Submitted:
20 April 2026
Posted:
21 April 2026
You are already at the latest version
Abstract
Keywords:
1. Introduction
2. Results
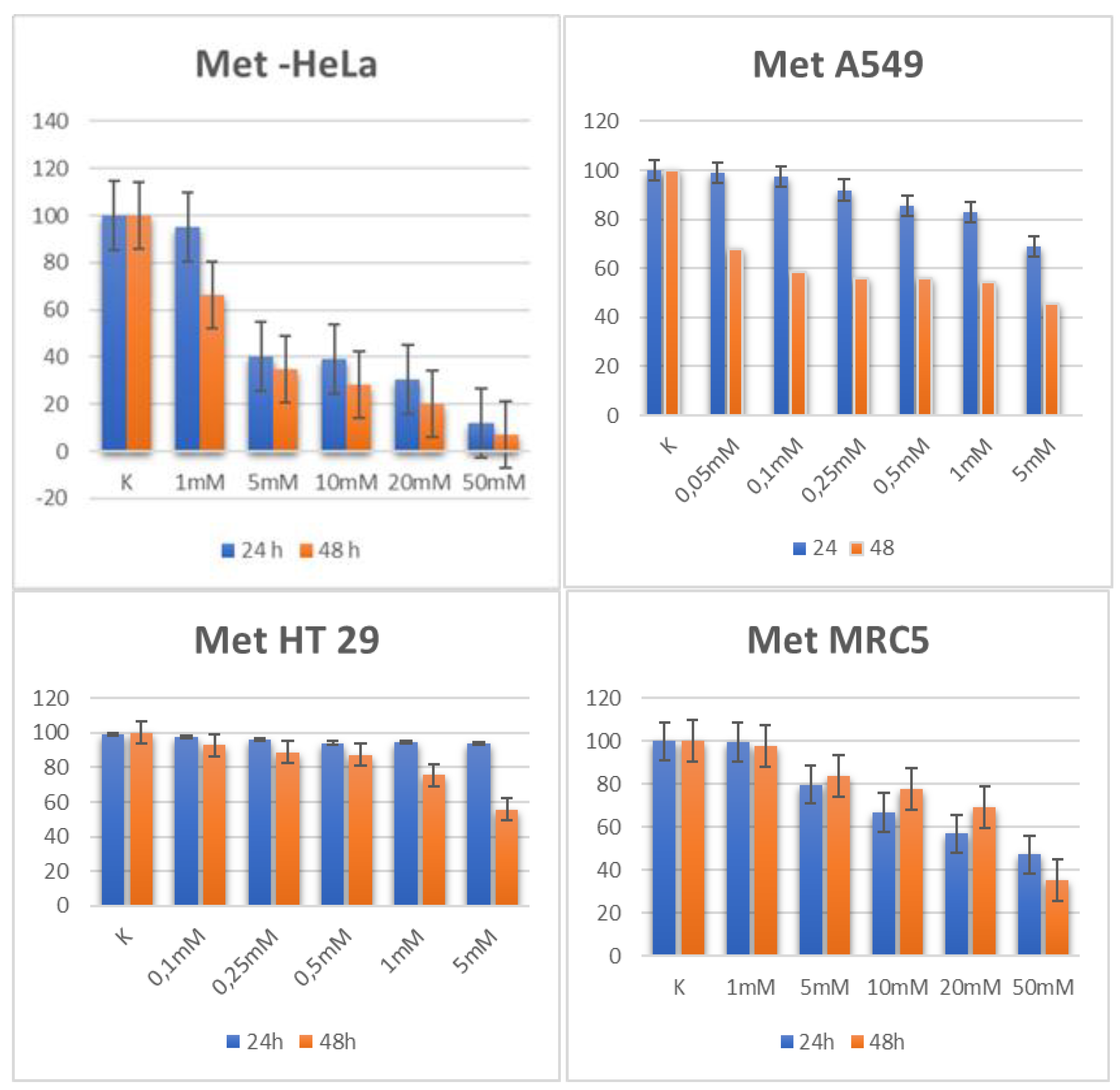
2.1. Metformin
2.1.1. Cytotoxic Activity
| Cell line | Metformin 24h | Metformin 48h |
|---|---|---|
| HeLa | 6.04 | 2.28 |
| A549 | 14.79 | 3.30 |
| HT29 | 26.53 | 10.54 |
| MRC-5 | 33.461 | 33.461 |
2.2. Caffeine
2.3. Combination of Metformin and Caffeine
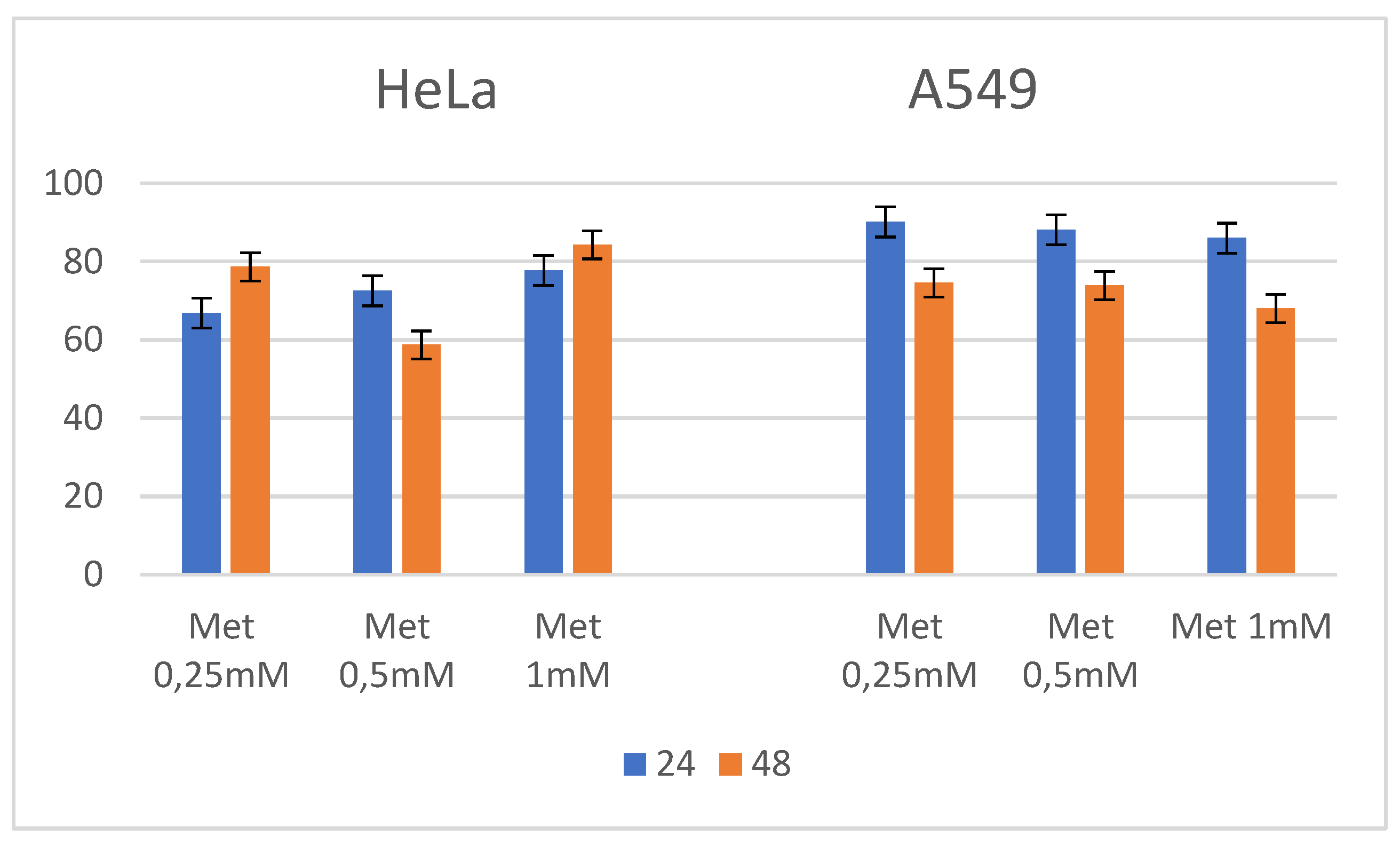
2.4. Mechanistic Studies
2.4.1. Flow Cytometry
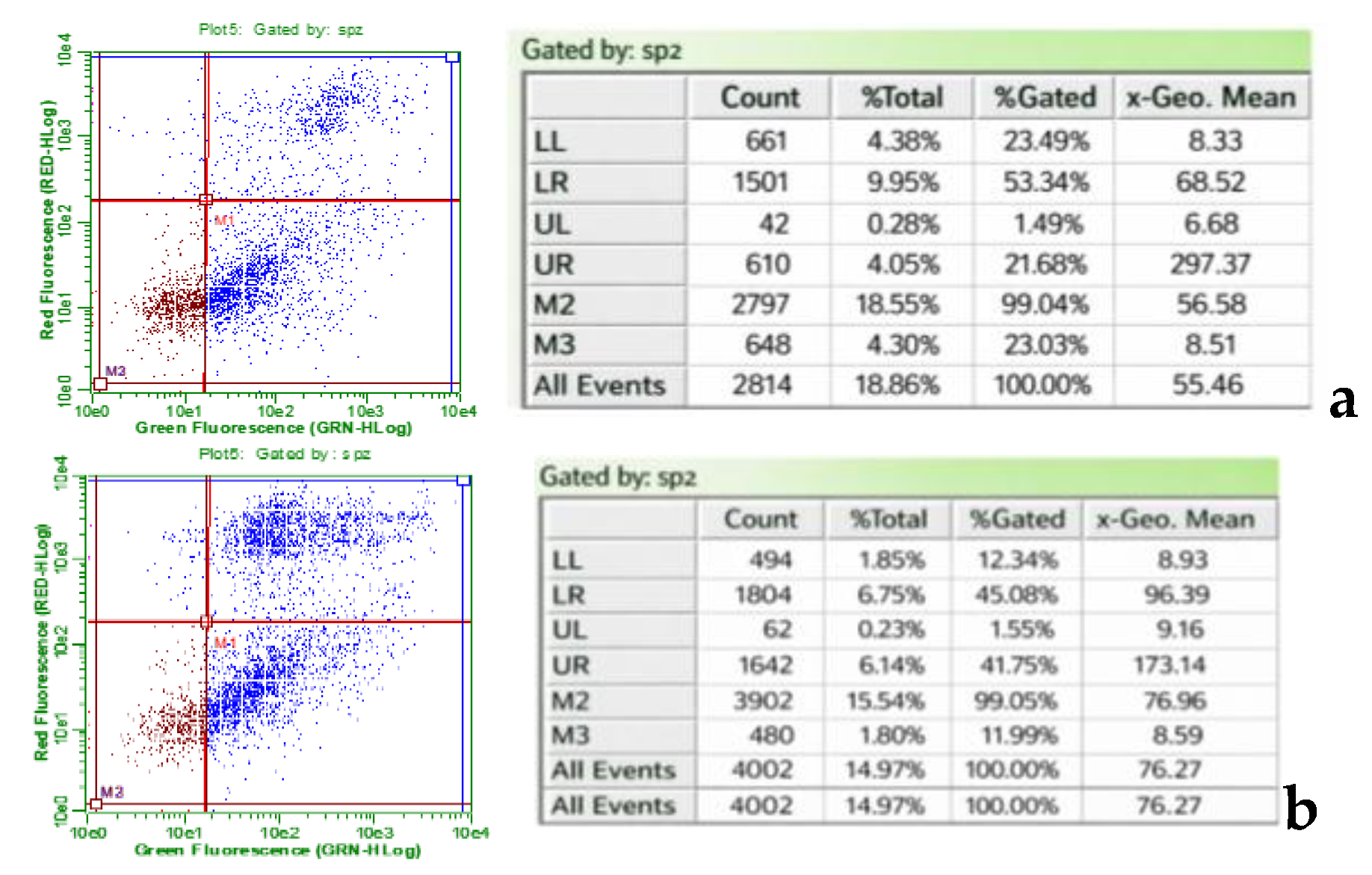
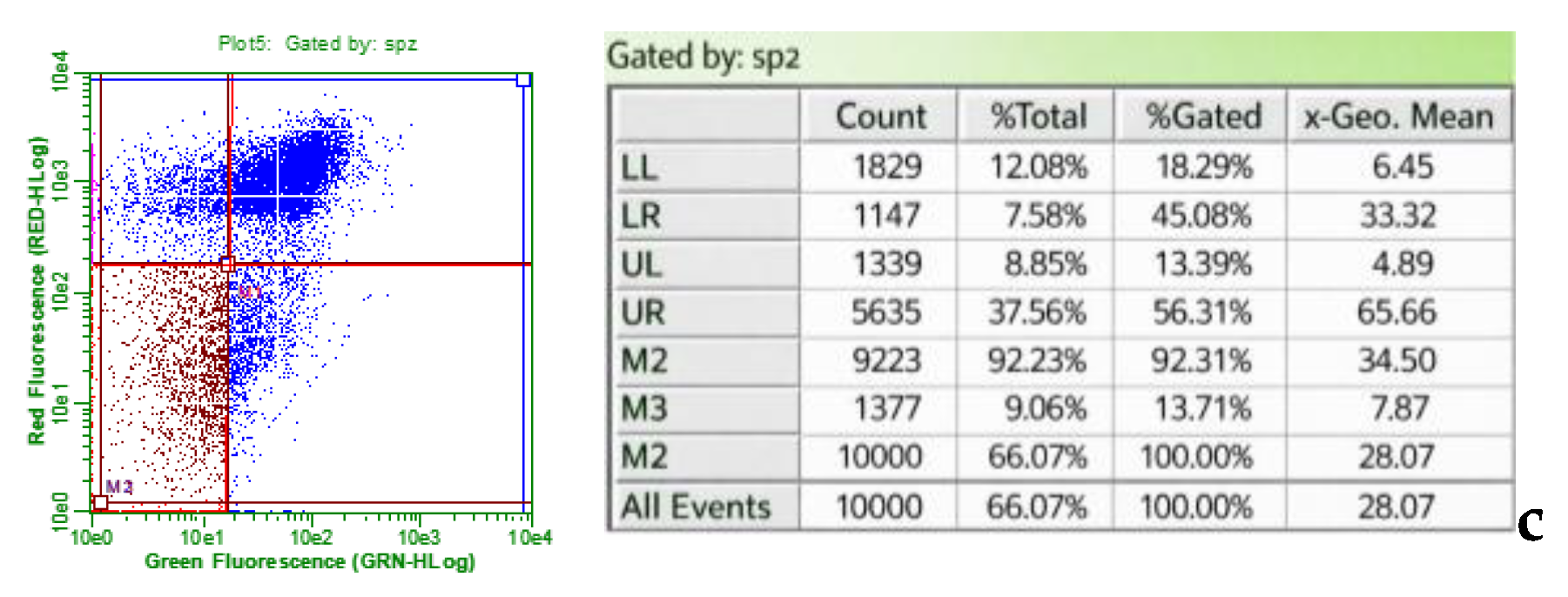
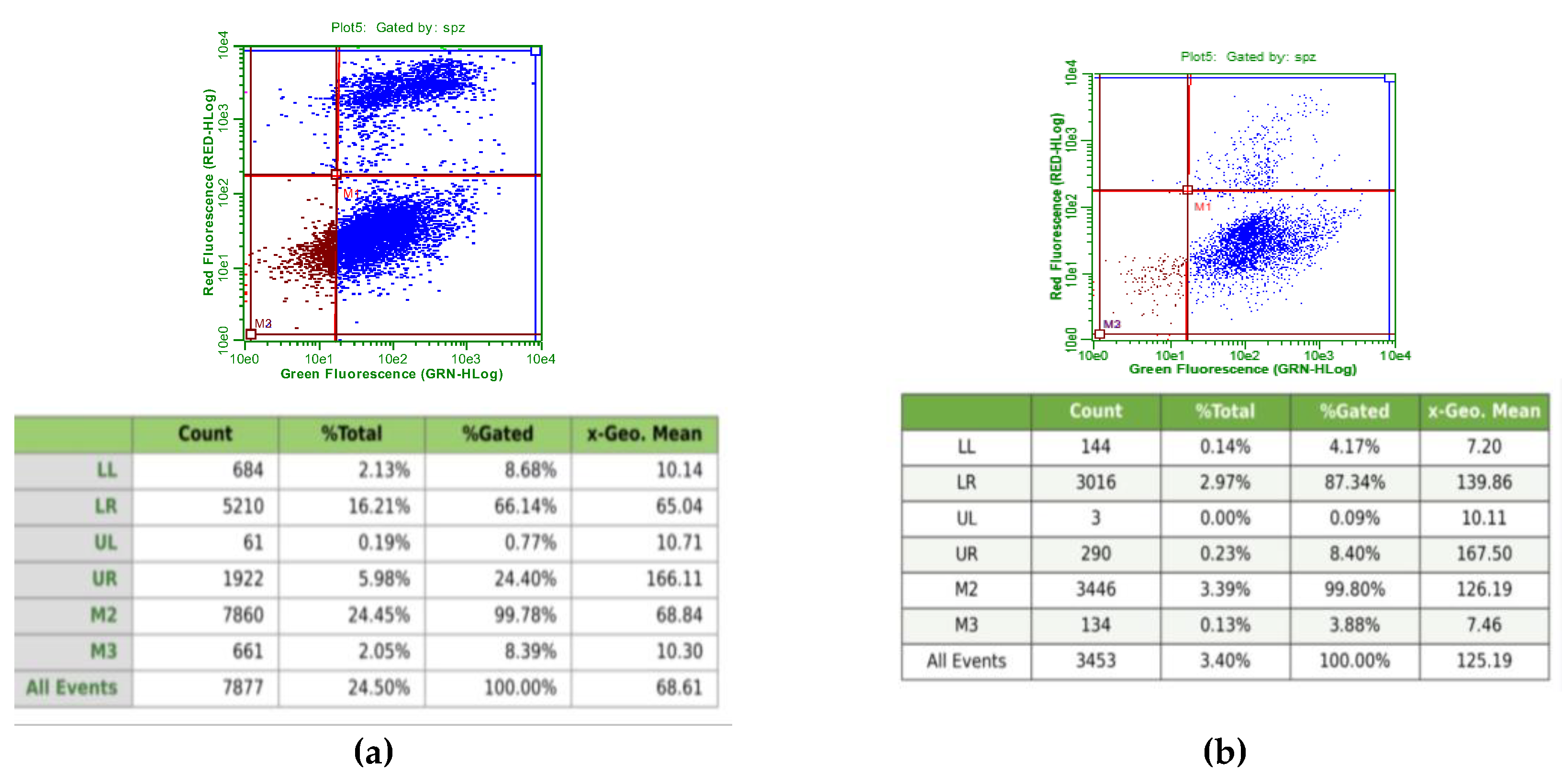
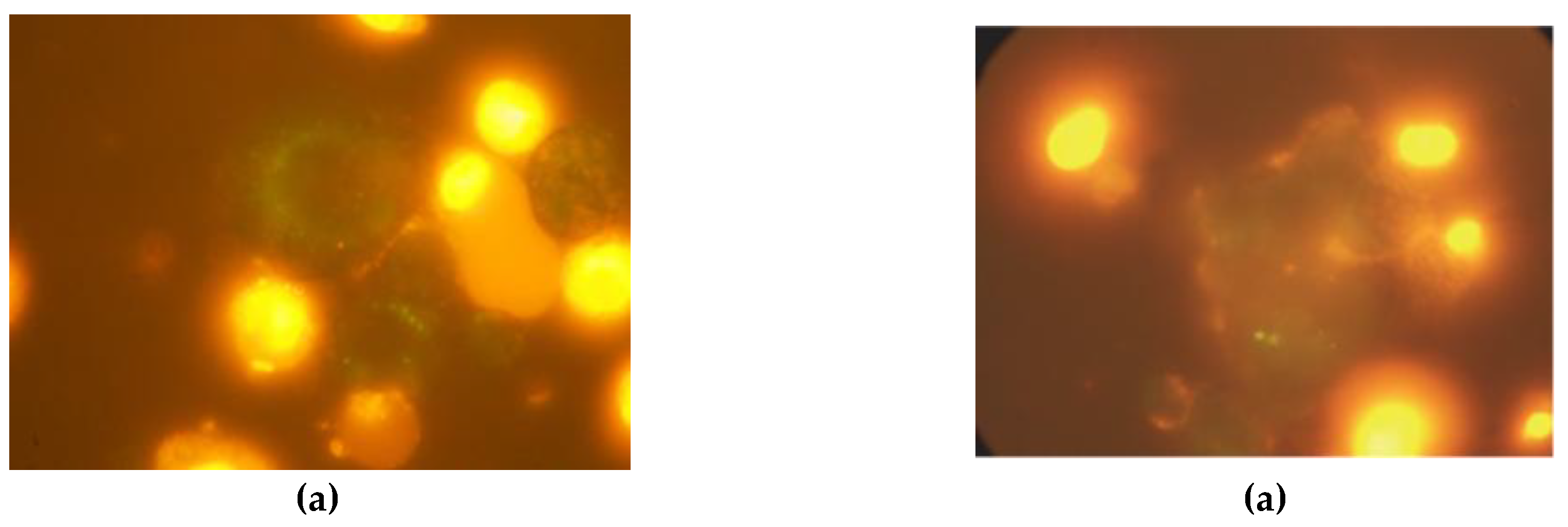
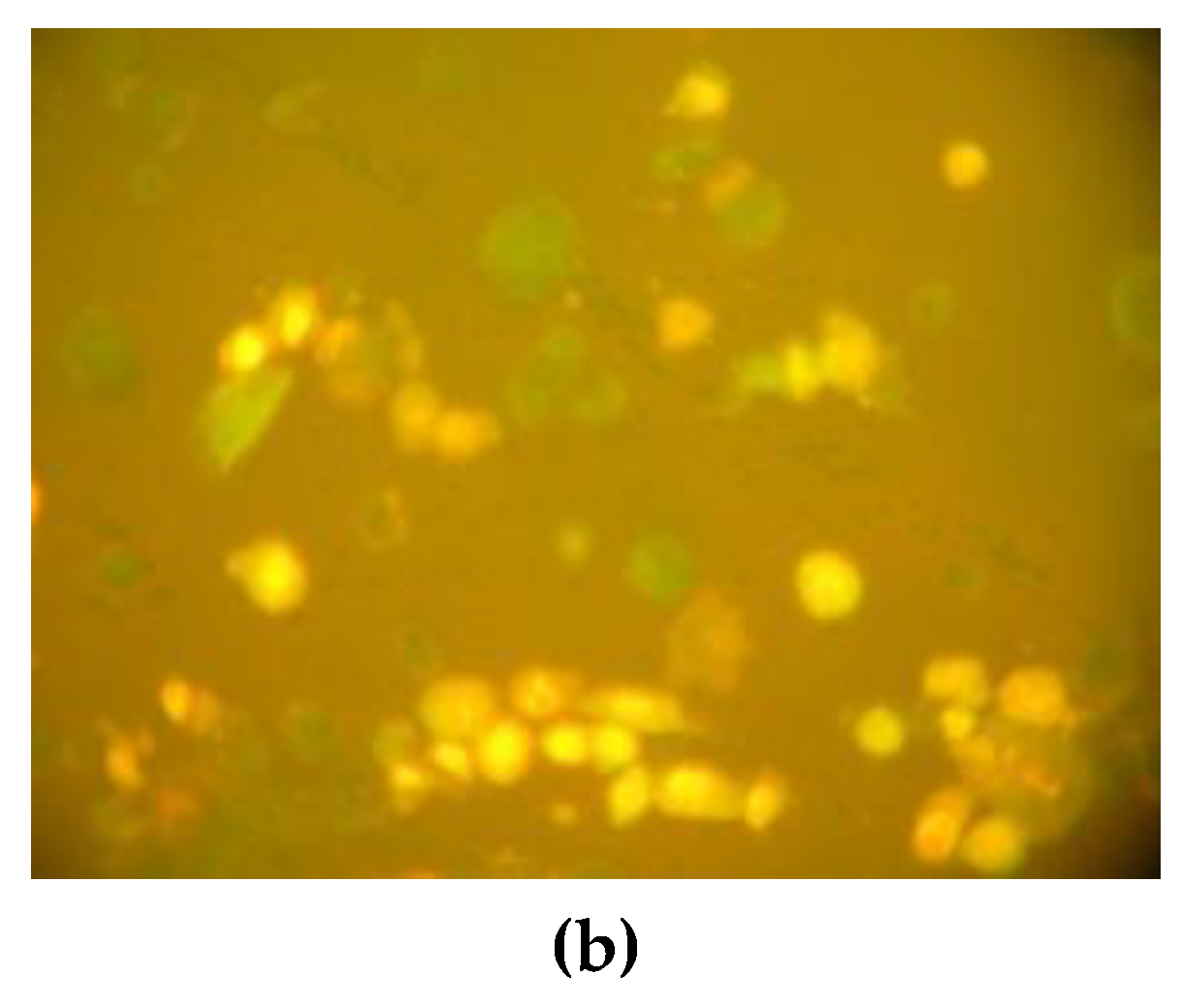
2.4.2. Immunofluorescence Microscopy Analysis of Apoptosis
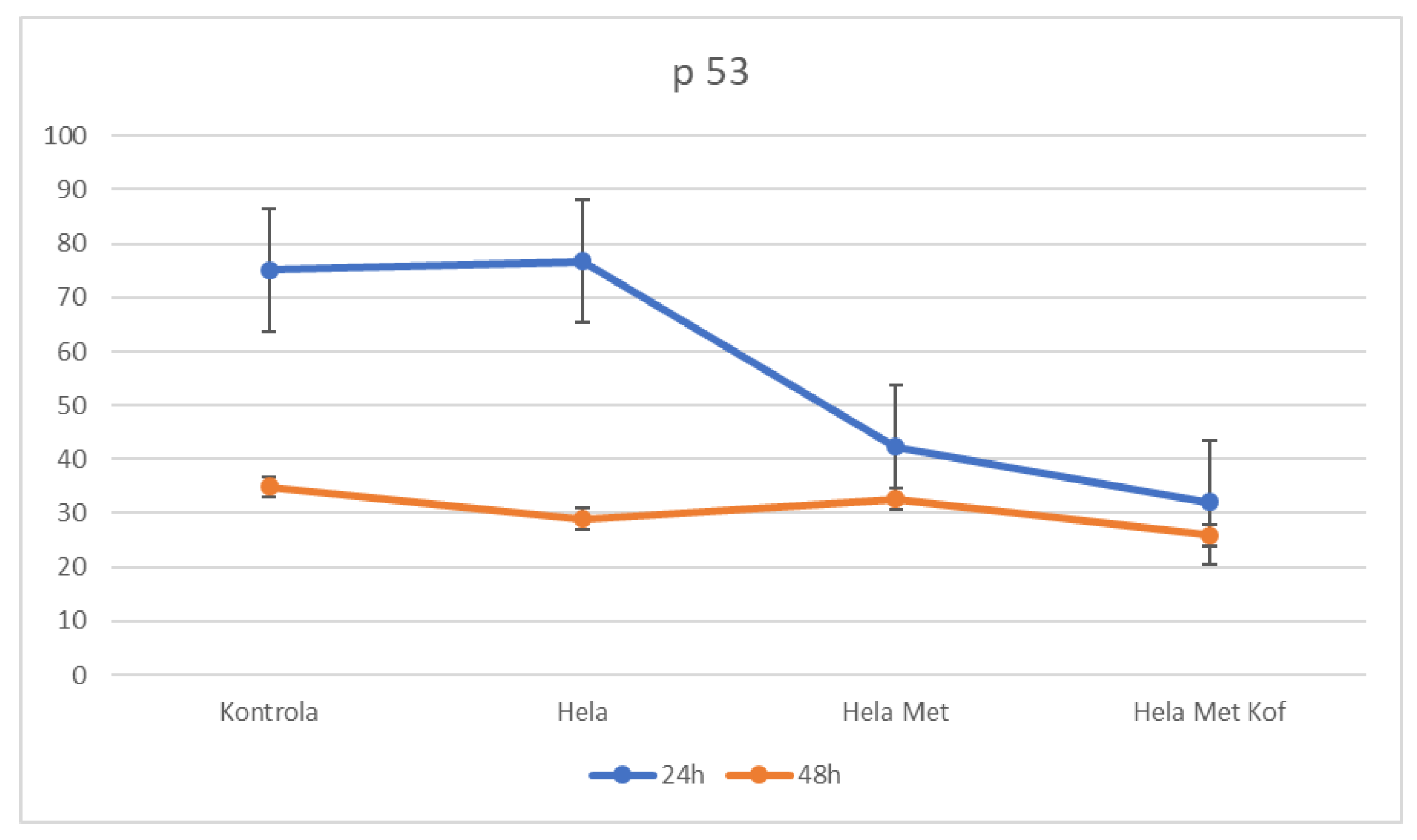
2.4.3. p53 ELISA: Modulation of Tumor Suppressor p53 Expression Following Metformin and Combined Metformin–Caffeine Treatment in Cervical Cancer Cells (HeLa cells)
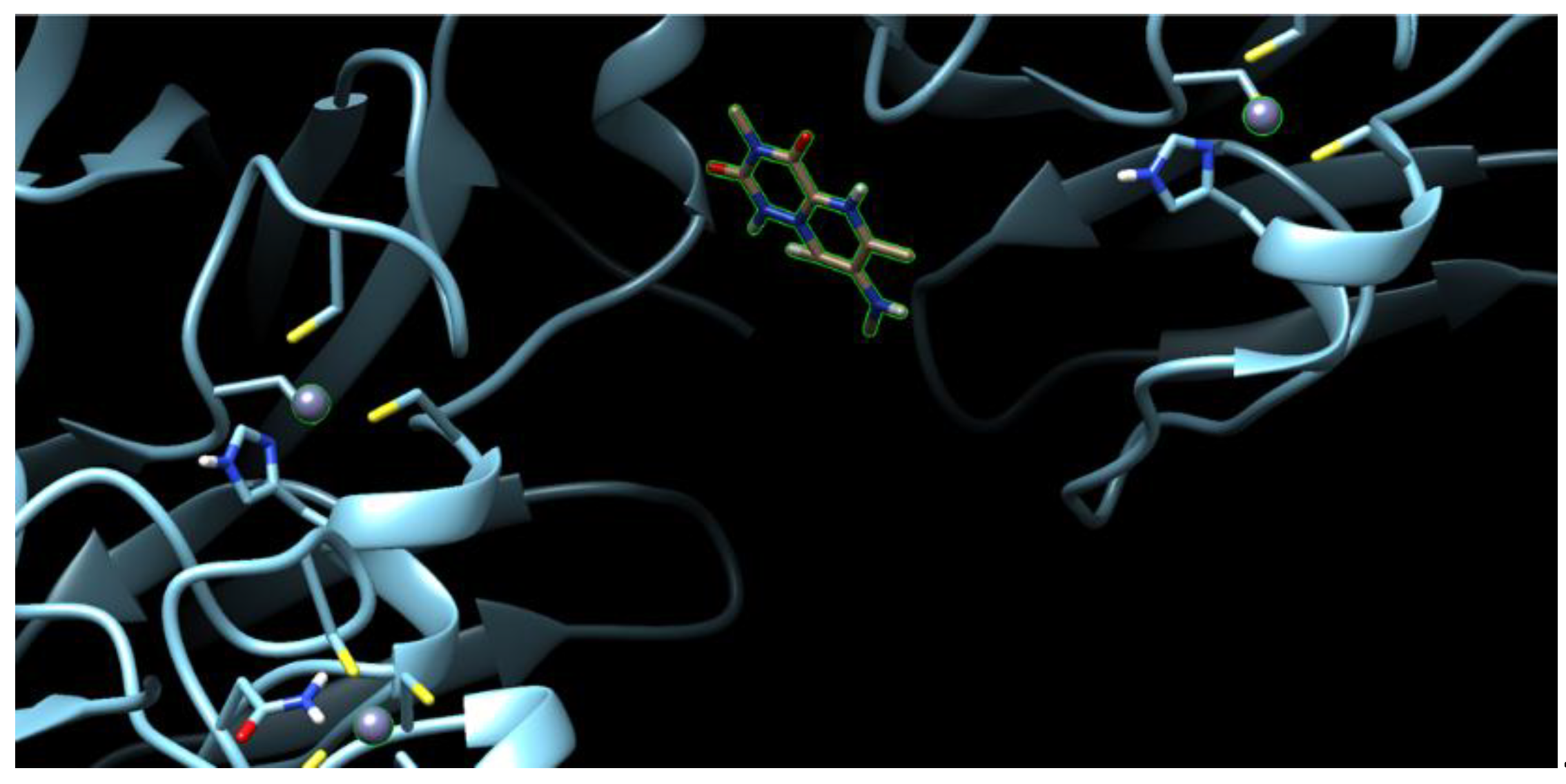
2.5. Molecular Docking Analysis
2.6. Drug Interaction Analysis by the Chou–Talalay Method
| 24h CI1 |
48h CI1 |
|---|---|
| MRC-5 0.96 | 1.53 |
| HeLa 0.78 | 1.55 |
| A549 1.13 | 2.34 |
| HT29 0.99 | 1.78 |
2.6.1. Isobologram Analysis and Theoretical Model
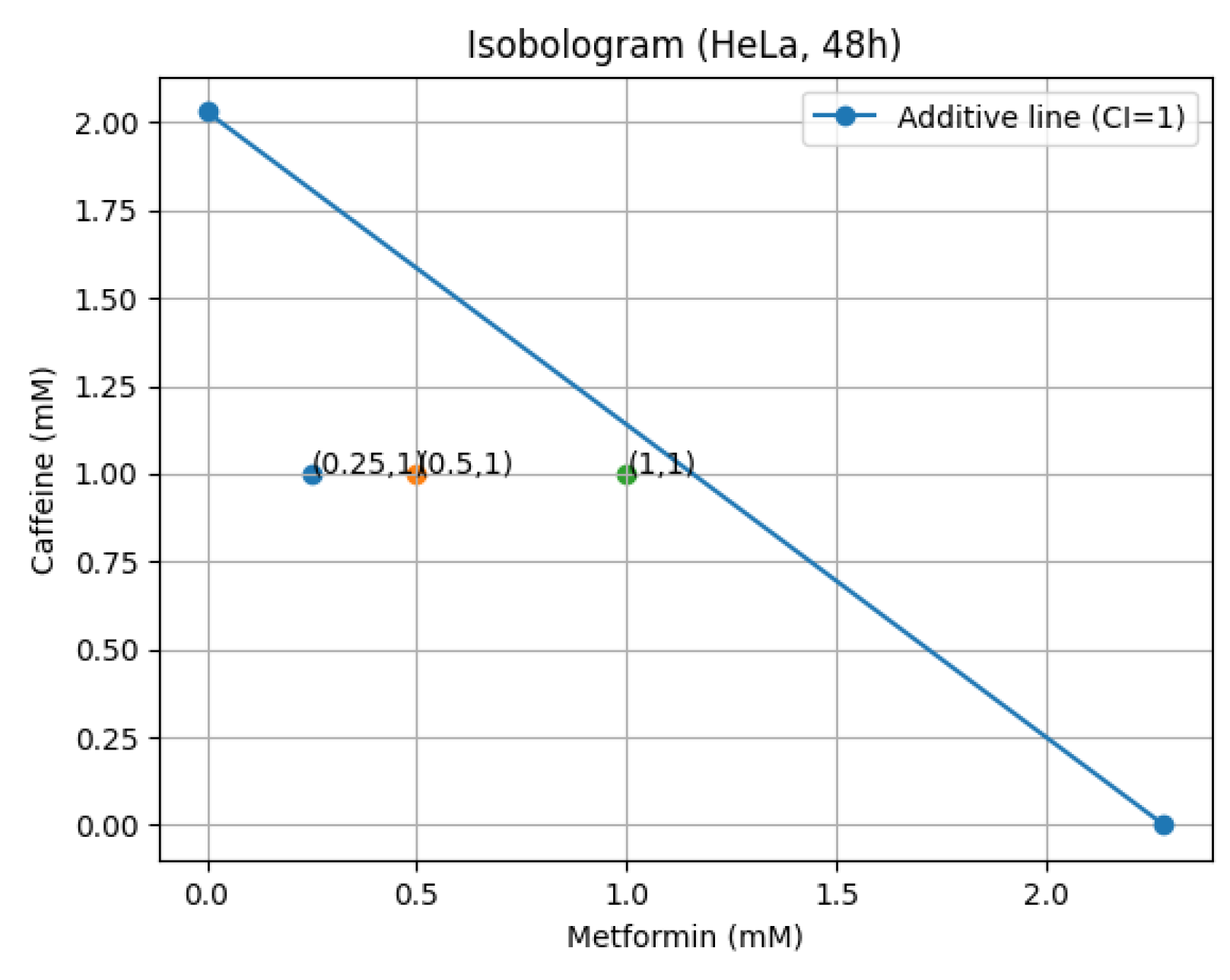
3. Discussion
4. Materials and Methods
4.1. Reagents
4.2. Cell Culture
4.3. Cell Viability and Proliferation Assays
4.3.1. Trypan Blue Exclusion Assay:
4.3.2. Sulforhodamine B (SRB) Assay
4.4. Apoptosis Analysis by Flow Cytometry
4.5. ELISA Assay: Determination of Tumor-Suppressor Genes for p53
4.6. Molecular Docking Analysis
4.7. Statistics
5. Conclusions
Supplementary Materials
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Acknowledgments
Conflicts of Interest
References
- World Health Organization. World Health Statistics 2024: Monitoring Health for the SDGs. WHO Press: Geneva, Switzerland, 2024. Available online: https://www.who.int/data/gho/publications/world-health-statistics(accessed on 2 September 2024).
- Wagle, N.S.; Nogueira, L.; Devasia, T.P.; et al. Cancer treatment and survivorship statistics, 2025. CA Cancer J. Clin. 2025, 75, 308–340. https://doi.org/10.3322/caac.70011. [CrossRef]
- Siegel, R.L.; Kratzer, T.B.; Giaquinto, A.N.; Sung, H.; Jemal, A. Cancer statistics, 2025. CA Cancer J. Clin. 2025, 75, 10–45. https://doi.org/10.3322/caac.21871. [CrossRef]
- Sung, H.; Ferlay, J.; Siegel, R.L.; Laversanne, M.; Soerjomataram, I.; Jemal, A.; Bray, F. Global cancer statistics 2020: GLOBOCAN estimates of incidence and mortality worldwide for 36 cancers in 185 countries. CA Cancer J. Clin. 2021, 71, 209–249. https://doi.org/10.3322/caac.21660. [CrossRef]
- Pryor, R.; Cabreiro, F. Repurposing metformin: an old drug with new tricks in its binding pockets. Biochem. J. 2015, 471(3):307-22. https://doi: 10.1042/BJ20150497. [CrossRef]
- Flory, J.; Lipska, K. Metformin in 2019. JAMA 2019, 321, 1926–1927.
- Evans, J.M.M.; Donnelly, L.A.; Emslie-Smith, A.M.; Alessi, D.R.; Morris, A.D. Metformin and reduced risk of cancer in diabetic patients. BMJ 2005, 330, 1304–1305. [CrossRef]
- Quinn, B.J.; Kitagawa, H.; Memmott, R.M.; Gills, J.J.; Dennis, P.A. Repositioning metformin for cancer prevention and treatment. Trends Endocrinol. Metab. 2013, 24, 469–480. [CrossRef]
- Vujovic, S.; Perovic, S.; Vlaovic, M.; Scepanovic, A.; Scepanovic, S. From metabolism to longevity: Molecular mechanisms underlying metformin’s anticancer and anti-aging effects. Curr. Issues Mol. Biol. 2026, 48, 286. [CrossRef]
- Saraei, P.; Asadi, I.; Kakar, M.A.; Moradi-Kor, N. The beneficial effects of metformin on cancer prevention and therapy: A comprehensive review of recent advances. Cancer Manag. Res. 2019, 11, 3295–3313. [CrossRef]
- Boroughs, L.K.; DeBerardinis, R.J. Metabolic pathways promoting cancer cell survival and growth. Nat. Cell Biol. 2015, 17, 351–359. [CrossRef]
- Morales, D.R.; Morris, A.D. Metformin in cancer treatment and prevention. Annu. Rev. Med. 2015, 66, 17–29.
- Zakikhani, M.; Dowling, R.; Fantus, I.; Sonenberg, N.; Pollak, M. Metformin is an AMPK-dependent growth inhibitor for breast cancer cells. Cancer Res. 2006, 66, 10269–10273. [CrossRef]
- Dowling, R.J.O.; Zakikhani, M.; Fantus, I.G.; Pollak, M.; Sonenberg, N. Metformin inhibits mTOR signaling in cancer cells. Cancer Res. 2007, 67, 10804–10812.
- Wang, Z.; Gu, C.; Wang, X.; Lang, Y.; Wu, Y.; Wu, X.; Zhu, X.; Wang, K.; Yang, H. Caffeine enhances the anti-tumor effect of 5-fluorouracil via increasing the production of reactive oxygen species in hepatocellular carcinoma. Med. Oncol.2019, 36, 97. https://doi.org/10.1007/s12032-019-1323-8. [CrossRef]
- Bode, A.M.; Dong, Z. The enigmatic effects of caffeine in cell cycle and cancer. Cancer Lett. 2007, 247, 26–39. [CrossRef]
- Weber, A.M.; Ryan, A.J.ATM and ATR as therapeutic targets in cancer. Pharmacol.Ther. 2015.149, 124–138. [CrossRef]
- Fagundes, T.R.; Madeira, T.B.; Melo, G.P.; Silva, C.R.; Silva, A.R.; Nunes, P.S.; de Oliveira, R.M.; de Souza, P.O.; Valverde, L.F.; de Souza, M.G.M. Caffeine improves the cytotoxic effect of dacarbazine on melanoma cells. Bioorg. Chem.2022, 120, 105576. https://doi.org/10.1016/j.bioorg.2022.105576. [CrossRef]
- Al-Lazikani, B.; Banerji, U.; Workman, P. Combinatorial drug therapy for cancer in the post-genomic era. Nat. Biotechnol.2012, 30, 679–692. https://doi.org/10.1038/nbt.2284. [CrossRef]
- Trott, O.; Olson, A.J. AutoDock Vina: Improving the speed and accuracy of docking. J. Comput. Chem.2010, 31, 455–461. https://doi.org/10.1002/jcc.21334. [CrossRef]
- Hardie, D.G. AMPK: A key regulator of energy balance in cells and organisms. Nat. Rev. Mol. Cell Biol. 2012, 13, 251–262.
- Wheaton, W.W.; Weinberg, S.E.; Hamanaka, R.B.; et al.Metformin inhibits mitochondrial complex I of cancer cells.eLife 2014, 3, e02242. [CrossRef]
- Heckman-Stoddard, B.; DeCensi, A.; Sahasrabuddhe, V.; Ford, L.Repurposing metformin for cancer prevention and treatment.Diabetologia2017, 60, 1639–1647.
- Pernicova, I.; Korbonits, M.Metformin—Mode of action and implications for cancer therapy.Endocr. Rev. 2014, 35, 688–719.
- Kasznicki, J.; Sliwinska, A.; Drzewoski, J.Metformin in cancer prevention and therapy. Ann. Transl. Med. 2018, 6, 1–11. [CrossRef]
- Rena, G.; Hardie, D.G.; Pearson, E.R.The mechanisms of action of metformin. Diabetologia 2017,60,1577-1585.
- Luengo, A.; Sullivan, L.B.; Vander Heiden, M.G. Understanding the complex-I-ty of metformin action: Limiting mitochondrial respiration to improve cancer therapy. BMC Biol.2014, 12, 82. [CrossRef]
- Lefranc, F.; Tabanca, N.; Kiss, R. Assessing the anticancer effects associated with food products and/or nutraceuticals using in vitro and in vivo preclinical development-related pharmacological tests. Semin. Cancer Biol.2017, 46, 14–32. [CrossRef]
- Vousden, K.H.; Lane, D.P. p53 in health and disease. Nature Reviews Molecular Cell Biology 2007.Nat Rev Mol Cell Biol. 2007 (4):275-83. doi: 10.1038/nrm2147. [CrossRef]
- Kalender A., Selvaraj A., Kim S.Y., Gulati P., BrûléS., ViolletB., Kalender, A.; et al. Metformin inhibits mTOR signaling. Cell Metabolism 2010; 11(5):390-401. doi: 10.1016/j.cmet.2010.03.014. [CrossRef]
- Bogdanović, G. Raletić-Savić, J. Marković N. In vitro assays for antitumor-drug screening on human tumor cell lines: dye exclusion test and colorimetric cytotoxicity assay. Arch Oncol. 1994; 2(4):181-4.
- Marciniak, M., Torbicki, A., Korpalski, M., Pawluczyk, M., Pawlikowski, K., Żygłowicz, M., Augustyn, D., Gaworek, P., Trybuła A. Metformin in Oncology- Its Effect on Cancer Development and Progression. Medical Science2025; 29: e132ms3665 doi: https://doi.org/10.54905/disssi.v29i162.e132ms3665. [CrossRef]
- Strober, W. Trypan blue exclusion test of cell viability. CurrProtoc Immunol. 2001; 21(1):A.3B.1-A.3B.2. [CrossRef]
- Shekan P., Storeng R., Scudiero D., Monks A., McMahon J.,Vistica D., et al. New colorimetric cytotoxicity assay for anticancer-drug screening. J Natl Cancer Inst. 1990; 82(13):1107-12.
- Higuchi, K.Mitsuhashi, N. Saitoh, J, Maebayashi, K. Sakurai, H, Akimoto, T. Niibe, H. Caffeine enhanced radiosensitivity of rat tumor cells with a mutant-type p53 by inducing apoptosis in a p53-independent manner. Cancer Lett. 2000; 152(2):157-62. doi: 10.1016/s0304-3835(99)00449-8. [CrossRef]
- Chou, TC.Talalay, P. Quantitative analysis of dose–effect relationships: the combined effects of multiple drugs or enzyme inhibitors. Adv Enzyme Regul. 1984; 22:27–55. doi: 10.1016/0065-2571(84)90007-4. [CrossRef]
- Chou, TC. Drug combination studies and their synergy quantification using the Chou–Talalay method. Cancer Res. 2010;70(2):440–446. doi:10.1158/0008-5472. CAN-14-3763. [CrossRef]
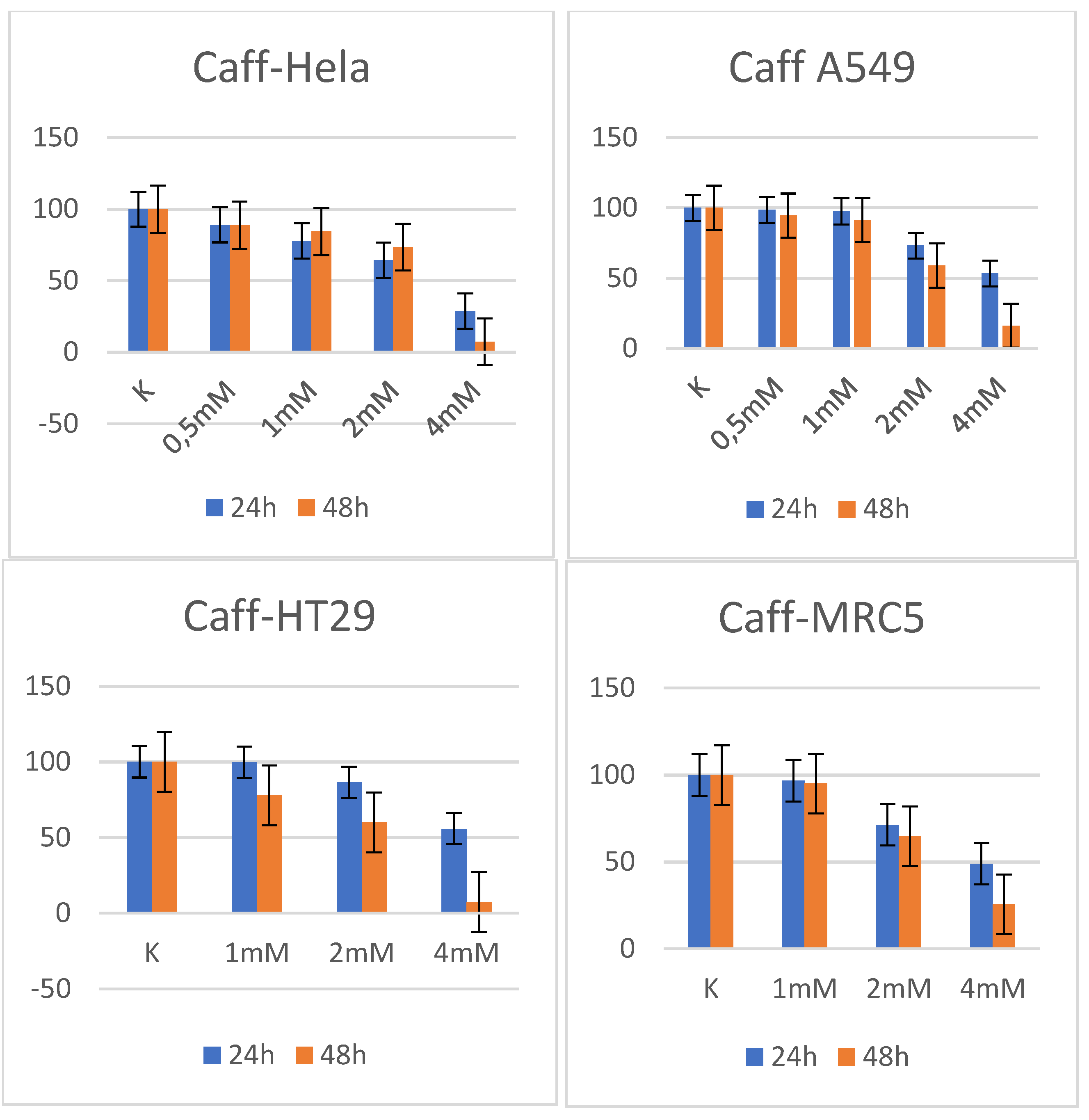


| Cell line | Caffeine24h | Caffeine48h |
|---|---|---|
| HeLa | 2.44 | 2.03 |
| A549 | 3.38 | 2.44 |
| HT29 | 3.41 | 2.01 |
| MRC-5 | 3.55 | 2.62 |
| Cell line | 24 h | 48h |
|---|---|---|
| HeLa | 2.23 | 2.40 |
| A549 | 12.39 | 6.36 |
| HT29 | 18.71 | 13.46 |
| MRC-5 | 22.60 | 38.30 |
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2026 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).




