Submitted:
20 April 2026
Posted:
20 April 2026
You are already at the latest version
Abstract
Keywords:
1. Introduction
2. Materials and Methods
2.1. Preparation of GRC Extract for GCN Synthesis
2.2. TAC (Total Antioxidant Capacity) Assay
2.3. High-Performance Liquid Chromatography (HPLC) Analysis for Adenosine
2.4. PM2.5 Sample Preparation
2.5. Cell Culture and Cell Viability Assay
2.6. ROS Assay
2.7. Acquisition of Microarray Data and Identification of Differentially Expressed Genes (DEGs)
2.8. Collection of PM- and LPS-Induced Lung Inflammation-Related Genes
2.9. Identification of Common and Hub Genes
2.10. Gene Ontology (GO) and Kyoto Encyclopedia of Genes and Genomes (KEGG) Pathway Functional Analysis
2.11. Reverse Transcriptase Polymerase Chain Reaction (RT-PCR)
2.12. Western Blot Analysis
2.13. Statistical Analysis
3. Results
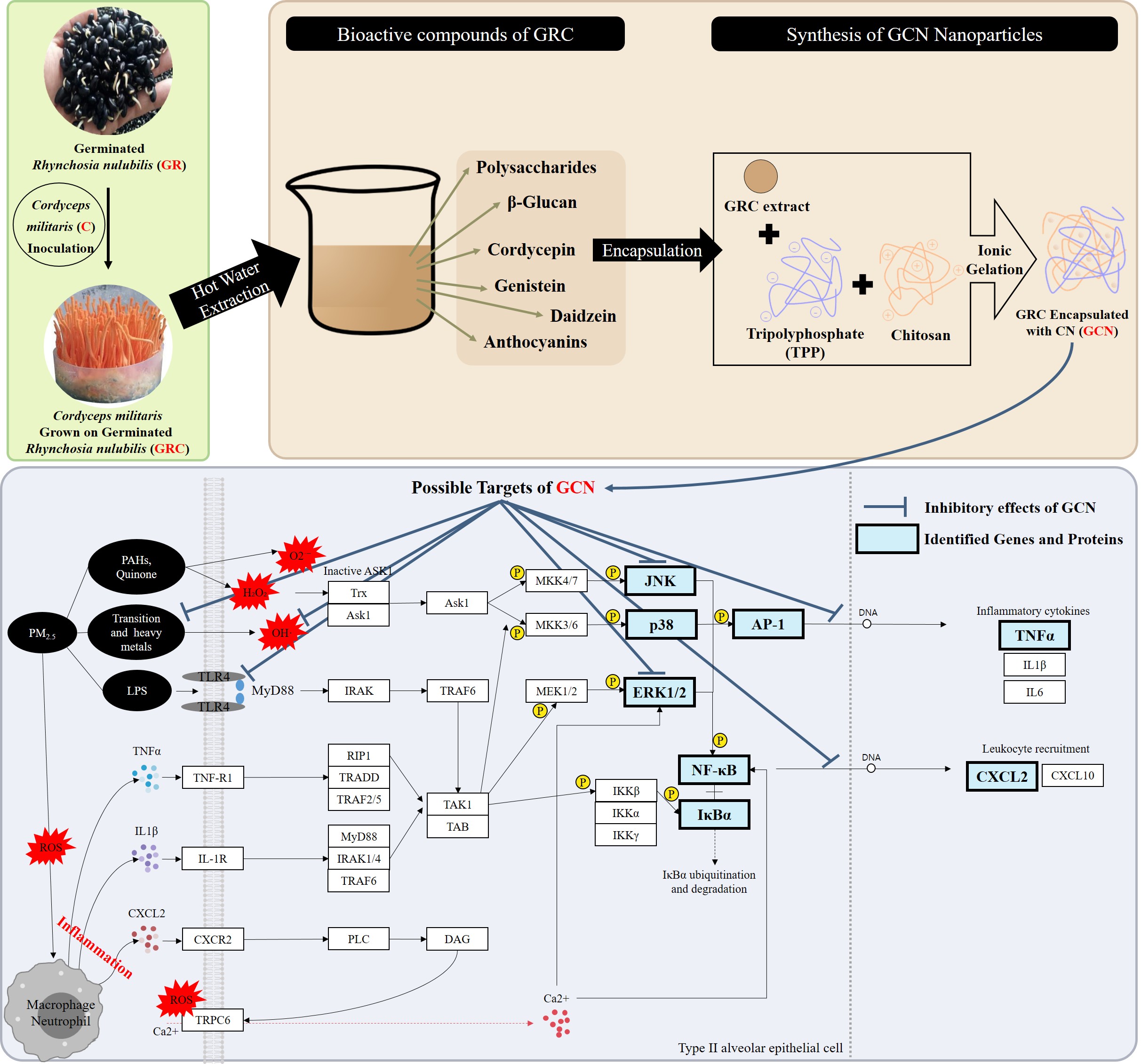
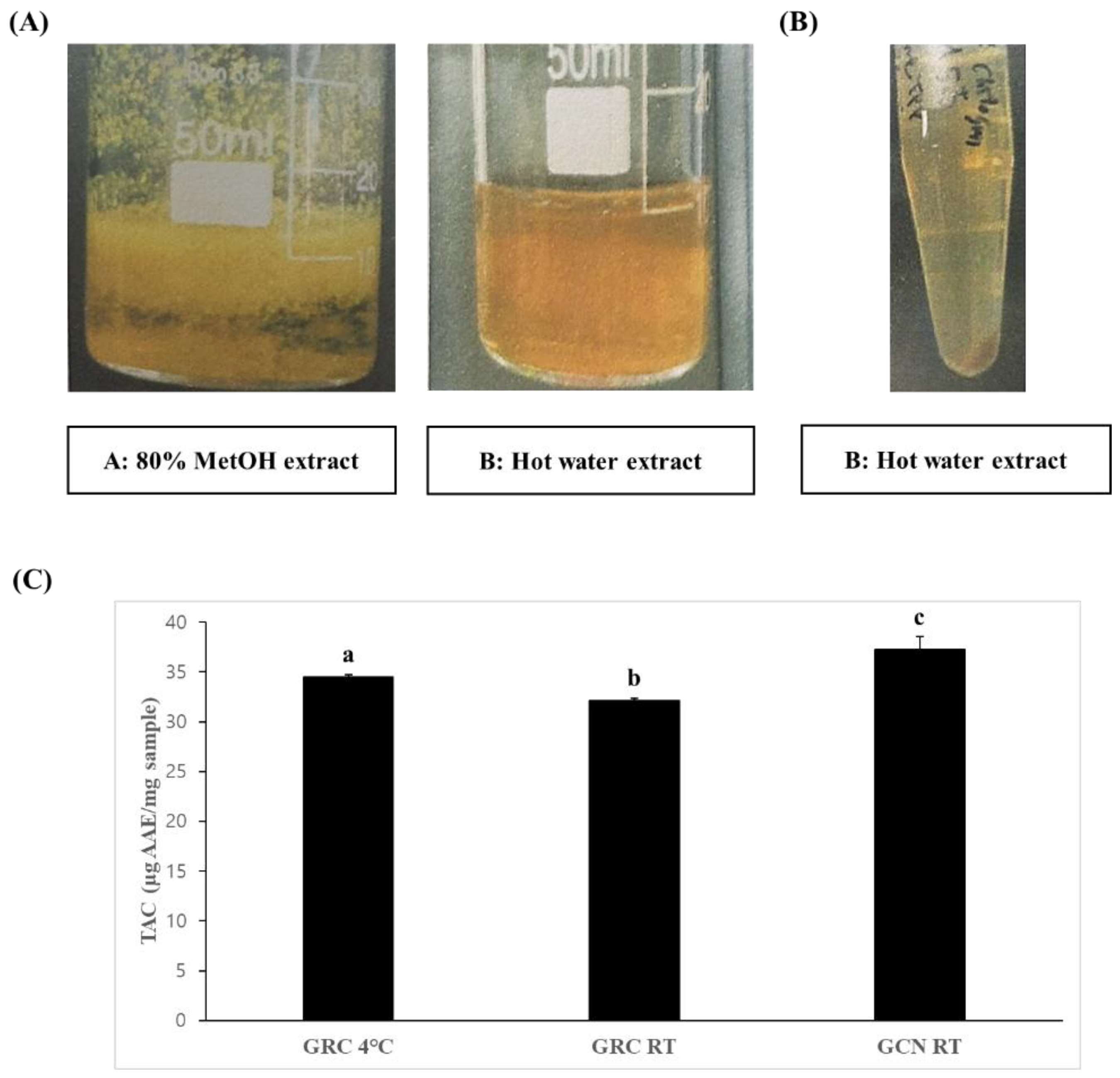
3.1. Formation of Chitosan Nanoparticles from Different GRC Extracts and Their Antioxidant Potential and Bioactive Compounds
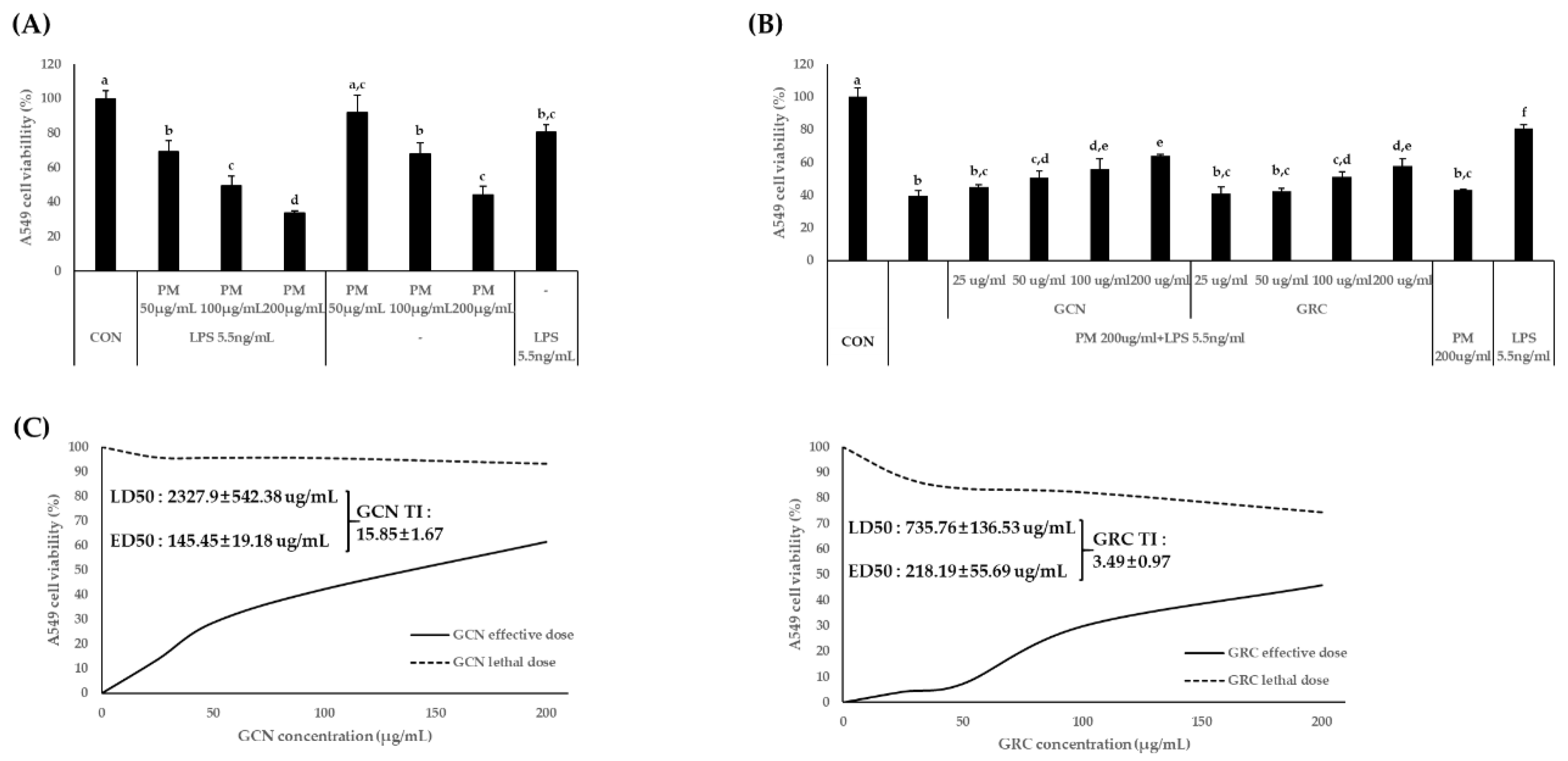
3.2. GCN Suppresses PM- and LPS-Induced A549 Cell Death and In Vivo Toxicological Evaluation
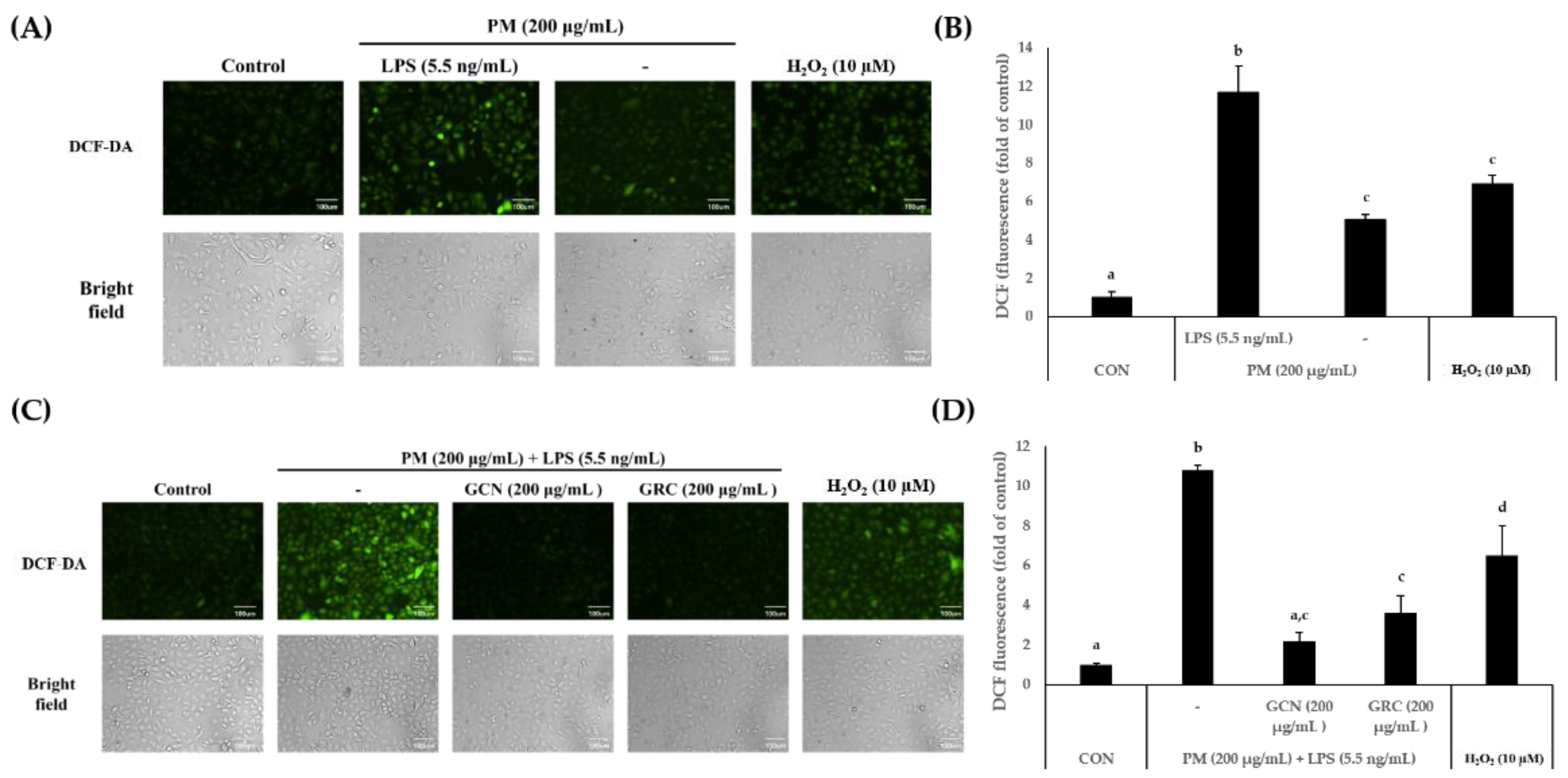
3.3. GCN Reduces PM- and LPS-Induced ROS Production in A549 Cells
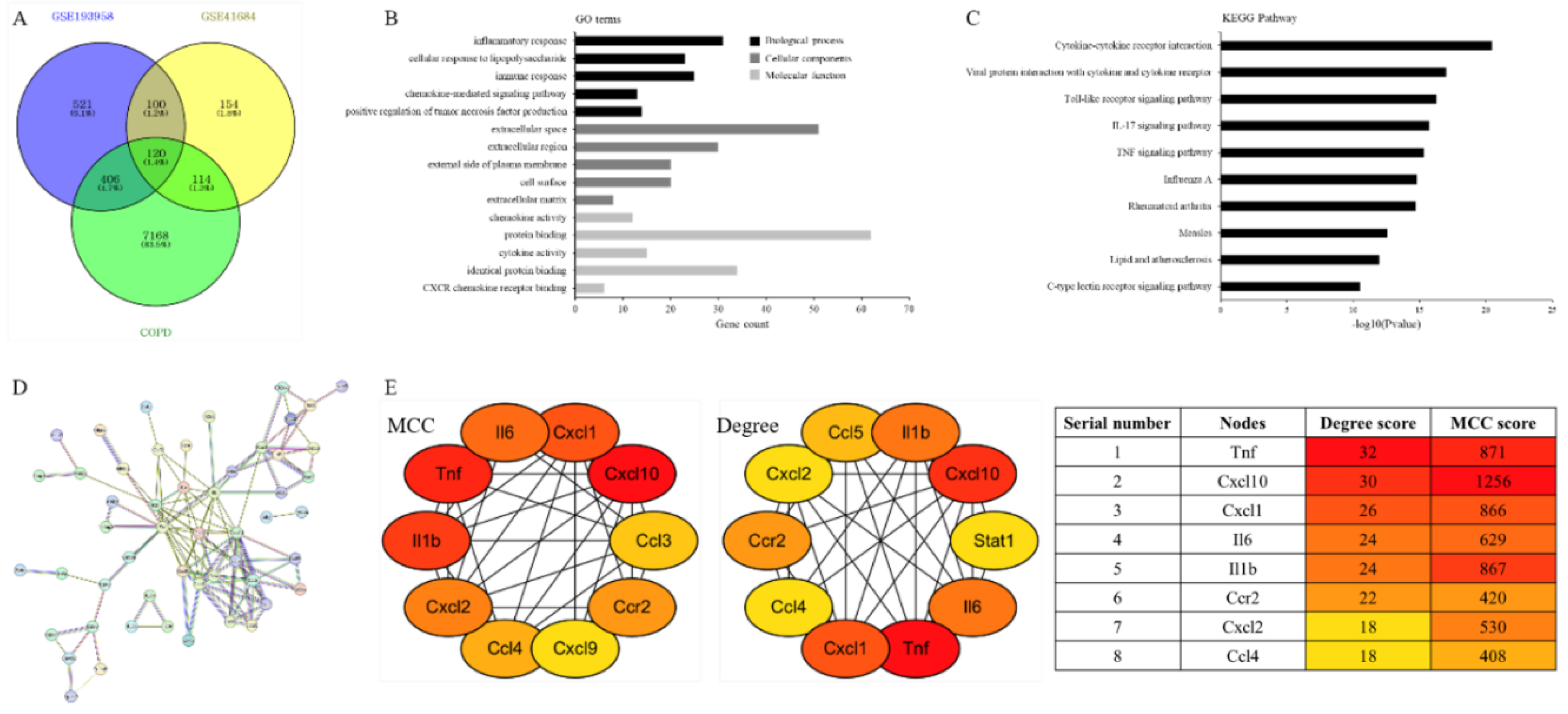
3.4. DEG Analysis Using GEO2R and DAVID for Biomarker Identification
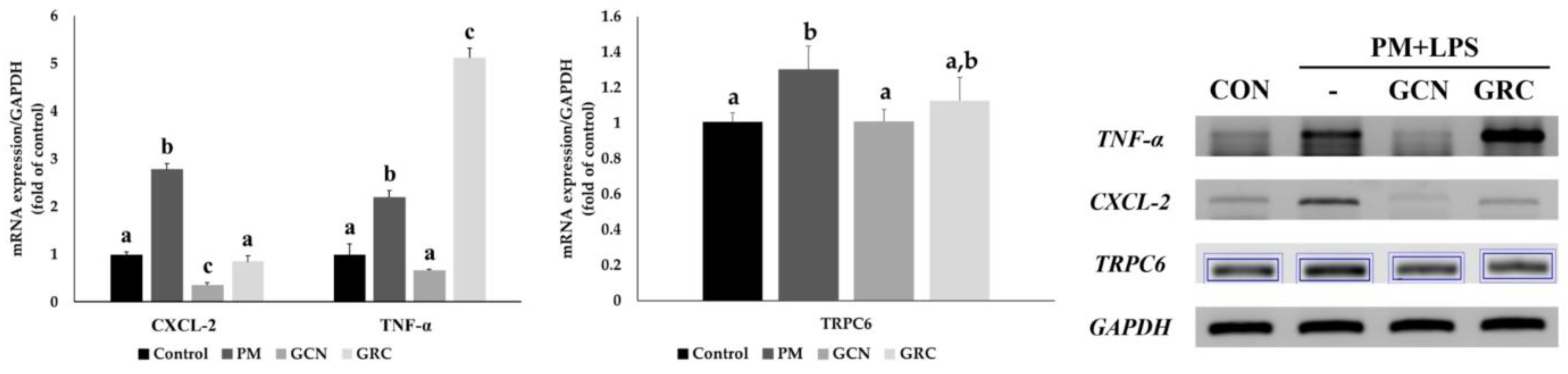
3.5. GCN Attenuates Cytokine and Chemokine Expression in PM- and LPS-Treated A549 Cells
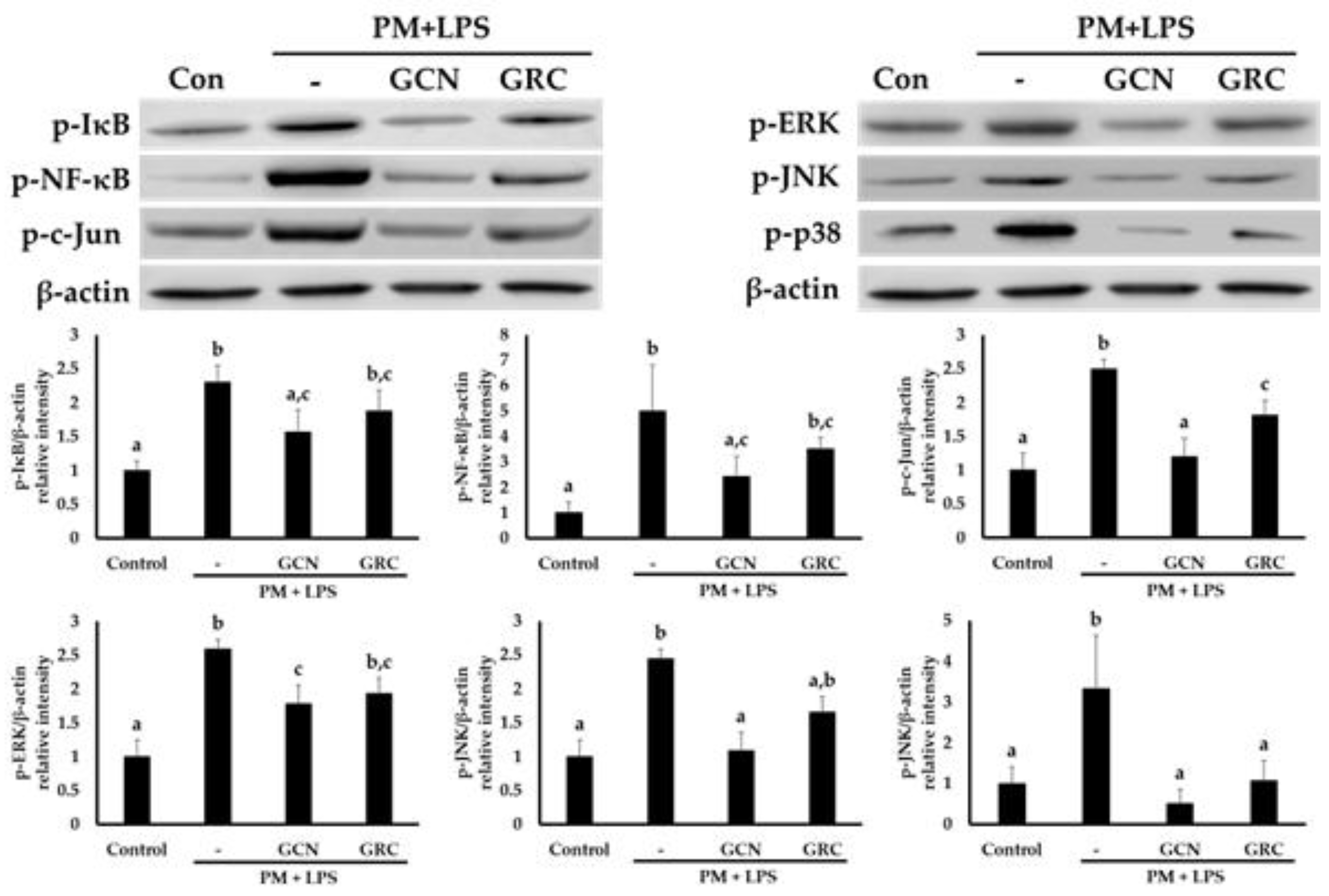
3.6. GCN Inhibits PM- and LPS-Induced Activation of MAPK-Mediated NF-κB/Activator Protein 1 (AP-1) Signaling Pathways
4. Discussion
5. Conclusions
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Acknowledgments
Conflicts of Interest
References
- Zomuansangi, R. Interaction of bacteria and inhalable particulate matter in respiratory infectious diseases caused by bacteria. Atmospheric Pollution Research 2024, 15(3), 102012. [Google Scholar] [CrossRef]
- Gualtieri, M. Winter fine particulate matter from Milan induces morphological and functional alterations in human pulmonary epithelial cells (A549). Toxicology letters 2009, 188(1), 52–62. [Google Scholar] [CrossRef] [PubMed]
- Dai, P. The roles of Nrf2 and autophagy in modulating inflammation mediated by TLR4-NFκB in A549 cell exposed to layer house particulate matter 2.5 (PM2. 5). Chemosphere 2019, 235, 1134–1145. [Google Scholar] [CrossRef] [PubMed]
- Wang, Q.; Liu, S. The effects and pathogenesis of PM2. 5 and its components on chronic obstructive pulmonary disease. International journal of chronic obstructive pulmonary disease 2023, 493–506. [Google Scholar] [CrossRef] [PubMed]
- Finnerty, K. Instillation of coarse ash particulate matter and lipopolysaccharide produces a systemic inflammatory response in mice. Journal of Toxicology and Environmental Health, Part A 2007, 70(23), 1957–1966. [Google Scholar] [CrossRef]
- Sang, S. The global burden of disease attributable to ambient fine particulate matter in 204 countries and territories, 1990–2019: A systematic analysis of the Global Burden of Disease Study 2019. Ecotoxicology and environmental safety 2022, 238, 113588. [Google Scholar] [CrossRef]
- Zhang, T. Protective effects of natural products against lung damage caused by fine particulate matter. Environmental Pollution 2025, 126942. [Google Scholar] [CrossRef]
- Wang, X. Mechanism of PM2. 5 induced/aggravated allergic diseases and its prevention and treatment. Allergy Medicine 2024. 2, 100012. [Google Scholar] [CrossRef]
- Yue, G.G.-L. Effects of Cordyceps sinensis, Cordyceps militaris and their isolated compounds on ion transport in Calu-3 human airway epithelial cells. Journal of Ethnopharmacology 2008, 117(1), 92–101. [Google Scholar] [CrossRef]
- Wang, X. Cordyceps militaris solid medium extract alleviates lipopolysaccharide-induced acute lung injury via regulating gut microbiota and metabolism. Frontiers in Immunology 2025. 15, 1528222. [Google Scholar] [CrossRef]
- Chou, Y.-C. Current progress regarding Cordyceps militaris, its metabolite function, and its production. Applied Sciences 2024, 14(11), 4610. [Google Scholar] [CrossRef]
- Das, S.K. Medicinal uses of the mushroom Cordyceps militaris: current state and prospects. Fitoterapia 2010, 81(8), 961–968. [Google Scholar] [CrossRef] [PubMed]
- Phull, A.-R.; Ahmed, M.; Park, H.-J. Cordyceps militaris as a bio functional food source: pharmacological potential, anti-inflammatory actions and related molecular mechanisms. Microorganisms 2022, 10(2), 405. [Google Scholar] [CrossRef] [PubMed]
- Zhang, J. Advance in Cordyceps militaris (Linn) Link polysaccharides: Isolation, structure, and bioactivities: A review. International journal of biological macromolecules 2019, 132, 906–914. [Google Scholar] [CrossRef]
- Park, H.-J. Influence of Culture Conditions on Bioactive Compounds in Cordyceps militaris: A Comprehensive Review. Foods 2025, 14(19), 3408. [Google Scholar] [CrossRef]
- Phoungthong, K. Utilization of corncob biochar in cultivation media for Cordycepin production and biomass of Cordyceps militaris. Sustainability 2022, 14(15), 9362. [Google Scholar] [CrossRef]
- Park, M.H.; Kim, M. Antioxidant and anti-inflammatory activity and cytotoxicity of ethanol extracts from Rhynchosia nulubilis cultivated with Ganoderma lucidum mycelium. Preventive nutrition and food science 2018, 23(4), p. 326. [Google Scholar] [CrossRef]
- Yu, S.; Park, H.; Kim, W. Anti-inflammaging effects of black soybean and black rice mixture extract by reprogramming of mitochondrial respirations in murine macrophages. Journal of Functional Foods 2022, 94, 105114. [Google Scholar] [CrossRef]
- Wu, T. Exploring the effect of boiling processing on the metabolic components of black beans through in vitro simulated digestion. LWT 2023, 184, 114987. [Google Scholar] [CrossRef]
- Kwon, H.-K.; Jo, W.-R.; Park, H.-J. Immune-enhancing activity of C. militaris fermented with Pediococcus pentosaceus (GRC-ON89A) in CY-induced immunosuppressed model. BMC complementary and alternative medicine 2018, 18(1), 75. [Google Scholar] [CrossRef]
- Phull, A.-R.; Dhong, K.-R.; Park, H.-J. Lactic acid bacteria fermented Cordyceps militaris (GRC-SC11) suppresses ige mediated mast cell activation and type i hypersensitive allergic murine model. Nutrients 2021, 13(11), 3849. [Google Scholar] [CrossRef]
- Park, D.K.; Park, H.-J. Ethanol Extract of Cordyceps militaris Grown on Germinated Soybeans Attenuates Dextran-Sodium-Sulfate-(DSS-) Induced Colitis by Suppressing the Expression of Matrix Metalloproteinases and Inflammatory Mediators. BioMed Research International 2013, 2013(1), p. 102918. [Google Scholar] [CrossRef]
- Lee, H.-J.; Park, H.-J. Germinated Rhynchosia nulubilis Fermented with Lactobacillus pentosus SC65 Reduces Particulate Matter Induced Type II Alveolar Epithelial Apoptotic Cell Death. International Journal of Molecular Sciences 2021, 22(7), 3660. [Google Scholar] [CrossRef] [PubMed]
- Zhang, D.-N. An efficient enzymatic modification of cordycepin in ionic liquids under ultrasonic irradiation. Ultrasonics sonochemistry 2014, 21(5), 1682–1687. [Google Scholar] [CrossRef] [PubMed]
- Gao, F. Unveiling the multifaceted roles of anthocyanins: a review of their bioavailability, impacts on gut and system health, and industrial implications. Current Research in Food Science 2025, 101137. [Google Scholar] [CrossRef] [PubMed]
- Alvarez-Suarez, J.M. Novel approaches in anthocyanin research-Plant fortification and bioavailability issues. Trends in Food Science & Technology 2021, 117, 92–105. [Google Scholar]
- Singla, A. Beta-glucan as a soluble dietary fiber source: Origins, biosynthesis, extraction, purification, structural characteristics, bioavailability, biofunctional attributes, industrial utilization, and global trade. Nutrients 2024, 16(6), 900. [Google Scholar] [CrossRef]
- Xiao, Y. Synthesis, characterization, and evaluation of genistein-loaded zein/carboxymethyl chitosan nanoparticles with improved water dispersibility, enhanced antioxidant activity, and controlled release property. Foods 2020, 9(11), 1604. [Google Scholar] [CrossRef]
- Scheepens, A.; Tan, K.; Paxton, J.W. Improving the oral bioavailability of beneficial polyphenols through designed synergies. Genes & nutrition 2010, 5(1), 75–87. [Google Scholar]
- Wu, Y. Chitosan nanoparticles efficiently enhance the dispersibility, stability and selective antibacterial activity of insoluble isoflavonoids. International Journal of Biological Macromolecules 2023, 232, 123420. [Google Scholar] [CrossRef]
- Zhang, W. Microencapsulation of anthocyanins extracted from black soybean peels by whey protein/fructo-oligosaccharide contributes to improved stability, bioavailability, and ability to regulate glycolipid metabolism. Food Frontiers 2024, 5(2), 570–583. [Google Scholar] [CrossRef]
- Marsup, P. Enhancement of chemical stability and dermal delivery of Cordyceps militaris extracts by nanoemulsion. Nanomaterials 2020, 10(8), 1565. [Google Scholar] [CrossRef] [PubMed]
- Guadarrama-Escobar, O.R. Chitosan nanoparticles as oral drug carriers. International journal of molecular sciences 2023, 24(5), 4289. [Google Scholar] [CrossRef] [PubMed]
- Cheng, X. Effects of a chitosan nanoparticles encapsulation on the properties of litchi polyphenols. Food Science and Biotechnology 2023, 32(13), 1861–1871. [Google Scholar] [CrossRef] [PubMed]
- Park, B.-J.; Dhong, K.-R.; Park, H.-J. Cordyceps militaris Grown on Germinated Rhynchosia nulubilis (GRC) Encapsulated in Chitosan Nanoparticle (GCN) Suppresses Particulate Matter (PM)-Induced Lung Inflammation in Mice. International Journal of Molecular Sciences 2024, 25(19), 10642. [Google Scholar] [CrossRef]
- Kim, H.-M. Chitosan Nanoparticle-Encapsulated Cordyceps militaris Grown on Germinated Rhynchosia nulubilis Reduces Type II Alveolar Epithelial Cell Apoptosis in PM2. 5-Induced Lung Injury. International Journal of Molecular Sciences 2025, 26(3), 1105. [Google Scholar]
- Wang, M. Preparing, optimising, and evaluating chitosan nanocapsules to improve the stability of anthocyanins from Aronia melanocarpa. RSC advances 2021, 11(1), 210–218. [Google Scholar] [CrossRef]
- Zheng, S. Application of network pharmacology in traditional Chinese medicine for the treatment of digestive system diseases. Frontiers in Pharmacology 2024, 15, 1412997. [Google Scholar] [CrossRef]
- Zhang, G.-b. Network pharmacology: a new approach for Chinese herbal medicine research. Evidence-Based Complementary and Alternative Medicine 2013, 2013(1), p. 621423. [Google Scholar] [CrossRef]
- Lee, H.-J.; Cho, H.-E.; Park, H.-J. Germinated black soybean fermented with Lactobacillus pentosus SC65 alleviates DNFB-induced delayed-type hypersensitivity in C57BL/6N mice. Journal of Ethnopharmacology 2021, 265, 113236. [Google Scholar] [CrossRef]
- Kim, H.-M. Chitosan Nanoparticle-Encapsulated Cordyceps militaris Grown on Germinated Rhynchosia nulubilis Reduces Type II Alveolar Epithelial Cell Apoptosis in PM2.5-Induced Lung Injury. International Journal of Molecular Sciences 2025, 26(3), 1105. [Google Scholar] [CrossRef]
- Zeng, Y. Screening of hub genes associated with pulmonary arterial hypertension by integrated bioinformatic analysis. BioMed research international 2021, 2021(1), 6626094. [Google Scholar] [CrossRef]
- Yin, Y. Mechanism of YuPingFeng in the treatment of COPD based on network pharmacology. BioMed Research International 2020, 2020(1), p. 1630102. [Google Scholar] [CrossRef] [PubMed]
- Harel, A. GIFtS: annotation landscape analysis with GeneCards. BMC bioinformatics 2009, 10, 1–11. [Google Scholar] [CrossRef] [PubMed]
- Aziz, F.; Shoaib, N.; Rehman, A. Hub genes identification and association of key pathways with hypoxia in cancer cells: A bioinformatics analysis. Saudi Journal of Biological Sciences 2023, 30(9), p. 103752. [Google Scholar] [CrossRef] [PubMed]
- Kumar, S.; Wee, J.-J.; Kumar, K.S. Identification of Common Hub Genes in COVID-19 and Comorbidities: Insights into Shared Molecular Pathways and Disease Severity. COVID 2025. 5, 7, 105. [Google Scholar] [CrossRef]
- Chin, C.-H. cytoHubba: identifying hub objects and sub-networks from complex interactome. BMC systems biology 2014, 8, 1–7. [Google Scholar] [CrossRef]
- Dhong, K.-R.; Kwon, H.-K.; Park, H.-J. Immunostimulatory Activity of Cordyceps militaris Fermented with Pediococcus pentosaceus SC11 Isolated from a Salted Small Octopus in Cyclophosphamide-Induced Immunocompromised Mice and Its Inhibitory Activity against SARS-CoV 3CL Protease. Microorganisms 2022, 10(12), 2321. [Google Scholar] [CrossRef]
- Kwon, H.-K.; Park, H.-J. Phellinus linteus Grown on Germinated Brown Rice Inhibits IgE-Mediated Allergic Activity through the Suppression of FcεRI-Dependent Signaling Pathway In Vitro and In Vivo. Evidence-Based Complementary and Alternative Medicine 2019. [Google Scholar] [CrossRef]
- Kwon, H.-K.; Park, H.-J. Phellinus linteus Grown on Germinated Brown Rice Inhibits IgE-Mediated Allergic Activity through the Suppression of FcεRI-Dependent Signaling Pathway In Vitro and In Vivo. Evidence-Based Complementary and Alternative Medicine 2019, 2019(1), p. 1485015. [Google Scholar] [CrossRef]
- Loffredo, C. PM2. 5 as a marker of exposure to tobacco smoke and other sources of particulate matter in Cairo, Egypt. The International Journal of Tuberculosis and Lung Disease 2016, 20(3), 417–422. [Google Scholar] [CrossRef] [PubMed]
- Fehrenbach, H. Alveolar epithelial type II cell: defender of the alveolus revisited. Respiratory Research 2001, 2(1), p. 33. [Google Scholar] [CrossRef] [PubMed]
- Yang, J. The impact of bacteria-derived ultrafine dust particles on pulmonary diseases. Experimental & molecular medicine 2020, 52(3), 338–347. [Google Scholar]
- Mushtaq, N. Adhesion of Streptococcus pneumoniae to human airway epithelial cells exposed to urban particulate matter. Journal of allergy and clinical immunology 2011, 127(5), 1236–1242. e2. [Google Scholar] [CrossRef]
- Kim, D.I.; Song, M.-K.; Lee, K. Diesel exhaust particulates enhances susceptibility of LPS-induced acute lung injury through upregulation of the IL-17 cytokine-derived TGF-β1/collagen i expression and activation of NLRP3 inflammasome signaling in mice. Biomolecules 2021, 11(1), 67. [Google Scholar] [CrossRef]
- Kwon, H.-K.; Jo, W.-R.; Park, H.-J. Immune-enhancing activity of C. militaris fermented with Pediococcus pentosaceus (GRC-ON89A) in CY-induced immunosuppressed model. BMC complementary and alternative medicine 2018, 18, 1–14. [Google Scholar] [CrossRef]
- Choi, J.N. Metabolomics revealed novel isoflavones and optimal cultivation time of Cordyceps militaris fermentation. Journal of Agricultural and Food Chemistry 2010, 58(7), 4258–4267. [Google Scholar] [CrossRef]
- Park, H.-J. The Genus Cordyceps Sensu Lato: Their Chemical Constituents, Biological Activities, and Therapeutic Effects on Air Pollutants Related to Lung and Vascular Diseases. Life 2025, 15(6), 935. [Google Scholar] [CrossRef]
- Zhang, D.-N. An efficient enzymatic modification of cordycepin in ionic liquids under ultrasonic irradiation. Ultrasonics sonochemistry 2014, 21(5), 1682–1687. [Google Scholar] [CrossRef]
- Scheepens, A.; Tan, K.; Paxton, J.W. Improving the oral bioavailability of beneficial polyphenols through designed synergies. Genes & nutrition 2010, 5(1), 75–87. [Google Scholar]
- Chen, M. Cordycepin: A review of strategies to improve the bioavailability and efficacy. Phytotherapy Research 2023, 37(9), 3839–3858. [Google Scholar] [CrossRef]
- Dyawanapelly, S. Improved mucoadhesion and cell uptake of chitosan and chitosan oligosaccharide surface-modified polymer nanoparticles for mucosal delivery of proteins. Drug Delivery and Translational Research 2016, 6(4), 365–379. [Google Scholar] [CrossRef] [PubMed]
- Kwon, H.-K. Pediococcus pentosaceus-fermented Cordyceps militaris inhibits inflammatory reactions and alleviates contact dermatitis. International Journal of Molecular Sciences 2018, 19(11), 3504. [Google Scholar] [CrossRef] [PubMed]
- Yan, Z. The cytotoxic effects of fine particulate matter (PM2. 5) from different sources at the air–liquid interface exposure on a549 cells. Toxics 2023, 12(1), 21. [Google Scholar] [CrossRef] [PubMed]
- Harrison, R.M. Airborne particulate matter. Philosophical Transactions of the Royal Society A 2020, 378(2183), p. 20190319. [Google Scholar] [CrossRef]
- Wang, J. Urban particulate matter triggers lung inflammation via the ROS-MAPK-NF-κB signaling pathway. Journal of thoracic disease 2017, 9(11), p. 4398. [Google Scholar] [CrossRef]
- Arlt, V.M. Pulmonary inflammation impacts on CYP1A1-mediated respiratory tract DNA damage induced by the carcinogenic air pollutant benzo [a] pyrene. Toxicological Sciences 2015, 146(2), 213–225. [Google Scholar] [CrossRef]
- Androutsopoulos, V.P.; Tsatsakis, A.M.; Spandidos, D.A. Cytochrome P450 CYP1A1: wider roles in cancer progression and prevention. BMC cancer 2009, 9, 1–17. [Google Scholar] [CrossRef]
- Longhin, E. Cell cycle alterations induced by urban PM2. 5 in bronchial epithelial cells: characterization of the process and possible mechanisms involved. Particle and fibre toxicology 2013, 10, 1–19. [Google Scholar] [CrossRef]
- Valavanidis, A. Electron paramagnetic resonance study of the generation of reactive oxygen species catalysed by transition metals and quinoid redox cycling by inhalable ambient particulate matter. Redox Report 2005, 10(1), 37–51. [Google Scholar] [CrossRef]
- Pardo, M. Particulate matter toxicity is Nrf2 and mitochondria dependent: the roles of metals and polycyclic aromatic hydrocarbons. Chemical research in toxicology 2020, 33(5), 1110–1120. [Google Scholar] [CrossRef]
- Valgimigli, L. The unusual reaction of semiquinone radicals with molecular oxygen. The Journal of organic chemistry 2008, 73(5), 1830–1841. [Google Scholar] [CrossRef]
- Park, S. Potential toxicity of inorganic ions in particulate matter: Ion permeation in lung and disruption of cell metabolism. Science of The Total Environment 2022, 824, 153818. [Google Scholar] [CrossRef] [PubMed]
- Scheres Firak, D. Ionic Strength Effect in Fenton Reactions in the Presence of Sulfate and Its Influence on the Aqueous Particle Phase. ACS Earth and Space Chemistry 2025. 9, 3, 662–670. [Google Scholar] [CrossRef]
- Li, H. Insight into urban PM2. 5 chemical composition and environmentally persistent free radicals attributed human lung epithelial cytotoxicity. Ecotoxicology and Environmental Safety 2022, 234, 113356. [Google Scholar] [CrossRef] [PubMed]
- Di, A. Chemical characterization of seasonal PM2. 5 samples and their cytotoxicity in human lung epithelial cells (A549). International Journal of Environmental Research and Public Health 2020, 17(12), 4599. [Google Scholar] [CrossRef]
- Xing, Y.-F. The impact of PM2. 5 on the human respiratory system. Journal of thoracic disease 2016, 8(1), p. E69. [Google Scholar]
- Wei, J.; Fang, T.; Shiraiwa, M. Effects of acidity on reactive oxygen species formation from secondary organic aerosols. ACS environmental Au 2022, 2(4), 336–345. [Google Scholar] [CrossRef]
- Shahpoury, P. Influence of aerosol acidity and organic ligands on transition metal solubility and oxidative potential of fine particulate matter in urban environments. Science of the Total Environment 2024. 906, 167405. [Google Scholar] [CrossRef]
- Fang, T. Oxidative potential of particulate matter and generation of reactive oxygen species in epithelial lining fluid. Environmental science & technology 2019, 53(21), 12784–12792. [Google Scholar]
- He, M. Role of iron and oxidative stress in the exacerbation of allergic inflammation in murine lungs caused by urban particulate matter<2.5 μm and desert dust. Journal of Applied Toxicology 2019, 39(6), 855–867. [Google Scholar]
- Birben, E. Oxidative stress and antioxidant defense. World allergy organization journal 2012, 5(1), 9–19. [Google Scholar] [CrossRef]
- Wang, Y. Poldip2/Nox4 Mediates Lipopolysaccharide-Induced Oxidative Stress and Inflammation in Human Lung Epithelial Cells. Mediators of Inflammation 2022, 2022(1), p. 6666022. [Google Scholar] [CrossRef] [PubMed]
- Sul, O.-J.; Ra, S.W. Quercetin Prevents LPS-Induced Oxidative Stress and Inflammation by Modulating NOX2/ROS/NF-kB in Lung Epithelial Cells. Molecules 2021, 26(22), 6949. [Google Scholar] [CrossRef] [PubMed]
- Mandaglio-Collados, D. Analysis of Key Proinflammatory Mechanisms in Cardiovascular Pathology through Stimulation with Lipopolysaccharide and Urban Particulate Matter in Mouse Atrial Cardiomyocytes. Environmental Toxicology and Pharmacology 2025, 104652. [Google Scholar] [CrossRef] [PubMed]
- Chen, W. Effects of real-ambient PM2. 5 exposure plus lipopolysaccharide on multiple organ damage in mice. Human & Experimental Toxicology 2022, 41, 09603271211061505. [Google Scholar]
- Chen, X. Urban particulate matter (PM) suppresses airway antibacterial defence. Respiratory research 2018, 19, 1–11. [Google Scholar] [CrossRef]
- Gamboa, J.M.; Leong, K.W. In vitro and in vivo models for the study of oral delivery of nanoparticles. Advanced drug delivery reviews 2013, 65(6), 800–810. [Google Scholar] [CrossRef]
- Olatunji, O.J. Neuroprotective effects of adenosine isolated from Cordyceps cicadae against oxidative and ER stress damages induced by glutamate in PC12 cells. Environmental toxicology and pharmacology 2016, 44, 53–61. [Google Scholar] [CrossRef]
- Wang, Z. Cordycepin prevents radiation ulcer by inhibiting cell senescence via NRF2 and AMPK in rodents. Nature communications 2019, 10(1), p. 2538. [Google Scholar] [CrossRef]
- Won, K.-J. Cordycepin attenuates neointimal formation by inhibiting reactive oxygen species–mediated responses in vascular smooth muscle cells in rats. Journal of pharmacological sciences 2009, 109(3), 403–412. [Google Scholar] [CrossRef] [PubMed]
- Yamashita, Y. Insights into the potential benefits of black soybean (Glycine max L.) polyphenols in lifestyle diseases. Food & Function 2020, 11(9), 7321–7339. [Google Scholar] [CrossRef] [PubMed]
- Yu, H.M. Comparison of protective effects between cultured Cordyceps militaris and natural Cordyceps sinensis against oxidative damage. Journal of Agricultural and Food Chemistry 2006, 54(8), 3132–3138. [Google Scholar] [CrossRef] [PubMed]
- Nguyen, T.Q. Cordyceps militaris-Derived Bioactive Gels: Therapeutic and Anti-Aging Applications in Dermatology. Gels 2025, 11(1), 33. [Google Scholar] [CrossRef]
- Yu, R. Structural elucidation and biological activity of a novel polysaccharide by alkaline extraction from cultured Cordyceps militaris. Carbohydrate polymers 2009, 75(1), 166–171. [Google Scholar] [CrossRef]
- Khan, A.A. Structural, rheological, antioxidant, and functional properties of β–glucan extracted from edible mushrooms Agaricus bisporus, Pleurotus ostreatus and Coprinus attrimentarius. Bioactive Carbohydrates and Dietary Fibre 2017, 11, 67–74. [Google Scholar] [CrossRef]
- Arunachalam, K.; Sreeja, P.S.; Yang, X. The antioxidant properties of mushroom polysaccharides can potentially mitigate oxidative stress, beta-cell dysfunction and insulin resistance. Frontiers in Pharmacology 2022, 13, 874474. [Google Scholar] [CrossRef]
- Ru, Q. Lipopolysaccharide accelerates fine particulate matter-induced cell apoptosis in human lung bronchial epithelial cells. International Journal of Occupational Medicine and Environmental Health 2017, 31(2), 173–183. [Google Scholar] [CrossRef]
- Wu, Y. Air pollutants and lung regeneration: impact on the fate of lung stem cells. Environment International 2025, 199, 109525. [Google Scholar] [CrossRef]
- Liu, Q. Attenuation of PM(2.5)-induced alveolar epithelial cells and lung injury through regulation of mitochondrial fission and fusion. Part Fibre Toxicol 2023, 20(1), p. 28. [Google Scholar] [CrossRef]
- Wang, F. LPS-induced inflammatory response and apoptosis are mediated by Fra-1 upregulation and binding to YKL-40 in A549 cells. Exp Ther Med 2021, 22(6), p. 1474. [Google Scholar] [CrossRef]
- Shu, J. Comparison and evaluation of two different methods to establish the cigarette smoke exposure mouse model of COPD. Scientific Reports 2017, 7(1), p. 15454. [Google Scholar] [CrossRef] [PubMed]
- Kim, D.I.; Song, M.K.; Lee, K. Diesel Exhaust Particulates Enhances Susceptibility of LPS-Induced Acute Lung Injury through Upregulation of the IL-17 Cytokine-Derived TGF-β(1)/Collagen I Expression and Activation of NLRP3 Inflammasome Signaling in Mice. Biomolecules 2021, 11(1). [Google Scholar] [CrossRef] [PubMed]
- Fonceca, A.M. Accumulation mode particles and LPS exposure induce TLR-4 dependent and independent inflammatory responses in the lung. Respiratory Research 2018, 19(1), p. 15. [Google Scholar] [CrossRef] [PubMed]
- Shim, S.; Yoo, H.S. The Application of Mucoadhesive Chitosan Nanoparticles in Nasal Drug Delivery. Mar Drugs 2020, 18(12). [Google Scholar] [CrossRef]
- Lee, H.J. Hair Growth Promoting Effect of 4HGF Encapsulated with PGA Nanoparticles (PGA-4HGF) by β-Catenin Activation and Its Related Cell Cycle Molecules. Int J Mol Sci 2019, 20(14). [Google Scholar] [CrossRef]
- Fetterman, J.L.; Sammy, M.J.; Ballinger, S.W. Mitochondrial toxicity of tobacco smoke and air pollution. Toxicology 2017, 391, 18–33. [Google Scholar] [CrossRef]
- Pappas, R.S. Toxic elements in tobacco and in cigarette smoke: inflammation and sensitization. Metallomics 2011, 3(11), 1181–1198. [Google Scholar] [CrossRef]
- Wang, Q.; Liu, S. The effects and pathogenesis of PM2. 5 and its components on chronic obstructive pulmonary disease. International Journal of Chronic Obstructive Pulmonary Disease 2023, 493–506. [Google Scholar] [CrossRef]
- Korsgren, M. Inhalation of LPS induces inflammatory airway responses mimicking characteristics of chronic obstructive pulmonary disease. Clinical physiology and functional imaging 2012, 32(1), 71–79. [Google Scholar] [CrossRef]
- Feng, Q.; Yu, Y.-Z.; Meng, Q.-H. Blocking tumor necrosis factor-α delays progression of chronic obstructive pulmonary disease in rats through inhibiting MAPK signaling pathway and activating SOCS3/TRAF1. Experimental and therapeutic medicine 2021, 22(5), p. 1311. [Google Scholar] [CrossRef] [PubMed]
- Ahmadi, A. p38 MAPK signaling in chronic obstructive pulmonary disease pathogenesis and inhibitor therapeutics. Cell Communication and Signaling 2023, 21(1), p. 314. [Google Scholar] [CrossRef] [PubMed]
- Schuliga, M. NF-kappaB signaling in chronic inflammatory airway disease. Biomolecules 2015, 5(3), 1266–1283. [Google Scholar] [CrossRef] [PubMed]
- Sidletskaya, K.; Vitkina, T.; Denisenko, Y. The role of toll-like receptors 2 and 4 in the pathogenesis of chronic obstructive pulmonary disease. International journal of chronic obstructive pulmonary disease 2020, 1481–1493. [Google Scholar] [CrossRef]
- Zhang, Q. scRNA-seq and scATAC-seq analyses highlight the role of TNF signaling pathway in chronic obstructive pulmonary disease model mice. PloS one 2025, 20(5), e0322538. [Google Scholar] [CrossRef]
- Yi, G. A large lung gene expression study identifying IL1B as a novel player in airway inflammation in COPD airway epithelial cells. Inflammation Research 2018, 67, 539–551. [Google Scholar] [CrossRef]
- Jing, H. Inhibition of CXC motif chemokine 10 (CXCL10) protects mice from cigarette smoke-induced chronic obstructive pulmonary disease. Medical Science Monitor: International Medical Journal of Experimental and Clinical Research 2018, 24, 5748. [Google Scholar] [CrossRef]
- Lappalainen, U. Interleukin-1β causes pulmonary inflammation, emphysema, and airway remodeling in the adult murine lung. American journal of respiratory cell and molecular biology 2005, 32(4), 311–318. [Google Scholar] [CrossRef]
- Liu, C.-W. PM 2.5-induced oxidative stress increases intercellular adhesion molecule-1 expression in lung epithelial cells through the IL-6/AKT/STAT3/NF-κB-dependent pathway. Particle and fibre toxicology 2018, 15, 1–16. [Google Scholar] [CrossRef]
- He, M. PM2. 5-induced lung inflammation in mice: Differences of inflammatory response in macrophages and type II alveolar cells. Journal of Applied Toxicology 2017, 37(10), 1203–1218. [Google Scholar] [CrossRef]
- Wu, Y.-F. Inactivation of MTOR promotes autophagy-mediated epithelial injury in particulate matter-induced airway inflammation. Autophagy 2020, 16(3), 435–450. [Google Scholar] [CrossRef]
- Yanagisawa, R. Gene expression analysis of murine lungs following pulmonary exposure to Asian sand dust particles. Experimental Biology and Medicine 2007, 232(8), 1109–1118. [Google Scholar] [CrossRef] [PubMed]
- Russo, R.C.; Ryffel, B. The chemokine system as a key regulator of pulmonary fibrosis: Converging pathways in human idiopathic pulmonary fibrosis (IPF) and the bleomycin-induced lung fibrosis model in mice. Cells 2024, 13(24), 2058. [Google Scholar] [CrossRef] [PubMed]
- Li, Y.-J. Nrf2 lowers the risk of lung injury via modulating the airway innate immune response induced by diesel exhaust in mice. Biomedicines 2020, 8(10), 443. [Google Scholar] [CrossRef] [PubMed]
- Liu, J. Increased alveolar epithelial TRAF6 via autophagy-dependent TRIM37 degradation mediates particulate matter-induced lung metastasis. Autophagy 2022, 18(5), 971–989. [Google Scholar] [CrossRef]
- Chen, Q. TRPC6-dependent Ca2+ signaling mediates airway inflammation in response to oxidative stress via ERK pathway. Cell Death & Disease 2020, 11(3), p. 170. [Google Scholar]
- Chen, Q.-Z. TRPC6 modulates adhesion of neutrophils to airway epithelial cells via NF-κB activation and ICAM-1 expression with ozone exposure. Experimental cell research 2019, 377(1–2), 56–66. [Google Scholar] [CrossRef]
- Vlag, J.; Nijenhuis, T. A Putative Role for TRPC6 in Immune-Mediated Kidney Injury. 2023.
- Lindemann, O. Intravascular adhesion and recruitment of neutrophils in response to CXCL1 depends on their TRPC6 channels. Journal of Molecular Medicine 2020, 98(3), 349–360. [Google Scholar] [CrossRef]
- Dhong, K.-R. Identification of TRPC6 as a novel diagnostic biomarker of PM-induced chronic obstructive pulmonary disease using machine learning models. Genes 2023, 14(2), 284. [Google Scholar] [CrossRef]
- Du, B. An insight into anti-inflammatory effects of fungal beta-glucans. Trends in Food Science & Technology 2015, 41(1), 49–59. [Google Scholar] [CrossRef]
- Moniruzzaman, M. A review on pharmacological insights of edible and medicinal mushroom based β-glucans. Applied Biological Chemistry 2025, 68(1), p. 41. [Google Scholar] [CrossRef]
- Zhu, W. β-Glucan modulates the lipopolysaccharide-induced innate immune response in rat mammary epithelial cells. International immunopharmacology 2013, 15(2), 457–465. [Google Scholar] [CrossRef]
- Smiderle, F.R. Anti-inflammatory properties of the medicinal mushroom Cordyceps militaris might be related to its linear (1→ 3)-β-D-glucan. PLoS One 2014, 9(10), e110266. [Google Scholar] [CrossRef]
- Chen, X.-Y. Black rice anthocyanins suppress metastasis of breast cancer cells by targeting RAS/RAF/MAPK pathway. BioMed research international 2015, 2015(1), 414250. [Google Scholar] [CrossRef] [PubMed]
- Kim, J.N. Black soybean anthocyanins attenuate inflammatory responses by suppressing reactive oxygen species production and mitogen activated protein kinases signaling in lipopolysaccharide-stimulated macrophages. Nutrition Research and Practice 2017, 11(5), 357–364. [Google Scholar] [CrossRef] [PubMed]
- Choi, Y.H.; Kim, G.-Y.; Lee, H.H. Anti-inflammatory effects of cordycepin in lipopolysaccharide-stimulated RAW 264.7 macrophages through Toll-like receptor 4-mediated suppression of mitogen-activated protein kinases and NF-κB signaling pathways. In Drug design, development and therapy; 2014; pp. 1941–1953. [Google Scholar]
- Lei, J. Cordycepin inhibits LPS-induced acute lung injury by inhibiting inflammation and oxidative stress. European Journal of Pharmacology 2018, 818, 110–114. [Google Scholar] [CrossRef] [PubMed]
- Feng, G.; Sun, B.; Li, T.-z. Daidzein attenuates lipopolysaccharide-induced acute lung injury via toll-like receptor 4/NF-kappaB pathway. International immunopharmacology 2015, 26(2), 392–400. [Google Scholar] [CrossRef]
- Kojima, K. Isoflavone aglycones attenuate cigarette smoke-induced emphysema via suppression of neutrophilic inflammation in a COPD murine model. Nutrients 2019, 11(9), 2023. [Google Scholar] [CrossRef]
| Compounds | Concentrations of analytes in 40mg/mL GRC (μg/mL) a | Contents (mg/g-GRC) b |
|---|---|---|
| Adenosine | 14.19±0.07 | 0.36±0.00 |
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2026 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).




