Submitted:
14 April 2026
Posted:
15 April 2026
You are already at the latest version
Abstract
Keywords:
1. Introduction
2. Literature Review and Key Issues
2.1. Vocal Biomarkers for ASD Detection
2.2. Self-Supervised and Contrastive Representation Learning
2.3. Generalization to Unseen Individuals: A Critical Challenge
2.4. Key Issues Addressed in This Work
- Limited robustness to inter-speaker variability: Existing supervised classifiers may overfit to cohort-specific patterns.
- Under-explored use of supervised contrastive learning in ASD vocal analysis.
- Need for cross-context modeling combining structured and spontaneous vocalizations.
3. Framework
3.1. Dataset
3.2. Preprocessing
- 1.
- Digitization: Audio signals are resampled at 16,000 Hz using the Librosa library.
- 2.
- Segmentation: Continuous recordings are sliced into 0.5s chunks with a 0.1s hop length, creating a high-density time series.
- 3.
- Domain Transformation: Each time-series segment is converted into its frequency-domain representation via Fast Fourier Transform (FFT).
- 4.
- Data Augmentation: Stochastic augmentations are applied to both domains to increase the diversity of the contrastive pairs.
- 5.
- Shuffling & Stratification: Data is shuffled and split using a stratified approach to preserve class distribution across batches ().
- Participant Distribution and Data Split.
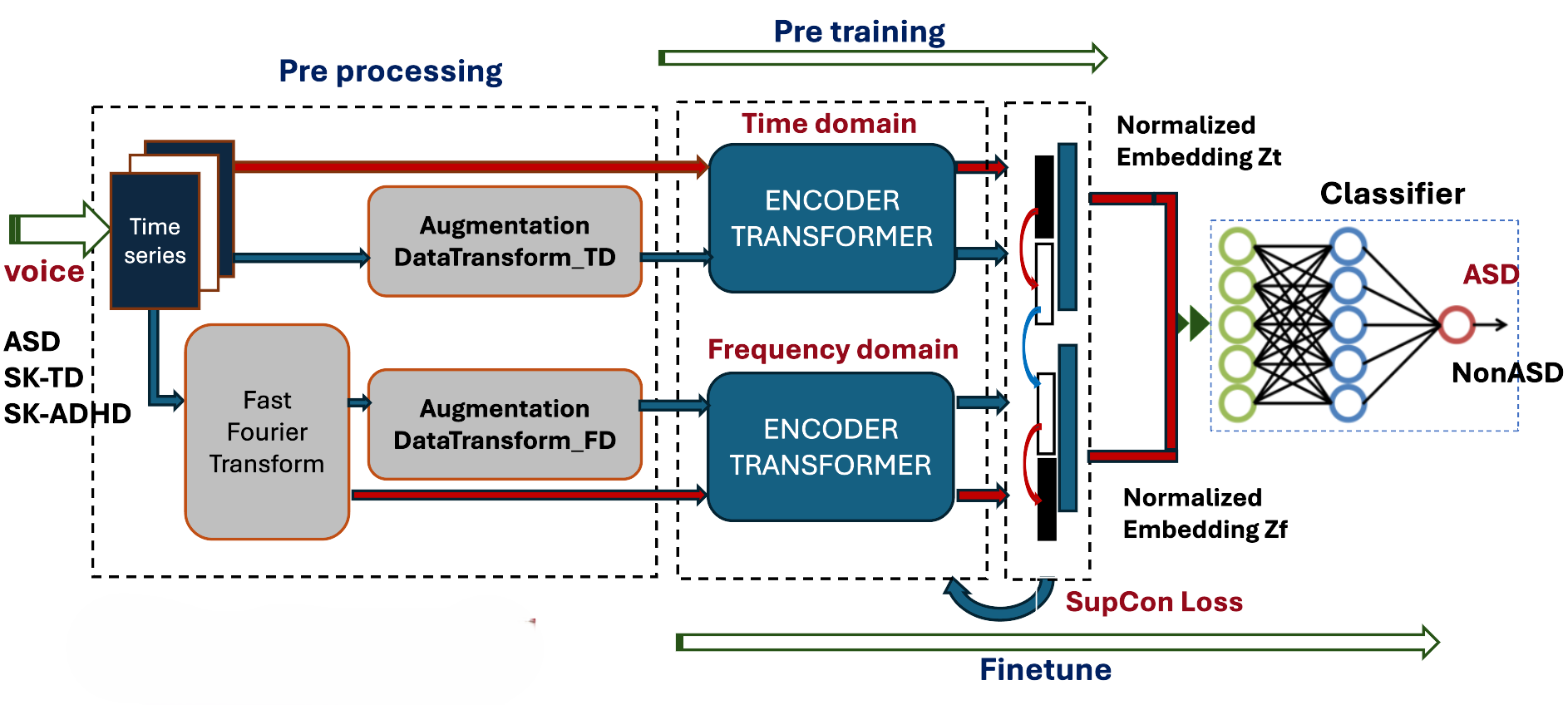
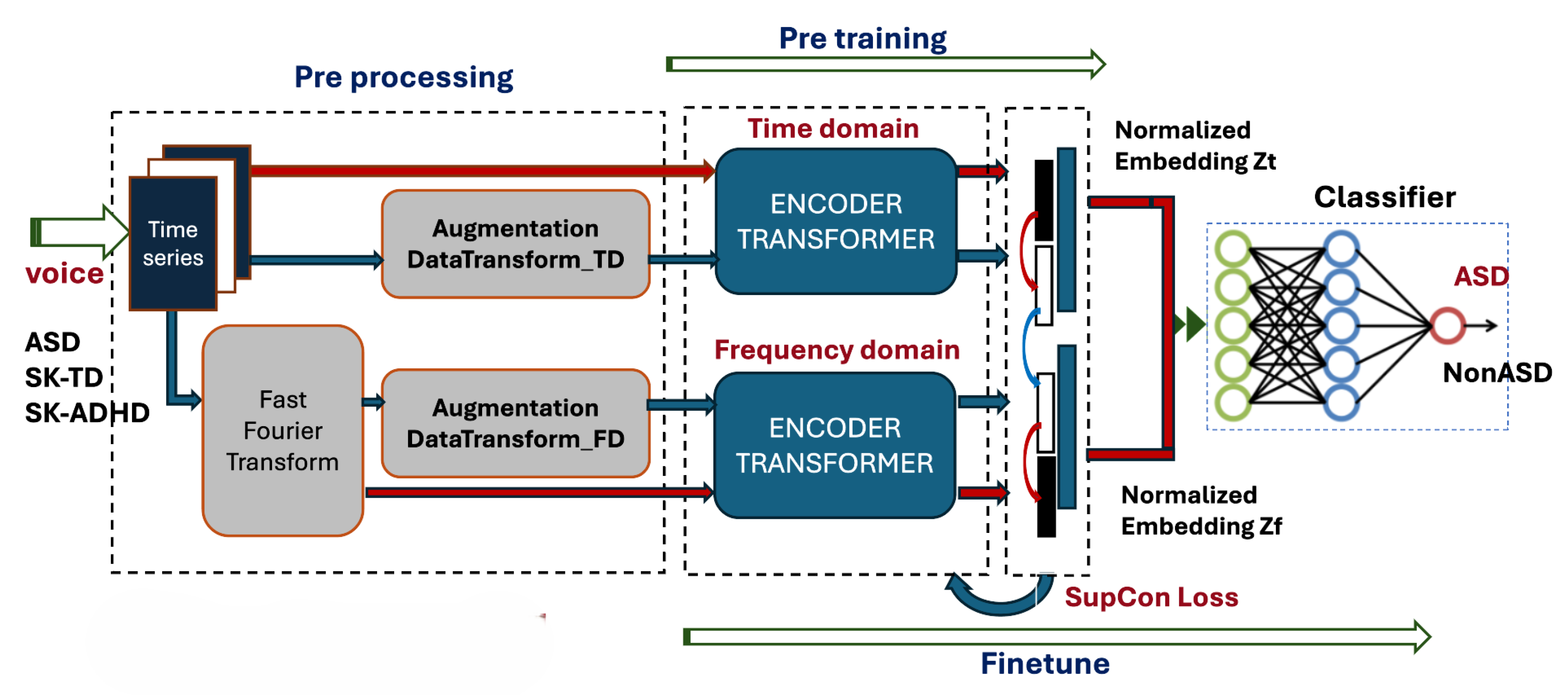
3.3. Contrastive Model Architecture
- Pre-training Phase: Two separate Transformer-based encoders process the time-series and the frequency spectra. The resulting latent vectors are normalized and projected into a joint embedding space. Instead of a self-supervised loss, we implement a Supervised Contrastive (SupCon) Loss, which encourages embeddings of the same class (e.g., ASD) to cluster together while pushing different classes apart.
- Fine-tuning Phase: The pre-trained encoders are coupled with a linear classifier. The final classification (presence or absence of ASD) is performed by concatenating the temporal and spectral representations, optimized again via the SupCon objective to refine decision boundaries.
3.3.1. Supervised Contrastive Loss (SupCon) in Pre-Training
- is the set of indices of all positives in the multiview batch relative to anchor i.
- is the set of all indices in the batch.
- is a scalar temperature parameter that controls the difficulty of the contrastive task.
3.3.2. Fine-Tuning and Classification
- Downstream Classifier Architecture.
- Fine-tuning Strategy.
3.4. Experimental Configuration
4. Results and Discussion
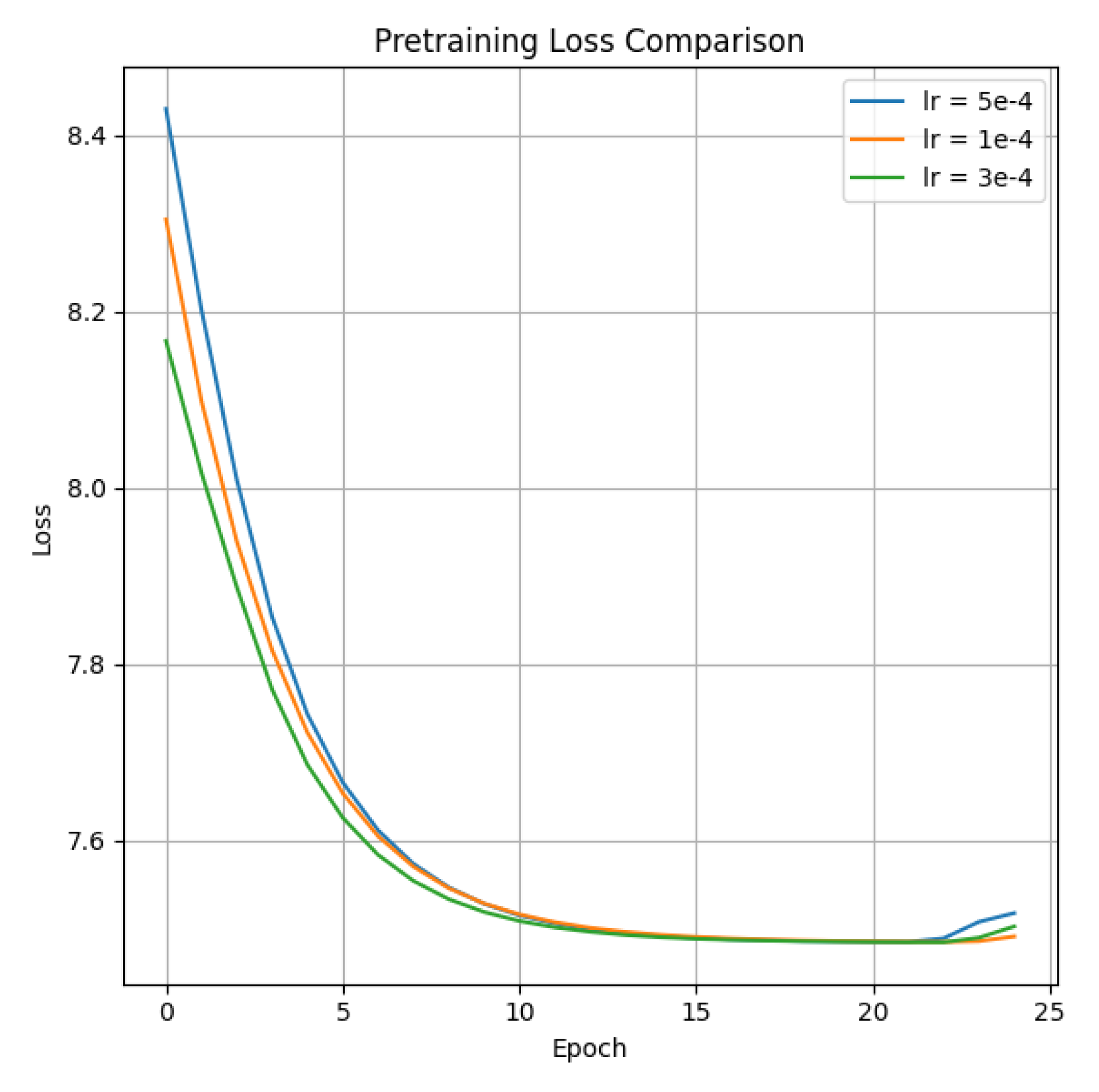
4.1. Pre-Training Results
- Training Dynamics.
- Learning Rate Impact.
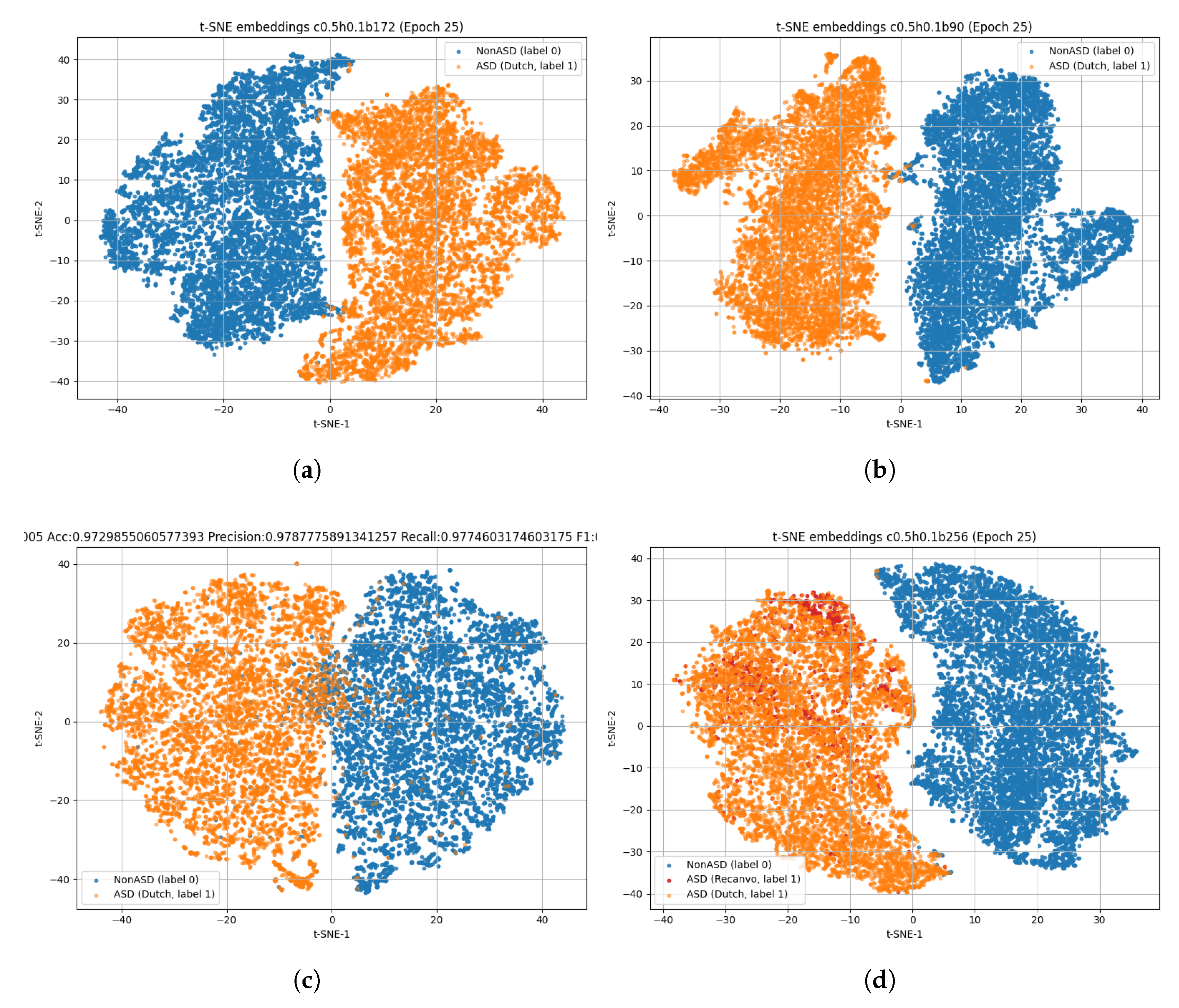
4.2. Fine-Tuning Results
- Embedding Space Analysis.
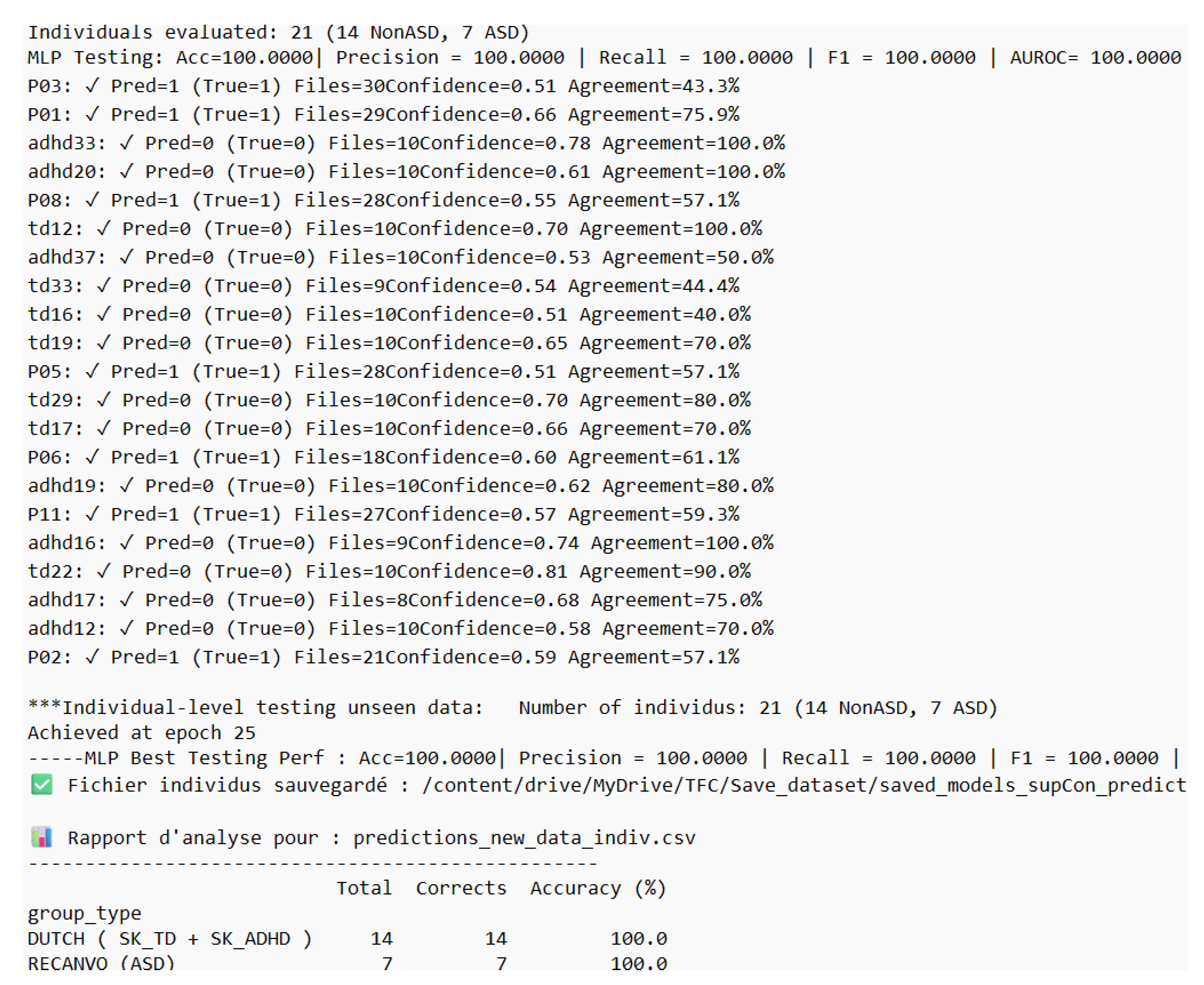
4.3. Inference Performance
4.3.1. Data Used in Training: Dutch + ReCANVo
4.3.2. Data Used in Training: Dutch Only
4.4. Comparison with Vocal Biomarker Techniques
4.5. Discussion
- Regularization through Data Heterogeneity and Learning Rate Calibration
- Analysis of Model Generalizability across Heterogeneous Cohorts
- Optimization of Class Separability and Intra-class Variance
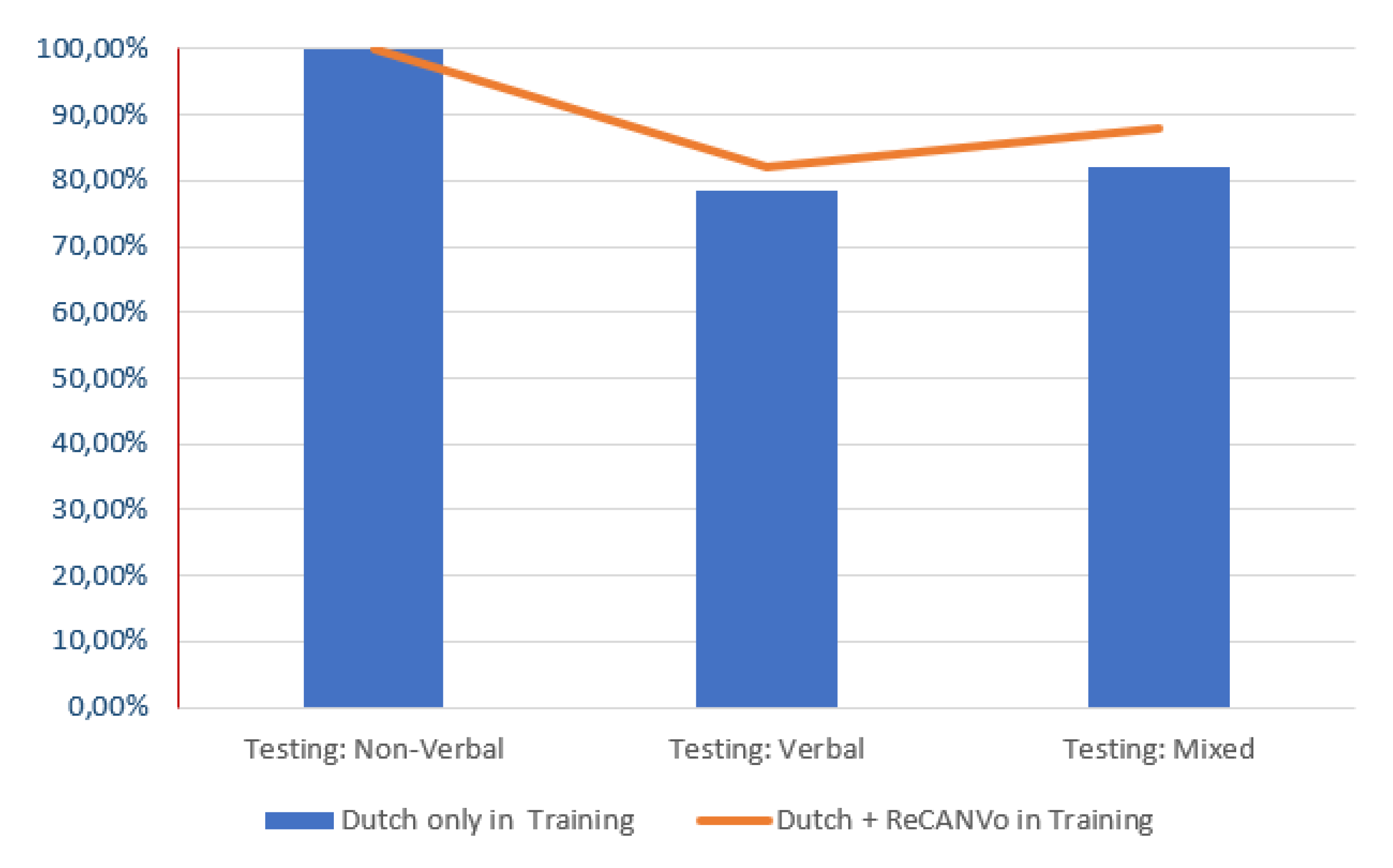
- Better prediction with non-verbal vocalizations
5. Conclusions
- Code Availability
Institutional Review Board Statement
Informed Consent Statement
Conflicts of Interest
References
- Zhang, X.; Zhao, Z.; Tsiligkaridis, T.; Zitnik, M. Self-Supervised Contrastive Pre-Training For Time Series via Time-Frequency Consistency. Proc. Neural Inf. Process. Syst. (NeurIPS), 2022. [Google Scholar]
- Khosla, P.; Teterwak, P.; Wang, C.; Sarna, A.; Tian, Y.; Isola, P.; Maschinot, A.; Liu, C.; Krishnan, D. Supervised Contrastive Learning. arXiv 2020, arXiv, 2004.11362. Available online: https://arxiv.org/abs/2004.11362.
- Hu, C.; Thrasher, J.; Li, W.; Ruan, M.; Yu, X.; Paul, L.; Wang, S.; Li, X. Exploring speech pattern disorders in autism using machine learning. arXiv 2024. [Google Scholar] [CrossRef]
- Sai, K.; Krishna, R.; Radha, K.; Rao, D.; Muneera, A. Automated ASD detection in children from raw speech using customized STFT–CNN model. Int. J. Speech Technol. 2024, 27, 701–716. [Google Scholar] [CrossRef]
- Vacca, J.; Brondino, N.; Dell’Acqua, F.; Vizziello, A.; Savazzi, P. Automatic Voice Classification of Autistic Subjects. arXiv 2024. [Google Scholar] [CrossRef]
- Deng, S.; Kosloski, E.; Patel, S.; Barnett, Z.; Nan, Y.; Kaplan, A.; Aarukapalli, S.; Doan, W.T.; Wang, M.; Singh, H.; et al. Hear Me, See Me, Understand Me: Audio–Visual Autism Behavior Recognition. arXiv 2024. [Google Scholar] [CrossRef]
- Chi, N.; Washington, P.; Kline, A.; Husic, A.; Hou, C.; He, C.; Dunlap, K.; Wall, D. Classifying autism from crowdsourced semi-structured speech recordings: A machine learning approach. arXiv 2022. [Google Scholar] [CrossRef]
- Murugaiyan, S.; Uyyala, S. Aspect-based sentiment analysis of customer speech data using deep CNN and BiLSTM. Cogn. Comput. 2023, 15, 914–931. [Google Scholar] [CrossRef]
- Rakotomanana, H.; Rouhafzay, G. A Scoping Review of AI-Based Approaches for Detecting Autism Traits Using Voice and Behavioral Data. Bioengineering contentReference[oaicite:0]index=0. 2025, 12, 1136. [Google Scholar] [CrossRef] [PubMed]
- Briend, F.; David, C.; Silleresi, S.; Malvy, J.; Ferré, S.; Latinus, M. Voice acoustics allow classifying autism spectrum disorder with high accuracy. Transl. Psychiatry 2023, 13, 250. [Google Scholar] [CrossRef] [PubMed]
- Lee, J.; Lee, G.; Bong, G.; Yoo, H.; Kim, H. Deep-learning–based detection of infants with autism spectrum disorder using autoencoder feature representation. Sensors 2020, 20, 6762. [Google Scholar] [CrossRef] [PubMed]
- Li, M.; Tang, D.; Zeng, J.; Zhou, T.; Zhu, H.; Chen, B.; Zou, X. An automated assessment framework for atypical prosody and stereotyped idiosyncratic phrases related to autism spectrum disorder. Comput. Speech Lang. 2019, 56, 80–94. [Google Scholar] [CrossRef]
- Rosales-Pérez, A.; Reyes-García, C.; Gonzalez, J.; Reyes-Galaviz, O.; Escalante, H.; Orlandi, S. Classifying infant cry patterns by the genetic selection of a fuzzy model. Biomed. Signal Process. Control 2015, 17, 38–46. [Google Scholar] [CrossRef]
- Asgari, M.; Chen, L.; Fombonne, E. Quantifying voice characteristics for detecting autism. Comput. Speech Lang. 2021, 12, 665096. [Google Scholar] [CrossRef] [PubMed]
- Lehnert-LeHouillier, H.; Terrazas, S.; Sandoval, S. Prosodic Entrainment in Conversations of Verbal Children and Teens on the Autism Spectrum. Front. Psychol. 2020, 11, 582221. [Google Scholar] [CrossRef] [PubMed]
- Mohanta, A.; Mittal, V. Analysis and classification of speech sounds of children with autism spectrum disorder using acoustic features. Comput. Speech Lang. 2022, 72, 101287. [Google Scholar] [CrossRef]
- Nature Digital Medicine. Reproducibility, external validation, and generalization in clinical machine learning. Nat. Digit. Med. 2023. [Google Scholar] [CrossRef]
- Psychiatry, Molecular. Cross-cohort generalization challenges in psychiatric biomarkers. Mol. Psychiatry 2023. [Google Scholar] [CrossRef]
- Laguna, A.; Pusil, S.; Paltrinieri, A. L.; Orlandi, S. Automatic Cry Analysis: Deep Learning for Screening of Autism Spectrum Disorder in Early Childhood. J. Autism Dev. Disord. 2025. [Google Scholar] [CrossRef] [PubMed]
- Chen, T.; Kornblith, S.; Norouzi, M.; Hinton, G. A Simple Framework for Contrastive Learning of Visual Representations. Proc. Mach. Learn. Res. 2020, 119, 1597–1607. Available online: https://arxiv.org/abs/2002.05709.
- Leng, Y.; Anwar, S. M.; Rekik, I.; He, S.; Lee, E.-J. Self-Supervised Graph Transformer with Contrastive Learning for Brain Connectivity Analysis Towards Improving Autism Detection. In Proc. IEEE Int. Symp. Biomed. Imaging (ISBI); IEEE: Houston, TX, USA, 2025; pp. 1–5. [Google Scholar] [CrossRef]
- MacWhinney, B. The CHILDES Project: Tools for Analyzing Talk, 3rd ed.; Lawrence Erlbaum Associates: Mahwah, NJ, USA, 2000; Available online: https://talkbank.org.
- Narain, J.; Johnson, K. T. ReCANVo: A Dataset of Real-World Communicative and Affective Nonverbal Vocalizations. Zenodo Dataset 2021. [Google Scholar] [CrossRef]

| Group | Total | Train/Val | Inference |
|---|---|---|---|
| SK-TD | 38 | 31 | 7 |
| SK-ADHD | 37 | 30 | 7 |
| SK-ASD (Dutch) | 46 | 39 | 7 |
| ReCANVo-ASD | 8 | - | 7 |
| Total | 129 | 100 | 28 |
| Hyperparameter | Value |
|---|---|
| Sampling Rate () | 16,000 Hz |
| Chunk Duration | 0.5 s |
| Hop Length | 0.1 s |
| Batch Size | 256 |
| Number of Epochs | 25 |
| Optimizer | Adam |
| Initial Learning Rate | |
| Temperature () | 0.07 |
| Random Seed | 42 |
| Hardware Acceleration | NVIDIA CUDA (Enabled) |
| Data Used in Training | LR | Accuracy | Precision | Recall | F1 |
|---|---|---|---|---|---|
| Dutch | 3E-04 | 99.62% | 100.00% | 100.00% | 100.00% |
| 82.14% | 76.47% | 92.85% | 83.87% * | ||
| Dutch | 1E-04 | 99.50% | 97.50% | 99.50% | 99.50% |
| 75.00% | 70.58% | 85.71% | 77.41% * | ||
| Dutch + ReCANVo | 1E-04 | 99.84% | 100.00% | 100.00% | 100.00% |
| 82.14% | 100.00% | 64.24% | 78.26% * | ||
| Dutch | 5E-04 | 97.30% | 98.55% | 98.55% | 98.55% |
| 64.28% | 59.09% | 92.85% | 72.22% * |
| Data used in Testing | Accuracy | Precision | Recall | F1 |
|---|---|---|---|---|
| ASD:ReCANVo + (TD,ADHD):Dutch | 100.00% | 100.00% | 100.00% | 100.00% |
| ASD:Dutch + (TD,ADHD):Dutch | 82.14% | 76.47% | 92.85% | 83.87% |
| ASD:(ReCANVo,Dutch) + (TD,ADHD):Dutch | 88.00% | 100.00% | 72.72% | 84.21% |
| Data used in testing | Accuracy | Precision | Recall | F1 |
|---|---|---|---|---|
| ASD:ReCANVo + (TD,ADHD):Dutch | 100.00% | 100.00% | 100.00% | 100.00% |
| ASD:Dutch + (TD,ADHD):Dutch | 78.57% | 78.57% | 78.57% | 78.57% |
| ASD:(ReCANVo,Dutch) + (TD,ADHD):Dutch | 82.14% | 76.47% | 92.85% | 83.87% |
| DUTCH in Training | Accuracy | Precision | Recall | F1 |
|---|---|---|---|---|
| ASD:ReCANVo + (TD,ADHD):Dutch | 67.71% | 50.00% | 6.45% | 11.43% |
| ASD:Dutch + (TD,ADHD):Dutch | 52.47% | 55.00% | 33.08% | 41.31% |
| ASD:(ReCANVo,Dutch) + (TD,ADHD):Dutch | 55.64% | 57.65% | 38.58% | 46.23% |
| MIXED in Training | Accuracy | Precision | Recall | F1 |
| ASD:ReCANVo + (TD,ADHD):Dutch | 92.71% | 86.36% | 91.94% | 89.06% |
| ASD:Dutch + (TD,ADHD):Dutch | 53.23% | 55.43% | 38.35% | 45.33% |
| ASD:(ReCANVo,Dutch) + (TD,ADHD):Dutch | 73.15% | 90.28% | 51.18% | 65.33% |
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2026 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license.




