Submitted:
13 April 2026
Posted:
15 April 2026
You are already at the latest version
Abstract
Keywords:
1. Introduction
2. Methods
2.1. Eligibility Criteria
2.2. Information Sources
2.3. Search Strategy
2.4. PRISMA Flow Diagram: Study Selection Process
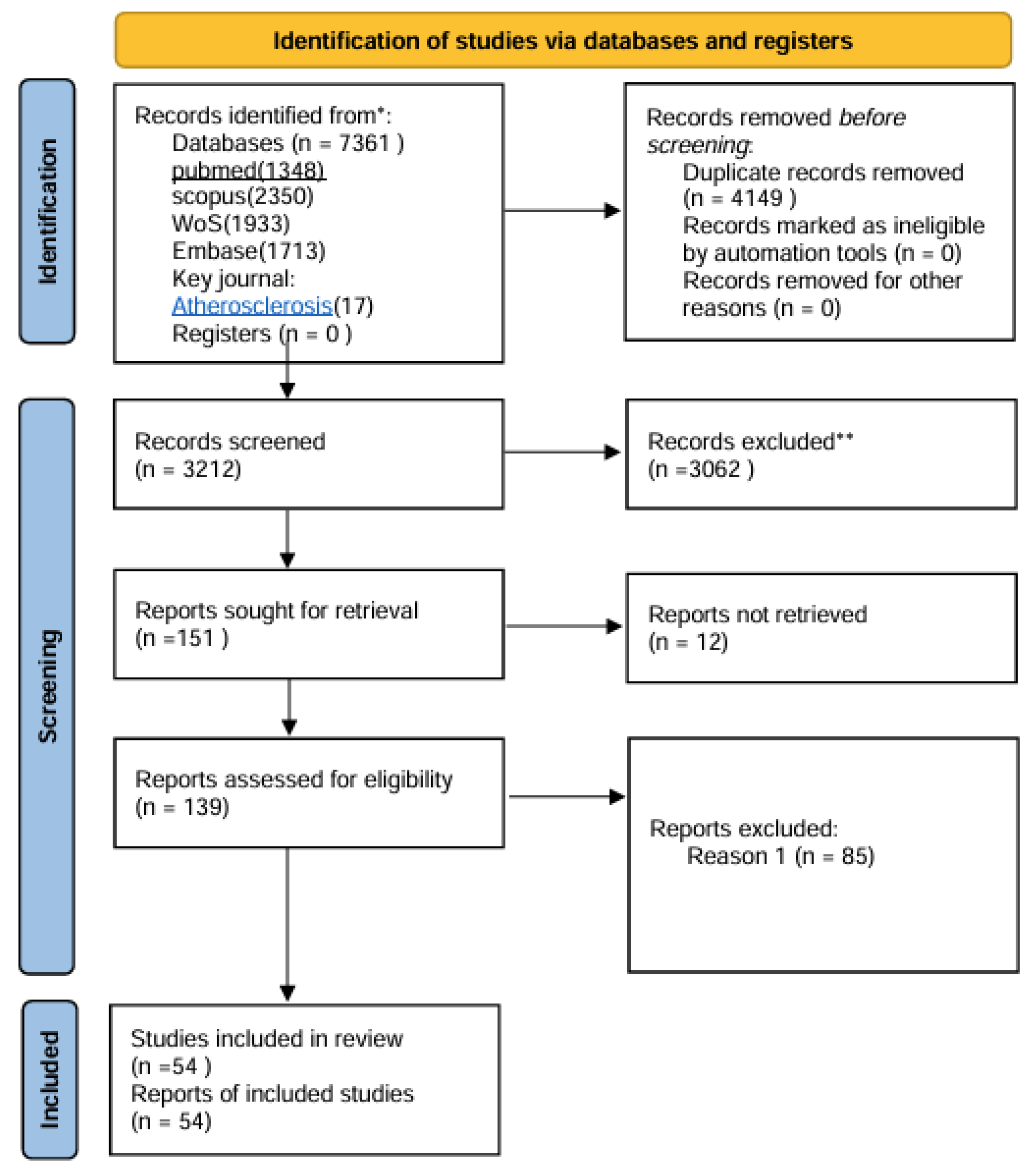
| authors | year | country | Details of study | Sample type | Detection method | Main findings | |
|---|---|---|---|---|---|---|---|
| 1 | Verheye, S. | 2007 | Belgium | Investigate whether stent-based delivery of an inhibitor of mammalian target of rapamycin (mTOR) can selectively clear macrophages in rabbit atherosclerotic plaques. |
White rabbits, murine macrophage cell line J774A.1. |
Taqman-PCR, TEM, Western blot. | The mTOR gene silencing was associated with selective induction of macrophage cell death, The Atg13 protein is phosphorylated in healthy cells but rapidly de- phosphorylated upon inhibition of TOR, stimulating its affinity to Atg1. |
| 2 | Inge De Meyer |
2012 | Belgium | Examination the in vitro and in vivo effects of TLR7 ligand imiquimod on the viability of cultured cells, target selectivity (macrophages), cell death characteristics, and the composition and size of established rabbit atherosclerotic plaques. |
murine macrophage cell line J774A.1, human U937 cells, Bone marrow-derived macrophages were obtained from C57BL/6, TLR7? and TLR7- mice, white rabbits. |
Immunoblot assays, Quantitative ELISAs, color image analysis system, Real time RT-PCR, TEM. |
TLR7 was required for imiquimod-induced Autophagy, imiquimod-treated cells with TLR7 secreted large amounts of several pro-inflammatory cytokines or chemokines including G-CSF, IL-6, IL-12, MCP-1, MCP-5, RANTES and TNF-a. |
| 3 | Shuang Mei |
2012 | North Carolina |
Examination of relationship between P38MAPK and formation of foam cells, an early step in the development of atherosclerosis. |
THP-1 cells. |
Immunoblot assays, Immunoprecipitation, transfection, Fluorescence Microscop, siRNA, Oil Red O Staining. |
LDL stimulated phosphorylation of p38 MAPK in a time- and dose-dependent manner, Activation of p38 MAPK Is Associated with Increased Accumulation of Cholesterol Esters in Macrophages, Exposure to LDL Inhibits Autophagy in Macrophages. |
| 4 | Ai Takeda Watanabe |
2012 | Japan | demonstrating that SIRT1 inactivation impairs starvation-induced autophagy leading to the accumulation of p62/Sqstm1 and resulting in inflammation through NF-kB activation. inhibition of SIRT1 contributes to the regulation of nutrient-sensing pathways by the activation of mTOR and the inhibiting AMPK activation, thereby causing the suppression of autophagy. |
THP-1 cell . |
Western blott, Quantitative RT-PCR, siRNA. |
inhibition of SIRT1 decreases starvation-induced autophagy and increases inflammation in THP-1 cells, The accumulation of p62/Sqstm1 is associated with the dysfunction of autophagy, Inhibition of SIRT1 induces inflammation by suppressing autophagy via mTOR activation. |
| 5 | Zufeng Ding |
2014 | china | Demonstrating that damaged mtDNA could trigger autophagy and NLRP3 inflammasome activation, and LOX-1 may play a critical role in this process. |
THP-1 cell. | siRNA, Western blot, Real-time quantitative PCR assay, fluorescence. |
LPS induces expression of LOX-1 and NLRP3 inflammasome, ROS generation, as well as autophagy and mtDNA damage. |
| 6 | Xuyang Feng |
2014 | china | understanding of the role of autophagy in the development of atherosclerosis, LPS-induced foam cell formation is regulated through Autophagy. |
THP-1 cell. |
enzymatic colorimetric assays, Transfection, fluorescence microscopy, or immunoblotting, Real-time PCR, Western blot analysis. |
LPS promotes foam cell formation by inducing lipid accumulation in macrophages, Autophagy was activated during LPS-induced foam cell formation, the level of ADRP in macrophages is positively correlated with the autophagic activity after treatment with LPS. |
| 7 | Eduardo J. Folco |
2014 | Boston |
Demonstratating IL-1β colocalizes with markers of hypoxia and with activated caspase-1 |
Cell culture(macrophages from PB) | Immunoblot, ELISA, Quantitative Polymerase Chain Reaction PCR, Immunofluorescence, Immunohistochemical Study. |
Hypoxia Induces IL-1β Expression by Stabilizing Pro-IL-1β Protein in Human Macrophages, Hypoxia Augments NLRP3 Induction and Inflammasome Activation. |
| 8 | Baoxin Liu |
2014 | china | Assess connection between enhancement in efferocytosis of apoptotic macrophages and autophagy mediated by Sirt1. |
A mouse macrophage-like RAW264.7 cell. |
western blot , Oil red O staining, flow cytometry, TEM, |
ox-LDL of appropriate concentration elevated the levels of Sirt1 and autophagy marker proteins such as Atg5, Atg7 and LC3-II/LC3-I at optimal time points, autophagy and apoptosis of RAW264.7 cells were triggered by incubation with 50 µM ox-LDL for 24 h, Upregulation of autophagy enhanced efferocytosis of apoptotic RAW264.7 cells. |
| 9 | Xiaochuang Wang |
2014 | china | providing an insight into how mTOR accelerates the pathological process of atherosclerosis. |
RAW 264.7 monocyte/macrophage-like cell. |
siRNA, Western blott, oil red O staining, HPLC, Fluorescence microscopy. |
The mTOR pathway was dramatically activated during foam cell formation, mTOR negatively regulated the activation Of the autophagic machinery responding to lipid loading in macrophage-derived foam cells, ULK1 was responsible for mTOR-regulated foam cell formation by autophagic pathway. |
| 10 | Baojun Huang | 2015 | china | Exploring how ox-LDL induce autophagy in macrophages, and the role of simvastatin in ox-LDL-induced macrophage autophagy and lipid accumulation. |
The J774A.1 murine macrophage Cell. |
Western blott, Confocal laser scanning microscopy, CHOD-PAP. |
Ox-LDL induces lipid accumulation in macrophages, Ox-LDL induces the transformation of LC3 Ⅰ to LC3 Ⅱ in Macrophages. |
| 11 | Guohua Li |
2015 | china | How DNA demethylation modifications and TET2 regulate ox-LDL-treated THP-1 macrophages autophagy. |
THP-1 monocyte cells. |
siRNA, Real-Time Quantitative PCR, Western Blot Analysis, Cell Immunofluorescence. |
ox-LDL inhibited macrophage autophagy, Beclin 1 and LC3-II Expression Was Inhibited by ox-LDL in THP-1-Derived Macrophages, TET2 mRNA levels decreased. |
| 12 | Ping-Ge Tian |
2015 | china | demonstrating that XBP1 mRNA splicing participates in macrophage proliferation, apoptosis, and autophagy via the transcriptional activation of Beclian-1. |
Bone marrow-derived macrophages were harvested from the femurs of wild-type C57BL/6 mice. |
cell proliferation assays, TEM, RT-PCR assays, Immunoblotting and immunohistochemistry staining assays. | The transient activation of XBP1 mRNA splicing triggers autophagy in macrophages, transient activation of XBP1 mRNA splicing for 24 or 48 h can induce autophagy in macrophages via the transcriptional regulation of Beclin-1. |
| 13 | Xi-Ming Yuan |
2016 | Sweden | Investigateing whether 7-oxysterols mixed in an atheroma-relevant proportion induces autophagy, whether autophagy induction influences 7-oxysterol-mediated apoptosis, and the underlying mechanisms, by focusing on cellular lipid levels, oxidative stress, and LMP in 7-oxysterol-treated-a human leukemic cell line (THP-1) cells. |
THP-1 cells . |
oil red O staining or acridine orange (AO) relocation test, flow cytometry, immunocytochemistry, confocal microscopy, Western blot analysis, TEM. |
7-Oxysterols induce lipid accumulation and dysfunction of autophagy. |
| 14 | Feng Zhou |
2016 | china | Evaluating autophagic effects of macrophages on MMP-9 expression, and the underlying mechanisms. |
THP-1 cells. | Oil red O staining, MTT cell apoptosis assay, siRNA, Western-blot analysis, real-time polymerase chain reaction analysis (PCR). |
oxLDL induces macrophages into defected autophagy, defected auto- phagy in macrophage foam cells was accompanied by decreased LC3 and increased p62/SQSTM1, autophagic Flux situation was involved in balance of MMP-9 formation and degradation in THP-1 derived macrophages, p62/SQSTM1 upregulates MMP-9 gene expression in dependent NF-kB signaling |
| 15 | Jiangping He | 2017 | china | SIRT6 overexpres- sion protected THP1 cells treated with oxidized low-density lipoprotein (ox-LDL) from forming foam cells, whereas SIRT6 knockdown aggravated foam cell formation. |
THP1 cell. | Immunoblot analysis, qRT-PCR, transfections, TEM, ORO staining, Flow cytometry. | SIRT6 and autophagy are inhibited in THP1 cells treated with ox-LDL. |
| 16 | Haofeng Ning | 2017 | china | investigating the correlation between p62 expression and oxLDL-induced foam cell formation. | THP-1 cell. | siRNA, Oil Red O staining, Western blott, RT-qPCR, ELISAM, MTT assay, FACS. | Prolonged oxLDL treatment induces p62 protein accumulation during foam cell formation, and increase IL-18 secretion and impaired autophagy. |
| 17 | Se-Jin Jeong | 2018 | korea | Investigating Prdx1 deficiency in macrophages led to increased susceptibility to oxidative stress and suppressed the clearance of modified LDL as aresult of impaired lipophagic flux. |
(ApoE)-deficient mice | confocal microscopy, fluorescence microscopy, flow cytometry analysis, Immunohistochemistry, Oil Red O staining, Quantitative real-time PCR analysis, Immunoblot analysis, Electron microscopy |
Prdx1 deficiency causes defective autophagic flux in macrophages, increases macrophage foam cells |
| 18 | Xing Liang | 2017 | china | Evaluation of feedback loop of the “EMMPRIN/NF-κB” pathway in atherosclerotic plaques via modulation of autophagy in macrophages. | (ApoE)-deficient mice, RAW264.7 cells. |
oil red O staining, haematoxylin and eosin (H&E), immunohistochemistry (IHC), Immunofluorescence staining, siRNA, flow cytometry, Real-time PCR, TEM, Western blotting analysis, ELISA. | Activation of the PI3K/Akt/mTOR signalling pathway plays a minor role in EMMPRIN-regulated macrophage autophagy, Down-regulation of EMMPRIN increases macrophage autophagy when treated with ox-LDL, EMMPRIN inhibits autophagy in oxLDL-stimulated macrophages primarily via NF-κB activation. |
| 19 | Wenhua Sun | 2018 | china | A link between autophagy and oxLDL. | Carotid artery tissues, THP-1 cells. | Western blot analysis, TUNEL. | Overexpression LC3 and beclin1 in macrophages during autophagy. |
| 20 | Sen Yang | 2018 | china | A link autophagy, inflammation and oxLDL. | The mouse macrophage-like cell line Raw264.7. | siRNA, Western blot analysis, ELISA. | Ox-LDL impairs autophagic flux in macrophages, ox-LDL significantly increased p62 , LC3-II expression levels in macrophages, Ox-LDL induce inflammation in macrophages. |
| 21 | Lu Zhang | 2018 | china | How CTRP9 can affect foam cell formation by activating autophagy. | THP-1 monocytes cells. | Oil Red O staining, Cholesterol Efflux Assay, Western Blot Assay | the ox-LDL group, the protein expression level of LC3 II was downregulated, whereas that of p62 was upregulated, ABCA1 and ABCG1 protein levels were downregulated, CTRP9 Induces Autophagy by Regulating the AMPK/mTOR Pathway in THP-1 Macrophage- Derived Foam Cells |
| 22 | Kathrin Ackermann | 2018 | German | investigating the in uence of GDF-15 in lipid homeostasis and autophagy in human MΦ during foam cell formation. |
THP-1 cells. | ELISA, Western blot, SDS-PAGE, RT-PCR, Oil Red O staining, Fluorometric assay, Immunocytofluorescence confocal laser scanning microscopy. | autophagy-relevant proteins/complexes ATG5, ATG12/ATG5 and p62 in THP-1 increased. |
| 23 | Shan Li | 2018 | china | Evaluating Programmed cell death protein 4 (PDCD4), a transcriptional Regulator’ effect on regulation of autophagy. |
Mice monocyte-macrophage leukemia-derived cell line RAW 264.7 and human monocytic cell line THP-1. |
Oil Red O staining, transfection, Real-time quantitative polymerase chain reaction, Western blots, Immunofluorescence, Flow cytometry. | PDCD4, negatively regulated autophagy, PDCD4-mediated autophagy is involved in the process of lipid hypolysis and efflux. |
| 24 | Xiuying Li | 2018 | china | Evaluating pivotal role of macrophage autophagy in the pathogenesis of atherosclerosis, assessment of dectin-1 binding by β-glucan converts RAW264.7 macrophages into an M1 phenotype via autophagy. |
RAW264.7 cells, ApoE-/- mice. | MTT assay, Flow cytometry, Western blott, RT-qPCR, TEM, Immunofluorescent histochemistry. | The ratio of LC3II/I decreased, β-glucan may suppress autophagy by downregulating the level of LC3-II/I and beclin-1. |
| 25 | Xiaofei Liang | 2019 | China | Assessment p62/mTOR/LXRa pathway inhibits cholesterol efflux mediated by ABCA1 and ABCG1 during autophagy blockage. | THP-1 cells. | siRNA, Western blot analysis, Fluorescence analysis, Oil red O staining. | Excess ox-LDL inhibited the expression of ABCA1 and ABCG1, and induced blockage of autophagy, p62 overexpression accelerates foam cell formation during autophagy blockage, A p62/mTOR/LXRa signaling pathway is involved in the formation of foam cells during autophagy blockage |
| 26 | Shanshan Zhong | 2019 | china | Assessing role of ALDH2 and LDLR in atherosclerosis. |
APOE–/– mice, HEK 293T cell lines, | Immunohistochemistry, Phagocytotic assay, Western blott, SDS-PAGE, GC-MS, Real-time PCR, Immunoprecipitation, Fluorescence microscopy, RNA-Seq analysis, ChIP assay, confocal microscopy, Luciferase assay. | AMPK plays an important role in regulating endocytosis and autophagy for LDLR-regulated ox-LDL metabolism in macrophages. |
| 27 | Xiaozhen Zhuo | 2019 | China | To explore the mechanism of how LSD1 regulates autophagy and the correlation between LSD1 and Ox-LDL-induced inflammation. | RAW264.7 murine macrophage cell. | RT-qPCR, Western blott, CCK-8 assay, siRNA, ELISA. | Ox-LDL upregulates the expression level of LSD1 in RAW264.7 cells, LSD1 inhibition activated autophagy via SESN2-mediated PI3K/Akt/Mtor pathway |
| 28 | Nicholas D. LeBlond | 2020 | canada | Foam Cell Induction Activates AMPK But Uncouples Its Regulation of Autophagy and Lysosomal Homeostasis |
Mice, BMDM mice. | Immunoblotting, Quantitative PCR, Immunofluorescent labeling, Immunofluorescent imaging, HPLC | Atherogenic Lipids Activate AMPK in Bone Marrow-Derived Macrophages, Treatment with oxLDL dramatically enhanced AMPK-specific ULK1 Ser555 phosphorylation and increased the conversion of LC3II in WT, but not CD36+/−cells, suggesting that CD36 plays a role in transmitting the atherogenic signal to AMPK, which in turn signals to regulate autophagy via TFEB programs. |
| 29 | Chao Wang | 2020 | China | How MicroRNA-761 modulates foam cell formation and inflammation through autophagy in the progression of atherosclerosis. |
THP-1 cell. | RNA transfection, Quantitative reverse transcription polymerase chain Reaction, Western blot (WB), bioinformatic analysis, Oil Red "O" staining, ELISA, | MiR-761 significantly elevated under the stimulation of ox-LD, Macrophage autophagy was promoted by miR-761 through mTOR-ULK1 pathway, miR-761 negatively affect the secretion of IL-1β and IL-18. |
| 30 | Qingqing Xiao | 2020 | china | How Macrophage autophagy regulates mitochondria-mediated Apoptosis. |
ApoE−/−C57BL/6 mice, RAW264.7 cells | haematoxylin and eosin (H&E, Sigma), Masson's trichrome (Sigma) and Oil Red O staining, Western blot analysis, Flow cytometric analysis, Real-time quantitative PCR, siRNA, Immunofluorescence staining, TUNNEL, TEM | Autophagy flux was blocked during 7-KC– induced macrophages apoptosis, accumulation of SQSTM1/P62 was paralleled to an increase in the ratio of LC3II/LC3I, reflecting impairment in the autophagy flux, Elevated MAPK and NF-κB activation with autophagy impairment. |
| 31 | MD Khurshidul Zahid | 2020 | USA | Evaluating macrophage foam cell formation in atherogenesis and inflammation, ER stress, and apoptosis and by promoting autophagy and inactivating mTOR. | RAW264.7 macrophage cells. | qPCR, Western blot analysis. | CEBPβ have a role in NFkB phosphorylation and NFkB and TNFα genes expression, increased ER stress, mTOR activation and apoptosis , decreased auophagy related genes in RAW264.7 macrophage cells, |
| 32 | Zhenfeng Zhou | 2020 | china | How K63 ubiquitin chains target NLRP3 inflammasome for autophagic degradation in ox-LDL stimulated THP-1 macrophages. | THP-1 cell. | Oil red O staining, siRNA, ELISA, Western blot, Immunoprecipitation, | Ox-LDLs activate NLRP3 inflammasome and secretion NLRP3, ASC, pro-caspase-1, pro-IL-1β, and activated caspase-1 in a time- and dose-dependent manner, ox-LDLs restrict autophagy in a time- and dose-dependent manner, p62 plays an important role in the foam-cell model, and mediates the regulation of NLRP3 inflammasome by autophagy, p62 mediates the regulation of NLRP3 inflammasomes through autophagy by recognizing the K63 polyubiquitin chains on NLRP3. |
| 33 | SeJeong Kim | 2021 | korea | Investigating the P62 role in macrophage autophagy. | Human umbilical vein endothelial cells, THP-1 cell. | siRNA, Quantitative Real-Time PCR (qPCR) Analysis, ELISA, BODIPY Staining, Immunocytochemistry, Western Blot Analysis. | There is abnormal autophagy in Atherosclerotic Conditions via increased p62. |
| 34 | Jiaru Liu | 2021 | china | Regulation of pyroptosis and autophagy via p62/Nrf2/ARE axis. | THP-1 cells. | CCK-8 assay, LDH release assay, ELISA, Flow cytometry analysis, Western blot analysis, qRT-PCR, Bioinformatic analyses. | ox-LDL induced THP-1 macrophages to pyroptosis in a concentration dependent manner, induced the both mRNA and protein expressions of pro-caspase-1 and GSDMD, Autophagy blockage triggered pyroptosis and inflammation in macrophages exposed to ox-LDL via Nrf2/ARE , p62/Nrf2/ARE pathway. |
| 35 | Jun Tao | 2021 | china | AMPK/FoxO1/TFEB signalling axis role in lysosomal biogenesis, foam cell formation and autophagy. |
murine macrophage RAW264.7 cell | RT-PCR, shRNA, Immunofluorescence, Western blotting analysis, Oil Red O staining. | Atherosclerotic macrophages have features of impaired autophagy and dysfunctional lysosomes, LC3 and SQSTM1/p62 elevated and dysregulated AMPK/ FoxO1/TFEB pathway. |
| 36 | Hui Zhang | 2021 | china | How ATG14 impact on inflammation and autophagosome lysosome fusion in macrophages. | Human artery samples, apoe−/- mice, Raw264.7 cells | Immunostaining, Western blott, flow cytometry, Electron microscopy, ELISA, RT qPCR assay. | Autophagy dysfunction with reduction of ATG14 expression was detected in macrophage in human atherosclerotic plaque, reduction of ATG14 induce inflammasome actiation. |
| 37 | Qi Peng | 2022 | china | Role of Nrf2 in autophagy and ferroptosis | THP-1 cell. | Western blot, CCK8 cell activity assay, | Nrf2 was increased by ox-LDL stimulation and xCT and GPX4 were decreased by ox-LDL stimulation, Inadequate autophagy of foam cells accelerates foam cell death via accumulation of p62 , LC3II/LC3I was significantly reduced, Insufficient autophagy in foam cells initiates Nrf2-induced ROS accumulation, The negative effect of Nrf2 further promotes ferroptosis in foam cells |
| 38 | Masataka Sano | 2022 | china | Neutrophil extracellular traps-mediated Beclin-1 suppression aggravates atherosclerosis by inhibiting macrophage autophagy |
apoE deficient mice, HL-60 cells, THP-1 cells | Immunoblot analyses, Enzyme-linked immunosorbent assay, Immunoprecipitation assay, Oil Red O staining , Immunocytochemistry, Immunohistochemistry, SEM, TEM, fluorescent assay |
NETs inhibit the kinase activity of Beclin-1-dependent PI3 kinase activity possibly through phosphorylating the tyrosine residues of Beclin-1 in the macrophages, Neutrophil extracellular traps negatively regulate autophagosome–lysosome fusion through upregulating Rubicon expression |
| 39 | Weihua Shao | 2022 | china | Identification of the relationship between the miR-29a-targeted PI3K signaling pathway and AS | ApoE−/− mice, mouse RAW264.7 cells | Bioinformatic analysis, histological analysis, MOVAT staining, Immunofluorescence and immunohistochemistry, Luciferase assay,TEM, Western blott | MiR-29a increased the expressions of IL-10, Mrc1 and Arginase-1 and decreased the expressions of IFN-γ, IL-1β and iNOS in vivo, MiR-29a overexpression increased autophagy and suppressed the PI3K/AKT/mTOR pathway |
| 40 | Zhen Tan | 2022 | china | Identification of the relationship between KLF2, Nrf2 and foam cell formation. | HUVECs, THP-1 monocytes | Transfection, RT-qPCR, Western blott, CCK-8 assay, ELISA, Oil Red O staining, | The expression of KLF2, Nrf2 in THP-1 macrophage-derived foam cells was decreased and impaired autophagy. |
| 41 | Wei Yu | 2022 | china | How can Uric Acid affect NRF2-Mediated Autophagy Dysfunction and Ferroptosis | ApoE−/− mice, THP-1 cells | Oil Red O Staining, Immunohistochemical Staining, Immunofluorescence Microscopy, CCK-8 assay, MDA assay, flow Cytometry, GSH assay, TEM, Immunoblotting , qPCR Analysis. |
HUA Inhibits the Protein Level of the NRF2/SLC7A11/ GPX4 Signaling Pathway in Macrophages in Atherosclerotic Plaques, NRF2-Mediated Autophagy Dysfunction and Ferroptosis Are Involved in Foam Cell Formation Induced by HUA |
| 42 | Liang Zheng | 2022 | china | Identification of relationship autophagy, NLRP3 and apoptosis | THP-1 monocytes | Transfection, TEM, Western blot analysis, Immunofluorescence assay, Oil Red O (ORO) staining, TUNNEL | ATG7 and Beclin1 increased, NLRP3 activity was increased |
| 43 | Guofu Hu | 2023 | china | Identification of relationship between autophagy and ferroptosis. | THP-1 monocytes, Human carotid atherosclerotic plaques, Apolipoprotein E knock out (ApoE-/-) mice | Bioinformatic analysis, Histological examination, Real-time PCR, Western blot, Immunofluorescent staining, Oil-red-o staining, TEM, GSH assay. | When autophagy decreased, ferroptosis been activated |
| 44 | Xuemei Hu | 2023 | china | Identification the role of Sialic acids ROS and autophagy blockage | RAW264.7 cells, APOE−/− mice | Real time PCR quantification, CCK-8 assay, Western blot, Flow cytometry, Immunofluorescence staining, flow cytometry | AC5-induced mitochondrial dysfunction and ROS production in macrophages is correlated with macrophage polarization, AC5 promotes autophagosome formation but decreases autophagic lysosomes fusion resulting in the autophagy flux blockage, |
| 45 | Shengmei Zeng | 2023 | china | How ATG5 affect macrophages Autophagy and inflammation under hypoxia |
RAW264.7 cells, C57BL/6 mice | Western blot, Immunofluorescence assay, siRNA, TEM, qRT-PCR, Co-immunoprecipitation | Hypoxia significantly upregulated macrophage inflammatory factor Levels, hypoxia-induced inflammation in RAW264.7 cells is mediated by autophagy, desialylation of ATG5 enhances ATG12–ATG5–ATG16L complex formation and thereby promotes hypoxia-induced autophagosome, ATG5 can affect its stability and promote the formation of the ATG5-ATG16L ATG12 complex. |
| 46 | Pengchen He, MM | 2024 | china | Effect of connexin 43 in LPS/IL-4-induced macrophage M1/M2 polarization |
RAW264.7 macrophages | Western blot, Immunofluorescence, Flow cytometry | M2-type polarization reduces autophagy cx43 protein expression was first decreased and then increased by IL-4 in RAW264.7 macrophages |
| 47 | Yuan-Mei Wang | 2024 | china | ASIC1/RIP1 accelerates atherosclerosis via disrupting lipophagy |
Human aortic plaques, ApoE-/- mice, RAW 264.7 macrophages, THP-1 cell | Bioinformatic analysis, Hematoxylin and eosin (H&E) staining, Immunofluorescence staining, ORO staining, Western blot analysis, TEM, | ASIC1 is abundantly expressed in macrophages within atherosclerotic Lesions, ASIC1 promotes RIP1 phosphorylation in both RAW 264.7 and THP-1 Macrophages, ASIC1-RIP1 association contributes to defective autophagy flux in both RAW 264.7 and THP-1 macrophages induced by ox-LDL, ASIC1/RIP1 facilitates lipid accumulation in RAW 264.7 macrophages by inhibiting lipophagy |
| 48 | Qianqian Wu | 2024 | china | The Effects of the oxLDL/β2GPI/anti-β2GPI Complex on Macrophage Autophagy and its Mechanism |
THP-1 cells | Western Blot Analysis, Adenoviral Transfection, TEM |
xpression of SQSTM1/P62 was significantly increased in the oxLDL/β2GPI/anti-β2GPI group, the oxLDL/β2GPI/anti-β2GPI treat- ment decreases macrophage autophagy, oxLDL/β2GPI/anti-β2GPI Complex Reduces the Autophagosomes and Blocks Autophagic Flux in Macrophages, oxLDL/β2GPI/anti-β2GPI Complex Enhances the Activity of PI3K/AKT/mTOR Pathway in Macrophages |
| 49 | Min Zeng | 2024 | china | How CTRP9 affect atherosclerosis progression through changing autophagic status of macrophages by activating USP22-mediated de-ubiquitination on Sirt1 |
Macrophages from PB | Cell transfection, Immunofluorescence, Oil Red O staining, ELISA, fluorescence assay, RT-qPCR, western blot | CTRP9 attenuated impaired cell viability, autophagy inhibition and increased lipid accumulation induced by ox-LDL, CTRP9 maintained Sirt1 protein level through enhancing its stability, CTRP9 triggered the de-ubiquitination of Sirt1 via up-regulating USP22 expression |
| 50 | Rui Bu |
2025 | china | Downregulation of ATP8B2 in atherosclerosis exacerbates foam cell- like pathological changes via impairing lysosomal membrane fusion |
THP-1 cells | Bioinformatics analysis, siRNA, RT-qPCR, Western blot, Immunofluorescence staining | Downregulation of ATP8B2 may promote the development of atherosclerosis, |
| 51 | Siyu Fan | 2025 | Netherlands | Detection of autophagy-lysosome regulation in an atherosclerosis cell model | RAW 264.7 macrophages | N.A | Increased ROS levels, The activation of TFEB might lead to autophagy-lysosomal biogenesis, early burst of lysosomal ROS triggers TFEB nuclear translocation |
| 52 | Qiujun Liu | 2025 | china | Identification of how Pyruvate dehydrogenase affect on macrophage autophagy | ApoE–/– mice, RAW264.7 macrophages | Transfection, Co-immunoprecipitation (Co-IP) assays, Western blot analysis, Immunofluorescence, bioinformatic analysis | Hcy inhibits autophagy by regulating glycolysis in macrophages via the inhibition of pyruvate dehydrogenase and AMPK/mTOR signaling pathway |
| 53 | Xiaodong Miao | 2025 | china | Down-regulation of ATP8B2 in Foam Cells Inhibits Autophagic Flux and ox-LDL Degradation in Atherosclerosis |
THP-1 human monocytic leukemia cell | Bioinformatic analysis, RT-PCR, Western Blot, Immunofluorescence | Downregulation of ATP8B2 Inhibits Autophagic Flux in Macrophages, Downregulation of ATP8B2 Inhibits ox-LDL Degradation and Mitochondrial Homeostasis in Macrophages |
| 54 | Kai Wen Wai | 2025 | Malaysia | How Nrf2 modulates macrophage foam cells senescence and autophagy activation |
THP-1 human monocytic cell | siRNA, Oil red O (ORO) assay, Autophagy assay, Senescence β-galactosidase staining, Western blot |
Increased Nrf2 , activate macrophage foam cell Formation and reduced macrophage autophagy |
3. Results
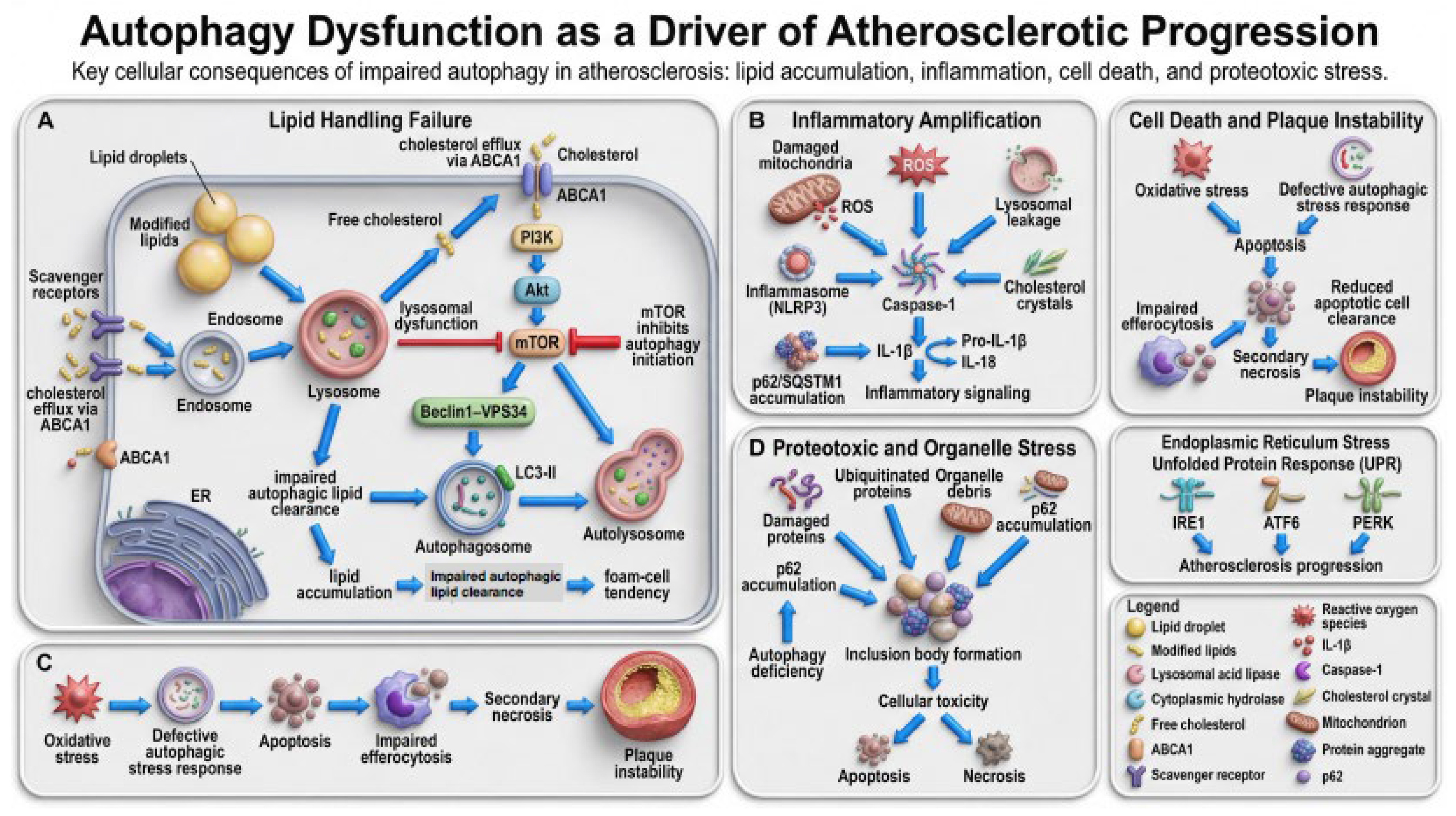
3.1. To Systematically Evaluate the Role of Macrophage Autophagy in the Initiation, Progression, and Stability of Atherosclerotic Plaques, Including its Impact on Lipid Metabolism and Foam Cell Formation
3.2. Macrophage Autophagy and Inflammatory Responses
3.3. Selective Autophagy Processes, like Mitophagy, Ferroptosis and Efferocytosis
3.4. Selective Autophagy Processes and miRNAs
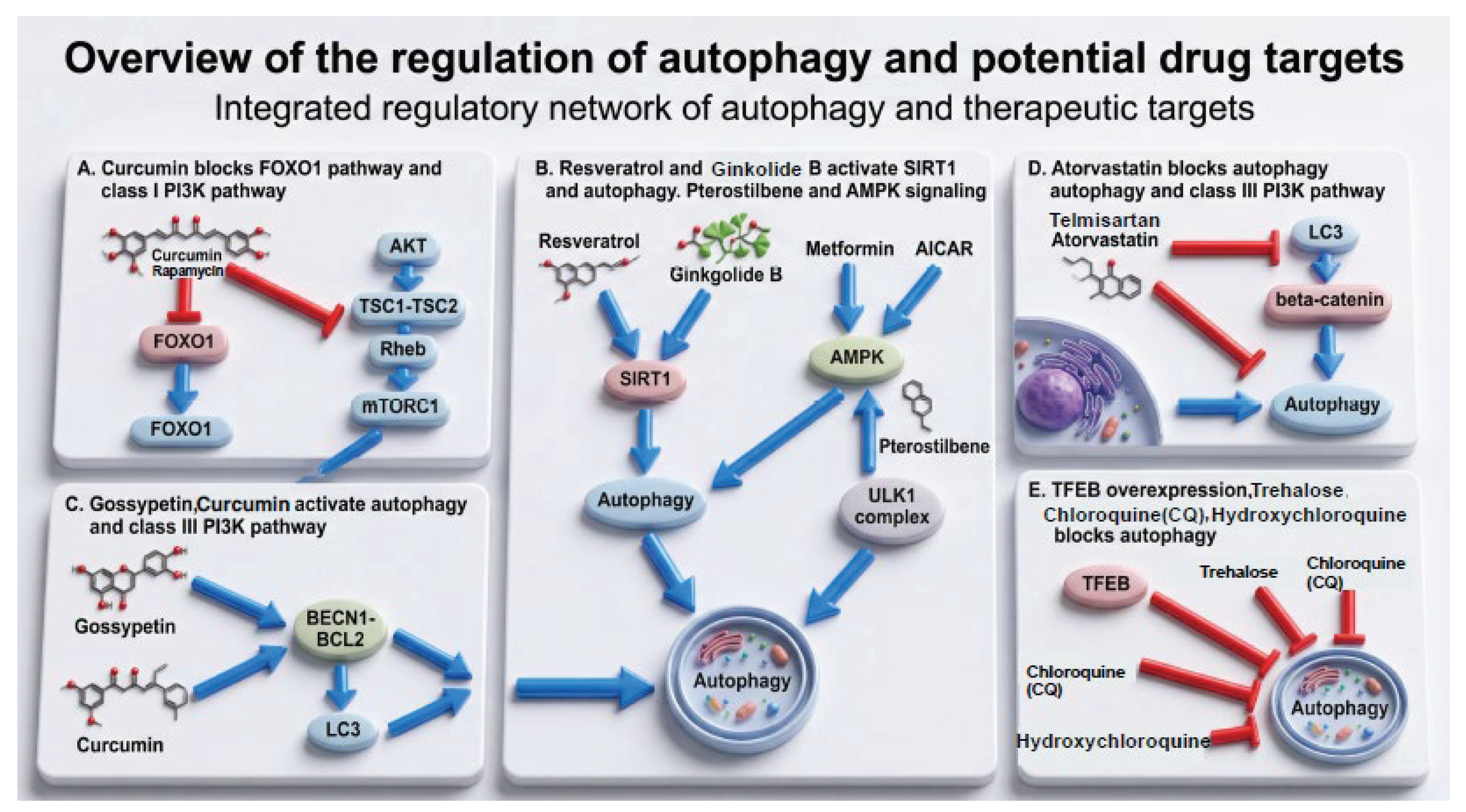
4. Discussion
5. Conclusion
Author Contributions
Funding
Availability of Data and Materials
Availability of Data and Materials
Competing interests
References
- Wirawan, E.; Berghe, T.V.; Lippens, S.; Agostinis, P.; Vandenabeele, P. Autophagy: for better or for worse. Cell research 2012, 22[1], 43–61. [Google Scholar] [CrossRef]
- Tukaj, C. The significance of macroautophagy in health and disease. Folia Morphologica 2013, 72[2], 87–93. [Google Scholar] [CrossRef] [PubMed]
- Yorimitsu, T.; Klionsky, D.J. Autophagy: molecular machinery for self-eating. Cell Death & Differentiation 2005, 12(2), 1542–52. [Google Scholar] [CrossRef] [PubMed]
- Mizushima, N. Autophagy: process and function. Genes & development 2007, 21(22), 2861–73. [Google Scholar]
- Badadani, M. Autophagy mechanism, regulation, functions, and disorders. International Scholarly Research Notices 2012, 2012(1), 927064. [Google Scholar] [CrossRef]
- Yang, Z.; Klionsky, D.J. An overview of the molecular mechanism of autophagy. In Autophagy in infection and immunity; 2009; pp. 1–32. [Google Scholar]
- Martinet, W.; Agostinis, P.; Vanhoecke, B.; Dewaele, M.; De Meyer, G.R. Autophagy in disease: a double-edged sword with therapeutic potential. Clinical science 2009, 116(9), 697–712. [Google Scholar] [CrossRef]
- Juretschke, T.; Beli, P. Causes and consequences of DNA damage-induced autophagy. Matrix Biology 2021, 100, 39–53. [Google Scholar] [CrossRef]
- Stead, E.R.; Castillo-Quan, J.I.; Miguel, V.E.M.; Lujan, C.; Ketteler, R.; Kinghorn, K.J.; et al. Agephagy–adapting autophagy for health during aging. Frontiers in cell and developmental biology 2019, 7, 308. [Google Scholar] [CrossRef]
- Glick, D.; Barth, S.; Macleod, K.F. Autophagy: cellular and molecular mechanisms. The Journal of pathology 2010, 221(1), 3–12. [Google Scholar] [CrossRef]
- Murrow, L.; Debnath, J. Autophagy as a stress-response and quality-control mechanism: implications for cell injury and human disease. Annual Review of Pathology: Mechanisms of Disease 2013, 8(1), 105–37. [Google Scholar] [CrossRef]
- Singh, R.B.; Mengi, S.A.; Xu, Y.-J.; Arneja, A.S.; Dhalla, N.S. Pathogenesis of atherosclerosis: A multifactorial process. Experimental & Clinical Cardiology 2002, 7(1), 40. [Google Scholar]
- Jebari-Benslaiman, S.; Galicia-García, U.; Larrea-Sebal, A.; Olaetxea, J.R.; Alloza, I.; Vandenbroeck, K.; et al. Pathophysiology of atherosclerosis. International journal of molecular sciences 2022, 23(6), 3346. [Google Scholar] [CrossRef]
- Ross, R. Cell biology of atherosclerosis. Annual review of physiology 1995, 57, 791–804. [Google Scholar] [CrossRef] [PubMed]
- Rafieian-Kopaei, M.; Setorki, M.; Doudi, M.; Baradaran, A.; Nasri, H. Atherosclerosis: process, indicators, risk factors and new hopes. International journal of preventive medicine 2014, 5(8), 927. [Google Scholar] [PubMed]
- van Rooy, M.-J.; Pretorius, E. Obesity, hypertension and hypercholesterolemia as risk factors for atherosclerosis leading to ischemic events. Current Medicinal Chemistry 2014, 21(19), 2121–9. [Google Scholar] [CrossRef] [PubMed]
- Batty, M.; Bennett, M.R.; Yu, E. The role of oxidative stress in atherosclerosis. Cells 2022, 11(23), 3843. [Google Scholar] [CrossRef]
- Poznyak, A.V.; Nikiforov, N.G.; Markin, A.M.; Kashirskikh, D.A.; Myasoedova, V.A.; Gerasimova, E.V.; et al. Overview of OxLDL and its impact on cardiovascular health: focus on atherosclerosis. Frontiers in Pharmacology 2021, 11, 613780. [Google Scholar] [CrossRef]
- Berliner, J.A.; Navab, M.; Fogelman, A.M.; Frank, J.S.; Demer, L.L.; Edwards, P.A.; et al. Atherosclerosis: basic mechanisms: oxidation, inflammation, and genetics. Circulation 1995, 91(9), 2488–96. [Google Scholar] [CrossRef]
- Hassanpour, M.; Rahbarghazi, R.; Nouri, M.; Aghamohammadzadeh, N.; Safaei, N.; Ahmadi, M. Role of autophagy in atherosclerosis: foe or friend? Journal of Inflammation 2019, 16, 1–10. [Google Scholar] [CrossRef]
- Martinet, W.; De Meyer, G.R. Autophagy in atherosclerosis. Curr Atheroscler Rep. 2008, 10(3), 216–23. [Google Scholar] [CrossRef]
- De Meyer, G.R.; Grootaert, M.O.; Michiels, C.F.; Kurdi, A.; Schrijvers, D.M.; Martinet, W. Autophagy in vascular disease. Circulation research 2015, 116(3), 468–79. [Google Scholar] [CrossRef]
- Henderson, J.M.; Weber, C.; Santovito, D. Beyond Self-Recycling: Cell-Specific Role of Autophagy in Atherosclerosis. Cells 2021, 10(3). [Google Scholar] [CrossRef] [PubMed]
- Perrotta, I.; Aquila, S. The role of oxidative stress and autophagy in atherosclerosis. Oxidative Medicine and Cellular Longevity 2015, 2015(1), 130315. [Google Scholar] [CrossRef] [PubMed]
- Grootaert, M.O.; Roth, L.; Schrijvers, D.M.; De Meyer, G.R.; Martinet, W. Defective autophagy in atherosclerosis: to die or to senesce? Oxidative medicine and cellular longevity 2018, 2018(1), 7687083. [Google Scholar] [CrossRef] [PubMed]
- Vindis, C. Autophagy: an emerging therapeutic target in vascular diseases. Br J Pharmacol. 2015, 172(9), 2167–78. [Google Scholar] [CrossRef]
- Moore, K.J.; Tabas, I. Macrophages in the pathogenesis of atherosclerosis. Cell. 2011, 145(3), 341–55. [Google Scholar] [CrossRef]
- Miller, Y.I.; Choi, S.-H.; Fang, L.; Tsimikas, S. Lipoprotein modification and macrophage uptake: role of pathologic cholesterol transport in atherogenesis. Cholesterol binding and cholesterol transport proteins: structure and function in health and disease 2010, 229–51. [Google Scholar]
- Gonzalez, L.; Trigatti, B.L. Macrophage apoptosis and necrotic core development in atherosclerosis: a rapidly advancing field with clinical relevance to imaging and therapy. Canadian Journal of Cardiology 2017, 33(3), 303–12. [Google Scholar] [CrossRef]
- Tabas, I. Consequences and therapeutic implications of macrophage apoptosis in atherosclerosis: the importance of lesion stage and phagocytic efficiency. Arteriosclerosis, thrombosis, and vascular biology 2005, 25(11), 2255–64. [Google Scholar] [CrossRef]
- Shao B-z, Han B-z, Zeng Y-x, Su D-f, Liu C. The roles of macrophage autophagy in atherosclerosis. Acta Pharmacologica Sinica, 2016; 37, 2, pp. 150–6. [CrossRef]
- Martinet, W.; De Meyer, G.R. Autophagy in atherosclerosis: a cell survival and death phenomenon with therapeutic potential. Circulation research 2009, 104(3), 304–17. [Google Scholar] [CrossRef]
- Meng, L.B.; Zhang, Y.M.; Luo, Y.; Gong, T.; Liu, D.P. Chronic Stress A Potential Suspect Zero of Atherosclerosis: A Systematic Review. Front Cardiovasc Med. 2021, 8, 738654. [Google Scholar] [CrossRef]
- Kan, J.X.; Zhu, J.P.; Wei, T.; Chen, H. Natural Products Ameliorate Non-Alcoholic Fatty Liver Disease and Atherosclerosis via the LKB1/AMPK Pathway. Nat Pro Comm. 2025, 20(11). [Google Scholar] [CrossRef]
- Page, M.J.; McKenzie, J.E.; Bossuyt, P.M.; Boutron, I.; Hoffmann, T.C.; Mulrow, C.D.; et al. The PRISMA 2020 statement: an updated guideline for reporting systematic reviews. BMJ 2021, 372, n71. [Google Scholar] [CrossRef]
- Feng, X.; Yuan, Y.; Wang, C.; Feng, J.; Yuan, Z.; Zhang, X.; et al. Autophagy involved in lipopolysaccharide-induced foam cell formation is mediated by adipose differentiation-related protein. Lipids Health Dis. 2014, 13(1). [Google Scholar] [CrossRef]
- Wu, Q.; Zhang, G.; Wang, T.; Zhou, H. The Effects of the oxLDL/β2GPI/anti-β2GPI Complex on Macrophage Autophagy and its Mechanism. Immun Inflamm Dis. 2024, 12(11), e70058. [Google Scholar] [CrossRef]
- Huang, B.; Jin, M.; Yan, H.; Cheng, Y.; Huang, D.; Ying, S.; et al. Simvastatin enhances oxidized-low density lipoprotein-induced macrophage autophagy and attenuates lipid aggregation. Mol Med Rep. 2015, 11(2), 1093–8. [Google Scholar] [CrossRef]
- Sun, W.; Lin, Y.; Chen, L.; Ma, R.; Cao, J.; Yao, J.; et al. Legumain suppresses OxLDL-induced macrophage apoptosis through enhancement of the autophagy pathway. Gene 2018, 652, 16–24. [Google Scholar] [CrossRef]
- Zheng, L.; Xu, H.; Zheng, F.; Lai, Y.; Li, J.; Lv, W.; et al. Intervention time decides the status of autophagy, NLRP3 activity and apoptosis in macrophages induced by ox-LDL. Lipids Health Dis. 2022, 21(1). [Google Scholar] [CrossRef]
- Zhang, H.; Ge, S.; Ni, B.; He, K.; Zhu, P.; Wu, X.; et al. Augmenting ATG14 alleviates atherosclerosis and inhibits inflammation via promotion of autophagosome-lysosome fusion in macrophages. Autophagy 2021, 17(12), 4218–30. [Google Scholar] [CrossRef]
- Tian, P.G.; Jiang, Z.X.; Li, J.H.; Zhou, Z.; Zhang, Q.H. Spliced XBP1 promotes macrophage survival and autophagy by interacting with Beclin-1. Biochem Biophys Res Commun. 2015, 463(4), 518–23. [Google Scholar] [CrossRef]
- He, P.; Dai, M.; Li, Z.; Wang, X.; Liu, H.; He, Y.; et al. Effect of connexin 43 in LPS/IL-4-induced macrophage M1/M2 polarization: An observational study. Medicine 2024, 103(15), E37811. [Google Scholar] [CrossRef]
- Li, G.; Peng, J.; Liu, Y.; Li, X.; Yang, Q.; Li, Y.; et al. Oxidized low-density lipoprotein inhibits THP-1-derived macrophage autophagy via TET2 down-regulation. Lipids 2015, 50(2), 177–83. [Google Scholar] [CrossRef]
- Li, X.; Luo, H.; Ye, Y.; Chen, X.; Zou, Y.; Duan, J.; et al. β-glucan, a dectin-1 ligand, promotes macrophage M1 polarization via NF-κB/autophagy pathway. Int J Oncol. 2019, 54(1), 271–82. [Google Scholar] [CrossRef]
- Bu, R.; Zhao, W.H.; Liang, R. Downregulation of ATP8B2 in atherosclerosis exacerbates foam cell-like pathological changes via impairing lysosomal membrane fusion. Mol Biol Rep. 2025, 52(1). [Google Scholar] [CrossRef]
- Miao, X.; Pan, R.; Chang, F. Down-regulation of ATP8B2 in Foam Cells Inhibits Autophagic Flux and ox-LDL Degradation in Atherosclerosis. Cell Biochem Biophys. 2025, 83(3), 3451–63. [Google Scholar] [CrossRef]
- Li, S.; Gao, G.; Wu, F.; Liu, D.; Zhao, H.; Ke, J.; et al. Programmed cell death protein 4 deficiency suppresses foam cell formation by activating autophagy in advanced glycation end-product low-density lipoprotein-induced macrophages. J Cell Biochem. 2019, 120(5), 7689–700. [Google Scholar] [CrossRef]
- Zhou, F.; Liu, D.; Ning, H.; Yu, X.; Guan, X. The roles of p62/SQSTM1 on regulation of matrix metalloproteinase-9 gene expression in response to oxLDL in atherosclerosis. Biochem Biophys Res Commun. 2016, 472(3), 451–8. [Google Scholar] [CrossRef]
- Ning, H.; Liu, D.; Yu, X.; Guan, X. Oxidized low-density lipoprotein-induced p62/SQSTM1 accumulation in THP-1-derived macrophages promotes IL-18 secretion and cell death. Exp Ther Med. 2017, 14(6), 5417–23. [Google Scholar] [CrossRef]
- Wang, Y.M.; Tang, H.; Tang, Y.J.; Liu, J.; Yin, Y.F.; Tang, Y.L.; et al. ASIC1/RIP1 accelerates atherosclerosis via disrupting lipophagy. J Adv Res. 2024, 63, 195–206. [Google Scholar] [CrossRef]
- Liu, Q.; Li, F.; Hu, S.; Ding, N.; Ma, F.; Hao, Y.; et al. Pyruvate dehydrogenase alleviates macrophage autophagy in Hcy-induced ApoE–/– mice. Acta Biochim Biophys Sin. 2025, 57(12), 2022–33. [Google Scholar] [CrossRef]
- He, J.; Zhang, G.; Pang, Q.; Yu, C.; Xiong, J.; Zhu, J.; et al. SIRT6 reduces macrophage foam cell formation by inducing autophagy and cholesterol efflux under ox-LDL condition. FEBS J. 2017, 284(9), 1324–37. [Google Scholar] [CrossRef]
- Yang, S.; He, J.; Li, X.; Liu, H.; Zhao, J.; Liu, M. Hydrogen attenuated oxidized low-density lipoprotein-induced inflammation through the stimulation of autophagy via sirtuin 1. Exp Ther Med. 2018, 16(5), 4042–8. [Google Scholar] [CrossRef]
- Zeng, M.; Yang, Y.; Wang, Z.; Zhao, X.; Zhu, D.; Wang, M.; et al. CTRP9 prevents atherosclerosis progression through changing autophagic status of macrophages by activating USP22 mediated-de-ubiquitination on Sirt1 in vitro. Mol Cell Endocrinol 2024, 584, 112161. [Google Scholar] [CrossRef]
- Yuan, X.M.; Sultana, N.; Siraj, N.; Ward, L.J.; Ghafouri, B.; Li, W. Autophagy Induction Protects Against 7-Oxysterol-induced Cell Death via Lysosomal Pathway and Oxidative Stress. J Cell Death 2016, 9, 1–7. [Google Scholar] [CrossRef]
- Liu, B.X.; Zhang, B.C.; Guo, R.; Li, S.; Xu, Y.W. Enhancement in efferocytosis of oxidized low-density lipoprotein-induced apoptotic RAW264.7 cells through Sirt1-mediated autophagy. Int J Mol Med. 2014, 33(3), 523–33. [Google Scholar] [CrossRef]
- Hu, X.; Li, Y.; Chen, Q.; wang, T.; Ma, L.; Zhang, W.; et al. Sialic acids promote macrophage M1 polarization and atherosclerosis by upregulating ROS and autophagy blockage. Int Immunopharmacol 2023, 120. [Google Scholar] [CrossRef]
- Ackermann, K.; Bonaterra, G.A.; Kinscherf, R.; Schwarz, A. Growth differentiation factor-15 regulates oxLDL-induced lipid homeostasis and autophagy in human macrophages. Atherosclerosis 2019, 281, 128–36. [Google Scholar] [CrossRef]
- Jeong, S.J.; Kim, S.; Park, J.G.; Jung, I.H.; Lee, M.N.; Jeon, S.; et al. Prdx1 (peroxiredoxin 1) deficiency reduces cholesterol efflux via impaired macrophage lipophagic flux. Autophagy 2018, 14(1), 120–33. [Google Scholar] [CrossRef]
- Tao, J.; Yang, P.; Xie, L.; Pu, Y.; Guo, J.; Jiao, J.; et al. Gastrodin induces lysosomal biogenesis and autophagy to prevent the formation of foam cells via AMPK-FoxO1-TFEB signalling axis. J Cell Mol Med. 2021, 25(12), 5769–81. [Google Scholar] [CrossRef]
- Liang, X.; Wang, C.; Sun, Y.; Song, W.; Lin, J.; Li, J.; et al. p62/mTOR/LXRα pathway inhibits cholesterol efflux mediated by ABCA1 and ABCG1 during autophagy blockage. Biochem Biophys Res Commun. 2019, 514(4), 1093–100. [Google Scholar] [CrossRef] [PubMed]
- Wang, X.; Li, L.; Niu, X.; Dang, X.; Li, P.; Qu, L.; et al. mTOR enhances foam cell formation by suppressing the autophagy pathway. DNA Cell Biol. 2014, 33(4), 198–204. [Google Scholar] [CrossRef]
- Verheye, S.; Martinet, W.; Kockx, M.M.; Knaapen, M.W.M.; Salu, K.; Timmermans, J.P.; et al. Selective Clearance of Macrophages in Atherosclerotic Plaques by Autophagy. J Am Coll Cardiol. 2007, 49(6), 706–15. [Google Scholar] [CrossRef]
- Zhang, L.; Liu, Q.; Zhang, H.; Wang, X.D.; Chen, S.Y.; Yang, Y.; et al. C1q/TNF-Related Protein 9 Inhibits THP-1 Macrophage Foam Cell Formation by Enhancing Autophagy. J Cardiovasc Pharmacol. 2018, 72(4), 167–75. [Google Scholar] [CrossRef]
- Zahid, M.K.; Rogowski, M.; Ponce, C.; Choudhury, M.; Moustaïd-Moussa, N.; Rahman, S.M. CCAAT/enhancer-binding protein beta (C/EBPβ) knockdown reduces inflammation, ER stress, and apoptosis, and promotes autophagy in oxLDL-treated RAW264.7 macrophage cells. Mol Cell Biochem. 2020, 463(1-2), 211–23. [Google Scholar] [CrossRef]
- Zhong, S.; Li, L.; Zhang, Y.L.; Zhang, L.; Lu, J.; Guo, S.; et al. Acetaldehyde dehydrogenase 2 interactions with LDLR and AMPK regulate foam cell formation. J Clin Invest. 2019, 129(1), 252–67. [Google Scholar] [CrossRef]
- LeBlond, N.D.; Nunes, J.R.C.; Smith, T.K.T.; O’Dwyer, C.; Robichaud, S.; Gadde, S.; et al. Foam cell induction activates ampk but uncouples its regulation of autophagy and lysosomal homeostasis. Int J Mol Sci. 2020, 21(23). [Google Scholar] [CrossRef]
- Mei, S.; Gu, H.; Ward, A.; Yang, X.; Guo, H.; Kahe, K.; et al. p38 mitogen-activated protein kinase (MAPK) promotes cholesterol ester accumulation in macrophages through inhibition of macroautophagy. J Biol Chem. 2012, 287(15), 11761–8. [Google Scholar] [CrossRef]
- Fan, S.; Gao, H.; Nieuwhof, W.; Mulder, T.; Fumelli, B.F.M.; Li, R.; et al. Quantum sensing of free radicals in macrophages reveals early autophagy-lysosome regulation in an atherosclerosis cell model. Redox Biol. 2025, 87. [Google Scholar] [CrossRef]
- Takeda-Watanabe, A.; Kitada, M.; Kanasaki, K.; Koya, D. SIRT1 inactivation induces inflammation through the dysregulation of autophagy in human THP-1 cells. Biochem Biophys Res Commun. 2012, 427(1), 191–6. [Google Scholar] [CrossRef]
- de Meyer, I.; Martinet, W.; Schrijvers, D.M.; Timmermans, J.P.; Bult, H.; De Meyer, G.R.Y. Toll-like receptor 7 stimulation by imiquimod induces macrophage autophagy and inflammation in atherosclerotic plaques. Basic Res Cardiol. 2012, 107(3). [Google Scholar] [CrossRef]
- Kim, S.J.; Lee, W.J.; Cho, K.J. P62 links the autophagy pathway and the ubiquitin–proteasome system in endothelial cells during atherosclerosis. Int J Mol Sci. 2021, 22(15). [Google Scholar] [CrossRef]
- Zhou, Z.; Zhu, X.; Yin, R.; Liu, T.; Yang, S.; Zhou, L.; et al. K63 ubiquitin chains target NLRP3 inflammasome for autophagic degradation in ox-LDL-stimulated THP-1 macrophages. Aging (Albany NY) 2020, 12(2), 1747–59. [Google Scholar] [CrossRef]
- Liu, J.; Wang, C.; Li, J.; Yu, Y.; Liu, Y.; Liu, H.; et al. Autophagy blockage promotes the pyroptosis of ox-LDL-treated macrophages by modulating the p62/Nrf2/ARE axis. J Physiol Biochem. 2021, 77(3), 419–29. [Google Scholar] [CrossRef]
- Sano, M.; Maejima, Y.; Nakagama, S.; Shiheido-Watanabe, Y.; Tamura, N.; Hirao, K.; et al. Neutrophil extracellular traps-mediated Beclin-1 suppression aggravates atherosclerosis by inhibiting macrophage autophagy. Front Cell Dev Biol. 2022, 10, 876147. [Google Scholar] [CrossRef]
- Zeng, S.; Wen, Y.; Yu, C. Desialylation of ATG5 by sialidase (NEU1) promotes macrophages autophagy and exacerbates inflammation under hypoxia. Cell Signal. 2023, 112, 110927. [Google Scholar] [CrossRef]
- Folco, E.J.; Sukhova, G.K.; Quillard, T.; Libby, P. Moderate hypoxia potentiates interleukin-1β production in activated human macrophages. Circ Res. 2014, 115(10), 875–83. [Google Scholar] [CrossRef]
- Wai, K.W.; Tang, Y.Q.; Foo, J.B.; Hing, B.H.; Yap, W.H. Nrf2 modulates macrophage foam cells senescence and autophagy activation. Toxicol Mechan Methods 2025. [Google Scholar] [CrossRef]
- Tan, Z.; Ren, H.; Liu, Y.; Yang, H.; Luo, Q.; Deng, X. KLF2 alleviates endothelial cell injury and inhibits the formation of THP-1 macrophage-derived foam cells by activating Nrf2 and enhancing autophagy. Exp Ther Med. 2022, 24(6), 737. [Google Scholar] [CrossRef]
- Liang, X.; Hou, X.; Yang, Y.; Liu, H.; Guo, R.; Yang, Z.; et al. The feedback loop of “EMMPRIN/NF-κB” worsens atherosclerotic plaque via suppressing autophagy in macrophage. J Mol Cell Cardiol. 2018, 114, 129–40. [Google Scholar] [CrossRef]
- Zhuo, X.Z.; Wu, Y.; Yang, Y.J.; Gao, L.; Qiao, X.R.; Chen, T. Knockdown of LSD1 meliorates Ox-LDL-stimulated NLRP3 activation and inflammation by promoting autophagy via SESN2-mesiated PI3K/Akt/mTOR signaling pathway. Life Sci. 2019, 233. [Google Scholar] [CrossRef]
- Hu, G.; Yuan, Z.; Wang, J. Autophagy inhibition and ferroptosis activation during atherosclerosis: Hypoxia-inducible factor 1α inhibitor PX-478 alleviates atherosclerosis by inducing autophagy and suppressing ferroptosis in macrophages. Biomed Pharmacother. 2023, 161, 114333. [Google Scholar] [CrossRef]
- Peng, Q.; Liu, H.; Luo, Z.; Zhao, H.; Wang, X.; Guan, X. Effect of autophagy on ferroptosis in foam cells via Nrf2. Mol Cell Biochem. 2022, 477(5), 1597–606. [Google Scholar] [CrossRef]
- Yu, W.; Liu, W.; Xie, D.; Wang, Q.; Xu, C.; Zhao, H.; et al. High Level of Uric Acid Promotes Atherosclerosis by Targeting NRF2-Mediated Autophagy Dysfunction and Ferroptosis. Oxid Med Cell Longev. 2022, 2022, 9304383. [Google Scholar] [CrossRef]
- Ding, Z.; Liu, S.; Wang, X.; Dai, Y.; Khaidakov, M.; Deng, X.; et al. LOX-1, mtDNA damage, and NLRP3 inflammasome activation inmacrophages: Implications in atherogenesis. Cardiovasc Res. 2014, 103(4), 619–28. [Google Scholar] [CrossRef] [PubMed]
- Xiao, Q.Q.; Che, X.Y.; Cai, B.; Tao, Z.Y.; Zhang, H.Y.; Shao, Q.; et al. Macrophage autophagy regulates mitochondria-mediated apoptosis and inhibits necrotic core formation in vulnerable plaques. J Cell Mol Med. 2020, 24(1), 260–75. [Google Scholar] [CrossRef] [PubMed]
- Shao, W.; Wang, S.; Wang, X.; Yao, L.; Yuan, X.; Huang, D.; et al. miRNA-29a inhibits atherosclerotic plaque formation by mediating macrophage autophagy via PI3K/AKT/mTOR pathway. Aging (Albany NY) 2022, 14(5), 2418–31. [Google Scholar] [CrossRef] [PubMed]
- Wang, C.; Yang, W.; Liang, X.; Song, W.; Lin, J.; Sun, Y.; et al. MicroRNA-761 modulates foam cell formation and inflammation through autophagy in the progression of atherosclerosis. Mol Cell Biochem. 2020, 474(1-2), 135–46. [Google Scholar] [CrossRef]
- Ni, D.; Lei, C.; Liu, M.; Peng, J.; Yi, G.; Mo, Z. Cell death in atherosclerosis. Cell Cycle 2024, 23(5), 495–518. [Google Scholar] [CrossRef]
- Xu, Y.; Zhang, Y.; Yu, W. Nano-therapeutics targeting the macrophage-based microenvironment in the treatment of atherosclerosis. J Transl Med. 2025, 23(1), 1171. [Google Scholar] [CrossRef]
- Wang, Z.; Li, X.; Moura, A.K.; Hu, J.Z.; Wang, Y.T.; Zhang, Y. Lysosome Functions in Atherosclerosis: A Potential Therapeutic Target. Cells 2025, 14(3). [Google Scholar] [CrossRef] [PubMed]
- Zhou, G.; Liu, Y.; Wu, H.; Zhang, D.; Yang, Q.; Li, Y. Research Progress on Histone Deacetylases Regulating Programmed Cell Death in Atherosclerosis. J Cardiovasc Transl Res. 2024, 17(2), 308–21. [Google Scholar] [CrossRef] [PubMed]
- Zhang, Y.; Wang, W.; Li, X.; Li, X.; Zheng, L. Transient receptor potential channels as key regulators of cell death in atherosclerosis. Front Immunol. 2025, 16, 1661805. [Google Scholar] [CrossRef] [PubMed]
- Yun, H.R.; Singh, M.K.; Han, S.; Ranbhise, J.S.; Ha, J.; Kim, S.S.; et al. Roles of Autophagy and Oxidative Stress in Cardiovascular Disease. Antioxidants (Basel) 2025, 14(10). [Google Scholar] [CrossRef]
- Jeong, S.J.; Zhang, X.; Rodriguez-Velez, A.; Evans, T.D.; Razani, B. P62/SQSTM1 and Selective Autophagy in Cardiometabolic Diseases. Antioxidants and Redox Signaling 2019, 31(6), 458–71. [Google Scholar] [CrossRef]
- Lin, L.; Zhang, M.X.; Zhang, L.; Zhang, D.; Li, C.; Li, Y.L. Autophagy, Pyroptosis, and Ferroptosis: New Regulatory Mechanisms for Atherosclerosis. In Frontiers in Cell and Developmental Biology; 2021. [Google Scholar]
- Liu, H.; Cao, Y.; Tong, T.; Shi, J.; Zhang, Y.; Yang, Y.; et al. Autophagy in atherosclerosis: a phenomenon found in human carotid atherosclerotic plaques. Chinese Medical Journal. 2015, 128(1), 69–74. [Google Scholar] [CrossRef]
- Jiang, T.; Harder, B.; De La Vega, M.R.; Wong, P.K.; Chapman, E.; Zhang, D.D. p62 links autophagy and Nrf2 signaling. Free Radical Biology and Medicine 2015, 88, 199–204. [Google Scholar] [CrossRef]
- Kerins, M.J.; Liu, P.; Tian, W.; Mannheim, W.; Zhang, D.D.; Ooi, A. Genome-wide CRISPR screen reveals autophagy disruption as the convergence mechanism that regulates the NRF2 transcription factor. Molecular and Cellular Biology 2019, 39(13), e00037–19. [Google Scholar] [CrossRef]
- Chistiakov, D.A.; Sobenin, I.A.; Orekhov, A.N.; Bobryshev, Y.V. Role of endoplasmic reticulum stress in atherosclerosis and diabetic macrovascular complications. BioMed research international 2014, 2014(1), 610140. [Google Scholar] [CrossRef]
- The UPR in atherosclerosis. In Seminars in immunopathology; Zhou, A.X., Tabas, I., Eds.; Springer, 2013. [Google Scholar]
- Yao, S.-T.; Qin, S.-C. The relationship of autophagy with endoplasmic reticulum stress and its role in pathogenesis, prevention and therapy of atherosclerosis. Sheng li xue bao:[Acta Physiologica Sinica] 2017, 69(4), 515–21. [Google Scholar]
- Zhan, S.; Guo, C.; Yan, H.; Zheng, G.; Yan, D. The multi-dimensional regulatory mechanism of Sirt6 in heart health: From cell death pathways to targeted therapy for cardiovascular diseases; Biochemical and Biophysical Research Communications, 2025; p. 782. [Google Scholar]
- Ni, L.; Yang, L.; Lin, Y. Recent progress of endoplasmic reticulum stress in the mechanism of atherosclerosis. In Frontiers in Cardiovascular Medicine; 2024. [Google Scholar]
- Zhou, J.; Tan, S.-H.; Codogno, P.; Shen, H.-M. Dual suppressive effect of MTORC1 on autophagy: tame the dragon by shackling both the head and the tail. Autophagy 2013, 9(5), 803–5. [Google Scholar] [CrossRef]
- Dossou, A.S.; Basu, A. The emerging roles of mTORC1 in macromanaging autophagy. Cancers 2019, 11(10), 1422. [Google Scholar] [CrossRef] [PubMed]
- Wang, X.; Li, L.; Niu, X.; Dang, X.; Li, P.; Qu, L.; et al. mTOR enhances foam cell formation by suppressing the autophagy pathway. DNA and cell biology 2014, 33(4), 198–204. [Google Scholar] [CrossRef] [PubMed]
- Zhu, L.; Liao, Y.; Jiang, B. Role of ROS and autophagy in the pathological process of atherosclerosis. J Physiol Biochem. 2024, 80(4), 743–56. [Google Scholar] [CrossRef] [PubMed]
- Li, Y.; Chen, Y. AMPK and autophagy; Biology and Diseases: Basic Science: Autophagy, 2019; pp. 85–108. [Google Scholar]
- Wang, S.; Li, H.; Yuan, M.; Fan, H.; Cai, Z. Role of AMPK in autophagy. Frontiers in physiology 2022, 13, 1015500. [Google Scholar] [CrossRef]
- Mihaylova, M.M.; Shaw, R.J. The AMPK signalling pathway coordinates cell growth, autophagy and metabolism. Nature cell biology 2011, 13(9), 1016–23. [Google Scholar] [CrossRef]
- Ge, Y.; Zhou, M.; Chen, C.; Wu, X.; Wang, X. Role of AMPK mediated pathways in autophagy and aging. Biochimie 2022, 195, 100–13. [Google Scholar] [CrossRef]
- Linton, M.F.; Moslehi, J.J.; Babaev, V.R. Akt signaling in macrophage polarization, survival, and atherosclerosis. International journal of molecular sciences 2019, 20(11), 2703. [Google Scholar] [CrossRef]
- Kma, L.; Baruah, T.J. The interplay of ROS and the PI3K/Akt pathway in autophagy regulation. Biotechnology and applied biochemistry 2022, 69(1), 248–64. [Google Scholar] [CrossRef]
- Yu, X.; Long, Y.C.; Shen, H.-M. Differential regulatory functions of three classes of phosphatidylinositol and phosphoinositide 3-kinases in autophagy. Autophagy 2015, 11(10), 1711–28. [Google Scholar] [CrossRef]
- Napolitano, G.; Ballabio, A. TFEB at a glance. Journal of cell science 2016, 129(13), 2475–81. [Google Scholar] [CrossRef] [PubMed]
- Settembre, C.; Di Malta, C.; Polito, V.A.; Arencibia, M.G.; Vetrini, F.; Erdin, S.; et al. TFEB links autophagy to lysosomal biogenesis. science 2011, 332(6036), 1429–33. [Google Scholar] [CrossRef] [PubMed]
- Emanuel, R.; Sergin, I.; Bhattacharya, S.; Turner, J.N.; Epelman, S.; Settembre, C.; et al. Induction of lysosomal biogenesis in atherosclerotic macrophages can rescue lipid-induced lysosomal dysfunction and downstream sequelae. Arteriosclerosis, thrombosis, and vascular biology 2014, 34(9), 1942–52. [Google Scholar] [CrossRef] [PubMed]
- Gan, Y.M.; Qin, N.S.; Ouyang, J.; Tang, Y.M.; Qin, J.H.; Wei, S.M.; et al. Cross-regulation of autophagy and pyroptosis: a new perspective on the inflammatory microenvironment of atherosclerosis. Apoptosis 2026, 31(1), 18. [Google Scholar] [CrossRef]
- Chen, J.; Zhao, S.; Su, Y.; Xu, L.; Chen, W.; Zhang, Z. Nanomedicine-Enabled/Augmented Cell Autophagy to Mitigate and Boycott the Development and Progression of Atherosclerotic Plaque. Int J Nanomed. 2026, 21, 1–20. [Google Scholar] [CrossRef]
- Li, P.H.; Jiang, W. A New Insight on Atherosclerosis Mechanism and Lipid-Lowering Drugs. REVIEWS IN CARDIOVASCULAR MEDICINE 2025, 26(3). [Google Scholar] [CrossRef]
- Kang, R.; Tang, D. Autophagy and ferroptosis—what is the connection? Current pathobiology reports 2017, 5, 153–9. [Google Scholar] [CrossRef]
- Liu, J.; Guo, Z.-N.; Yan, X.-L.; Huang, S.; Ren, J.-X.; Luo, Y.; et al. Crosstalk between autophagy and ferroptosis and its putative role in ischemic stroke. Frontiers in Cellular Neuroscience 2020, 14, 577403. [Google Scholar] [CrossRef]
- Zhang, Y.; Yang, J.; Ouyang, C.; Meng, N. The association between ferroptosis and autophagy in cardiovascular diseases. Cell Biochem Funct. 2024, 42(2). [Google Scholar] [CrossRef]
- Xiao, Q.; Che, X.; Cai, B.; Tao, Z.; Zhang, H.; Shao, Q.; et al. Macrophage autophagy regulates mitochondria-mediated apoptosis and inhibits necrotic core formation in vulnerable plaques. Journal of cellular and molecular medicine 2020, 24(1), 260–75. [Google Scholar] [CrossRef]
- Ni, D.; Mo, Z.; Yi, G. Recent insights into atherosclerotic plaque cell autophagy. Experimental Biology and Medicine 2021, 246(24), 2553–8. [Google Scholar] [CrossRef]
- Luo, Y.; Lu, S.; Zhou, P.; Ai, Q.-D.; Sun, G.-B.; Sun, X.-B. Autophagy: an exposing therapeutic target in atherosclerosis. Journal of cardiovascular pharmacology 2016, 67(3), 266–74. [Google Scholar] [CrossRef] [PubMed]
- Owaki, R.; Aoki, H.; Toriuchi, K.; Inoue, Y.; Hayashi, H.; Takeshita, S.; et al. AMPK activators suppress cholesterol accumulation in macrophages via suppression of the mTOR pathway. Experimental Cell Research 2023, 432(1), 113784. [Google Scholar] [CrossRef] [PubMed]
- Martinet, W.; Verheye, S.; De Meyer, G.R. Everolimus-induced mTOR inhibition selectively depletes macrophages in atherosclerotic plaques by autophagy. Autophagy 2007, 3(3), 241–4. [Google Scholar] [CrossRef] [PubMed]
- Luo, Z.; Xu, W.; Ma, S.; Qiao, H.; Gao, L.; Zhang, R.; et al. Moderate autophagy inhibits vascular smooth muscle cell senescence to stabilize progressed atherosclerotic plaque via the mTORC1/ULK1/ATG13 signal pathway. Oxidative Medicine and Cellular Longevity 2017, 2017(1), 3018190. [Google Scholar] [CrossRef]
- Evans, T.D.; Jeong, S.-J.; Zhang, X.; Sergin, I.; Razani, B. TFEB and trehalose drive the macrophage autophagy-lysosome system to protect against atherosclerosis. Autophagy 2018, 14(4), 724–6. [Google Scholar] [CrossRef]
- Qiao, L.; Ma, J.; Zhang, Z.; Sui, W.; Zhai, C.; Xu, D.; et al. Deficient Chaperone-Mediated Autophagy Promotes Inflammation and Atherosclerosis. Circ Res. 2021, 129(12), 1141–57. [Google Scholar] [CrossRef]
- Qiao, L.; Wang, H.F.; Xiang, L.; Ma, J.; Zhu, Q.; Xu, D.; et al. Deficient Chaperone-Mediated Autophagy Promotes Lipid Accumulation in Macrophage. J Cardiovasc Transl Res. 2021, 14(4), 661–9. [Google Scholar] [CrossRef]
- Qiao, L.; Wang, H-f; Xiang, L.; Ma, J.; Zhu, Q.; Xu, D.; et al. Deficient chaperone-mediated autophagy promotes lipid accumulation in macrophage. Journal of cardiovascular translational research 2021, 14, 661–9. [Google Scholar] [CrossRef]
- Wang, S.; Yuan, R.; Liu, M.; Zhang, Y.; Jia, B.; Ruan, J.; et al. Targeting autophagy in atherosclerosis: Advances and therapeutic potential of natural bioactive compounds from herbal medicines and natural products. Biomedicine & Pharmacotherapy 2022, 155, 113712. [Google Scholar] [CrossRef]
- Schrijvers, D.M.; De Meyer, G.R.; Martinet, W. Autophagy in atherosclerosis: a potential drug target for plaque stabilization. Arteriosclerosis, thrombosis, and vascular biology 2011, 31(12), 2787–91. [Google Scholar] [CrossRef] [PubMed]
- Grootaert, M.O.; da Costa Martins, P.A.; Bitsch, N.; Pintelon, I.; De Meyer, G.R.; Martinet, W.; et al. Defective autophagy in vascular smooth muscle cells accelerates senescence and promotes neointima formation and atherogenesis. Autophagy 2015, 11(11), 2014–32. [Google Scholar] [CrossRef] [PubMed]
- Kurdi, A.; De Meyer, G.R.; Martinet, W. Potential therapeutic effects of mTOR inhibition in atherosclerosis. British journal of clinical pharmacology 2016, 82(5), 1267–79. [Google Scholar] [CrossRef]
- Ma, A.; Wang, J.; Yang, L.; An, Y.; Zhu, H. AMPK activation enhances the anti-atherogenic effects of high density lipoproteins in apoE−/− mice. Journal of lipid research 2017, 58(8), 1536–47. [Google Scholar] [CrossRef]
- Zhang, W.; Xu, W.; Chen, W.; Zhou, Q. Interplay of autophagy inducer rapamycin and proteasome inhibitor MG132 in reduction of foam cell formation and inflammatory cytokine expression. Cell Transplantation 2018, 27(8), 1235–48. [Google Scholar] [CrossRef]
- Robichaud, S.; Rochon, V.; Emerton, C.; Ouimet, M.I. Autophagy Activation Promotes Atherosclerosis Regression. Arteriosclerosis, Thrombosis, and Vascular Biology 2023, 43 (Suppl_1), A704–A. [Google Scholar] [CrossRef]
- Sergin, I.; Evans, T.D.; Zhang, X.; Bhattacharya, S.; Stokes, C.J.; Song, E.; et al. Exploiting macrophage autophagy-lysosomal biogenesis as a therapy for atherosclerosis. Nature communications 2017, 8(1), 15750. [Google Scholar] [CrossRef]
- Zhang, Y.; Cao, X.; Zhu, W.; Liu, Z.; Liu, H.; Zhou, Y.; et al. Resveratrol enhances autophagic flux and promotes ox-LDL degradation in HUVECs via upregulation of SIRT1. Oxidative medicine and cellular longevity 2016, 2016(1), 7589813. [Google Scholar] [CrossRef]
- Feng, T.; Liu, P.; Xu, Y.; Si, S. SIRT1 activators and their effects on atherosclerosis progression. Cardiol Res Cardiovasc Med 2018. [Google Scholar] [CrossRef]
- Kitada, M.; Ogura, Y.; Koya, D. The protective role of Sirt1 in vascular tissue: its relationship to vascular aging and atherosclerosis. Aging (Albany NY) 2016, 8(10), 2290. [Google Scholar] [CrossRef]
- Mauthe, M.; Orhon, I.; Rocchi, C.; Zhou, X.; Luhr, M.; Hijlkema, K.-J.; et al. Chloroquine inhibits autophagic flux by decreasing autophagosome-lysosome fusion. Autophagy 2018, 14(8), 1435–55. [Google Scholar] [CrossRef]
- Zhao, X.; Li, K.; Liu, Q.; Jiang, M.; Luo, Y. Oral Exposure To SiO2 Nanoparticles Promotes Foam Cell Areas in Aortic Sinus of ApoE-/- Mice and Epigallocatechin Gallate Attenuates the Effects via Metabolic Restoration. Cardiovasc Toxicol. 2025, 25(12), 1837–49. [Google Scholar] [CrossRef] [PubMed]
- Tang, C.; Wang, H.; Guo, L.; Cui, Y.; Zou, C.; Hu, J.; et al. Multifunctional Nanomedicine for Targeted Atherosclerosis Therapy: Activating Plaque Clearance Cascade and Suppressing Inflammation. ACS Nano 2025, 19(3), 3339–61. [Google Scholar] [CrossRef] [PubMed]
- Ouimet, M.; Franklin, V.; Mak, E.; Liao, X.; Tabas, I.; Marcel, Y.L. Autophagy regulates cholesterol efflux from macrophage foam cells via lysosomal acid lipase. Cell metabolism 2011, 13(6), 655–67. [Google Scholar] [CrossRef] [PubMed]
- Maiuri, M.C.; Grassia, G.; Platt, A.M.; Carnuccio, R.; Ialenti, A.; Maffia, P. Macrophage autophagy in atherosclerosis. Mediators of inflammation 2013, 2013(1), 584715. [Google Scholar] [CrossRef]
- Li, W.; Sultana, N.; Siraj, N.; Ward, L.J.; Pawlik, M.; Levy, E.; et al. Autophagy dysfunction and regulatory cystatin C in macrophage death of atherosclerosis. J Cell Mol Med. 2016, 20(9), 1664–72. [Google Scholar] [CrossRef]
- Marchio, P.; Guerra-Ojeda, S.; Vila, J.M.; Aldasoro, M.; Victor, V.M.; Mauricio, M.D. Targeting early atherosclerosis: a focus on oxidative stress and inflammation. Oxidative medicine and cellular longevity 2019, 2019(1), 8563845. [Google Scholar] [CrossRef]
- Madaudo, C.; Coppola, G.; Parlati, A.L.M.; Corrado, E. Discovering Inflammation in Atherosclerosis: Insights from Pathogenic Pathways to Clinical Practice. International Journal of Molecular Sciences 2024, 25(11), 6016. [Google Scholar] [CrossRef]
- Xu, W.Q.; Yang, Y.X.; Ma, Y.; Deng, S.A.; Zhang, Z.Z.; Huang, X.F.; et al. A bibliometric analysis of autophagy research in atherosclerosis over the past decade. Medicine (Baltimore) 2026, 105(3), e46969. [Google Scholar] [CrossRef]
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2026 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).




