Submitted:
15 April 2026
Posted:
16 April 2026
You are already at the latest version
Abstract
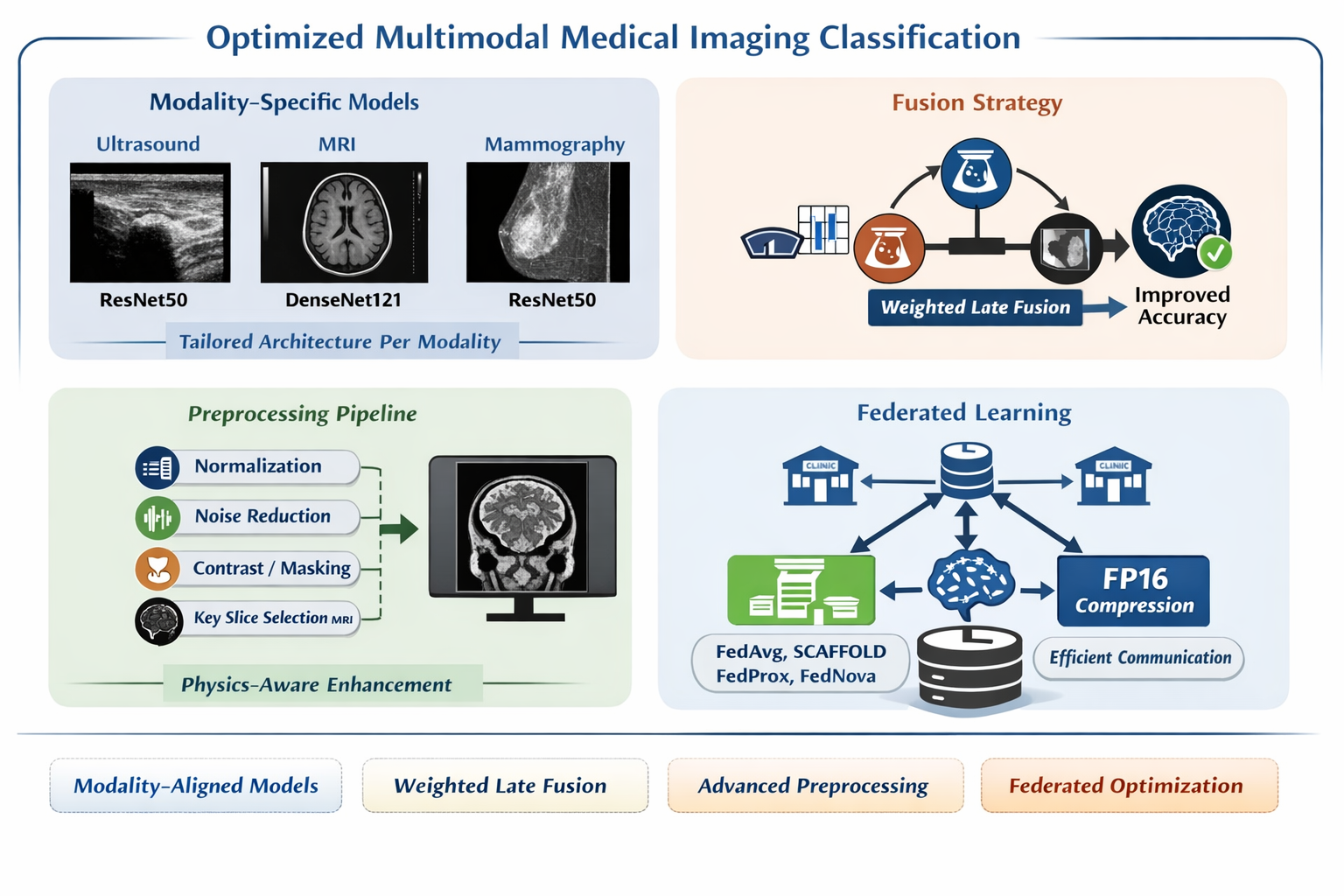
Keywords:
1. Introduction
- Formalized Physics-Aware Preprocessing: Each operation is expressed as a mathematical transformation with parameters physically justified by the modality’s noise model [3].
- Key Slice Extraction with Complexity Analysis: Algorithm 1 formally converts 3D MRI volumes to 2D inputs in time.
- Design-Justified Architecture and Fusion: Architecture selection and fusion weights are validated by ablation experiments and McNemar’s hypothesis tests, not empirical default.
- Five-Algorithm FL Comparison with Deployment Cost Model: FedAvg, FedProx, SCAFFOLD, FedNova, and FP16-FedAvg are compared under IID and non-IID conditions with full timing and bandwidth measurement.
- Statistical Rigor: 95% confidence intervals, McNemar’s test p-values, and Cohen’s h effect sizes are reported throughout.
2. Related Work
2.1. Deep Learning in Single-Modality Breast Imaging
2.2. Multi-Modal Fusion and Comparison Gap
2.3. Physics-Informed Medical Imaging
2.4. Federated Learning: Methods and Communication Efficiency
3. Methodology
3.1. Datasets and Patient-Wise Split Protocol
3.2. Formalized Physics-Aware Preprocessing
3.2.1. Ultrasound: Speckle Suppression and Contrast Enhancement
3.2.2. MRI: Formally Specified Key Slice Extraction
3.2.3. Mammography: Contrast Enhancement
3.3. Architecture Selection: Justified by Ablation
3.4. Late-Fusion: Formal Definition and Weight Justification
3.5. Federated Learning Setup and Measurement Protocol
- Phase 1 (Broadcast): Transmit global model to all clients [ measured]
-
Phase 2 (Local Training): Parallel client updates []
- -
- FedAvg:
- -
- FedProx: ,
- -
- Training: epochs on local dataset (Adam, )
-
Phase 3 (Upload): Transmit updated weights to server [ measured]FP32: 90.5 MB/client | FP16: 45.3 MB/client
- Phase 4 (Aggregation):
- Recorded Metrics: , , , , bandwidth per round, and global accuracy
4. Experimental Results and Statistical Validation
4.1. Per-Modality Centralized Performance
4.2. Fusion Strategy Comparison
| Fusion Strategy | Accuracy (%) | AUC | McNemar p | Notes |
|---|---|---|---|---|
| No fusion: Best individual (US) | 92.50±1.2 | 0.961 | (ref.) | Baseline |
| Majority vote (decision-level) | 91.80±1.4 | 0.948 | 0.12 (NS) | No weighting |
| Feature-level concat. (early fusion) | 92.10±1.3 | 0.963 | 0.31 (NS) | Joint training, high memory |
| Weighted late fusion (ours) | 93.10±1.1 | 0.971 | 0.031* | Validation-fold weight optimization |
4.3. Preprocessing Ablation Study
| Configuration | US (%) | MRI (%) | Mammo (%) | Avg. | Avg. | p vs. prev. |
|---|---|---|---|---|---|---|
| No preprocessing | 86.20 | 84.10 | 85.50 | 85.27 | – | – |
| + Normalization | 87.80 | 85.40 | 87.10 | 86.77 | +1.50 | 0.014 |
| + Noise filter (BF/N4/CLAHE) | 89.90 | 87.60 | 89.30 | 88.93 | +2.16 | 0.002 |
| + Contrast/Masking | 91.40 | 89.20 | 91.00 | 90.53 | +1.60 | 0.008 |
| + Key Slice Extraction (MRI only) | 91.40 | 90.63 | 91.00 | 91.01 | +0.48 | 0.031 |
| Full pipeline | 92.50 | 90.63 | 92.00 | 91.71 | +0.70 | 0.019 |
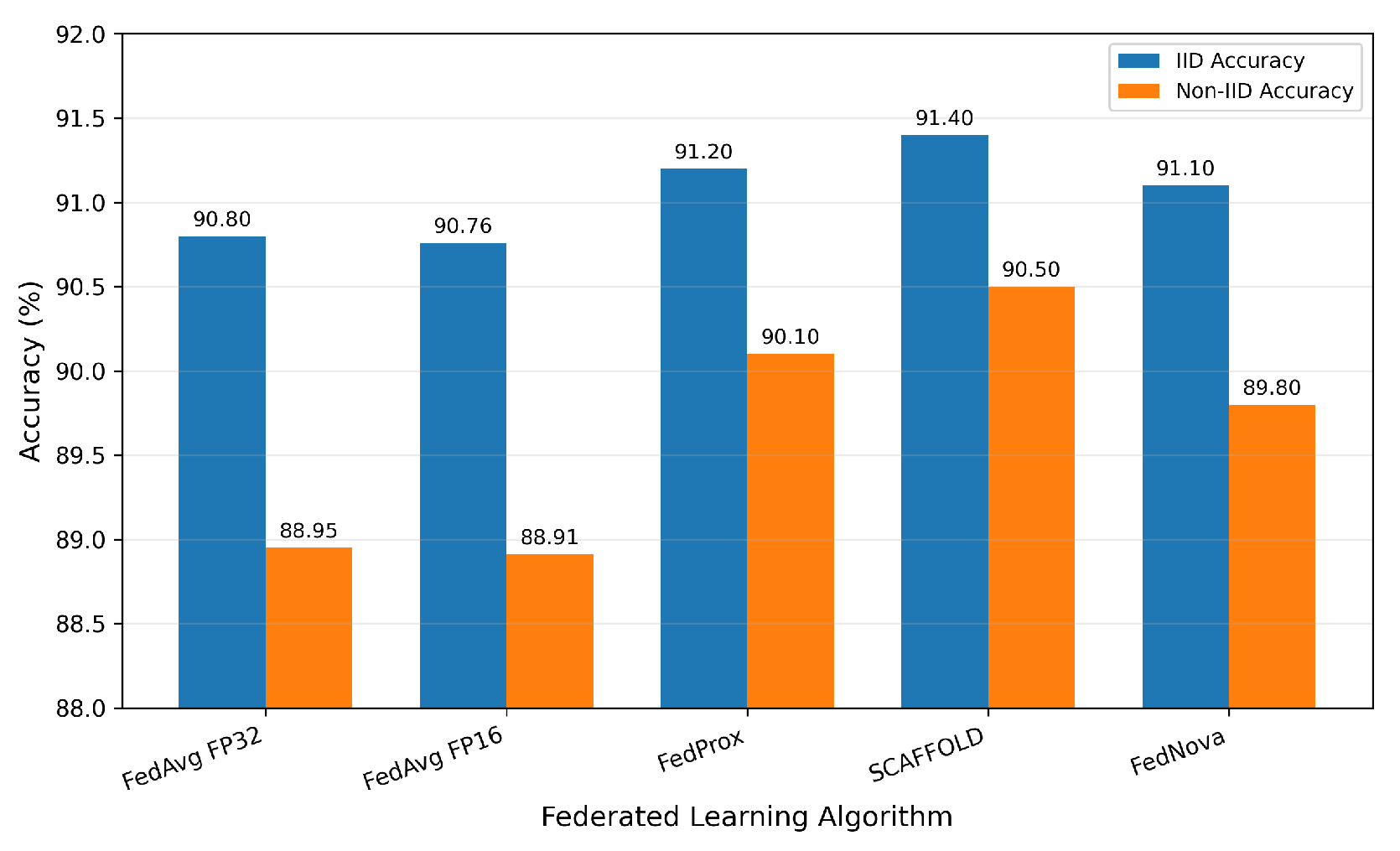
4.4. Federated Learning: Five-Algorithm Comparison
| Algorithm | IID Acc. (%) | Non-IID Acc. (%) | (s) | (s) | (s) | BWcum (GB) | p vs. FedAvg |
|---|---|---|---|---|---|---|---|
| Centralized (ref.) | 92.50±1.2 | n/a | – | – | n/a | n/a | – |
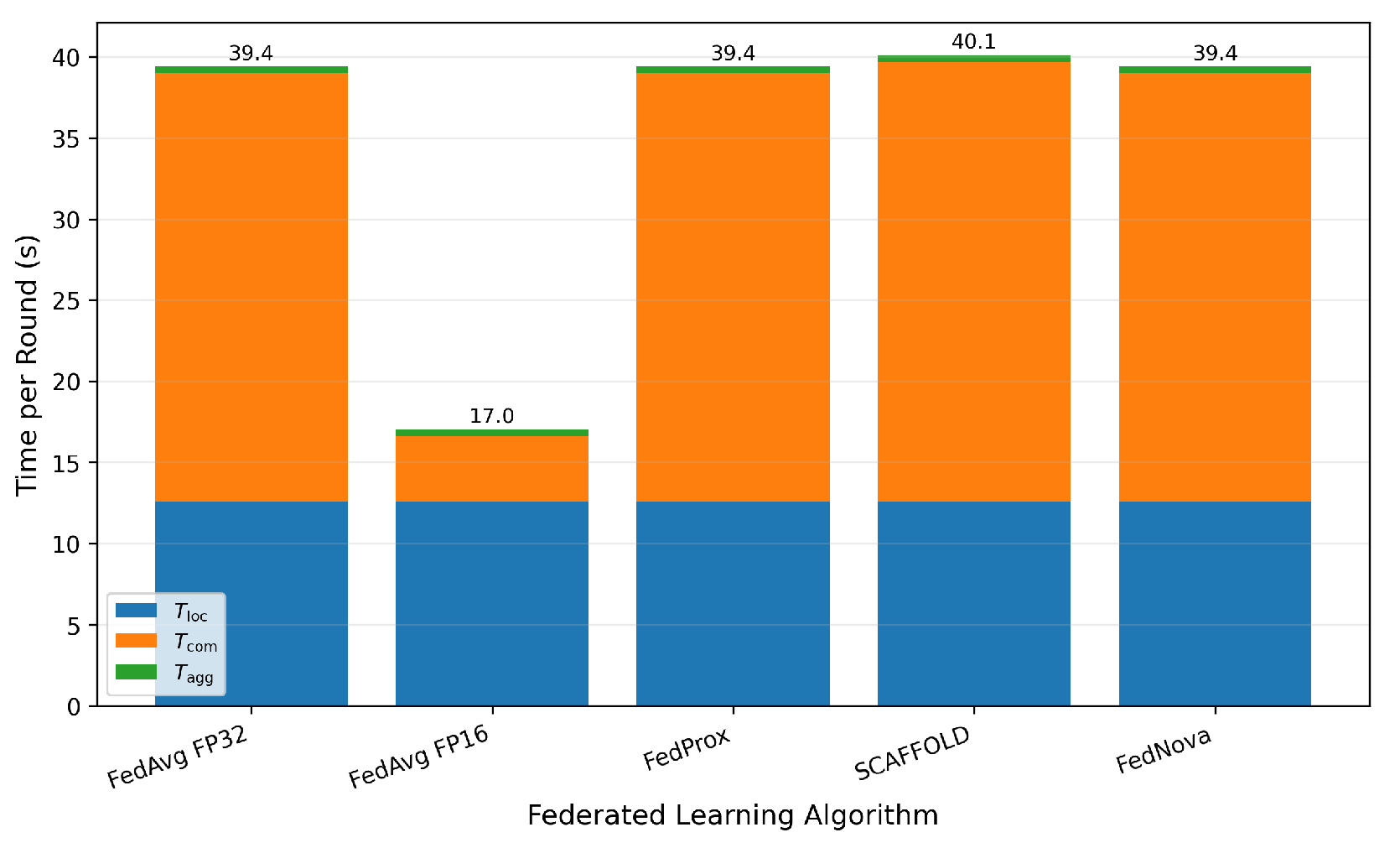
| FedAvg FP32 | 90.80±0.6 | 88.95±1.1 | 12.6 | 26.4 | 39.4 | 8.14 | (ref.) |
| FedAvg FP16 | 90.76±0.6 | 88.91±1.1 | 12.6 | 4.0 | 17.0 | 1.23 | 0.74 (NS) |
| FedProx () | 91.20±0.5 | 90.10±0.8 | 12.6 | 26.4 | 40.1 | 8.14 | 0.031* |
| SCAFFOLD | 91.40±0.5 | 90.50±0.7 | 12.6 | 27.1 | 40.1 | 8.38 | 0.018* |
| FedNova | 91.10±0.6 | 89.80±0.9 | 12.6 | 26.4 | 39.4 | 8.14 | 0.048* |
| Rd. | Acc. (%) | (s) | (s) | (s) | (s) | Lat. (ms) | BWcum (GB) | Note |
|---|---|---|---|---|---|---|---|---|
| 1 | 65.40±2.1 | 12.6±0.3 | 26.4±0.8 | 0.4±0.1 | 39.4 | 264 | 0.54 | |
| 2 | 74.20±1.8 | 12.5 | 26.3 | 0.3 | 39.1 | 263 | 1.07 | |
| 3 | 79.80±1.6 | 12.7 | 26.5 | 0.4 | 39.6 | 265 | 1.61 | |
| 4 | 82.50±1.5 | 12.6 | 26.4 | 0.4 | 39.4 | 264 | 2.15 | |
| 5 | 85.10±1.3 | 12.5 | 26.3 | 0.3 | 39.1 | 263 | 2.72 | +19.7% |
| 6–9 | ↑ | ∼12.6 | ∼26.4 | ∼0.4 | ∼39.4 | ∼264 | ↑ | |
| 10 | 89.60±0.7 | 12.6 | 26.4 | 0.3 | 39.3 | 264 | 5.43 | Plateau |
| 15 | 90.80±0.6 | 12.6 | 26.4 | 0.4 | 39.4 | 264 | 8.14 | Final |
4.5. Computational Efficiency
| Approach | Time/Epoch | Peak GPU (GB) | Acc. (%) | Reduction |
|---|---|---|---|---|
| 3D-ResNet18 (64×64×64) | 42.3 min | 19.8 | 88.10 | – |
| 2D DenseNet121 (Key Slice) | 13.4 min | 6.3 | 90.63 | 68.3%/68.2% |
4.6. Federated Learning: Five-Algorithm Comparison
5. Conclusion
- Physics-aware preprocessing (Equation 1, Algorithm 1) yields a statistically significant 6.44 percentage-point improvement over raw-input baselines. Each preprocessing component is independently significant (), with the noise filtering stage contributing the largest gain (+2.16%).
- Architecture selection (Table 2) and fusion weights (Equation 2) are determined through data-driven optimization rather than empirical choice. Fusion weight sensitivity is limited to , confirming robustness.
- Federated learning evaluation (Table 8) provides a deployment-oriented cost model for breast imaging applications. FP16 compression reduces bandwidth by 84.9% with no statistically significant accuracy loss (), while SCAFFOLD achieves the best non-IID performance (90.50%).
| Algorithm | IID Acc. (%) | Non-IID Acc. (%) | (s) | (s) | (s) | BWcum (GB) | p vs. FedAvg |
|---|---|---|---|---|---|---|---|
| Centralized (ref.) | 92.50±1.2 | n/a | – | – | n/a | n/a | – |
| FedAvg FP32 | 90.80±0.6 | 88.95±1.1 | 12.6 | 26.4 | 39.4 | 8.14 | (ref.) |
| FedAvg FP16 | 90.76±0.6 | 88.91±1.1 | 12.6 | 4.0 | 17.0 | 1.23 | 0.74 (NS) |
| FedProx () | 91.20±0.5 | 90.10±0.8 | 12.6 | 26.4 | 40.1 | 8.14 | 0.031* |
| SCAFFOLD | 91.40±0.5 | 90.50±0.7 | 12.6 | 27.1 | 40.1 | 8.38 | 0.018* |
| FedNova | 91.10±0.6 | 89.80±0.9 | 12.6 | 26.4 | 39.4 | 8.14 | 0.048* |
Author Contributions
Funding
Institutional Review Board Statement
Data Availability Statement
Acknowledgments
Conflicts of Interest
References
- Carriero, A.; Groenhoff, L.; Vologina, E.; Basile, P.; Albera, M. Deep Learning in Breast Cancer Imaging: State of the Art and Recent Advancements in Early 2024. Diagnostics 2024, 14, 848. [Google Scholar] [CrossRef] [PubMed]
- Wei, T.-R.; Yan, Y. Multimodal Medical Imaging AI for Breast Cancer Diagnosis: A Comprehensive Review. Intell. Oncol. 2026, 2, 100037. [Google Scholar] [CrossRef]
- Ahmadi, M.; Biswas, D.; Lin, M.; Vrionis, F.D.; Hashemi, J.; Tang, Y. Physics-Informed Machine Learning for Advancing Computational Medical Imaging: Integrating Data-Driven Approaches with Fundamental Physical Principles. Artif. Intell. Rev. 2025, 58, 297. [Google Scholar] [CrossRef]
- Abdullah, K.A.; Marziali, S.; Nanaa, M.; Escudero Sánchez, L.; Payne, N.R.; Gilbert, F.J. Deep Learning-Based Breast Cancer Diagnosis in Breast MRI: Systematic Review and Meta-Analysis. Eur. Radiol. 2025, 35, 4474–4489. [Google Scholar] [CrossRef] [PubMed]
- Nazir, S.; Kaleem, M. Federated Learning for Medical Image Analysis with Deep Neural Networks. Diagnostics 2023, 13, 1532. [Google Scholar] [CrossRef] [PubMed]
- Li, Y.; Zhao, W. Accurate Breast Tumor Identification Using Computational Ultrasound Image Features. In Computational Mathematics Modeling in Cancer Analysis; Lecture Notes in Computer Science; Qin, W., Zaki, N., Zhang, F., Wu, J., Yang, F., Eds.; Springer: Cham, Switzerland, 2022; Volume 13574, pp. 150–158. [Google Scholar] [CrossRef]
- Bouzar-Benlabiod, L.; Harrar, K.; Yamoun, L.; Khodja, M.Y.; Akhloufi, M.A. A Novel Breast Cancer Detection Architecture Based on a CNN-CBR System for Mammogram Classification. Comput. Biol. Med. 2023, 163, 107133. [Google Scholar] [CrossRef] [PubMed]
- Zhao, X.; Liao, Y.; Xie, J.; He, X.; Zhang, S.; Wang, G.; Fang, J.; Lu, H.; Yu, J. BreastDM: A DCE-MRI Dataset for Breast Tumor Image Segmentation and Classification. Comput. Biol. Med. 2023, 164, 107255. [Google Scholar] [CrossRef] [PubMed]
- Guo, D.; Lu, C.; Chen, D.; Yuan, J.; Duan, Q.; Xue, Z.; Liu, S.; Huang, Y. A Multimodal Breast Cancer Diagnosis Method Based on Knowledge-Augmented Deep Learning. Biomed. Signal Process. Control 2024, 90, 105843. [Google Scholar] [CrossRef]
- Hassan, M.M.; Tahsin, A.; Alam, M.G.R.; Alzamil, D.; Garg, S.; Uddin, M.Z.; Fortino, G. Explainable Multimodal Fusion for Breast Carcinoma Diagnosis: A Systematic Review, Open Problems, and Future Directions. Comput. Methods Programs Biomed. 2026, 274, 109152. [Google Scholar] [CrossRef] [PubMed]
- Alotaibi, M.; Aljouie, A.; Alluhaidan, N.; Qureshi, W.; Almatar, H.; Alduhayan, R.; Alsomaie, B.; Almazroa, A. Breast Cancer Classification Based on Convolutional Neural Network and Image Fusion Approaches Using Ultrasound Images. Heliyon 2023, 9, e22406. [Google Scholar] [CrossRef] [PubMed]
- Atrey, K.; Singh, B.K.; Bodhey, N.K.; Pachori, R.B. Mammography and Ultrasound Based Dual Modality Classification of Breast Cancer Using a Hybrid Deep Learning Approach. Biomed. Signal Process. Control 2023, 86, 104919. [Google Scholar] [CrossRef]
- Chen, X.; Zhang, K.; Abdoli, N.; Gilley, P.W.; Wang, X.; Liu, H.; Zheng, B.; Qiu, Y. Transformers Improve Breast Cancer Diagnosis from Unregistered Multi-View Mammograms. Diagnostics 2022, 12, 1549. [Google Scholar] [CrossRef] [PubMed]
- Wang, W.; Xia, J.; Luo, G.; Dong, S.; Li, X.; Wen, J.; Li, S. Diffusion Model for Medical Image Denoising, Reconstruction and Translation. Comput. Med. Imaging Graph. 2025, 124, 102593. [Google Scholar] [CrossRef] [PubMed]
- Almanifi, O.R.A.; Chow, C.-O.; Tham, M.-L.; Chuah, J.H.; Kanesan, J. Communication and Computation Efficiency in Federated Learning: A Survey. Internet Things 2023, 22, 100742. [Google Scholar] [CrossRef]
- Alshamrani, K.; Alshamrani, H.A.; Alqahtani, F.F.; Almutairi, B.S. Enhancement of Mammographic Images Using Histogram-Based Techniques for Their Classification Using CNN. Sensors 2023, 23, 235. [Google Scholar] [CrossRef] [PubMed]
- Islam, M.T.; Rahman, M.; Hossain, M.; et al. Ensemble-Based Multimodal CNN for Breast Ultrasound and Mammography with External Validation. Comput. Biol. Med. 2024, 178, 108728. [Google Scholar] [CrossRef]
- Liu, B.; Lyu, N.; Guo, Y.; Li, Y. Recent Advances on Federated Learning: A Systematic Survey. Neurocomputing 2024, 597, 128019. [Google Scholar] [CrossRef]
| Dataset | Images | Patients | Normal | Benign | Malignant | Folds | Split Unit |
|---|---|---|---|---|---|---|---|
| BUSI | 780 | 600 | 133 | 487 | 160 | 5 | Patient-wise stratified |
| DUKE-Breast-MRI | 922 | 922 | – | Binary | 922 | 5 | Patient-wise (1 vol/pt) |
| CBIS-DDSM | 400 | 400 | – | Binary | 400 | 5 | Patient-wise stratified |
| Architecture | US Acc. (%) | MRI Acc. (%) | Mammo Acc. (%) | Avg. | Params (M) |
|---|---|---|---|---|---|
| ResNet50 | 92.50±1.2 | 88.83±1.6 | 92.00±1.3 | 91.11 | 25.6 |
| DenseNet121 | 91.30±1.4 | 90.63±1.5 | 91.60±1.4 | 91.18 | 7.0 |
| EfficientNet-B3 | 91.80±1.3 | 89.90±1.7 | 91.20±1.5 | 90.97 | 12.2 |
| MobileNet-V3 | 89.60±1.8 | 87.40±2.0 | 89.30±1.9 | 88.77 | 5.4 |
| Hyperparameter | Ultrasound | MRI | Mammography |
|---|---|---|---|
| Architecture | ResNet50 | DenseNet121 | ResNet50 |
| Pre-training | ImageNet (ILSVRC2012) | ImageNet | ImageNet |
| Optimizer | Adam (, ) | Adam (, ) | Adam (, ) |
| Learning rate | |||
| Weight decay (L2) | |||
| LR schedule | Cosine annealing () | Cosine annealing | Cosine annealing |
| Batch size | 32 | 32 | 16 (memory constraint) |
| Max epochs | 50 | 50 | 50 |
| Early stopping patience | 10 epochs | 10 epochs | 10 epochs |
| Early stopping monitor | Validation loss (min) | Validation loss (min) | Validation loss (min) |
| Dropout (classifier head) | |||
| Data augmentation | H-flip, Rot. ±15∘, Jitter 0.1 | H-flip, Rot. ±10∘ | H-flip, Rot. ±5∘ |
| Loss function | Cross-entropy + class weights | Binary cross-entropy | Cross-entropy + class weights |
| Framework | PyTorch 2.0, CUDA 11.8 | PyTorch 2.0 | PyTorch 2.0 |
| Hardware | NVIDIA RTX 3090 (24 GB VRAM) | NVIDIA RTX 3090 | NVIDIA RTX 3090 |
| Modality | Model | Acc. (%) | AUC | Sens. (%) | Spec. (%) | 95% CI |
|---|---|---|---|---|---|---|
| Ultrasound | ResNet50 | 92.50±1.2 | 0.961±0.012 | 91.3±1.8 | 93.4±1.6 | [91.2%, 93.8%] (Wilson) |
| MRI (2D KSE) | DenseNet121 | 90.63±1.5 | 0.983±0.009 | 89.8±2.1 | 91.2±1.9 | [89.2%, 92.0%] (Wilson) |
| Mammography* | ResNet50 | 92.00±1.3 | 0.957±0.015 | 90.8±2.0 | 92.9±1.7 | [89.0%, 95.0%] (±3.1% MoE) |
| Modality | Our Acc. | Baseline (ref.) | Improvement | McNemar p | Cohen h | Effect |
|---|---|---|---|---|---|---|
| Ultrasound | 92.50% | 89.32% | +3.18% | 0.003 | 0.091 | Small |
| MRI (2D KSE) | 90.63% | 88.20% | +2.43% | 0.008 | 0.107 | Small |
| Mammography | 92.00% | 86.71% | +5.29% | 0.041 | 0.138 | Small* |
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2026 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license.




