Submitted:
17 April 2026
Posted:
20 April 2026
You are already at the latest version
Abstract
Keywords:
1. Introduction
2. Results and Discussion
2.1. Circular Dichroism (CD) Studies
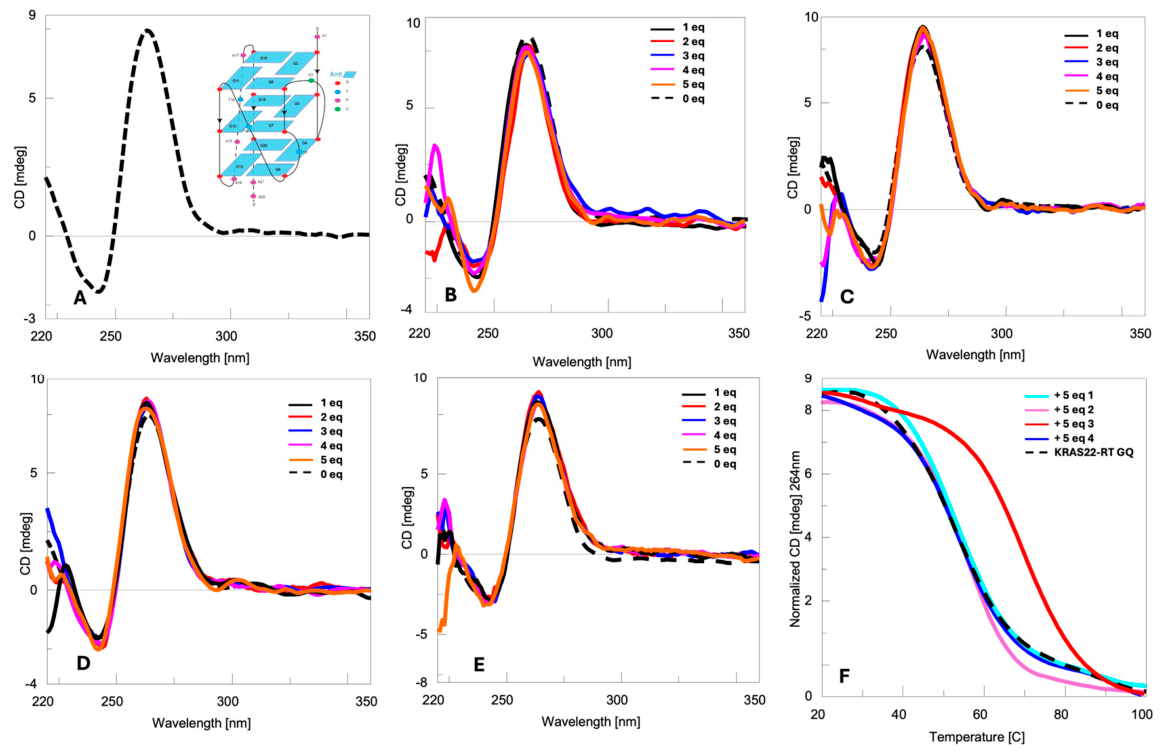
2.2. NMR Studies
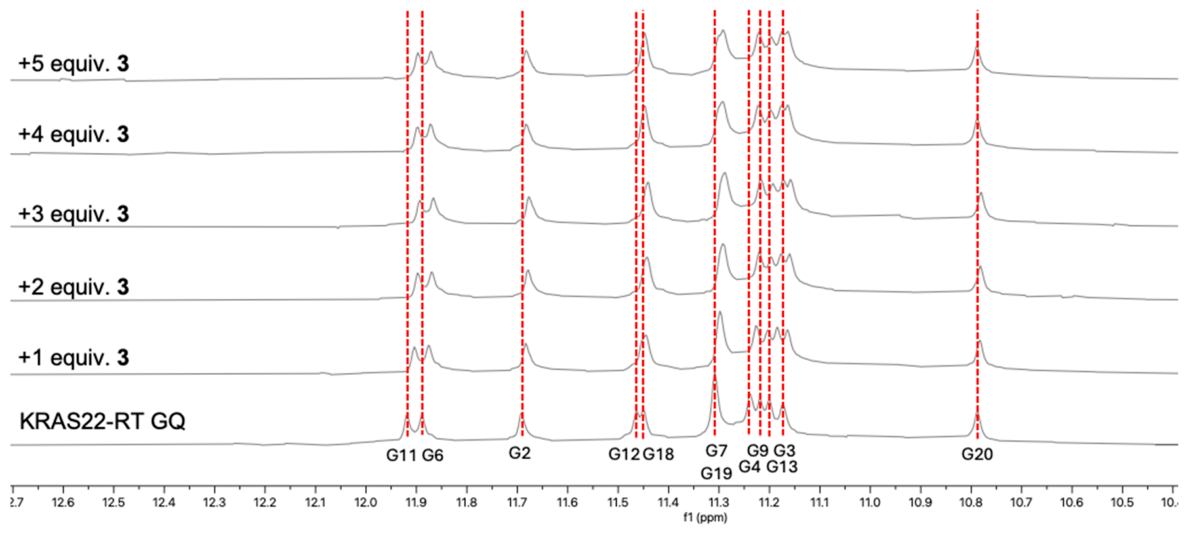
2.3. Evaluation of the Antiproliferative Activity of Compounds 1–4
3. Materials and Methods
3.1. Preparation of Oligonucleotide KRAS-22RT
3.2. Preparation of KRAS-22RT GQ
3.3. Circular Dichroism (CD) Experiments
3.4. NMR Titration and Melting Experiments
3.5. Cell Lines and Culture Conditions
3.6. Cell Proliferation Assay
4. Conclusions
Supplementary Materials
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Acknowledgments
Conflicts of Interest
Abbreviations
References
- Barbieri, F.; Tabanelli, G.; Braschi, G.; Bassi, D.; Morandi, S.; Šimat, V.; Čagalj, M.; Gardini, F.; Montanari, C. Mediterranean Plants and Spices as a Source of Bioactive Essential Oils for Food Applications: Chemical Characterisation and In Vitro Activity. Int. J. Mol. Sci. 2025, 26, 3875. [Google Scholar] [CrossRef]
- Chileh-Chelh, T.; Ezzaitouni, M.; Belarbi, E.-H.; Guil-Guerrero, J.L. Mediterranean Wild Edible Plants as Promising Sources of Less-Polar Bioactive Compounds: Fatty Acids, Carotenoids, Tocopherols, and Sterols. Journal of Food Composition and Analysis 2025, 148, 108370. [Google Scholar] [CrossRef]
- Vella, F.M.; Pignone, D.; Laratta, B. The Mediterranean Species Calendula Officinalis and Foeniculum Vulgare as Valuable Source of Bioactive Compounds. Molecules 2024, 29, 3594. [Google Scholar] [CrossRef] [PubMed]
- Gao, L.; Liu, X.; Luo, X.; Lou, X.; Li, P.; Li, X.; Liu, X. Antiaging Effects of Dietary Supplements and Natural Products. Front. Pharmacol. 2023, 14. [Google Scholar] [CrossRef] [PubMed]
- Bjørklund, G.; Shanaida, M.; Lysiuk, R.; Butnariu, M.; Peana, M.; Sarac, I.; Strus, O.; Smetanina, K.; Chirumbolo, S. Natural Compounds and Products from an Anti-Aging Perspective. Molecules 2022, 27, 7084. [Google Scholar] [CrossRef]
- Latif, R.; Nawaz, T. Medicinal Plants and Human Health: A Comprehensive Review of Bioactive Compounds, Therapeutic Effects, and Applications. Phytochemistry Reviews 2026, 25, 2299–2342. [Google Scholar] [CrossRef]
- Twaij, B.M.; Hasan, Md.N. Bioactive Secondary Metabolites from Plant Sources: Types, Synthesis, and Their Therapeutic Uses. International Journal of Plant Biology 2022, 13, 4–14. [Google Scholar] [CrossRef]
- Aware, C.B.; Patil, D.N.; Suryawanshi, S.S.; Mali, P.R.; Rane, M.R.; Gurav, R.G.; Jadhav, J.P. Natural Bioactive Products as Promising Therapeutics: A Review of Natural Product-Based Drug Development. South African Journal of Botany 2022, 151, 512–528. [Google Scholar] [CrossRef]
- Wang, K.-B.; Wang, Y.; Dickerhoff, J.; Yang, D. DNA G-Quadruplexes as Targets for Natural Product Drug Discovery. Engineering 2024, 38, 39–51. [Google Scholar] [CrossRef]
- Mazzini, S.; Princiotto, S.; Musso, L.; Passarella, D.; Beretta, G.L.; Perego, P.; Dallavalle, S. Synthesis and Investigation of the G-Quadruplex Binding Properties of Kynurenic Acid Derivatives with a Dihydroimidazoquinoline-3,5-Dione Core. Molecules 2022, 27, 2791. [Google Scholar] [CrossRef]
- Marzano, M.; Nolli, M.G.; D’Errico, S.; Falanga, A.P.; Terracciano, M.; Dardano, P.; De Stefano, L.; Piccialli, G.; Borbone, N.; Oliviero, G. Enhancing G-Quadruplex-Based DNA Nanotechnology: New Lipophilic DNA G-Quadruplexes with TBDPS Modifications. RSC Adv. 2025, 15, 17933–17945. [Google Scholar] [CrossRef] [PubMed]
- Spiegel, J.; Adhikari, S.; Balasubramanian, S. The Structure and Function of DNA G-Quadruplexes. Trends Chem. 2020, 2, 123–136. [Google Scholar] [CrossRef] [PubMed]
- Lam, E.Y.N.; Beraldi, D.; Tannahill, D.; Balasubramanian, S. G-Quadruplex Structures Are Stable and Detectable in Human Genomic DNA. Nat. Commun. 2013, 4. [Google Scholar] [CrossRef]
- Kosiol, N.; Juranek, S.; Brossart, P.; Heine, A.; Paeschke, K. G-Quadruplexes: A Promising Target for Cancer Therapy. Mol. Cancer 2021, 20, 40. [Google Scholar] [CrossRef]
- Xu, J.; Huang, H.; Zhou, X. G-Quadruplexes in Neurobiology and Virology: Functional Roles and Potential Therapeutic Approaches. JACS Au 2021, 1, 2146–2161. [Google Scholar] [CrossRef]
- Yan, M.P.; Wee, C.E.; Yen, K.P.; Stevens, A.; Wai, L.K. G-Quadruplex Ligands as Therapeutic Agents against Cancer, Neurological Disorders and Viral Infections. Future Med. Chem. 2023, 15, 1987–2009. [Google Scholar] [CrossRef] [PubMed]
- Xu, Y.; Komiyama, M. G-Quadruplexes in Human Telomere: Structures, Properties, and Applications. Molecules 2023, 29, 174. [Google Scholar] [CrossRef]
- Falanga, A.P.; Piccialli, I.; Greco, F.; D’Errico, S.; Nolli, M.G.; Borbone, N.; Oliviero, G.; Roviello, G.N. Nanostructural Modulation of G-Quadruplex in Neurodegeneration: Orotate Interaction Revealed Through Experimental and Computational Approaches. J. Neurochem. 2025, 169. [Google Scholar] [CrossRef]
- Platella, C.; Ghirga, F.; Zizza, P.; Pompili, L.; Marzano, S.; Pagano, B.; Quaglio, D.; Vergine, V.; Cammarone, S.; Botta, B.; et al. Identification of Effective Anticancer G-Quadruplex-Targeting Chemotypes through the Exploration of a High Diversity Library of Natural Compounds. Pharmaceutics 2021, 13, 1611. [Google Scholar] [CrossRef]
- Chen, J.; He, Y.; Xu, Y.; Umer, M.; Anwar, N.; Wei, S.; Liu, W.; Wang, Z.; Gao, C. Higher Selective Targeting of Telomeric Multimeric G-Quadruplex by Natural Product Berberine. Curr. Med. Chem. 2026, 33, 676–688. [Google Scholar] [CrossRef]
- Das, A.; Dutta, S. Binding Studies of Aloe-Active Compounds with G-Quadruplex Sequences. ACS Omega 2021, 6, 18344–18351. [Google Scholar] [CrossRef]
- Pirota, V.; Stasi, M.; Benassi, A.; Doria, F. An Overview of Quadruplex Ligands: Their Common Features and Chemotype Diversity. In Annual Reports in Medicinal Chemistry; Academic Press Inc., 2020; Vol. 54, pp. 163–196. [Google Scholar]
- Zhang, H.; Xiang, J.; Hu, H.; Liu, Y.; Yang, F.; Shen, G.; Tang, Y.; Chen, C. Selective Recognition of Specific G-Quadruplex vs. Duplex DNA by a Phenanthroline Derivative. Int. J. Biol. Macromol. 2015, 78, 149–156. [Google Scholar] [CrossRef]
- Hashimoto, Y.; Imagawa, Y.; Nagano, K.; Maeda, R.; Nagahama, N.; Torii, T.; Kinoshita, N.; Takamiya, N.; Kawauchi, K.; Tatesishi-Karimata, H.; et al. Simple and Fast Screening for Structure-Selective G-Quadruplex Ligands. Chemical Communications 2023, 59, 4891–4894. [Google Scholar] [CrossRef]
- Asamitsu, S.; Obata, S.; Yu, Z.; Bando, T.; Sugiyama, H. Recent Progress of Targeted G-Quadruplex-Preferred Ligands Toward Cancer Therapy. Molecules 2019, 24, 429. [Google Scholar] [CrossRef]
- Choucair, K.; Imtiaz, H.; Uddin, M.H.; Nagasaka, M.; Al-Hallak, M.N.; Philip, P.A.; El-Rayes, B.; Pasche, B.C.; Azmi, A.S. Targeting KRAS Mutations: Orchestrating Cancer Evolution and Therapeutic Challenges. Signal Transduct. Target. Ther. 2025, 10, 385. [Google Scholar] [CrossRef] [PubMed]
- Uniyal, P.; Kashyap, V.K.; Behl, T.; Parashar, D.; Rawat, R. KRAS Mutations in Cancer: Understanding Signaling Pathways to Immune Regulation and the Potential of Immunotherapy. Cancers (Basel). 2025, 17, 785. [Google Scholar] [CrossRef] [PubMed]
- Hyman, D.M.; Taylor, B.S.; Baselga, J. Implementing Genome-Driven Oncology. Cell 2017, 168, 584–599. [Google Scholar] [CrossRef] [PubMed]
- Lefort, S.; Tan, S.; Balani, S.; Rafn, B.; Pellacani, D.; Hirst, M.; Sorensen, P.H.; Eaves, C.J. Initiation of Human Mammary Cell Tumorigenesis by Mutant KRAS Requires YAP Inactivation. Oncogene 2020, 39, 1957–1968. [Google Scholar] [CrossRef]
- Singh, A.; Greninger, P.; Rhodes, D.; Koopman, L.; Violette, S.; Bardeesy, N.; Settleman, J. A Gene Expression Signature Associated with “K-Ras Addiction” Reveals Regulators of EMT and Tumor Cell Survival. Cancer Cell 2009, 15, 489–500. [Google Scholar] [CrossRef]
- Cox, A.D.; Fesik, S.W.; Kimmelman, A.C.; Luo, J.; Der, C.J. Drugging the Undruggable RAS: Mission Possible? Nat. Rev. Drug Discov. 2014, 13, 828–851. [Google Scholar] [CrossRef]
- Ou, A.; Schmidberger, J.W.; Wilson, K.A.; Evans, C.W.; Hargreaves, J.A.; Grigg, M.; O’Mara, M.L.; Iyer, K.S.; Bond, C.S.; Smith, N.M. High Resolution Crystal Structure of a KRAS Promoter G-Quadruplex Reveals a Dimer with Extensive Poly-A π-Stacking Interactions for Small-Molecule Recognition. Nucleic Acids Res. 2020, 48, 5766–5776. [Google Scholar] [CrossRef]
- Kerkour, A.; Marquevielle, J.; Ivashchenko, S.; Yatsunyk, L.A.; Mergny, J.L.; Salgado, G.F. High-Resolution Three-Dimensional NMR Structure of the KRAS Proto-Oncogene Promoter Reveals Key Features of a G-Quadruplex Involved in Transcriptional Regulation. Journal of Biological Chemistry 2017, 292, 8082–8091. [Google Scholar] [CrossRef] [PubMed]
- Wang, X.D.; Nie, Q.W.; Hu, M.H. A Small-Sized Imidazole-Derived Ligand Binds to the KRAS Promoter G-Quadruplex and Inhibits Cancer Growth with Enhanced Immunomodulation. Journal of Biological Chemistry 2025, 301. [Google Scholar] [CrossRef]
- Fiorentino, A.; D’Abrosca, B.; Pacifico, S.; Iacovino, R.; Izzo, A.; Uzzo, P.; Russo, A.; Di Blasio, B.; Monaco, P. Carexanes from Carex Distachya Desf.: Revised Stereochemistry and Characterization of Four Novel Polyhydroxylated Prenylstilbenes. Tetrahedron 2008, 64, 7782–7786. [Google Scholar] [CrossRef]
- Fiorentino, A.; D’Abrosca, B.; Pacifico, S.; Iacovino, R.; Mastellone, C.; Di Blasio, B.; Monaco, P. Distachyasin: A New Antioxidant Metabolite from the Leaves of Carex Distachya. Bioorg. Med. Chem. Lett. 2006, 16, 6096–6101. [Google Scholar] [CrossRef]
- Okon, E.; Kukula-Koch, W.; Halasa, M.; Jarzab, A.; Baran, M.; Dmoszynska-Graniczka, M.; Angelis, A.; Kalpoutzakis, E.; Guz, M.; Stepulak, A.; et al. Magnoflorine—Isolation and the Anticancer Potential against NCI-H1299 Lung, MDA-MB-468 Breast, T98G Glioma, and TE671 Rhabdomyosarcoma Cancer Cells. Biomolecules 2020, 10, 1532. [Google Scholar] [CrossRef]
- Xu, T.; Kuang, T.; Du, H.; Li, Q.; Feng, T.; Zhang, Y.; Fan, G. Magnoflorine: A Review of Its Pharmacology, Pharmacokinetics and Toxicity. Pharmacol. Res. 2020, 152, 104632. [Google Scholar] [CrossRef] [PubMed]
- del Villar-Guerra, R.; Trent, J.O.; Chaires, J.B. G-Quadruplex Secondary Structure Obtained from Circular Dichroism Spectroscopy. Angewandte Chemie 2018, 130, 7289–7293. [Google Scholar] [CrossRef]
- Marzano, M.; Prencipe, F.; Delre, P.; Mangiatordi, G.F.; Travagliante, G.; Ronga, L.; Piccialli, G.; Saviano, M.; D’Errico, S.; Tesauro, D.; et al. A CD Study of a Structure-Based Selection of N-Heterocyclic Bis-Carbene Gold(I) Complexes as Potential Ligands of the G-Quadruplex-Forming Human Telomeric HTel23 Sequence. Molecules 2024, 29, 5446. [Google Scholar] [CrossRef]
- Carvalho, J.; Queiroz, J.A.; Cruz, C. Circular Dichroism of G-Quadruplex: A Laboratory Experiment for the Study of Topology and Ligand Binding. J. Chem. Educ. 2017, 94, 1547–1551. [Google Scholar] [CrossRef]
- Falanga, A.P.; D’Urso, A.; Travagliante, G.; Gangemi, C.M.A.; Marzano, M.; D’Errico, S.; Terracciano, M.; Greco, F.; De Stefano, L.; Dardano, P.; et al. Higher-Order G-Quadruplex Structures and Porphyrin Ligands: Towards a Non-Ambiguous Relationship. Int. J. Biol. Macromol. 2024, 268, 131801. [Google Scholar] [CrossRef]
- Fracchioni, G.; Vailati, S.; Grazioli, M.; Pirota, V. Structural Unfolding of G-Quadruplexes: From Small Molecules to Antisense Strategies. Molecules 2024, 29, 3488. [Google Scholar] [CrossRef]
- Santos, T.; Salgado, G.F.; Cabrita, E.J.; Cruz, C. G-Quadruplexes and Their Ligands: Biophysical Methods to Unravel G-Quadruplex/Ligand Interactions. Pharmaceuticals 2021, 14, 769. [Google Scholar] [CrossRef]
- D’Aria, F.; Pagano, B.; Petraccone, L.; Giancola, C. KRAS Promoter G-Quadruplexes from Sequences of Different Length: A Physicochemical Study. Int. J. Mol. Sci. 2021, 22, 448. [Google Scholar] [CrossRef]
- Dickerhoff, J.; Yang, D. NMR Structure Study of DNA G-Quadruplexes and Ligand Complexes. Prog. Nucl. Magn. Reson. Spectrosc. 2026, 154–155, 101597. [Google Scholar] [CrossRef]
- Marzano, M.; D’Errico, S.; Greco, F.; Falanga, A.P.; Terracciano, M.; Di Prisco, D.; Piccialli, G.; Borbone, N.; Oliviero, G. Polymorphism of G-Quadruplexes Formed by Short Oligonucleotides Containing a 3′-3′ Inversion of Polarity: From G:C:G:C Tetrads to π–π Stacked G-Wires. Int. J. Biol. Macromol. 2023, 253, 127062. [Google Scholar] [CrossRef]
- Marzano, M.; Falanga, A.P.; Dardano, P.; D’Errico, S.; Rea, I.; Terracciano, M.; De Stefano, L.; Piccialli, G.; Borbone, N.; Oliviero, G. π–π Stacked DNA G-Wire Nanostructures Formed by a Short G-Rich Oligonucleotide Containing a 3′–3′ Inversion of Polarity Site. Organic Chemistry Frontiers 2020, 7, 2187–2195. [Google Scholar] [CrossRef]
- Reed, C.R.; Kennedy, S.D.; Horowitz, R.H.; Keedakkatt Puthenpeedikakkal, A.M.; Stern, H.A.; Mathews, D.H. Modeling and NMR Data Elucidate the Structure of a G-Quadruplex-Ligand Interaction for a Pu22T-Cyclometalated Iridium(III) System. Journal of Physical Chemistry B 2024, 128, 11634–11643. [Google Scholar] [CrossRef]
- Wang, K.-B.; Liu, Y.; Li, J.; Xiao, C.; Wang, Y.; Gu, W.; Li, Y.; Xia, Y.-Z.; Yan, T.; Yang, M.-H.; et al. Structural Insight into the Bulge-Containing KRAS Oncogene Promoter G-Quadruplex Bound to Berberine and Coptisine. Nat. Commun. 2022, 13, 6016. [Google Scholar] [CrossRef] [PubMed]
- Coban, T.; Robertson, C.; Schwikkard, S.; Singer, R.; Legresley, A. Synthesis and Evaluation of Bis(Imino)Anthracene Derivatives as G-Quadruplex Ligands. RSC Med. Chem. 2021, 12, 751–757. [Google Scholar] [CrossRef] [PubMed]
- Dallavalle, S.; Musso, L.; Artali, R.; Aviñó, A.; Scaglioni, L.; Eritja, R.; Gargallo, R.; Mazzini, S. G-Quadruplex Binding Properties of a Potent PARP-1 Inhibitor Derived from 7-Azaindole-1-Carboxamide. Sci. Rep. 2021, 11. [Google Scholar] [CrossRef]
- Carvalho, J.; Pereira, E.; Marquevielle, J.; Campello, M.P.C.; Mergny, J.-L.; Paulo, A.; Salgado, G.F.; Queiroz, J.A.; Cruz, C. Fluorescent Light-up Acridine Orange Derivatives Bind and Stabilize KRAS-22RT G-Quadruplex. Biochimie 2018, 144, 144–152. [Google Scholar] [CrossRef] [PubMed]
- Lin, C.; Wu, G.; Wang, K.; Onel, B.; Sakai, S.; Shao, Y.; Yang, D. Molecular Recognition of the Hybrid-2 Human Telomeric G-Quadruplex by Epiberberine: Insights into Conversion of Telomeric G-Quadruplex Structures. Angewandte Chemie 2018, 130, 11054–11059. [Google Scholar] [CrossRef]
- Falanga, A.P.; Cremonini, M.; Bartocci, A.; Nolli, M.G.; Terracciano, M.; Volpi, S.; Dumont, E.; Piccialli, G.; Casnati, A.; Sansone, F.; et al. Calixarenes Meet (TG4T)4 G-Quadruplex: Exploring Reciprocal Interactions to Develop Innovative Biotechnological Applications. Int. J. Biol. Macromol. 2025, 305. [Google Scholar] [CrossRef] [PubMed]
- Guarra, F.; Marzo, T.; Ferraroni, M.; Papi, F.; Bazzicalupi, C.; Gratteri, P.; Pescitelli, G.; Messori, L.; Biver, T.; Gabbiani, C. Interaction of a Gold(I) Dicarbene Anticancer Drug with Human Telomeric DNA G-Quadruplex: Solution and Computationally Aided X-Ray Diffraction Analysis. Dalton Transactions 2018, 47, 16132–16138. [Google Scholar] [CrossRef]
- Canale, T.D.; Sen, D. Hemin-Utilizing G-Quadruplex DNAzymes Are Strongly Active in Organic Co-Solvents. Biochimica et Biophysica Acta (BBA) - General Subjects 2017, 1861, 1455–1462. [Google Scholar] [CrossRef]
| Entry | λmax (nm) | λmin (nm) | TM (°C) | ΔTM (°C) * |
|---|---|---|---|---|
| KRAS22-RT GQ | 264 | 243 | 52 | - |
| KRAS22-RT GQ + 1 | 264 | 243 | 52 | 0 |
| KRAS22-RT GQ + 2 | 264 | 243 | 55 | +3 |
| KRAS22-RT GQ + 3 | 264 | 243 | 68 | +16 |
| KRAS22-RT GQ + 4 | 264 | 243 | 55 | +3 |
| MCF-7 IC50 (µg/mL) ± SE | MCF-7R IC50 (µg/mL) ± SE | MDA-MB-231 IC50 (µg/mL) ± SE | HMEpiC IC50 (µg/mL) ± SE |
|
|---|---|---|---|---|
| 1 | 38.5 ± 2.2 | 17.3 ± 1.1 | 24.3 ± 3.1 | 26.2 ± 0.9 |
| 2 | 39.2 ± 6.9 | 30.7 ± 2.3 | 28.5 ± 1.2 | 20.0 ± 0.0 |
| 3 | 28.3 ± 1.8 | 16.7 ± 0.6 | 22.5 ± 3.2 | 14.0 ± 0.7 |
| 4 | 28.7 ± 1.5 | 13.3 ± 0.7 | 25.7 ± 1.8 | < 10.0 |
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2026 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).