Submitted:
21 April 2026
Posted:
22 April 2026
You are already at the latest version
Abstract
Keywords:
Introduction
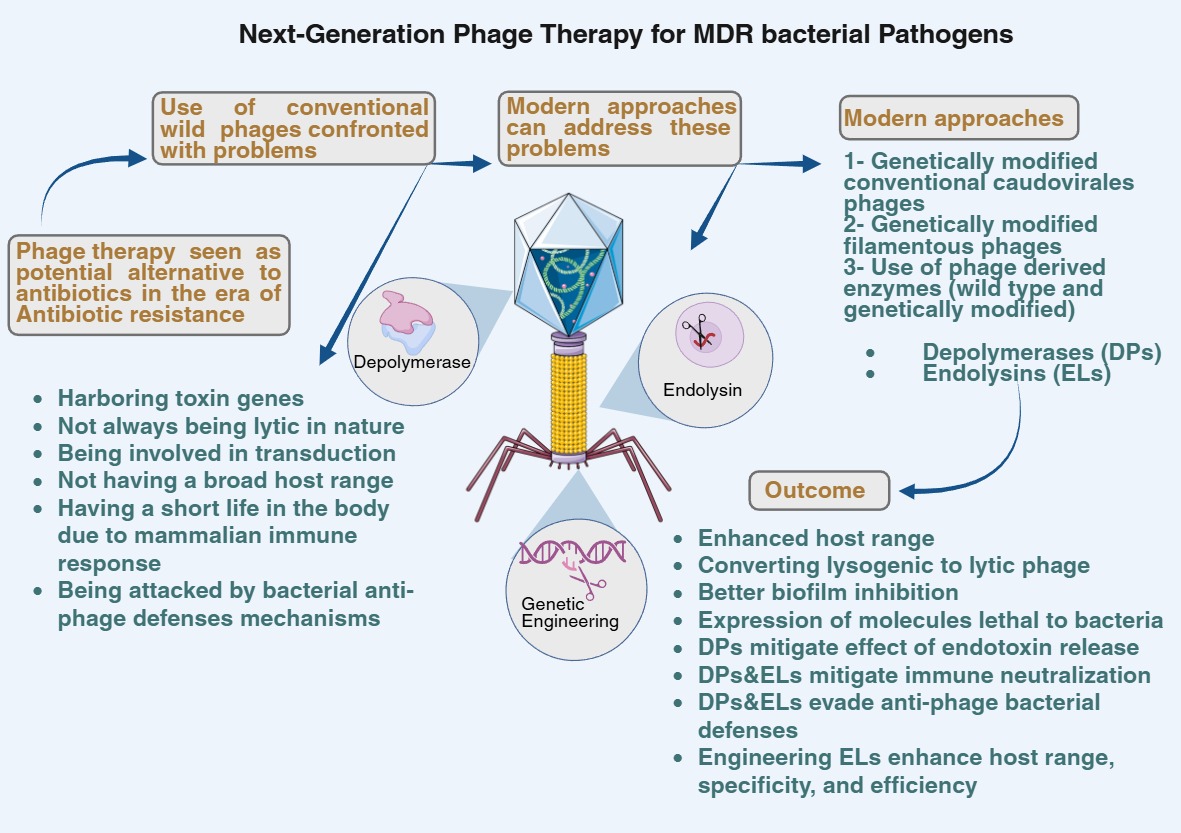
Advantages and limitations of phage therapy
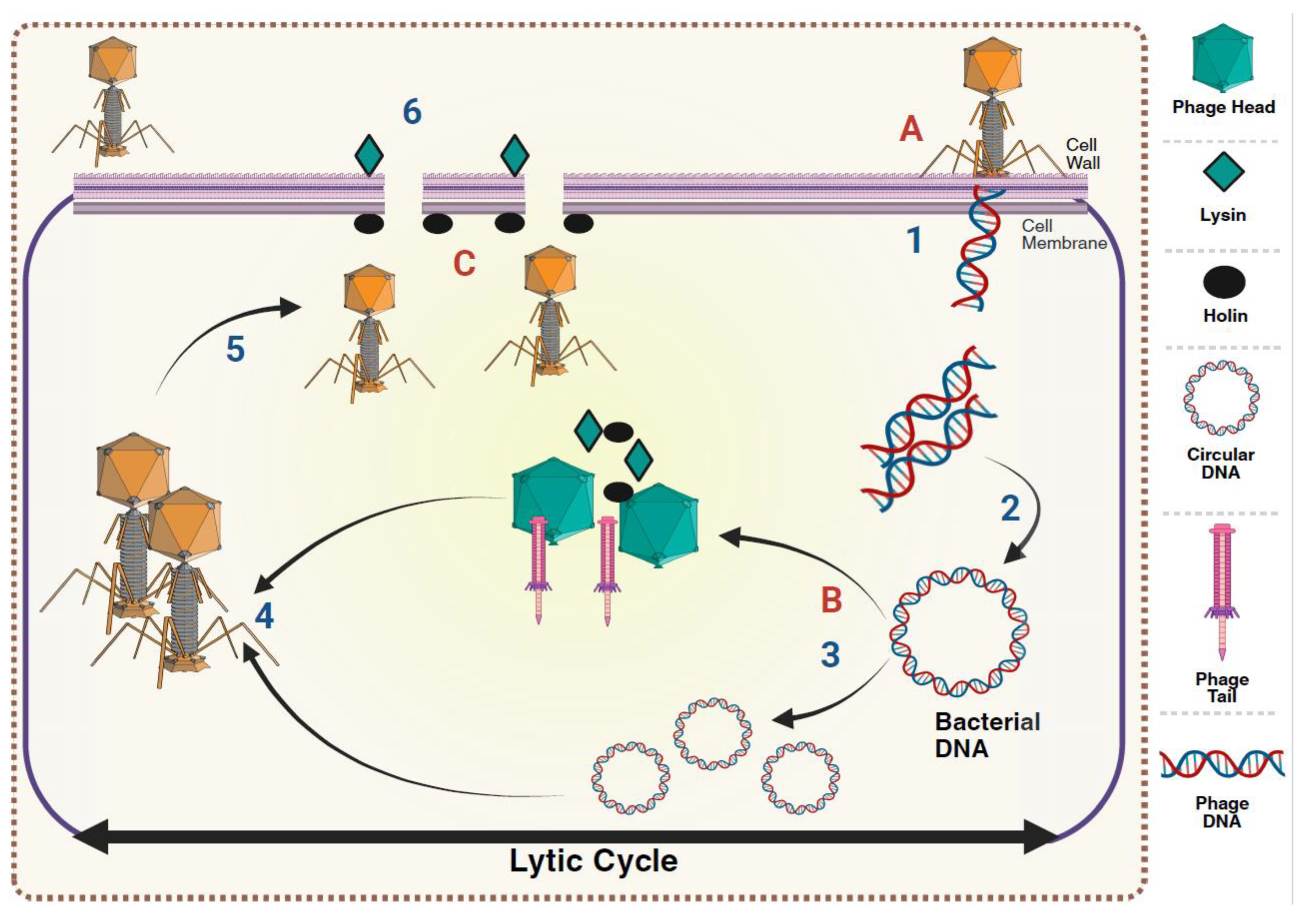
Life cycle of phages and window for genetic modifications
Genetic engineering of non-filamentous phages for use in phage therapy
Genetic engineering and potential use of filamentous phages for phage therapy
Phage-derived enzymes as potential alternatives to antibiotics
Depolymerases as anti-infective agents
Engineered endolysins as anti-infective
Conclusions
Author Contributions
Funding
Acknowledgments
Conflicts of interest
Abbreviations
| AMP | antimicrobial peptides |
| CBD | wall binding domain |
| CPS | Capsular polysaccharide |
| EAD | enzymatically active domain |
| EPS | Exopolysaccharide |
| LAB | Lactic acid bacteria |
| LPS | Lipopolysaccharide |
| MRSA | Methicillin Resistant S. aureus |
| Psa | Pseudomonas syringae pv. actinidiae |
| RBPs | Receptor binding proteins |
| SMPA | sheep myeloid 29-amino acid peptide |
| VALs | Virion-associated lysins |
References
- Elfadadny A, Ragab RF, Abou Shehata MA, Elfadadny MR, Farag A, Abd El-Aziz AH, et al. Exploring bacteriophage applications in medicine and beyond. Acta microbiol hell. 2024;69:167-79. [CrossRef]
- Sahoo K, Meshram S, Sahoo Jr K. The Evolution of Phage Therapy: A Comprehensive Review of Current Applications and Future Innovations. Cureus. 2024;16:e70414. [CrossRef]
- Grigson SR, Giles SK, Edwards RA, Papudeshi B. Knowing and naming: phage annotation and nomenclature for phage therapy. Clin Infect Dis. 2023;77:S352-S9. [CrossRef]
- Guo Z, Yuan M, Chai J. Mini review advantages and limitations of lytic phages compared with chemical antibiotics to combat bacterial infections. Heliyon. 2024;10:e34849. [CrossRef]
- Mobarezi Z, Esfandiari AH, Abolbashari S, Meshkat Z. Efficacy of phage therapy in controlling staphylococcal biofilms: a systematic review. Eur J Med Res. 2025;30:605. [CrossRef]
- Diallo K, Dublanchet A. Benefits of combined phage–antibiotic therapy for the control of antibiotic-resistant bacteria: A literature review. Antibiotics. 2022;11:839. [CrossRef]
- Merabishvili M, Pirnay J-P, De Vos D. Guidelines to compose an ideal bacteriophage cocktail. Bacteriophage Therapy: From Lab to Clinical Practice. 2018;1693:99-110. [CrossRef]
- Fernández L, Gutiérrez D, García P, Rodríguez A. The perfect bacteriophage for therapeutic applications—a quick guide. Antibiotics. 2019;8:126. [CrossRef]
- Nayab S, Idrees K, Aslam MA. Synergism of phages and antimicrobial peptides for treating multidrug resistant bacterial pathogens. Explor Drug Sci. 2025;3:1008133. [CrossRef]
- Keen EC, Dantas G. Close encounters of three kinds: bacteriophages, commensal bacteria, and host immunity. Trends Microbiol. 2018;26:943-54. [CrossRef]
- Ingmer H, Gerlach D, Wolz C. Temperate phages of Staphylococcus aureus. Microbiol Spectr. 2019;7:10.1128/microbiolspec. gpp3-0058-2018. [CrossRef]
- Schicklmaier P, Schmieger H. Frequency of generalized transducing phages in natural isolates of the Salmonella typhimurium complex. Appl Environ Microbiol. 1995;61:1637-40. [CrossRef]
- Penadés JR, Chen J, Quiles-Puchalt N, Carpena N, Novick RP. Bacteriophage-mediated spread of bacterial virulence genes. Curr Opin Microbiol. 2015;23:171-8. [CrossRef]
- Chiang YN, Penadés JR, Chen J. Genetic transduction by phages and chromosomal islands: the new and noncanonical. PLoS Pathog. 2019;15:e1007878. [CrossRef]
- Bertozzi Silva J, Storms Z, Sauvageau D. Host receptors for bacteriophage adsorption. FEMS Microbiol Lett. 2016;363:fnw002. [CrossRef]
- Doron S, Melamed S, Ofir G, Leavitt A, Lopatina A, Keren M, et al. Systematic discovery of antiphage defense systems in the microbial pangenome. Science. 2018;359:eaar4120. [CrossRef]
- Ofir G, Sorek R. Contemporary phage biology: from classic models to new insights. Cell. 2018;172:1260-70. [CrossRef]
- Bernheim A, Sorek R. The pan-immune system of bacteria: antiviral defence as a community resource. Nature Reviews Microbiology. 2020;18:113-9. [CrossRef]
- Hampton HG, Watson BN, Fineran PC. The arms race between bacteria and their phage foes. Nature. 2020;577:327-36. [CrossRef]
- Rostøl JT, Marraffini L. (Ph) ighting phages: how bacteria resist their parasites. Cell Host and Microbe. 2019;25:184-94. [CrossRef]
- Leprince A, Mahillon J. Phage adsorption to gram-positive bacteria. Viruses. 2023;15:196. [CrossRef]
- Samir S. Molecular machinery of the triad Holin, Endolysin, and Spanin: key players orchestrating bacteriophage-induced cell lysis and their therapeutic applications. Protein Peptide Lett. 2024;31:85-96. [CrossRef]
- Pirnay J-P, De Vos D, Verbeken G, Merabishvili M, Chanishvili N, Vaneechoutte M, et al. The phage therapy paradigm: prêt-à-porter or sur-mesure? Pharm Res. 2011;28:934-7. [CrossRef]
- Cui Z, Guo X, Feng T, Li L. Exploring the whole standard operating procedure for phage therapy in clinical practice. J Transl Med. 2019;17:1-7. [CrossRef]
- Gibson SB, Green SI, Liu CG, Salazar KC, Clark JR, Terwilliger AL, et al. Constructing and characterizing bacteriophage libraries for phage therapy of human infections. Front Microbiol. 2019;10:2537. [CrossRef]
- Friman VP, Soanes-Brown D, Sierocinski P, Molin S, Johansen HK, Merabishvili M, et al. Pre-adapting parasitic phages to a pathogen leads to increased pathogen clearance and lowered resistance evolution with Pseudomonas aeruginosa cystic fibrosis bacterial isolates. J Evol Biol. 2016;29:188-98. [CrossRef]
- Mahichi F, Synnott AJ, Yamamichi K, Osada T, Tanji Y. Site-specific recombination of T2 phage using IP008 long tail fiber genes provides a targeted method for expanding host range while retaining lytic activity. FEMS Microbiol Lett. 2009;295:211-7. [CrossRef]
- Ando H, Lemire S, Pires DP, Lu TK. Engineering modular viral scaffolds for targeted bacterial population editing. Cell systems. 2015;1:187-96. [CrossRef]
- Yehl K, Lemire S, Yang AC, Ando H, Mimee M, Torres MDT, et al. Engineering phage host-range and suppressing bacterial resistance through phage tail fiber mutagenesis. Cell. 2019;179:459-69. e9. [CrossRef]
- Yoichi M, Abe M, Miyanaga K, Unno H, Tanji Y. Alteration of tail fiber protein gp38 enables T2 phage to infect Escherichia coli O157: H7. J Biotechnol. 2005;115:101-7. [CrossRef]
- Lynch KH, Seed KD, Stothard P, Dennis JJ. Inactivation of Burkholderia cepacia complex phage KS9 gp41 identifies the phage repressor and generates lytic virions. J Virol. 2010;84:1276-88. [CrossRef]
- Kilcher S, Studer P, Muessner C, Klumpp J, Loessner MJ. Cross-genus rebooting of custom-made, synthetic bacteriophage genomes in L-form bacteria. Proc Natl Acad Sci. 2018;115:567-72. [CrossRef]
- Zhang H, Fouts D, DePew J, Stevens R. Genetic modifications to temperate Enterococcus faecalis phage ϕEf11 that abolish the establishment of lysogeny and sensitivity to repressor, and increase host range and productivity of lytic infection. Microbiology. 2013;159:1023-35. [CrossRef]
- Dedrick RM, Guerrero-Bustamante CA, Garlena RA, Russell DA, Ford K, Harris K, et al. Engineered bacteriophages for treatment of a patient with a disseminated drug-resistant Mycobacterium abscessus. Nat Med. 2019;25:730-3. [CrossRef]
- Lu TK, Collins JJ. Dispersing biofilms with engineered enzymatic bacteriophage. Proc Natl Acad Sci. 2007;104:11197-202. [CrossRef]
- Born Y, Fieseler L, Thöny V, Leimer N, Duffy B, Loessner MJ. Engineering of bacteriophages Y2:: dpoL1-C and Y2:: luxAB for efficient control and rapid detection of the fire blight pathogen, Erwinia amylovora. Appl Environ Microbiol. 2017;83:e00341-17. [CrossRef]
- Pei R, Lamas-Samanamud GR. Inhibition of biofilm formation by T7 bacteriophages producing quorum-quenching enzymes. Appl Environ Microbiol. 2014;80:5340-8. [CrossRef]
- Paul VD, Sundarrajan S, Rajagopalan SS, Hariharan S, Kempashanaiah N, Padmanabhan S, et al. Lysis-deficient phages as novel therapeutic agents for controlling bacterial infection. BMC Microbiol. 2011;11:1-9. [CrossRef]
- Matsuda T, Freeman TA, Hilbert DW, Duff M, Fuortes M, Stapleton PP, et al. Lysis-deficient bacteriophage therapy decreases endotoxin and inflammatory mediator release and improves survival in a murine peritonitis model. Surgery. 2005;137:639-46. [CrossRef]
- Hagens S, Habel A, Von Ahsen U, Von Gabain A, Bläsi U. Therapy of experimental Pseudomonas infections with a nonreplicating genetically modified phage. Antimicrob Agents Chemother. 2004;48:3817-22. [CrossRef]
- Møller-Olsen C, Ho SFS, Shukla RD, Feher T, Sagona AP. Engineered K1F bacteriophages kill intracellular Escherichia coli K1 in human epithelial cells. Sci Rep. 2018;8:17559. [CrossRef]
- Mai-Prochnow A, Hui JGK, Kjelleberg S, Rakonjac J, McDougald D, Rice SA. Big things in small packages: the genetics of filamentous phage and effects on fitness of their host. FEMS Microbiol Rev. 2015;39:465-87. [CrossRef]
- Rakonjac J, Russel M, Khanum S, Brooke SJ, Rajič M. Filamentous phage: structure and biology. Recombinant antibodies for infectious diseases. 2017;1053:1-20. [CrossRef]
- Yacoby I, Shamis M, Bar H, Shabat D, Benhar I. Targeting antibacterial agents by using drug-carrying filamentous bacteriophages. Antimicrob Agents Chemother. 2006;50:2087-97. [CrossRef]
- Yang Y, Kang D, Mihalache B, Vexler S, Jain S, Peng H, et al. Metagenome-inspired libraries to engineer phage M13 for targeted killing of Gram-negative bacterial species. Nucleic Acids Res. 2025;53:gkaf984. [CrossRef]
- Krom RJ, Bhargava P, Lobritz MA, Collins JJ. Engineered phagemids for nonlytic, targeted antibacterial therapies. Nano Lett. 2015;15:4808-13. [CrossRef]
- Westwater C, Kasman LM, Schofield DA, Werner PA, Dolan JW, Schmidt MG, et al. Use of genetically engineered phage to deliver antimicrobial agents to bacteria: an alternative therapy for treatment of bacterial infections. Antimicrob Agents Chemother. 2003;47:1301-7. [CrossRef]
- Hagens S, Bläsi U. Genetically modified filamentous phage as bactericidal agents: a pilot study. Lett Appl Microbiol. 2003;37:318-23. [CrossRef]
- Lu TK, Collins JJ. Engineered bacteriophage targeting gene networks as adjuvants for antibiotic therapy. Proc Natl Acad Sci. 2009;106:4629-34. [CrossRef]
- Bhattarai SR, Yoo SY, Lee S-W, Dean D. Engineered phage-based therapeutic materials inhibit Chlamydia trachomatis intracellular infection. Biomaterials. 2012;33:5166-74. [CrossRef]
- Smith HW, Huggins MB, Shaw KM. Factors influencing the survival and multiplication of bacteriophages in calves and in their environment. Microbiology. 1987;133:1127-35. [CrossRef]
- Dabrowska K, Switała-Jelen K, Opolski A, Weber-Dabrowska B, Gorski A. Bacteriophage penetration in vertebrates. J Appl Microbiol. 2005;98:7-13. [CrossRef]
- Cisek AA, Dąbrowska I, Gregorczyk KP, Wyżewski Z. Phage therapy in bacterial infections treatment: one hundred years after the discovery of bacteriophages. Curr Microbiol. 2017;74:277-83. [CrossRef]
- Maciejewska B, Olszak T, Drulis-Kawa Z. Applications of bacteriophages versus phage enzymes to combat and cure bacterial infections: an ambitious and also a realistic application? Appl Microbiol Biotechnol. 2018;102:2563-81. [CrossRef]
- Lewis K. Platforms for antibiotic discovery. Nat Rev Drug Discov. 2013;12:371-87. 10.1038/nrd3975.
- Drulis-Kawa Z, Majkowska-Skrobek G, Maciejewska B. Bacteriophages and phage-derived proteins–application approaches. Curr Med Chem. 2015;22:1757-73. [CrossRef]
- Olszak T, Shneider MM, Latka A, Maciejewska B, Browning C, Sycheva LV, et al. The O-specific polysaccharide lyase from the phage LKA1 tailspike reduces Pseudomonas virulence. Sci Rep. 2017;7:16302. [CrossRef]
- Zou X, Xiao X, Mo Z, Ge Y, Jiang X, Huang R, et al. Systematic strategies for developing phage resistant Escherichia coli strains. Nat commun. 2022;13:4491. [CrossRef]
- Latka A, Maciejewska B, Majkowska-Skrobek G, Briers Y, Drulis-Kawa Z. Bacteriophage-encoded virion-associated enzymes to overcome the carbohydrate barriers during the infection process. Appl Microbiol Biotechnol. 2017;101:3103-19. [CrossRef]
- Chen Y, Wang S, Wang Y, Zhangxiang L, Chen H, Li X, et al. Complete genome sequence of the novel phage vB_EcoS_PHB17, which infects Shiga-toxin-producing Escherichia coli. Arch Virol. 2019;164:3111-3. [CrossRef]
- Ma YX, Wang CY, Li YY, Li J, Wan QQ, Chen JH, et al. Considerations and caveats in combating ESKAPE pathogens against nosocomial infections. Advanced Science. 2020;7:1901872. [CrossRef]
- Pires DP, Oliveira H, Melo LD, Sillankorva S, Azeredo J. Bacteriophage-encoded depolymerases: their diversity and biotechnological applications. Appl Microbiol Biotechnol. 2016;100:2141-51. [CrossRef]
- Lin H, Paff ML, Molineux IJ, Bull JJ. Antibiotic therapy using phage depolymerases: robustness across a range of conditions. Viruses. 2018;10:622. [CrossRef]
- Chen Y, Sun E, Yang L, Song J, Wu B. Therapeutic application of bacteriophage PHB02 and its putative depolymerase against Pasteurella multocida capsular type A in mice. Front Microbiol. 2018;9:1678. [CrossRef]
- Lin H, Paff ML, Molineux IJ, Bull JJ. Therapeutic application of phage capsule depolymerases against K1, K5, and K30 capsulated E. coli in mice. Front Microbiol. 2017;8:2257. [CrossRef]
- Lin T-L, Hsieh P-F, Huang Y-T, Lee W-C, Tsai Y-T, Su P-A, et al. Isolation of a bacteriophage and its depolymerase specific for K1 capsule of Klebsiella pneumoniae: implication in typing and treatment. J Infect Dis. 2014;210:1734-44. [CrossRef]
- Chen Y, Li X, Wang S, Guan L, Li X, Hu D, et al. A novel tail-associated O91-specific polysaccharide depolymerase from a podophage reveals lytic efficacy of Shiga toxin-producing Escherichia coli. Appl Environ Microbiol. 2020;86:e00145-20. [CrossRef]
- Olsen NM, Thiran E, Hasler T, Vanzieleghem T, Belibasakis GN, Mahillon J, et al. Synergistic removal of static and dynamic Staphylococcus aureus biofilms by combined treatment with a bacteriophage endolysin and a polysaccharide depolymerase. Viruses. 2018;10:438. [CrossRef]
- Wu Y, Wang R, Xu M, Liu Y, Zhu X, Qiu J, et al. A novel polysaccharide depolymerase encoded by the phage SH-KP152226 confers specific activity against multidrug-resistant Klebsiella pneumoniae via biofilm degradation. Front Microbiol. 2019;10:2768. [CrossRef]
- Latka A, Drulis-Kawa Z. Advantages and limitations of microtiter biofilm assays in the model of antibiofilm activity of Klebsiella phage KP34 and its depolymerase. Sci Rep. 2020;10:20338. [CrossRef]
- Husain FM, Zahra A, Ali A, Kamthan M, Al-Shabib NA, Farooqui Z, et al. Bacteriophages and Their Enzymes: Allies Against Microbial Biofilms. Pharmaceuticals. 2025;18:1771. [CrossRef]
- Wang I-N, Smith DL, Young R. Holins: the protein clocks of bacteriophage infections. Annu Rev Microbiol. 2000;54:799-825. [CrossRef]
- Loessner MJ. Bacteriophage endolysins—current state of research and applications. Curr Opin Microbiol. 2005;8:480-7. [CrossRef]
- Abedon ST. Lysis from without. Bacteriophage. 2011;1:46-9. [CrossRef]
- Yang H, Yu J, Wei H. Engineered bacteriophage lysins as novel anti-infectives. Front Microbiol. 2014;5:542. [CrossRef]
- Seijsing J, Sobieraj AM, Keller N, Shen Y, Zinkernagel AS, Loessner MJ, et al. Improved biodistribution and extended serum half-life of a bacteriophage endolysin by albumin binding domain fusion. Front Microbiol. 2018;9:2927. [CrossRef]
- Yan G, Yang R, Fan K, Dong H, Gao C, Wang S, et al. External lysis of Escherichia coli by a bacteriophage endolysin modified with hydrophobic amino acids. Amb Express. 2019;9:106. [CrossRef]
- Briers Y, Walmagh M, Van Puyenbroeck V, Cornelissen A, Cenens W, Aertsen A, et al. Engineered endolysin-based “Artilysins” to combat multidrug-resistant gram-negative pathogens. MBio. 2014;5:10.1128/mbio. 01379-14. [CrossRef]
- Mancoš M, Šramková Z, Peterková D, Vidová B, Godány A. Functional expression and purification of tailor-made chimeric endolysin with the broad antibacterial spectrum. Biologia. 2020;75:2031-43. [CrossRef]
- Hong H-W, Kim YD, Jang J, Kim MS, Song M, Myung H. Combination effect of engineered endolysin EC340 with antibiotics. Front Microbiol. 2022;13:821936. [CrossRef]
- Islam MM, Kim D, Kim K, Park S-J, Akter S, Kim J, et al. Engineering of lysin by fusion of antimicrobial peptide (cecropin A) enhances its antibacterial properties against multidrug-resistant Acinetobacter baumannii. Front Microbiol. 2022;13:988522. [CrossRef]
- Abdelkader K, Gutierrez D, Tames-Caunedo H, Ruas-Madiedo P, Safaan A, Khairalla AS, et al. Engineering a Lysin with Intrinsic Antibacterial Activity (LysMK34) by Cecropin A Fusion Enhances Its Antibacterial Properties against Acinetobacter baumannii. Appl Environ Microbiol. 2022;88:e0151521. [CrossRef]
- Briers Y, Walmagh M, Grymonprez B, Biebl M, Pirnay J-P, Defraine V, et al. Art-175 is a highly efficient antibacterial against multidrug-resistant strains and persisters of Pseudomonas aeruginosa. Antimicrob Agents Chemother. 2014;58:3774-84. [CrossRef]
- Basit A, Qadir S, Qureshi S, Rehman SU. Cloning and expression analysis of fused holin-endolysin from RL bacteriophage; Exhibits broad activity against multi drug resistant pathogens. Enzyme Microb Technol. 2021;149:109846. [CrossRef]
- Heselpoth RD, Euler CW, Schuch R, Fischetti VA. Lysocins: bioengineered antimicrobials that deliver lysins across the outer membrane of Gram-negative bacteria. Antimicrob Agents Chemother. 2019;63:10.1128/aac. 00342-19. [CrossRef]
- Lukacik P, Barnard TJ, Keller PW, Chaturvedi KS, Seddiki N, Fairman JW, et al. Structural engineering of a phage lysin that targets Gram-negative pathogens. Proc Natl Acad Sci. 2012;109:9857-62. [CrossRef]
- Yan G, Liu J, Ma Q, Zhu R, Guo Z, Gao C, et al. The N-terminal and central domain of colicin A enables phage lysin to lyse Escherichia coli extracellularly. Antonie Van Leeuwenhoek. 2017;110:1627-35. [CrossRef]
- Oliveira H, Melo LD, Santos SB, Nóbrega FL, Ferreira EC, Cerca N, et al. Molecular aspects and comparative genomics of bacteriophage endolysins. J Virol. 2013;87:4558-70. [CrossRef]
- Walmagh M, Briers Y, Santos SBd, Azeredo J, Lavigne R. Characterization of modular bacteriophage endolysins from Myoviridae phages OBP, 201φ2-1 and PVP-SE1. PLoS ONE. 2012;7:e36991. [CrossRef]
- Briers Y, Schmelcher M, Loessner MJ, Hendrix J, Engelborghs Y, Volckaert G, et al. The high-affinity peptidoglycan binding domain of Pseudomonas phage endolysin KZ144. Biochem Biophys Res Commun. 2009;383:187-91. [CrossRef]
- Díez-Martínez R, De Paz HD, García-Fernández E, Bustamante N, Euler CW, Fischetti VA, et al. A novel chimeric phage lysin with high in vitro and in vivo bactericidal activity against Streptococcus pneumoniae. J Antimicrob Chemother. 2015;70:1763-73. [CrossRef]
- Becker SC, Foster-Frey J, Stodola AJ, Anacker D, Donovan DM. Differentially conserved staphylococcal SH3b_5 cell wall binding domains confer increased staphylolytic and streptolytic activity to a streptococcal prophage endolysin domain. Gene. 2009;443:32-41. [CrossRef]
- Dong Q, Wang J, Yang H, Wei C, Yu J, Zhang Y, et al. Construction of a chimeric lysin Ply187N-V12C with extended lytic activity against staphylococci and streptococci. Microbial biotechnology. 2015;8:210-20. [CrossRef]
- Fernandes S, Proença D, Cantante C, Silva FA, Leandro C, Lourenço S, et al. Novel chimerical endolysins with broad antimicrobial activity against methicillin-resistant Staphylococcus aureus. Microb Drug Resist. 2012;18:333-43. [CrossRef]
- Mao J, Schmelcher M, Harty WJ, Foster-Frey J, Donovan DM. Chimeric Ply187 endolysin kills Staphylococcus aureus more effectively than the parental enzyme. FEMS Microbiol Lett. 2013;342:30-6. [CrossRef]
- Landlinger C, Tisakova L, Oberbauer V, Schwebs T, Muhammad A, Latka A, et al. Engineered phage endolysin eliminates Gardnerella biofilm without damaging beneficial bacteria in bacterial vaginosis ex vivo. Pathogens. 2021;10:54. [CrossRef]
- Daniel A, Euler C, Collin M, Chahales P, Gorelick KJ, Fischetti VA. Synergism between a novel chimeric lysin and oxacillin protects against infection by methicillin-resistant Staphylococcus aureus. Antimicrob Agents Chemother. 2010;54:1603-12. [CrossRef]
- Donovan DM, Dong S, Garrett W, Rousseau GM, Moineau S, Pritchard DG. Peptidoglycan hydrolase fusions maintain their parental specificities. Appl Environ Microbiol. 2006;72:2988-96. [CrossRef]
- Behera M, Singh G, Vats A, Roshan M, Gautam D, Rana C, et al. Expression and characterization of novel chimeric endolysin CHAPk-SH3bk against biofilm-forming methicillin-resistant Staphylococcus aureus. Int J Biol Macromol. 2024;254:127969. [CrossRef]
- Behera M, Singh P, Verma AK, De S, Ghorai SM. Activity of GS-linked chimeric endolysin CHAPk-SH3bk against methicillin-resistant Staphylococcus aureus biofilms: an in-vitro, ex-vivo and in-vivo study. npj Biofilms and Microbiomes. 2025;11:94. [CrossRef]
- Momen S, Soleimani N, Azizmohseni F, Ahmadbeigi Y, Borhani S, Amini-Bayat Z. Characterization and bioinformatic analysis of a new chimeric endolysin against MRSA with great stability. AMB Express. 2024;14:143. [CrossRef]
- Wang S, Gu J, Lv M, Guo Z, Yan G, Yu L, et al. The antibacterial activity of E. coli bacteriophage lysin lysep3 is enhanced by fusing the Bacillus amyloliquefaciens bacteriophage endolysin binding domain D8 to the C-terminal region. J Microbiol. 2017;55:403-8. [CrossRef]
- Cheng Q, Fischetti VA. Mutagenesis of a bacteriophage lytic enzyme PlyGBS significantly increases its antibacterial activity against group B streptococci. Appl Microbiol Biotechnol. 2007;74:1284-91. [CrossRef]
- Mayer MJ, Garefalaki V, Spoerl R, Narbad A, Meijers R. Structure-based modification of a Clostridium difficile-targeting endolysin affects activity and host range. J Bacteriol. 2011;193:5477-86. [CrossRef]
- Dahal RH, Choi Y-J, Kim S, Kim J. CHAPSAP26-161, A Truncated Protein and Enzymatic Active Domain of Endolysin LysSAP26, as a Potential Therapeutic Agent to Combat Clostridioides difficile Infection. J Bacteriol Virol. 2024;54:107-21. [CrossRef]
- Wang Y, Wang X, Liu X, Lin B. Research progress on strategies for improving the enzyme properties of bacteriophage endolysins. J Microbiol Biotechnol. 2024;34:1189. [CrossRef]
- Schmelcher M, Tchang VS, Loessner MJ. Domain shuffling and module engineering of Listeria phage endolysins for enhanced lytic activity and binding affinity. Microb Biotechnol. 2011;4:651-62. [CrossRef]
- Rodríguez-Rubio L, Martínez B, Rodríguez A, Donovan DM, García P. Enhanced staphylolytic activity of the Staphylococcus aureus bacteriophage vB_SauS-phiIPLA88 HydH5 virion-associated peptidoglycan hydrolase: fusions, deletions, and synergy with LysH5. Appl Environ Microbiol. 2012;78:2241-8. [CrossRef]
- Díez-Martínez R, de Paz H, Bustamante N, García E, Menéndez M, García P. Improving the lethal effect of Cpl-7, a pneumococcal phage lysozyme with broad bactericidal activity, by inverting the net charge of its cell wall-binding module. Antimicrob Agents Chemother. 2013;57:5355-65. [CrossRef]
- Love MJ, Coombes D, Manners SH, Abeysekera GS, Billington C, Dobson RC. The molecular basis for Escherichia coli O157: H7 phage FAHEc1 endolysin function and protein engineering to increase thermal stability. Viruses. 2021;13:1101. [CrossRef]
- Jansson MK, Nguyen DT, Mikkat S, Warnke C, Janssen MB, Warnke P, et al. Synthetic mRNA delivered to human cells leads to expression of Cpl-1 bacteriophage-endolysin with activity against Streptococcus pneumoniae. Mol Ther Nucleic Acids. 2024;35:102145. [CrossRef]
- Gervasi T, Horn N, Wegmann U, Dugo G, Narbad A, Mayer MJ. Expression and delivery of an endolysin to combat Clostridium perfringens. Appl Microbiol Biotechnol. 2014;98:2495-505. [CrossRef]
- Chandran C, Tham HY, Rahim RA, Lim SHE, Yusoff K, Song AA-L. Lactococcus lactis secreting phage lysins as a potential antimicrobial against multi-drug resistant Staphylococcus aureus. PeerJ. 2022;10:e12648. [CrossRef]
- Dzuvor CK, Shanbhag BK, Younas T, Shen H-H, Haritos VS, He L. Engineering self-assembled endolysin nanoparticles against antibiotic-resistant bacteria. ACS Applied Bio Materials. 2022;5:4993-5003. [CrossRef]
- Nobrega FL, Costa AR, Santos JF, Siliakus MF, Van Lent JW, Kengen SW, et al. Genetically manipulated phages with improved pH resistance for oral administration in veterinary medicine. Sci Rep. 2016;6:39235. [CrossRef]
- Peng H, Borg RE, Dow LP, Pruitt BL, Chen IA. Controlled phage therapy by photothermal ablation of specific bacterial species using gold nanorods targeted by chimeric phages. Proc Natl Acad Sci. 2020;117:1951-61. [CrossRef]
- Gattuboyena N, Tsai Y-C, Lin L-C. Therapeutic and Diagnostic Potential of a Novel K1 Capsule Dependent Phage, JSSK01, and Its Depolymerase in Multidrug-Resistant Escherichia coli Infections. Int J Mol Sci. 2024;25:12497. [CrossRef]
- Napolitano V, Privitera M, Drulis-Kawa Z, Marasco D, Fallarini S, Berisio R, et al. Structural and functional features of Klebsiella pneumoniae capsular degradation by the phage depolymerase KP32gp38: Implications for vaccination against K. pneumoniae. Int J Antimicrob Agents. 2025:107596. [CrossRef]
- Chen Y, Fan Z, Fu T, Li Z, Feng J, Cui X, et al. Characterization of a Phage-Encoded Depolymerase Against Klebsiella pneumoniae K30 Capsular Type and Its Therapeutic Application in a Murine Model of Aspiration Pneumonia. Viruses. 2025;17:1446. [CrossRef]
- Li P, Guo G, Zheng X, Xu S, Zhou Y, Qin X, et al. Therapeutic efficacy of a K5-specific phage and depolymerase against Klebsiella pneumoniae in a mouse model of infection. Vet Res. 2024;55:59. [CrossRef]
- Wang R, Liu Y, Zhang Y, Yu S, Zhuo H, Huang Y, et al. Identification and characterization of the capsule depolymerase Dpo27 from phage IME-Ap7 specific to Acinetobacter pittii. Front Cell Infect Microbiol. 2024;14:1373052. [CrossRef]
- Borzilov AI, Volozhantsev NV, Korobova OV, Kolupaeva LV, Pereskokova ES, Kombarova TI, et al. Bacteriophage and phage-encoded depolymerase exhibit antibacterial activity against K9-type Acinetobacter baumannii in mouse sepsis and burn skin infection models. Viruses. 2025;17:70. [CrossRef]
- Yang P, Shan B, Hu X, Xue L, Song G, He P, et al. Identification of a novel phage depolymerase against ST11 K64 carbapenem-resistant Klebsiella pneumoniae and its therapeutic potential. J Bacteriol. 2025;207:e00387-24. [CrossRef]
- Zhao J, Wang J, Zhang C, Xu S, Ren H, Zou L, et al. Characterization of a Salmonella abortus equi phage 4FS1 and its depolymerase. Frontiers in Veterinary Science. 2024;11:1496684. [CrossRef]
- Zelmer A, Martin MJ, Gundogdu O, Birchenough G, Lever R, Wren BW, et al. Administration of capsule-selective endosialidase E minimizes upregulation of organ gene expression induced by experimental systemic infection with Escherichia coli K1. Microbiology. 2010;156:2205-15. [CrossRef]
- Waseh S, Hanifi-Moghaddam P, Coleman R, Masotti M, Ryan S, Foss M, et al. Orally administered P22 phage tailspike protein reduces Salmonella colonization in chickens: prospects of a novel therapy against bacterial infections. PLoS ONE. 2010;5:e13904. [CrossRef]
- Li M, Li P, Chen L, Guo G, Xiao Y, Chen L, et al. Identification of a phage-derived depolymerase specific for KL64 capsule of Klebsiella pneumoniae and its anti-biofilm effect. Virus Genes. 2021;57:434-42. [CrossRef]
- V. Volozhantsev N, M. Shpirt A, I. Borzilov A, V. Komisarova E, M. Krasilnikova V, S. Shashkov A, et al. Characterization and therapeutic potential of bacteriophage-encoded polysaccharide depolymerases with β galactosidase activity against Klebsiella pneumoniae K57 capsular type. Antibiotics. 2020;9:732. [CrossRef]
- Chen X, Liu M, Zhang P, Xu M, Yuan W, Bian L, et al. Phage-derived depolymerase as an antibiotic adjuvant against multidrug-resistant Acinetobacter baumannii. Frontiers in microbiology. 2022;13:845500. [CrossRef]
- Liu Y, Mi Z, Mi L, Huang Y, Li P, Liu H, et al. Identification and characterization of capsule depolymerase Dpo48 from Acinetobacter baumannii phage IME200. PeerJ. 2019;7:e6173. [CrossRef]
- Rice CJ, Kelly SA, O’Brien SC, Melaugh EM, Ganacias JC, Chai ZH, et al. Novel phage-derived depolymerase with activity against Proteus mirabilis biofilms. Microorganisms. 2021;9:2172. [CrossRef]
- Tung CW, Thapa K, Phan A, Mohapatra A, Hashmi M, Bleich K, et al. Antibacterial Activity of a Fused Endolysin ENDO-1252/KL9P Against Multiple Serovars of Salmonella enterica. Microbiol Biotechnol. 2025;18:e70237. [CrossRef]
- Kogawa M, Yoda T, Matsuhashi A, Matsushita A, Otsuka Y, Shibagaki S, et al. Development of chimera AMP–Endolysin with wider spectra against Gram-Negative Bacteria using High-Throughput assay. Viruses. 2025;17:200. [CrossRef]
- Kim D, Kim J, Kim M. Potential of antimicrobial peptide-fused endolysin LysC02 as therapeutics for infections and disinfectants for food contact surfaces to control Cronobacter sakazakii. Food Control. 2024;157:110190. [CrossRef]
- Lim J, Myung H, Lim D, Song M. Antimicrobial peptide thanatin fused endolysin PA90 (Tha-PA90) for the control of Acinetobacter baumannii infection in mouse model. J Biomed Sci. 2024;31:36. [CrossRef]
- Antonova N, Grigoriev I, Lendel A, Usacheva O, Klimova A, Usachev E, et al. Engineering of recombinant endolysin LysSi3 to increase its antibacterial properties. Appl Biochem Microbiol. 2024;60:802-11. [CrossRef]
- Vasina DV, Antonova NP, Gushchin VA, Aleshkin AV, Fursov MV, Fursova AD, et al. Development of novel antimicrobials with engineered endolysin LysECD7-SMAP to combat Gram-negative bacterial infections. J Biomed Sci. 2024;31:75. [CrossRef]
- Antonova NP, Abdullaeva SD, Vasina DV, Grigoriev IV, Usachev EV, Usacheva OV, et al. Modifying Pharmacokinetic Properties of the Gram-Negative Bacteria Targeting Endolysin ML06 Without Affecting Antibacterial Activity. Int J Mol Sci. 2025;26:4376. [CrossRef]
- Wang Y, Li H, Cui Y, Li Y, Yu Y, Zhang X, et al. Enhancing food system protection: synergistic action of engineered endolysin LYSMP and berberine hydrochloride against Streptococcus suis on pork. Food Res Int. 2025;221:117214. [CrossRef]
- Li X, Shangguan W, Yang X, Hu X, Li Y, Zhao W, et al. Influence of lipopolysaccharide-interacting peptides fusion with endolysin LysECD7 and fatty acid derivatization on the efficacy against Acinetobacter baumannii infection in vitro and in vivo. Viruses. 2024;16:760. [CrossRef]
- Rani R, Poria V, Jaglan AB, Kapoor P, Bhutani K, Sharma P, et al. Improved lytic action of engineered phage-encoded endolysin and docking insights into its action on bacterial peptidoglycan. Int J Biol Macromol. 2025:146105. [CrossRef]
- Amiryan A, Sannikova E, Serkina A, Gubaidullin I, Bulushova N, Kozlov D. Optimization of the Structure of the Staphylococcus aureus Phage K Endolysin CHAP Domain to Increase Lytic Activity. Appl Biochem Microbiol. 2024;60:1565-74. [CrossRef]
- Easwaran M, Govindaraj RG, Naderi M, Brylinski M, De Zoysa M, Shin H-J. Evaluating the antibacterial activity of engineered phage ФEcSw endolysin against multi-drug-resistant Escherichia coli strain Sw1. Int J Antimicrob Agents. 2025;65:107395. [CrossRef]
- Mortazavi N, Aliakbarlu J, Imani M. The antibacterial effects of engineered Salmonella phage PVP-SE1 endolysin on the planktonic cells and biofilms of food-borne pathogens and its antibacterial activity in milk. Food Bioscience. 2025:107274. [CrossRef]
- Park W, Park M, Chun J, Hwang J, Kim S, Choi N, et al. Delivery of endolysin across outer membrane of Gram-negative bacteria using translocation domain of botulinum neurotoxin. Int J Antimicrob Agents. 2024;64:107216. [CrossRef]
- Chen W, Han L-M, Chen X-Z, Yi P-C, Li H, Ren Y-Y, et al. Engineered endolysin of Klebsiella pneumoniae phage is a potent and broad-spectrum bactericidal agent against “ESKAPEE” pathogens. Front Microbiol. 2024;15:1397830. [CrossRef]
- Krishnan M, Tham HY, Wan Nur Ismah WAK, Yusoff K, Song AA-L. Effect of Domain Manipulation in the Staphylococcal Phage Endolysin, Endo88, on Lytic Efficiency and Host Range. Mol Biotechnol. 2025;67:2536-44. [CrossRef]
- Warring SL, Sisson HM, Randall G, Grimon D, Dams D, Gutiérrez D, et al. Engineering an antimicrobial chimeric endolysin that targets the phytopathogen Pseudomonas syringae pv. actinidiae. J Biol Chem. 2025:110224. [CrossRef]
- Munetomo S, Uchiyama J, Takemura-Uchiyama I, Wanganuttara T, Yamamoto Y, Tsukui T, et al. Examination of yield, bacteriolytic activity and cold storage of linker deletion mutants based on endolysin S6_ORF93 derived from Staphylococcus giant bacteriophage S6. PLoS ONE. 2024;19:e0310962. [CrossRef]
- Lee J, Cho H, Kim K-s. Surface-displaying protein from Lacticaseibacillus paracasei–derived extracellular vesicles: Identification and utilization in the fabrication of an endolysin-displaying platform against Staphylococcus aureus. Chem Eng J. 2025;512:162196. [CrossRef]
- Sáez Moreno D, Cunha J, Melo LDR, Tanaka K, Bamba T, Hasunuma T, et al. CRISPR-Cas9 engineered Saccharomyces cerevisiae for endolysin delivery to combat Listeria monocytogenes. Appl Microbiol Biotechnol. 2025; 109(1), 81. [CrossRef]
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2026 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).



