Submitted:
13 April 2026
Posted:
14 April 2026
You are already at the latest version
Abstract
Keywords:
1. Introduction
2. Theoretical Foundation
2.1. Problem Categorization
- Static Covariate & Static Treatment The classic static ITE estimation problem in which baseline (pre-treatment) covariates are used to estimate the effect of a single fixed treatment. This is the fundamental scenario for answering questions, such as “Does Drug A work better than a placebo for patients given the baseline disease severity?”
- Time-Varying Covariate & Static Treatment A single treatment is administered, but the decision to treat (or not) is based on a sequence of covariates. For example, deciding whether and when to perform an organ transplant is a one-time decision based on past lab results and health vitals.
- Static Covariate & Time-Varying Treatment A sequence of treatments is administered based solely on baseline covariates, without adaptation to intermediate observations. Although uncommon in healthcare, this setting arises in industrial processes where, for example, a fixed sequence of temperature and pressure set points is predetermined from initial material properties and executed without adjustment during processing.
- Time-Varying Covariate & Time-Varying Treatment Both covariates and treatments are subject to change and may influence one another over time. This is the most common and complex scenario, such as the management of a patient in the Intensive Care Unit (ICU), where a physician repeatedly measures vital signs, uses them to administer a dose of a blood pressure drug, and then observes the new vital signs to decide on the next dose, with the intention of optimizing a downstream outcome such as 30-day survival.
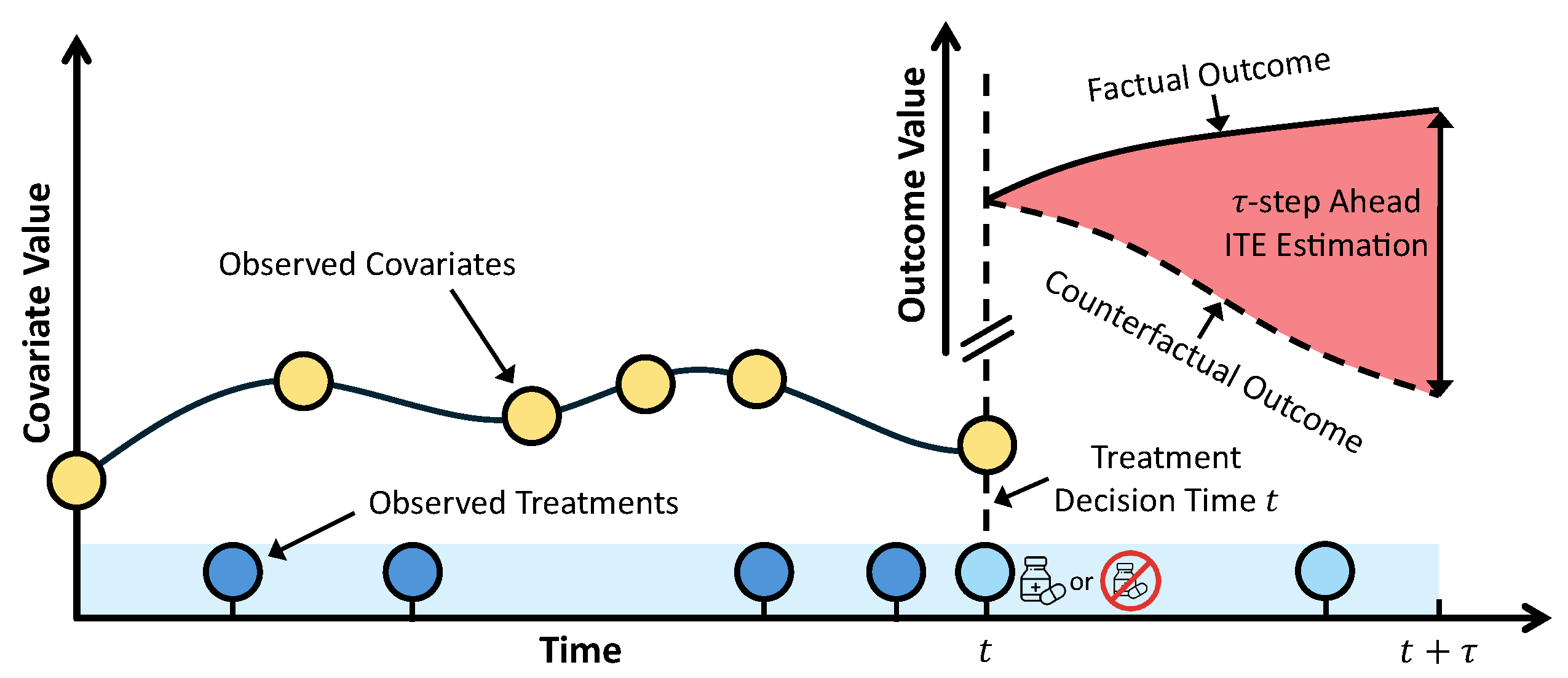
2.2. Formalization and Notation
- (Pre-treatment) Covariates: A -dimensional random vector of patient features at time k. The covariate space may comprise both continuous (e.g., vital signs, lab results) and discrete (e.g., diagnosis codes) components. Each covariate feature may be static (i.e., remaining constant across all time steps such as sex or ethnicity), or time-varying, changing in response to disease progression and prior treatments. A particular realized covariate vector is denoted .
- Treatment: The treatment assignment at time k is a random vector taking values in the treatment space . The treatment space may be discrete (e.g., a finite set of medication options), continuous (e.g., drug dosage), or multivariate when multiple treatments are administered concurrently. A realized treatment vector at time k is denoted by .
- Outcome : The outcome of interest, measured steps after the decision time t. The outcome space may be binary (e.g., for mortality) or continuous (e.g., for blood pressure). A realized outcome is denoted .
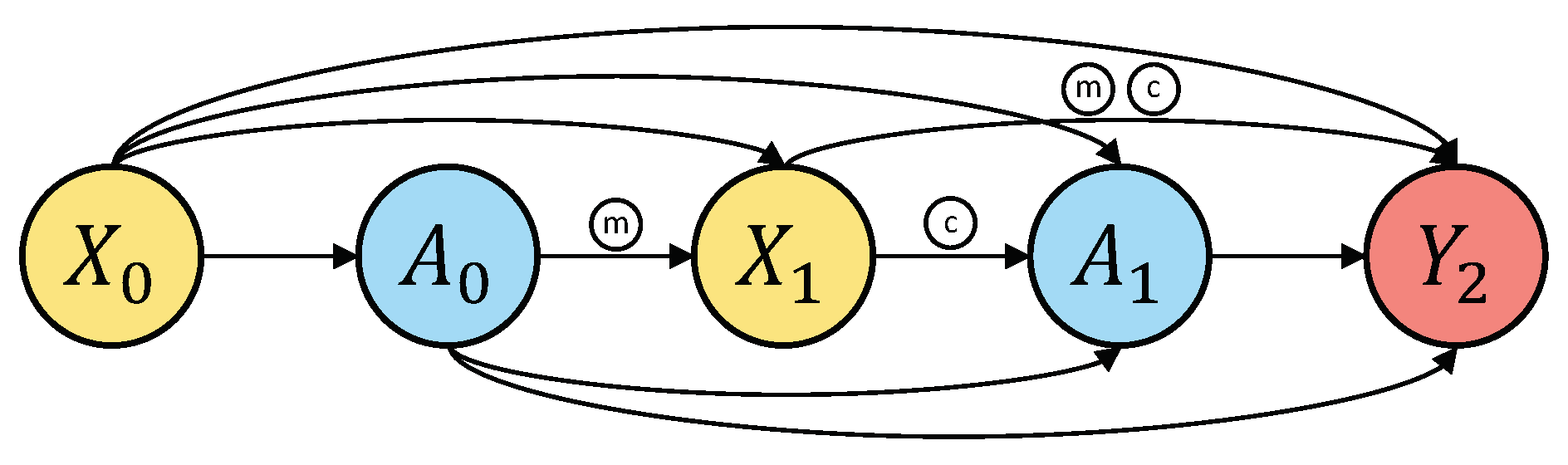
2.3. The Challenge of Time-Dependent Confounding
- Mediator: is caused by and, in turn, affects , forming the mediated path (). This is a valid component of the total causal effect of treatment on .
- Confounder: is a common cause of and , creating a spurious association between them through the path (), which does not reflect the causal effect of treatment .
2.4. Identifiability of Time-Varying ITE
-
Consistency: Let denote a fixed full treatment sequence, where is a particular realized past treatment sequence (a value of ) and is a prescribed future treatment sequence. For an individual, if the observed treatment sequence matches , then the observed outcome is equal to the corresponding potential outcome:This implies that there are no multiple versions of the treatment, i.e., consistency requires that the treatment definition be sufficiently precise so that technical variations do not change the potential outcome. In the static ITE estimation setting, the corresponding assumption is the Stable Unit Treatment Value Assumption (SUTVA) [23], which consists of two components: (i) consistency as defined above, and (ii) no interference which states that an individual’s potential outcomes do not depend on the treatment assignments of others. Both components remain necessary in the sequential setting. However, in modern deep learning literature, no interference is typically absorbed into the statistical setup rather than stated as a standalone assumption. Seminal works [8,24] explicitly list consistency as an assumption, but encode no interference implicitly by defining the dataset as a collection of N independent and identically distributed (i.i.d.) patient trajectories.
- Sequential Positivity: For each time step , every treatment value must be assignable with positive probability, conditional on any history with :or, when is continuous, . More intuitively, the effect of a given treatment sequence cannot be identified for histories where that sequence (or parts of it) is impossible under the observed data. If a particular treatment is never administered to patients with a history , then we cannot use the observed data to learn the counterfactual outcome under that action in that stratum.
-
Sequential Exchangeability (or Ignorability): At each time step from baseline to end of the treatment horizon , after conditioning on the history , treatment is independent of future potential outcomes under any treatment sequence. For the horizon- outcome, this can be written asThis assumption must hold throughout history , not only during the future treatment window . If unmeasured confounders influenced both treatment and covariates in any previous step , then the observed history itself depends on these confounders. Conditioning on is therefore insufficient to make treatment independent of potential outcomes, and the resulting treatment effect estimates will be biased.
3. From Theory to Practice
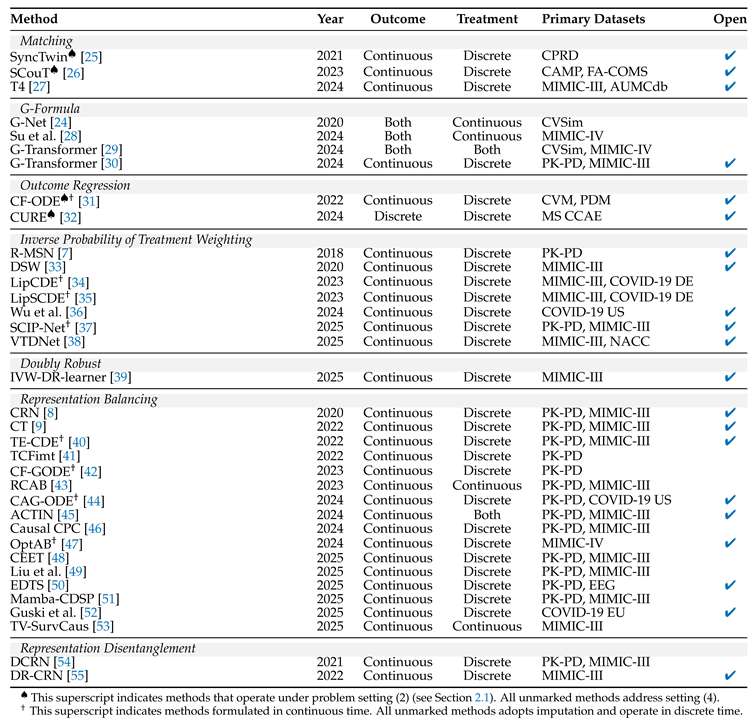 |
4. Time-Varying ITE Estimation Methodology
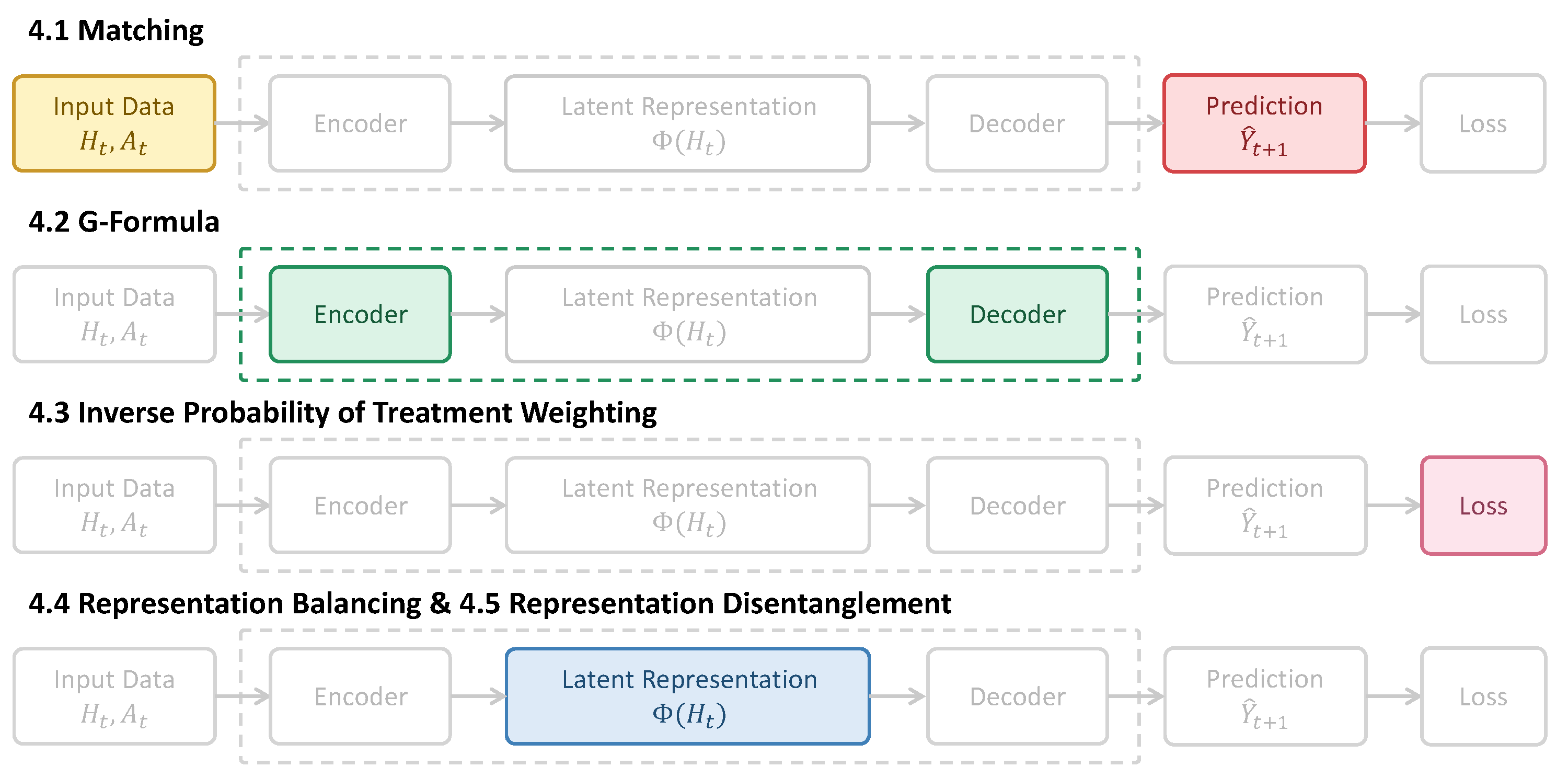
4.1. Matching
4.2. G-Formula
4.3. Inverse Probability of Treatment Weighting (IPTW)
4.4. Representation Balancing
4.4.1. Predictive Balancing
4.4.2. Distribution Alignment
4.5. Representation Disentanglement
5. Clinical Application Scenarios
6. Evaluation
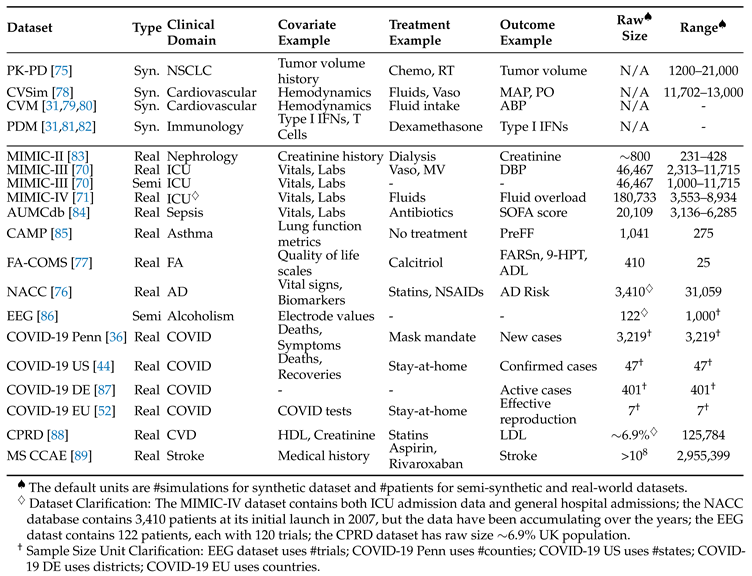 |
6.1. Datasets
6.1.1. Synthetic Datasets
6.1.2. Semi-Synthetic Datasets
- Extract Covariates: Real-world healthcare time series data are used to ensure that the complexity of the feature space is realistic. Vital signs and demographics of the patient are extracted from MIMIC-III to serve as time-varying and static covariates, respectively.
- Simulate Untreated Outcomes: The untreated outcome represents the outcome that would be observed if no treatment were administered. It is generated by combining a longitudinal trend that captures the patient’s endogenous progression over time, a covariate-driven component representing exogenous fluctuations caused by current physiological states, and additive noise .
- Simulate Treatment Vector: We introduce time-dependent confounding by modeling the conditional distribution of each treatment component based on the patient’s history. This creates a biased assignment mechanism in which the intensity or probability of the treatment depends on the severity of the past.where denotes the logistic sigmoid function, mapping the history representation to the probability interval . The realization is sampled from a distribution specific to the domain . For binary treatments (i.e., ): represents Bernoulli probabilities and .
- Calculate the Cumulative Treatment Effect: Treatments are modeled to have a delayed and decaying impact (e.g., inverse-square decay) rather than merely an immediate influence. The influence is scaled by the assignment probability to correlate treatment efficacy with patient severity.where is a decay function (e.g., inverse-square), and computes the influence of treatment over time u given the realized treatment vector and the probability vector .
- Simulate Treated Outcomes: Combine the untreated outcome with the cumulative treatment effects to obtain the treated outcome.
6.1.3. Real-World Datasets
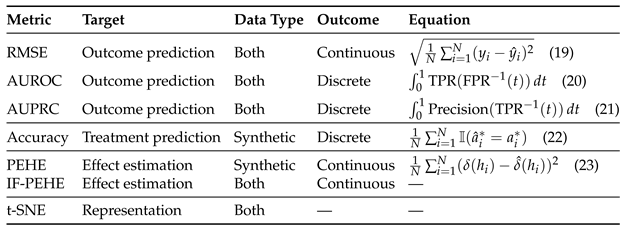
6.2. Evaluation Metrics
6.2.1. Outcome Prediction
6.2.2. Treatment Prediction
6.2.3. Treatment Effect Estimation
6.2.4. Representation Evaluation
6.3. Baselines
7. Current Challenges
7.1. Challenges From Data
7.1.1. Continuous Time Dynamics
7.1.2. Irregularities
7.2. Evaluation Challenges
7.2.1. Unobservable Counterfactuals
7.2.2. Reproducibility
8. Future Directions
8.1. Time-to-Event Outcomes in Time-Varying ITE
8.2. Multi-Modality for Time-Varying ITE
8.3. Digital Twin in Time-Varying ITE
9. Conclusions
Acknowledgments
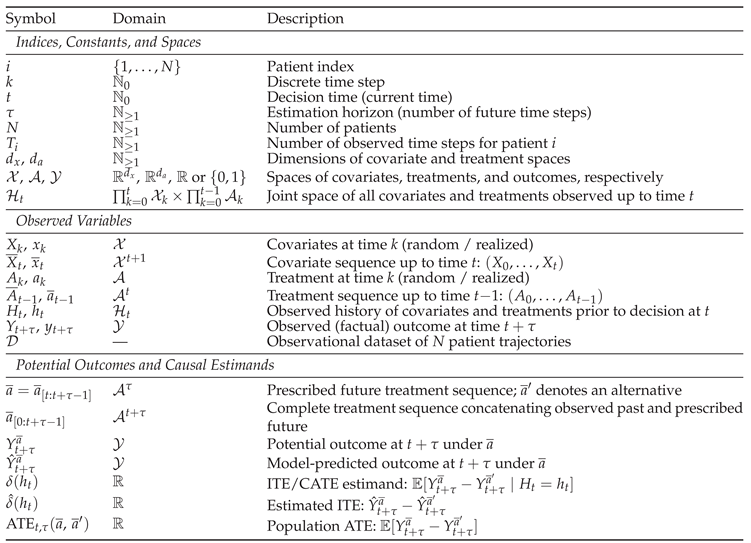
Appendix A. Notations
 |
References
- Marques, L.; Costa, B.; Pereira, M.; Silva, A.; Santos, J.; Saldanha, L.; Silva, I.; Magalhães, P.; Schmidt, S.; Vale, N. Advancing precision medicine: a review of innovative in silico approaches for drug development, clinical pharmacology and personalized healthcare. Pharmaceutics 2024, 16, 332. [Google Scholar] [CrossRef]
- Pearl, J. Causality, 2 ed.; Cambridge University Press, 2009. [Google Scholar]
- Rubin, D.B. Causal inference using potential outcomes: Design, modeling, decisions. Journal of the American statistical Association 2005, 100, 322–331. [Google Scholar] [CrossRef]
- Shalit, U.; Johansson, F.D.; Sontag, D. Estimating individual treatment effect: generalization bounds and algorithms. In Proceedings of the ICML. PMLR, 2017; pp. 3076–3085. [Google Scholar]
- Robins, J.M.; Hernan, M.A.; Brumback, B. Marginal structural models and causal inference in epidemiology; 2000. [Google Scholar]
- Robins, J. A new approach to causal inference in mortality studies with a sustained exposure period—application to control of the healthy worker survivor effect. Mathematical modelling 1986, 7, 1393–1512. [Google Scholar] [CrossRef]
- Lim, B. Forecasting treatment responses over time using recurrent marginal structural networks. NeurIPS 2018, 31. [Google Scholar]
- Bica, I.; Alaa, A.M.; Jordon, J.; Van Der Schaar, M. Estimating counterfactual treatment outcomes over time through adversarially balanced representations. ICLR, 2020. [Google Scholar]
- Melnychuk, V.; Frauen, D.; Feuerriegel, S. Causal transformer for estimating counterfactual outcomes. In Proceedings of the ICML. PMLR, 2022; pp. 15293–15329. [Google Scholar]
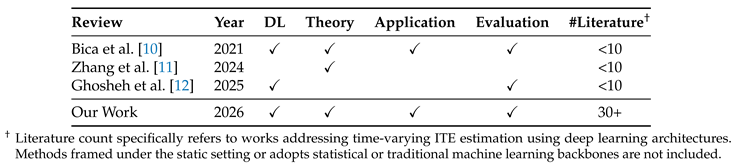
- Bica, I.; Alaa, A.M.; Lambert, C.; Van Der Schaar, M. From real-world patient data to individualized treatment effects using machine learning: current and future methods to address underlying challenges. Clinical Pharmacology & Therapeutics 2021, 109, 87–100. [Google Scholar]
- Zhang, Y.; Kreif, N.; Gc, V.S.; Manca, A. Machine learning methods to estimate individualized treatment effects for use in health technology assessment. Medical Decision Making 2024, 44, 756–769. [Google Scholar] [CrossRef]
- Ghosheh, G.O.; Gögl, M.; Zhu, T. A perspective on individualized treatment effects estimation from time-series health data. JAMIA 2025, ocae323. [Google Scholar] [CrossRef]
- Munroe, E.S.; Spicer, A.; Castellvi-Font, A.; Zalucky, A.; Dianti, J.; Linck, E.G.; Talisa, V.; Urner, M.; Angus, D.C.; Baedorf-Kassis, E.; et al. Evidence-based personalised medicine in critical care: a framework for quantifying and applying individualised treatment effects in patients who are critically ill. The Lancet Respiratory Medicine 2025, 13, 556–568. [Google Scholar] [CrossRef]
- Feuerriegel, S.; Frauen, D.; Melnychuk, V.; Schweisthal, J.; Hess, K.; Curth, A.; Bauer, S.; Kilbertus, N.; Kohane, I.S.; van der Schaar, M. Causal machine learning for predicting treatment outcomes. Nature Medicine 2024, 30, 958–968. [Google Scholar] [CrossRef]
- Shi, J.; Norgeot, B. Learning causal effects from observational data in healthcare: a review and summary. Frontiers in Medicine 2022, 9, 864882. [Google Scholar] [CrossRef]
- Neyman, J. On the application of probability theory to agricultural experiments. Essay on principles. Ann. Agricultural Sciences 1923, 1–51.
- Rubin, D.B. Estimating causal effects of treatments in randomized and nonrandomized studies. Journal of educational Psychology 1974, 66, 688. [Google Scholar] [CrossRef]
- Imbens, G.W.; Rubin, D.B. Causal inference in statistics, social, and biomedical sciences; Cambridge university press, 2015. [Google Scholar]
- Hernan, M.; Robins, J. Causal Inference: What If . In Chapman & Hall/CRC Monographs on Statistics & Applied Probab; CRC Press, 2025. [Google Scholar]
- Holland, P.W. Statistics and causal inference. Journal of the American statistical Association 1986, 81, 945–960. [Google Scholar] [CrossRef]
- Angrist, J.D.; Pischke, J.S. Mostly harmless econometrics: An empiricist’s companion; Princeton university press, 2009. [Google Scholar]
- Rosenbaum, P.R.; Rubin, D.B. The central role of the propensity score in observational studies for causal effects. Biometrika 1983, 70, 41–55. [Google Scholar] [CrossRef]
- Rubin, D.B. Randomization analysis of experimental data: The Fisher randomization test comment. Journal of the American statistical association 1980, 75, 591–593. [Google Scholar] [CrossRef]
- Li, R.; Hu, S.; Lu, M.; Utsumi, Y.; Chakraborty, P.; Sow, D.M.; Madan, P.; Li, J.; Ghalwash, M.; Shahn, Z.; et al. G-net: a recurrent network approach to g-computation for counterfactual prediction under a dynamic treatment regime. In Proceedings of the Machine Learning for Health. PMLR, 2021; pp. 282–299. [Google Scholar]
- Qian, Z.; Zhang, Y.; Bica, I.; Wood, A.; van der Schaar, M. Synctwin: Treatment effect estimation with longitudinal outcomes. NeurIPS 2021, 34, 3178–3190. [Google Scholar]
- Dedhia, B.; Balasubramanian, R.; Jha, N.K. SCouT: synthetic counterfactuals via spatiotemporal transformers for actionable healthcare. ACM Transactions on Computing for Healthcare 2023, 4, 1–28. [Google Scholar] [CrossRef]
- Liu, R.; Hunold, K.M.; Caterino, J.M.; Zhang, P. Estimating treatment effects for time-to-treatment antibiotic stewardship in sepsis. Nature machine intelligence 2023, 5, 421–431. [Google Scholar] [CrossRef]
- Su, M.; Hu, S.; Xiong, H.; Kassis, E.B.; Lehman, L.w.H. Counterfactual Sepsis Outcome Prediction Under Dynamic and Time-Varying Treatment Regimes. AMIA Summits on Translational Science Proceedings 2024, 2024, 285. [Google Scholar]
- Xiong, H.; Wu, F.; Deng, L.; Su, M.; Shahn, Z.; Lehman, L.w.H. G-transformer: Counterfactual outcome prediction under dynamic and time-varying treatment regimes. PMLR 2024, 252, https–proceedings. [Google Scholar]
- Hess, K.; Frauen, D.; Melnychuk, V.; Feuerriegel, S. G-transformer for conditional average potential outcome estimation over time. arXiv 2024. arXiv:2405.21012. [CrossRef]
- De Brouwer, E.; Gonzalez, J.; Hyland, S. Predicting the impact of treatments over time with uncertainty aware neural differential equations. In Proceedings of the AISTATS. PMLR, 2022; pp. 4705–4722. [Google Scholar]
- Liu, R.; Chen, P.Y.; Zhang, P. CURE: A deep learning framework pre-trained on large-scale patient data for treatment effect estimation. Patterns 2024, 5. [Google Scholar] [CrossRef]
- Liu, R.; Yin, C.; Zhang, P. Estimating individual treatment effects with time-varying confounders. In Proceedings of the ICDM. IEEE, 2020; pp. 382–391. [Google Scholar]
- Cao, D.; Enouen, J.; Wang, Y.; Song, X.; Meng, C.; Niu, H.; Liu, Y. Estimating treatment effects from irregular time series observations with hidden confounders. Proceedings of the AAAI 2023, Vol. 37, 6897–6905. [Google Scholar] [CrossRef]
- Cao, D.; Enouen, J.; Liu, Y. Estimating treatment effects in continuous time with hidden confounders. arXiv 2023. arXiv:2302.09446. [CrossRef]
- Wu, S.; Zhou, W.; Chen, M.; Zhu, S. Counterfactual generative models for time-varying treatments. In Proceedings of the KDD, 2024; pp. 3402–3413. [Google Scholar]
- Hess, K.; Feuerriegel, S. Stabilized neural prediction of potential outcomes in continuous time. ICLR; 2025. [Google Scholar]
- Dai, H.; Huang, Y.; Liu, Y.; He, X.; Guo, J.; Prosperi, M.; Bian, J. Variational temporal deconfounder network for individualized treatment effect estimation with longitudinal observational data. Journal of Biomedical Informatics 2025, 104880. [Google Scholar] [CrossRef] [PubMed]
- Frauen, D.; Hess, K.; Feuerriegel, S. Model-agnostic meta-learners for estimating heterogeneous treatment effects over time. ICLR; 2025. [Google Scholar]
- Seedat, N.; Imrie, F.; Bellot, A.; Qian, Z.; van der Schaar, M. Continuous-time modeling of counterfactual outcomes using neural controlled differential equations. ICML, 2022. [Google Scholar]
- Xi, P.; Wang, G.; Hu, Z.; Xiong, Y.; Gong, M.; Huang, W.; Wu, R.; Ding, Y.; Lv, T.; Fan, C.; et al. TCFimt: Temporal Counterfactual Forecasting from Individual Multiple Treatment Perspective. arXiv 2022, arXiv:2212.08890. [Google Scholar] [CrossRef]
- Jiang, S.; Huang, Z.; Luo, X.; Sun, Y. CF-GODE: Continuous-time causal inference for multi-agent dynamical systems. In Proceedings of the KDD, 2023; pp. 997–1009. [Google Scholar]
- Zhu, P.; Li, Z.; Ogino, M. Recurrent Continuous Adversarial Balancing for Estimating Individual Treatment Effect Over Time. International Journal of Computational Intelligence Systems 2023, 16, 82. [Google Scholar] [CrossRef]
- Huang, Z.; Hwang, J.; Zhang, J.; Baik, J.; Zhang, W.; Wodarz, D.; Sun, Y.; Gu, Q.; Wang, W. Causal graph ode: Continuous treatment effect modeling in multi-agent dynamical systems. In Proceedings of the WWW, 2024; pp. 4607–4617. [Google Scholar]
- Wang, X.; Lyu, S.; Yang, L.; Zhan, Y.; Chen, H. A dual-module framework for counterfactual estimation over time. In Proceedings of the ICML, 2024. [Google Scholar]
- El Bouchattaoui, M.; Tami, M.; Lepetit, B.; Cournède, P.H. Causal contrastive learning for counterfactual regression over time. NeurIPS 2024, 37, 1333–1369. [Google Scholar]
- Wendland, P.; Schenkel-Haeger, C.; Wenningmann, I.; Kschischo, M. An optimal antibiotic selection framework for Sepsis patients using Artificial Intelligence. npj Digital Medicine 2024, 7, 343. [Google Scholar] [CrossRef]
- Zhang, W.; Peng, H.; He, H. CEET: Time-Series Casual Effect Estimation via Transformer Based Representation Learning. In Proceedings of the ISCAIT. IEEE, 2025; pp. 369–373. [Google Scholar]
- Liu, Y.; Dong, J.; Fu, C.; Shi, W.; Jiang, Z.; Hua, Z.; Long, B.; Carlson, D. Counterfactual Outcome Estimation in Time Series via Sub-treatment Group Alignment and Random Temporal Masking. 2025. [Google Scholar]
- Norimatsu, Y.; Imaizumi, M. Encode-Decoder-based GAN for Estimating Counterfactual Outcomes under Sequential Selection Bias and Combinatorial Explosion. In Proceedings of the CLeaR, 2025. [Google Scholar]
- Wang, H.; Li, H.; Zou, H.; Chi, H.; Lan, L.; Huang, W.; Yang, W. Effective and Efficient Time-Varying Counterfactual Prediction with State-Space Models. In Proceedings of the ICLR, 2025. [Google Scholar]
- Guski, J.; Botz, J.; Fröhlich, H. Estimating the causal impact of non-pharmaceutical interventions on COVID-19 spread in seven EU countries via machine learning. Scientific Reports 2025, 15, 9203. [Google Scholar] [CrossRef] [PubMed]
- Abraich, A. TV-SurvCaus: Dynamic Representation Balancing for Causal Survival Analysis. arXiv arXiv:2505.01785.
- Berrevoets, J.; Curth, A.; Bica, I.; McKinney, E.; van der Schaar, M. Disentangled counterfactual recurrent networks for treatment effect inference over time. arXiv arXiv:2112.03811. [CrossRef]
- Chu, J.; Zhang, Y.; Huang, F.; Si, L.; Huang, S.; Huang, Z. Disentangled representation for sequential treatment effect estimation. Computer Methods and Programs in Biomedicine 2022, 226, 107175. [Google Scholar] [CrossRef]
- Thomas, L.E.; Yang, S.; Wojdyla, D.; Schaubel, D.E. Matching with time-dependent treatments: a review and look forward. Statistics in medicine 2020, 39, 2350–2370. [Google Scholar] [CrossRef]
- Abadie, A.; L’hour, J. A penalized synthetic control estimator for disaggregated data. Journal of the American Statistical Association 2021, 116, 1817–1834. [Google Scholar] [CrossRef]
- Taubman, S.L.; Robins, J.M.; Mittleman, M.A.; Hernán, M.A. Intervening on risk factors for coronary heart disease: an application of the parametric g-formula. International journal of epidemiology 2009, 38, 1599–1611. [Google Scholar] [CrossRef]
- Imbens, G.W. Nonparametric estimation of average treatment effects under exogeneity: A review. Review of Economics and statistics 2004, 86, 4–29. [Google Scholar] [CrossRef]
- Chen, X.; Hu, L.; Li, F. A flexible Bayesian g-formula for causal survival analyses with time-dependent confounding: X. Chen et al. Lifetime Data Analysis 2025, 31, 394–421. [Google Scholar] [CrossRef]
- Bica, I.; Alaa, A.; Van Der Schaar, M. Time series deconfounder: Estimating treatment effects over time in the presence of hidden confounders. In Proceedings of the ICML. PMLR, 2020; pp. 884–895. [Google Scholar]
- Bang, H.; Robins, J.M. Doubly robust estimation in missing data and causal inference models. Biometrics 2005, 61, 962–973. [Google Scholar] [CrossRef]
- Robins, J.M.; Rotnitzky, A.; Zhao, L.P. Estimation of regression coefficients when some regressors are not always observed. Journal of the American statistical Association 1994, 89, 846–866. [Google Scholar] [CrossRef]
- Xiao, Y.; Moodie, E.E.; Abrahamowicz, M. Comparison of approaches to weight truncation for marginal structural Cox models. Epidemiologic Methods 2013, 2, 1–20. [Google Scholar] [CrossRef]
- Léger, M.; Chatton, A.; Le Borgne, F.; Pirracchio, R.; Lasocki, S.; Foucher, Y. Causal inference in case of near-violation of positivity: comparison of methods. Biometrical Journal 2022, 64, 1389–1403. [Google Scholar] [CrossRef] [PubMed]
- Testa, L.; Chiaromonte, F.; Roeder, K. Rescuing double robustness: safe estimation under complete misspecification. arXiv arXiv:2509.22446. [CrossRef]
- Cheng, P.; Hao, W.; Dai, S.; Liu, J.; Gan, Z.; Carin, L. Club: A contrastive log-ratio upper bound of mutual information. In Proceedings of the ICML. PMLR, 2020; pp. 1779–1788. [Google Scholar]
- Melnychuk, V.; Frauen, D.; Feuerriegel, S. Bounds on representation-induced confounding bias for treatment effect estimation. ICLR, 2024. [Google Scholar]
- Welch, R.; Zhang, J.; Uhler, C. Identifiability guarantees for causal disentanglement from purely observational data. NeurIPS 2024, 37, 102796–102821. [Google Scholar]
- Johnson, A.E.; Pollard, T.J.; Shen, L.; Lehman, L.w.H.; Feng, M.; Ghassemi, M.; Moody, B.; Szolovits, P.; Anthony Celi, L.; Mark, R.G. MIMIC-III, a freely accessible critical care database. Scientific data 2016, 3, 1–9. [Google Scholar] [CrossRef]
- Johnson, A.E.; Bulgarelli, L.; Shen, L.; Gayles, A.; Shammout, A.; Horng, S.; Pollard, T.J.; Hao, S.; Moody, B.; Gow, B.; et al. MIMIC-IV, a freely accessible electronic health record dataset. Scientific data 2023, 10, 1. [Google Scholar] [CrossRef]
- Kuzmanovic, M.; Hatt, T.; Feuerriegel, S. Deconfounding Temporal Autoencoder: estimating treatment effects over time using noisy proxies. In Proceedings of the Machine learning for health. PMLR, 2021; pp. 143–155. [Google Scholar]
- Vasan, N.; Baselga, J.; Hyman, D.M. A view on drug resistance in cancer. Nature 2019, 575, 299–309. [Google Scholar] [CrossRef]
- Wang, X.; Lyu, S.; Luo, C.; Zhou, X.; Chen, H. Variational counterfactual intervention planning to achieve target outcomes. In Proceedings of the ICML, 2025. [Google Scholar]
- Geng, C.; Paganetti, H.; Grassberger, C. Prediction of treatment response for combined chemo-and radiation therapy for non-small cell lung cancer patients using a bio-mathematical model. Scientific reports 2017, 7, 13542. [Google Scholar] [CrossRef]
- Beekly, D.L.; Ramos, E.M.; Lee, W.W.; Deitrich, W.D.; Jacka, M.E.; Wu, J.; Hubbard, J.L.; Koepsell, T.D.; Morris, J.C.; Kukull, W.A.; et al. The National Alzheimer’s Coordinating Center (NACC) database: the uniform data set. Alzheimer Disease & Associated Disorders 2007, 21, 249–258. [Google Scholar]
- Regner, S.R.; Wilcox, N.S.; Friedman, L.S.; Seyer, L.A.; Schadt, K.A.; Brigatti, K.W.; Perlman, S.; Delatycki, M.; Wilmot, G.R.; Gomez, C.M.; et al. Friedreich ataxia clinical outcome measures: natural history evaluation in 410 participants. Journal of child neurology 2012, 27, 1152–1158. [Google Scholar] [CrossRef]
- Heldt, T.; Mukkamala, R.; Moody, G.B.; Mark, R.G. CVSim: an open-source cardiovascular simulator for teaching and research. The open pacing, electrophysiology & therapy journal 2010, 3, 45. [Google Scholar]
- Zenker, S.; Rubin, J.; Clermont, G. From inverse problems in mathematical physiology to quantitative differential diagnoses. PLoS computational biology 2007, 3, e204. [Google Scholar] [CrossRef] [PubMed]
- Linial, O.; Ravid, N.; Eytan, D.; Shalit, U. Generative ode modeling with known unknowns. In Proceedings of the Proceedings of the Conference on Health, Inference, and Learning, 2021; pp. 79–94. [Google Scholar]
- Dai, W.; Rao, R.; Sher, A.; Tania, N.; Musante, C.J.; Allen, R. A prototype QSP model of the immune response to SARS-CoV-2 for community development. CPT: pharmacometrics & systems pharmacology 2021, 10, 18–29. [Google Scholar]
- Qian, Z.; Zame, W.; Fleuren, L.; Elbers, P.; van der Schaar, M. Integrating expert ODEs into neural ODEs: pharmacology and disease progression. NeurIPS 2021, 34, 11364–11383. [Google Scholar]
- Saeed, M.; Lieu, C.; Raber, G.; Mark, R.G. MIMIC II: a massive temporal ICU patient database to support research in intelligent patient monitoring. In Proceedings of the Computers in cardiology. IEEE, 2002; pp. 641–644. [Google Scholar]
- Thoral, P.J.; Peppink, J.M.; Driessen, R.H.; Sijbrands, E.J.; Kompanje, E.J.; Kaplan, L.; Bailey, H.; Kesecioglu, J.; Cecconi, M.; Churpek, M.; et al. Sharing ICU patient data responsibly under the society of critical care medicine/European society of intensive care medicine joint data science collaboration: the Amsterdam university medical centers database (AmsterdamUMCdb) example. Critical care medicine 2021, 49, e563–e577. [Google Scholar] [CrossRef]
- Group, C.A.M.P.R.; et al. The childhood asthma management program (CAMP): design, rationale, and methods. Controlled clinical trials 1999, 20, 91–120. [Google Scholar]
- Zhang, X.L.; Begleiter, H.; Porjesz, B.; Wang, W.; Litke, A. Event related potentials during object recognition tasks. Brain research bulletin 1995, 38, 531–538. [Google Scholar] [CrossRef]
- Steiger, E.; Mußgnug, T.; Kroll, L.E. Causal analysis of COVID-19 observational data in German districts reveals effects of mobility, awareness, and temperature. medRxiv 2020, 2020–07. [Google Scholar] [CrossRef]
- Herrett, E.; Gallagher, A.M.; Bhaskaran, K.; Forbes, H.; Mathur, R.; Van Staa, T.; Smeeth, L. Data resource profile: clinical practice research datalink (CPRD). International journal of epidemiology 2015, 44, 827–836. [Google Scholar] [CrossRef]
- Merative. MarketScan Research Databases 2023.
- Hatt, T.; Feuerriegel, S. Sequential deconfounding for causal inference with unobserved confounders. In Proceedings of the Causal Learning and Reasoning. PMLR, 2024; pp. 934–956. [Google Scholar]
- Hill, J.L. Bayesian nonparametric modeling for causal inference. Journal of Computational and Graphical Statistics 2011, 20, 217–240. [Google Scholar] [CrossRef]
- Alaa, A.; Van Der Schaar, M. Validating causal inference models via influence functions. In Proceedings of the ICML. PMLR, 2019; pp. 191–201. [Google Scholar]
- Maaten, L.; Hinton, G. Visualizing data using t-SNE. Journal of machine learning research 2008, 9, 2579–2605. [Google Scholar]
- Kacprzyk, K.; Holt, S.; Berrevoets, J.; Qian, Z.; van der Schaar, M. ODE discovery for longitudinal heterogeneous treatment effects inference. ICLR, 2024. [Google Scholar]
- Che, Z.; Purushotham, S.; Cho, K.; Sontag, D.; Liu, Y. Recurrent neural networks for multivariate time series with missing values. Scientific reports 2018, 8, 6085. [Google Scholar] [CrossRef]
- Chen, R.T.; Rubanova, Y.; Bettencourt, J.; Duvenaud, D.K. Neural ordinary differential equations. NeurIPS 2018, 31. [Google Scholar]
- Kim, S.; Ji, W.; Deng, S.; Ma, Y.; Rackauckas, C. Stiff neural ordinary differential equations. Chaos: An Interdisciplinary Journal of Nonlinear Science 2021, 31. [Google Scholar] [CrossRef]
- Singer, M.; Deutschman, C.S.; Seymour, C.W.; Shankar-Hari, M.; Annane, D.; Bauer, M.; Bellomo, R.; Bernard, G.R.; Chiche, J.D.; Coopersmith, C.M.; et al. The third international consensus definitions for sepsis and septic shock (Sepsis-3). Jama 2016, 315, 801–810. [Google Scholar] [CrossRef]
- Sisk, R.; Lin, L.; Sperrin, M.; Barrett, J.K.; Tom, B.; Diaz-Ordaz, K.; Peek, N.; Martin, G.P. Informative presence and observation in routine health data: a review of methodology for clinical risk prediction. JAMIA 2021, 28, 155–166. [Google Scholar] [CrossRef]
- Rubin, D.B. Inference and missing data. Biometrika 1976, 63, 581–592. [Google Scholar] [CrossRef]
- Agniel, D.; Kohane, I.S.; Weber, G.M. Biases in electronic health record data due to processes within the healthcare system: retrospective observational study. BMJ 2018, 361. [Google Scholar] [CrossRef]
- Vanderschueren, T.; Curth, A.; Verbeke, W.; Van Der Schaar, M. Accounting for informative sampling when learning to forecast treatment outcomes over time. In Proceedings of the ICML. PMLR, 2023; pp. 34855–34874. [Google Scholar]
- Gentzel, A.; Garant, D.; Jensen, D. The case for evaluating causal models using interventional measures and empirical data. NeurIPS 2019, 32. [Google Scholar]
- Parikh, H.; Varjao, C.; Xu, L.; Tchetgen, E.T. Validating causal inference methods. In Proceedings of the ICML. PMLR, 2022; pp. 17346–17358. [Google Scholar]
- Neal, B.; Huang, C.W.; Raghupathi, S. Realcause: Realistic causal inference benchmarking. arXiv arXiv:2011.15007. [CrossRef]
- Gögl, M.; Liu, Y.; Yau, C.; Watkinson, P.; Zhu, T. DoseSurv: Predicting Personalized Survival Outcomes under Continuous-Valued Treatments. 2025.
- D’Amour, A.; Ding, P.; Feller, A.; Lei, L.; Sekhon, J. Overlap in observational studies with high-dimensional covariates. Journal of Econometrics 2021, 221, 644–654. [Google Scholar] [CrossRef]
- Corral-Acero, J.; Margara, F.; Marciniak, M.; Rodero, C.; Loncaric, F.; Feng, Y.; Gilbert, A.; Fernandes, J.F.; Bukhari, H.A.; Wajdan, A.; et al. The ‘Digital Twin’to enable the vision of precision cardiology. European heart journal 2020, 41, 4556–4564. [Google Scholar] [CrossRef]
- Kiciman, E.; Ness, R.; Sharma, A.; Tan, C. Causal reasoning and large language models: Opening a new frontier for causality. TMLR, 2023. [Google Scholar]
- Deng, L.; Xiong, H.; Wu, F.; Kapoor, S.; Ghosh, S.; Shahn, Z.; Lehman, L.w.H. Uncertainty Quantification for Conditional Treatment Effect Estimation under Dynamic Treatment Regimes. PMLR 2024, 259, 248. [Google Scholar]
- Hess, K.; Melnychuk, V.; Frauen, D.; Feuerriegel, S. Bayesian neural controlled differential equations for treatment effect estimation. ICLR, 2023. [Google Scholar]
| 1 | This should be distinguished from selection bias, which arises from conditioning on a common effect (collider) of treatment and outcome [19]. |
| 2 | Following standard practice in the literature, the outcome index is shifted to to reflect the assumption that a treatment administered at time k affects the outcome observed at time . |
 and
and  icon separately denotes trainable and frozen model parameters. Solid and dashed circular outlines distinguishes between observed data and either prescribed or predicted values.
icon separately denotes trainable and frozen model parameters. Solid and dashed circular outlines distinguishes between observed data and either prescribed or predicted values.
 and
and  icon separately denotes trainable and frozen model parameters. Solid and dashed circular outlines distinguishes between observed data and either prescribed or predicted values.
icon separately denotes trainable and frozen model parameters. Solid and dashed circular outlines distinguishes between observed data and either prescribed or predicted values.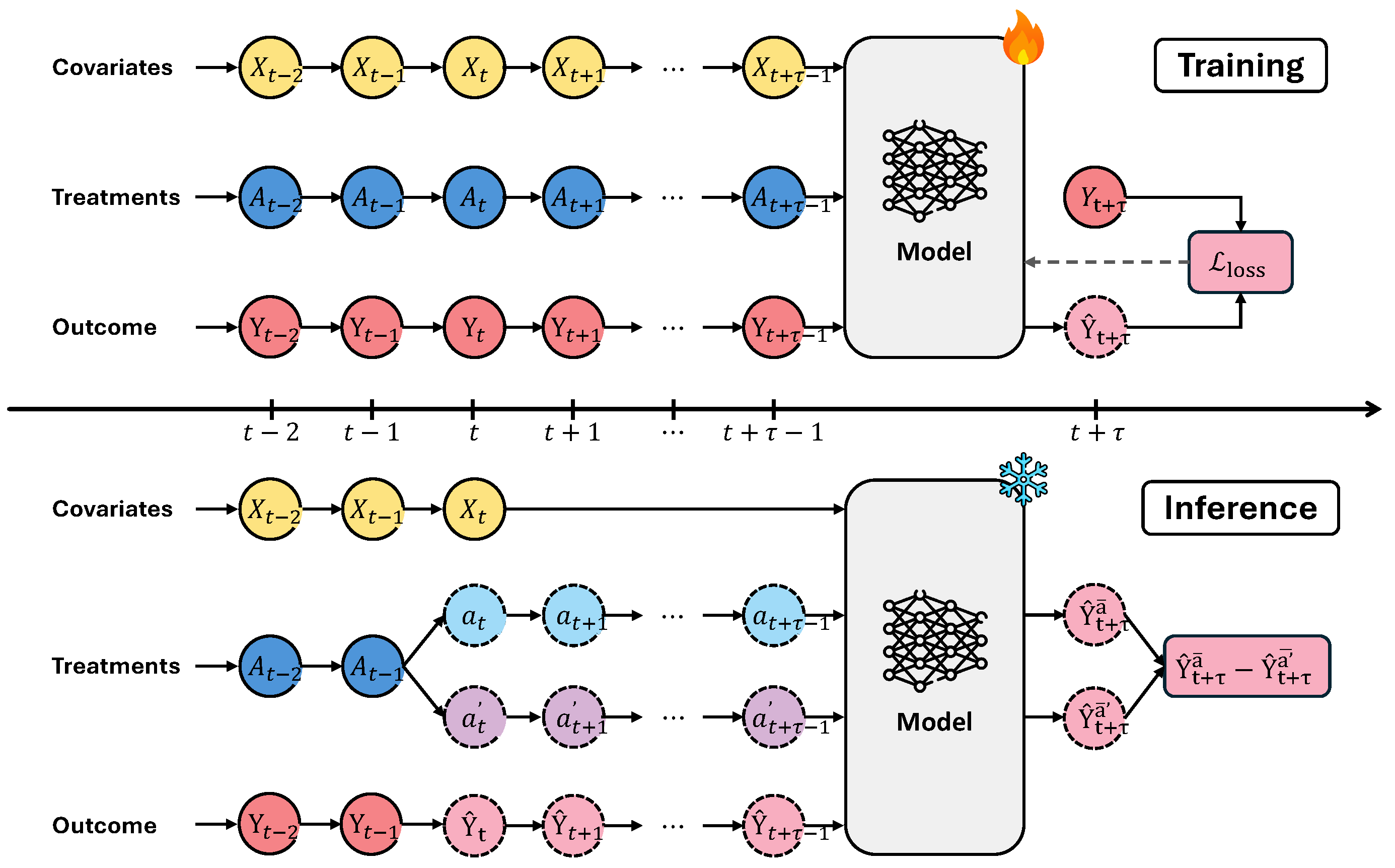
 |
 |
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2026 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).




