Submitted:
14 April 2026
Posted:
15 April 2026
You are already at the latest version
Abstract
Keywords:
1. Introduction
2. Materials and Methods
2.1. Collection of Grain Samples
2.2. DNA Extraction and Amplicon Sequencing
2.3. Bioinformatics and Statistical Analysis
3. Results
3.1. Changes in pH During Fermentation
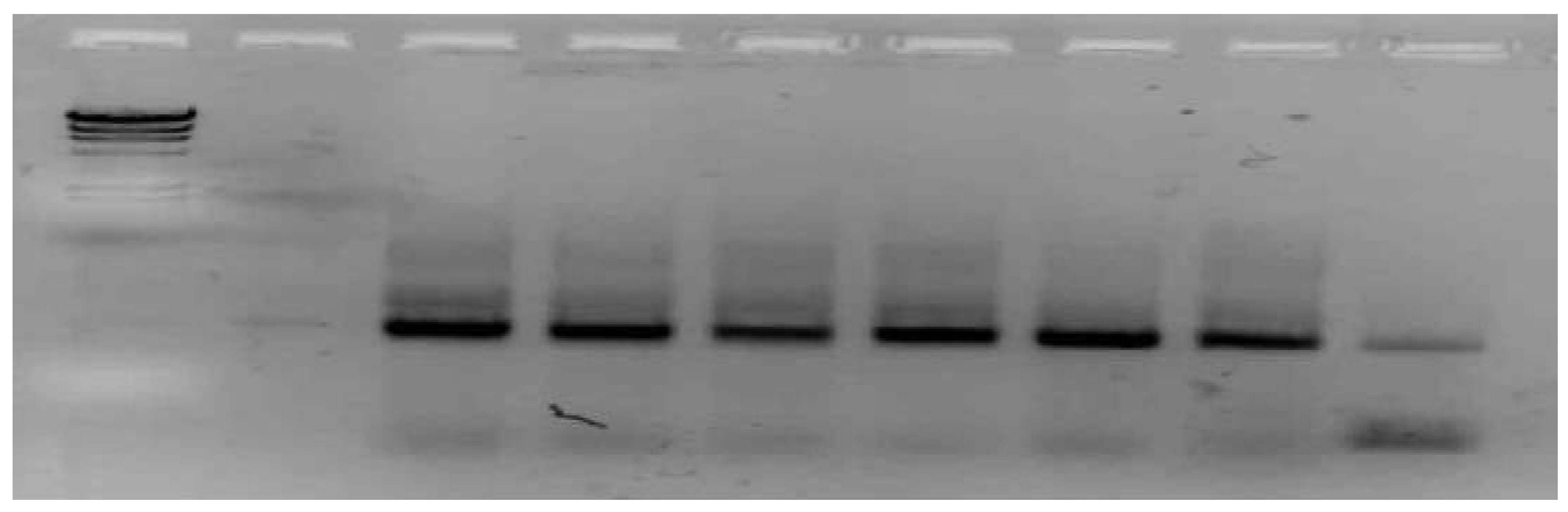
3.2. PCR Amplification and Sequencing Quality
3.3. DNA Yield and Quality
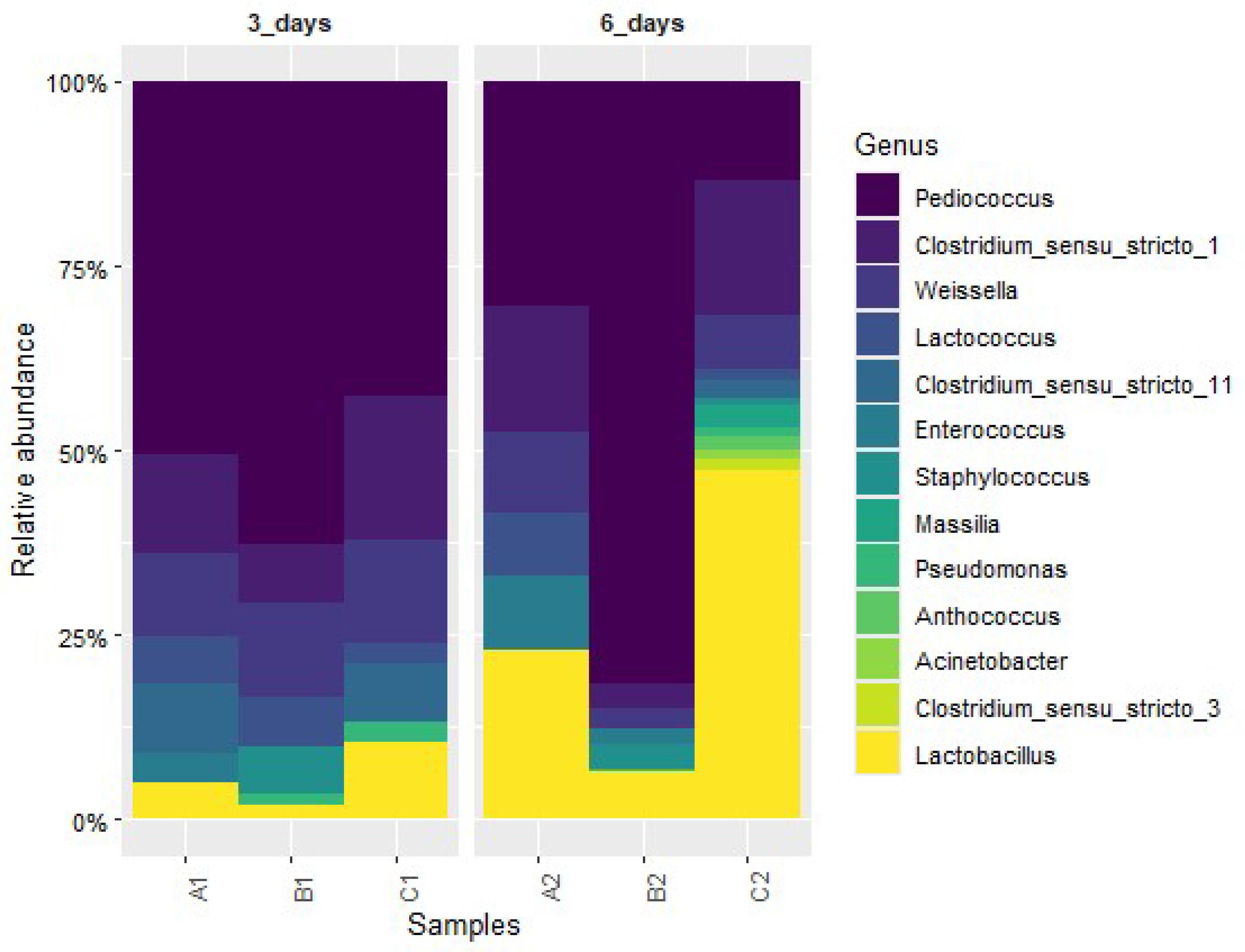
3.3. Genus-Level Microbial Community Composition
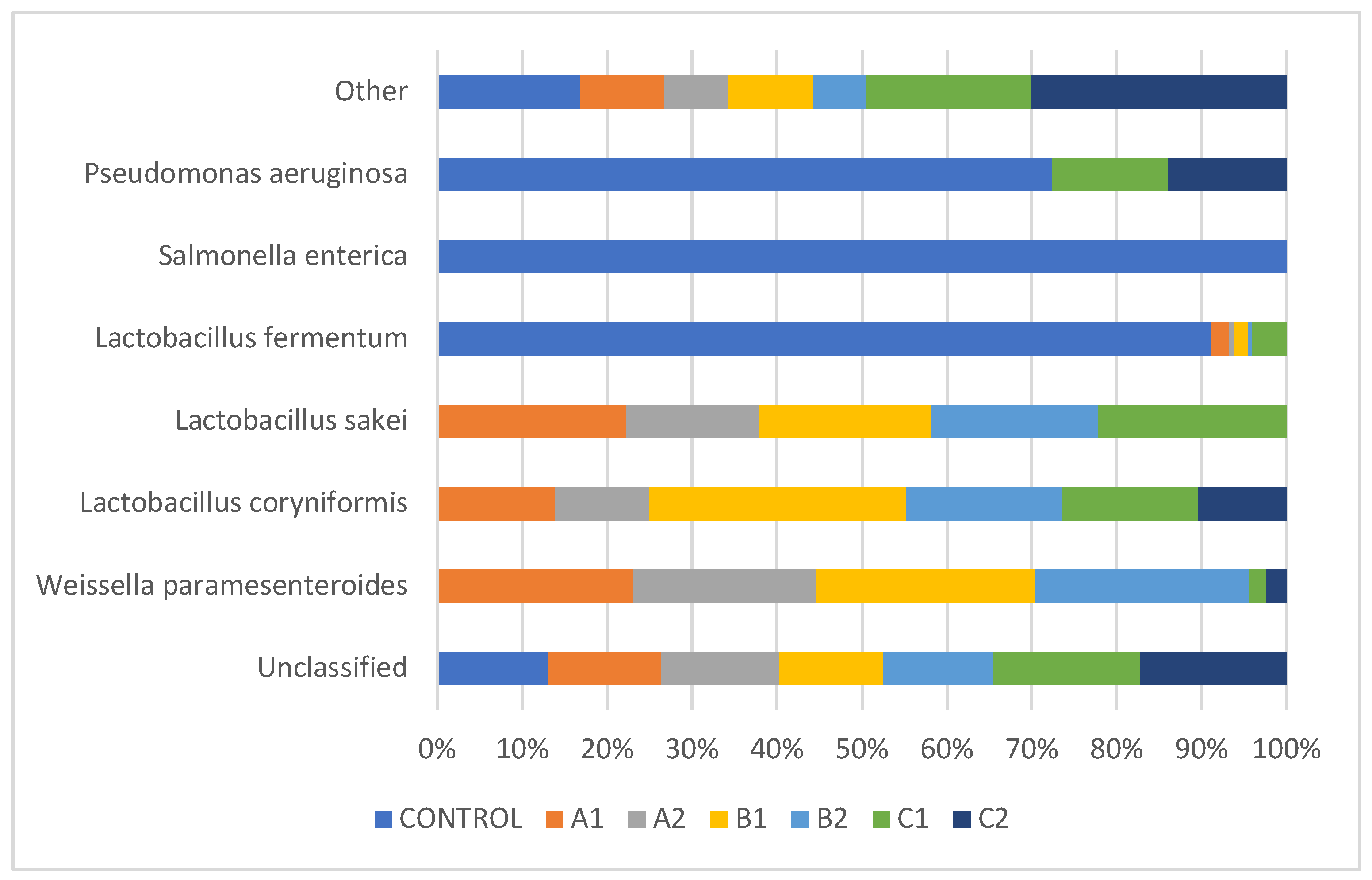
3.4. Species-Level Microbial Community Composition
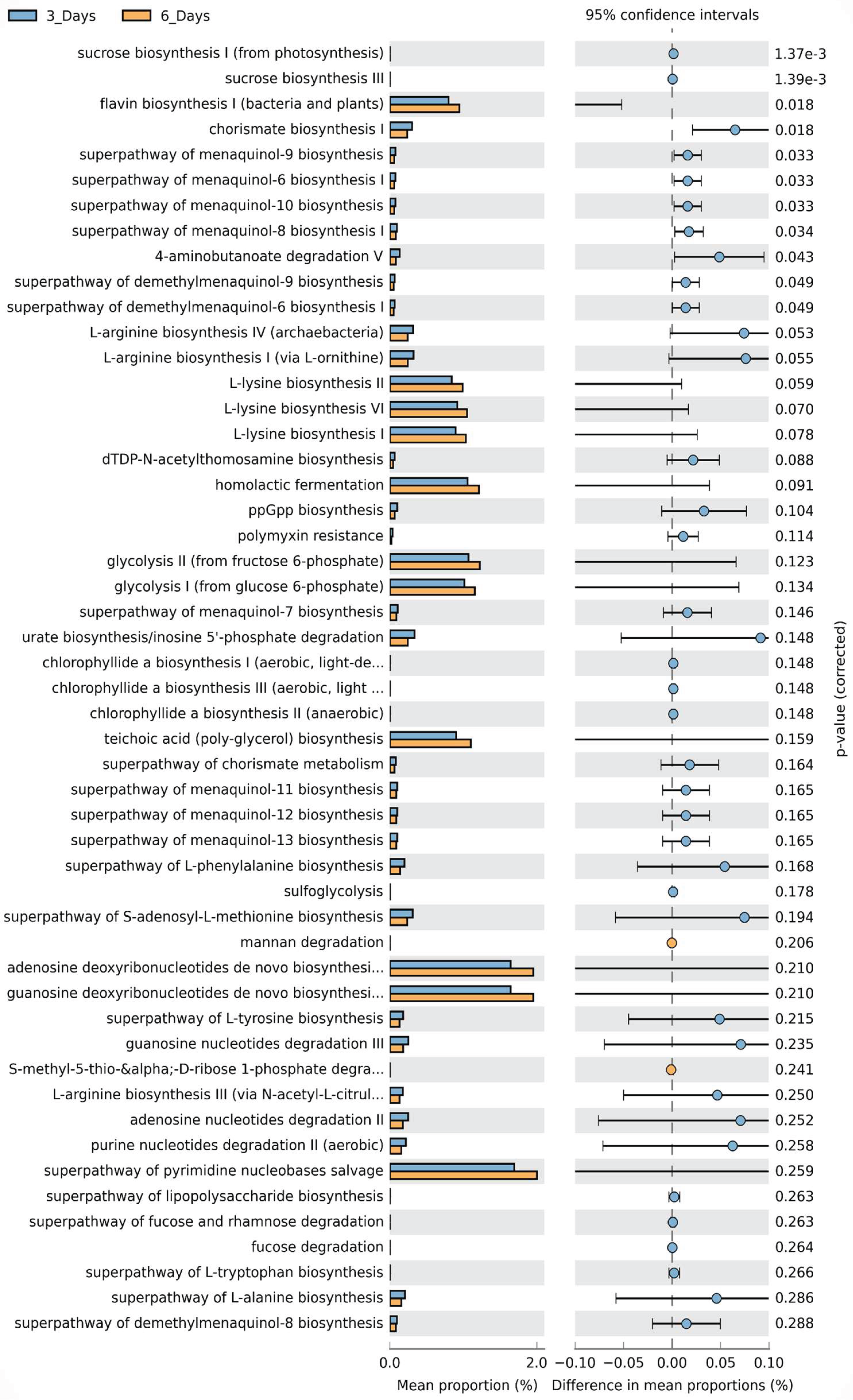
3.5. Functional Preddiction of Microbial Communities
4. Discussion
5. Conclusions
Supplementary Materials
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Acknowledgments
Conflicts of Interest
Abbreviations
| ASV | Amplicon Sequence Variant |
| DNA | Deoxyribonucleic Acid |
| LAB | Lactic Acid Bacteria |
| PCR | Polymerase Chain Reaction |
| rRNA | Ribosomal Ribonucleic Acid |
| NanoDrop | Spectrophotometric instrument used for nucleic acid quantification |
| SILVA | Ribosomal RNA gene database used for taxonomic classification |
| PICRUSt2 | Phylogenetic Investigation of Communities by Reconstruction of Unobserved States (version 2) |
References
- Prajapati, J.; Nair, B. The history of fermented foods in Fermented Functional Foods. Handb Fermented Foods 2003, 1–25. [Google Scholar]
- Gänzle, M. The periodic table of fermented foods: limitations and opportunities. Appl. Microbiol. Biotechnol. 2022, 106, 2815–2826. [Google Scholar] [CrossRef] [PubMed]
- Pswarayi, F.; Gänzle, M. African cereal fermentations: A review on fermentation processes and microbial composition of non-alcoholic fermented cereal foods and beverages. Int. J. Food Microbiol. 2022, 378, 109815. [Google Scholar] [CrossRef] [PubMed]
- Obafemi, Y.D.; Oranusi, S.U.; Ajanaku, K.O.; Akinduti, P.A.; Leech, J.; Cotter, P.D. African fermented foods: overview, emerging benefits, and novel approaches to microbiome profiling. npj Sci. Food 2022, 6, 1–9. [Google Scholar] [CrossRef]
- Borrelli, G.M.; Ficco, D.B.M. Cereals and Cereal-Based Foods: Nutritional, Phytochemical Characterization and Processing Technologies. Foods 2025, 14, 1234. [Google Scholar] [CrossRef]
- Gunathunga, C.; Senanayake, S.; Jayasinghe, M.A.; Brennan, C.S.; Truong, T.; Marapana, U.; Chandrapala, J. Germination effects on nutritional quality: A comprehensive review of selected cereals and pulses changes. J. Food Compos. Anal. 2024, 128. [Google Scholar] [CrossRef]
- Garrido-Galand, S.; Asensio-Grau, A.; Calvo-Lerma, J.; Heredia, A.; Andrés, A. The potential of fermentation on nutritional and technological improvement of cereal and legume flours: A review. Food Res. Int. 2021, 145, 110398. [Google Scholar] [CrossRef]
- Iqbal, B.; Alabbosh, K.F.; Raheem, A.; Ullah, I.; Khan, A.R.; Memon, M.S. Microbial transformation: the role of fermentation in advancing nutritional quality and human health. Arch. Microbiol. 2025, 207, 1–13. [Google Scholar] [CrossRef]
- Atter, A.; Diaz, M.; Tano-Debrah, K.; Kunadu, A.P.-H.; Mayer, M.J.; Sayavedra, L.; Misita, C.; Amoa-Awua, W.; Narbad, A. The predominant lactic acid bacteria and yeasts involved in the spontaneous fermentation of millet during the production of the traditional porridge Hausa koko in Ghana. BMC Microbiol. 2024, 24, 1–17. [Google Scholar] [CrossRef]
- Kandler, O. Carbohydrate metabolism in lactic acid bacteria. Antonie van Leeuwenhoek 1983, 49, 209–224. [Google Scholar] [CrossRef]
- Gautam, A.; Poopalarajah, R.; Ahmad, A.R.; Rana, B.N.; Denekew, T.W.; Ahn, N.; Utenova, L.; Kunwor, Y.S.; Bhandari, N.N.; Jha, A.R. Ecological factors that drive microbial communities in culturally diverse fermented foods. BMC Microbiol. 2025, 25, 1–20. [Google Scholar] [CrossRef]
- Arsov, A.; Tsigoriyna, L.; Batovska, D.; Armenova, N.; Mu, W.; Zhang, W.; Petrov, K.; Petrova, P. Bacterial Degradation of Antinutrients in Foods: The Genomic Insight. Foods 2024, 13, 2408. [Google Scholar] [CrossRef]
- Kazemi, S.; Homayouni-Rad, A.; Kafil, H.S.; Sarabi-Aghdam, V.; Zeynolabedini, P.; pour Agha, B.; Madadi, S.A. Selection of appropriate probiotic yeasts for use in dairy products: a narrative review. Food Prod. Process. Nutr. 2025, 7, 1–28. [Google Scholar] [CrossRef]
- Sharma, S.; Sharma, R.; Govindaraj, M.; Mahala, R.S.; Satyavathi, C.T.; Srivastava, R.K.; Gumma, M.K.; Kilian, B. Harnessing wild relatives of pearl millet for germplasm enhancement: Challenges and opportunities. Crop. Sci. 2020, 61, 177–200. [Google Scholar] [CrossRef]
- Srivastava, R.K.; Singh, R.B.; Pujarula, V.L.; Bollam, S.; Pusuluri, M.; Chellapilla, T.S.; Yadav, R.S.; Gupta, R. Genome-Wide Association Studies and Genomic Selection in Pearl Millet: Advances and Prospects. Front. Genet. 2020, 10, 1389. [Google Scholar] [CrossRef] [PubMed]
- Rai, K.; Singh, P.; Ameta, V.; Gupta, S.K.; Jayalekha, A.; Mahala, R.; Pareek, S.; Swami, M.; Verma, Y. Seed set variability under high temperatures during flowering period in pearl millet (Pennisetum glaucum L. (R.) Br.). Field Crop. Res. 2015, 171, 41–53. [Google Scholar] [CrossRef]
- Krishnan, R.; Meera, M.S. Pearl millet minerals: effect of processing on bioaccessibility. J. Food Sci. Technol. 2018, 55, 3362–3372. [Google Scholar] [CrossRef]
- Boncompagni, E.; Orozco-Arroyo, G.; Cominelli, E.; Gangashetty, P.I.; Grando, S.; Zu, T.T.K.; Daminati, M.G.; Nielsen, E.; Sparvoli, F. Antinutritional factors in pearl millet grains: Phytate and goitrogens content variability and molecular characterization of genes involved in their pathways. PLOS ONE 2018, 13, e0198394. [Google Scholar] [CrossRef]
- Rault, A.; Bouix, M.; Béal, C. Fermentation pH Influences the Physiological-State Dynamics of Lactobacillus bulgaricus CFL1 during pH-Controlled Culture. Appl. Environ. Microbiol. 2009, 75, 4374–4381. [Google Scholar] [CrossRef] [PubMed]
- Savoie, S.; Champagne, C.; Chiasson, S.; Audet, P. Media and process parameters affecting the growth, strain ratios and specific acidifying activities of a mixed lactic starter containing aroma-producing and probiotic strains. J. Appl. Microbiol. 2006, 103, 163–174. [Google Scholar] [CrossRef] [PubMed]
- Admassie, M. A Review on Food Fermentation and the Biotechnology of Lactic Acid Bacteria. World J. Food Sci. Technol. 2018, 2, 19. [Google Scholar] [CrossRef]
- Zapaśnik, A.; Sokołowska, B.; Bryła, M. Role of Lactic Acid Bacteria in Food Preservation and Safety. Foods 2022, 11, 1283. [Google Scholar] [CrossRef] [PubMed]
- Atuna, R.A.; Akabanda, F.; Bletz, S.; Scherff, N.; Makurat, J.; Bordewick-Dell, U.; Lamping, M.; Ritter, G.; Mellmann, A.; Amagloh, F.K. Bacterial community dynamics in spontaneously fermented Ghanaian cereals, legumes, root, tuber and plantain flours. BMC Microbiol. 2025, 25, 1–12. [Google Scholar] [CrossRef]
- Zhao, X.; Xiang, F.; Tang, F.; Cai, W.; Guo, Z.; Hou, Q.; Yang, X.; Song, W.; Shan, C. Bacterial Communities and Prediction of Microbial Metabolic Pathway in Rice Wine Koji From Different Regions in China. Front. Microbiol. 2022, 12, 748779. [Google Scholar] [CrossRef]
- Abedfar, A.; Hosseininezhad, M.; Sadeghi, A.; Raeisi, M.; Feizy, J. Investigation on “spontaneous fermentation” and the productivity of microbial exopolysaccharides by Lactobacillus plantarum and Pediococcus pentosaceus isolated from wheat bran sourdough. LWT 2018, 96, 686–693. [Google Scholar] [CrossRef]
- Han, D.; Yan, Q.; Liu, J.; Jiang, Z.; Yang, S. Transcriptomic Analysis of Pediococcus pentosaceus Reveals Carbohydrate Metabolic Dynamics Under Lactic Acid Stress. Front. Microbiol. 2021, 12. [Google Scholar] [CrossRef]
- Liu, L.; Zhu, T.; Yang, J.; Zhao, J.; Jiang, J.; Zhang, W.; Liu, M.; Shi, Y.; Li, H.; Hu, Y. Metabolic network of lactic acid bacteria driving the formation of characteristic flavor compounds in traditional Yunnan fermented soy whey (FSW) and their formation mechanisms. LWT 2025, 230. [Google Scholar] [CrossRef]
- Deatraksa, J.; Sunthornthummas, S.; Rangsiruji, A.; Sarawaneeyaruk, S.; Suwannasai, N.; Pringsulaka, O. Isolation of folate-producing Weissella spp. from Thai fermented fish (Plaa Som Fug). LWT 2018, 89, 388–391. [Google Scholar] [CrossRef]
- Liu, M.; Li, X.; Ye, T.; Zhao, L.; Zhang, X. Safety evaluation of Weissella confusa SY628 and the effect of its fermentation on the taste and quality of soy yogurt. Front. Microbiol. 2025, 16, 1567399. [Google Scholar] [CrossRef] [PubMed]
- Zagorec, M.; Champomier-Vergès, M.-C. Lactobacillus sakei: A Starter for Sausage Fermentation, a Protective Culture for Meat Products. Microorganisms 2017, 5, 56. [Google Scholar] [CrossRef] [PubMed]
- O’cAllaghan, J.L.; Willner, D.; Buttini, M.; Huygens, F.; Pelzer, E.S. Limitations of 16S rRNA Gene Sequencing to Characterize Lactobacillus Species in the Upper Genital Tract. Front. Cell Dev. Biol. 2021, 9, 641921. [Google Scholar] [CrossRef] [PubMed]
- Niya, B.; Yaakoubi, K.; Beraich, F.Z.; Arouch, M.; Kadmiri, I.M. Current status and future developments of assessing microbiome composition and dynamics in anaerobic digestion systems using metagenomic approaches. Heliyon 2024, 10, e28221. [Google Scholar] [CrossRef]
- Raes, E.J.; Karsh, K.; Sow, S.L.S.; Ostrowski, M.; Brown, M.V.; van de Kamp, J.; Franco-Santos, R.M.; Bodrossy, L.; Waite, A.M. Metabolic pathways inferred from a bacterial marker gene illuminate ecological changes across South Pacific frontal boundaries. Nat. Commun. 2021, 12, 1–12. [Google Scholar] [CrossRef]
- Naylor, D.; McClure, R.; Jansson, J. Trends in Microbial Community Composition and Function by Soil Depth. Microorganisms 2022, 10, 540. [Google Scholar] [CrossRef] [PubMed]
- Dania, M.I.; Faraji, B.; Wachira, J. Micronutrient Biosynthesis Potential of Spontaneous Grain Fermentation Microbiomes. Int. J. Environ. Res. Public Heal. 2022, 19, 16621. [Google Scholar] [CrossRef] [PubMed]
| Parameter | 0h | 72h | 144h |
|---|---|---|---|
| pH (mean, n=3) | 6.50 | 4.09 | 3.59 |
| Sample | Fermentation Time (hours) | DNA Concentration (ng/µL) by Nanodrop |
|---|---|---|
| A1 | 72 | 13.8 |
| B1 | 72 | 59.7 |
| C1 | 72 | 7.1 |
| A2 | 144 | 28.4 |
| B2 | 144 | 18.9 |
| C2 | 144 | 18.2 |
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2026 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).




