Submitted:
14 April 2026
Posted:
15 April 2026
You are already at the latest version
Abstract
Keywords:
1. Introduction
2. Materials and Methods
2.1. Study Design and Population
2.2. Data Collection and Procedures
2.3. Laboratory Analysis
2.4. Statistical Analysis
3. Results
3.1. Cohort Characteristics
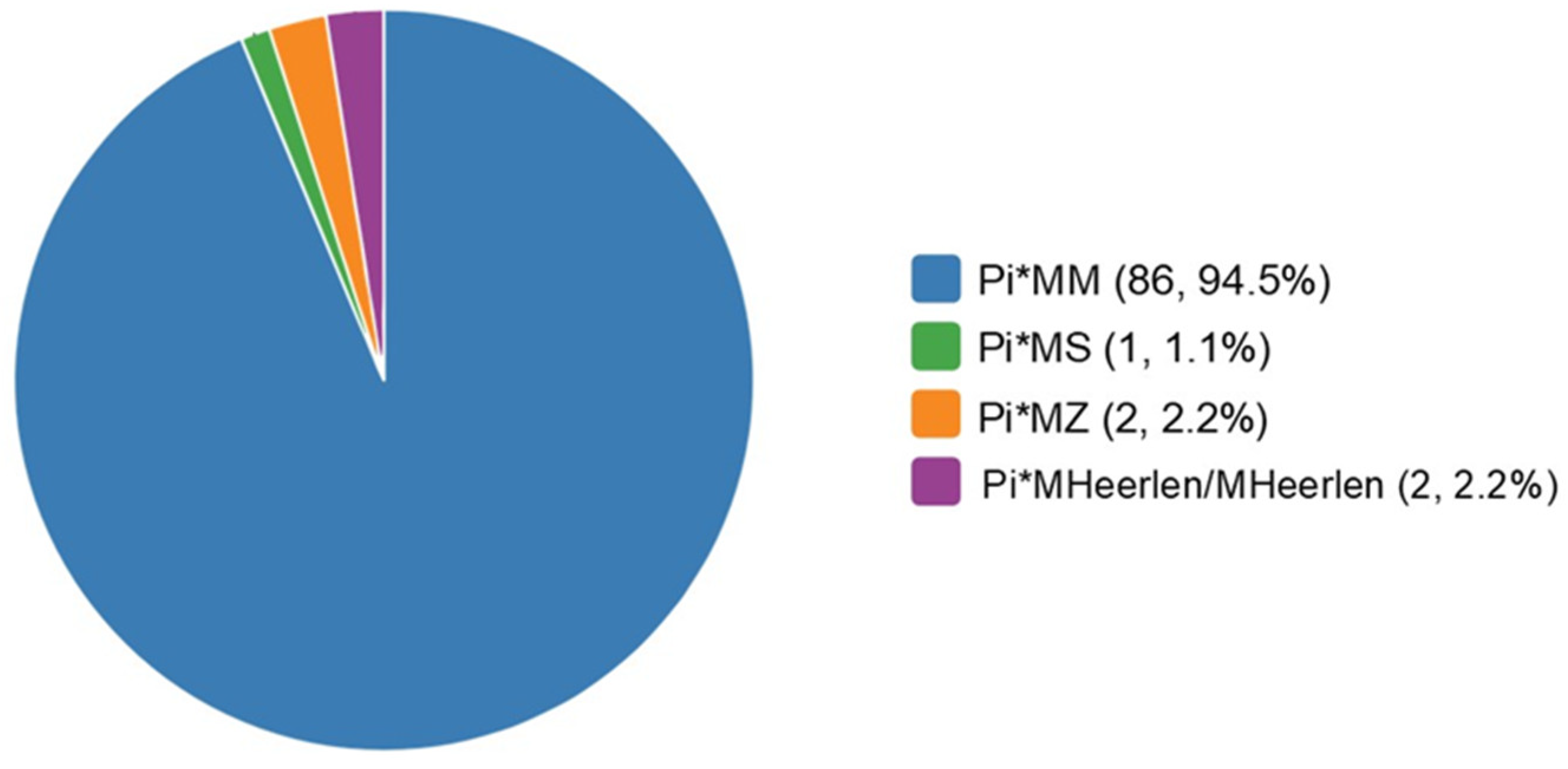
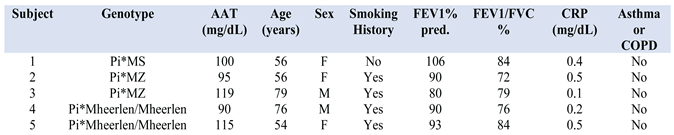
3.2. Genetic Screening Results
4. Discussion
5. Conclusions
Supplementary Materials
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Acknowledgments
Conflicts of Interest
Abbreviations
| AAT | Alpha-1 Antitrypsin |
| AATD | Alpha-1 Antitrypsin Deficiency |
| ATS | American Thoracic Society |
| COPD | Chronic Obstructive Pulmonary Disease |
| CRP | C-reactive protein |
| DNA | deoxyribonucleic acid |
| EARCO | European Alpha-1 Research Collaboration |
| ERS | European Respiratory Society |
| FEV1 | forced expiratory volume in 1 s |
| FVC | forced vital capacity |
| HCC | hepatocellular carcinoma |
| IEF | isoeletric focusing |
| IQR | interquartile range (first–third quartile) |
| IRCCS | Scientific Institute for Research, Hospitalization and Healthcare |
| Pi | Protease inhibitor |
| PiMM | normal AAT phenotype (M/M alleles) |
| PiMS | heterozygous AAT phenotype (M/S alleles) |
| PiZZ | severe AAT phenotype (Z/Z alleles) |
References
- Strnad, P.; McElvaney, N.G.; Lomas, D.A. Alpha1-Antitrypsin Deficiency. N. Engl. J. Med. 2020, 382, 1443–1455. [Google Scholar] [CrossRef] [PubMed]
- Greene, C.M.; Marciniak, S.J.; Teckman, J.; Ferrarotti, I.; Brantly, M.L.; Lomas, D.A. α1-Antitrypsin deficiency. Nat. Rev. Dis. Primers 2016, 2, 16051. [Google Scholar]
- Bouchecareilh, M. Alpha-1 antitrypsin deficiency-mediated liver toxicity: Why do some patients do poorly? What do we know so far? Chronic Obstr. Pulm. Dis. 2020, 7, 172–181. [Google Scholar] [CrossRef]
- Foil, K.E. Variants of SERPINA1 and the increasing complexity of testing for alpha-1 antitrypsin deficiency. Ther. Adv. Chronic Dis. 2021, 12. [Google Scholar] [CrossRef]
- Ferrarotti, I.; Ottaviani, S.; De Silvestri, A.; Corsico, A.G. Update on α1-antitrypsin deficiency. Breathe 2018, 14, e17–e24. [Google Scholar] [CrossRef]
- Stoller, J.K.; Aboussouan, L.S. A review of alpha1-antitrypsin deficiency. Am. J. Respir. Crit. Care Med. 2012, 185, 246–259. [Google Scholar] [CrossRef]
- Horváth, I.; Canotilho, M.; Chlumský, J.; Chorostowska-Wynimko, J.; Corda, L.; Derom, E.; Ficker, J.H.; Kneussl, M.; Miravitlles, M.; Sucena, M.; et al. Diagnosis and management of α1-antitrypsin deficiency in Europe: An expert survey. ERJ Open Res. 2019, 5, 00171-2018. [Google Scholar] [CrossRef] [PubMed]
- Miravitlles, M.; Dirksen, A.; Ferrarotti, I.; Koblizek, V.; Lange, P.; Mahadeva, R.; McElvaney, N.G.; Parr, D.; Piitulainen, E.; Roche, N.; et al. European Respiratory Society statement: Diagnosis and treatment of pulmonary disease in alpha1-antitrypsin deficiency. Eur. Respir. J. 2017, 50, 1700610. [Google Scholar] [PubMed]
- Chapman, K.R.; Burdon, J.G.; Piitulainen, E.; Sandhaus, R.A.; Seersholm, N.; Stocks, J.; Stoel, B.C.; Huang, L.; Zhang, Z.; Edelman, J.M.; et al. Intravenous augmentation treatment and lung density in severe alpha1 antitrypsin deficiency (RAPID): A randomised, double-blind, placebo-controlled trial. Lancet 2015, 386, 360–368. [Google Scholar] [CrossRef]
- McElvaney, N.G.; Burdon, J.; Holmes, M.; Glanville, A.; Wark, P.A.B.; Thompson, P.J.; Hernandez, P.; Chlumsky, J.; Teschler, H.; Ficker, J.H.; et al. Long-term efficacy and safety of α1 proteinase inhibitor treatment for emphysema caused by severe α1 antitrypsin deficiency: An open-label extension trial (RAPID-OLE). Lancet Respir. Med. 2017, 5, 51–60. [Google Scholar]
- McElvaney, G.N.; Al-Alawi, M.; Miravitlles, M. Clinical considerations in individuals with α1-antitrypsin PI*SZ genotype. Eur. Respir. J. 2020, 55, 1902410. [Google Scholar] [CrossRef]
- de Serres, F.J. Worldwide racial and ethnic distribution of alpha1-antitrypsin deficiency: Summary of an analysis of published genetic epidemiologic surveys. Chest 2002, 122, 1818–1829. [Google Scholar] [CrossRef]
- Greulich, T.; Altraja, A.; Barrecheguren, M.; Chlumský, J.; Chorostowska-Wynimko, J.; Corda, L.; Ferrarotti, I.; Giotaki, S.; Horváth, I.; Ilic, D.; et al. Protocol for the EARCO Registry: A pan-European observational study in patients with alpha1-antitrypsin deficiency. ERJ Open Res. 2020, 6, 00181-2019. [Google Scholar] [CrossRef]
- Luisetti, M.; Ferrarotti, I.; Corda, L.; Ottaviani, S.; Gatta, N.; Tinelli, C.; Bruletti, G.; Bertella, E.; Balestroni, G.; Confalonieri, M.; et al. Italian registry of patients with alpha-1 antitrypsin deficiency: General data and quality of life evaluation. COPD 2015, 12, 52–57. [Google Scholar] [CrossRef]
- Miravitlles, M.; Turner, A.M.; Torres-Duran, M.; Tanash, H.; Rodríguez-García, C.; López-Campos, J.L.; Chlumsky, J.; Guimaraes, C.; Rodríguez-Hermosa, J.L.; Corsico, A.; et al. Clinical and functional characteristics of individuals with alpha-1 antitrypsin deficiency: EARCO international registry. Respir. Res. 2022, 23, 352. [Google Scholar] [CrossRef] [PubMed]
- Pini, L.; Corda, L.; Malerba, M.; Montanelli, A.; Favagrossa, G.M.; Terzi, A.; Marcobruni, N.; Magoni, M.; Franchini, D.; Radaeli, A.; et al. Il deficit di alpha 1-antitripsina: Lo studio clinico di Brescia [Alpha 1-antitrypsin deficiency: The Brescia clinical study]. Recent. Prog. Med. 2000, 91, 352–361. [Google Scholar]
- Graham, B.L.; Steenbruggen, I.; Miller, M.R.; Barjaktarevic, I.Z.; Cooper, B.G.; Hall, G.L.; Hallstrand, T.S.; Helgeson, K.C.; Haynes, J.M.; Culver, B.H.; et al. Standardization of Spirometry 2019 Update. An Official American Thoracic Society and European Respiratory Society Technical Statement. Am. J. Respir. Crit. Care Med. 2019, 200, e70–e88. [Google Scholar] [CrossRef] [PubMed]
- Ottaviani, S.; Barzon, V.; Buxens, A.; Gorrini, M.; Pagani, B.; Belmonte, I.; Rodriguez-Frias, F.; Miravitlles, M.; Janciauskiene, S.; Luisetti, M.; et al. Molecular diagnosis of alpha1-antitrypsin deficiency: A new method based on Luminex technology. J. Clin. Lab. Anal. 2020, 34, e23272. [Google Scholar] [CrossRef]
- Lopez-Campos, J.L.; Rapun, N.; Czischke, K.; Barroso, J.; Rodriguez-Frias, F.; Belmonte, I.; Lara, B.; Gorrini, M.; Ferrarotti, I.; Ottaviani, S.; et al. Distribution of alpha1 antitrypsin rare alleles in six countries: Results from the Progenika diagnostic network. Hum. Genomics 2023, 17, 47. [Google Scholar] [CrossRef]
- De Serres, F.J.; Luisetti, M.; Ferrarotti, I.; Blanco, I.; Fernández-Bustillo, E. Alpha-1 antitrypsin deficiency in Italy: Regional differences of the PIS and PIZ deficiency alleles. Monaldi Arch. Chest Dis. 2005, 63, 133–141. [Google Scholar] [CrossRef]
- Seixas, S.; Marques, P.I. Known Mutations at the Cause of Alpha-1 Antitrypsin Deficiency an Updated Overview of SERPINA1 Variation Spectrum. Appl. Clin. Genet. 2021, 14, 173–194. [Google Scholar] [CrossRef] [PubMed]
- Poller, W.; Merklein, F.; Faber, J.P.; Weidinger, S. Molecular characterisation of the defective alpha 1-antitrypsin alleles PI Mwurzburg (Pro369Ser), Mheerlen (Pro369Leu), and Q0lisbon (Thr68Ile). Eur. J. Hum. Genet. 1999, 7, 321–331. [Google Scholar] [CrossRef] [PubMed]
- Ferrarotti, I.; Baccheschi, J.; Zorzetto, M.; Tinelli, C.; Campo, I.; Genzini, C.; Scabini, R.; Gorrini, M.; Mazzola, P.; Luisetti, M.; et al. Prevalence and phenotype of subjects carrying rare variants in the Italian registry for alpha1-antitrypsin deficiency. J. Med. Genet. 2005, 42, 282–287. [Google Scholar] [CrossRef] [PubMed]
- Meseeha, M.; Sankari, A.; Attia, M. Alpha-1 Antitrypsin Deficiency. In StatPearls; StatPearls Publishing: Treasure Island, FL, USA, 2024; Available online: https://www.ncbi.nlm.nih.gov/books/NBK442030/ (accessed on 2 April 2026).
| Variable | All Subjects (n = 91) |
|---|---|
| Age, years Male sex, n (%) Smoking history, n (%) FEV1, % predicted FEV1/FVC, % AAT, mg/dL CRP, mg/dL Asthma, n (%) COPD, n (%) |
61 (54–67) 34 (37.4) 61 (67.0) 96 (90–102) 78 (74–84) 120 (104–140) 0.2 (0.2–0.4) 2 (2.2) 0 (0) |
 |
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2026 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).




