Submitted:
17 April 2026
Posted:
20 April 2026
You are already at the latest version
Abstract
Keywords:
1. Introduction
2. Materials and Methods
2.1. Clinical Samples
2.2. Cell Culture and Transfection
2.3. Gene Expression Profiling Data Acquisition and Processing
2.4. Differentially Expressed Gene Identification
2.5. GO and KEGG Enrichment Analysis
2.6. Identification of Hub MEMRG-DEGs
2.7. Validation of Hub MEMRG-DEGs Expression and Evaluation of Diagnostic Efficacy
2.8. Single-Sample Gene Set Enrichment Analysis
2.9. Single-Cell Transcriptomic Data Analysis
2.10. Western Blot (WB)
2.11. Immunofluorescence (IF) Staining
2.12. Measurement of Mitochondrial Membrane Potential
2.13. Statistical Analysis
3. Results
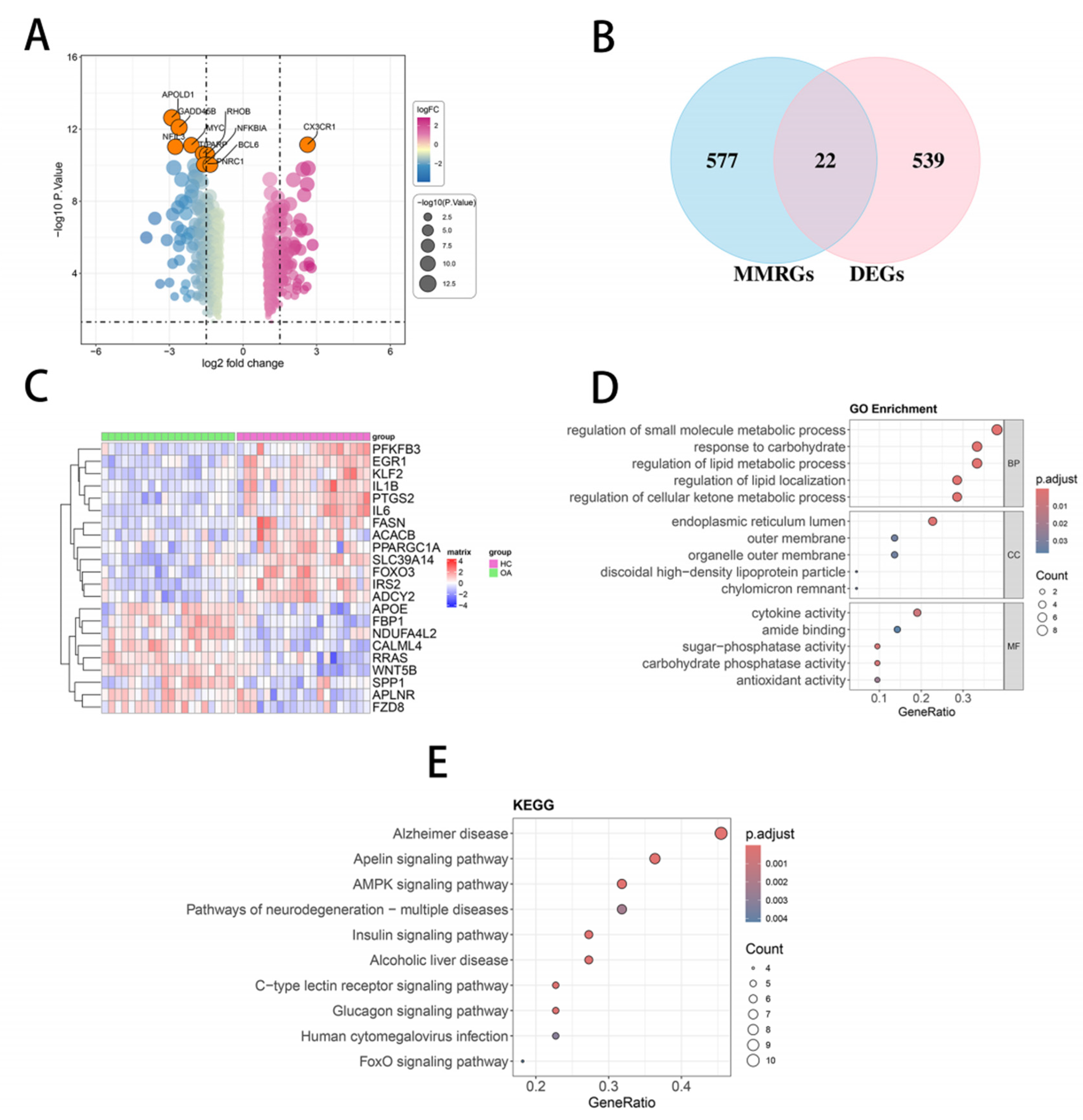
3.1. Identification and Functional Enrichment of MEMRG-DEGs
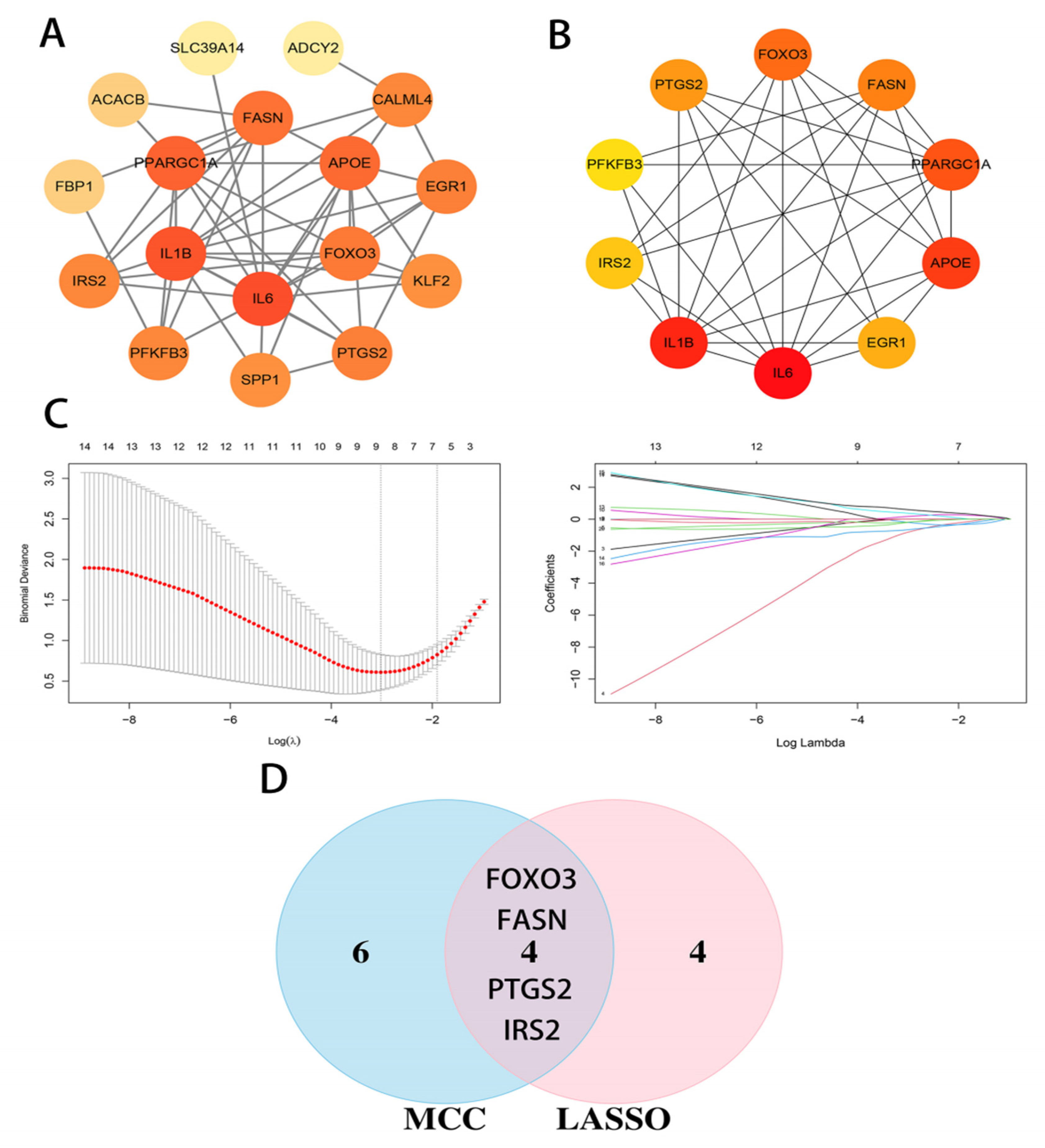
3.2. Protein-Protein Interaction Network and Hub Gene Screens
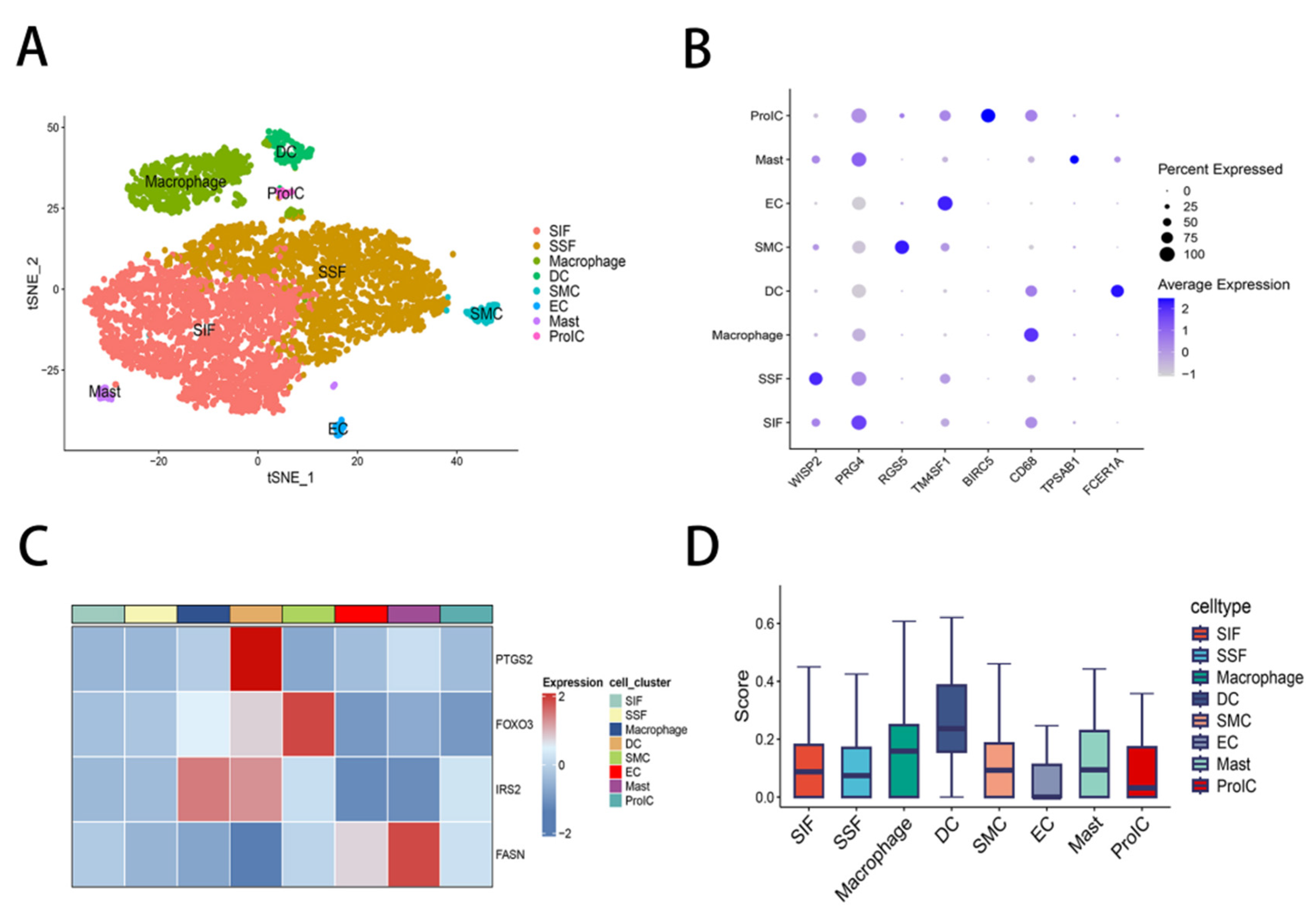
3.3. scRNA-Seq Identified Localization of Hub MEMRG-DEGs
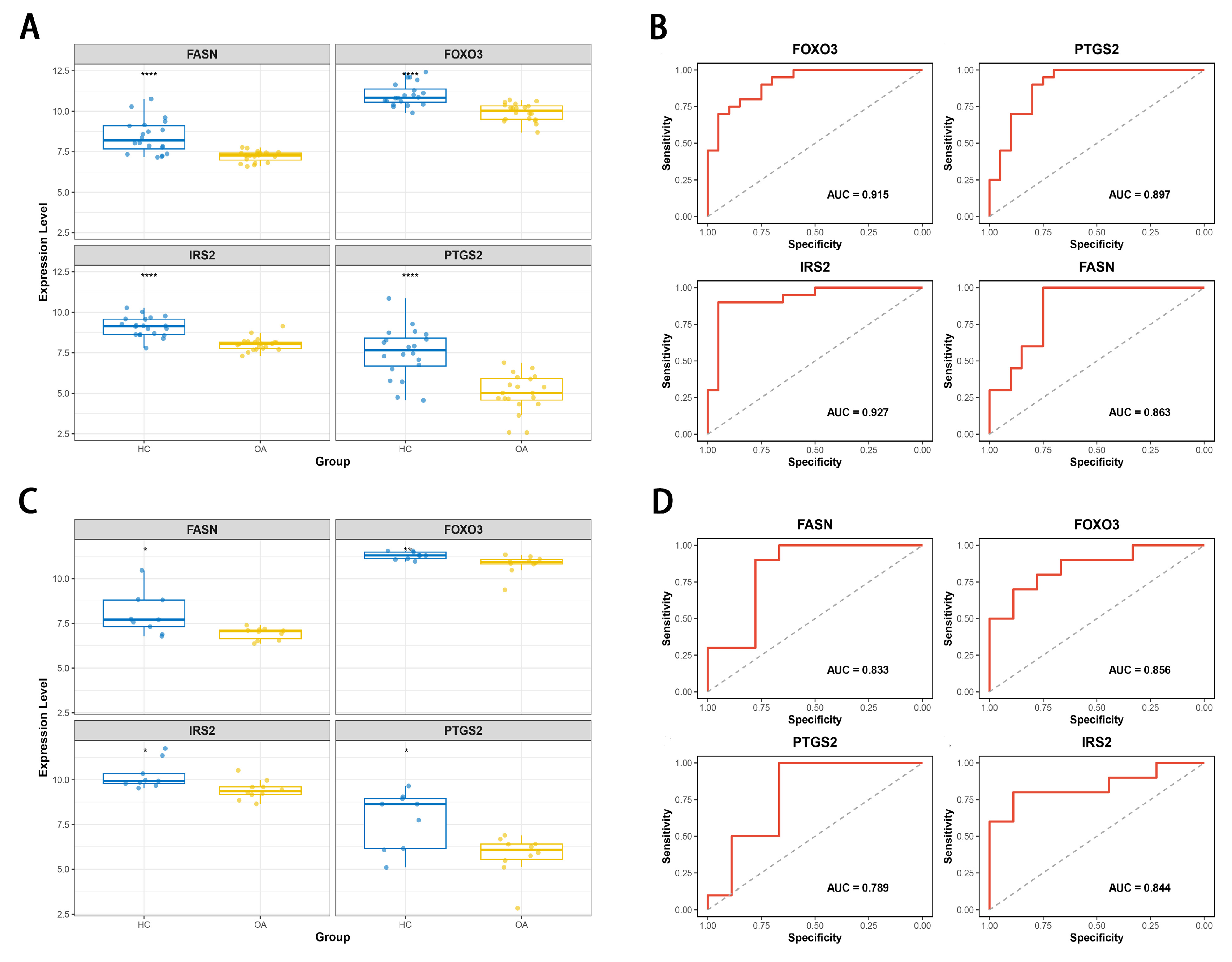
3.4. Expression Validation and Evaluation of Diagnostic Efficacy for Hub MEMRG-DEGs
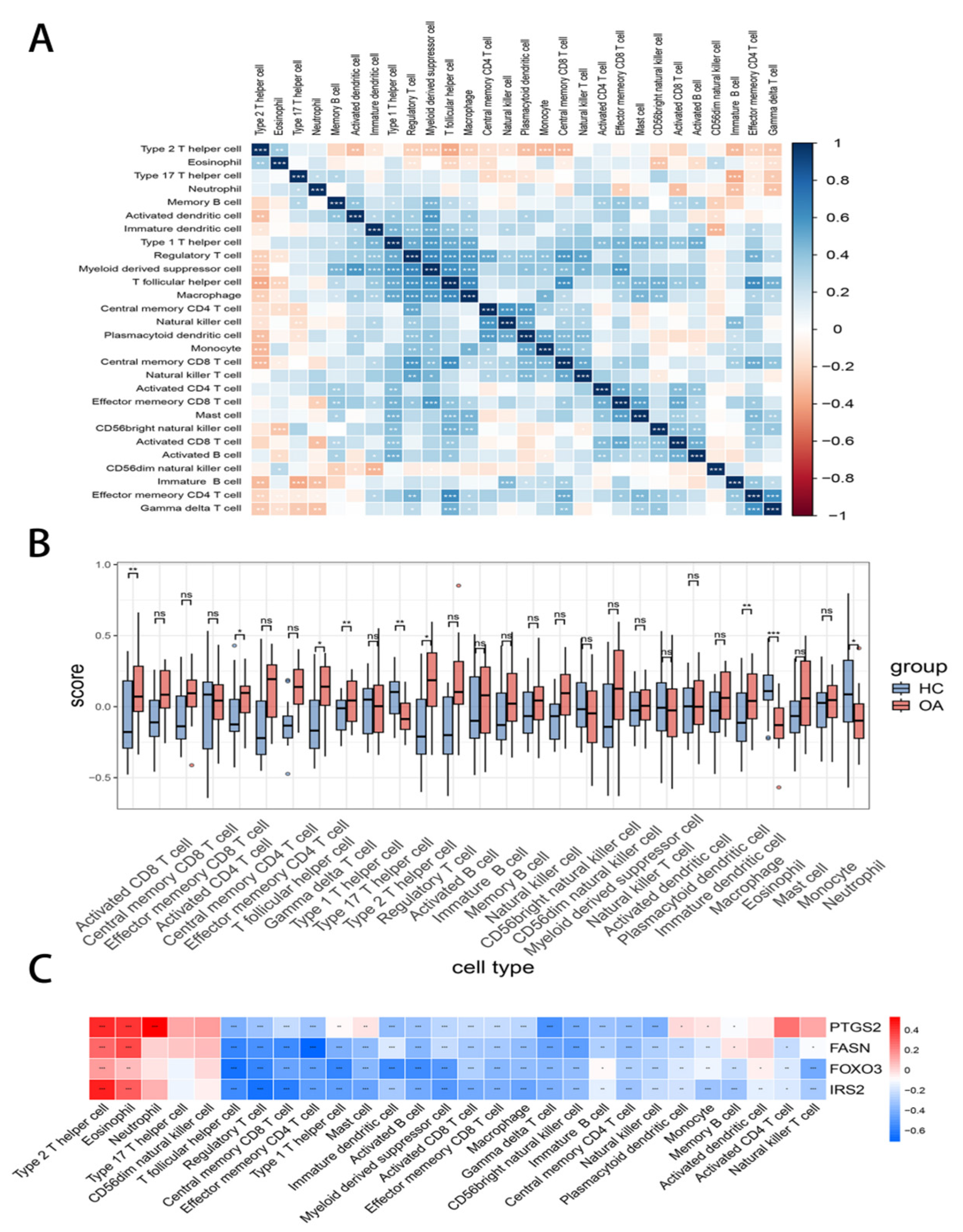
3.5. IRS2 Demonstrated Significant Correlation with Macrophages
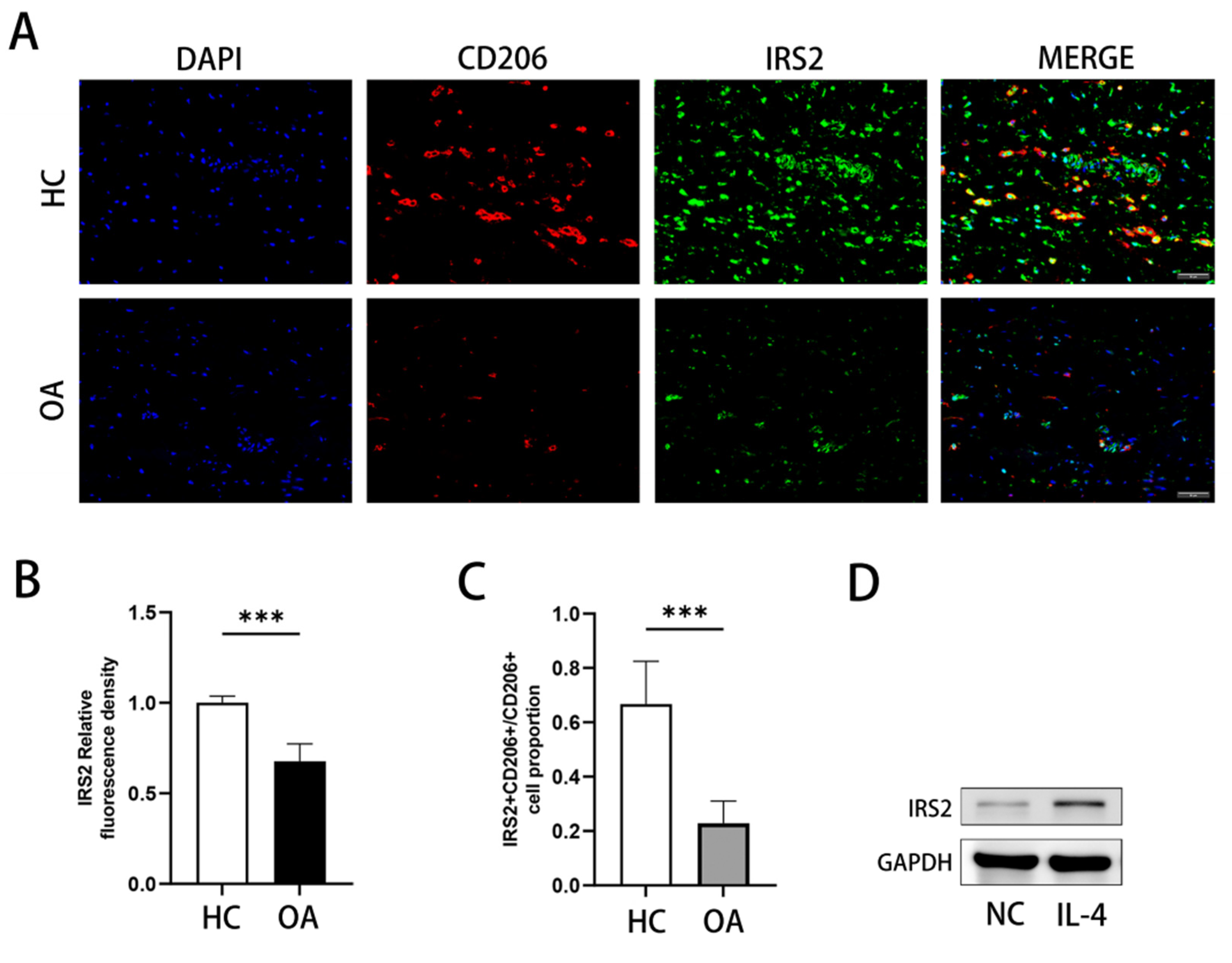
3.6. IRS2 Was Downregulated in OA Synovium
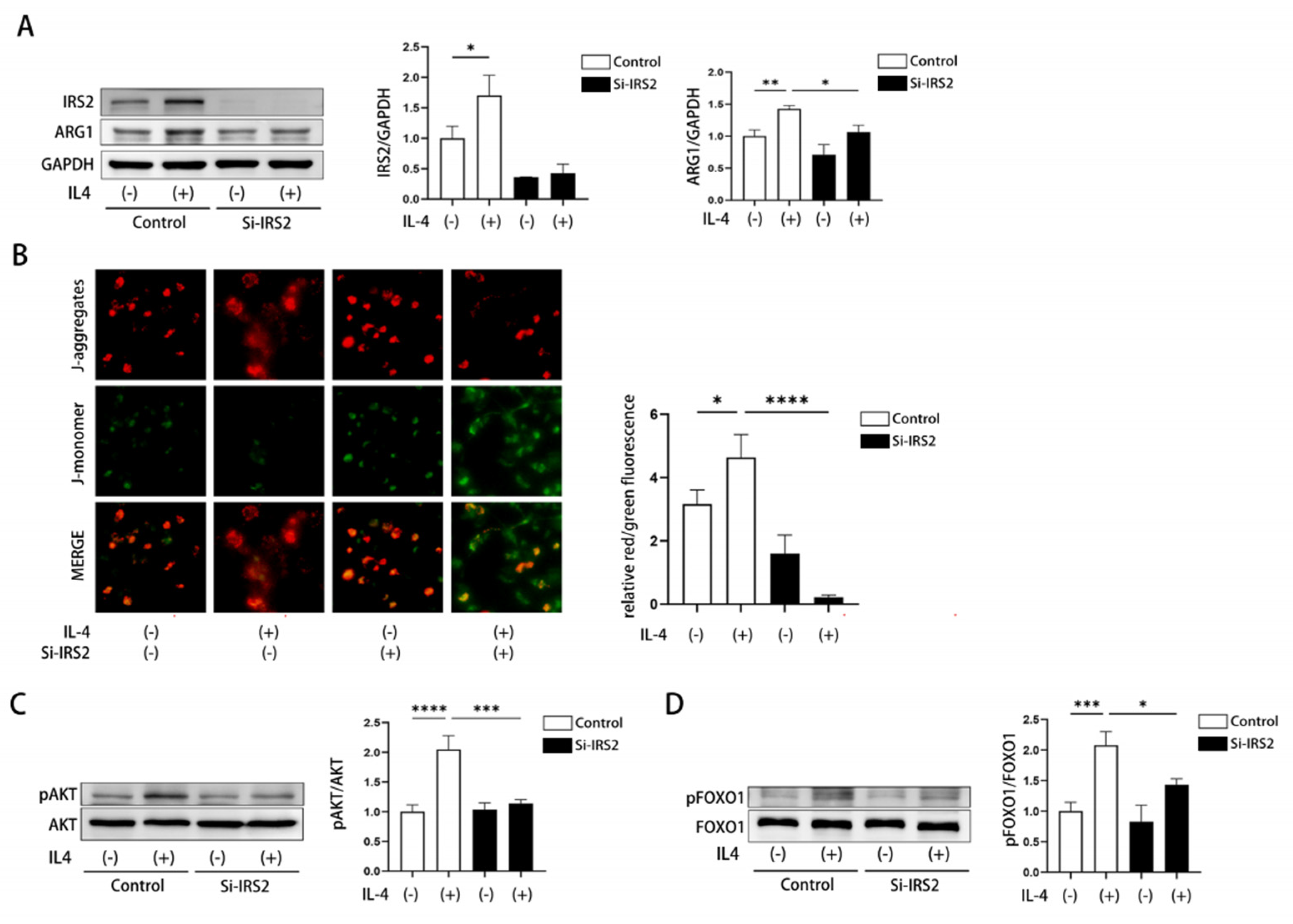
3.7. Downregulation of IRS2 Impaired M2 Macrophage Polarization
4. Discussion
5. Conclusions
Supplementary Materials
Author Contributions
Funding
Institutional Review Board Statement
Data Availability Statement
Acknowledgments
Conflicts of Interest
References
- Loeser RF, Goldring SR, Scanzello CR, Goldring MB. Osteoarthritis: a disease of the joint as an organ. Arthritis Rheum. 2012;64(6):1697–707. [CrossRef]
- Courties A, Kouki I, Soliman N, Mathieu S, Sellam J. Osteoarthritis year in review 2024: epidemiology and therapy. Osteoarthr Cartil. 2024;32(11):1397–404. [CrossRef]
- Leifer VP, Katz JN, Losina E. The burden of OA-health services and economics. Osteoarthr Cartil. 2022;30(1):10–6. [CrossRef]
- Benito MJ, Veale DJ, Fitzgerald O, van den Berg WB, Bresnihan B. Synovial tissue inflammation in early and late osteoarthritis. Ann Rheum Dis. 2005;64(9):1263–7. [CrossRef]
- Atukorala I, Kwoh CK, Guermazi A, Roemer FW, Boudreau RM, Hannon MJ, et al. Synovitis in knee osteoarthritis: a precursor of disease? Ann Rheum Dis. 2016;75(2):390–5. [CrossRef]
- Leopoldino AO, Machado GC, Ferreira PH, Pinheiro MB, Day R, McLachlan AJ, et al. Paracetamol versus placebo for knee and hip osteoarthritis. Cochrane Database Syst Rev. 2019;(2). [CrossRef]
- Klein-Wieringa IR, de Lange-Brokaar BJE, Yusuf E, Andersen SN, Kwekkeboom JC, Kroon HM, et al. Inflammatory cells in patients with endstage knee osteoarthritis: a comparison between the synovium and the infrapatellar fat pad. J Rheumatol. 2016;43(4):771–8. [CrossRef]
- Klein-Wieringa IR, Kloppenburg M, Bastiaansen-Jenniskens YM, Yusuf E, Kwekkeboom JC, El-Bannoudi H, et al. The infrapatellar fat pad of patients with osteoarthritis has an inflammatory phenotype. Ann Rheum Dis. 2011;70(5):851–7. [CrossRef]
- Zhang HY, Lin CX, Zeng C, Wang ZY, Wang H, Lu JS, et al. Synovial macrophage M1 polarisation exacerbates experimental osteoarthritis partially through R-spondin-Ann Rheum Dis. 2018;77(10):1524–34. [CrossRef]
- Xue CC, Tian J, Cui ZP, Liu Y, Sun DW, Xiong MT, et al. Reactive oxygen species (ROS)-mediated M1 macrophage-dependent nanomedicine remodels inflammatory microenvironment for osteoarthritis recession. Bioact Mater. 2024;33:545–61. [CrossRef]
- Van den Bossche J, Baardman J, Otto NA, van der Velden S, Neele AE, van den Berg SM, et al. Mitochondrial dysfunction prevents repolarization of inflammatory macrophages. Cell Rep. 2016;17(3):684–96. [CrossRef]
- Protasoni M, Zeviani M. Mitochondrial structure and bioenergetics in normal and disease conditions. Int J Mol Sci. 2021;22(2). [CrossRef]
- Loftus RM, Finlay DK. Immunometabolism: cellular metabolism turns immune regulator. J Biol Chem. 2016;291(1):1–10. [CrossRef]
- Gauthier T, Yao C, Dowdy T, Jin WW, Lim YJ, Patino LC, et al. TGF-β uncouples glycolysis and inflammation in macrophages and controls survival during sepsis. Sci Signal. 2023;16(797). [CrossRef]
- Palmieri EM, Holewinski R, McGinity CL, Pierri CL, Maio N, Weiss JM, et al. Pyruvate dehydrogenase operates as an intramolecular nitroxyl generator during macrophage metabolic reprogramming. Nat Commun. 2023;14(1). [CrossRef]
- Peace CG, O’Carroll SM, O’Neill LAJ. Fumarate hydratase as a metabolic regulator of immunity. Trends Cell Biol. 2024;34(6):442–50. [CrossRef]
- He XX, Huang YJ, Hu CL, Xu QQ, Wei QJ. Songorine modulates macrophage polarization and metabolic reprogramming to alleviate inflammation in osteoarthritis. Front Immunol. 2024;15. [CrossRef]
- Stump CS, Short KR, Bigelow ML, Schimke JM, Nair KS. Effect of insulin on human skeletal muscle mitochondrial ATP production, protein synthesis, and mRNA transcripts. Proc Natl Acad Sci USA. 2003;100(13):7996–8001. [CrossRef]
- Yu TZ, Robotham JL, Yoon Y. Increased production of reactive oxygen species in hyperglycemic conditions requires dynamic change of mitochondrial morphology. Proc Natl Acad Sci USA. 2006;103(8):2653–8. [CrossRef]
- Cheng ZY, Tseng Y, White MF. Insulin signaling meets mitochondria in metabolism. Trends Endocrinol Metab. 2010;21(10):589–98. [CrossRef]
- Kubota T, Inoue M, Kubota N, Takamoto I, Mineyama T, Iwayama K, et al. Downregulation of macrophage Irs2 by hyperinsulinemia impairs IL-4-indeuced M2a-subtype macrophage activation in obesity. Nat Commun. 2018;9. [CrossRef]
- Leek JT, Johnson WE, Parker HS, Jaffe AE, Storey JD. The sva package for removing batch effects and other unwanted variation in high-throughput experiments. Bioinformatics. 2012;28(6):882–3. [CrossRef]
- Zhang ZW, Jin GW, Zhao J, Deng SQ, Chen F, Wuyun G, et al. Mitochondrial energy metabolism correlates with an immunosuppressive tumor microenvironment and poor prognosis in esophageal squamous cell carcinoma. Comput Struct Biotechnol J. 2023;21:4118–33. [CrossRef]
- Ritchie ME, Phipson B, Wu D, Hu YF, Law CW, Shi W, et al. limma powers differential expression analyses for RNA-sequencing and microarray studies. Nucleic Acids Res. 2015;43(7). [CrossRef]
- Yu GC, Wang LG, Han YY, He QY. clusterProfiler: an R package for comparing biological themes among gene clusters. OMICS. 2012;16(5):284–7. [CrossRef]
- Ashburner M, Ball CA, Blake JA, Botstein D, Butler H, Cherry JM, et al. Gene ontology: tool for the unification of biology. Nat Genet. 2000;25(1):25–9. [CrossRef]
- Kanehisa M, Goto S. KEGG: Kyoto Encyclopedia of Genes and Genomes. Nucleic Acids Res. 2000;28(1):27–30. [CrossRef]
- Szklarczyk D, Gable AL, Lyon D, Junge A, Wyder S, Huerta-Cepas J, et al. STRING v11: protein-protein association networks with increased coverage, supporting functional discovery in genome-wide experimental datasets. Nucleic Acids Res. 2019;47(D1):D607–13. [CrossRef]
- Shannon P, Markiel A, Ozier O, Baliga NS, Wang JT, Ramage D, et al. Cytoscape: a software environment for integrated models of biomolecular interaction networks. Genome Res. 2003;13(11):2498–504. [CrossRef]
- Chin CH, Chen SH, Wu HH, Ho CW, Ko MT, Lin CY. cytoHubba: identifying hub objects and sub-networks from complex interactome. BMC Syst Biol. 2014;8(Suppl 4):S11. [CrossRef]
- Hänzelmann S, Castelo R, Guinney J. GSVA: gene set variation analysis for microarray and RNA-seq data. BMC Bioinformatics. 2013;14:7. [CrossRef]
- Mangiola S, Doyle MA, Papenfuss AT. Interfacing Seurat with the R tidy universe. Bioinformatics. 2021;37(22):4100–7. [CrossRef]
- Korsunsky I, Millard N, Fan J, Slowikowski K, Zhang F, Wei K, et al. Fast, sensitive and accurate integration of single-cell data with Harmony. Nat Methods. 2019;16(12):1289–96. [CrossRef]
- Hu CX, Li TY, Xu YQ, Zhang XX, Li F, Bai J, et al. CellMarker 2.0: an updated database of manually curated cell markers in human/mouse and web tools based on scRNA-seq data. Nucleic Acids Res. 2023;51(D1):D870–6. [CrossRef]
- Aibar S, González-Blas CB, Moerman T, Van AHT, Imrichova H, Hulselmans G, et al. SCENIC: single-cell regulatory network inference and clustering. Nat Methods. 2017;14(11):1083–6. [CrossRef]
- Sellam J, Berenbaum F. The role of synovitis in pathophysiology and clinical symptoms of osteoarthritis. Nat Rev Rheumatol. 2010;6(11):625–35. [CrossRef]
- Li YS, Luo W, Zhu SA, Lei GH. T cells in osteoarthritis: alterations and beyond. Front Immunol. 2017;8:356. [CrossRef]
- Zhang H, Cai D, Bai X. Macrophages regulate the progression of osteoarthritis. Osteoarthr Cartil. 2020;28(5):555–61. [CrossRef]
- Zhao K, Ruan JQ, Nie LY, Ye XM, Li JB. Effects of synovial macrophages in osteoarthritis. Front Immunol. 2023;14:1164137. [CrossRef]
- Liu DD, Qin HW, Gao Y, Sun MY, Wang MN. Cardiovascular disease: mitochondrial dynamics and mitophagy crosstalk mechanisms with novel programmed cell death and macrophage polarisation. Pharmacol Res. 2024;206:107258. [CrossRef]
- Wang WY, Chu YR, Lu YK, Xu J, Zhao WX, Liang Z, et al. Skatole alleviates osteoarthritis by reprogramming macrophage polarization and protecting chondrocytes. Research. 2025;8:0604. [CrossRef]
- Xie JW, Huang ZY, Yu XJ, Zhou L, Pei FX. Clinical implications of macrophage dysfunction in the development of osteoarthritis of the knee. Cytokine Growth Factor Rev. 2019;46:36–44. [CrossRef]
- Qi ZH, Zhu JP, Cai WS, Lou CB, Li ZY. The role and intervention of mitochondrial metabolism in osteoarthritis. Mol Cell Biochem. 2024;479(6):1513–24. [CrossRef]
- Kumar P, Liu C, Hsu JW, Chacko S, Minard C, Jahoor F, et al. Glycine and N-acetylcysteine (GlyNAC) supplementation in older adults improves glutathione deficiency, oxidative stress, mitochondrial dysfunction, inflammation, insulin resistance, endothelial dysfunction, genotoxicity, muscle strength, and cognition: results of a pilot clinical trial. Clin Transl Med. 2021;11(3):e372. [CrossRef]
- Tan ZK, Liu QQ, Chen HJ, Zhang ZY, Wang Q, Mu YS, et al. Pectolinarigenin alleviated septic acute kidney injury via inhibiting JAK2/STAT3 signaling and mitochondria dysfunction. Biomed Pharmacother. 2023;159:114286. [CrossRef]
- Luo H, Li LF, Han S, Liu T. The role of monocyte/macrophage chemokines in pathogenesis of osteoarthritis: a review. Int J Immunogenet. 2024;51(3):130–42. [CrossRef]
- de Lange-Brokaar BJE, Ioan-Facsinay A, van Osch GJVM, Zuurmond AM, Schoones J, Toes REM, et al. Synovial inflammation, immune cells and their cytokines in osteoarthritis: a review. Osteoarthr Cartil. 2012;20(12):1484–99. [CrossRef]
- Liu BL, Zhang MQ, Zhao JM, Zheng M, Yang H. Imbalance of M1/M2 macrophages is linked to severity level of knee osteoarthritis. Exp Ther Med. 2018;16(6):5009–14. [CrossRef]
- Van den Bossche J, O’Neill LA, Menon D. Macrophage immunometabolism: where are we (going)? Trends Immunol. 2017;38(6):395–406. [CrossRef]
- Zhang L, Chen X, Cai PQ, Sun H, Shen SY, Guo BS, et al. Reprogramming mitochondrial metabolism in synovial macrophages of early osteoarthritis by a camouflaged meta-defensome. Adv Mater. 2022;34(30):e2202715. [CrossRef]
- Heller NM, Qi XL, Junttila IS, Shirey KA, Vogel SN, Paul WE, et al. Type I IL-4Rs selectively activate IRS-2 to induce target gene expression in macrophages. Sci Signal. 2008;1(51):ra17. [CrossRef]
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2026 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).



