Submitted:
16 April 2026
Posted:
17 April 2026
You are already at the latest version
Abstract
Keywords:
Introduction
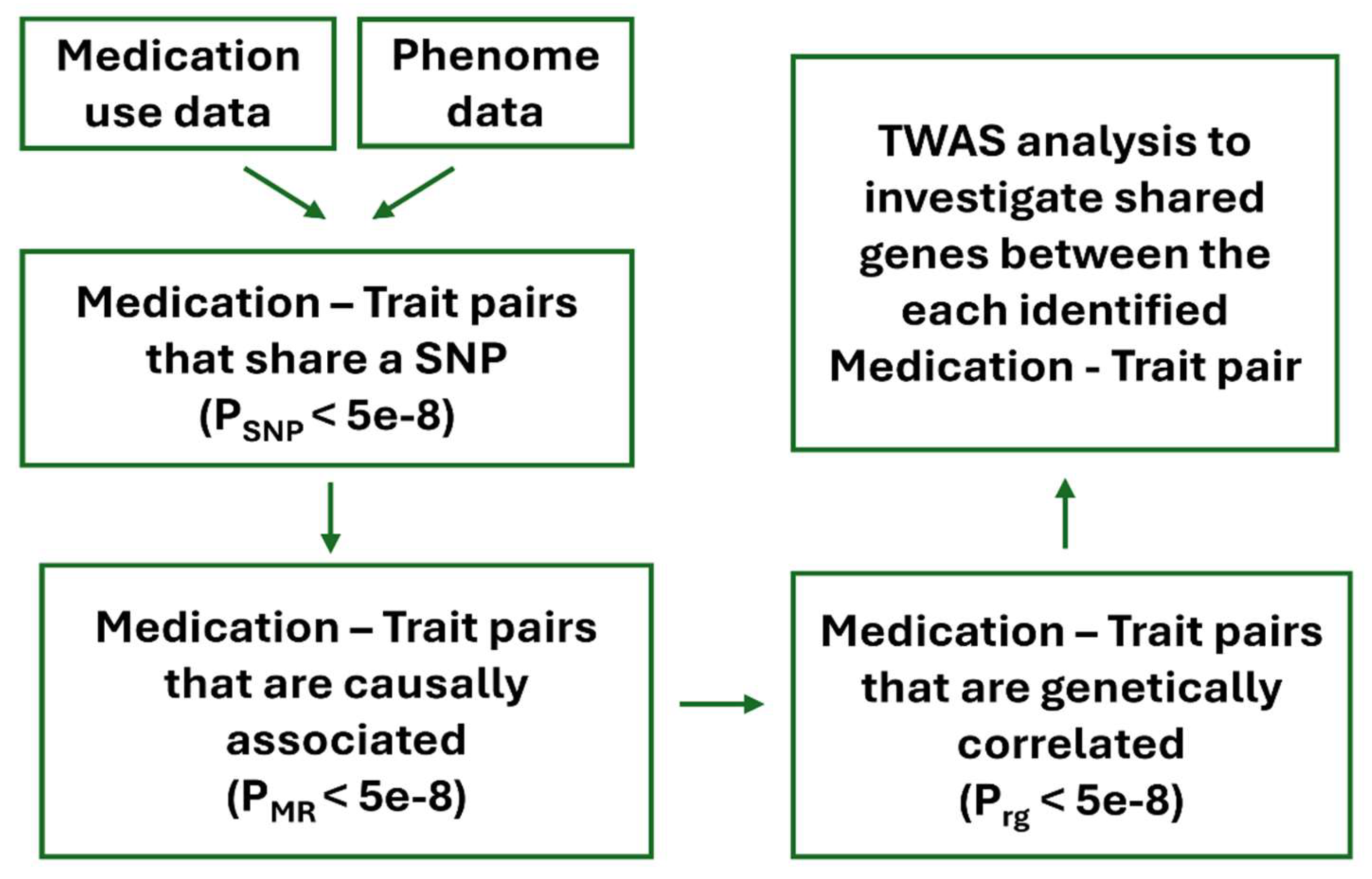
Methods
- a)
- SNPs must be associated with exposure at GWAS significance level (P < 5e-8).
- b)
- The degree of linkage disequilibrium between SNPs must not exceed r2>0.05.
- c)
- SNPs must not show pleiotropic effect (i.e., Exposure ← SNP → Outcome). For this purpose, pleiotropic SNPs were excluded using HEIDI test (P<0.01) implemented in GSRM algorithm.
Results
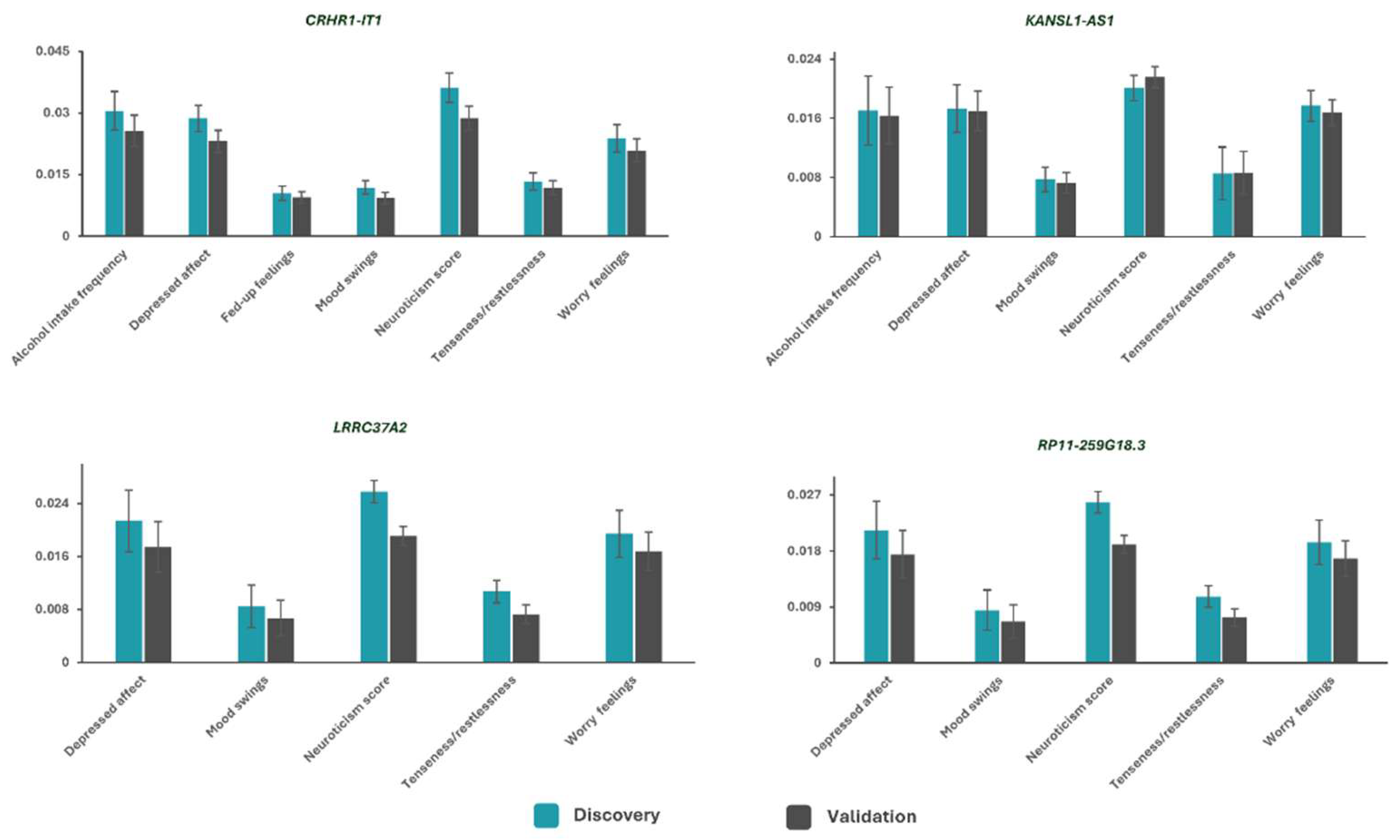
Paracetamol and Affective Traits
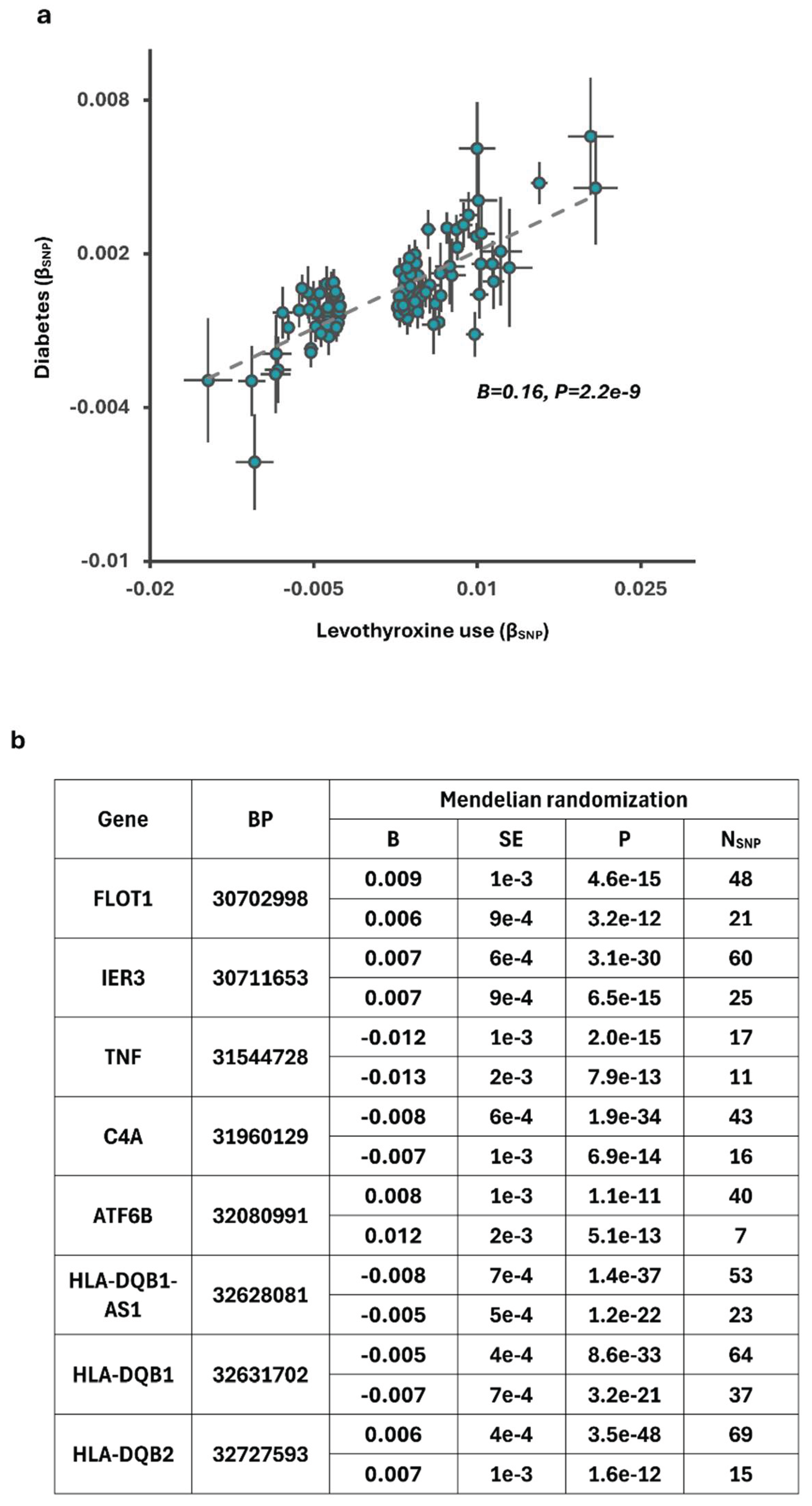
Levothyroxine and Cardiometabolic Traits
Discussion
Supplementary Materials
References
- Nelson, M. R.; et al. The support of human genetic evidence for approved drug indications. Nat. Genet. 2015, 47, 856–860. [Google Scholar] [CrossRef] [PubMed]
- King, E. A.; Davis, J. W.; Degner, J. F. Are drug targets with genetic support twice as likely to be approved? Revised estimates of the impact of genetic support for drug mechanisms on the probability of drug approval. PLoS Genet. 2019, 15, e1008489. [Google Scholar] [CrossRef] [PubMed]
- Wu, Y.; et al. Genome-wide association study of medication-use and associated disease in the UK Biobank. Nat. Commun. 2019, 10, 1891. [Google Scholar] [CrossRef] [PubMed]
- Zhu, Z.; et al. Causal associations between risk factors and common diseases inferred from GWAS summary data. Nat. Commun. 2018, 9, 224. [Google Scholar] [CrossRef] [PubMed]
- Ning, Z.; Pawitan, Y.; Shen, X. High-definition likelihood inference of genetic correlations across human complex traits. Nat. Genet. 2020, 52, 859–864. [Google Scholar] [CrossRef] [PubMed]
- Võsa, U.; et al. Large-scale cis- and trans-eQTL analyses identify thousands of genetic loci and polygenic scores that regulate blood gene expression. Nat. Genet. 2021, 53, 1300–1310. [Google Scholar] [CrossRef] [PubMed]
- Xu, Y.; et al. An atlas of genetic scores to predict multi-omic traits. Nature 2023, 616, 123–131. [Google Scholar] [CrossRef] [PubMed]
- Koolen-de Vries syndrome: MedlinePlus Genetics. Available online: https://medlineplus.gov/genetics/condition/koolen-de-vries-syndrome/.
- Noble, J. A.; Erlich, H. A. Genetics of type 1 diabetes. Cold Spring Harb. Perspect. Med. 2012, 2, a007732. [Google Scholar] [CrossRef] [PubMed]
- Borst, S. E. The role of TNF-α in insulin resistance. Endocrine 2004, 23, 177–182. [Google Scholar] [CrossRef] [PubMed]
- Aydemir, O.; et al. Polymorphisms in intron 1 of HLA-DRA differentially associate with type 1 diabetes and celiac disease and implicate involvement of complement system genes C4A and C4B. eLife 2026, 12, RP89068. [Google Scholar] [CrossRef] [PubMed]
- DeWall, C. N.; et al. Acetaminophen reduces social pain: Behavioral and neural evidence. Psychol. Sci. 2010, 21, 931–937. [Google Scholar] [CrossRef] [PubMed]
- Durso, G. R. O.; Luttrell, A.; Way, B. M. Over-the-Counter Relief From Pains and Pleasures Alike: Acetaminophen Blunts Evaluation Sensitivity to Both Negative and Positive Stimuli. Psychol. Sci. 2015, 26, 750–758. [Google Scholar] [CrossRef] [PubMed]
- Mischkowski, D.; Crocker, J.; Way, B. M. A Social Analgesic? Acetaminophen (Paracetamol) Reduces Positive Empathy. Front. Psychol. 2019, 10, 538. [Google Scholar] [CrossRef] [PubMed]
- Mallet, C.; Desmeules, J.; Pegahi, R.; Eschalier, A. An Updated Review on the Metabolite (AM404)-Mediated Central Mechanism of Action of Paracetamol (Acetaminophen): Experimental Evidence and Potential Clinical Impact. J. Pain Res. 2023, 16, 1081–1094. [Google Scholar] [CrossRef] [PubMed]
- Pearce, E. N. Thyroid hormone and obesity. Curr. Opin. Endocrinol. Diabetes Obes. 2012, 19, 408. [Google Scholar] [CrossRef] [PubMed]
- Ostadrahimi, A.; et al. Effects of levothyroxine replacement therapy on insulin resistance in patients with untreated primary hypothyroidism. BMC Res. Notes 2023, 16, 237. [Google Scholar] [CrossRef] [PubMed]
| Trait | Source | Mendelian randomization | Genetic correlation | |||||
| Beta | SE | P | NSNPs | rg | SE | P | ||
| Depressed affect | PMID: 29942085 | 1.0 | 0.1 | 1.2E-13 | 12 | 0.5 | 0.03 | 1.5E-82 |
| Worry feelings | PMID: 29942085 | 1.0 | 0.1 | 4.0E-20 | 15 | 0.3 | 0.02 | 1.2E-31 |
| Mood swings | UKBB | 0.7 | 0.1 | 6.9E-29 | 13 | 0.5 | 0.02 | 6.1E-100 |
| Fed-up feelings | UKBB | 0.6 | 0.1 | 1.8E-24 | 15 | 0.5 | 0.03 | 4.9E-87 |
| Neuroticism score | UKBB | 1.2 | 0.2 | 1.3E-14 | 11 | 0.4 | 0.02 | 1.4E-69 |
| Frequency of tenseness / restlessness in last 2 weeks | UKBB | 0.7 | 0.1 | 1.1E-20 | 14 | 0.4 | 0.03 | 1.1E-55 |
| Alcohol intake frequency | UKBB | 1.7 | 0.2 | 1.0E-22 | 14 | 0.3 | 0.02 | 1.8E-62 |
| Gene information | Mendelian randomization | |||||
| Chr | bp | Symbol | B | SE | P | NSNP |
| 6 | 30885127 | VARS2 | 0.011a | 0.002 | 2.5E-10 | 18 |
| 0.016b | 0.002 | 1.1E-11 | 21 | |||
| 6 | 31635566 | CSNK2B | -0.026 | 0.005 | 4.6E-08 | 5 |
| -0.038 | 0.007 | 2.8E-08 | 8 | |||
| 17 | 43711638 | CRHR1-IT1 | 0.007 | 0.001 | 2.9E-09 | 35 |
| 0.008 | 0.001 | 4.6E-09 | 31 | |||
| 17 | 44272515 | KANSL1-AS1 | 0.005 | 0.001 | 6.5E-10 | 27 |
| 0.004 | 0.001 | 1.8E-08 | 36 | |||
| 17 | 44337444 | RP11-259G18.3 | 0.009 | 0.002 | 4.2E-08 | 13 |
| 0.005 | 0.001 | 3.4E-09 | 35 | |||
| 17 | 44610946 | LRRC37A2 | 0.005 | 0.001 | 9.1E-10 | 26 |
| 0.005 | 0.001 | 4.1E-08 | 32 | |||
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2026 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).




