Submitted:
16 April 2026
Posted:
20 April 2026
You are already at the latest version
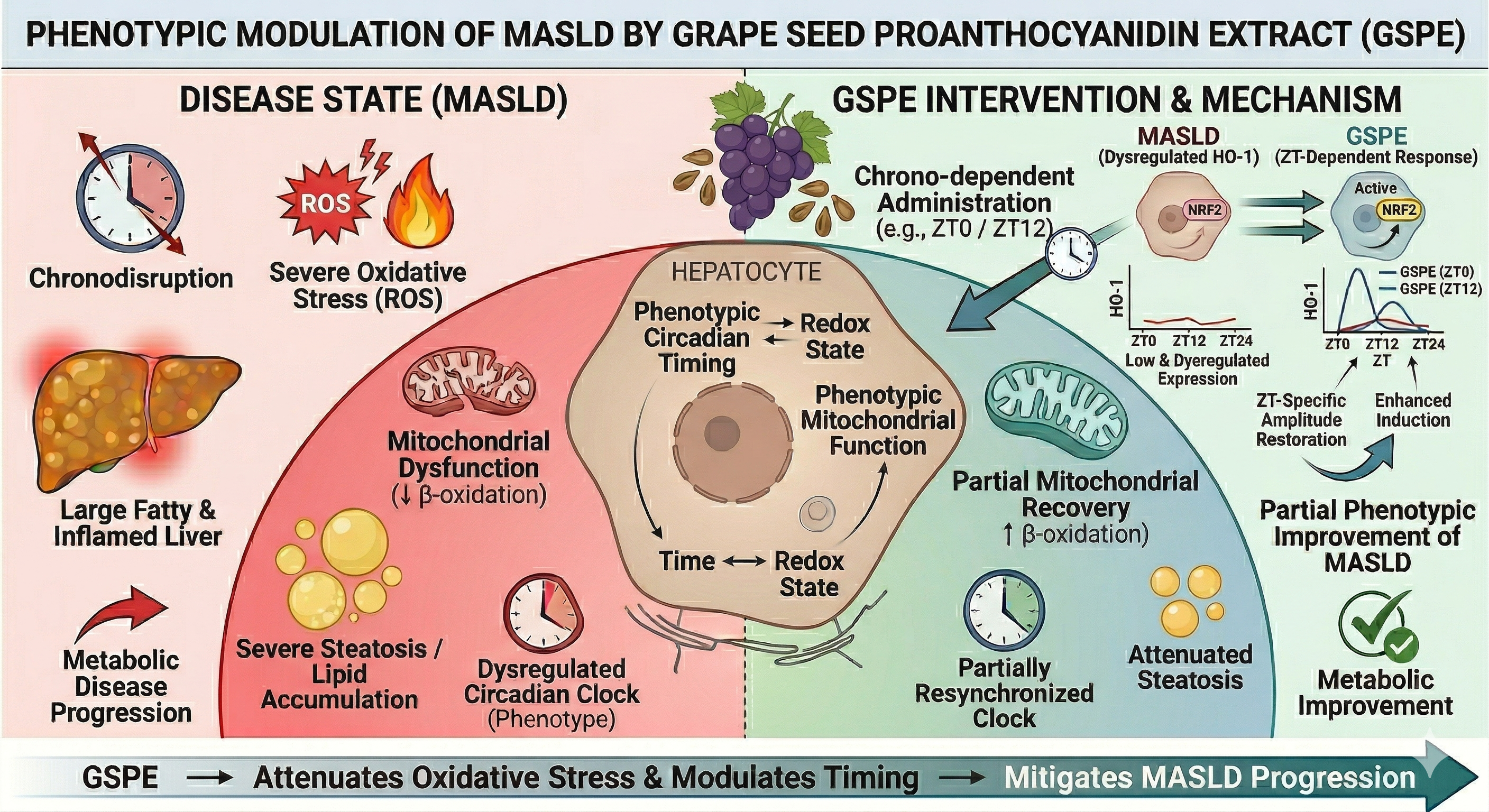
Abstract
Keywords:
1. Introduction
2. Materials and Methods
2.1. Experimental Procedure
2.2. Western Blot Analysis
2.3. Gene-Expression Analysis
2.4. Metabolomic Analysis
2.5. Liver Lipid Profiling
2.6. Statistical analysis
3. Results
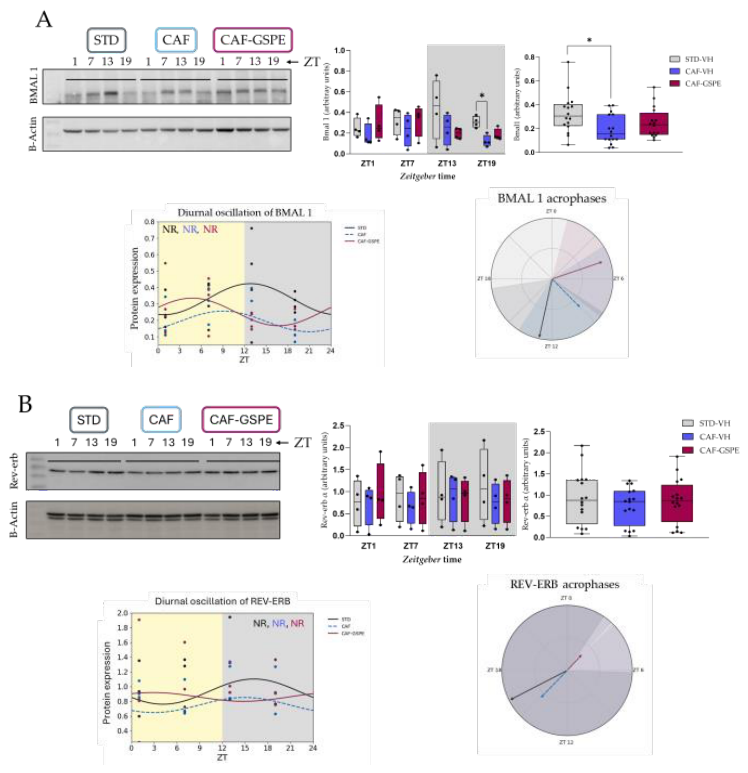
3.1. CAF Diet Effects on Hepatic Circadian Clock
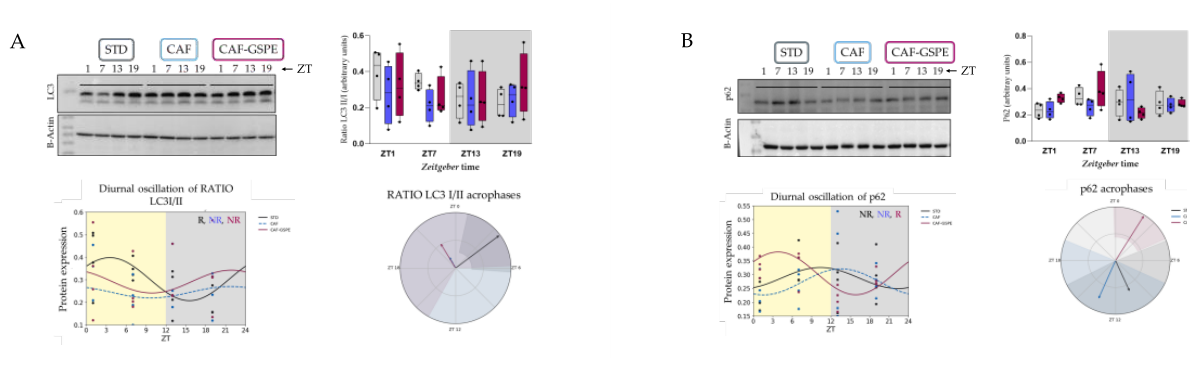
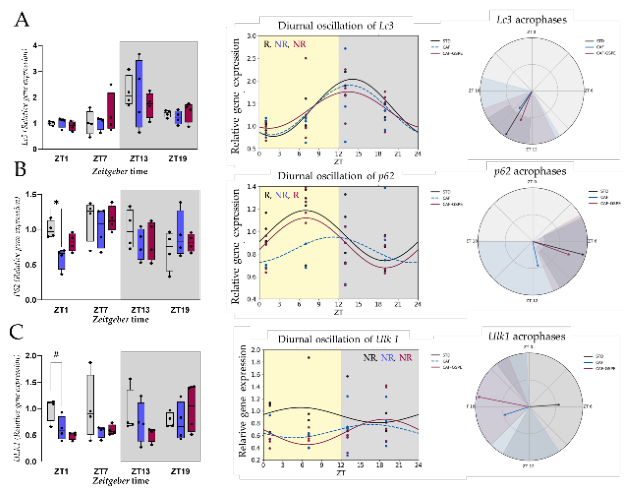
3.2. Effect of the CAF Diet and GSPE Supplementation on the Circadian Regulation of Hepatic Autophagy
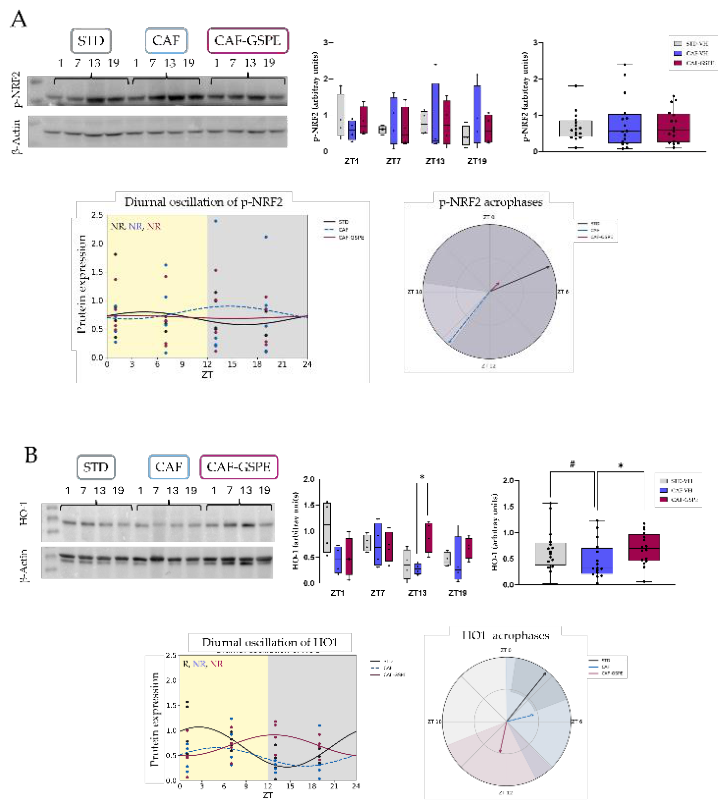
3.3. CAF Diet Effects on Hepatic Antioxidant Responses (NRF2/HO-1 Axis)
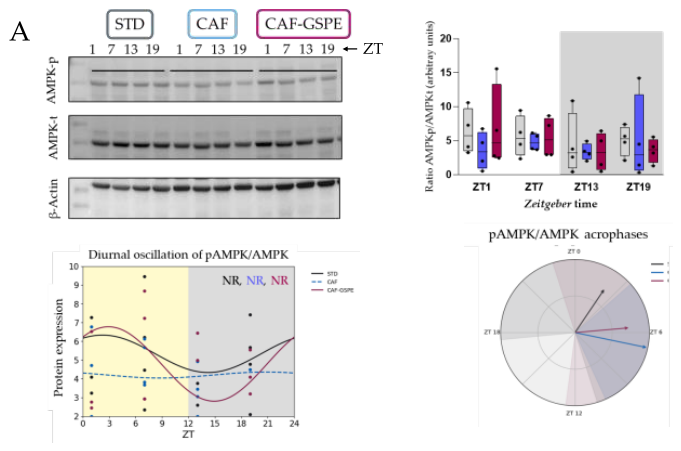
3.4. CAF Diet and GSPE Effects on AMPK Activation
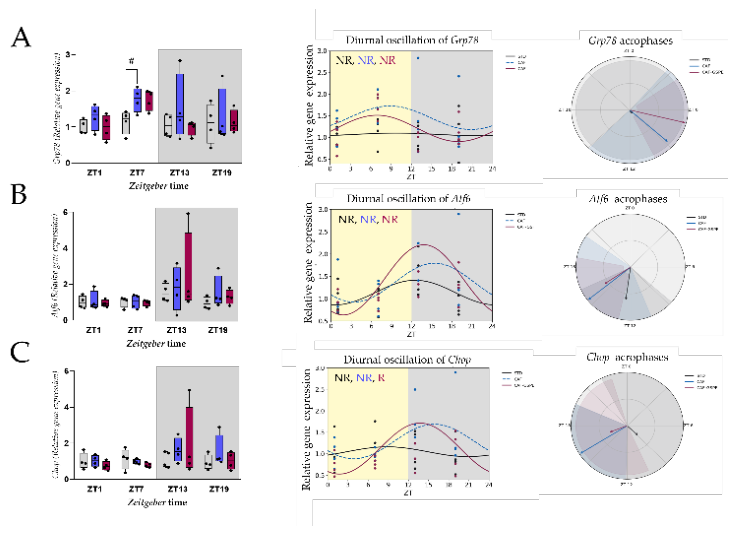
3.5. CAF Diet and GSPE Effects on ER-Stress Related Genes
3.6. Lipid Liver Profile
3.7. Liver Metabolomics Profile
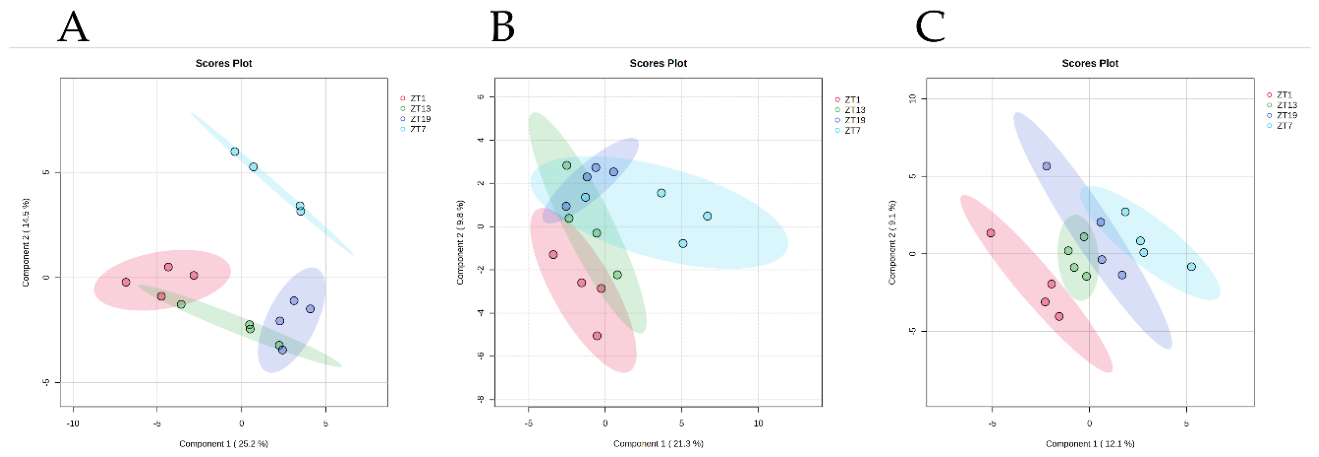
3.7.1. Global Metabolic Structure and Circadian Organization.
3.7.2. Identification of Discriminant Metabolites.
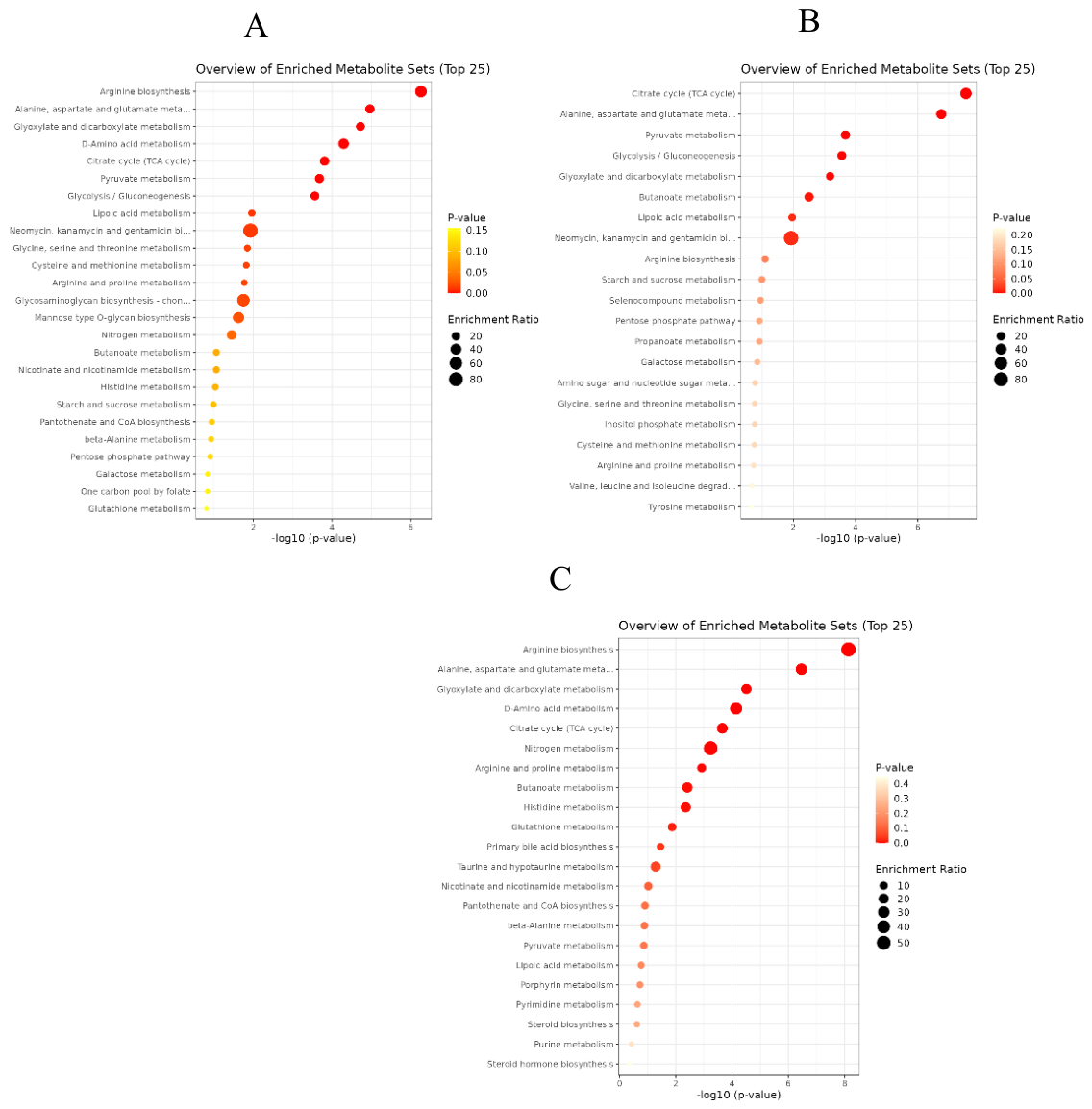
3.7.3. Metabolic Pathway Representation
4. Discussion
Supplementary Materials
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Acknowledgments
Conflicts of Interest
Abbreviations
| α-KG | Alpha-ketoglutaric acid |
| AMPK | AMP-activated protein kinase |
| AP-1 | Activator protein 1 |
| ARNT | Aryl hydrocarbon receptor nuclear translocator |
| Atf6 | Activating transcription factor 6 |
| BCA | Bicinchoninic acid |
| BMAL1 | Brain and muscle ARNT-like 1 |
| CAF | Cafeteria diet |
| cDNA | Complementary DNA |
| Chop | C/EBP homologous protein |
| CLOCK | Circadian locomotor output cycles kaput |
| COS | Centre for Omic Sciences |
| CRY | Cryptochrome |
| ECL | Enhanced chemiluminescence |
| ER | Endoplasmic reticulum |
| FDR | False discovery rate |
| FOXO | Forkhead box class O |
| GC-qTOF | Gas chromatography coupled with quadrupole time-of-flight mass spectrometry |
| Grp78 | Glucose-regulated protein 78 |
| GSPE | Grape seed proanthocyanidin extract |
| HO-1 | Heme Oxygenase-1 |
| KEAP1 | Kelch-like ECH-associated protein 1 |
| LC3 | Microtubule-associated protein 1A/1B-light chain 3 |
| LC3-I | Cytosolic form of LC3 |
| LC3-II | Lipidated form of LC3 |
| MASH | Metabolic dysfunction-associated steatohepatitis |
| MASLD | Metabolic dysfunction-associated steatotic liver disease |
| MESOR | Midline estimating statistic of rhythm |
| mTORC1 | Mechanistic target of rapamycin complex 1 |
| NAFLD | Non-alcoholic fatty liver disease |
| NR1D1 | Nuclear receptor subfamily 1 group D member 1 |
| NRF2 | Nuclear factor erythroid 2–related factor 2 |
| p-NRF2 | Phosphorylated Nuclear Factor Erythroid 2-Related Factor 2 |
| PCA | Principal component analysis |
| PER | Period protein |
| PERMANOVA | Permutational multivariate analysis of variance |
| PIC | Protease inhibitor cocktail |
| PLS-DA | Partial least squares discriminant analysis |
| PMSF | Phenylmethanesulfonyl fluoride |
| Ppia | Peptidylprolyl isomerase A |
| p-62 | Ubiquitin-binding protein p62 |
| PVDF | Polyvinylidene difluoride |
| qPCR | Quantitative polymerase chain reaction |
| REV-ERBα | Nuclear receptor subfamily 1 group D member 1 |
| RIPA | Radioimmunoprecipitation assay buffer |
| ROR | Retinoic acid-related orphan receptor |
| ROS | Reactive oxygen species. |
| RT-qPCR | Reverse transcription quantitative polymerase chain reaction |
| SEM | Standard error of the mean |
| SDS | Sodium dodecyl sulfate |
| SQSTM1 | Sequestosome 1 |
| STD | Standard diet |
| TAG | Triacylglycerol |
| TBST | Tris-buffered saline with Tween 20 |
| TCA cycle | Tricarboxylic acid cycle |
| UPR | Unfolded protein response |
| Ulk1 | Unc-51 like autophagy activating kinase 1 |
| VH | Vehicle |
| VIP | Variable importance in projection |
| ZT | Zeitgeber time |
References
- Eslam, M.; Newsome, P.N.; Sarin, S.K.; Anstee, Q.M.; Targher, G.; Romero-Gomez, M.; Zelber-Sagi, S.; Wai-Sun Wong, V.; Dufour, J.F.; Schattenberg, J.M.; et al. A New Definition for Metabolic Dysfunction-Associated Fatty Liver Disease: An International Expert Consensus Statement. J. Hepatol. 2020, 73, 202–209. [Google Scholar] [CrossRef] [PubMed]
- Rinella, M.E.; Lazarus, J. V.; Ratziu, V.; Francque, S.M.; Sanyal, A.J.; Kanwal, F.; Romero, D.; Abdelmalek, M.F.; Anstee, Q.M.; Arab, J.P.; et al. A Multisociety Delphi Consensus Statement on New Fatty Liver Disease Nomenclature. Hepatology 2023, 78, 1966–1986. [Google Scholar] [CrossRef]
- Reinke, H.; Asher, G. Crosstalk between Metabolism and Circadian Clocks. Nat. Rev. Mol. Cell Biol. 2019, 20, 227–241. [Google Scholar] [CrossRef]
- Rudic, R.D.; McNamara, P.; Curtis, A.M.; Boston, R.C.; Panda, S.; Hogenesch, J.B.; FitzGerald, G.A. BMAL1 and CLOCK, Two Essential Components of the Circadian Clock, Are Involved in Glucose Homeostasis. PLoS Biol. 2004, 2, e377. [Google Scholar] [CrossRef]
- Shi, D.; Chen, J.; Wang, J.; Yao, J.; Huang, Y.; Zhang, G.; Bao, Z. Circadian Clock Genes in the Metabolism of Non-Alcoholic Fatty Liver Disease. Front. Physiol. 2019, 10, 423. [Google Scholar] [CrossRef]
- Bai, P.; Cantó, C.; Oudart, H.; Brunyánszki, A.; Cen, Y.; Thomas, C.; Yamamoto, H.; Huber, A.; Kiss, B.; Houtkooper, R.H.; et al. PARP-1 Inhibition Increases Mitochondrial Metabolism through SIRT1 Activation. Cell Metab. 2011, 13, 461–468. [Google Scholar] [CrossRef]
- Singh, R.; Kaushik, S.; Wang, Y.; Xiang, Y.; Novak, I.; Komatsu, M.; Tanaka, K.; Cuervo, A.M.; Czaja, M.J. Autophagy Regulates Lipid Metabolism. Nature 2009, 458, 1131–1135. [Google Scholar] [CrossRef]
- Ueno, T.; Komatsu, M. Autophagy in the Liver: Functions in Health and Disease. Nature Reviews Gastroenterology & Hepatology 2017, 14, 170–184. [Google Scholar] [CrossRef] [PubMed]
- Henkel, A.S. Unfolded Protein Response Sensors in Hepatic Lipid Metabolism and Nonalcoholic Fatty Liver Disease. Semin. Liver Dis. 2018, 38, 320–332. [Google Scholar] [CrossRef] [PubMed]
- Ipsen, D.H.; Lykkesfeldt, J.; Tveden-Nyborg, P. Molecular Mechanisms of Hepatic Lipid Accumulation in Non-Alcoholic Fatty Liver Disease. Cellular and Molecular Life Sciences 2018, 75 75, 3313–3327. [Google Scholar] [CrossRef]
- Cuadrado, A.; Rojo, A.I.; Wells, G.; Hayes, J.D.; Cousin, S.P.; Rumsey, W.L.; Attucks, O.C.; Franklin, S.; Levonen, A.L.; Kensler, T.W.; et al. Therapeutic Targeting of the NRF2 and KEAP1 Partnership in Chronic Diseases. Nature Reviews Drug Discovery 2018, 18:4 18, 295–317. [Google Scholar] [CrossRef] [PubMed]
- Jain, A.; Lamark, T.; Sjøttem, E.; Larsen, K.B.; Awuh, J.A.; Øvervatn, A.; McMahon, M.; Hayes, J.D.; Johansen, T. P62/SQSTM1 Is a Target Gene for Transcription Factor NRF2 and Creates a Positive Feedback Loop by Inducing Antioxidant Response Element-Driven Gene Transcription. Journal of Biological Chemistry 2010, 285, 22576–22591. [Google Scholar] [CrossRef] [PubMed]
- Komatsu, M.; Kurokawa, H.; Waguri, S.; Taguchi, K.; Kobayashi, A.; Ichimura, Y.; Sou, Y.S.; Ueno, I.; Sakamoto, A.; Tong, K.I.; et al. The Selective Autophagy Substrate P62 Activates the Stress Responsive Transcription Factor Nrf2 through Inactivation of Keap1. Nature Cell Biology 2010, 12:3 2010(12), 213–223. [Google Scholar] [CrossRef]
- Wu, L.; Xu, W.; Li, H.; Dong, B.; Geng, H.; Jin, J.; Han, D.; Liu, H.; Zhu, X.; Yang, Y.; et al. Vitamin C Attenuates Oxidative Stress, Inflammation, and Apoptosis Induced by Acute Hypoxia through the Nrf2/Keap1 Signaling Pathway in Gibel Carp (Carassius Gibelio). Antioxidants 2022, Vol. 11 11, 935. [Google Scholar] [CrossRef]
- Pekovic-Vaughan, V.; Gibbs, J.; Yoshitane, H.; Yang, N.; Pathiranage, D.; Guo, B.; Sagami, A.; Taguchi, K.; Bechtold, D.; Loudon, A.; et al. The Circadian Clock Regulates Rhythmic Activation of the NRF2/Glutathione-Mediated Antioxidant Defense Pathway to Modulate Pulmonary Fibrosis. Genes Dev. 2014, 28, 548. [Google Scholar] [CrossRef] [PubMed]
- Miani, M.; Le Naour, J.; Waeckel-Enée, E.; Verma, S. chand; Straube, M.; Emond, P.; Ryffel, B.; van Endert, P.; Sokol, H.; Diana, J. Gut Microbiota-Stimulated Innate Lymphoid Cells Support β-Defensin 14 Expression in Pancreatic Endocrine Cells, Preventing Autoimmune Diabetes. Cell Metab. 2018, 28, 557–572.e6. [Google Scholar] [CrossRef]
- Xu, Y.Q.; Zhang, D.; Jin, T.; Cai, D.J.; Wu, Q.; Lu, Y.; Liu, J.; Klaassen, C.D. Diurnal Variation of Hepatic Antioxidant Gene Expression in Mice. PLoS One 2012, 7, e44237. [Google Scholar] [CrossRef]
- Baselga-Escudero, L.; Bladé, C.; Ribas-Latre, A.; Casanova, E.; Salvadó, M.J.; Arola, L.; Arola-Arnal, A. Grape Seed Proanthocyanidins Repress the Hepatic Lipid Regulators MiR-33 and MiR-122 in Rats. In Mol. Nutr. Food Res.; WGROUP:STRING:PUBLICATION, 2012; Volume 56, pp. 1636–1646. [Google Scholar] [CrossRef]
- Hosseini, H.; Teimouri, M.; Shabani, M.; Koushki, M.; Babaei Khorzoughi, R.; Namvarjah, F.; Izadi, P.; Meshkani, R. Resveratrol Alleviates Non-Alcoholic Fatty Liver Disease through Epigenetic Modification of the Nrf2 Signaling Pathway. Int. J. Biochem. Cell Biol. 2020, 119, 105667. [Google Scholar] [CrossRef]
- Fan, C.; Ling-Hu, A.; Sun, D.; Gao, W.; Zhang, C.; Duan, X.; Li, H.; Tian, W.; Yu, Q.; Ke, Z. Nobiletin Ameliorates Hepatic Lipid Deposition, Oxidative Stress, and Inflammation by Mechanisms That Involve the Nrf2/NF-ΚB Axis in Nonalcoholic Fatty Liver Disease. J. Agric. Food Chem. 2023, 71, 20105–20117. [Google Scholar] [CrossRef]
- Terra, X.; Montagut, G.; Bustos, M.; Llopiz, N.; Ardèvol, A.; Bladé, C.; Fernández-Larrea, J.; Pujadas, G.; Salvadó, J.; Arola, L.; et al. Grape-Seed Procyanidins Prevent Low-Grade Inflammation by Modulating Cytokine Expression in Rats Fed a High-Fat Diet. J. Nutr. Biochem. 2009, 20, 210–218. [Google Scholar] [CrossRef]
- Manocchio, F.; Soliz-Rueda, J.R.; Ribas-Latre, A.; Bravo, F.I.; Arola-Arnal, A.; Suarez, M.; Muguerza, B. Grape Seed Proanthocyanidins Modulate the Hepatic Molecular Clock via MicroRNAs. Mol. Nutr. Food Res. 2022, 66, 2200443. [Google Scholar] [CrossRef]
- Cortés-Espinar, A.J.; Ibarz-Blanch, N.; Soliz-Rueda, J.R.; Calvo, E.; Bravo, F.I.; Mulero, M.; Ávila-Román, J. Abrupt Photoperiod Changes Differentially Modulate Hepatic Antioxidant Response in Healthy and Obese Rats: Effects of Grape Seed Proanthocyanidin Extract (GSPE). Int. J. Mol. Sci. 2023, 24, 17057. [Google Scholar] [CrossRef]
- Rodríguez, R.M.; de Assis, L.V.M.; Calvo, E.; Colom-Pellicer, M.; Quesada-Vázquez, S.; Cruz-Carrión, Á; Escoté, X.; Oster, H.; Aragonès, G.; Mulero, M. Grape-Seed Proanthocyanidin Extract (GSPE) Modulates Diurnal Rhythms of Hepatic Metabolic Genes and Metabolites, and Reduces Lipid Deposition in Cafeteria-Fed Rats in a Time-of-Day-Dependent Manner. Mol. Nutr. Food Res. 2024, 68, 2400554. [Google Scholar] [CrossRef] [PubMed]
- Cortés-Espinar, A.J.; Ibarz-Blanch, N.; Soliz-Rueda, J.R.; Bonafos, B.; Feillet-Coudray, C.; Casas, F.; Bravo, F.I.; Calvo, E.; Ávila-Román, J.; Mulero, M. Rhythm and ROS: Hepatic Chronotherapeutic Features of Grape Seed Proanthocyanidin Extract Treatment in Cafeteria Diet-Fed Rats. Antioxidants 2023, 12, 1606. [Google Scholar] [CrossRef]
- Hernandez-Baixauli, J.; Chomiciute, G.; Tracey, H.; Mora, I.; Cortés-Espinar, A.J.; Ávila-Román, J.; Abasolo, N.; Palacios-Jordan, H.; Foguet-Romero, E.; Suñol, D.; et al. Exploring Metabolic and Gut Microbiome Responses to Paraquat Administration in Male Wistar Rats: Implications for Oxidative Stress. Antioxidants 2024, 13, 67. [Google Scholar] [CrossRef] [PubMed]
- Rodríguez, R.M.; Cortés-Espinar, A.J.; Soliz-Rueda, J.R.; Feillet-Coudray, C.; Casas, F.; Colom-Pellicer, M.; Aragonès, G.; Avila-Román, J.; Muguerza, B.; Mulero, M.; et al. Time-of-Day Circadian Modulation of Grape-Seed Procyanidin Extract (GSPE) in Hepatic Mitochondrial Dynamics in Cafeteria-Diet-Induced Obese Rats. Nutrients 2022, 14, 774. [Google Scholar] [CrossRef]
- Livak, K.J.; Schmittgen, T.D. Analysis of Relative Gene Expression Data Using Real-Time Quantitative PCR and the 2−ΔΔCT Method. Methods 2001, 25, 402–408. [Google Scholar] [CrossRef]
- Cajka, T.; Fiehn, O. Toward Merging Untargeted and Targeted Methods in Mass Spectrometry-Based Metabolomics and Lipidomics. Anal. Chem. 2015, 88, 524–545. [Google Scholar] [CrossRef] [PubMed]
- Moškon, M. CosinorPy: A Python Package for Cosinor-Based Rhythmometry. BMC Bioinformatics 2020, 21, 1–12. [Google Scholar] [CrossRef]
- Montagner, A.; Korecka, A.; Polizzi, A.; Lippi, Y.; Blum, Y.; Canlet, C.; Tremblay-Franco, M.; Gautier-Stein, A.; Burcelin, R.; Yen, Y.C.; et al. Hepatic Circadian Clock Oscillators and Nuclear Receptors Integrate Microbiome-Derived Signals. Scientific Reports 2016, 6:1 2016(6), 20127. [Google Scholar] [CrossRef]
- Jacobi, D.; Liu, S.; Burkewitz, K.; Kory, N.; Knudsen, N.H.; Alexander, R.K.; Unluturk, U.; Li, X.; Kong, X.; Hyde, A.L.; et al. Hepatic Bmal1 Regulates Rhythmic Mitochondrial Dynamics and Promotes Metabolic Fitness. Cell Metab. 2015, 22, 709–720. [Google Scholar] [CrossRef]
- Ma, D.; Panda, S.; Lin, J.D. Temporal Orchestration of Circadian Autophagy Rhythm by C/EBPβ. EMBO J. 2011, 30, 4642. [Google Scholar] [CrossRef] [PubMed]
- Ma, X.; Ye, Y.; Sun, J.; Ji, J.; Wang, J.S.; Sun, X. Coexposure of Cyclopiazonic Acid with Aflatoxin B1 Involved in Disrupting Amino Acid Metabolism and Redox Homeostasis Causing Synergistic Toxic Effects in Hepatocyte Spheroids. J. Agric. Food Chem. 2022, 70, 5166–5176. [Google Scholar] [CrossRef] [PubMed]
- Sunny, N.E.; Bril, F.; Cusi, K. Mitochondrial Adaptation in Nonalcoholic Fatty Liver Disease: Novel Mechanisms and Treatment Strategies. Trends in Endocrinology and Metabolism 2017, 28, 250–260. [Google Scholar] [CrossRef] [PubMed]
- Dodson, M.; De La Vega, M.R.; Cholanians, A.B.; Schmidlin, C.J.; Chapman, E.; Zhang, D.D. Modulating NRF2 in Disease: Timing Is Everything. Annu. Rev. Pharmacol. Toxicol. 2018, 59, 555. [Google Scholar] [CrossRef]
- Kasai, S.; Shimizu, S.; Tatara, Y.; Mimura, J.; Itoh, K. Regulation of Nrf2 by Mitochondrial Reactive Oxygen Species in Physiology and Pathology. Biomolecules 2020, 10, 320. [Google Scholar] [CrossRef]
- Lee, Y.J.; Kim, S.W.; Jung, M.H.; Kim, Y.S.; Kim, K.S.; Suh, D.S.; Kim, K.H.; Choi, E.H.; Kim, J.; Kwon, B.S. Plasma-Activated Medium Inhibits Cancer Stem Cell-like Properties and Exhibits a Synergistic Effect in Combination with Cisplatin in Ovarian Cancer. Free Radic. Biol. Med. 2022, 182, 276–288. [Google Scholar] [CrossRef]
- Fujimaki, J.; Sayama, N.; Shiotani, S.; Suzuki, T.; Nonaka, M.; Uezono, Y.; Oyabu, M.; Kamei, Y.; Nukaya, H.; Wakabayashi, K.; et al. The Steroidal Alkaloid Tomatidine and Tomatidine-Rich Tomato Leaf Extract Suppress the Human Gastric Cancer-Derived 85As2 Cells In Vitro and In Vivo via Modulation of Interferon-Stimulated Genes. Nutrients 2022, Vol. 14 14, 1023. [Google Scholar] [CrossRef]
- Arola-Arnal, A.; Cruz-Carrión, Á; Torres-Fuentes, C.; Ávila-Román, J.; Aragonès, G.; Mulero, M.; Bravo, F.I.; Muguerza, B.; Arola, L.; Suárez, M. Chrononutrition and Polyphenols: Roles and Diseases. Nutrients 2019, Vol. 11 11 Page 2602 2019, 2602. [Google Scholar] [CrossRef]
- Franzago, M.; Alessandrelli, E.; Notarangelo, S.; Stuppia, L.; Vitacolonna, E. Chrono-Nutrition: Circadian Rhythm and Personalized Nutrition. International Journal of Molecular Sciences 2023, Vol. 24 24, 2571. [Google Scholar] [CrossRef]
- Koronowski, K.B.; Sassone-Corsi, P. Communicating Clocks Shape Circadian Homeostasis. Science 2021, 371, eabd0951. [Google Scholar] [CrossRef] [PubMed]
- Longo, V.D.; Panda, S. Fasting, Circadian Rhythms, and Time Restricted Feeding in Healthy Lifespan. Cell Metab. 2016, 23, 1048. [Google Scholar] [CrossRef] [PubMed]
| Gene | Forward primer (5' to 3') | Reverse Primer (5' to 3') |
| Grp78 | TCGACTTGGGGACCACCTAT | GCCCTGATCGTTGGCTATGA |
| Atf6 | GGACCAGGTGGTGTCAGAG | GACAGCTCTGCGCTTTGGG |
| Chop | ACCACCACACCTGAAAGCAG | AGCTGGACACTGTCTCAAAG |
| Lc3 | GGTCCAGTTGTGCCTTTATTGA | GTGTGTGGGTTGTGTACGTCG |
| Sqstm1 | CTAGGCATCGAGGTTGACATT | CTTGGCTGAGTACCACTCTTATC |
| Ulk1 | GGCTTACAGACTGCCATTGA | GATACCACGCTGGCCTTATAC |
| Ppia | TCAAACACAAATGGTTCCCAGT | ATTCCTGGACCCAAAACGCT |
| Parameter | Group | ||
| STD-VH | CAF-VH | CAF-GSPE | |
| TAG (mg/g) | 3.755 (3.373-3.215) | 11,686 (11,335-12,118)* | 9,914 (9,633-9,985)& |
| Cholesterol (mg/g) | 1,207 (1,130-1,251) | 1,395 (1,314-1,762)* | 1,390 (1,247-1,481) |
| Total lipids (mg/g) | 63,719 (54,201-72,582) | 89,127(77,189-105,920)* | 81,662 (75,031-88,872) |
| Group | Metabolite | VIP score |
| STD-VH | Glutamine Serine Lactic acid Malic acid Glucose-6-phosphate α-Ketoglutaric acid Pyruvic acid Ornithine Aspartic acid Xylonic acid |
1.98 1.90 1.82 1.50 1.47 1.38 1.32 1.30 1.23 1.03 |
| CAF-VH | Pyruvic acid Lactic acid Alanine Citric acid 3-Hydroxyisobutyric acid α-Ketoglutaric acid Succinic acid Glucose-6-phosphate Malic acid |
2.43 2.01 1.53 1.45 1.40 1.37 1.13 1.09 1.01 |
| CAF-GSPE | Cholesterol Ornithine |
2.11 1.92 |
| 4-Hydroxyproline | 1.63 | |
| Taurine | 1.52 | |
| Malic acid | 1.41 | |
| Aspartic acid | 1.32 | |
| Glycerol-1-phosphate | 1.31 | |
| α-Ketoglutaric acid | 1.17 | |
| Citric acid | 1.13 | |
| Glutamine | 1.13 | |
| Glutamic acid | 1.11 |
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2026 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).




