Submitted:
12 April 2026
Posted:
14 April 2026
You are already at the latest version
Abstract
Keywords:
1. Introduction
2. Materials and Methods
Ethics Statement
Pleural Effusion Collection and Sample Processing
Patient-Derived Organoid Culture
Establishing Cisplatin Tolerant Organoids
Immunohistochemistry
Ki67 Proliferation Index
Cell Viability and Half-Maximal Inhibitory Concentration Determination
RNA Isolation from Patient-Derived Organoids
RNA Sequencing
Metabolic Flux Analysis
Data Analysis
3. Results
3.1. Morphological and Histological Characterisation of Mesothelioma Organoid Models
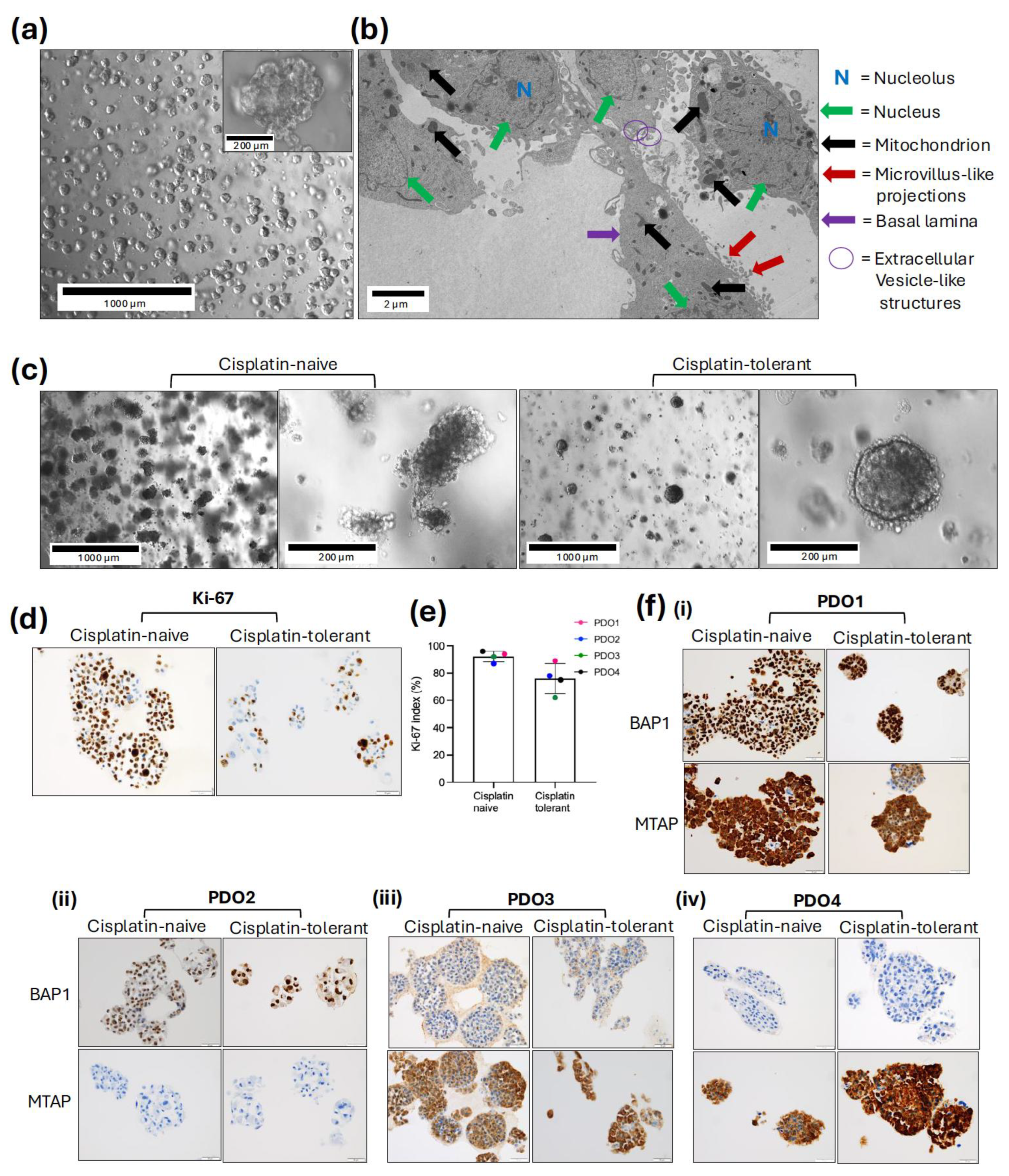
3.2. Distinct Molecular Landscapes of Mesothelioma PDO Models
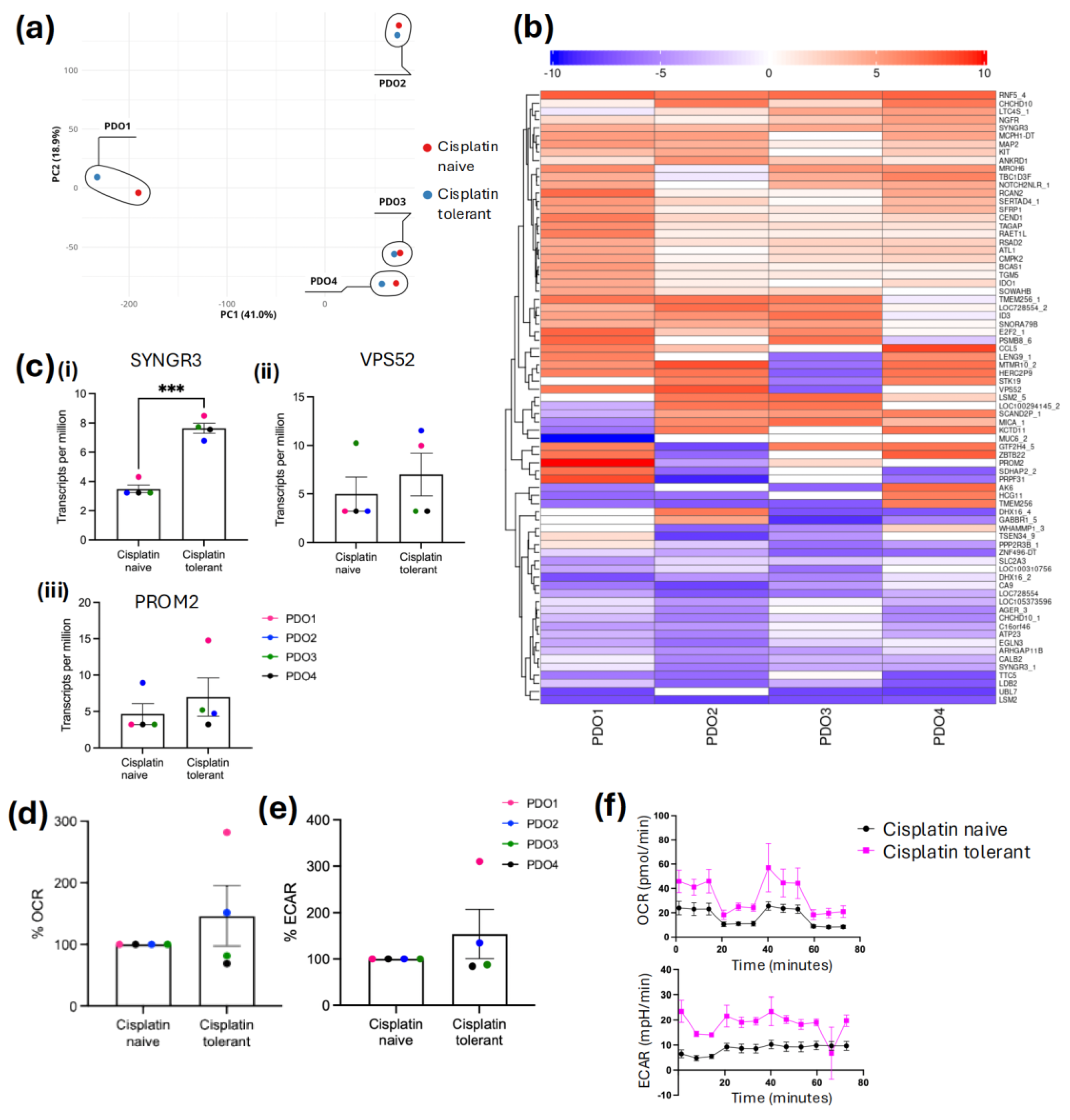
4. Discussion
5. Conclusions
Supplementary Materials
Author Contributions
Funding
Data Availability Statement
Acknowledgments
Conflicts of Interest
Abbreviations
| ACK: | Ammonium-Chloride-Potassium |
| AdVDMEM/F12: | Advanced Dulbecco’s Modified Eagle Medium/Ham’s/F-12 |
| ALKBH5: | AlkB homolog 5 |
| ALM: | Acral lentiginous melanoma |
| DDB2: | Damage-specific DNA-binding protein 2 |
| DTP: | Drug-tolerant persister |
| ECAR: | Extracellular acidification rate |
| EV: | Extracellular vesicle |
| FCCP: | Carbonyl cyanide-p-trifluoromethoxyphenylhydrazone |
| GARP: | Golgi-associated retrograde protein |
| IC50: | Half-maximal inhibitory concentration |
| IHC: | Immunohistochemistry |
| m6A: | N6-methyladenosine |
| NATA: | National Association of Testing Authorities |
| NSCLC: | Non-small cell lung cancer |
| OCR: | Oxygen consumption rate |
| PCA: | Principal Component Analysis |
| PDO: | Patient-derived organoid |
| PM: | Pleural mesothelioma |
| PRMT5: | Protein Arginine Methyltransferase 5 |
| PTMs: | Posttranslational modifications |
| RCPA: | Royal College of Pathologists of Australia |
| RI: | Resistance Index |
| SD: | Standard deviation |
| SEM: | Standard error of the mean |
| TEM: | Transmission electron microscopy |
| WT: | Wild type |
References
- Kindler HL, Ismaila N, Bazhenova L, Chu Q, Churpek JE, Dagogo-Jack I, et al. Treatment of Pleural Mesothelioma: ASCO Guideline Update. Journal of Clinical Oncology. 2025;43(8):1006-38. [CrossRef]
- Oehl K, Vrugt B, Wagner U, Kirschner MB, Meerang M, Weder W, et al. Alterations in BAP1 Are Associated with Cisplatin Resistance through Inhibition of Apoptosis in Malignant Pleural Mesothelioma. Clinical Cancer Research. 2021;27(8):2277-91.
- Mikubo M, Inoue Y, Liu G, Tsao M-S. Mechanism of Drug Tolerant Persister Cancer Cells: The Landscape and Clinical Implication for Therapy. Journal of Thoracic Oncology. 2021;16(11):1798-809. [CrossRef]
- Volpini L, Monaco F, Casalone E, Herman EJ, Filomena R, Manzotti S, et al. Use of patient-derived organoids for pleural mesothelioma 3-D modeling. Am J Physiol Lung Cell Mol Physiol. 2025;329(6):L756-l69. [CrossRef]
- Lin RZ, Chang HY. Recent advances in three-dimensional multicellular spheroid culture for biomedical research. Biotechnol J. 2008;3(9-10):1172-84.
- Shen S, Vagner S, Robert C. Persistent Cancer Cells: The Deadly Survivors. Cell. 2020;183(4):860-74. [CrossRef]
- Marine JC, Dawson SJ, Dawson MA. Non-genetic mechanisms of therapeutic resistance in cancer. Nat Rev Cancer. 2020;20(12):743-56. [CrossRef]
- Ramirez M, Rajaram S, Steininger RJ, Osipchuk D, Roth MA, Morinishi LS, et al. Diverse drug-resistance mechanisms can emerge from drug-tolerant cancer persister cells. Nat Commun. 2016;7:10690. [CrossRef]
- Sharma SV, Lee DY, Li B, Quinlan MP, Takahashi F, Maheswaran S, et al. A chromatin-mediated reversible drug-tolerant state in cancer cell subpopulations. Cell. 2010;141(1):69-80. [CrossRef]
- Hata AN, Niederst MJ, Archibald HL, Gomez-Caraballo M, Siddiqui FM, Mulvey HE, et al. Tumor cells can follow distinct evolutionary paths to become resistant to epidermal growth factor receptor inhibition. Nat Med. 2016;22(3):262-9. [CrossRef]
- Peyre L, Péré M, Meyer M, Bian B, Moureau-Barbato M, Djema W, et al. Transition between cell states of sensitivity reveals molecular vulnerability of drug-tolerant cells. Mol Syst Biol. 2025;21(12):1702-30. [CrossRef]
- Vlachogiannis G, Hedayat S, Vatsiou A, Jamin Y, Fernández-Mateos J, Khan K, et al. Patient-derived organoids model treatment response of metastatic gastrointestinal cancers. Science. 2018;359(6378):920-6. [CrossRef]
- Sharick JT, Walsh CM, Sprackling CM, Pasch CA, Pham DL, Esbona K, et al. Metabolic Heterogeneity in Patient Tumor-Derived Organoids by Primary Site and Drug Treatment. Front Oncol. 2020;10:553. [CrossRef]
- Jia D, Lu M, Jung KH, Park JH, Yu L, Onuchic JN, et al. Elucidating cancer metabolic plasticity by coupling gene regulation with metabolic pathways. Proceedings of the National Academy of Sciences. 2019;116(9):3909-18. [CrossRef]
- Emert BL, Cote CJ, Torre EA, Dardani IP, Jiang CL, Jain N, et al. Variability within rare cell states enables multiple paths toward drug resistance. Nat Biotechnol. 2021;39(7):865-76. [CrossRef]
- Viale A, Pettazzoni P, Lyssiotis CA, Ying H, Sánchez N, Marchesini M, et al. Oncogene ablation-resistant pancreatic cancer cells depend on mitochondrial function. Nature. 2014;514(7524):628-32. [CrossRef]
- Cedres S, Valdivia A, Priano I, Rocha P, Iranzo P, Pardo N, et al. BAP1 Mutations and Pleural Mesothelioma: Genetic Insights, Clinical Implications, and Therapeutic Perspectives. Cancers (Basel). 2025;17(9). [CrossRef]
- Husain AN, Chapel DB, Attanoos R, Beasley MB, Brcic L, Butnor K, et al. Guidelines for Pathologic Diagnosis of Mesothelioma: 2023 Update of the Consensus Statement From the International Mesothelioma Interest Group. Arch Pathol Lab Med. 2024;148(11):1251-71. [CrossRef]
- Hocking AJ, Mortimer LA, Farrall AL, Russell PA, Klebe S. Establishing mesothelioma patient-derived organoid models from malignant pleural effusions. Lung Cancer. 2024;191:107542. [CrossRef]
- Harada K, Sakamoto N, Ukai S, Yamamoto Y, Pham QT, Taniyama D, et al. Establishment of oxaliplatin-resistant gastric cancer organoids: importance of myoferlin in the acquisition of oxaliplatin resistance. Gastric Cancer. 2021;24(6):1264-77. [CrossRef]
- Ewels PA, Peltzer A, Fillinger S, Patel H, Alneberg J, Wilm A, et al. The nf-core framework for community-curated bioinformatics pipelines. Nature Biotechnology. 2020;38(3):276-8. [CrossRef]
- Grüning B, Dale R, Sjödin A, Chapman BA, Rowe J, Tomkins-Tinch CH, et al. Bioconda: sustainable and comprehensive software distribution for the life sciences. Nature Methods. 2018;15(7):475-6. [CrossRef]
- da Veiga Leprevost F, Grüning BA, Alves Aflitos S, Röst HL, Uszkoreit J, Barsnes H, et al. BioContainers: an open-source and community-driven framework for software standardization. Bioinformatics. 2017;33(16):2580-2.
- Di Tommaso P, Chatzou M, Floden EW, Barja PP, Palumbo E, Notredame C. Nextflow enables reproducible computational workflows. Nature Biotechnology. 2017;35(4):316-9. [CrossRef]
- Love MI, Huber W, Anders S. Moderated estimation of fold change and dispersion for RNA-seq data with DESeq2. Genome Biology. 2014;15(12):550.
- Louw A, Panou V, Szejniuk WM, Meristoudis C, Chai SM, van Vliet C, et al. BAP1 Loss by Immunohistochemistry Predicts Improved Survival to First-Line Platinum and Pemetrexed Chemotherapy for Patients With Pleural Mesothelioma: A Validation Study. J Thorac Oncol. 2022;17(7):921-30. [CrossRef]
- Pulford E, Huilgol K, Moffat D, Henderson DW, Klebe S. Malignant Mesothelioma, BAP1 Immunohistochemistry, and VEGFA: Does BAP1 Have Potential for Early Diagnosis and Assessment of Prognosis? Dis Markers. 2017;2017:1310478. [CrossRef]
- Lalloo F, Kulkarni A, Chau C, Nielsen M, Sheaff M, Steele J, et al. Clinical practice guidelines for the diagnosis and surveillance of BAP1 tumour predisposition syndrome. Eur J Hum Genet. 2023;31(11):1261-9. [CrossRef]
- Brown CW, Amante JJ, Chhoy P, Elaimy AL, Liu H, Zhu LJ, et al. Prominin2 Drives Ferroptosis Resistance by Stimulating Iron Export. Dev Cell. 2019;51(5):575-86.e4. [CrossRef]
- Schindler C, Chen Y, Pu J, Guo X, Bonifacino JS. EARP is a multisubunit tethering complex involved in endocytic recycling. Nat Cell Biol. 2015;17(5):639-50. [CrossRef]
- Murphy RM, Tasoulas J, Porrello A, Carper MB, Tsai Y-H, Coffey AR, et al. Tumor Cell Extrinsic Synaptogyrin 3 Expression as a Diagnostic and Prognostic Biomarker in Head and Neck Cancer. Cancer Research Communications. 2022;2(9):987-1004. [CrossRef]
- Hangauer MJ, Viswanathan VS, Ryan MJ, Bole D, Eaton JK, Matov A, et al. Drug-tolerant persister cancer cells are vulnerable to GPX4 inhibition. Nature. 2017;551(7679):247-50. [CrossRef]
- Peng-Winkler Y, Fendt SM. Metabolic adaptations in cancer progression. Physiol Rev. 2026;106(1):1-51. [CrossRef]
- Davidson B, Holth A, Hummel C, Flatmark K, Torgunrud A. Different Frequency and Clinical Role for MTAP Loss in Pleural and Peritoneal Mesothelioma. Appl Immunohistochem Mol Morphol. 2024;32(6):280-4. [CrossRef]
- Yu F, Zheng S, Yu C, Gao S, Shen Z, Nar R, et al. KRAS mutants confer platinum resistance by regulating ALKBH5 posttranslational modifications in lung cancer. J Clin Invest. 2025;135(6). [CrossRef]
- Conibear E, Stevens TH. Vps52p, Vps53p, and Vps54p form a novel multisubunit complex required for protein sorting at the yeast late Golgi. Mol Biol Cell. 2000;11(1):305-23.
- Li L, Ho PW, Liu H, Pang SY, Chang EE, Choi ZY, et al. Transcriptional Regulation of the Synaptic Vesicle Protein Synaptogyrin-3 (SYNGR3) Gene: The Effects of NURR1 on Its Expression. Int J Mol Sci. 2022;23(7). [CrossRef]
- Rehman SK, Haynes J, Collignon E, Brown KR, Wang Y, Nixon AML, et al. Colorectal Cancer Cells Enter a Diapause-like DTP State to Survive Chemotherapy. Cell. 2021;184(1):226-42.e21. [CrossRef]
- Shrestha R, Nabavi N, Lin YY, Mo F, Anderson S, Volik S, et al. BAP1 haploinsufficiency predicts a distinct immunogenic class of malignant peritoneal mesothelioma. Genome Med. 2019;11(1):8. [CrossRef]
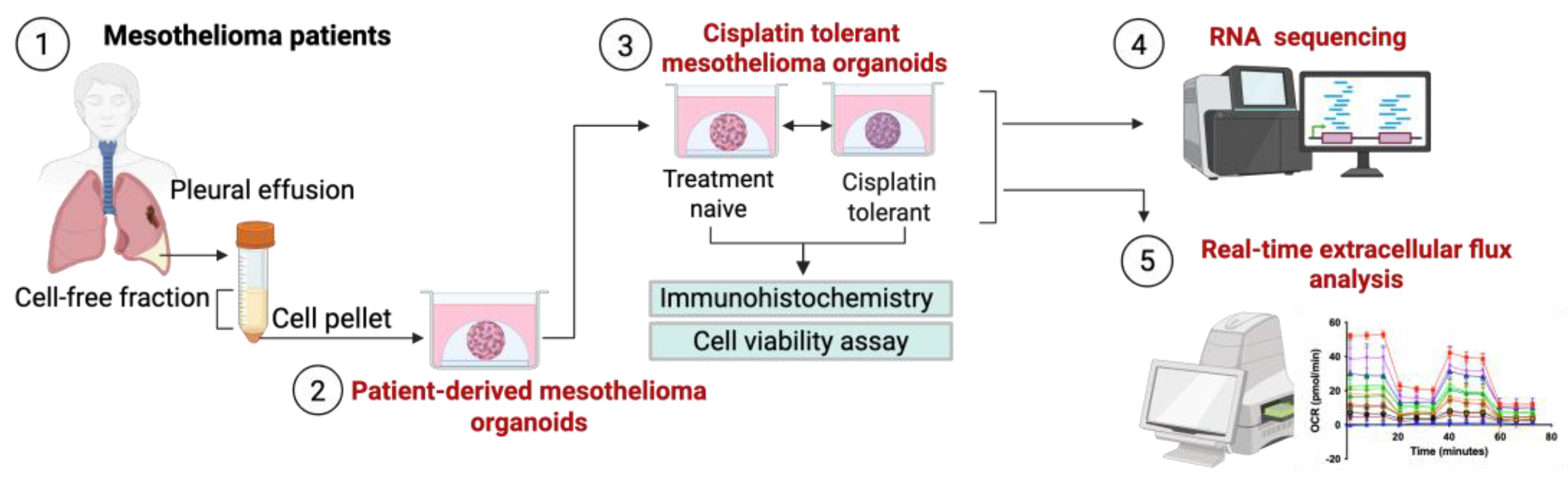
| PDO ID | Age at diagnosis (years) | Sex | Survival (months) |
BAP1 Status | MTAP Status | KRAS Status | Metabolic Phenotype | VPS52 | SYNGR3 | PROM2 |
|---|---|---|---|---|---|---|---|---|---|---|
| PDO 1 |
47 |
Male |
1 |
+ |
+ |
+ |
Hyper metabolic/hybrid state. Concurrent high increase in bioenergetic flux. | ↑ | ↑ | ↑ |
| PDO2 |
77 |
Male |
1 |
+ |
- |
N/A |
Hyper metabolic/hybrid state. Concurrent increase in bioenergetic flux. | ↑ | ↑ | ↓ |
| PDO3 |
74 |
Male |
20 |
- |
+ |
N/A |
High reliance on mitochondrial respiration; predicted convergent stable baseline. | ↓ | ↑ | ↑ |
| PDO4 |
80 |
Female |
5 |
- |
+ |
N/A |
↔ | ↑ | ↔ |
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2026 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).




