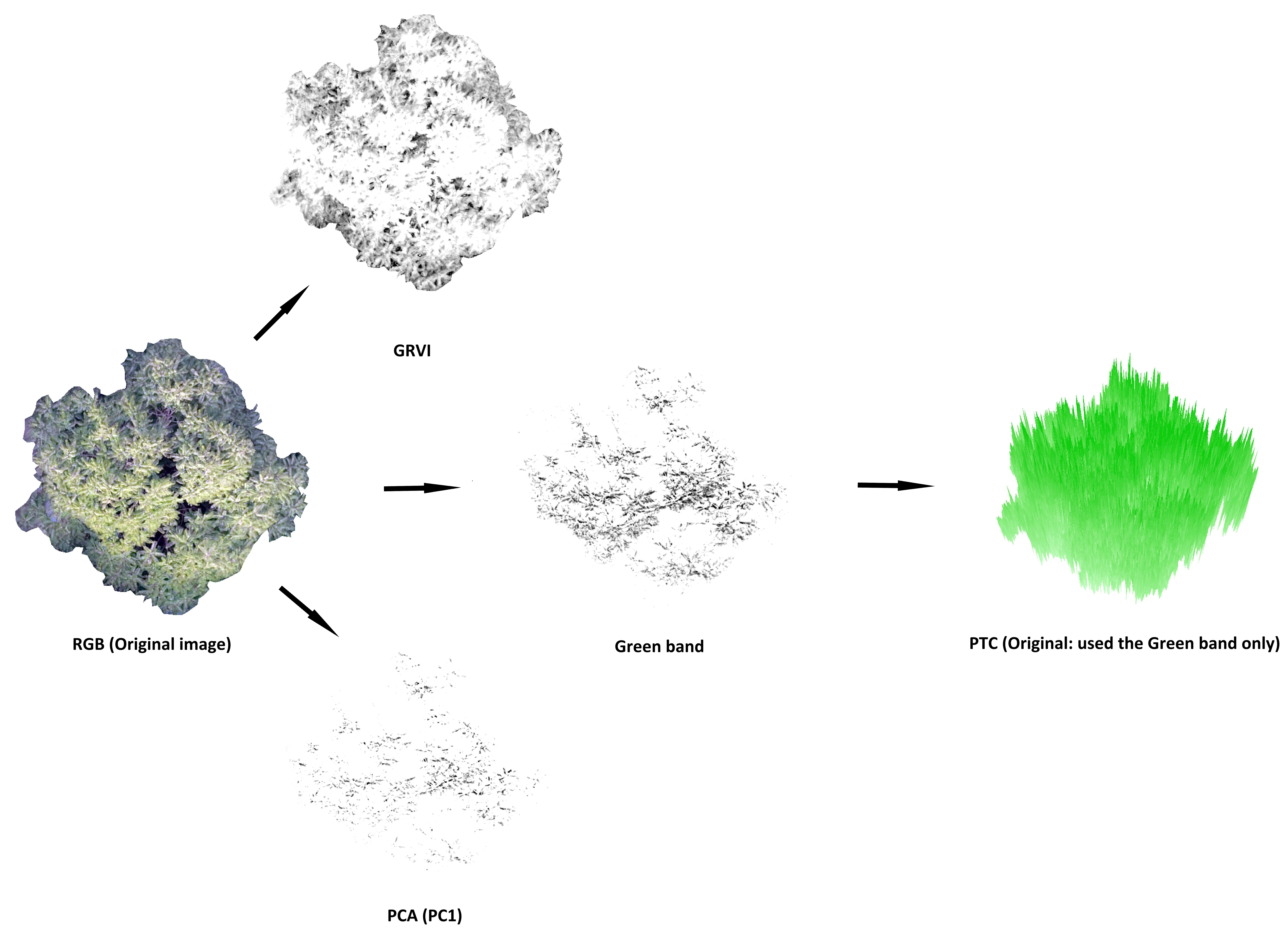
Individual tree classification has a long history of diverse development, with recent trends focusing on the adoption of machine learning and deep learning approaches. A simple and powerful approach that lets the model auto-pilot, but weakens the need for physical characteristic understanding. Over more than a decade of our research, we have focused on establishing a direct representation of individual trees that bridges 2D top-down imagery and true 3D models. In this study, we investigated the fundamental question of the influence of the input data on these ML/DL models. In 2024, we introduced a novel data transformation method, the Pseudo Tree Crown (PTC), which provides a pseudo-3D pixel-value perspective that enhances the informational richness of images and significantly improves classification performance. Our original implementation was successfully tested on urban and deciduous trees in 2024 and was later extended to Canadian natural conifer species under snow conditions in 2025. However, the original PTC relied on the green band, limiting its applicability to green-leaf species. In this study, we analyzed and compared the performance of different data variations and transformations, such as the Green–Red Vegetation Index (GRVI) and Principal Component Analysis (PCA), as direct input and used their PTC forms. Classifications were conducted using Random Forest, ResNet50, and YOLOv10. The results confirmed the effectiveness of the PTC, which consistently improves classification accuracy by at least 7% without introducing additional computational time or complexity. Furthermore, PTC exhibits robust, consistent behaviour across all data forms, demonstrating its strong resilience and reliability.