Submitted:
09 April 2026
Posted:
15 April 2026
You are already at the latest version
Abstract
Keywords:
1. Introduction
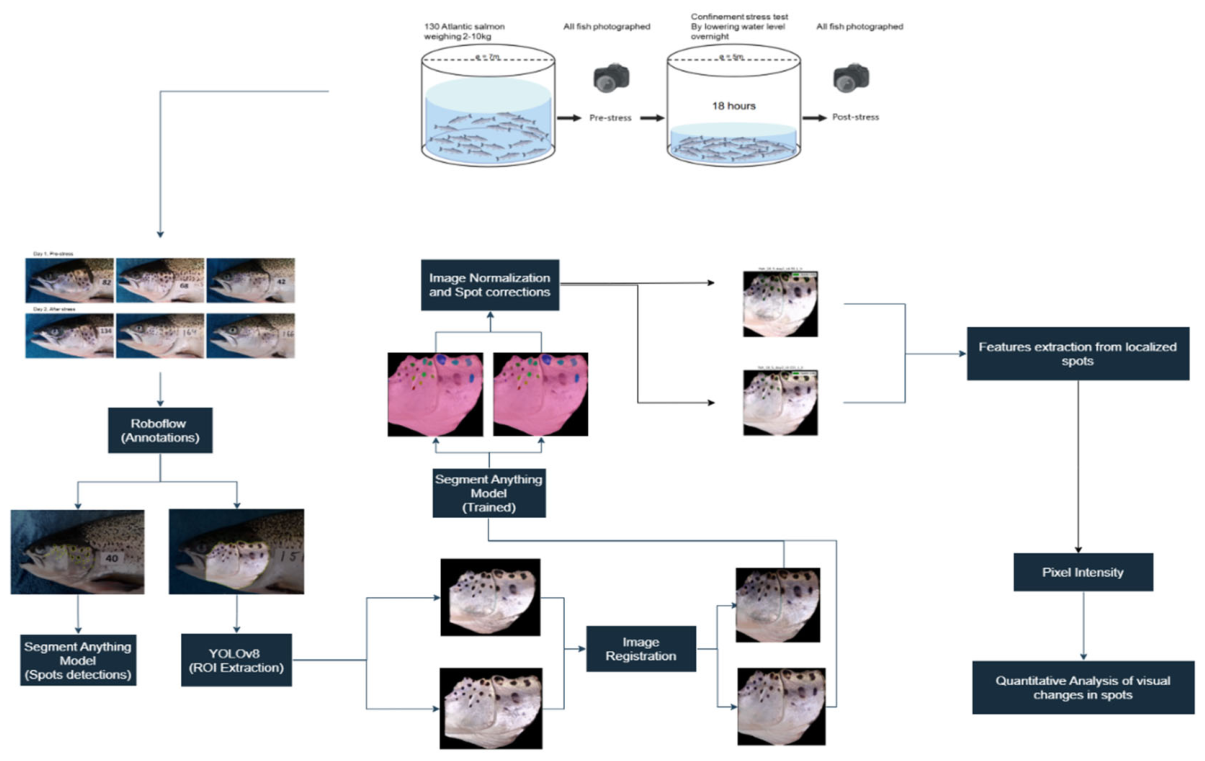
- We apply a computer vision-based methodology to quantify melanin-based skin spots on the operculum of Atlantic salmon (salmo salar).
- We propose a semi-automated methodological pipeline for operculum detection, spot segmentation, and grayscale-based quantification of temporal changes in spot appearance.
- We describe an imaging and annotation workflow that may support future studies of opercular pigmentation dynamics in salmon.
2. Related Works
2.1. Fish Disease Detection
2.2. Fish Detection and Monitoring
2.3. Fish Re-Identification
3. Materials and Methods
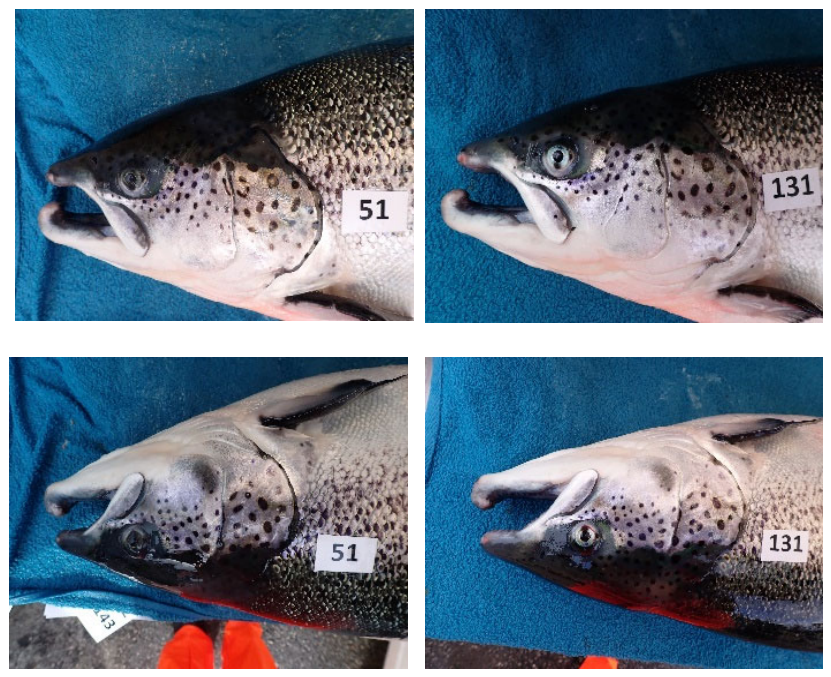
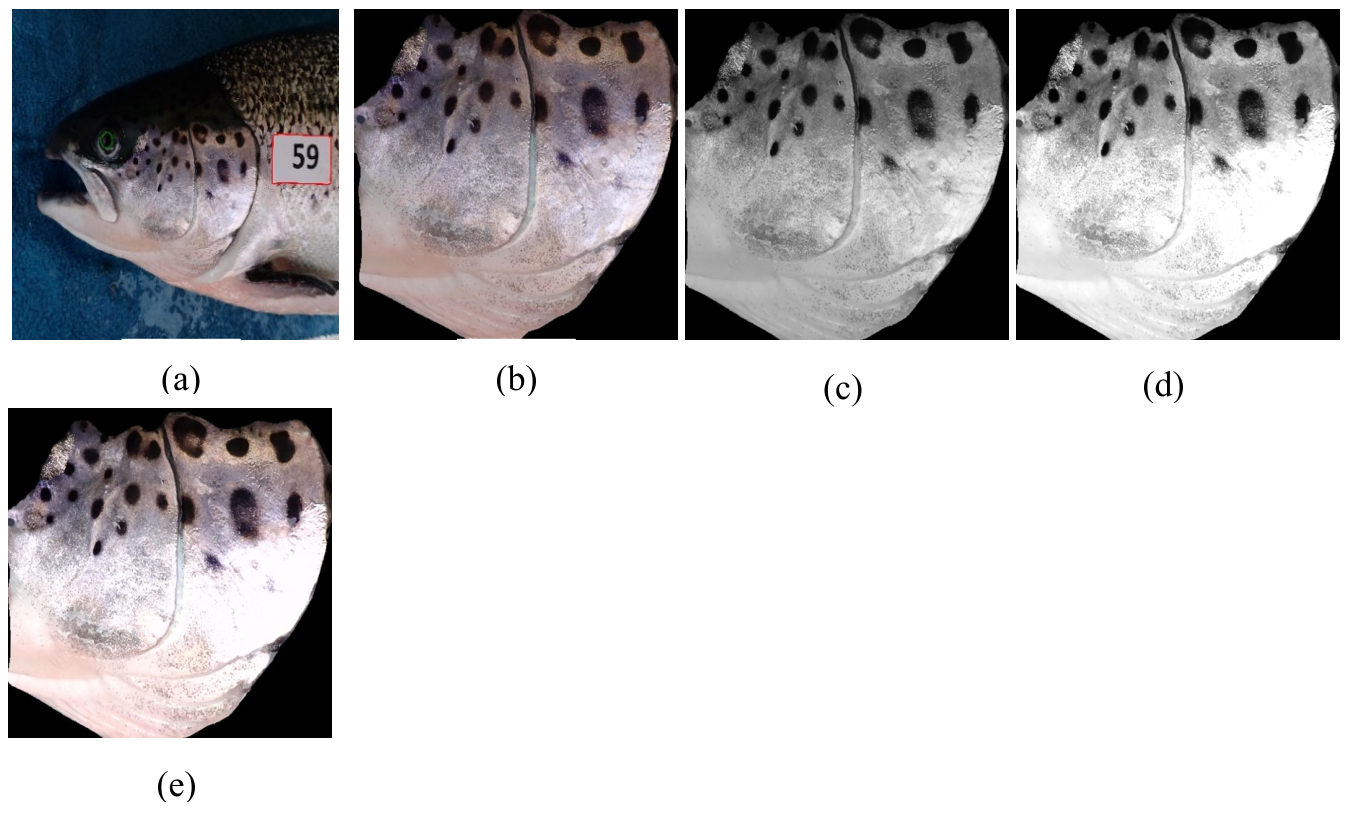
3.1. Experimental Setup and Dataset
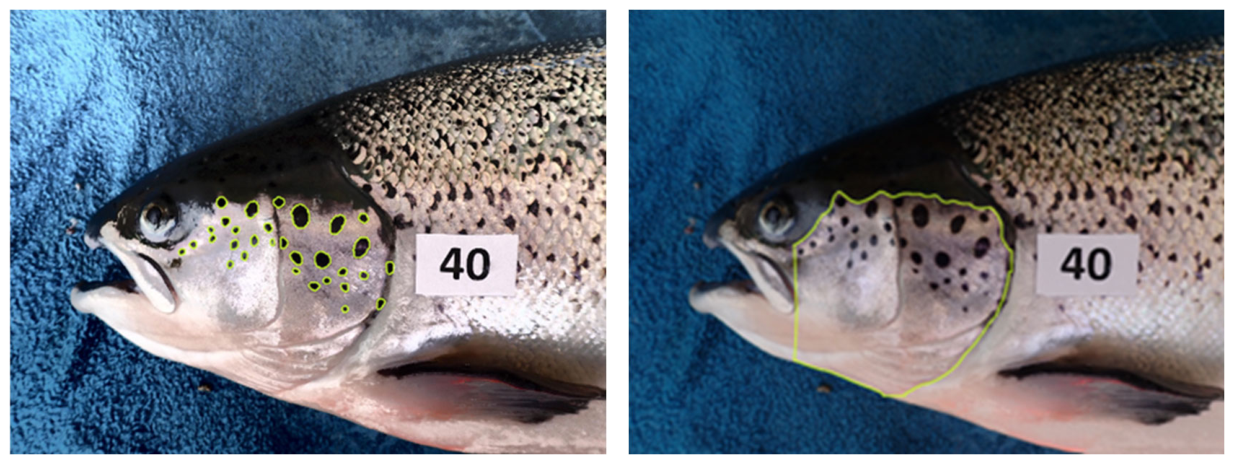
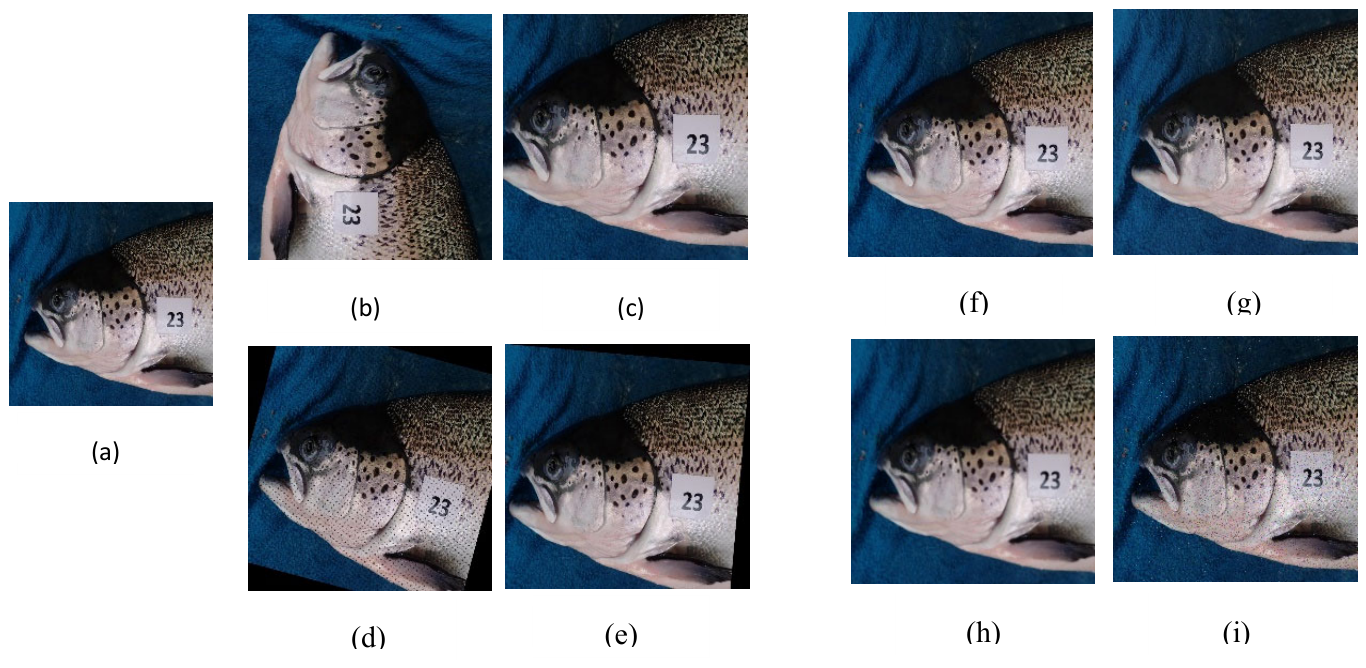
3.2. Image Annotations and Augmentations


3.2.1. Geometric Transformations
- Reflection: Clockwise, counterclockwise, upside-down reflections were generated with their respective transformation matrices using the following equation:
- Rotation: Images were randomly rotated using angles (ϴ) randomly sampled from a given range = [] using the following transformation equation:where is the rotation transformation matrix.
- Shear: Shearing was applied in both horizontal and vertical directions using angles (ϴ) randomly sampled from a given range= []
- Crops: Crops were generated with a randomly sampled zoom factor percentage (zf_perc) within a given range= [0, 10%] using the following transformation:where idx is the zoom factor index along which pixels in horizontal (w) and vertical (h) directions of Image (I) are sampled for the crop.
3.2.2. Pixel Intensity Transformations
- Exposure: Image exposure in both directions was randomly adjusted by randomly sampling a threshold from a given range = [-2%, 2%] using the following transformation equations:
- Brightness: Brightness adjustments were like exposure adjustments except instead of utilizing the LAB color space, the RGB images were converted to HSV color space (Busin et al., 2008), and the value channel of the converted images were adjusted.
- Blur and Noise: Blur was introduced using a gaussian filter (Bradski et al., 2000; Nelsonet al., 2020) while for noise salt-and-pepper noise (Bradski et al., 2000; Azzeh et al., 2018) was used.
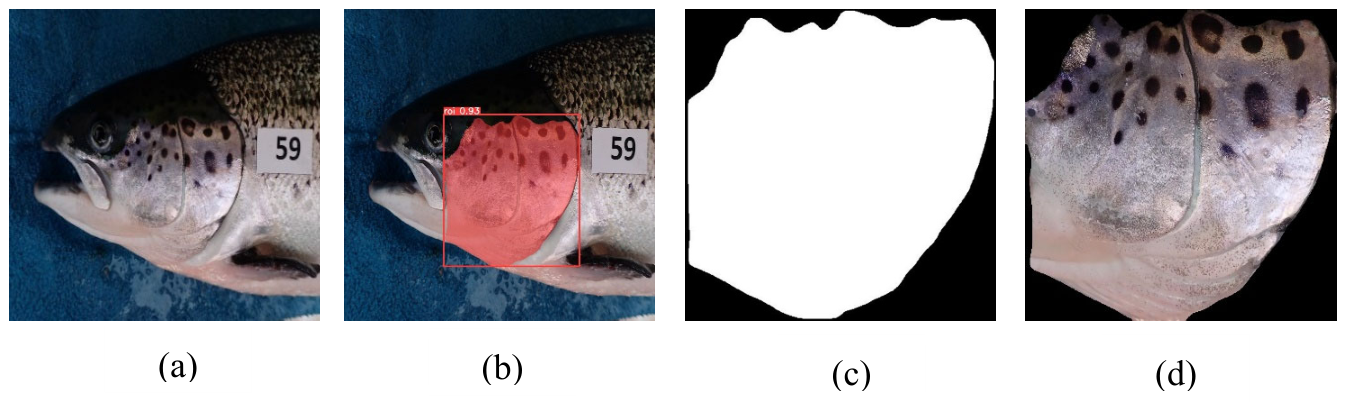
3.3. Detection and Segmentation of the Operculum Region and Spots
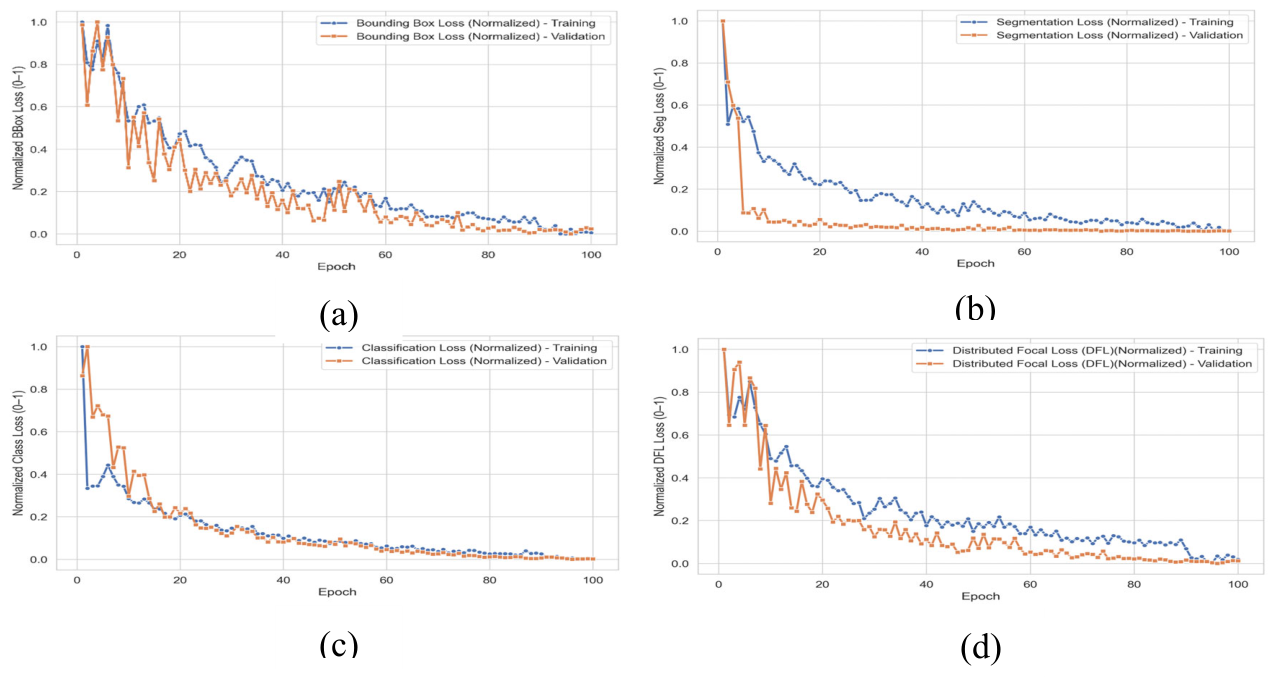
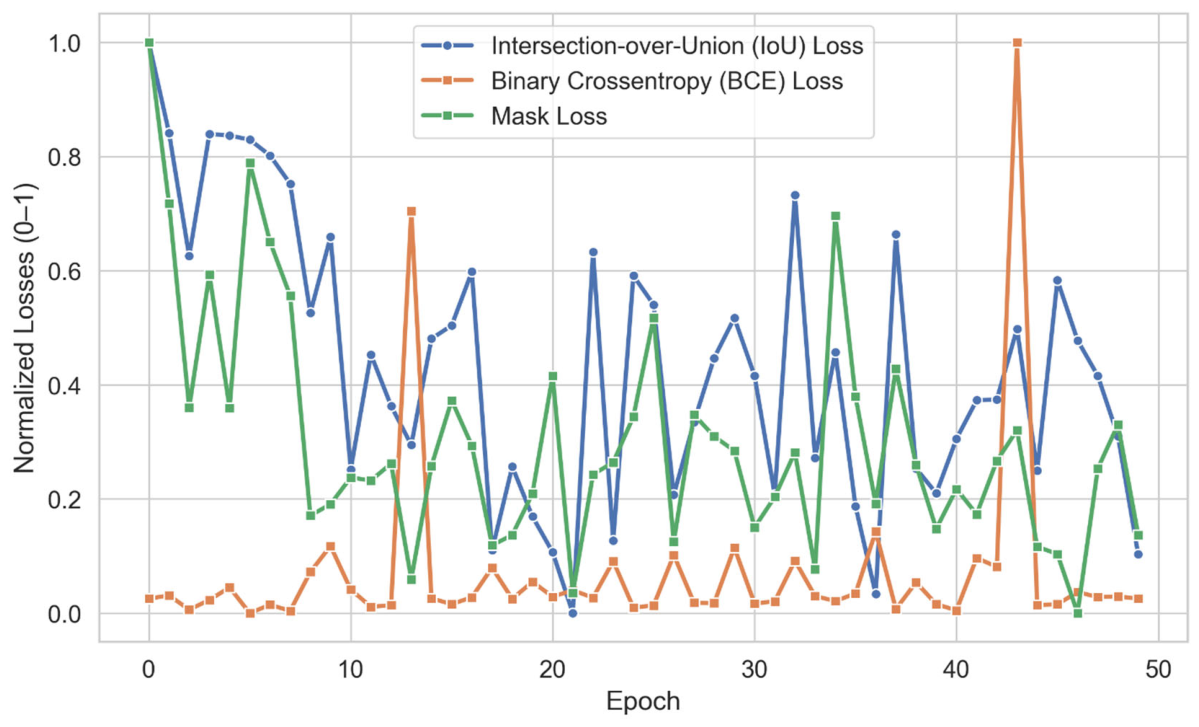
3.3.1. YOLOv8 and SAM2.1 Training
3.4. Inference
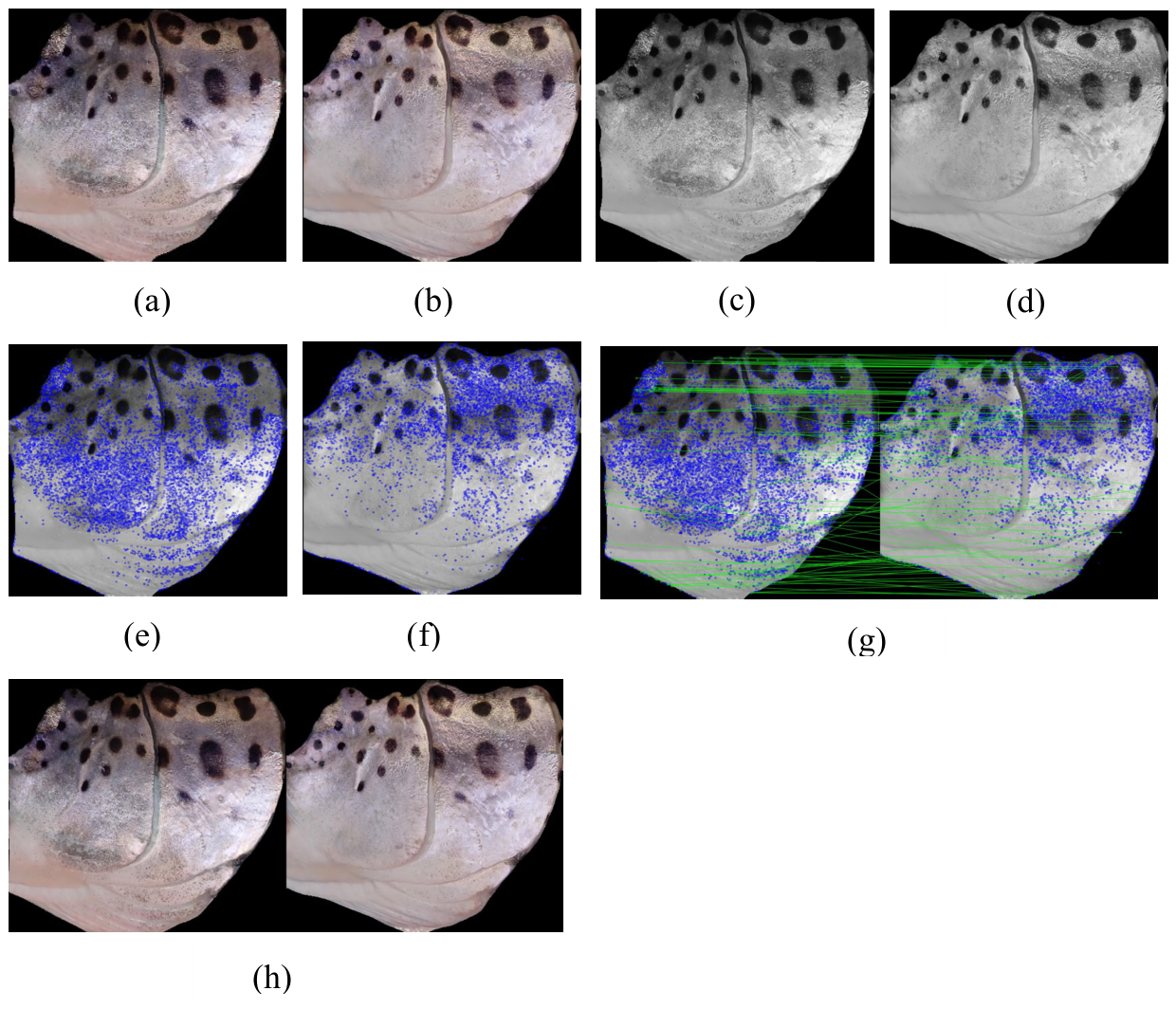
3.4.1. Operculum Regions Registration
3.4.2. Operculum Regions Normalization
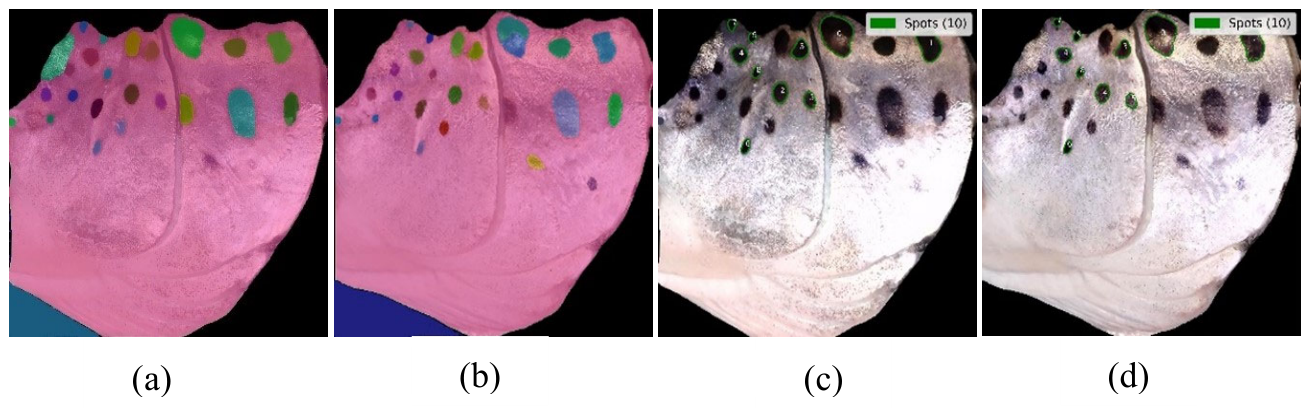
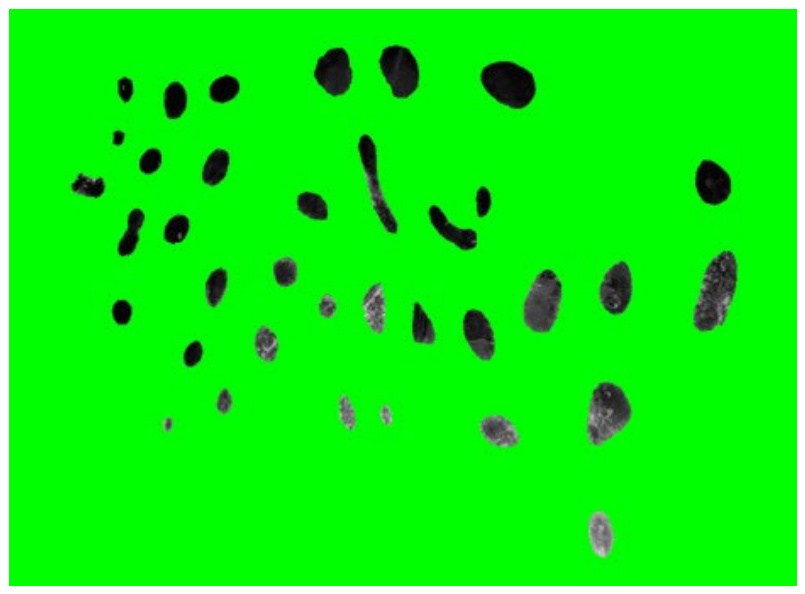
3.4.3. Operculum Region Spots Segmentation
3.4.4. Feature Extraction
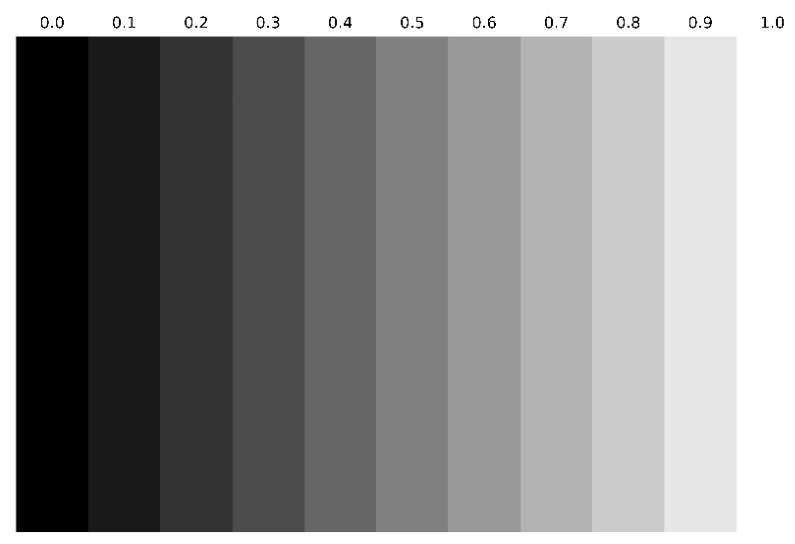
3.4.5. Statistical Analysis (Grayscale Intensity)
4. Results
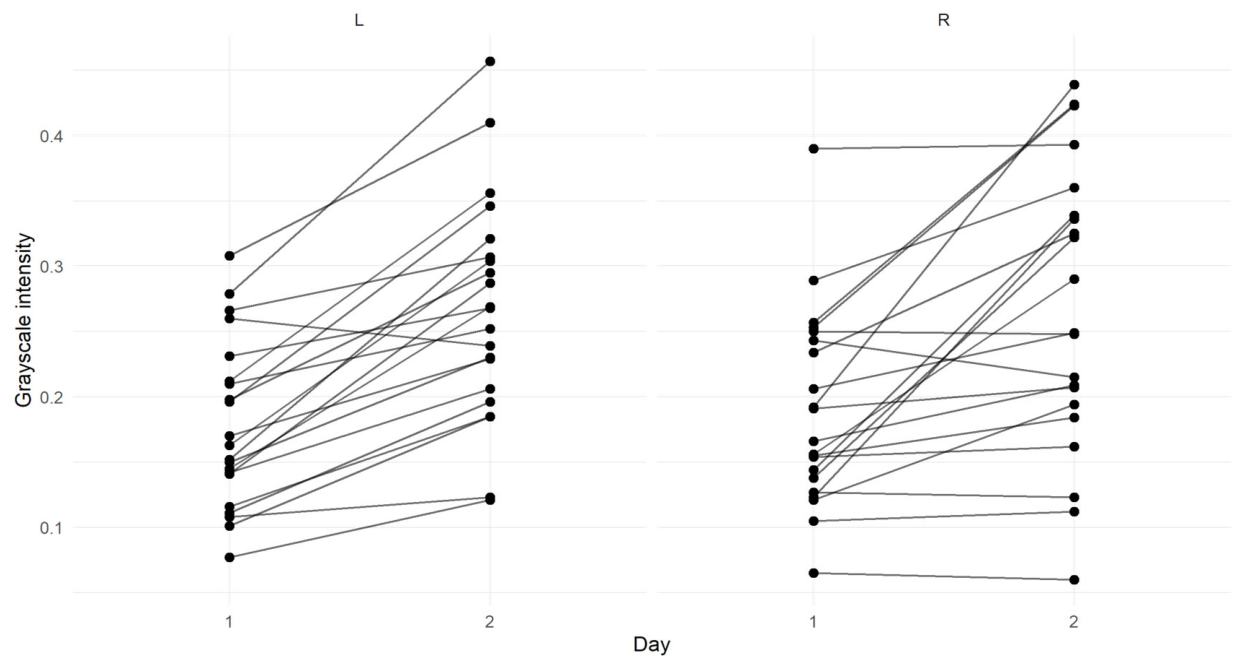
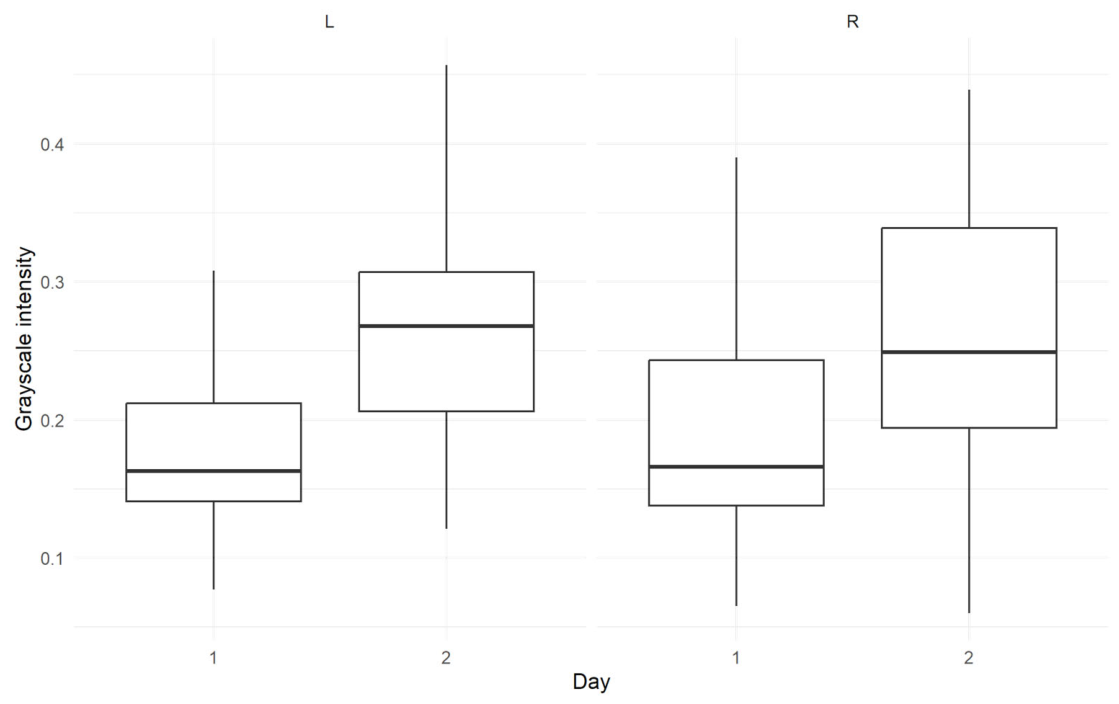
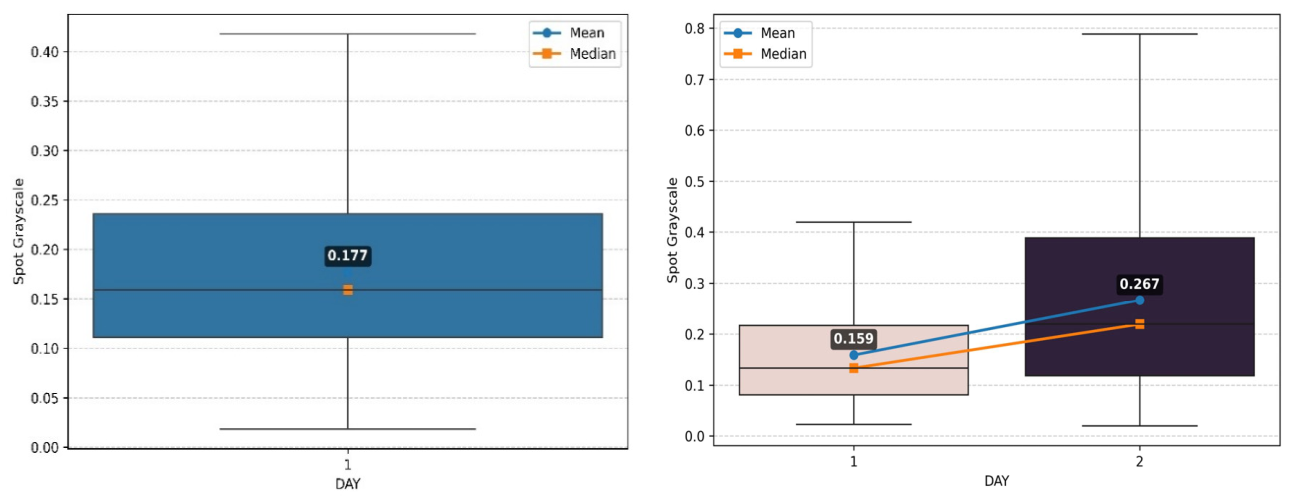
4.1. Effect of Treatment on the Spot Pixel Intensities
4.2. Grayscale Intensities of Treated and Control Groups
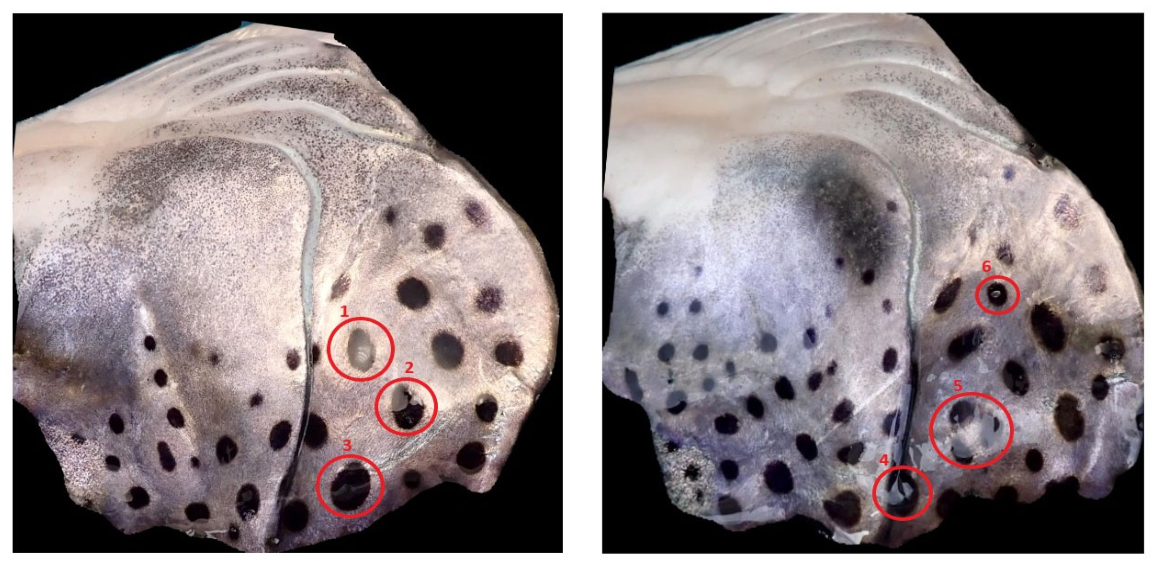
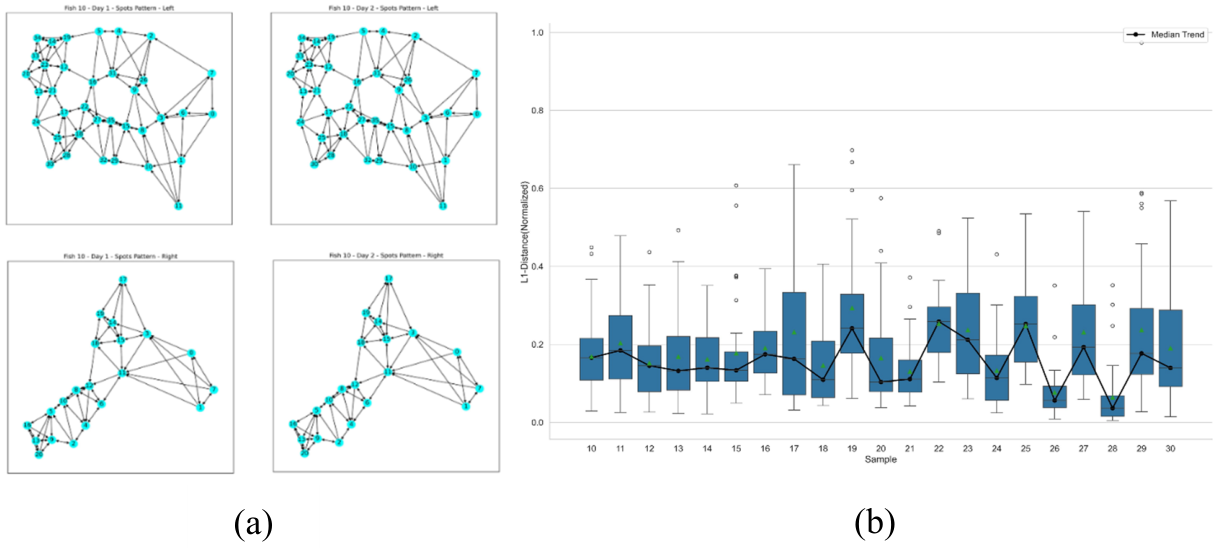
4.3. Neighborhood Based Grayscale Intensity Analysis
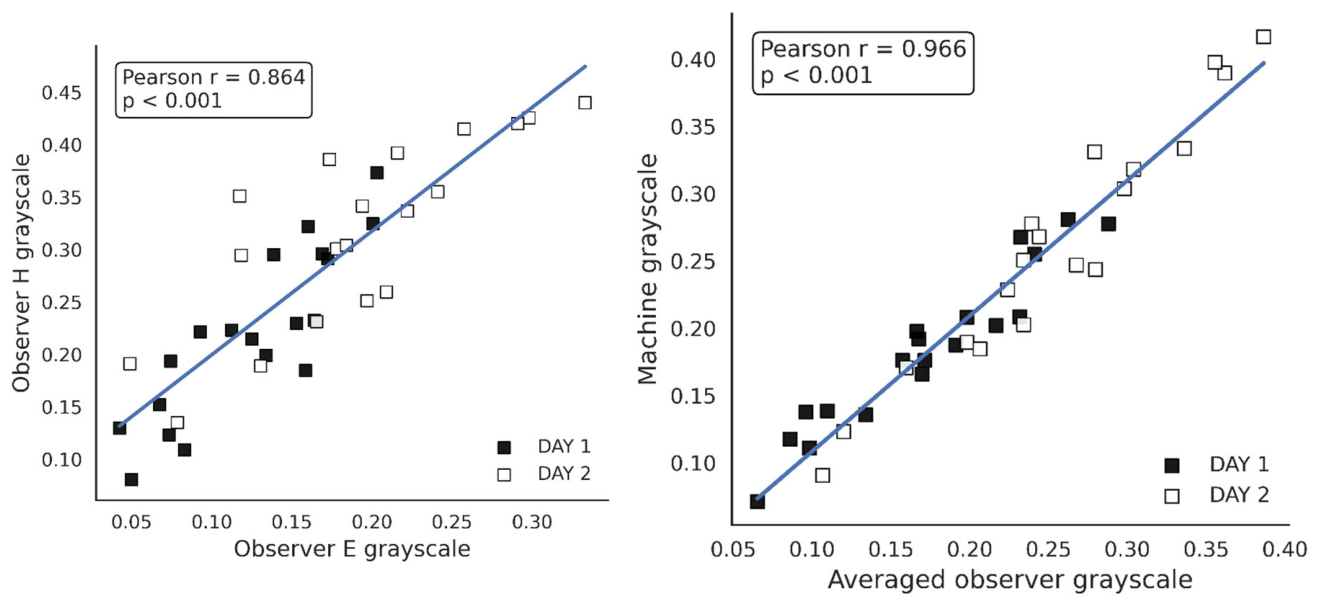
4.4. Manual Scoring of Spots on a Grayscale by Observers
| Fish Sample |
Observer (E) |
Observer (H) |
||
| Change left (%) |
Change right (%) | Change left (%) | Change right (%) | |
| 10 | 4.5 | 156.07 | 13.44 | 125.44 |
| 11 | 45.14 | 96.34 | 16.80 | 59.44 |
| 12 | 23.12 | 32.25 | -5.66 | 31.19 |
| 13 | 36.5 | 3.41 | 65.47 | -8.90 |
| 14 | 86.5 | 41.95 | 32.01 | 37.03 |
| 15 | 11.37 | 5.22 | 41.66 | 0.87 |
| 16 | 12.8 | 43.08 | 18.32 | 4.26 |
| 17 | 59.5 | 51.11 | 76 | 67.64 |
| 18 | 118.7 | 132.83 | 75 | 104.71 |
| 19 | 76.9 | 0 | 127.27 | 0 |
| 20 | 66.36 | 14.47 | 37.85 | 0 |
| 21 | 65.16 | 91.30 | 133.54 | 128.08 |
| 22 | 41.24 | 164.72 | 23.32 | 97.76 |
| 23 | N/A | N/A | N/A | N/A |
| 24 | 57.47 | 61.29 | 25.31 | 94.66 |
| 25 | N/A | N/A | N/A | N/A |
| 26 | 40.84 | 96.55 | 65.76 | 72 |
| 27 | 86.82 | 235.08 | 56.52 | 110 |
| 28 | 36.58 | -4.54 | 36.60 | 63.20 |
| 29 | 59.01 | 0 | 28.04 | 0 |
| 30 | 113 | 16.66 | 133.91 | 9.68 |
| Average | 54.83 | 65.15 | 52.69 | 52.52 |
5. Discussion and Future Research
6. Conclusion
Funding
Data Availability Statement
Acknowledgments
Ethics Statement
Conflicts of Interest
References
- Sommerset, I., Wiik-Nielsen, J., Moldal, T., Oliveira, V. H. S., Svendsen, J. C., Haukaas, A., & Brun, E. (2024). Fish health report 2023. Norwegian Veterinary Institute: Ås, Norway.
- Tvete, I.F.; Aldrin, M.; Jensen, B.B. Towards better survival: Modeling drivers for daily mortality in Norwegian Atlantic salmon farming. Prev. Veter- Med. 2022, 210, 105798. [CrossRef]
- Overton, K.; Dempster, T.; Oppedal, F.; Kristiansen, T.S.; Gismervik, K.; Stien, L.H. Salmon lice treatments and salmon mortality in Norwegian aquaculture: a review. Rev. Aquac. 2018, 11, 1398–1417. [CrossRef]
- Department of Energy, "Sustainable manufacturing and circular economy," Tech. Rep., 3 2023. https://www. energy.gov/eere/articles/sustainable.
- Stien, L.H.; Tørud, B.; Gismervik, K.; Lien, M.E.; Medaas, C.; Osmundsen, T.; Kristiansen, T.S.; Størkersen, K.V. Governing the welfare of Norwegian farmed salmon: Three conflict cases. Mar. Policy 2020, 117. [CrossRef]
- Keihani, R.; Gomes, A.S.; Balseiro, P.; Handeland, S.O.; Gorissen, M.; Arukwe, A. Evaluation of stress in farmed Atlantic salmon (Salmo salar) using different biological matrices. Comp. Biochem. Physiol. Part A: Mol. Integr. Physiol. 2024, 298, 111743. [CrossRef]
- E Adams, C.; Turnbull, J.F.; Bell, A.; E Bron, J.; A Huntingford, F. Multiple determinants of welfare in farmed fish: stocking density, disturbance, and aggression in Atlantic salmon (Salmo salar). Can. J. Fish. Aquat. Sci. 2007, 64, 336–344. [CrossRef]
- Volpato, G.L.; Gonçalves-De-Freitas, E.; Fernandes-De-Castilho, M. Insights into the concept of fish welfare. Dis. Aquat. Org. 2007, 75, 165–171. [CrossRef]
- Cao, Y.; Tveten, A.-K.; Stene, A. Establishment of a non-invasive method for stress evaluation in farmed salmon based on direct fecal corticoid metabolites measurement. Fish Shellfish. Immunol. 2017, 66, 317–324. [CrossRef]
- Leclercq, E.; Taylor, J.F.; Migaud, H. Morphological skin colour changes in teleosts. Fish Fish. 2010, 11, 159–193. [CrossRef]
- Césarini, J.; I.N.S.E.R.M Melanins and their possible roles through biological evolution. Adv. Space Res. 1996, 18, 35–40. [CrossRef]
- Riley, P. A. (1997). Melanin. The international journal of biochemistry & cell biology, 29(11), 1235-1239.
- A Mackintosh, J. The Antimicrobial Properties of Melanocytes, Melanosomes and Melanin and the Evolution of Black Skin. J. Theor. Biol. 2001, 211, 101–113. [CrossRef]
- Roulin, A. The evolution, maintenance and adaptive function of genetic colour polymorphism in birds. Biol. Rev. 2004, 79, 815–848. [CrossRef]
- E Hoekstra, H. Genetics, development and evolution of adaptive pigmentation in vertebrates. Heredity 2006, 97, 222–234. [CrossRef]
- Kittilsen, S.; Schjolden, J.; Beitnes-Johansen, I.; Shaw, J.; Pottinger, T.; Sørensen, C.; Braastad, B.; Bakken, M.; Øverli, Ø. Melanin-based skin spots reflect stress responsiveness in salmonid fish. Horm. Behav. 2009, 56, 292–298. [CrossRef]
- Kittilsen, S.; Johansen, I.B.; Braastad, B.O.; Øverli, Ø. Pigments, Parasites and Personalitiy: Towards a Unifying Role for Steroid Hormones?. PLOS ONE 2012, 7, e34281. [CrossRef]
- Khan, U.W.; Øverli, Ø.; Hinkle, P.M.; Pasha, F.A.; Johansen, I.B.; Berget, I.; Silva, P.I.M.; Kittilsen, S.; Höglund, E.; Omholt, S.W.; et al. A novel role for pigment genes in the stress response in rainbow trout (Oncorhynchus mykiss). Sci. Rep. 2016, 6, 28969. [CrossRef]
- Yi, M.; Lu, H.; Du, Y.; Sun, G.; Shi, C.; Li, X.; Tian, H.; Liu, Y. The color change and stress response of Atlantic salmon (Salmo salar L.) infected with Aeromonas salmonicida. Aquac. Rep. 2021, 20. [CrossRef]
- Milinski, M. (1990). Parasites and host decision-making.
- Höglund, E.; Balm, P.H.M.; Winberg, S. Skin Darkening, A Potential Social Signal in Subordinate Arctic Charr (Salvelinus Alpinus): The Regulatory Role of Brain Monoamines and Pro-Opiomelanocortin-Derived Peptides. J. Exp. Biol. 2000, 203, 1711–1721. [CrossRef]
- Maan, M.E.; Seehausen, O.; Söderberg, L.; Johnson, L.; Ripmeester, E.A.P.; Mrosso, H.D.J.; Taylor, M.I.; van Dooren, T.J.M.; van Alphen, J.J.M. Intraspecific sexual selection on a speciation trait, male coloration, in the Lake Victoria cichlidPundamilia nyererei. Proc. R. Soc. B: Biol. Sci. 2004, 271, 2445–2452. [CrossRef]
- Yasir, I.; Qin, J.G. Impact of Background on Color Performance of False Clownfish, Amphiprion ocellaris, Cuvier. J. World Aquac. Soc. 2009, 40, 724–734. [CrossRef]
- Logan, D. W., Burn, S. F., & Jackson, I. J. (2006). Regulation of pigmentation in zebrafish.
- melanophores. Pigment cell research, 19(3), 206-213.
- Mills, M.G.; Patterson, L.B. Not just black and white: Pigment pattern development and evolution in vertebrates. Semin. Cell Dev. Biol. 2009, 20, 72–81. [CrossRef]
- Ludwig, D.S.; Mountjoy, K.G.; Tatro, J.B.; Gillette, J.A.; Frederich, R.C.; Flier, J.S.; Maratos-Flier, E. Melanin-concentrating hormone: a functional melanocortin antagonist in the hypothalamus. Am. J. Physiol. Metab. 1998, 274, E627–E633. [CrossRef]
- Willard, D.H.; Bodnar, W.; Harris, C.; Kiefer, L.; Nichols, J.S.; Blanchard, S.; Hoffman, C.; Moyer, M.; Burkhart, W. Agouti structure and function: characterization of a potent.alpha.-melanocyte stimulating hormone receptor antagonist. Biochemistry 1995, 34, 12341–12346. [CrossRef]
- Kawauchi, H. Functions of melanin-concentrating hormone in fish. J. Exp. Zoöl. Part A: Comp. Exp. Biol. 2006, 305A, 751–760. [CrossRef]
- Gröneveld, D.; Balm, P.H.; Bonga, S.E.W. Biphasic effect of MCH on α-MSH release from the tilapia (Oreochromis mossambicus) pituitary. Peptides 1995, 16, 945–949. [CrossRef]
- Shimomura, Y.; Mori, M.; Sugo, T.; Ishibashi, Y.; Abe, M.; Kurokawa, T.; Onda, H.; Nishimura, O.; Sumino, Y.; Fujino, M. Isolation and Identification of Melanin-Concentrating Hormone as the Endogenous Ligand of the SLC-1 Receptor. Biochem. Biophys. Res. Commun. 1999, 261, 622–626. [CrossRef]
- Lu, D.; Willard, D.; Patel, I.R.; Kadwell, S.; Overton, L.; Kost, T.; Luther, M.; Chen, W.; Woychik, R.P.; Wilkison, W.O.; et al. Agouti protein is an antagonist of the melanocyte-stimulating-hormone receptor. Nature 1994, 371, 799–802. [CrossRef]
- Cerdá-Reverter, J.M.; Agulleiro, M.J.; R, R.G.; Sánchez, E.; Ceinos, R.; Rotllant, J. Fish melanocortin system. Eur. J. Pharmacol. 2011, 660, 53–60. [CrossRef]
- Cerdá-Reverter, J.M.; Haitina, T.; Schiöth, H.B.; Peter, R.E. Gene Structure of the Goldfish Agouti-Signaling Protein: A Putative Role in the Dorsal-Ventral Pigment Pattern of Fish. Endocrinology 2005, 146, 1597–1610. [CrossRef]
- Chaki, J.; Dey, N. A Beginner's Guide to Image Preprocessing Techniques; Taylor & Francis: London, United Kingdom, 2018; ISBN:.
- Mohanaiah, P., Sathyanarayana, P., & GuruKumar, L. (2013). Image texture feature extraction using GLCM approach. International journal of scientific and research publications, 3(5), 1-5.
- Meena, K.S.; Suriya, S. A Survey on Supervised and Unsupervised Learning Techniques. International Conference on Artificial Intelligence, Smart Grid and Smart City Applications. Cham: Springer International Publishing; pp. 627–644.
- Ahmed, M. S., Aurpa, T. T., & Azad, M. A. K. (2022). Fish disease detection using image based machine learning technique in aquaculture. Journal of King Saud University-Computer and Information Sciences, 34(8), 5170-5182.
- Shetty, A. K., Saha, I., Sanghvi, R. M., Save, S. A., & Patel, Y. J. (2021, April). A review: Object detection models. In 2021 6th International Conference for Convergence in Technology (I2CT) (pp. 1-8). IEEE.
- Zhang, C.; Bracke, M.; Torres, R.d.S.; Gansel, L.C. Rapid detection of salmon louse larvae in seawater based on machine learning. Aquaculture 2024, 592. [CrossRef]
- Wu, J. (2017). Introduction to convolutional neural networks. National Key Lab for Novel Software Technology. Nanjing University. China, 5(23), 495.
- Simonyan, K., & Zisserman, A. (2014). Very deep convolutional networks for large-scale image recognition. arXiv preprint arXiv:1409.1556.
- Gupta, A.; Bringsdal, E.; Knausgård, K.M.; Goodwin, M. Accurate Wound and Lice Detection in Atlantic Salmon Fish Using a Convolutional Neural Network. Fishes 2022, 7, 345. [CrossRef]
- Liang, X.; Hu, P.; Zhang, L.; Sun, J.; Yin, G. MCFNet: Multi-Layer Concatenation Fusion Network for Medical Images Fusion. IEEE Sensors J. 2019, 19, 7107–7119. [CrossRef]
- Vaswani, A., Shazeer, N., Parmar, N., Uszkoreit, J., Jones, L., Gomez, A. N.,... & Polosukhin, I. (2017). Attention is all you need. Advances in neural information processing systems, 30.
- Huang, G. B., Liang, N. Y., Rong, H. J., Saratchandran, P., & Sundararajan, N. (2005). On-line sequential extreme learning machine. Computational Intelligence, 2005, 232-237.
- Huang, Y.-P.; Khabusi, S.P. A CNN-OSELM Multi-Layer Fusion Network With Attention Mechanism for Fish Disease Recognition in Aquaculture. IEEE Access 2023, 11, 58729–58744. [CrossRef]
- Bochkovskiy, A., Wang, C. Y., & Liao, H. Y. M. (2020). Yolov4: Optimal speed and accuracy of object detection. arXiv preprint arXiv:2004.10934.
- Banno, K.; Kaland, H.; Crescitelli, A.; Tuene, S.; Aas, G.; Gansel, L. A novel approach for wild fish monitoring at aquaculture sites: wild fish presence analysis using computer vision. Aquac. Environ. Interactions 2022, 14, 97–112. [CrossRef]
- Mogdans, J.; Bleckmann, H. Coping with flow: behavior, neurophysiology and modeling of the fish lateral line system. Biol. Cybern. 2012, 106, 627–642. [CrossRef]
- Jocher, Glenn, et al. (2020). “ultralytics/yolov5: v3. 0.” Zenodo.
- Khanam, R., & Hussain, M. (2024). What is YOLOv5: A deep look into the internal features of the popular object detector. arXiv preprint arXiv:2407.20892.
- Yu, H.; Wang, Z.; Qin, H.; Chen, Y. An Automatic Detection and Counting Method for Fish Lateral Line Scales of Underwater Fish Based on Improved YOLOv5. IEEE Access 2023, 11, 143616–143627. [CrossRef]
- Ellis, T.; Berrill, I.; Lines, J.; Turnbull, J.F.; Knowles, T.G. Mortality and fish welfare. Fish Physiol. Biochem. 2011, 38, 189–199. [CrossRef]
- Cao, K.; Liu, Y.; Meng, G.; Sun, Q. An Overview on Edge Computing Research. IEEE Access 2020, 8, 85714–85728. [CrossRef]
- Li, S., Xu, L. D., & Zhao, S. (2015). The internet of things: a survey. Information systems frontiers, 17(2), 243-259.
- Wang, C.-Y.; Bochkovskiy, A.; Liao, H.-Y.M. YOLOv7: Trainable Bag-of-Freebies Sets New State-of-the-Art for Real-Time Object Detectors. 2023 IEEE/CVF Conference on Computer Vision and Pattern Recognition (CVPR). Vancouver, BC, Canada, 17-24 June 2023; pp. 7464–7475.
- Ranjan, R.; Sharrer, K.; Tsukuda, S.; Good, C. MortCam: An Artificial Intelligence-aided fish mortality detection and alert system for recirculating aquaculture. Aquac. Eng. 2023, 102. [CrossRef]
- Farnoush, R., & ZAR, P. B. (2008). Image segmentation using Gaussian mixture model.
- He, K., Gkioxari, G., Dollár, P., & Girshick, R. (2017). Mask r-cnn. In Proceedings of the IEEE international conference on computer vision (pp. 2961-2969).
- Ding, S., Zhang, J., Xu, X., & Zhang, Y. (2016). A wavelet extreme learning machine. Neural Computing and Applications, 27(4), 1033-1040.
- Al Duhayyim, M.; Alshahrani, H.M.; Al-Wesabi, F.N.; Alamgeer, M.; Hilal, A.M.; Hamza, M.A. Intelligent Deep Learning Based Automated Fish Detection Model for UWSN. Comput. Mater. Contin. 2022, 70, 5871–5887. [CrossRef]
- O’shea, K., & Nash, R. (2015). An introduction to convolutional neural networks. arXiv preprint arXiv:1511.08458.
- Lowe, D.G. Distinctive Image Features from Scale-Invariant Keypoints. Int. J. Comput. Vis. 2004, 60, 91–110. [CrossRef]
- Cisar, P.; Bekkozhayeva, D.; Movchan, O.; Saberioon, M.; Schraml, R. Computer vision based individual fish identification using skin dot pattern. Sci. Rep. 2021, 11, 1–12. [CrossRef]
- Bekkozhayeva, D.; Cisar, P. Image-Based Automatic Individual Identification of Fish without Obvious Patterns on the Body (Scale Pattern). Appl. Sci. 2022, 12, 5401. [CrossRef]
- Ren, S.; He, K.; Girshick, R.; Sun, J. Faster R-CNN: Towards real-time object detection with region proposal networks. IEEE Trans. Pattern Anal. Mach. Intell. 2017, 39, 1137–1149. [CrossRef]
- Lin, T.-Y., Maire, M., Belongie, S., Hays, J., Perona, P., Ramanan, D., Dollár, P., Zitnick, C.L. (2014). Microsoft Coco: Common Objects in Context. In Proceedings of the Computer Vision–ECCV 2014: 13th European Conference, Zurich, Switzerland, September 6-12, 2014; pp. 740–755. [CrossRef]
- Kuznetsova, A., Rom, H., Alldrin, N., Uijlings, J., Krasin, I., Pont-Tuset, J.,... & Ferrari, V. (2020). The open images dataset v4: Unified image classification, object detection, and visual relationship detection at scale. International journal of computer vision, 128(7), 1956-1981.
- He, K.; Zhang, X.; Ren, S.; Sun, J. Deep Residual Learning for Image Recognition. In Proceedings of the IEEE on compute Rvision and pattern recognition, 2015, abs/1512.03385.
- Levy, A.; Shalom, B.R.; Chalamish, M. A guide to similarity measures and their data science applications. J. Big Data 2025, 12, 1–57. [CrossRef]
- Zhou, Z.; Hitt, N.P.; Letcher, B.H.; Shi, W.; Li, S. Pigmentation-based Visual Learning for Salvelinus fontinalis Individual Re-identification. 2022 IEEE International Conference on Big Data (Big Data); pp. 6850–6852.
- Chen, T., Kornblith, S., Norouzi, M., & Hinton, G. (2020, November). A simple framework for contrastive learning of visual representations. In International conference on machine learning (pp. 1597-1607). PmLR.
- Gehring, J.; Auli, M.; Grangier, D.; Dauphin, Y. A Convolutional Encoder Model for Neural Machine Translation. In Proceedings of the 55th Annual Meeting of the Association for Computational Linguistics, Berlin, Germany, 7–12 August 2016; pp. 123–135. [CrossRef]
- Shi, W.; Zhou, Z.; Letcher, B.H.; Hitt, N.; Kanno, Y.; Futamura, R.; Kishida, O.; Morita, K.; Li, S. Aging Contrast: A Contrastive Learning Framework for Fish Re-identification Across Seasons and Years. Australasian Joint Conference on Artificial Intelligence. Singapore: Springer Nature Singapore; pp. 252–264.
- Dwyer, B., Nelson, J., Hansen, T., et al. (2025). Roboflow (Version 1.0) [Software]. Available from https://roboflow.com. Computer vision.
- Hafiz, A.M.; Bhat, G.M. A survey on instance segmentation: state of the art. Int. J. Multimedia Inf. Retr. 2020, 9, 171–189. [CrossRef]
- Kirillov, A., Mintun, E., Ravi, N., Mao, H., Rolland, C., Gustafson, L.,... & Girshick, R. (2023). Segment anything. In Proceedings of the IEEE/CVF international conference on computer vision (pp. 4015-4026).
- Xu, M.; Yoon, S.; Fuentes, A.; Park, D.S. A Comprehensive Survey of Image Augmentation Techniques for Deep Learning. Pattern Recognit. 2023, 137. [CrossRef]
- OpenCV. Open Source Computer Vision Library, 2015. Available online: https://opencv.org (accessed on 14 December 2022).
- Busin, L., Vandenbroucke, N., & Macaire, L. (2008). Color spaces and image segmentation. Advances in imaging and electron physics, 151(1), 1.
- Nelson, J. (2020). The Importance of Blur as an Image Augmentation Technique. Roboflow Blog: https://blog.roboflow.com/using-blurin- computer-vision-preprocessing/.
- Azzeh, J.; Zahran, B.; Alqadi, Z. Salt and Pepper Noise: Effects and Removal. JOIV : Int. J. Informatics Vis. 2018, 2, 252–256. [CrossRef]
- Goodfellow, I. J., Shlens, J., & Szegedy, C. (2014). Explaining and harnessing adversarial examples. arXiv preprint arXiv:1412.6572.
- Dwyer, B., Nelson, J., Hansen, T., et al. (2025). Image Augmentation. Roboflow Documentation. https://docs.roboflow.com/datasets/datasetversions/ image-augmentation/.
- Ioffe, S., & Szegedy, C. (2015, June). Batch normalization: Accelerating deep network training by reducing internal covariate shift. In International conference on machine learning (pp. 448-456). pmlr.
- Ramachandran, P., Zoph, B., & Le, Q. V. (2017). Swish: a self-gated activation function. arXiv preprint arXiv:1710.05941, 7(1), 5.
- Zeiler, M. D., Krishnan, D., Taylor, G. W., & Fergus, R. (2010, June). Deconvolutional networks. In 2010 IEEE Computer Society Conference on computer vision and pattern recognition (pp. 2528-2535). IEEE.
- Roodschild, M.; Sardiñas, J.G.; Will, A. A new approach for the vanishing gradient problem on sigmoid activation. Prog. Artif. Intell. 2020, 9, 351–360. [CrossRef]
- He, K.; Zhang, X.; Ren, S.; Sun, J. Spatial Pyramid Pooling in Deep Convolutional Networks for Visual Recognition. IEEE Trans. Pattern Anal. Mach. Intell. 2015, 37, 1904–1916. [CrossRef]
- Ren, J., Bi, Z., Niu, Q., Liu, J., Peng, B., Zhang, S.,... & Liu, M. (2024). Deep Learning and Machine Learning--Object Detection and Semantic Segmentation: From Theory to Applications. arXiv preprint arXiv:2410.15584.
- Ravi, N., Gabeur, V., Hu, Y. T., Hu, R., Ryali, C., Ma, T.,... & Feichtenhofer, C. (2024). Sam 2: Segment anything in images and videos. arXiv preprint arXiv:2408.00714.
- Ryali, C., Hu, Y. T., Bolya, D., Wei, C., Fan, H., Huang, P. Y.,... & Feichtenhofer, C. (2023, July).Hiera: A hierarchical vision transformer without the bells-and-whistles. In International conference on machine learning (pp. 29441-29454). PMLR.
- Dosovitskiy, A., Beyer, L., Kolesnikov, A., Weissenborn, D., Zhai, X., Unterthiner, T.,... & Houlsby, N. (2020). An image is worth 16x16 words: Transformers for image recognition at scale. arXiv preprint arXiv:2010.11929.
- Lin, T.Y.; Dollár, P.; Girshick, R.; He, K.; Hariharan, B.; Belongie, S. Feature pyramid networks for object detection. In Proceedings of the IEEE Conference on Computer Vision and Pattern Recognition, Honolulu, HI, USA, 21–26 July 2017. [CrossRef]
- Turner, R. E. (2023). An introduction to transformers. arXiv preprint arXiv:2304.10557.
- Shaw, P.; Uszkoreit, J.; Vaswani, A. Self-Attention with Relative Position Representations. Assoc. Comput. Linguist. 2018, 2, 464–468. [CrossRef]
- Gheini, M.; Ren, X.; May, J. Cross-Attention is All You Need: Adapting Pretrained Transformers for Machine Translation. arXiv preprint arXiv:2104.08771.
- Li, Q.; Yan, M.; Xu, J. Optimizing Convolutional Neural Network Performance by Mitigating Underfitting and Overfitting. 2021 IEEE/ACIS 19th International Conference on Computer and Information Science (ICIS). IEEE; pp. 126–131.
- Adam, K. D. B. J. (2014). A method for stochastic optimization. arXiv preprint arXiv:1412.6980, 1412(6).
- Lin, T.-Y., Maire, M., Belongie, S., Hays, J., Perona, P., Ramanan, D., Dollár, P., Zitnick, C.L. (2014). Microsoft Coco: Common Objects in Context. In Proceedings of the Computer Vision–ECCV 2014: 13th European Conference, Zurich, Switzerland, September 6-12, 2014; pp. 740–755. [CrossRef]
- Buslaev, A.; Iglovikov, V.I.; Khvedchenya, E.; Parinov, A.; Druzhinin, M.; Kalinin, A.A. Albumentations: Fast and Flexible Image Augmentations. Information 2020, 11, 125. [CrossRef]
- Padilla, R.; Passos, W.L.; Dias, T.L.B.; Netto, S.L.; da Silva, E.A.B. A Comparative Analysis of Object Detection Metrics with a Companion Open-Source Toolkit. Electronics 2021, 10, 279. [CrossRef]
- Gallagher, J. How to Fine-Tune SAM-2.1 on a Custom Dataset. (2024). Roboflow Blog: https://blog.roboflow.com/fine-tune-sam-2- 1/.
- Muja, M.; Lowe, D.G. FAST APPROXIMATE NEAREST NEIGHBORS WITH AUTOMATIC ALGORITHM CONFIGURATION. International Conference on Computer Vision Theory and Applications. pp. 331–340.
- Lowe, D.G. Distinctive Image Features from Scale-Invariant Keypoints. Int. J. Comput. Vis. 2004, 60, 91–110. [CrossRef]
- Fischler, M.A.; Bolles, R.C. Random Sample Consensus: A Paradigm for Model Fitting with Applications to Image Analysis and Automated Cartography. In Readings in Computer Vision; Fischler, M.A., Firschein, O., Eds.; Morgan Kaufmann: San Francisco (CA), 1987; pp. 726–740 ISBN 978-0-08-051581-6.
- Dubrofsky, E. (2009). Homography estimation. Diplomová práce. Vancouver: Univerzita Britské Kolumbie, 5.
- Haralick, R. M., Shanmugam, K., & Dinstein, I. H. (2007). Textural features for image classification. IEEE Transactions on systems, man, and cybernetics, (6), 610-621.
- Van der Walt, S., Schönberger, J. L., Nunez-Iglesias, J., Boulogne, F., Warner, J. D., Yager, N.,.. & Yu, T. (2014). scikit-image: image processing in Python. PeerJ, 2, e453.
- Hagberg, A.A.; Schult, D.A.; Swart, P.J. Exploring Network Structure, Dynamics, and Function using NetworkX. Python in Science Conference. pp. 11–15.
- Shapiro, S. S., & Wilk, M. B. (1965). An analysis of variance test for normality (complete samples). Biometrika, 52(3-4), 591-611.
- Marden, J.I. Positions and QQ Plots. Stat. Sci. 2004, 19, 606–614. [CrossRef]
- Hotelling, H. (1960). Contributions to probability and statistics: essays in honor of Harold Hotelling. Stanford University Press.
- Abdi, H. (2010). The greenhouse-geisser correction. Encyclopedia of research design, 1(1), 544-548.
- Ducrest, A.; Keller, L.; Roulin, A. Pleiotropy in the melanocortin system, coloration and behavioural syndromes. Trends Ecol. Evol. 2008, 23, 502–510. [CrossRef]
- Järvi, T.; Bakken, M. The function of the variation in the breast stripe of the great tit (Parus major). Anim. Behav. 1984, 32, 590–596. [CrossRef]
- Bagnara, J.T.; Matsumoto, J. (2006). Comparative anatomy and physiology of pigment cells in nonmammalian tissues. The pigmentary system: physiology and pathophysiology, 11-59.
- Sohan, M., Sai Ram, T., & Rami Reddy, C. V. (2024). A review on yolov8 and its advancements. In International Conference on Data Intelligence and Cognitive Informatics (pp. 529- 545). Springer, Singapore.
- Yoon, K.; Lim, C. LayerAct: Advanced Activation Mechanism for Robust Inference of CNNs. Proc. AAAI Conf. Artif. Intell. 2025, 39, 22200–22207. [CrossRef]
- Ramanath, R., & Drew, M. S. (2014). White balance. In Computer Vision (pp. 885-888). Springer, Boston, MA.
- Voulodimos, A.; Doulamis, N.; Doulamis, A.; Protopapadakis, E. Deep Learning for Computer Vision: A Brief Review. Comput. Intell. Neurosci. 2018, 2018, 7068349. [CrossRef]

| Augmentation Type | Upper Limit Value |
| Translate—Translates the image horizontally and vertically by a fraction of the image size [0.0-1.0] | 0.015 |
| Scaling—Scales the image by a gain factor [0-1] | 0.15 |
| BGR channels alteration—Flips the image channels from RGB to BGR with the specified probability [0.0-1.0] | 0.1 |
| Image Mosaic—Combines four training images into one with the specified probability [0.0-1.0] | 0.3 |
| Flip Up and down—Flips the image upside down with the specified probability [0.0-1.0] | 0.5 |
| Flip right and left—Flips the image left to right with the specified probability [0.0-1.0] | 0.5 |
| Cutmix—Combines portions of two images with probability [0.0-1.0] | 0.015 |
| Copy_paste—Copies and pastes objects across images to increase object instances with probability [0.0-1.0] | 0.0 |
| Shearing – Shearing image randomly between [0o-180o] | 3o |
| Degrees – Rotating image randomly between [0o,180o] | 5o |
| Hue – hue of the image randomly between [0.0-1.0] | 0.01 |
| Saturation – saturation of the image randomly between [0.0-1.0] | 0.5 |
| Value – brightness of the image randomly between [0.0-1.0] | 0.4 |
| Model | Precision (Bbox) | Recall (Bbox) | Precision (Mask) | Recall (Mask) |
mAP50 (Mask) |
mAP50-95 (Mask) |
| Training | 0.95 | 0.97 | 0.95 | 0.97 | 0.995 | 0.796 |
| Validation | 0.998 | 1.00 | 0.998 | 1.00 | 0.99 | 0.76 |
| Mode | BCE Loss | IoU Loss | Mask Loss |
| Training | 0.008 | 0.1665 | 0.0025 |
| Fish Sample | Change Left (%) | Change Right (%) |
| 10 | 16.01 | 135.41 |
| 11 | 33.11 | 67.19 |
| 12 | 20 | 20.87 |
| 13 | 48.98 | 5.19 |
| 14 | 53.33 | 38.88 |
| 15 | -8.07 | -0.8 |
| 16 | 15.41 | 24.56 |
| 17 | 34.70 | 6.6 |
| 18 | 83.16 | 60.33 |
| 19 | 111.18 | -11.52 |
| 20 | 85.51 | 0.76 |
| 21 | 76.57 | 25.90 |
| 22 | 63.79 | 133.33 |
| 23 | 45.07 | 64.98 |
| 24 | 59.48 | 18.70 |
| 25 | 86.50 | 173.17 |
| 26 | 57.14 | -7.69 |
| 27 | 76.53 | 85.89 |
| 28 | 13.88 | -3.14 |
| 29 | 67.92 | 128.64 |
| 30 | 103.54 | 8.37 |
| Average | 54.46 | 46.46 |
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2026 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license.




