Submitted:
21 April 2026
Posted:
22 April 2026
You are already at the latest version
Abstract
Keywords:
1. Introduction
2. Cellular State Landscapes in Human Disease
2.1. Technologies Enabling Cellular State Profiling
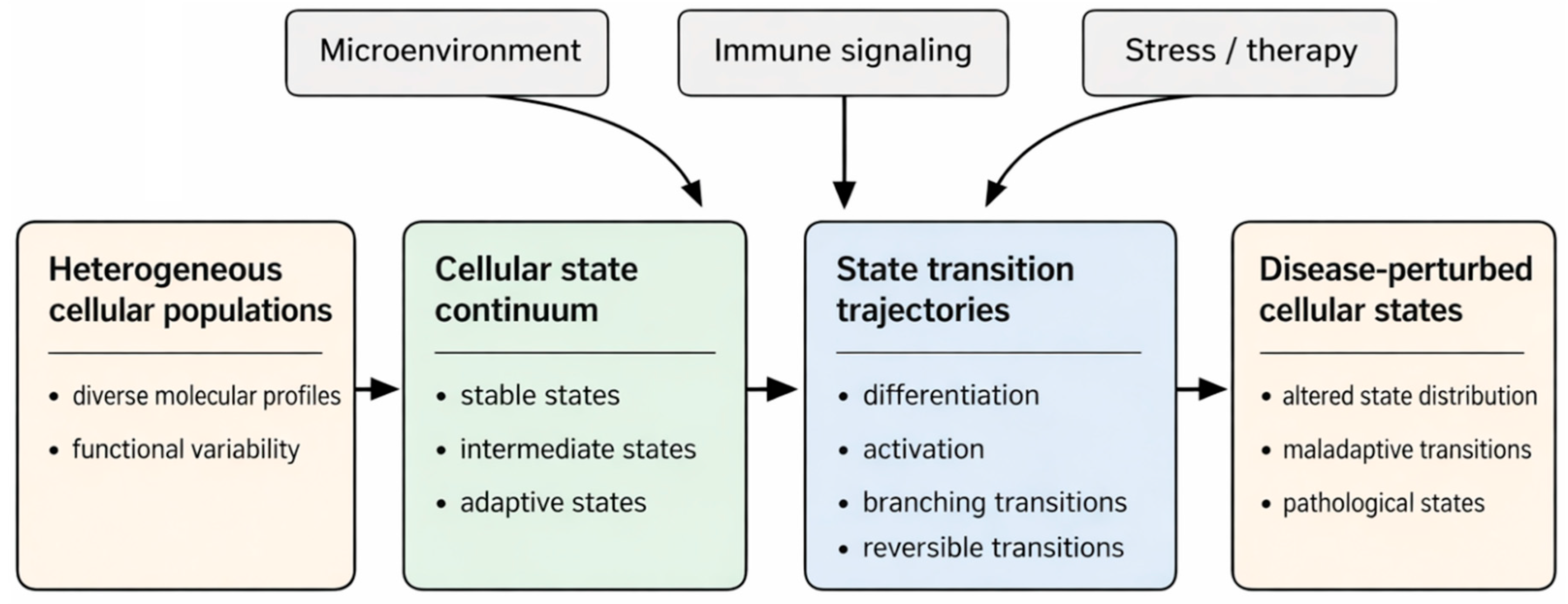
2.2. Cellular State Continuums and Landscape Frameworks
2.3. Disease as a Perturbation of Cellular State Landscapes
2.4. Dynamics, Quantification, and Transition Modeling
2.5. Limitations and Challenges in Cellular State Mapping
3. Artificial Intelligence for Cellular State Mapping
3.1. Computational Challenges in Cellular State Analysis
3.2. Machine Learning for Representation and Feature Extraction
3.3. Trajectory Inference and Modeling of State Transitions
3.4. Graph-Based and Network Modeling Approaches
3.5. Multimodal Integration and Inference of Cellular State Transitions
4. Neural Circuits as Dynamic Disease Networks
4.1. Organization and Functional Principles of Neural Circuits
4.2. Experimental Mapping of Circuit Structure and Dynamics
4.3. Neural Circuit Dysfunction and Disease Signatures
4.4. Dynamic Circuit States, Cellular Interactions, and Challenges
5. AI-Driven Neural Circuit Decoding
5.1. Decoding, Classification, and Prediction of Neural Activity
5.2. AI Analysis of Brain Imaging and Large-Scale Circuit Data
5.3. Brain–Machine Interfaces, Closed-Loop Systems, and Modeling of Circuit Dynamics
6. Linking Cellular States and Neural Circuits
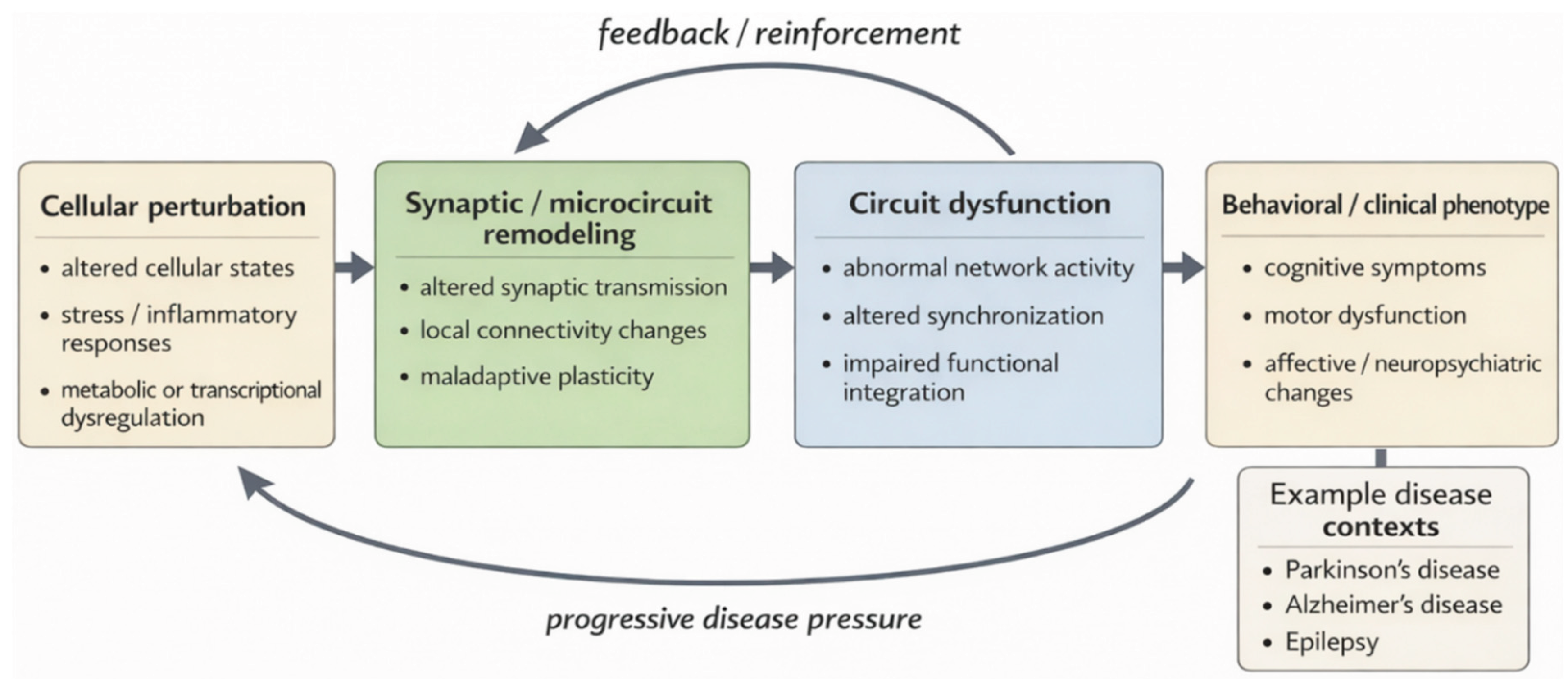
6.1. Cellular–Circuit Interactions Across Scales
6.2. Toward Integrative Multiscale Models of Disease
7. Toward AI-Integrated Multiscale Medicine
7.1. Multiscale Organization of Disease and the Need for Integration
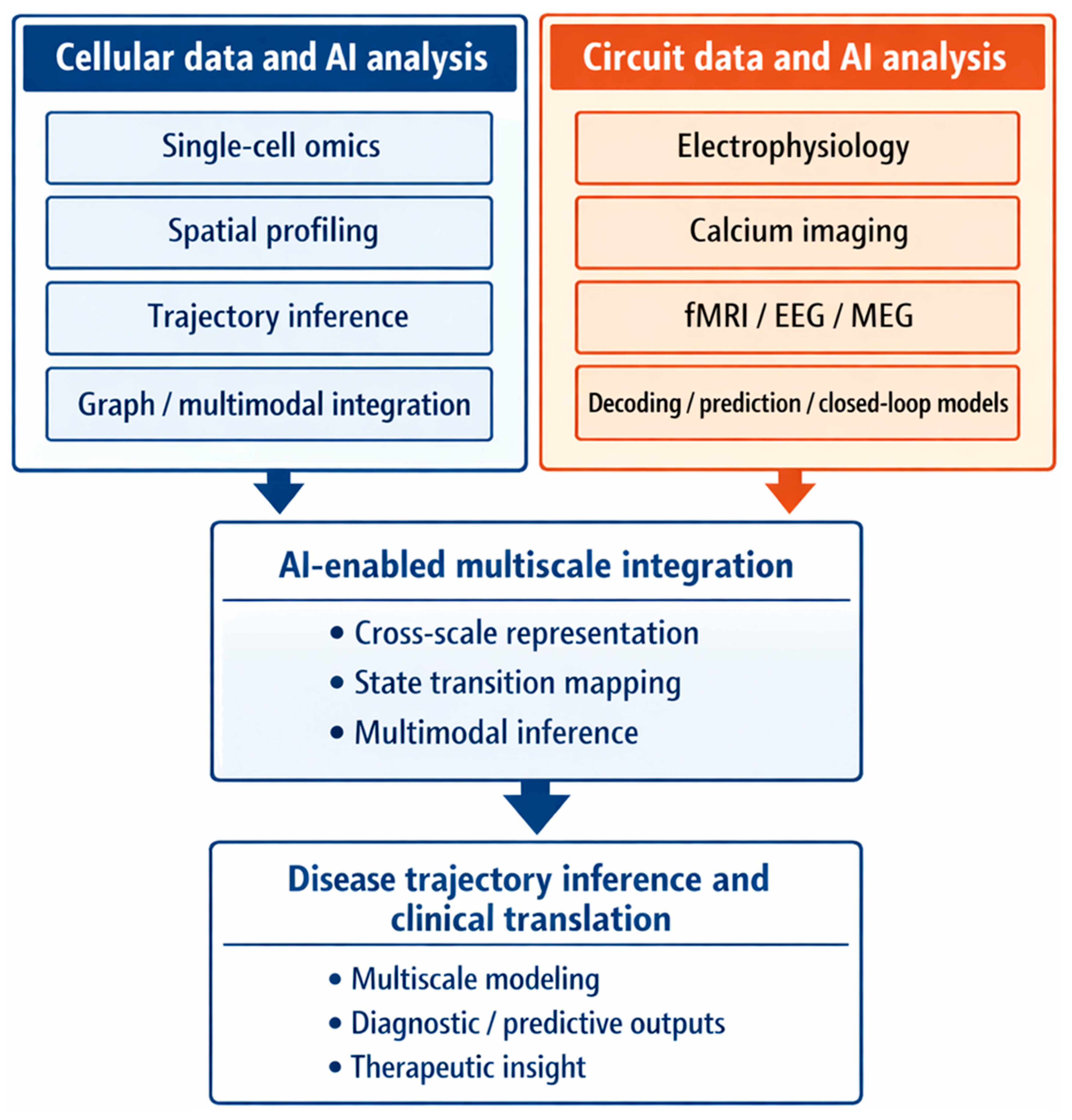
7.2. Artificial Intelligence as a Framework for Multiscale Integration
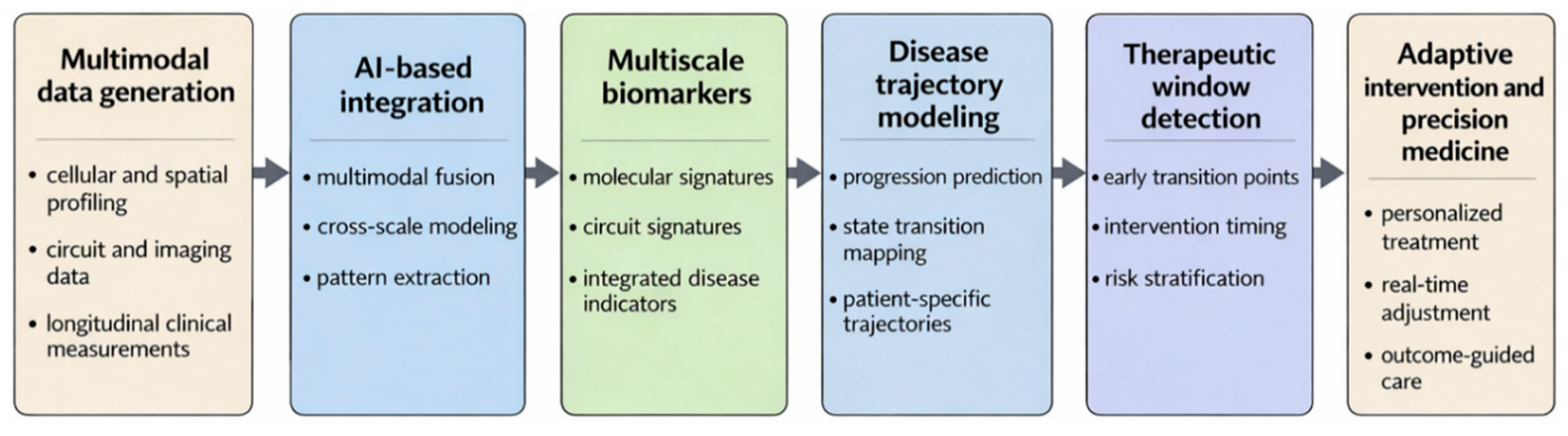
7.3. Implications for Diagnosis, Prediction, and Therapeutic Strategies
8. Future Directions
Conclusions
References
- Chen, J. Towards time-resolved multiscale and multimodal imaging. Npj Imaging 2025, 3, 67. [Google Scholar] [CrossRef]
- Khan, A.F.; Iturria-Medina, Y. Beyond the usual suspects: multi-factorial computational models in the search for neurodegenerative disease mechanisms. Transl Psychiatry 2024, 14, 386. [Google Scholar] [CrossRef]
- Liu, L.; Chen, A.; Li, Y.; Mulder, J.; Heyn, H.; Xu, X. Spatiotemporal omics for biology and medicine. Cell 2024, 187, 4488–4519. [Google Scholar] [CrossRef]
- Rafelski, S.M.; Theriot, J.A. Establishing a conceptual framework for holistic cell states and state transitions. Cell 2024, 187, 2633–2651. [Google Scholar] [CrossRef] [PubMed]
- Short, S.; Garcia-Tejera, R.; Schumacher, L.J.; Coutu, D.L. Next generation lineage tracing and its applications to unravel development. NPJ Syst Biol Appl 2025, 11, 60. [Google Scholar] [CrossRef]
- Zhang, Z.; Wang, Z.; Sun, Y.; Shen, J.; Peng, Q.; Li, T.; Zhou, P. Deciphering cell-fate trajectories using spatiotemporal single-cell transcriptomic data. NPJ Syst Biol Appl 2025, 12, 2. [Google Scholar] [CrossRef]
- Demmings, M.D.; da Silva Chagas, L.; Traetta, M.E.; Rodrigues, R.S.; Acutain, M.F.; Barykin, E.; Datusalia, A.K.; German-Castelan, L.; Mattera, V.S.; Mazengenya, P.; et al. (Re)building the nervous system: A review of neuron-glia interactions from development to disease. J Neurochem 2025, 169, e16258. [Google Scholar] [CrossRef]
- Tizabi, Y.; Getachew, B.; Hauser, S.R.; Tsytsarev, V.; Manhaes, A.C.; da Silva, V.D.A. Role of Glial Cells in Neuronal Function, Mood Disorders, and Drug Addiction. Brain Sci 2024, 14. [Google Scholar] [CrossRef]
- Zhang, W.; Xiao, D.; Mao, Q.; Xia, H. Role of neuroinflammation in neurodegeneration development. Signal Transduct Target Ther 2023, 8, 267. [Google Scholar] [CrossRef] [PubMed]
- Naffaa, M.M. Significance of the anterior cingulate cortex in neurogenesis plasticity: Connections, functions, and disorders across postnatal and adult stages. Bioessays 2024, 46, e2300160. [Google Scholar] [CrossRef] [PubMed]
- Bunne, C.; Roohani, Y.; Rosen, Y.; Gupta, A.; Zhang, X.; Roed, M.; Alexandrov, T.; AlQuraishi, M.; Brennan, P.; Burkhardt, D.B.; et al. How to build the virtual cell with artificial intelligence: Priorities and opportunities. Cell 2024, 187, 7045–7063. [Google Scholar] [CrossRef]
- Nam, Y.; Kim, J.; Jung, S.H.; Woerner, J.; Suh, E.H.; Lee, D.G.; Shivakumar, M.; Lee, M.E.; Kim, D. Harnessing Artificial Intelligence in Multimodal Omics Data Integration: Paving the Path for the Next Frontier in Precision Medicine. Annu Rev Biomed Data Sci 2024, 7, 225–250. [Google Scholar] [CrossRef]
- Onciul, R.; Tataru, C.I.; Dumitru, A.V.; Crivoi, C.; Serban, M.; Covache-Busuioc, R.A.; Radoi, M.P.; Toader, C. Artificial Intelligence and Neuroscience: Transformative Synergies in Brain Research and Clinical Applications. J Clin Med 2025, 14. [Google Scholar] [CrossRef]
- Tchatchoua Ngassam, B.; Niu, H.; Pang, S.; Shydlouskaya, V.; Andrews, T.S. Applications of AI to single-cell and spatial transcriptomics: current state-of-the-art and challenges. Front Bioinform 2025, 5, 1715821. [Google Scholar] [CrossRef]
- Bonev, B.; Castelo-Branco, G.; Chen, F.; Codeluppi, S.; Corces, M.R.; Fan, J.; Heiman, M.; Harris, K.; Inoue, F.; Kellis, M.; et al. Opportunities and challenges of single-cell and spatially resolved genomics methods for neuroscience discovery. Nat Neurosci 2024, 27, 2292–2309. [Google Scholar] [CrossRef] [PubMed]
- Chang, E.H. Bridging complexity through integrative systems neuroscience. Front Syst Biol 2024, 4, 1487298. [Google Scholar] [CrossRef] [PubMed]
- Hanchate, N.K. Single-cell genomics meets systems neuroscience: Insights from mapping the brain circuitry of stress. J Neuroendocrinol 2025, 37, e70005. [Google Scholar] [CrossRef] [PubMed]
- Barnett, S.N.; Cujba, A.M.; Yang, L.; Maceiras, A.R.; Li, S.; Kedlian, V.R.; Pett, J.P.; Polanski, K.; Miranda, A.M.A.; Xu, C.; et al. An organotypic atlas of human vascular cells. Nat Med 2024, 30, 3468–3481. [Google Scholar] [CrossRef]
- Lai, Y.; Ramirez-Pardo, I.; Isern, J.; An, J.; Perdiguero, E.; Serrano, A.L.; Li, J.; Garcia-Dominguez, E.; Segales, J.; Guo, P.; et al. Multimodal cell atlas of the ageing human skeletal muscle. Nature 2024, 629, 154–164. [Google Scholar] [CrossRef]
- Lake, B.B.; Menon, R.; Winfree, S.; Hu, Q.; Melo Ferreira, R.; Kalhor, K.; Barwinska, D.; Otto, E.A.; Ferkowicz, M.; Diep, D.; et al. An atlas of healthy and injured cell states and niches in the human kidney. Nature 2023, 619, 585–594. [Google Scholar] [CrossRef] [PubMed]
- Walchli, T.; Ghobrial, M.; Schwab, M.; Takada, S.; Zhong, H.; Suntharalingham, S.; Vetiska, S.; Gonzalez, D.R.; Wu, R.; Rehrauer, H.; et al. Single-cell atlas of the human brain vasculature across development, adulthood and disease. Nature 2024, 632, 603–613. [Google Scholar] [CrossRef]
- Carstens, J.L.; Krishnan, S.N.; Rao, A.; Sorace, A.G.; Seeley, E.H.; Ferri-Borgogno, S.; Burks, J.K. Spatial Multiplexing and Omics. Nat Rev Methods Primers 2024, 4. [Google Scholar] [CrossRef] [PubMed]
- Chelebian, E.; Avenel, C.; Wahlby, C. Combining spatial transcriptomics with tissue morphology. Nat Commun 2025, 16, 4452. [Google Scholar] [CrossRef] [PubMed]
- Wang, H.; Huang, R.; Nelson, J.; Gao, C.; Tran, M.; Yeaton, A.; Krishna, S.; Felt, K.; Pfaff, K.L.; Bowman, T.; et al. Systematic benchmarking of imaging spatial transcriptomics platforms in FFPE tissues. Nat Commun 2025, 16, 10215. [Google Scholar] [CrossRef] [PubMed]
- Yapp, C.; Nirmal, A.J.; Zhou, F.; Wong, A.Y.H.; Tefft, J.B.; Lu, Y.D.; Shang, Z.; Maliga, Z.; Llopis, P.M.; Murphy, G.F.; et al. Highly multiplexed 3D profiling of cell states and immune niches in human tumors. Nat Methods 2025, 22, 2180–2193. [Google Scholar] [CrossRef]
- Hutton, A.; Meyer, J.G. Trajectory Inference for Single Cell Omics; 2025. [Google Scholar]
- Sashittal, P.; Zhang, R.Y.; Law, B.K.; Schmidt, H.; Strzalkowski, A.; Bolondi, A.; Chan, M.M.; Raphael, B.J. Inferring cell differentiation maps from lineage tracing data. Nat Methods 2026, 23, 532–541. [Google Scholar] [CrossRef]
- Weiler, P.; Lange, M.; Klein, M.; Pe'er, D.; Theis, F. CellRank 2: unified fate mapping in multiview single-cell data. Nat Methods 2024, 21, 1196–1205. [Google Scholar] [CrossRef]
- Zhou, Y.; Hou, S.; Miao, X.; Zhang, G.; Li, Z.; Zhang, D.; Lin, Y.; Lin, Y. Decoding cell fate: integrated experimental and computational analysis at the single-cell level. Bioinformatics 2025, 41. [Google Scholar] [CrossRef]
- Ianevski, A.; Nader, K.; Driva, K.; Senkowski, W.; Bulanova, D.; Moyano-Galceran, L.; Ruokoranta, T.; Kuusanmaki, H.; Ikonen, N.; Sergeev, P.; et al. Single-cell transcriptomes identify patient-tailored therapies for selective co-inhibition of cancer clones. Nat Commun 2024, 15, 8579. [Google Scholar] [CrossRef]
- Jeong, D.; Danielli, S.G.; Maass, K.K.; Ghasemi, D.R.; Tetzlaff, S.K.; Reyhan, E.; Jiang, L.; Katiyar, S.; Sundheimer, J.K.; Lo Cascio, C.; et al. Multidimensional profiling of heterogeneity in supratentorial ependymomas. Nature 2026. [Google Scholar] [CrossRef]
- Otsuji, K.; Takahashi, Y.; Osako, T.; Kobayashi, T.; Takano, T.; Saeki, S.; Yang, L.; Baba, S.; Kumegawa, K.; Suzuki, H.; et al. Serial single-cell RNA sequencing unveils drug resistance and metastatic traits in stage IV breast cancer. NPJ Precis Oncol 2024, 8, 222. [Google Scholar] [CrossRef] [PubMed]
- Tirosh, I.; Suva, M.L. Cancer cell states: Lessons from ten years of single-cell RNA-sequencing of human tumors. Cancer Cell 2024, 42, 1497–1506. [Google Scholar] [CrossRef] [PubMed]
- Ferrari-Souza, J.P.; Povala, G.; Rahmouni, N.; Bellaver, B.; Ferreira, P.C.L.; De Bastiani, M.A.; Leffa, D.T.; Lussier, F.Z.; Aguzzoli, C.S.; Brum, W.S.; et al. Microglia modulate Abeta-dependent astrocyte reactivity in Alzheimer's disease. Nat Neurosci 2026, 29, 81–87. [Google Scholar] [CrossRef]
- Martins-Ferreira, R.; Calafell-Segura, J.; Leal, B.; Rodriguez-Ubreva, J.; Martinez-Saez, E.; Mereu, E.; Pinho, E.C.P.; Laguna, A.; Ballestar, E. The Human Microglia Atlas (HuMicA) unravels changes in disease-associated microglia subsets across neurodegenerative conditions. Nat Commun 2025, 16, 739. [Google Scholar] [CrossRef]
- Prater, K.E.; Green, K.J.; Mamde, S.; Sun, W.; Cochoit, A.; Smith, C.L.; Chiou, K.L.; Heath, L.; Rose, S.E.; Wiley, J.; et al. Human microglia show unique transcriptional changes in Alzheimer's disease. Nat Aging 2023, 3, 894–907. [Google Scholar] [CrossRef]
- Serrano-Pozo, A.; Li, H.; Li, Z.; Munoz-Castro, C.; Jaisa-Aad, M.; Healey, M.A.; Welikovitch, L.A.; Jayakumar, R.; Bryant, A.G.; Noori, A.; et al. Astrocyte transcriptomic changes along the spatiotemporal progression of Alzheimer's disease. Nat Neurosci 2024, 27, 2384–2400. [Google Scholar] [CrossRef]
- Almet, A.A.; Tsai, Y.C.; Watanabe, M.; Nie, Q. Inferring pattern-driving intercellular flows from single-cell and spatial transcriptomics. Nat Methods 2024, 21, 1806–1817. [Google Scholar] [CrossRef]
- Sun, Y.; Shao, C.; Duan, H.; Wang, Z.; Xu, S.; Wang, C.; Xiu, J.; Liu, J.; Wang, X.; Yao, X.; et al. Dynamic Evolution of the Tumor Immune Microenvironment in Malignant Tumors and Emerging Therapeutic Paradigms. MedComm (2020) 2025, 6. [CrossRef]
- Lim, H.S.; Qiu, P. Quantifying the clusterness and trajectoriness of single-cell RNA-seq data. PLoS Comput Biol 2024, 20, e1011866. [Google Scholar] [CrossRef]
- Ranek, J.S.; Stallaert, W.; Milner, J.J.; Redick, M.; Wolff, S.C.; Beltran, A.S.; Stanley, N.; Purvis, J.E. DELVE: feature selection for preserving biological trajectories in single-cell data. Nat Commun 2024, 15, 2765. [Google Scholar] [CrossRef] [PubMed]
- Millard, N.; Chen, J.H.; Palshikar, M.G.; Pelka, K.; Spurrell, M.; Price, C.; He, J.; Hacohen, N.; Raychaudhuri, S.; Korsunsky, I. Batch correcting single-cell spatial transcriptomics count data with Crescendo improves visualization and detection of spatial gene patterns. Genome Biol 2025, 26, 36. [Google Scholar] [CrossRef]
- Nguyen, H.C.T.; Baik, B.; Yoon, S.; Park, T.; Nam, D. Benchmarking integration of single-cell differential expression. Nat Commun 2023, 14, 1570. [Google Scholar] [CrossRef]
- Zhu, Q.; Jiang, Z.; Zuckerman, B.; Weinberger, L.; Thomson, M.; Gartner, Z.J. Revealing a coherent cell-state landscape across single-cell datasets with CONCORD. Nat Biotechnol 2026. [Google Scholar] [CrossRef]
- Hevdeli, O.; Petrenko, E.; Aran, D. CellMentor: cell-type aware dimensionality reduction for single-cell RNA-sequencing data. Nat Commun 2025, 17, 396. [Google Scholar] [CrossRef]
- Hughes, S.; Hamilton, T.; Kolokotrones, T.; Deeds, E.J. DeepAtlas: a tool for effective manifold learning. bioRxiv 2025. [Google Scholar] [CrossRef]
- Nicol, P.B.; Miller, J.W. Model-based dimensionality reduction for single-cell RNA-seq using generalized bilinear models. Biostatistics 2025, 26. [Google Scholar] [CrossRef] [PubMed]
- Cao, Y.; Zhao, X.; Tang, S.; Jiang, Q.; Li, S.; Li, S.; Chen, S. scButterfly: a versatile single-cell cross-modality translation method via dual-aligned variational autoencoders. Nat Commun 2024, 15, 2973. [Google Scholar] [CrossRef]
- Joas, M.J.; Jurenaite, N.; Prascevic, D.; Scherf, N.; Ewald, J. AUTOENCODIX: a generalized and versatile framework to train and evaluate autoencoders for biological representation learning and beyond. Nat Comput Sci 2026, 6, 96–108. [Google Scholar] [CrossRef]
- Kang, M.; Gulati, G.S.; Brown, E.L.; Qi, Z.; Avagyan, S.; Armenteros, J.J.A.; Gleyzer, R.; Zhang, W.; Steen, C.B.; D'Silva, J.P.; et al. Improved reconstruction of single-cell developmental potential with CytoTRACE 2. Nat Methods 2025, 22, 2258–2263. [Google Scholar] [CrossRef] [PubMed]
- Klein, D.; Palla, G.; Lange, M.; Klein, M.; Piran, Z.; Gander, M.; Meng-Papaxanthos, L.; Sterr, M.; Saber, L.; Jing, C.; et al. Mapping cells through time and space with moscot. Nature 2025, 638, 1065–1075. [Google Scholar] [CrossRef] [PubMed]
- Sha, Y.; Qiu, Y.; Zhou, P.; Nie, Q. Reconstructing growth and dynamic trajectories from single-cell transcriptomics data. Nat Mach Intell 2024, 6, 25–39. [Google Scholar] [CrossRef] [PubMed]
- Sumanaweera, D.; Suo, C.; Cujba, A.M.; Muraro, D.; Dann, E.; Polanski, K.; Steemers, A.S.; Lee, W.; Oliver, A.J.; Park, J.E.; et al. Gene-level alignment of single-cell trajectories. Nat Methods 2025, 22, 68–81. [Google Scholar] [CrossRef]
- Ali, M.; Richter, S.; Erturk, A.; Fischer, D.S.; Theis, F.J. Graph neural networks learn emergent tissue properties from spatial molecular profiles. Nat Commun 2025, 16, 8419. [Google Scholar] [CrossRef] [PubMed]
- Yang, W.; Wang, P.; Xu, S.; Wang, T.; Luo, M.; Cai, Y.; Xu, C.; Xue, G.; Que, J.; Ding, Q.; et al. Deciphering cell-cell communication at single-cell resolution for spatial transcriptomics with subgraph-based graph attention network. Nat Commun 2024, 15, 7101. [Google Scholar] [CrossRef] [PubMed]
- Zhang, T.; Zhang, Z.; Li, L.; Dong, B.; Wang, G.; Zhang, D. GTAD: a graph-based approach for cell spatial composition inference from integrated scRNA-seq and ST-seq data. Brief Bioinform 2023, 25. [Google Scholar] [CrossRef]
- Ritter, M.; Blume, C.; Tang, Y.; Patel, A.; Patel, B.; Berghaus, N.; Kada Benotmane, J.; Kueckelhaus, J.; Yabo, Y.; Zhang, J.; et al. Spatially resolved transcriptomics and graph-based deep learning improve accuracy of routine CNS tumor diagnostics. Nat Cancer 2025, 6, 292–306. [Google Scholar] [CrossRef]
- Chen, S.; Zhu, B.; Huang, S.; Hickey, J.W.; Lin, K.Z.; Snyder, M.; Greenleaf, W.J.; Nolan, G.P.; Zhang, N.R.; Ma, Z. Integration of spatial and single-cell data across modalities with weakly linked features. Nat Biotechnol 2024, 42, 1096–1106. [Google Scholar] [CrossRef]
- Dong, C.; Meng, X.; Zhang, T.; Guo, Z.; Liu, Y.; Wu, P.; Chen, S.; Zhou, F.; Ma, Y.; Xiong, H.; et al. Single-cell EpiChem jointly measures drug-chromatin binding and multimodal epigenome. Nat Methods 2024, 21, 1624–1633. [Google Scholar] [CrossRef]
- He, Z.; Hu, S.; Chen, Y.; An, S.; Zhou, J.; Liu, R.; Shi, J.; Wang, J.; Dong, G.; Shi, J.; et al. Mosaic integration and knowledge transfer of single-cell multimodal data with MIDAS. Nat Biotechnol 2024, 42, 1594–1605. [Google Scholar] [CrossRef]
- Koladiya, A.; Good, Z.; Varra, S.R.; Bendall, S.C.; Davis, K.L. CellFuse Enables Multi-modal Integration of Single-cell and Spatial Proteomics data. bioRxiv 2025. [Google Scholar] [CrossRef]
- Mitra, S.; Malik, R.; Wong, W.; Rahman, A.; Hartemink, A.J.; Pritykin, Y.; Dey, K.K.; Leslie, C.S. Single-cell multi-ome regression models identify functional and disease-associated enhancers and enable chromatin potential analysis. Nat Genet 2024, 56, 627–636. [Google Scholar] [CrossRef]
- Ma, Y.; Yao, Y.; Zhou, Y.; Dai, W.; Li, J.; Gui, Y.; Sun, H.; Zhu, Z.; Jiang, D.; Chen, C.; et al. Integrating polygenic signals and single-cell multiomics identifies cell-type-specific regulomes critical for immune- and aging-related diseases. Nat Aging 2026, 6, 270–289. [Google Scholar] [CrossRef] [PubMed]
- Boyeau, P.; Hong, J.; Gayoso, A.; Kim, M.; McFaline-Figueroa, J.L.; Jordan, M.I.; Azizi, E.; Ergen, C.; Yosef, N. Deep generative modeling of sample-level heterogeneity in single-cell genomics. Nat Methods 2025, 22, 2264–2274. [Google Scholar] [CrossRef] [PubMed]
- Song, K.; Zheng, Y.; Zhao, B.; Eidelman, D.H.; Tang, J.; Ding, J. DOLPHIN advances single-cell transcriptomics beyond gene level by leveraging exon and junction reads. Nat Commun 2025, 16, 6202. [Google Scholar] [CrossRef] [PubMed]
- Schuster, V.; Dann, E.; Krogh, A.; Teichmann, S.A. multiDGD: A versatile deep generative model for multi-omics data. Nat Commun 2024, 15, 10031. [Google Scholar] [CrossRef]
- Tayyebi, Z.; Pine, A.R.; Leslie, C.S. Scalable and unbiased sequence-informed embedding of single-cell ATAC-seq data with CellSpace. Nat Methods 2024, 21, 1014–1022. [Google Scholar] [CrossRef]
- Chen, Y.; Fan, X.; Shi, C.; Shi, Z.; Wang, C. A joint analysis of single cell transcriptomics and proteomics using transformer. NPJ Syst Biol Appl 2025, 11, 1. [Google Scholar] [CrossRef]
- Liu, C.; Ding, S.; Kim, H.J.; Long, S.; Xiao, D.; Ghazanfar, S.; Yang, P. Multitask benchmarking of single-cell multimodal omics integration methods. Nat Methods 2025, 22, 2449–2460. [Google Scholar] [CrossRef]
- Yang, Y.; Cui, Y.; Zeng, X.; Zhang, Y.; Loza, M.; Park, S.J.; Nakai, K. STAIG: Spatial transcriptomics analysis via image-aided graph contrastive learning for domain exploration and alignment-free integration. Nat Commun 2025, 16, 1067. [Google Scholar] [CrossRef]
- Magness, A.; Colliver, E.; Enfield, K.S.S.; Lee, C.; Shimato, M.; Daly, E.; Moore, D.A.; Sivakumar, M.; Valand, K.; Levi, D.; et al. Deep cell phenotyping and spatial analysis of multiplexed imaging with TRACERx-PHLEX. Nat Commun 2024, 15, 5135. [Google Scholar] [CrossRef]
- Zohora, F.T.; Paliwal, D.; Flores-Figueroa, E.; Li, J.; Gao, T.; Notta, F.; Schwartz, G.W. CellNEST reveals cell-cell relay networks using attention mechanisms on spatial transcriptomics. Nat Methods 2025, 22, 1505–1519. [Google Scholar] [CrossRef] [PubMed]
- Zhu, F.; Grier, H.A.; Tandon, R.; Cai, C.; Agarwal, A.; Giovannucci, A.; Kaufman, M.T.; Pandarinath, C. A deep learning framework for inference of single-trial neural population dynamics from calcium imaging with subframe temporal resolution. Nat Neurosci 2022, 25, 1724–1734. [Google Scholar] [CrossRef]
- Gosztolai, A.; Peach, R.L.; Arnaudon, A.; Barahona, M.; Vandergheynst, P. MARBLE: interpretable representations of neural population dynamics using geometric deep learning. Nat Methods 2025, 22, 612–620. [Google Scholar] [CrossRef]
- Angelaki, D.; Benson, B.; Benson, J.; Birman, D.; Bonacchi, N.; Bougrova, K.; Bruijns, S.A.; Carandini, M.; Catarino, J.A.; et al.; International Brain; L A brain-wide map of neural activity during complex behaviour. Nature 2025, 645, 177–191. [Google Scholar] [CrossRef]
- Ajmera, S.; Khan, R.A.; Jain, N.; Kim, G.; Castro, A.; Berenbaum, H.; Husain, F.T. Sparse machine learning of resting-state fMRI reveals brain-wide dysconnectivity in hyperacusis. Commun Med (Lond) 2025, 6, 55. [Google Scholar] [CrossRef]
- Brown, J.A.; Lee, A.J.; Fernhoff, K.; Pistone, T.; Pasquini, L.; Wise, A.B.; Staffaroni, A.M.; Mandelli, M.L.; Lee, S.E.; Boxer, A.L.; et al. Functional network collapse in neurodegenerative disease. Nat Commun 2025, 16, 10273. [Google Scholar] [CrossRef]
- Rassoulou, F.; Sharma, A.; Steina, A.; Butz, M.; Hartmann, C.J.; Bahners, B.H.; Vesper, J.; Schnitzler, A.; Hirschmann, J. Electrophysiological signatures predict the therapeutic window of deep brain stimulation electrode contacts. NPJ Digit Med 2025, 8, 635. [Google Scholar] [CrossRef]
- Waldthaler, J.; Comarovschii, I.; Lundqvist, D. Magnetoencephalography-based prediction of longitudinal symptom progression in Parkinson's disease. NPJ Parkinsons Dis 2026, 12, 29. [Google Scholar] [CrossRef]
- Oby, E.R.; Degenhart, A.D.; Grigsby, E.M.; Motiwala, A.; McClain, N.T.; Marino, P.J.; Yu, B.M.; Batista, A.P. Dynamical constraints on neural population activity. Nat Neurosci 2025, 28, 383–393. [Google Scholar] [CrossRef] [PubMed]
- Merk, T.; Kohler, R.M.; Brotons, T.M.; Vossberg, S.R.; Peterson, V.; Lyra, L.F.; Vanhoecke, J.; Chikermane, M.; Binns, T.S.; Li, N.; et al. Invasive neurophysiology and whole brain connectomics for neural decoding in patients with brain implants. Nat Biomed Eng 2025. [Google Scholar] [CrossRef] [PubMed]
- Topalovic, U.; Barclay, S.; Ling, C.; Alzuhair, A.; Yu, W.; Hokhikyan, V.; Chandrakumar, H.; Rozgic, D.; Jiang, W.; Basir-Kazeruni, S.; et al. A wearable platform for closed-loop stimulation and recording of single-neuron and local field potential activity in freely moving humans. Nat Neurosci 2023, 26, 517–527. [Google Scholar] [CrossRef]
- Tejada-Lapuerta, A.; Schaar, A.C.; Gutgesell, R.; Palla, G.; Halle, L.; Minaeva, M.; Vornholz, L.; Dony, L.; Drummer, F.; Richter, T.; et al. Nicheformer: a foundation model for single-cell and spatial omics. Nat Methods 2025, 22, 2525–2538. [Google Scholar] [CrossRef] [PubMed]
- Khilkevich, A.; Lohse, M.; Low, R.; Orsolic, I.; Bozic, T.; Windmill, P.; Mrsic-Flogel, T.D. Brain-wide dynamics linking sensation to action during decision-making. Nature 2024, 634, 890–900. [Google Scholar] [CrossRef] [PubMed]
- Safaai, H.; Wang, A.Y.; Kira, S.; Blanco Malerba, S.; Panzeri, S.; Harvey, C.D. Specialized structure of neural population codes in parietal cortex outputs. Nat Neurosci 2025, 28, 2550–2560. [Google Scholar] [CrossRef]
- Consortium, M.I. Functional connectomics spanning multiple areas of mouse visual cortex. Nature 2025, 640, 435–447. [Google Scholar] [CrossRef]
- Jin, L.; Sullivan, H.A.; Zhu, M.; Lavin, T.K.; Matsuyama, M.; Fu, X.; Lea, N.E.; Xu, R.; Hou, Y.; Rutigliani, L.; et al. Long-term labeling and imaging of synaptically connected neuronal networks in vivo using double-deletion-mutant rabies viruses. Nat Neurosci 2024, 27, 373–383. [Google Scholar] [CrossRef] [PubMed]
- Rivera, J.F.; Huang, H.; Weng, W.; Sohn, H.; Girasole, A.E.; Li, S.; Albanese, M.A.; Qin, M.; Tao, C.; Klug, M.E.; et al. ATLAS: a rationally designed anterograde transsynaptic tracer. Nat Methods 2025, 22, 1101–1111. [Google Scholar] [CrossRef]
- Shapson-Coe, A.; Januszewski, M.; Berger, D.R.; Pope, A.; Wu, Y.; Blakely, T.; Schalek, R.L.; Li, P.H.; Wang, S.; Maitin-Shepard, J.; et al. A petavoxel fragment of human cerebral cortex reconstructed at nanoscale resolution. Science 2024, 384, eadk4858. [Google Scholar] [CrossRef]
- Vafaii, H.; Mandino, F.; Desrosiers-Gregoire, G.; O'Connor, D.; Markicevic, M.; Shen, X.; Ge, X.; Herman, P.; Hyder, F.; Papademetris, X.; et al. Multimodal measures of spontaneous brain activity reveal both common and divergent patterns of cortical functional organization. Nat Commun 2024, 15, 229. [Google Scholar] [CrossRef]
- Ren, J.; Zhang, W.; Dahmani, L.; Gordon, E.M.; Li, S.; Zhou, Y.; Long, Y.; Huang, J.; Zhu, Y.; Guo, N.; et al. Parkinson's disease as a somato-cognitive action network disorder. Nature 2026, 651, 1030–1038. [Google Scholar] [CrossRef]
- Sumarac, S.; Youn, J.; Fearon, C.; Zivkovic, L.; Keerthi, P.; Flouty, O.; Popovic, M.; Hodaie, M.; Kalia, S.; Lozano, A.; et al. Clinico-physiological correlates of Parkinson's disease from multi-resolution basal ganglia recordings. NPJ Parkinsons Dis 2024, 10, 175. [Google Scholar] [CrossRef]
- Wang, Y.; Jiang, Z.; Chu, C.; Zhang, Z.; Wang, J.; Li, D.; He, N.; Fietkiewicz, C.; Zhou, C.; Kaiser, M.; et al. Push-pull effects of basal ganglia network in Parkinson's disease inferred by functional MRI. NPJ Parkinsons Dis 2024, 10, 224. [Google Scholar] [CrossRef]
- Jamieson, A.J.; Leonards, C.A.; Davey, C.G.; Harrison, B.J. Major depressive disorder associated alterations in the effective connectivity of the face processing network: a systematic review. Transl Psychiatry 2024, 14, 62. [Google Scholar] [CrossRef] [PubMed]
- Myers, J.; Xiao, J.; Mathura, R.K.; Shofty, B.; Gates, V.; Adkinson, J.; Allawala, A.B.; Anand, A.; Gadot, R.; Najera, R.; et al. Intracranial directed connectivity links subregions of the prefrontal cortex to major depression. Nat Commun 2025, 16, 6309. [Google Scholar] [CrossRef] [PubMed]
- Zhang, H.; Li, C.; Shi, K.; Xia, Y.; Song, Y.; Feng, J.; Wang, Z.; Wang, K.; Yang, Y. Early treatment-related changes in dorsolateral prefrontal cortex activity and functional connectivity as potential biomarkers for antidepressant response in major depressive disorder. Transl Psychiatry 2025, 15, 350. [Google Scholar] [CrossRef]
- Chari, A.; Hernan, A.E.; Mahoney, J.M.; Thornton, R.; Tahir, M.Z.; Tisdall, M.M.; Scott, R.C. Single unit-derived connectivity networks in tuberous sclerosis complex reveal propensity for network hypersynchrony driven by tuber-tuber interactions. Sci Rep 2024, 14, 31654. [Google Scholar] [CrossRef]
- Raghavan, M.; Pilet, J.; Carlson, C.; Anderson, C.T.; Mueller, W.; Lew, S.; Ustine, C.; Shah-Basak, P.; Youssofzadeh, V.; Beardsley, S.A. Gamma amplitude-envelope correlations are strongly elevated within hyperexcitable networks in focal epilepsy. Sci Rep 2024, 14, 17736. [Google Scholar] [CrossRef]
- Ereira, S.; Waters, S.; Razi, A.; Marshall, C.R. Early detection of dementia with default-mode network effective connectivity. Nat Ment Health 2024, 2, 787–800. [Google Scholar] [CrossRef]
- Rodemer, W.; Ra, I.; Gujral, J.; Jia, E.; Juul, H.; Zhang, B.; Hoxha, K.; Xing, B.; Mehta, S.; Farag, M.; et al. Network Dysfunction Precedes Neurodegeneration in a dox-Regulatable TDP-43 Mouse Model of ALS-FTD. J Neurosci 2025, 45. [Google Scholar] [CrossRef] [PubMed]
- Yan, Y.; Song, D.; Li, G.; Li, J.; Tang, Y.; Li, D.; Mao, J.; Li, H.; Liu, X.; Yu, D.; et al. The dynamic impairment of synaptic transmission in the PCx-IL engram circuit contributes to early olfactory memory decline in Alzheimer's disease. Mol Psychiatry 2026. [Google Scholar] [CrossRef]
- Akella, S.; Ledochowitsch, P.; Siegle, J.H.; Belski, H.; Denman, D.D.; Buice, M.A.; Durand, S.; Koch, C.; Olsen, S.R.; Jia, X. Deciphering neuronal variability across states reveals dynamic sensory encoding. Nat Commun 2025, 16, 1768. [Google Scholar] [CrossRef]
- Luo, T.Z.; Kim, T.D.; Gupta, D.; Bondy, A.G.; Kopec, C.D.; Elliott, V.A.; DePasquale, B.; Brody, C.D. Transitions in dynamical regime and neural mode during perceptual decisions. Nature 2025, 646, 1156–1166. [Google Scholar] [CrossRef]
- Faust, T.E.; Lee, Y.H.; O'Connor, C.D.; Boyle, M.A.; Gunner, G.; Duran-Laforet, V.; Ferrari, L.L.; Murphy, R.E.; Badimon, A.; Sakers, K.; et al. Microglia-astrocyte crosstalk regulates synapse remodeling via Wnt signaling. Cell 2025, 188, 5212–5230 e5221. [Google Scholar] [CrossRef]
- Kim, E.; Du, H.; Tan, Y.; Brown, B.L.; Chang, Y.; Diaz-Castro, B.; Sweedler, J.V.; Yu, X. Prefrontal cortex astrocytes modulate distinct neuronal populations to control anxiety-like behavior. Nat Commun 2025, 16, 7819. [Google Scholar] [CrossRef]
- Marty-Lombardi, S.; Lu, S.; Ambroziak, W.; Schrenk-Siemens, K.; Wang, J.; DePaoli-Roach, A.A.; Hagenston, A.M.; Wende, H.; Tappe-Theodor, A.; Simonetti, M.; et al. Neuron-astrocyte metabolic coupling facilitates spinal plasticity and maintenance of inflammatory pain. Nat Metab 2024, 6, 494–513. [Google Scholar] [CrossRef]
- Sancho, L.; Boisvert, M.M.; Eddy, T.; Burgado, J.; Contreras, M.; Labarta-Bajo, L.; Wang, E.; Tatsumi, L.; Allen, N.J. Astrocyte CCN1 stabilizes neural circuits in the adult brain. Nature 2026, 649, 948–958. [Google Scholar] [CrossRef] [PubMed]
- Naffaa, M.M. Mechanisms of astrocytic and microglial purinergic signaling in homeostatic regulation and implications for neurological disease. Exploration of Neuroscience 2025, 4. [Google Scholar] [CrossRef]
- Findling, C.; Hubert, F.; Acerbi, L.; Benson, B.; Benson, J.; Birman, D.; Bonacchi, N.; Buchanan, E.K.; Bruijns, S.; et al.; International Brain; L Brain-wide representations of prior information in mouse decision-making. Nature 2025, 645, 192–200. [Google Scholar] [CrossRef] [PubMed]
- Mathis, M.W.; Perez Rotondo, A.; Chang, E.F.; Tolias, A.S.; Mathis, A. Decoding the brain: From neural representations to mechanistic models. Cell 2024, 187, 5814–5832. [Google Scholar] [CrossRef] [PubMed]
- Zhang, Y.; Lyu, H.; Hurwitz, C.; Wang, S.; Findling, C.; Wang, Y.; Hubert, F.; Pouget, A.; Varol, E.; et al.; International Brain; L Exploiting correlations across trials and behavioral sessions to improve neural decoding. Neuron 2026, 114, 536–551 e511. [Google Scholar] [CrossRef]
- Luppi, A.I.; Gellersen, H.M.; Liu, Z.Q.; Peattie, A.R.D.; Manktelow, A.E.; Adapa, R.; Owen, A.M.; Naci, L.; Menon, D.K.; Dimitriadis, S.I.; et al. Systematic evaluation of fMRI data-processing pipelines for consistent functional connectomics. Nat Commun 2024, 15, 4745. [Google Scholar] [CrossRef]
- Safari, M.; Shalbaf, R.; Bagherzadeh, S.; Shalbaf, A. Classification of mental workload using brain connectivity and machine learning on electroencephalogram data. Sci Rep 2024, 14, 9153. [Google Scholar] [CrossRef]
- Bruin, W.B.; Abe, Y.; Alonso, P.; Anticevic, A.; Backhausen, L.L.; Balachander, S.; Bargallo, N.; Batistuzzo, M.C.; Benedetti, F.; Bertolin Triquell, S.; et al. The functional connectome in obsessive-compulsive disorder: resting-state mega-analysis and machine learning classification for the ENIGMA-OCD consortium. Mol Psychiatry 2023, 28, 4307–4319. [Google Scholar] [CrossRef]
- Mocilnik, V.; Rutar Gorisek, V.; Sajovic, J.; Pretnar Oblak, J.; Drevensek, G.; Rogelj, P. Integrating EEG and Machine Learning to Analyze Brain Changes during the Rehabilitation of Broca's Aphasia. Sensors (Basel) 2024, 24. [Google Scholar] [CrossRef]
- Dong, Y.; Wang, S.; Huang, Q.; Berg, R.W.; Li, G.; He, J. Neural Decoding for Intracortical Brain-Computer Interfaces. Cyborg Bionic Syst 2023, 4, 0044. [Google Scholar] [CrossRef] [PubMed]
- Pontes, E.D.; Pinto, M.; Lopes, F.; Teixeira, C. Concept-drifts adaptation for machine learning EEG epilepsy seizure prediction. Sci Rep 2024, 14, 8204. [Google Scholar] [CrossRef] [PubMed]
- Sourmpis, C.; Petersen, C.C.H.; Gerstner, W.; Bellec, G. Biologically informed cortical models predict optogenetic perturbations. Elife 2026, 14. [Google Scholar] [CrossRef] [PubMed]
- Ng, B.; Tasaki, S.; Greathouse, K.M.; Walker, C.K.; Zhang, A.; Covitz, S.; Cieslak, M.; Weber, A.J.; Adamson, A.B.; Andrade, J.P.; et al. Integration across biophysical scales identifies molecular and cellular correlates of person-to-person variability in human brain connectivity. Nat Neurosci 2024, 27, 2240–2252. [Google Scholar] [CrossRef]
- Sharf, T.; van der Molen, T.; Glasauer, S.M.K.; Guzman, E.; Buccino, A.P.; Luna, G.; Cheng, Z.; Audouard, M.; Ranasinghe, K.G.; Kudo, K.; et al. Functional neuronal circuitry and oscillatory dynamics in human brain organoids. Nat Commun 2022, 13, 4403. [Google Scholar] [CrossRef]
- Verduzco Espinoza, A.P.; Na, N.; Campanati, L.; Ngo, P.; Baldwin, K.K.; Cline, H.T. Microglia-to-neuron signaling links APOE4 and inflammation to enhanced neuronal lipid metabolism and network activity. Proc Natl Acad Sci U S A 2025, 122, e2516103122. [Google Scholar] [CrossRef]
- He, X.; Chen, J.; Zhong, Y.; Cen, P.; Shen, L.; Huang, F.; Wang, J.; Jin, C.; Zhou, R.; Zhang, X.; et al. Forebrain neural progenitors effectively integrate into host brain circuits and improve neural function after ischemic stroke. Nat Commun 2025, 16, 5132. [Google Scholar] [CrossRef] [PubMed]
- Rasetto, N.B.; Giacomini, D.; Berardino, A.A.; Waichman, T.V.; Beckel, M.S.; Di Bella, D.J.; Brown, J.; Davies-Sala, M.G.; Gerhardinger, C.; Lie, D.C.; et al. Transcriptional dynamics orchestrating the development and integration of neurons born in the adult hippocampus. Sci Adv 2024, 10, eadp6039. [Google Scholar] [CrossRef] [PubMed]
- Yee, C.; Xiao, Y.; Chen, H.; Reddy, A.R.; Xu, B.; Medwig-Kinney, T.N.; Zhang, W.; Boyle, A.P.; Herbst, W.A.; Xiang, Y.K.; et al. An activity-regulated transcriptional program directly drives synaptogenesis. Nat Neurosci 2024, 27, 1695–1707. [Google Scholar] [CrossRef]
- Naffaa, M.M.; Yin, H.H. A cholinergic signaling pathway underlying cortical circuit activation of quiescent neural stem cells in the lateral ventricle. Sci Signal 2024, 17, eadk8810. [Google Scholar] [CrossRef]
- Naffaa, M.M.; Khan, R.R.; Kuo, C.T.; Yin, H.H. Cortical regulation of neurogenesis and cell proliferation in the ventral subventricular zone. Cell Rep 2023, 42, 112783. [Google Scholar] [CrossRef]
- Chater, T.E.; Eggl, M.F.; Goda, Y.; Tchumatchenko, T. Competitive processes shape multi-synapse plasticity along dendritic segments. Nat Commun 2024, 15, 7572. [Google Scholar] [CrossRef]
- Kim, C.M.; Finkelstein, A.; Chow, C.C.; Svoboda, K.; Darshan, R. Distributing task-related neural activity across a cortical network through task-independent connections. Nat Commun 2023, 14, 2851. [Google Scholar] [CrossRef]
- Theparambil, S.M.; Kopach, O.; Braga, A.; Nizari, S.; Hosford, P.S.; Sagi-Kiss, V.; Hadjihambi, A.; Konstantinou, C.; Esteras, N.; Gutierrez Del Arroyo, A.; et al. Adenosine signalling to astrocytes coordinates brain metabolism and function. Nature 2024, 632, 139–146. [Google Scholar] [CrossRef]
- Heir, R.; Abuzgaya, M.; Adaidi, H.; Franquin, M.; Abbasi, Z.; Stellwagen, D. A screen of immune signaling molecules regulated by neuronal activity identifies interferon-gamma as a modulator of synaptic function and anxiety-like behavior. Brain Behav Immun 2025, 130, 106066. [Google Scholar] [CrossRef]
- Jin, H.; Li, M.; Jeong, E.; Castro-Martinez, F.; Zuker, C.S. A body-brain circuit that regulates body inflammatory responses. Nature 2024, 630, 695–703. [Google Scholar] [CrossRef] [PubMed]
- Song, L.; Chen, W.; Hou, J.; Guo, M.; Yang, J. Spatially resolved mapping of cells associated with human complex traits. Nature 2025, 641, 932–941. [Google Scholar] [CrossRef]
- Naffaa, M.M.; Yin, H.H. Lateral Ventricular Neural Stem Cells Provide Negative Feedback to Circuit Activation Through GABAergic Signaling. Cells 2025, 14. [Google Scholar] [CrossRef]
- Frigerio, I.; Morella, M.L.; Winkelaar, A.B.; Vorgia, D.T.; Breve, J.J.; Boiten, W.A.; van der Gaag, B.L.; Calandri, I.L.; Jonker, A.J.; Berendse, H.W.; et al. Synaptic enrichment of pSer129 alpha-synuclein correlates with dopaminergic denervation in early-stage Parkinson's disease. Nat Commun 2025, 16, 6630. [Google Scholar] [CrossRef] [PubMed]
- Rodriguez-Aller, R.; Romero-Quineche, B.; Morissette, M.; Sheta, R.; Di Paolo, T.; Saghatelyan, A.; Oueslati, A. Optogenetic-induced alpha-synuclein accumulation reveals early synaptic dysfunction in experimental models of Parkinson's disease. NPJ Parkinsons Dis 2025, 11, 356. [Google Scholar] [CrossRef] [PubMed]
- Fagiani, F.; Pedrini, E.; Martire, M.S.; Gastoldi, G.; Vanden Bulcke, C.; Lin, J.P.; Maric, D.; Brambilla, E.; Ruffini, F.; Peri, C.; et al. Spatially-restricted inflammation-induced senescent-like glia in multiple sclerosis and patient-derived organoids. Nat Commun 2025, 16, 8477. [Google Scholar] [CrossRef]
- Miquel-Rio, L.; Jerico-Escolar, J.; Sarries-Serrano, U.; Yanes-Castilla, C.; Torres-Lopez, M.; Argibay, U.; Paz, V.; Casal, C.; Munoz-Moreno, E.; Lopez-Gil, X.; et al. Early synaptic changes and reduced brain connectivity in PD-like mice with depressive phenotype. NPJ Parkinsons Dis 2025, 11, 242. [Google Scholar] [CrossRef]
- Bahl, E.; Chatterjee, S.; Mukherjee, U.; Elsadany, M.; Vanrobaeys, Y.; Lin, L.C.; McDonough, M.; Resch, J.; Giese, K.P.; Abel, T.; et al. Using deep learning to quantify neuronal activation from single-cell and spatial transcriptomic data. Nat Commun 2024, 15, 779. [Google Scholar] [CrossRef] [PubMed]
- Caporale, N.; Castaldi, D.; Rigoli, M.T.; Cheroni, C.; Valenti, A.; Stucchi, S.; Lessi, M.; Bulgheresi, D.; Trattaro, S.; Pezzali, M.; et al. Multiplexing cortical brain organoids for the longitudinal dissection of developmental traits at single-cell resolution. Nat Methods 2025, 22, 358–370. [Google Scholar] [CrossRef]
- Kam, T.I.; Hinkle, J.T.; Dawson, T.M.; Dawson, V.L. Microglia and astrocyte dysfunction in parkinson's disease. Neurobiol Dis 2020, 144, 105028. [Google Scholar] [CrossRef]
- Carrasco-Gomez, M.; Garcia-Colomo, A.; Cabrera-Alvarez, J.; Bruna, R.; Santos, A.; Maestu, F. Dynamics of brain connectivity across the Alzheimer's disease spectrum through magnetoencephalography. Sci Rep 2026, 16, 3905. [Google Scholar] [CrossRef]
- Rohden, F.; Ferreira, P.C.L.; Bellaver, B.; Ferrari-Souza, J.P.; Aguzzoli, C.S.; Soares, C.; Abbas, S.; Zalzale, H.; Povala, G.; Lussier, F.Z.; et al. Glial reactivity correlates with synaptic dysfunction across aging and Alzheimer's disease. Nat Commun 2025, 16, 5653. [Google Scholar] [CrossRef]
- Thergarajan, P.; O'Brien, T.J.; Jones, N.C.; Ali, I. Ligand-receptor interactions: A key to understanding microglia and astrocyte roles in epilepsy. Epilepsy Behav 2025, 163, 110219. [Google Scholar] [CrossRef] [PubMed]
- Zhang, S.; Zhou, Y.; Ren, Y.; Shang, J.; Jiang, A.; Hong, Y.; Hong, S.; Wang, C.; Wang, W.; Chen, J. Microglial activation drives neuronal dysregulation in alcohol-induced seizure susceptibility. Sci Rep 2025, 15, 38389. [Google Scholar] [CrossRef]
- Liang, Q.; Zhou, Z.; Chen, S.; Lin, S.; Lin, X.; Li, Y.; Zhang, Y.; Peng, B.; Hou, G.; Qiu, Y. Symptom-based depression subtypes: brain dynamic specificity and its association with gene expression profiles. Transl Psychiatry 2025, 15, 33. [Google Scholar] [CrossRef] [PubMed]
- Wang, J.; Chen, C.; Zhang, X.; Wang, S.; Hao, R.; Xu, Y.; Fang, N.; Teng, H.; Zhang, B. Symptom-specific neural circuits and corresponding transcriptomic profiles in depression. Behav Brain Res 2026, 500, 115966. [Google Scholar] [CrossRef] [PubMed]
- Garton, T.; Gadani, S.P.; Gill, A.J.; Calabresi, P.A. Neurodegeneration and demyelination in multiple sclerosis. Neuron 2024, 112, 3231–3251. [Google Scholar] [CrossRef]
- Luoma, A.; Matilainen, M.; Tuisku, J.M.; Aarnio, R.; Nikkila, T.; Laaksonen, S.; Koivumaki, M.; Honkonen, E.; Nylund, M.; Wahlroos, S.; et al. Synaptic Density in Multiple Sclerosis: An In Vivo Study Using [(11)C]UCB-J-PET Imaging. Neurol Neuroimmunol Neuroinflamm 2025, 12, e200435. [Google Scholar] [CrossRef]
- Abou-El-Hassan, H.; Yahya, T.; Zusman, B.E.; Albastaki, O.; Imkamp, H.T.; Ye, J.J.; Percopo, F.; Christenson, J.R.; Al Mansi, M.H.; Lu, K.J.; et al. Astrocyte activation persists one year after TBI: a dynamic shift from inflammation to neurodegeneration. Commun Biol 2025, 8, 1745. [Google Scholar] [CrossRef]
- Leskinen, S.; Mehta, N.H.; Shah, H.A.; Quelle, M.; Woodworth, R.; Dituri, G.; Balucani, C.; D'Amico, R.S. Functional Connectivity Changes in Traumatic Brain Injury: A Systematic Review and Coordinate-Based Meta-Analysis of fMRI Studies. Neurology 2025, 105, e214298. [Google Scholar] [CrossRef]
- Kale, M.B.; Wankhede, N.L.; Bishoyi, A.K.; Ballal, S.; Kalia, R.; Arya, R.; Kumar, S.; Khalid, M.; Gulati, M.; Umare, M.; et al. Emerging biophysical techniques for probing synaptic transmission in neurodegenerative disorders. Neuroscience 2025, 565, 63–79. [Google Scholar] [CrossRef]
- Hickey, J.W.; Agmon, E.; Horowitz, N.; Tan, T.K.; Lamore, M.; Sunwoo, J.B.; Covert, M.W.; Nolan, G.P. Integrating multiplexed imaging and multiscale modeling identifies tumor phenotype conversion as a critical component of therapeutic T cell efficacy. Cell Syst 2024, 15, 322–338 e325. [Google Scholar] [CrossRef]
- Shi, Q.; Chen, Y.; Li, Y.; Qin, S.; Yang, Y.; Gao, Y.; Zhu, L.; Wang, D.; Zhang, Z. Cross-tissue multicellular coordination and its rewiring in cancer. Nature 2025, 643, 529–538. [Google Scholar] [CrossRef]
- Wang, E.; Yu, K.; Cao, J.; Wang, M.; Katsel, P.; Song, W.M.; Wang, Z.; Li, Y.; Wang, X.; Wang, Q.; et al. Multiscale proteomic modeling reveals protein networks driving Alzheimer's disease pathogenesis. Cell 2025, 188, 6186–6204 e6113. [Google Scholar] [CrossRef]
- Wang, Q.; Antone, J.; Alsop, E.; Reiman, R.; Funk, C.; Bendl, J.; Dudley, J.T.; Liang, W.S.; Karr, T.L.; Roussos, P.; et al. Single cell transcriptomes and multiscale networks from persons with and without Alzheimer's disease. Nat Commun 2024, 15, 5815. [Google Scholar] [CrossRef]
- Ge, Q.; Wang, Z.; Wang, Y.; Chu, T.; Wang, Z.; Zhu, W.; Zhang, Y.; Chen, Y.; Ye, D.; Xu, W.; et al. Integrative single-cell and spatial transcriptomics with explainable AI reveal lethal prognostic axis in prostate cancer. NPJ Digit Med 2026, 9, 119. [Google Scholar] [CrossRef]
- Klughammer, J.; Abravanel, D.L.; Segerstolpe, A.; Blosser, T.R.; Goltsev, Y.; Cui, Y.; Goodwin, D.R.; Sinha, A.; Ashenberg, O.; Slyper, M.; et al. A multi-modal single-cell and spatial expression map of metastatic breast cancer biopsies across clinicopathological features. Nat Med 2024, 30, 3236–3249. [Google Scholar] [CrossRef]
- Gao, Y.; Ventura-Diaz, S.; Wang, X.; He, M.; Xu, Z.; Weir, A.; Zhou, H.Y.; Zhang, T.; van Duijnhoven, F.H.; Han, L.; et al. An explainable longitudinal multi-modal fusion model for predicting neoadjuvant therapy response in women with breast cancer. Nat Commun 2024, 15, 9613. [Google Scholar] [CrossRef] [PubMed]
- Li, Z.; Li, Y.; Xiang, J.; Wang, X.; Yang, S.; Zhang, X.; Eweje, F.; Chen, Y.; Luo, X.; Li, Y.; et al. AI-enabled virtual spatial proteomics from histopathology for interpretable biomarker discovery in lung cancer. Nat Med 2026, 32, 231–244. [Google Scholar] [CrossRef]
- Mataraso, S.J.; Espinosa, C.A.; Seong, D.; Reincke, S.M.; Berson, E.; Reiss, J.D.; Kim, Y.; Ghanem, M.; Shu, C.H.; James, T.; et al. A machine learning approach to leveraging electronic health records for enhanced omics analysis. Nat Mach Intell 2025, 7, 293–306. [Google Scholar] [CrossRef] [PubMed]
- Jasodanand, V.H.; Kowshik, S.S.; Puducheri, S.; Romano, M.F.; Xu, L.; Au, R.; Kolachalama, V.B. AI-driven fusion of multimodal data for Alzheimer's disease biomarker assessment. Nat Commun 2025, 16, 7407. [Google Scholar] [CrossRef] [PubMed]
- Xue, C.; Kowshik, S.S.; Lteif, D.; Puducheri, S.; Jasodanand, V.H.; Zhou, O.T.; Walia, A.S.; Guney, O.B.; Zhang, J.D.; Poesy, S.; et al. AI-based differential diagnosis of dementia etiologies on multimodal data. Nat Med 2024, 30, 2977–2989. [Google Scholar] [CrossRef]
- Akbarialiabad, H.; Pasdar, A.; Murrell, D.F.; Mostafavi, M.; Shakil, F.; Safaee, E.; Leachman, S.A.; Haghighi, A.; Tarbox, M.; Bunick, C.G.; et al. Enhancing randomized clinical trials with digital twins. NPJ Syst Biol Appl 2025, 11, 110. [Google Scholar] [CrossRef]
- Estrada, H.; Chen, Y.; Lemaire, T.; Davoudi, N.; Ozbek, A.; Parduzi, Q.; Shoham, S.; Razansky, D. Holographic transcranial ultrasound neuromodulation enhances stimulation efficacy by cooperatively recruiting distributed brain circuits. Nat Biomed Eng 2026, 10, 6–15. [Google Scholar] [CrossRef] [PubMed]
- McCallum, S.; Suresh, K.B.; Islam, T.S.; Tripathi, M.K.; Saustad, A.W.; Shelest, O.; Patil, A.; Lee, D.; Kwon, B.; Leitholf, K.; et al. Lesion-remote astrocytes govern microglia-mediated white matter repair. Nature 2026, 649, 959–970. [Google Scholar] [CrossRef]
- Provenza, N.R.; Reddy, S.; Allam, A.K.; Rajesh, S.V.; Diab, N.; Reyes, G.; Caston, R.M.; Katlowitz, K.A.; Gandhi, A.D.; Bechtold, R.A.; et al. Disruption of neural periodicity predicts clinical response after deep brain stimulation for obsessive-compulsive disorder. Nat Med 2024, 30, 3004–3014. [Google Scholar] [CrossRef]
- Yeh, C.Y.; Aguirre, K.; Laveroni, O.; Kim, S.; Wang, A.; Liang, B.; Zhang, X.; Han, L.M.; Valbuena, R.; Bassik, M.C.; et al. Mapping spatial organization and genetic cell-state regulators to target immune evasion in ovarian cancer. Nat Immunol 2024, 25, 1943–1958. [Google Scholar] [CrossRef]
- Gabitto, M.I.; Travaglini, K.J.; Rachleff, V.M.; Kaplan, E.S.; Long, B.; Ariza, J.; Ding, Y.; Mahoney, J.T.; Dee, N.; Goldy, J.; et al. Integrated multimodal cell atlas of Alzheimer's disease. Nat Neurosci 2024, 27, 2366–2383. [Google Scholar] [CrossRef]
- Xiang, H.; Xiao, Y.; Li, F.; Li, C.; Liu, L.; Deng, T.; Yan, C.; Zhou, F.; Wang, X.; Ou, J.; et al. Development and validation of an interpretable model integrating multimodal information for improving ovarian cancer diagnosis. Nat Commun 2024, 15, 2681. [Google Scholar] [CrossRef] [PubMed]
- Yang, P.; Jin, K.; Yao, Y.; Jin, L.; Shao, X.; Li, C.; Lu, X.; Fan, X. Spatial integration of multi-omics single-cell data with SIMO. Nat Commun 2025, 16, 1265. [Google Scholar] [CrossRef] [PubMed]
- Busch, J.L.; Kaplan, J.; Behnke, J.K.; Witzig, V.S.; Drescher, L.; Habets, J.G.V.; Kuhn, A.A. Chronic adaptive deep brain stimulation for Parkinson's disease: clinical outcomes and programming strategies. NPJ Parkinsons Dis 2025, 11, 264. [Google Scholar] [CrossRef]
- De Domenico, M.; Allegri, L.; Caldarelli, G.; d'Andrea, V.; Di Camillo, B.; Rocha, L.M.; Rozum, J.; Sbarbati, R.; Zambelli, F. Challenges and opportunities for digital twins in precision medicine from a complex systems perspective. NPJ Digit Med 2025, 8, 37. [Google Scholar] [CrossRef]
- Sadee, C.; Testa, S.; Barba, T.; Hartmann, K.; Schuessler, M.; Thieme, A.; Church, G.M.; Okoye, I.; Hernandez-Boussard, T.; Hood, L.; et al. Medical digital twins: enabling precision medicine and medical artificial intelligence. Lancet Digit Health 2025, 7, 100864. [Google Scholar] [CrossRef]
- Dehghanghanatkaman, A. Dynamic context-aware multi-modal deep learning for longitudinal prediction of Parkinson's disease progression. Sci Rep 2025, 16, 2104. [Google Scholar] [CrossRef]
- Korbmacher, M.; Lie, I.A.; Wesnes, K.; Westman, E.; Espeseth, T.; Specht, K.; Andreassen, O.A.; Westlye, L.T.; Wergeland, S.; Myhr, K.M.; et al. Multimodal predictors of disability progression and processing speed decline in relapsing-remitting multiple sclerosis. Sci Rep 2025, 15, 36156. [Google Scholar] [CrossRef]
- Shmatko, A.; Jung, A.W.; Gaurav, K.; Brunak, S.; Mortensen, L.H.; Birney, E.; Fitzgerald, T.; Gerstung, M. Learning the natural history of human disease with generative transformers. Nature 2025, 647, 248–256. [Google Scholar] [CrossRef] [PubMed]
- Acosta, J.N.; Falcone, G.J.; Rajpurkar, P.; Topol, E.J. Multimodal biomedical AI. Nat Med 2022, 28, 1773–1784. [Google Scholar] [CrossRef] [PubMed]
- Azarfar, G.; Naimimohasses, S.; Rambhatla, S.; Komorowski, M.; Ferro, D.; Lewis, P.R.; Gates, D.; Shara, N.; Gascon, G.M.; Chang, A.; et al. Responsible adoption of multimodal artificial intelligence in health care: promises and challenges. Lancet Digit Health 2025, 7, 100917. [Google Scholar] [CrossRef]
- Park, W.Y.; Sippel Schmidt, T.; Salvador, G.; O'Donnell, K.; Genereaux, B.; Jeon, K.; You, S.C.; Dewey, B.E.; Nagy, P.; Alzheimer's Disease Neuroimaging, I. Breaking data silos: incorporating the DICOM imaging standard into the OMOP CDM to enable multimodal research. J Am Med Inform Assoc 2025, 32, 1533–1541. [Google Scholar] [CrossRef]
- Lekadir, K.; Frangi, A.F.; Porras, A.R.; Glocker, B.; Cintas, C.; Langlotz, C.P.; Weicken, E.; Asselbergs, F.W.; Prior, F.; Collins, G.S.; et al. FUTURE-AI: international consensus guideline for trustworthy and deployable artificial intelligence in healthcare. BMJ 2025, 388, e081554. [Google Scholar] [CrossRef] [PubMed]
- Mousavi, R.; Mustafa Ali, M.K.; Lobo, D. Discovery of Dynamic Models for AML Disease Progression from Longitudinal Multi-Modal Clinical Data Using Explainable Machine Learning. medRxiv 2025. [Google Scholar] [CrossRef]
- Soladoye, A.A.; Aderinto, N.; Osho, D.; Olawade, D.B. Explainable machine learning models for early Alzheimer's disease detection using multimodal clinical data. Int J Med Inform 2025, 204, 106093. [Google Scholar] [CrossRef]
- Zhang, S.; Cornet, R.; Benis, N. Cross-Standard Health Data Harmonization using Semantics of Data Elements. Sci Data 2024, 11, 1407. [Google Scholar] [CrossRef]
- Zhang, S.; Yuan, J.; Sun, Y.; Wu, F.; Liu, Z.; Zhai, F.; Zhang, Y.; Somekh, J.; Peleg, M.; Zhu, Y.C.; et al. Machine learning on longitudinal multi-modal data enables the understanding and prognosis of Alzheimer's disease progression. iScience 2024, 27, 110263. [Google Scholar] [CrossRef]
- Rood, J.E.; Hupalowska, A.; Regev, A. Toward a foundation model of causal cell and tissue biology with a Perturbation Cell and Tissue Atlas. Cell 2024, 187, 4520–4545. [Google Scholar] [CrossRef]
- Weinstock, J.S.; Arce, M.M.; Freimer, J.W.; Ota, M.; Marson, A.; Battle, A.; Pritchard, J.K. Gene regulatory network inference from CRISPR perturbations in primary CD4(+) T cells elucidates the genomic basis of immune disease. Cell Genom 2024, 4, 100671. [Google Scholar] [CrossRef]
- Chen, V.; Yang, M.; Cui, W.; Kim, J.S.; Talwalkar, A.; Ma, J. Applying interpretable machine learning in computational biology-pitfalls, recommendations and opportunities for new developments. Nat Methods 2024, 21, 1454–1461. [Google Scholar] [CrossRef]
- Wu, X.; Zhang, S.; Zhang, Z.; He, Z.; Xu, Z.; Wang, W.; Jin, Z.; You, J.; Guo, Y.; Zhang, L.; et al. Biologically interpretable multi-task deep learning pipeline predicts molecular alterations, grade, and prognosis in glioma patients. NPJ Precis Oncol 2024, 8, 181. [Google Scholar] [CrossRef] [PubMed]
- Kang, J.; Park, H.J. Integration of partially observed multimodal and multiscale neural signals for estimating a neural circuit using dynamic causal modeling. PLoS Comput Biol 2024, 20, e1012655. [Google Scholar] [CrossRef] [PubMed]
- Sourlos, N.; Vliegenthart, R.; Santinha, J.; Klontzas, M.E.; Cuocolo, R.; Huisman, M.; van Ooijen, P. Recommendations for the creation of benchmark datasets for reproducible artificial intelligence in radiology. Insights Imaging 2024, 15, 248. [Google Scholar] [CrossRef] [PubMed]
- Yang, J.; Dung, N.T.; Thach, P.N.; Phong, N.T.; Phu, V.D.; Phu, K.D.; Yen, L.M.; Thy, D.B.X.; Soltan, A.A.S.; Thwaites, L.; et al. Generalizability assessment of AI models across hospitals in a low-middle and high income country. Nat Commun 2024, 15, 8270. [Google Scholar] [CrossRef]
- Ong Ly, C.; Unnikrishnan, B.; Tadic, T.; Patel, T.; Duhamel, J.; Kandel, S.; Moayedi, Y.; Brudno, M.; Hope, A.; Ross, H.; et al. Shortcut learning in medical AI hinders generalization: method for estimating AI model generalization without external data. NPJ Digit Med 2024, 7, 124. [Google Scholar] [CrossRef]
- Zhu, H.; Bai, J.; Li, N.; Li, X.; Liu, D.; Buckeridge, D.L.; Li, Y. FedWeight: mitigating covariate shift of federated learning on electronic health records data through patients re-weighting. NPJ Digit Med 2025, 8, 286. [Google Scholar] [CrossRef]
- Sokol, K.; Fackler, J.; Vogt, J.E. Artificial intelligence should genuinely support clinical reasoning and decision making to bridge the translational gap. NPJ Digit Med 2025, 8, 345. [Google Scholar] [CrossRef] [PubMed]
- You, J.G.; Hernandez-Boussard, T.; Pfeffer, M.A.; Landman, A.; Mishuris, R.G. Clinical trials informed framework for real world clinical implementation and deployment of artificial intelligence applications. NPJ Digit Med 2025, 8, 107. [Google Scholar] [CrossRef]
- Finster, M.; Wenzel, M.; Taghizadeh, E. Common data models and data standards for tabular health data: a systematic review. BMC Med Inform Decis Mak 2025, 25, 422. [Google Scholar] [CrossRef]
- Gilani, K.; Wei, W.; Peters, C.; Verket, M.; Rocca, H.B.; Nicolis, E.; Colombo, M.; Marx-Schutt, K.; Urovi, V.; Dumontier, M. CMEO: a metadata-centric ontology for clinical studies exploration and harmonization assessment. BMC Med Inform Decis Mak 2025, 26, 8. [Google Scholar] [CrossRef]
- Wang, E.H.; Myles, P.; Foraker, R.; Toh, S.; Mosquera, L.; El Emam, K.; Burcu, M. Crossing borders securely: synthetic data and federated networks for privacy-preserving access to real-world data and emerging use cases. NPJ Digit Med 2025, 8, 758. [Google Scholar] [CrossRef] [PubMed]
- Welzel, C.; Ostermann, M.; Smith, H.L.; Minssen, T.; Kirsten, T.; Gilbert, S. Enabling secure and self determined health data sharing and consent management. NPJ Digit Med 2025, 8, 560. [Google Scholar] [CrossRef] [PubMed]
- Alderman, J.E.; Palmer, J.; Laws, E.; McCradden, M.D.; Ordish, J.; Ghassemi, M.; Pfohl, S.R.; Rostamzadeh, N.; Cole-Lewis, H.; Glocker, B.; et al. Tackling algorithmic bias and promoting transparency in health datasets: the STANDING Together consensus recommendations. Lancet Digit Health 2025, 7, e64–e88. [Google Scholar] [CrossRef]
- Hasanzadeh, F.; Josephson, C.B.; Waters, G.; Adedinsewo, D.; Azizi, Z.; White, J.A. Bias recognition and mitigation strategies in artificial intelligence healthcare applications. NPJ Digit Med 2025, 8, 154. [Google Scholar] [CrossRef]
- Badani, A.; de Moraes, F.Y.; Vollmuth, P.; Chung, C.; Mansouri, A. AI and innovation in clinical trials. NPJ Digit Med 2025, 8, 683. [Google Scholar] [CrossRef] [PubMed]
- Wells, B.J.; Nguyen, H.M.; McWilliams, A.; Pallini, M.; Bovi, A.; Kuzma, A.; Kramer, J.; Chou, S.H.; Hetherington, T.; Corn, P.; et al. A practical framework for appropriate implementation and review of artificial intelligence (FAIR-AI) in healthcare. NPJ Digit Med 2025, 8, 514. [Google Scholar] [CrossRef] [PubMed]
| Domain | Data modality | Method / AI approach | Primary analytical purpose | What it can reveal | Main limitations | References |
|---|---|---|---|---|---|---|
| Cellular state profiling | Single-cell RNA sequencing (scRNA-seq) | Dimensionality reduction, deep latent embedding, autoencoders, variational autoencoders | Representation of cellular heterogeneity and state structure | Cellular subpopulations, rare states, gradients of cellular variation, latent state organization | Sparsity, batch effects, snapshot nature of data, limited direct temporal information | [64,65] |
| Cellular state profiling | Single-cell ATAC-seq / epigenomic profiling | Multimodal integration, latent variable models, matrix factorization, deep generative models | Characterization of chromatin-defined cell states and regulatory landscapes | Regulatory programs, chromatin accessibility dynamics, epigenetic constraints on state transitions | Sparse signal, noisy data, integration challenges with transcriptomic modalities | [66,67] |
| Cellular state profiling | Single-cell proteomic / multimodal assays | Joint embedding, multimodal deep learning, probabilistic integration | Integration of protein and transcript features for refined cellular state definition | Concordance or divergence between transcriptional and protein states, functional phenotype refinement | Limited feature depth, cross-platform variability, incomplete multimodal correspondence | [68,69] |
| Spatial state mapping | Spatial transcriptomics | Graph-based integration, spatially aware deep learning, neighborhood modeling | Mapping cellular states within tissue architecture | Spatial organization of states, tissue niches, local cellular interactions, state–environment relationships | Resolution limits, mixed spots in some platforms, complex spatial normalization | [57,70] |
| Spatial state mapping | Multiplexed imaging / spatial proteomics | Graph neural networks, image-based feature extraction, spatial clustering | Identification of state distributions and cell–cell interactions in situ | Tissue microenvironment structure, signaling neighborhoods, spatial coupling of phenotypes | Imaging noise, segmentation errors, limited multiplexing in some platforms | [54,71] |
| Trajectory reconstruction | Cross-sectional single-cell datasets | Trajectory inference, pseudotime modeling, diffusion-based methods, graph trajectory models | Reconstruction of developmental or disease-related state transitions | Differentiation pathways, branching trajectories, transitional intermediates, candidate progression routes | Pseudotime may not reflect true biological time, sensitivity to preprocessing and sampling density | [51,52] |
| Lineage and transition modeling | Lineage tracing + omics integration | Probabilistic lineage inference, graph modeling, multimodal temporal alignment | Reconstruction of lineage relationships and directional transitions | Cell fate relationships, clonal dynamics, validation of inferred trajectories | Experimental complexity, incomplete lineage capture, integration difficulty across modalities | [27,51] |
| Cellular interaction modeling | Single-cell + spatial neighborhood data | Graph neural networks, cell–cell interaction inference, network learning | Modeling relationships among cells and their microenvironments | Local signaling structure, interaction networks, microenvironmental regulation of state transitions | Dependency on graph construction choices, uncertainty in inferred interactions | [54,72] |
| Neural population decoding | Calcium imaging | Supervised learning, deep neural networks, temporal decoding models | Decoding behavioral or cognitive variables from neural activity | Population-level activity patterns, behavioral state prediction, stimulus representation | Indirect measurement of activity, temporal resolution constraints, signal preprocessing dependencies | [73] |
| Neural population decoding | Electrophysiological recordings | Supervised decoding, recurrent neural networks, state-space models | Mapping neural firing patterns to behavior, cognition, or circuit states | High-temporal-resolution representations of neural dynamics, transitions between functional states | Limited sampling of full networks, variability across sessions and individuals | [74,75] |
| Brain-wide functional analysis | fMRI | Machine learning classifiers, representation learning, connectivity modeling | Identification of large-scale functional signatures and disease-associated patterns | Functional connectivity structure, distributed network states, disease classification signals | Indirect hemodynamic signal, limited temporal resolution, susceptibility to confounds | [76,77] |
| Brain-wide functional analysis | EEG / MEG | Time-series classification, spectral feature learning, deep temporal models | Characterization of neural dynamics across time and disease states | Oscillatory signatures, transient network states, seizure-related or disease-related patterns | Noise sensitivity, source localization ambiguity, inter-subject variability | [78,79] |
| Circuit modeling | Neural population time series / multimodal neural data | Recurrent neural networks, dynamical systems models, latent state models | Simulation and prediction of circuit dynamics over time | Temporal state transitions, network stability, responses to perturbation, hidden dynamic structure | Limited biological interpretability, possible mismatch between model structure and circuit biology | [74,80] |
| Closed-loop circuit analysis | Real-time neural recordings | Adaptive decoding, reinforcement learning, closed-loop control algorithms | Real-time detection of circuit states and adaptive intervention | Online state estimation, seizure detection, adaptive stimulation targets, responsive modulation | Generalizability, safety, interpretability, dependence on real-time signal quality | [81,82] |
| Multiscale integration | Single-cell, spatial, synaptic, circuit, and clinical datasets | Multimodal deep learning, shared latent space models, cross-modal alignment, integrative graph frameworks | Linking biological information across scales | Relationships between cellular state transitions, tissue organization, circuit dynamics, and clinical outcomes | Heterogeneous data structures, temporal misalignment, incomplete sampling, limited causal interpretability | [58,83] |
| Disease context | Cellular state changes | Synaptic / microcircuit alterations | Circuit-level dysfunction | Behavioral / clinical consequence | Relevance to multiscale modeling | References |
|---|---|---|---|---|---|---|
| Parkinson’s disease | Degeneration of dopaminergic neurons; altered neuronal stress-response and metabolic states; reactive astrocytic and microglial changes | Altered nigrostriatal and corticostriatal synaptic transmission; disturbed inhibitory–excitatory balance within basal ganglia microcircuits | Abnormal basal ganglia network activity, disrupted oscillatory dynamics, and impaired motor circuit coordination | Bradykinesia, rigidity, tremor, and progressive motor dysfunction | Illustrates how neuronal state changes propagate through synaptic and microcircuit disruption to large-scale motor circuit abnormalities and clinical symptoms | [134,140] |
| Alzheimer’s disease | Neuronal stress and degeneration; microglial activation; astrocytic reactivity; altered inflammatory and homeostatic cellular states | Synaptic loss, impaired synaptic plasticity, and disruption of local hippocampal and cortical microcircuits | Progressive disruption of large-scale network connectivity and reduced functional integration across memory-related circuits | Memory impairment, cognitive decline, and progressive loss of executive function | Demonstrates how glial and neuronal state transitions interact with synaptic degeneration and network disorganization during disease progression | [141,142] |
| Epilepsy | Altered neuronal excitability states; reactive astrocytes and microglia; inflammatory and metabolic shifts in local tissue environments | Enhanced excitatory transmission, impaired inhibitory control, maladaptive synaptic remodeling, and destabilized local microcircuits | Pathological hypersynchrony, recurrent seizure-generating network states, and impaired circuit stability | Recurrent seizures, cognitive impairment, and variable neuropsychiatric symptoms | Highlights how cellular and inflammatory state changes can reduce network stability and drive recurrent pathological circuit transitions | [143,144] |
| Depression | Altered neuronal and glial functional states; stress-associated transcriptional changes; impaired cellular resilience; neuroimmune dysregulation | Reduced synaptic plasticity, altered local connectivity within limbic and prefrontal microcircuits, and disturbed neuromodulatory balance | Dysregulated limbic–prefrontal circuit activity and altered functional connectivity in mood-related networks | Persistent low mood, anhedonia, cognitive slowing, and affective dysregulation | Supports a model in which stress-related cellular changes scale upward to synaptic and circuit dysfunction underlying behavioral symptoms | [145,146] |
| Neuroinflammatory disorders | Immune cell activation; reactive microglial and astrocytic states; altered neuronal homeostasis under inflammatory conditions | Cytokine-driven modulation of synaptic transmission, impaired synaptic stability, and disrupted local circuit interactions | Abnormal network activity, reduced circuit adaptability, and impaired functional integration across affected regions | Cognitive impairment, sensory or motor dysfunction, fatigue, and neurobehavioral abnormalities | Emphasizes the role of immune–neural coupling in linking inflammatory cellular states to synaptic and circuit-level dysfunction | [147,148] |
| Traumatic brain injury / injury-related disorders | Neuronal stress responses; glial activation; altered repair-associated and inflammatory cellular programs | Synaptic disruption, loss of local connectivity, maladaptive plasticity, and unstable microcircuit reorganization | Impaired network communication, abnormal circuit reconfiguration, and altered activity propagation | Cognitive deficits, sensory–motor impairments, mood changes, and persistent neurological dysfunction | Illustrates how acute cellular injury responses can evolve into persistent multiscale dysfunction affecting circuit organization and behavior | [149,150] |
| Neurodegenerative disorders broadly | Progressive neuronal vulnerability states; glial activation; metabolic and proteostatic dysregulation | Synaptic weakening or loss, impaired local plasticity, and deterioration of microcircuit integrity | Large-scale circuit disconnection, reduced adaptability, and abnormal functional dynamics | Cognitive, motor, and behavioral decline depending on the affected systems | Provides a general framework for understanding disease progression as coupled transitions across cellular, synaptic, and network scales | [77,151] |
| Challenge | Significance | Current limitations | Future opportunities | Potential clinical implications | References |
|---|---|---|---|---|---|
| Data heterogeneity across modalities | Multiscale disease modeling depends on integrating molecular, cellular, circuit, imaging, and clinical datasets that differ substantially in format, scale, and resolution | Data are generated using different platforms, preprocessing pipelines, feature spaces, and quality standards, limiting comparability and joint analysis | Development of harmonized preprocessing workflows, cross-platform integration methods, and shared multimodal data standards | More robust biomarker discovery, improved generalizability of predictive models, and stronger cross-study reproducibility | [183] |
| Temporal misalignment across biological scales | Disease progression unfolds over time, but cellular, synaptic, circuit, and clinical data are often collected at different and nonaligned timepoints | Cross-sectional designs and uneven sampling make it difficult to reconstruct temporal relationships and state transitions across scales | Longitudinal multimodal study designs, time-aware computational models, and improved temporal alignment strategies | Earlier detection of disease transitions, better prediction of progression, and improved identification of therapeutic windows | [158,184] |
| Limited causal inference | Predictive associations across scales are informative, but mechanistic understanding requires distinguishing correlation from causation | Many AI models infer statistical relationships from observational data without resolving directional or mechanistic dependencies | Integration of AI with perturbation experiments, causal inference frameworks, and mechanistically informed modeling | More reliable target identification and greater confidence in intervention strategies | [185,186] |
| Model interpretability | Scientific and clinical adoption depends on understanding how model outputs relate to biological mechanisms and disease processes | Complex deep learning models may achieve strong performance while remaining difficult to interpret biologically | Development of interpretable AI methods, biologically constrained architectures, and explanation tools linked to experimental validation | Improved clinician trust, clearer biological insight, and stronger translational potential | [187,188] |
| Incomplete multiscale coverage | Disease processes often span levels that are incompletely sampled, leaving important gaps between molecular, cellular, synaptic, circuit, and clinical domains | Many studies capture only one or two biological levels, limiting the construction of integrated disease models | Expanded multimodal datasets spanning multiple scales, including synaptic, microcircuit, and longitudinal clinical measurements | More complete disease modeling and better linkage between mechanistic biology and patient outcomes | [157,189] |
| External validation and reproducibility | Models that perform well in one dataset or institution may not generalize to independent cohorts or real-world settings | Independent validation is often limited, and results may be sensitive to site-specific protocols, cohort composition, and analytic choices | Multi-center validation studies, benchmark datasets, transparent reporting practices, and reproducible computational pipelines | Greater reliability of AI tools in biomedical research and stronger readiness for clinical deployment | [190,191] |
| Generalizability across patient populations | Multiscale AI models must perform across diverse individuals, disease stages, and healthcare environments | Many models are trained on restricted cohorts that do not adequately represent biological, demographic, or clinical diversity | Inclusion of more diverse patient populations, stratified evaluation, and federated or distributed learning approaches | Fairer and more broadly applicable models for diagnosis, prognosis, and therapeutic guidance | [192,193] |
| Clinical workflow integration | Even accurate models have limited value if they do not align with clinical workflows, decision timing, and interpretability needs | Model outputs are often not presented in formats that support routine clinical use or time-sensitive decision-making | Development of clinician-facing decision-support systems, workflow-aware interfaces, and prospective implementation studies | Greater adoption in practice, improved decision support, and more effective patient management | [194,195] |
| Standardization of multimodal data infrastructure | Scalable multiscale medicine requires interoperable data ecosystems across research and healthcare settings | Fragmented databases, inconsistent metadata, and lack of shared ontologies hinder integration, reuse, and comparison across studies | Shared repositories, interoperable metadata standards, and common data models across modalities and institutions | Faster model development, stronger collaboration, and improved reproducibility across settings | [196,197] |
| Ethical, legal, and privacy considerations | Multiscale AI models often rely on sensitive, longitudinal, and potentially identifiable datasets, raising governance and trust concerns | Risks include insufficient consent frameworks, privacy breaches, data misuse, and limited clarity around governance responsibilities | Privacy-preserving learning, transparent governance structures, and ethically informed data-sharing frameworks | More responsible implementation, stronger public trust, and safer clinical adoption | [198,199] |
| Algorithmic bias and equity | Unequal representation in training data can produce systematic underperformance for certain populations | Bias may arise from cohort selection, measurement variability, social determinants of health, and healthcare system disparities | Bias auditing, fairness-aware modeling, representative datasets, and continuous monitoring after deployment | Reduced inequities in AI-supported diagnosis, prognosis, and treatment recommendations | [200,201] |
| Prospective clinical translation | Clinical value depends on demonstrating benefit in real-world patient care rather than only retrospective performance | Many AI models remain proof-of-concept and lack prospective evaluation in clinically relevant environments | Prospective clinical studies, adaptive evaluation frameworks, and integration with patient outcome analyses | Stronger evidence for clinical utility and improved translation from computational modeling to therapeutic benefit | [202,203] |
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2026 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).




