Submitted:
15 April 2026
Posted:
16 April 2026
You are already at the latest version
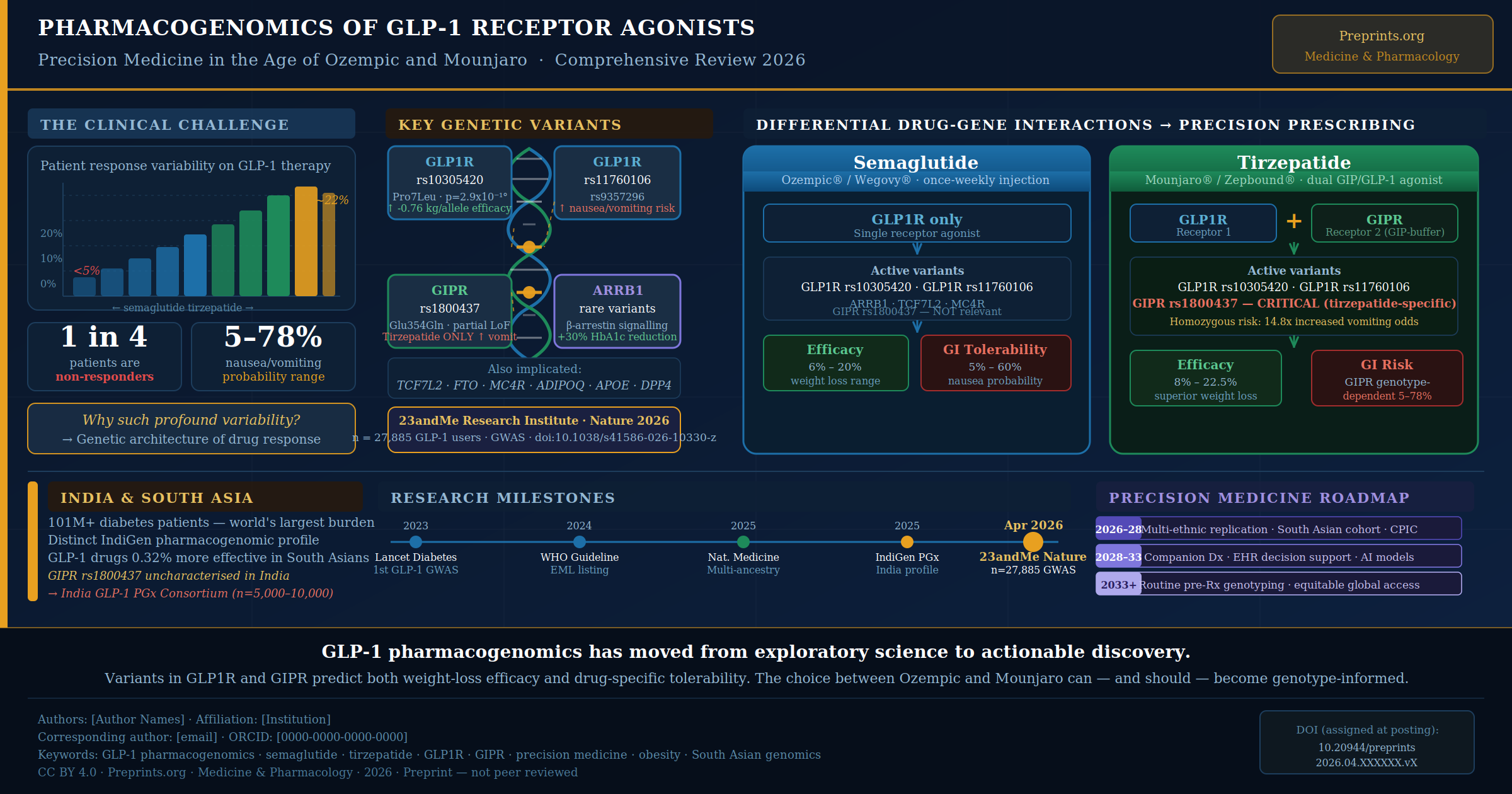
Abstract
Keywords:
1. Introduction: The GLP-1 Revolution and the Variability Problem
2. Mechanistic Foundations: How GLP-1 Drugs Work and Why Genetics Matters
2.1. Semaglutide (Ozempic® / Wegovy®): GLP-1 Receptor Agonism
2.2. Tirzepatide (Mounjaro® / Zepbound®): Dual GIP/GLP-1 Agonism
2.3. Why Genetics Shapes Response
3. Key Genetic Variants: The Emerging Pharmacogenomic Map
4. Landmark Research: From Hypothesis to Genome-Wide Evidence
4.1. The Lancet Diabetes & Endocrinology (2023): The First GLP-1 GWAS

4.2. Nature Medicine (2025): Multi-Ancestry Biobank Analysis
4.3. Nature (April 2026): The 23andMe GWAS — A Field-Defining Study
- Weight-loss efficacy: A missense variant in GLP1R (rs10305420, p.Pro7Leu), located in the receptor’s signal peptide, was associated with approximately 0.76 kg additional weight loss per allele copy (P = 2.9 × 10⁻¹⁰), attributed to enhanced receptor trafficking to the cell surface.
- GI side effects (both drugs): Variants near GLP1R (rs11760106 for vomiting, P = 2.5×10⁻²⁷; rs9357296 for nausea, P = 2.6×10⁻²⁸) were linked to nausea and vomiting across both drug cohorts.
- Tirzepatide-specific vomiting: A partial loss-of-function variant in GIPR (rs1800437, p.Glu354Gln) was associated with increased vomiting risk exclusively in tirzepatide users — not in those on semaglutide. The biological rationale: GIP receptor co-activation normally buffers GLP-1-induced nausea; when GIPR function is impaired, that buffering is lost.
- Compound risk: Individuals homozygous for risk alleles at both GLP1R and GIPR showed 14.8-fold increased odds of tirzepatide-mediated vomiting — a finding potentially enabling genotype-informed drug selection before treatment initiation.
- Predictive modelling: Integrating genetic, demographic, and clinical factors stratified patients into groups with 6%–20% expected weight loss and nausea probability ranging from 5% to 78% — a range with direct clinical significance.

4.4. Tufts University: Next-Generation Quad-Agonist Design (2025)
4.5. AI-Driven Pharmacogenomics (2025)
5. Expanding Clinical Frontiers: Beyond Diabetes and Obesity
6. The Biopharma Perspective
6.1. Novo Nordisk: Biomarker Investments and Precision Obesity Medicine
6.2. Eli Lilly: Tirzepatide Pharmacogenomics and Next-Generation Pipeline
6.3. 23andMe Research Institute: Consumer Genomics Meets Pharmacogenomics
6.4. PGxAI and the AI-Pharmacogenomics Ecosystem
7. India and South Asia: A Critical but Underrepresented Frontier
7.1. The Indian Diabetes and Obesity Burden
7.2. Indian-Specific Pharmacogenomic Profile
7.3. Evidence for Enhanced GLP-1 Efficacy in South Asians
7.4. Indian Expert Perspectives
7.5. Indian Research Institutions and Infrastructure
- CSIR-Institute of Genomics and Integrative Biology (CSIR-IGIB), New Delhi — led the IndiGen pharmacogenomics study; holds the largest Indian whole-genome dataset with GLP1R/GIPR variant frequencies
- Madras Diabetes Research Foundation (MDRF), Chennai — world-class diabetes clinical cohorts; Dr. V. Mohan’s group is the natural home for prospective GLP-1 pharmacogenomic studies
- St. John’s Research Institute, Bangalore — co-led the 2024 BMJ pharmacogenomics paper; has biobank infrastructure for South Asian metabolic disease
- Translational Health Science and Technology Institute (THSTI), Faridabad — national translational research with genomics capabilities
- GenomeIndia project — CSIR-led initiative sequencing 10,000 Indian whole genomes across diverse linguistic/geographic groups; an ideal platform for population-level GLP-1 pharmacogenomics

8. Safety Signals with Pharmacogenomic Implications
8.1. Gastrointestinal Toxicity
8.2. Neuropsychiatric Risk
8.3. Drug–Drug Interaction: Oral Contraception (Tirzepatide)
8.4. Lean Mass Loss
9. Translation Challenges: From Discovery to Clinic
9.1. The Ancestry Diversity Gap
9.2. Self-Report Limitations and Real-World Validation
9.3. Effect Sizes and Polygenic Prediction
9.4. Regulatory and Implementation Pathways
10. Roadmap: Precision Medicine for GLP-1 Pharmacogenomics
Near-Term (2026–2028): Replication and Diversity
- Large-scale replication of Nature 2026 findings in non-European cohorts — specifically South Asian, East Asian, and African ancestry populations
- Prospective India GLP-1 pharmacogenomics cohort study (5,000–10,000 patients; MDRF/CSIR-IGIB partnership; GenomeIndia alignment)
- CPIC guideline development for GLP-1 receptor agonists based on GLP1R and GIPR genotype — the first precision prescribing tool in metabolic medicine
- Integration of GLP-1 pharmacogenomics into existing pre-emptive pharmacogenomic panels (PGx-Passport, YouScript, OneOme RightMed)
Medium-Term (2028–2033): Clinical Implementation
- Companion diagnostic approval (FDA/EMA/CDSCO) for GLP1R/GIPR panel as a prescribing aid for semaglutide vs. tirzepatide selection
- EHR-integrated clinical decision support (building on PGxAI/HealthShare model) flagging high-risk tirzepatide genotypes at point of prescribing
- Pharmacogenomic sub-analyses of GLP-1 trials in new indications (neurodegeneration, addiction, MASH)
- AI-powered multi-omics models integrating genomics, metabolomics, gut microbiome, and clinical variables for holistic response prediction
Long-Term (2033–2040): Routine Genotype-Guided Prescribing
- Standard pre-treatment genotyping for GLP-1 drugs in patients with complex obesity or non-response — analogous to CYP2C19 testing before clopidogrel or BRCA testing before PARP inhibitors
- Next-generation drug design: quad-agonists and CNS-targeted compounds engineered for pharmacogenomic resilience across known genetic backgrounds
- National pharmacogenomics programs in India (GenomeIndia) generating actionable prescribing data for the world’s largest diabetes population
- Equitable access frameworks ensuring precision tools reach low- and middle-income countries where the GLP-1 disease burden is greatest
11. Conclusions
Artificial Intelligence Disclosure
Informed Consent Statement
Data Availability Statement
References
- Auton A et al. (23andMe Research Institute) Genetic predictors of GLP1 receptor agonist weight loss and side effects; Nature, 8 Apr 2026. [Google Scholar] [CrossRef]
- Dawed, A.Y.; et al. Pharmacogenomics of GLP-1 receptor agonists: a genome-wide analysis of observational data and large randomised controlled trials. Lancet Diabetes Endocrinol. 2023, 11(1), 33–41. [Google Scholar] [CrossRef] [PubMed]
- German, J.; et al. Association between plausible genetic factors and weight loss from GLP1-RA and bariatric surgery. Nat Med. 2025, 31, 2269–2276. [Google Scholar] [CrossRef]
- Han, R.; He, P. Pharmacogenomics of Tirzepatide: Genomic Insights into Dual GIP/GLP-1 Agonist Response in Type 2 Diabetes and Atherosclerosis. Pharmaceuticals 2025, 18(9), 1261. [Google Scholar] [CrossRef] [PubMed]
- Sivadas, A.; Sahana, S.; Jolly, B.; et al. Landscape of pharmacogenetic variants associated with non-insulin antidiabetic drugs in the Indian population. BMJ Open Diabetes Res Care 2024, 12(2), e003769. [Google Scholar] [CrossRef]
- Celletti, F.; Farrar, J.; De Regil, L. WHO Guideline on the Use and Indications of GLP-1 Therapies for the Treatment of Obesity in Adults. JAMA 2026, 335(5), 434–438. [Google Scholar] [CrossRef]
- Dinsmore, T.; Kumar, K.; et al. Novel tetra-functional GLP-1/GIP/glucagon/PYY chimeric peptide for weight management. J Am Chem Soc. Full citation to be confirmed upon publication. 2025. [Google Scholar]
- Wilding, J.P.H.; et al. Once-Weekly Semaglutide in Adults with Overweight or Obesity (STEP 1). N Engl J Med. 2021, 384(11), 989–1002. [Google Scholar] [CrossRef] [PubMed]
- Cao, M.; et al. Pathophysiology of type 2 diabetes: A focus on the metabolic differences among southeast Asian, Chinese and Indian populations. Diabetes Obes Metab 2025. [Google Scholar] [CrossRef]
- Lincoff, A.M.; et al. Semaglutide and Cardiovascular Outcomes in Obesity without Diabetes (SELECT). N Engl J Med. 2023, 389(24), 2221–2232. [Google Scholar] [CrossRef]
- Jastreboff, A.M.; et al. Tirzepatide once weekly for the treatment of obesity (SURMOUNT-1). N Engl J Med. 2022, 387(3), 205–216. [Google Scholar] [CrossRef] [PubMed]
- Perkovic, V.; et al. Semaglutide and Kidney Outcomes in Type 2 Diabetes (FLOW). N Engl J Med 2024. [Google Scholar] [CrossRef] [PubMed]
- Pharmacogenomics of anti-obesity drugs: A bioinformatics approach. Research Square 2025. [CrossRef] [PubMed]
- Multi-author. Emerging Frontiers in GLP-1 Therapeutics: A Comprehensive Evidence Base. Pharmaceutics 2025, 17(8), 1036. [CrossRef]
- Additional references 15–50 to be added from manuscript citations — full reference list to be compiled before submission.
| Gene | Variant (rs ID) | Associated Effect | Drug(s) | Evidence |
|---|---|---|---|---|
| GLP1R | rs10305420 (Pro7Leu) | Increased weight loss (~0.76 kg/allele) | Both | Strong: Nature 2026 GWAS (n=27,885) |
| GLP1R | rs6923761 (Gly168Ser) | Altered glycaemic response; sex-specific weight effects | GLP-1 RAs broadly | Moderate: multiple cohort studies |
| GLP1R | rs11760106 / rs9357296 | Increased risk of nausea / vomiting | Both | Strong: Nature 2026 GWAS |
| GLP1R | rs367543060 | Reduced glycaemic response | GLP-1 RAs broadly | Moderate: Kyriakidou et al. |
| GIPR | rs1800437 (Glu354Gln) | Increased vomiting (partial loss-of-function) | Tirzepatide ONLY | Strong: Nature 2026 GWAS (drug-specific) |
| GIPR | rs2287019 (C>T) | Greater fasting glucose improvement | Tirzepatide | Moderate: POUNDS LOST trial |
| ARRB1 | Multiple rare variants | 30% greater HbA1c reduction | GLP-1 RAs broadly | Strong: Lancet Diabetes 2023 (n=4,571) |
| TCF7L2 | rs7903146 | Modifies beta-cell glycaemic response | GLP-1 RAs broadly | Moderate: multiple GWAS |
| FTO | rs9939609 | Adiposity modulation; energy homeostasis | Tirzepatide | Emerging |
| MC4R | Multiple variants | Appetite regulation; weight loss prediction | Both | Emerging — bioinformatics |
| ADIPOQ | Multiple variants | BMI reduction prediction | Tirzepatide | Emerging |
| APOE | e2/e3/e4 | Cardiovascular/lipid response modulation | Tirzepatide (CV) | Emerging |
| DPP4 | Expression eQTLs | Incretin degradation; drug interaction | All GLP-1 RAs | Emerging — GTEx data |
| Indication | Key Evidence | Status (2024–26) |
|---|---|---|
| Type 2 Diabetes | ADA first-line injectable; HbA1c −1.5–2.0% | Fully approved (all agents) |
| Obesity | STEP/SURMOUNT: 7–24% weight loss | Semaglutide, Tirzepatide approved |
| Cardiovascular Risk | SELECT trial: 20% MACE reduction | Semaglutide FDA-approved (Mar 2024) |
| Chronic Kidney Disease | FLOW trial: reduced kidney failure progression | Semaglutide FDA-approved (Jan 2025) |
| Obstructive Sleep Apnea | SURMOUNT-OSA: significant AHI reduction | Tirzepatide FDA-approved (Dec 2024) |
| MASH (Liver Disease) | Phase 3 data emerging | Under regulatory review |
| Osteoarthritis | STEP-9: 14.1-point WOMAC improvement vs placebo | Strong signals; off-label |
| Neurodegeneration (AD/PD) | 70% reduced AD risk vs insulin (US EHR, n>1M) | Phase 2 trials ongoing |
| Substance Use Disorders | Lower opioid overdose/alcohol intoxication (n=1.3M) | Phase 2 trials ongoing |
| Adolescent Obesity | STEP TEENS; 600% Rx increase 2020–2024 | Semaglutide/Liraglutide approved ≥12y |
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2026 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).




