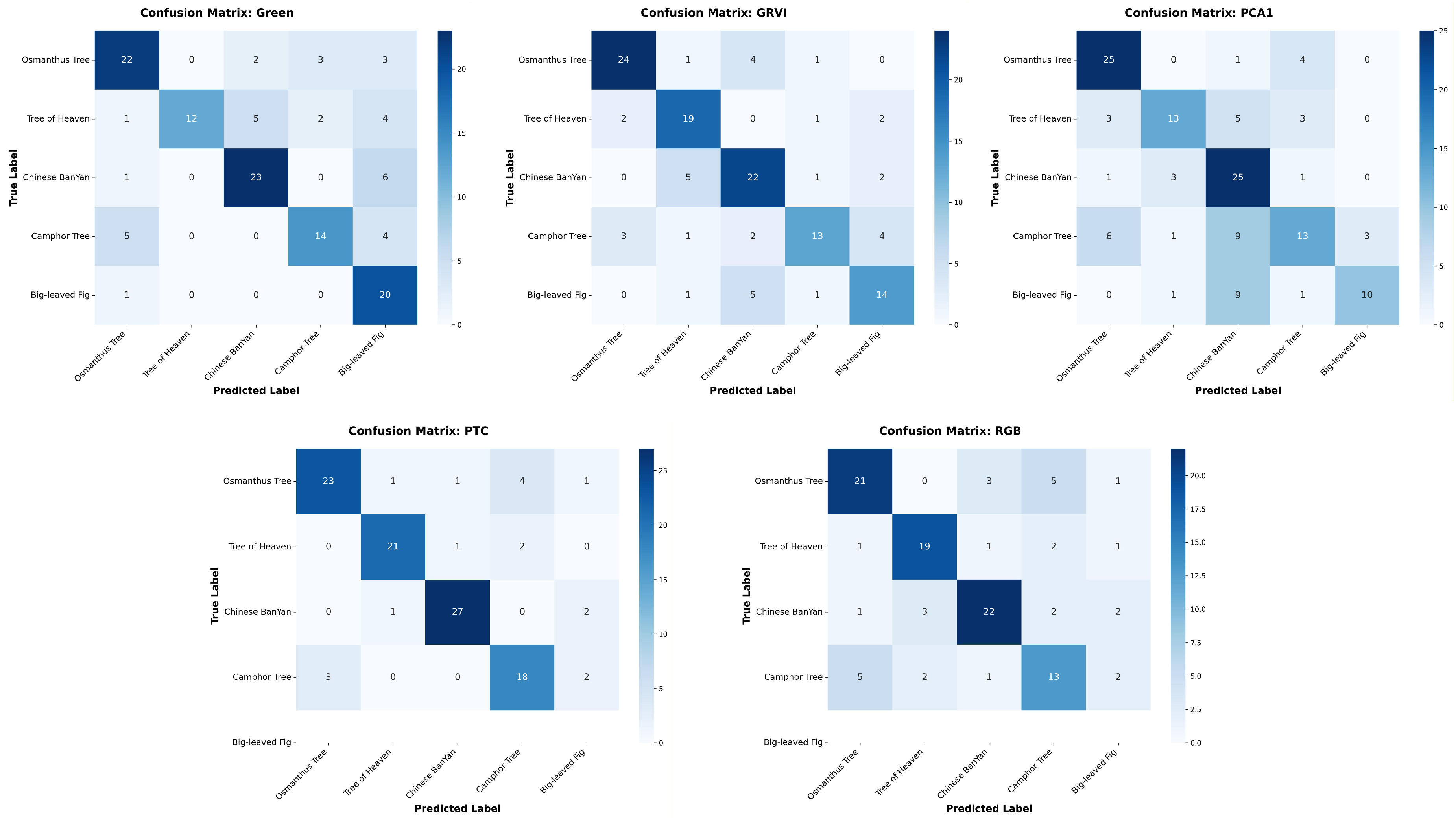
| Model | Data | Tree Species | Precision | Recall | F1-score | IoU |
|---|---|---|---|---|---|---|
| RF | RGB | Tree of Heaven | 0.4570 | 0.4513 | 0.4506 | 0.2910 |
| Osmanthus Tree | 0.4778 | 0.3888 | 0.4216 | 0.2672 | ||
| Chinese Banyan | 0.4922 | 0.5218 | 0.5024 | 0.3379 | ||
| Big-leaved Fig | 0.4644 | 0.4366 | 0.4466 | 0.2875 | ||
| Campphor Tree | 0.4211 | 0.4821 | 0.4439 | 0.2870 | ||
| Green | Tree of Heaven | 0.4346 | 0.3894 | 0.4083 | 0.2606 | |
| Osmanthus Tree | 0.5150 | 0.5439 | 0.5269 | 0.3590 | ||
| Chinese Banyan | 0.5150 | 0.5439 | 0.5269 | 0.3590 | ||
| Big-leaved Fig | 0.5354 | 0.4341 | 0.4740 | 0.3133 | ||
| Campphor Tree | 0.4726 | 0.5044 | 0.4857 | 0.3212 | ||
| GRVI | Tree of Heaven | 0.4501 | 0.4360 | 0.4383 | 0.2812 | |
| Osmanthus Tree | 0.4833 | 0.3783 | 0.4133 | 0.2610 | ||
| Chinese Banyan | 0.4905 | 0.5283 | 0.5046 | 0.3376 | ||
| Big-leaved Fig | 0.4833 | 0.4880 | 0.4812 | 0.3187 | ||
| Campphor Tree | 0.4000 | 0.4451 | 0.4146 | 0.2629 | ||
| PCA1 | Tree of Heaven | 0.4468 | 0.4572 | 0.4489 | 0.2936 | |
| Osmanthus Tree | 0.5080 | 0.5283 | 0.5167 | 0.3533 | ||
| Chinese Banyan | 0.4950 | 0.5283 | 0.5113 | 0.3499 | ||
| Big-leaved Fig | 0.4327 | 0.3945 | 0.4106 | 0.2616 | ||
| Campphor Tree | 0.4886 | 0.4630 | 0.4764 | 0.3155 | ||
| PTC | Tree of Heaven | 0.6337 | 0.6097 | 0.6196 | 0.4517 | |
| Osmanthus Tree | 0.6381 | 0.6514 | 0.6431 | 0.4769 | ||
| Chinese Banyan | 0.6948 | 0.6847 | 0.6881 | 0.5303 | ||
| Big-leaved Fig | 0.6280 | 0.6752 | 0.6486 | 0.4830 | ||
| Campphor Tree | 0.5427 | 0.5354 | 0.5374 | 0.3758 | ||
| ResNet50 | RGB | Tree of Heaven | 0.7241 | 0.7000 | 0.7119 | 0.5526 |
| Osmanthus Tree | 0.7917 | 0.7917 | 0.7917 | 0.6552 | ||
| Chinese Banyan | 0.6471 | 0.7333 | 0.6875 | 0.5238 | ||
| Big-leaved Fig | 0.6667 | 0.5714 | 0.6154 | 0.4444 | ||
| Campphor Tree | 0.5652 | 0.5652 | 0.5652 | 0.3939 | ||
| Green | Tree of Heaven | 1.0000 | 0.5000 | 0.6667 | 0.5000 | |
| Osmanthus Tree | 0.7333 | 0.7333 | 0.7333 | 0.5789 | ||
| Chinese Banyan | 0.7667 | 0.7667 | 0.7667 | 0.6216 | ||
| Big-leaved Fig | 0.5405 | 0.9524 | 0.6897 | 0.5263 | ||
| Campphor Tree | 0.7368 | 0.6087 | 0.6667 | 0.5000 | ||
| GRVI | Tree of Heaven | 0.778 | 0.5833 | 0.6667 | 0.5000 | |
| Osmanthus Tree | 0.5946 | 0.7333 | 0.6567 | 0.4889 | ||
| Chinese Banyan | 0.5429 | 0.6333 | 0.5846 | 0.4130 | ||
| Big-leaved Fig | 0.6500 | 0.6190 | 0.6341 | 0.4643 | ||
| Campphor Tree | 0.6667 | 0.5217 | 0.5854 | 0.4138 | ||
| PCA1 | Tree of Heaven | 0.7222 | 0.5417 | 0.6190 | 0.4483 | |
| Osmanthus Tree | 0.7143 | 0.8333 | 0.7692 | 0.6250 | ||
| Chinese Banyan | 0.6250 | 0.8333 | 0.7143 | 0.5556 | ||
| Big-leaved Fig | 0.7692 | 0.4762 | 0.5882 | 0.4167 | ||
| Campphor Tree | 0.5909 | 0.5652 | 0.5882 | 0.4062 | ||
| PTC | Tree of Heaven | 0.9130 | 0.8750 | 0.8936 | 0.8077 | |
| Osmanthus Tree | 0.8846 | 0.7667 | 0.8214 | 0.6970 | ||
| Chinese Banyan | 0.9000 | 0.9000 | 0.9000 | 0.8182 | ||
| Big-leaved Fig | 0.8000 | 0.9524 | 0.8696 | 0.7692 | ||
| Campphor Tree | 0.7500 | 0.7826 | 0.7660 | 0.6207 | ||
| YOLOv10 | RGB | Tree of Heaven | 0.8662 | 0.6064 | 0.7006 | 0.5444 |
| Osmanthus Tree | 0.7289 | 0.5067 | 0.5929 | 0.4240 | ||
| Chinese Banyan | 0.7116 | 0.4999 | 0.6029 | 0.4330 | ||
| Big-leaved Fig | 0.4950 | 0.6299 | 0.5535 | 0.3875 | ||
| Campphor Tree | 0.4276 | 0.6081 | 0.5019 | 0.3361 | ||
| Green | Tree of Heaven | 0.7802 | 0.6384 | 0.6917 | 0.5454 | |
| Osmanthus Tree | 0.7503 | 0.6686 | 0.6591 | 0.5030 | ||
| Chinese Banyan | 0.8103 | 0.6291 | 0.7029 | 0.5566 | ||
| Big-leaved Fig | 0.7319 | 0.7510 | 0.7265 | 0.5863 | ||
| Campphor Tree | 0.4769 | 0.6317 | 0.5219 | 0.3576 | ||
| GRVI | Tree of Heaven | 0.8675 | 0.6058 | 0.7002 | 0.5430 | |
| Osmanthus Tree | 0.7259 | 0.5072 | 0.5937 | 0.4240 | ||
| Chinese Banyan | 0.7120 | 0.5431 | 0.6031 | 0.4335 | ||
| Big-leaved Fig | 0.4945 | 0.6324 | 0.5536 | 0.3882 | ||
| Campphor Tree | 0.4283 | 0.6184 | 0.5014 | 0.3363 | ||
| PCA1 | Tree of Heaven | 0.7639 | 0.5413 | 0.6336 | 0.4807 | |
| Osmanthus Tree | 0.6721 | 0.5850 | 0.6233 | 0.4559 | ||
| Chinese Banyan | 0.5803 | 0.4826 | 0.5247 | 0.3664 | ||
| Big-leaved Fig | 0.5136 | 0.5149 | 0.5109 | 0.3661 | ||
| Campphor Tree | 0.5380 | 0.6325 | 0.5819 | 0.4152 | ||
| PTC | Tree of Heaven | 0.9359 | 0.6063 | 0.7774 | 0.5675 | |
| Osmanthus Tree | 0.7548 | 0.8750 | 0.8030 | 0.6721 | ||
| Chinese Banyan | 0.8449 | 0.7674 | 0.8080 | 0.6779 | ||
| Big-leaved Fig | 0.9853 | 0.7210 | 0.8104 | 0.7077 | ||
| Campphor Tree | 0.4820 | 0.5787 | 0.5186 | 0.3732 |
| Input Data | Random Forest | ResNet50 | YOLOv10 |
|---|---|---|---|
| RGB | |||
| Green | |||
| GRVI | |||
| PCA1 | |||
| PTC |
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2026 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).




