Submitted:
11 April 2026
Posted:
14 April 2026
You are already at the latest version
Abstract
Keywords:
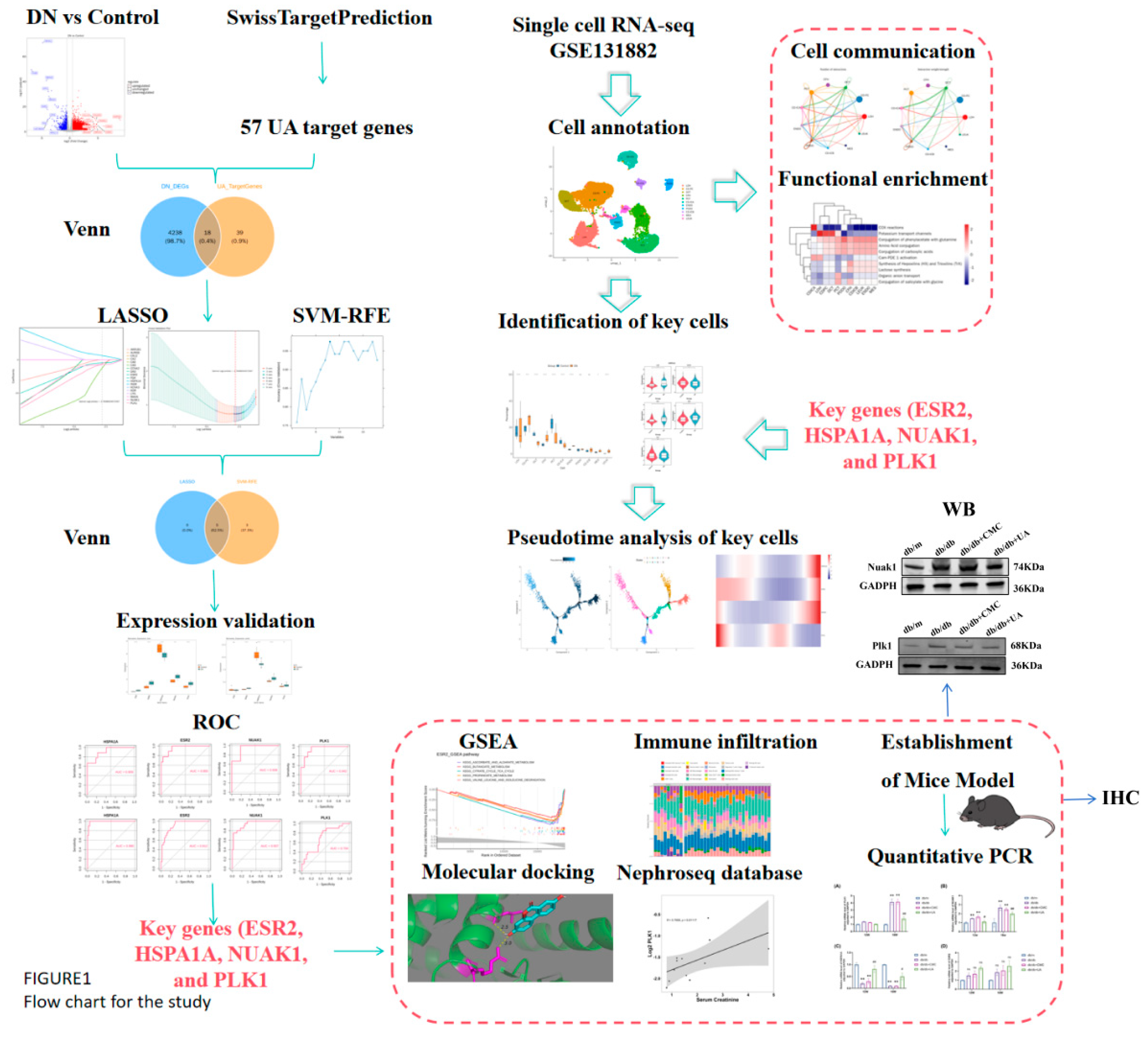
1. Introduction
2. Results
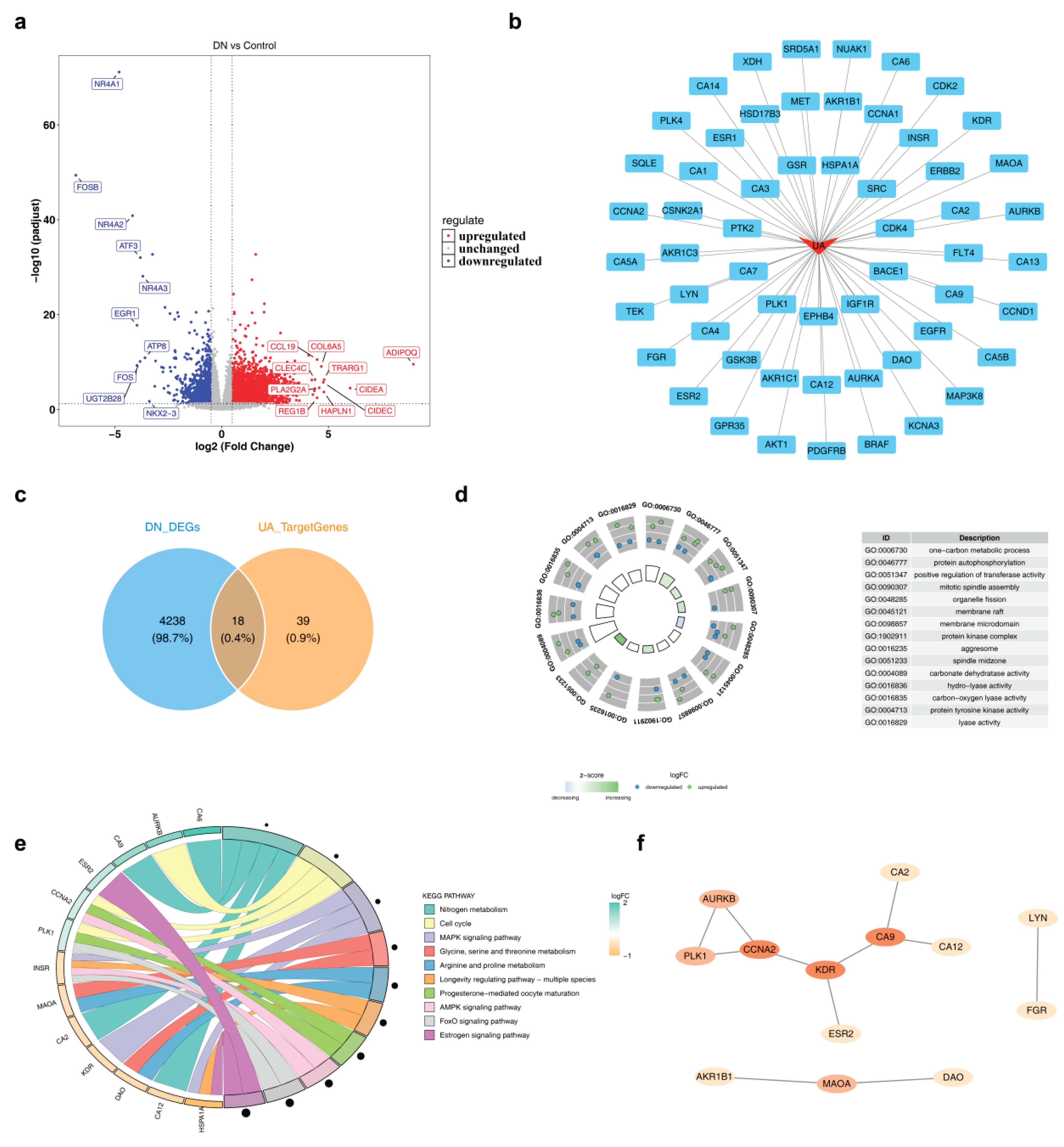
2.1. 18 Candidate genes affecting DKD development
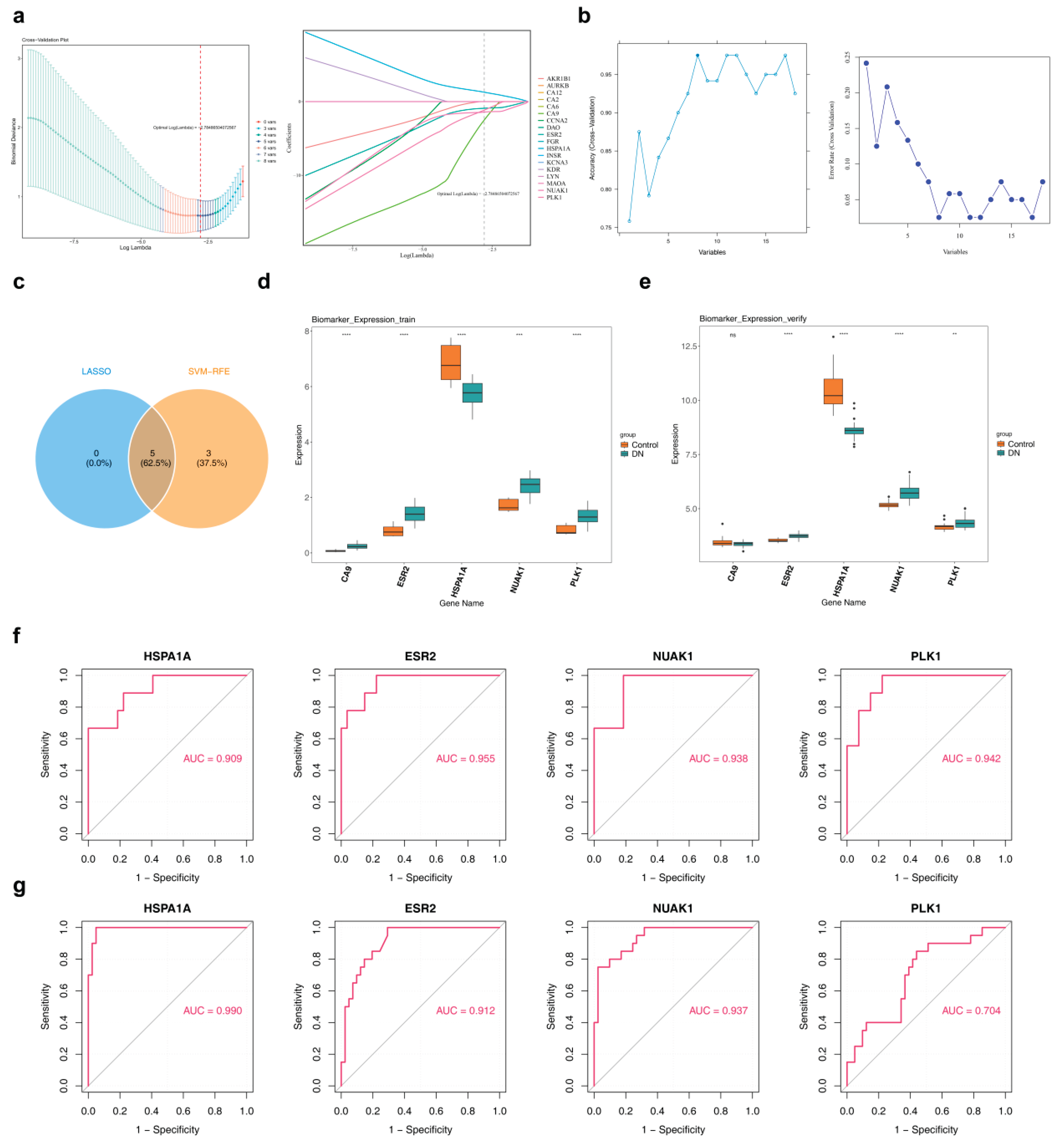
2.2. A total of 4 biomarkers that treat DKD were discovered
2.3. Functional enrichment and immunoregulation of biomarkers in DKD
2.4. Biomarker-UA binding affinity and clinical renal correlation
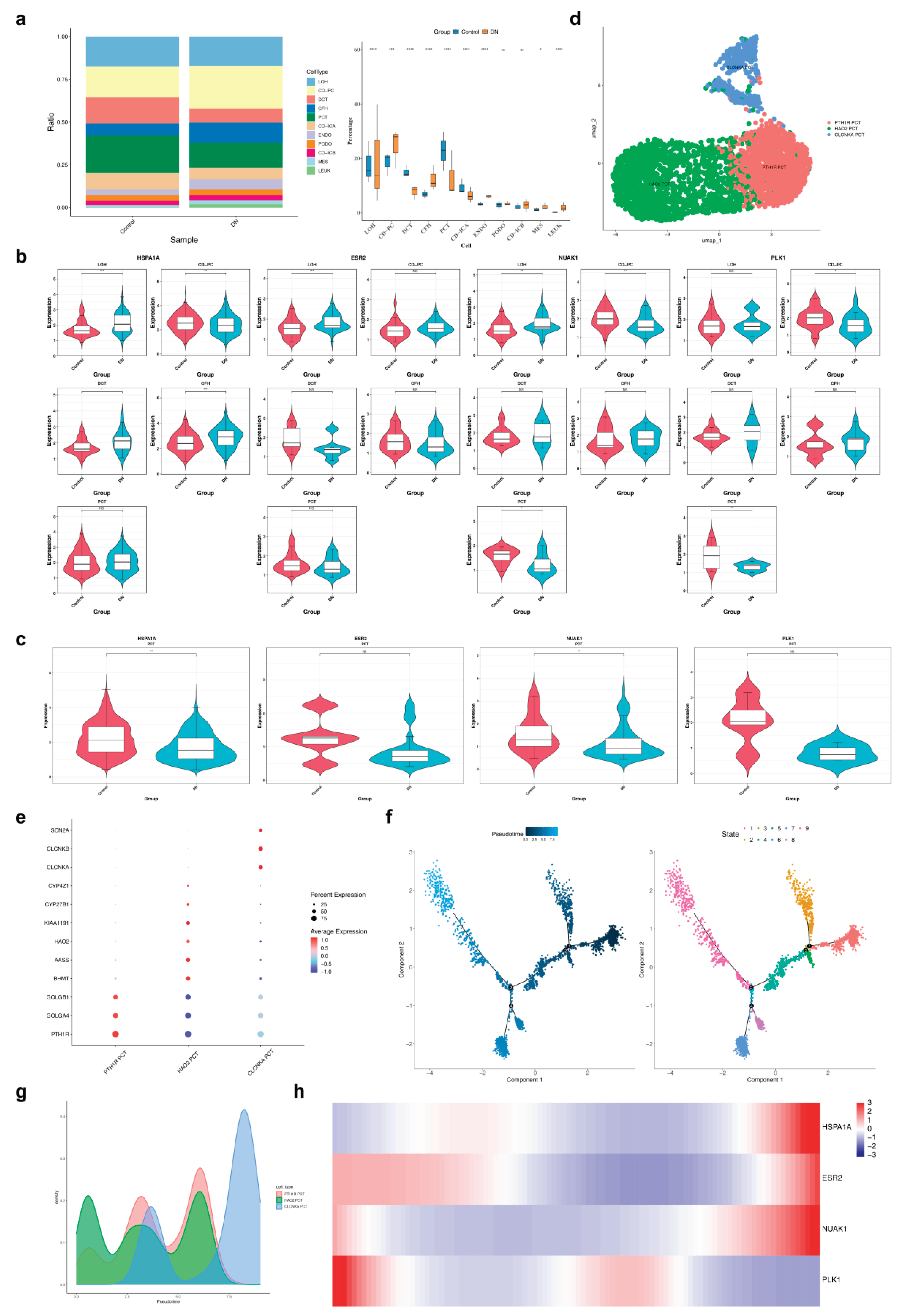
2.5. Identification of 11 renal cell types from scRNA-seq data
2.6. Functional enrichment and cell communication in DKD
2.7. Biomarkers associated with proximal tubule differentiation in DKD
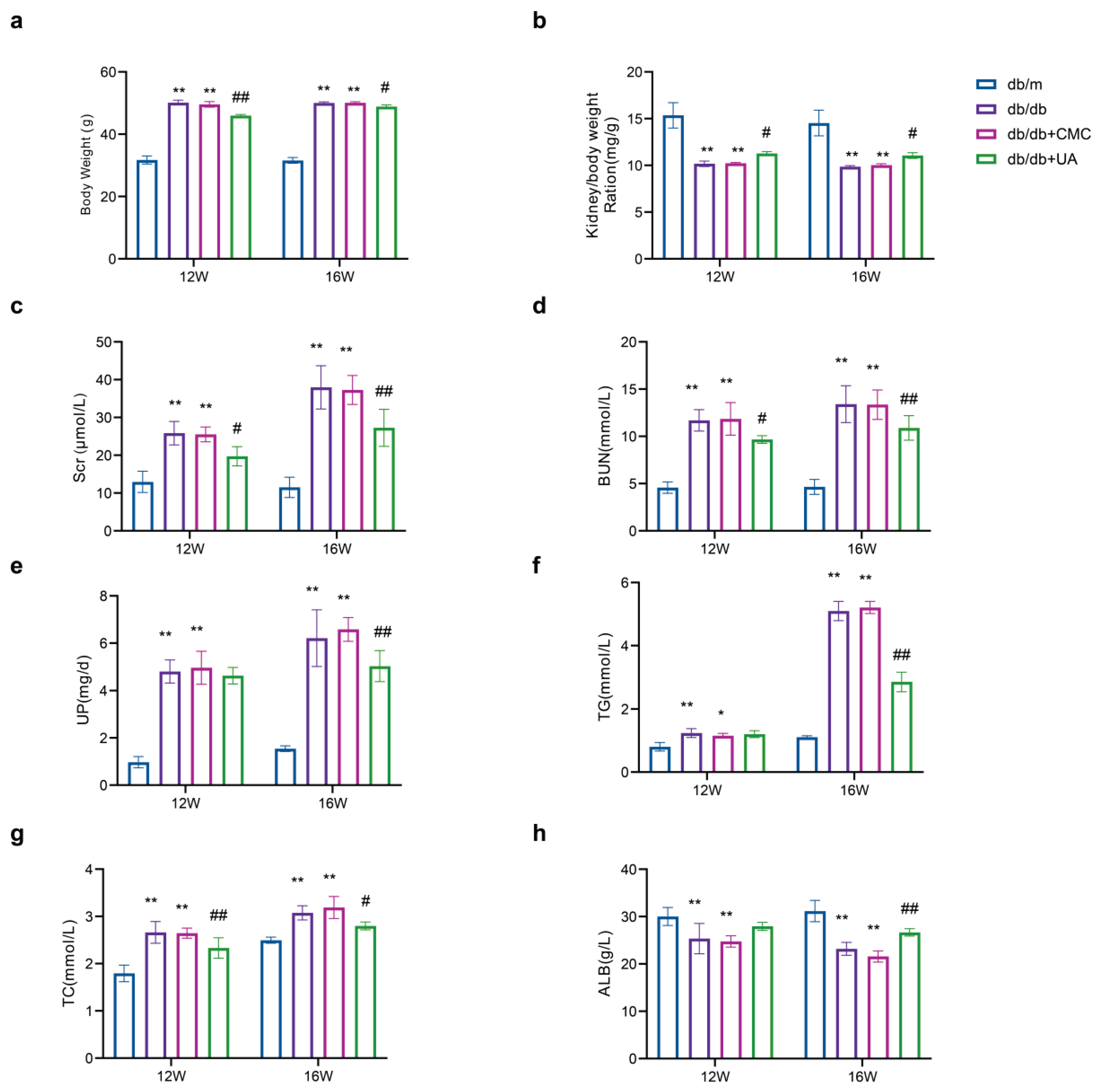
2.8. General condition and biochemical parameters in diabetic mice
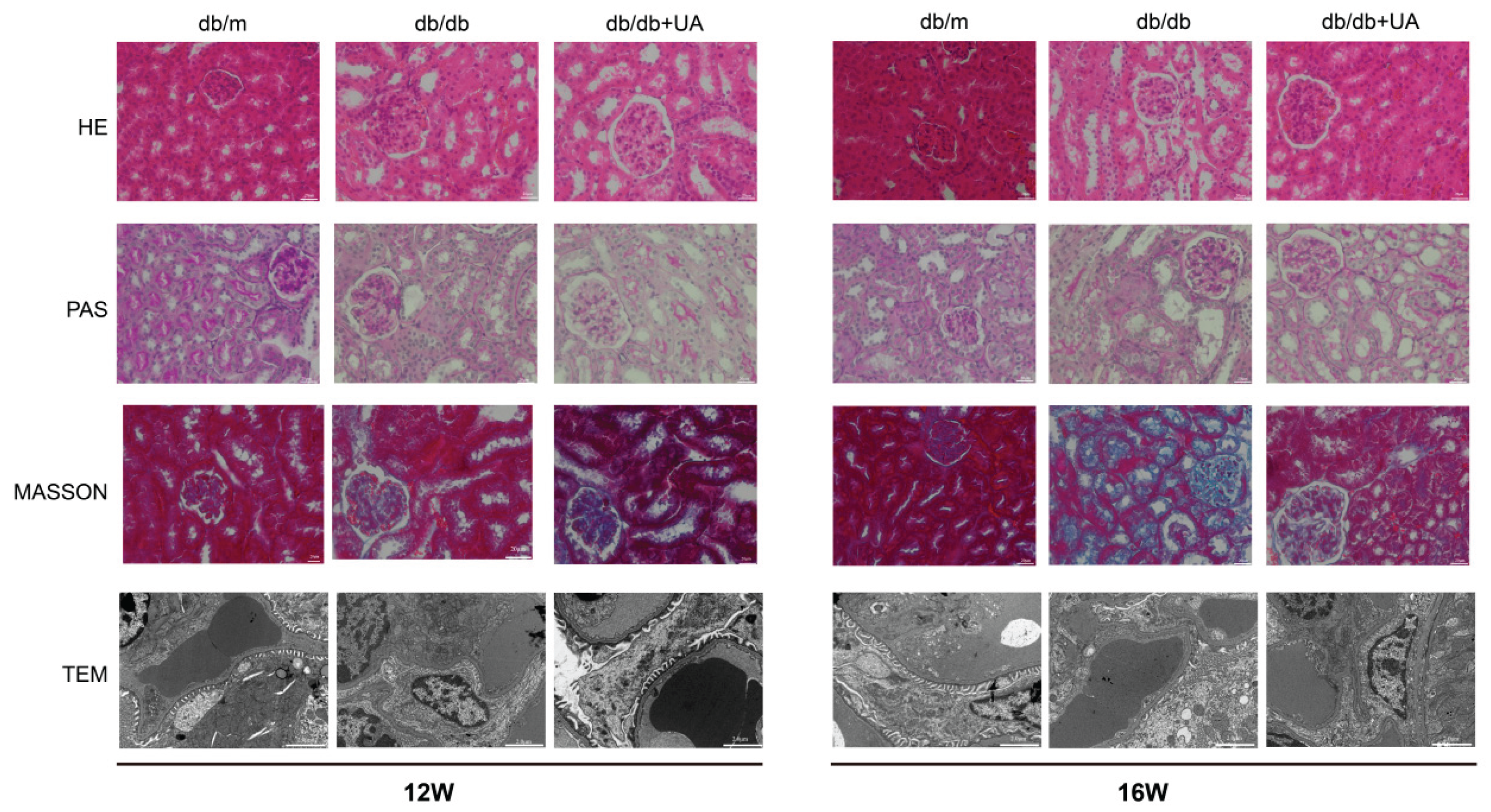
2.9. UA treatment ameliorates renal morphological alterations in diabetic mice
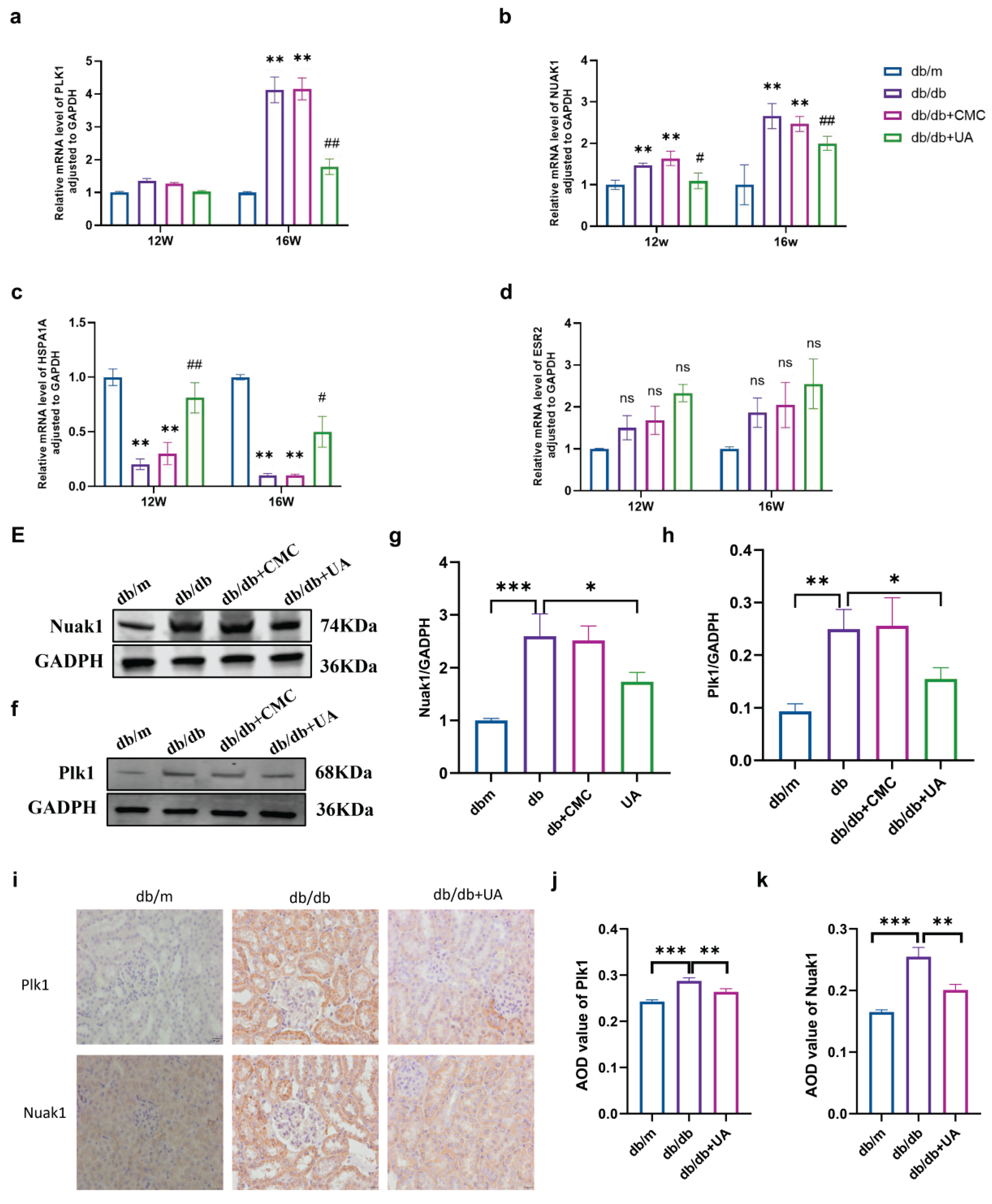
2.10. Expression level of hub genes in diabetic mice renal tissue
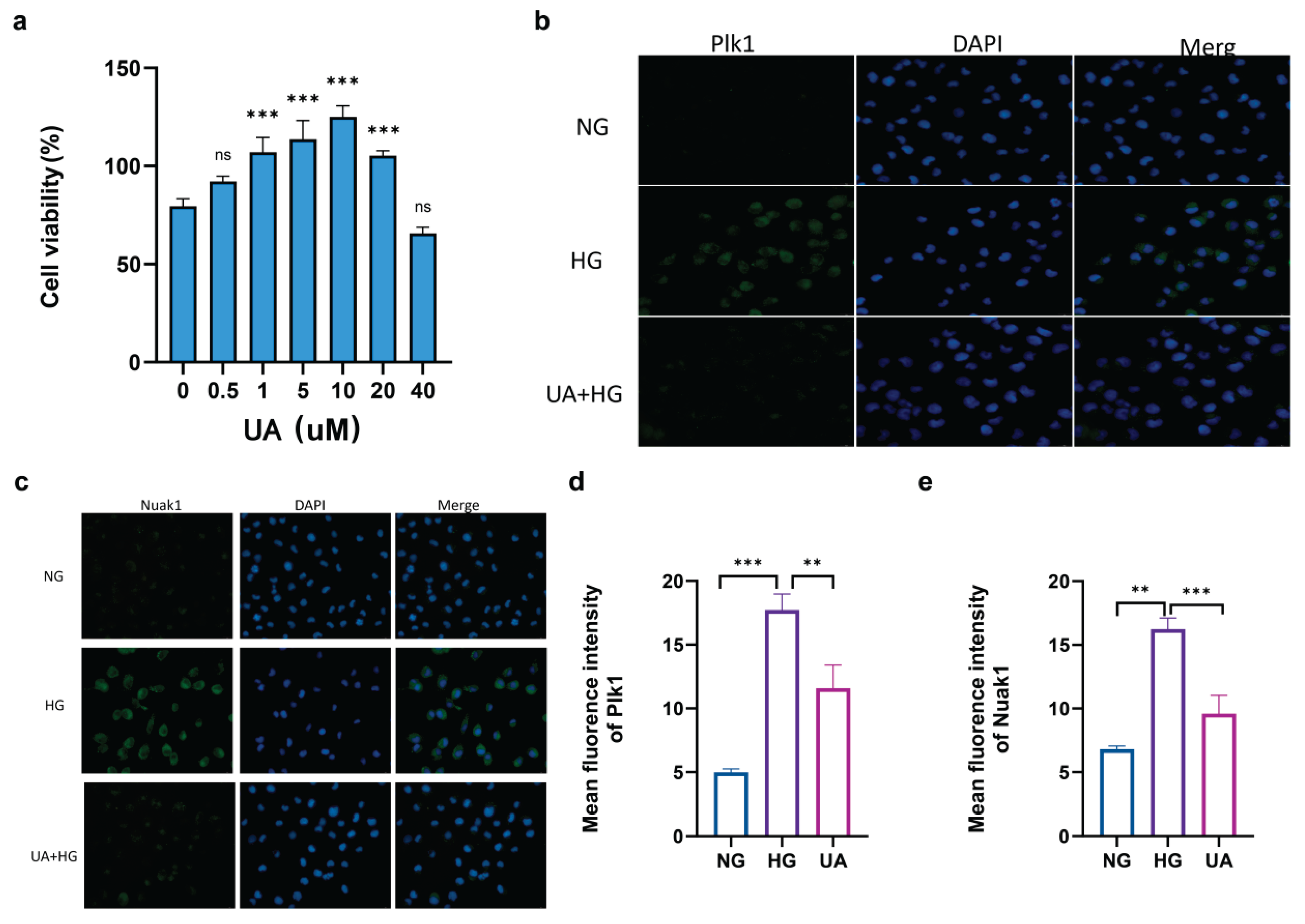
2.11. UA restored the altered expression of NUAK1 and PLK1 in HK-2 cells exposed to HG
3. Discussion
4. Conclusions
5. Materials and methods
5.1. Data collection
5.2. Differential expression analysis
5.3. Identification and functional analysis of urolithin A-related DEGs
5.4. Identification and validation of biomarkers
5.5. Gene set enrichment analysis (GSEA)
5.6. Immune infiltration analysis
5.7. Molecular docking
5.8. NephroSeq database analysis
5.9. Single-cell data quality control and high-variable gene screening
5.10. Cell dimension-reduction clustering, and annotation
5.11. Functional enrichment analysis and cell communication analysis
5.12. Identification of key cells and pseudotiming analysis
5.13. Animal experiments
5.14. Biochemical assays
5.15. Electron microscopy
5.16. Pathological and immunohistochemical staining
5.17. Western blot analysis
5.18. Cell culture
5.19. Cell viability assay
5.20. Immunofluorescence
5.21. RNA extraction and quantitative PCR
5.22. Statistical analysis
Supplementary Materials
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Acknowledgments
Conflicts of Interest
Abbreviations
| Abbreviations | Full Name |
| UA | urolithin A |
| DKD | diabetic kidney diseas |
| ROC | receiver operating characteristic |
| GSEA | gene set enrichment analysis |
| PCT | proximal convoluted tubules |
| ESRD | end-stage renal disease |
| GO | Gene Ontology |
| KEGG | Kyoto Encyclopedia of Genes and Genomes |
| PPI | protein-protein interaction |
| STRING | search tool for the retrieval of interacting genes |
| LASSO | least absolute shrinkage and selection operator |
| SVM-RFE | support vector machine recursive feature elimination |
| AUC | area under the curve |
| MSigDB | Molecular Signatures Database |
| PDB | Protein Data Bank |
| GFR | glomerular filtration rate |
| UMI | unique molecular identifier |
| HVGs | highly variable genes |
| UMAP | uniform manifold approximation and projection |
| PCT | proximal convoluted tubules |
| ENDO | endothelial cells |
| PODO | podocytes |
| MES | mesangial cells |
| CD-ICA | collecting duct-intercalated A cells |
| CD-ICB | collecting duct-intercalated B cells |
| LOH | loops of Henle |
| CDPC | collecting duct-principal cells |
| DCT | distal convoluted tubules |
| CFH | complement factor H |
| AMPK | AMP-activated protein kinase |
| ROS | reactive oxygen species |
| HSP70 | HSPA1A encodes heat shock protein 70 |
| Erβ AA |
ESR2 encodes estrogen receptor beta Asiatic acid |
References
- Brosius, F. C.; Cherney, D.; Gee, P. O.; Harris, R. C.; Kliger, A. S.; Tuttle, K. R.; Quaggin, S. E. Transforming the Care of Patients with Diabetic Kidney Disease. Clin J Am Soc Nephrol 2021, 16, 1590–1600. [Google Scholar] [CrossRef]
- Gupta, S.; Dominguez, M.; Golestaneh, L. Diabetic Kidney Disease: An Update. Med Clin North Am 2023, 107, 689–705. [Google Scholar] [CrossRef]
- Winiarska, A.; Knysak, M.; Nabrdalik, K.; Gumprecht, J.; Stompor, T. Inflammation and Oxidative Stress in Diabetic Kidney Disease: The Targets for SGLT2 Inhibitors and GLP-1 Receptor Agonists. Int J Mol Sci 2021, 22. [Google Scholar] [CrossRef]
- Samsu, N. Diabetic Nephropathy: Challenges in Pathogenesis, Diagnosis, and Treatment. Biomed Res Int 2021, 2021, 1497449. [Google Scholar] [CrossRef] [PubMed]
- Espín, J. C.; Larrosa, M.; García-Conesa, M. T.; Tomás-Barberán, F. Biological significance of urolithins, the gut microbial ellagic Acid-derived metabolites: the evidence so far. Evid Based Complement Alternat Med 2013, 2013, 270418. [Google Scholar] [CrossRef] [PubMed]
- Doyle, B.; Griffiths, L. A. The metabolism of ellagic acid in the rat. Xenobiotica 1980, 10, 247–56. [Google Scholar] [CrossRef] [PubMed]
- Singh, A.; D’Amico, D.; Andreux, P. A.; Fouassier, A. M.; Blanco-Bose, W.; Evans, M.; Aebischer, P.; Auwerx, J.; Rinsch, C. Urolithin A improves muscle strength, exercise performance, and biomarkers of mitochondrial health in a randomized trial in middle-aged adults. Cell Rep Med 2022, 3, 100633. [Google Scholar] [CrossRef]
- Larrosa, M.; Gonzalez-Sarrias, A.; Yanez-Gascon, M. J.; Selma, M. V.; Azorin-Ortuno, M.; Toti, S.; Tomas-Barberan, F.; Dolara, P.; Espin, J. C. Anti-inflammatory properties of a pomegranate extract and its metabolite urolithin-A in a colitis rat model and the effect of colon inflammation on phenolic metabolism. J Nutr Biochem 2010, 21, 717–25. [Google Scholar] [CrossRef]
- Zhou, B.; Li, Q.; Wang, J.; Chen, P.; Jiang, S. Ellagic acid attenuates streptozocin induced diabetic nephropathy via the regulation of oxidative stress and inflammatory signaling. Food Chem Toxicol 2019, 123, 16–27. [Google Scholar] [CrossRef]
- Kotewicz, M.; Krauze-Baranowska, M.; Daca, A.; Ploska, A.; Godlewska, S.; Kalinowski, L.; Lewko, B. Urolithins Modulate the Viability, Autophagy, Apoptosis, and Nephrin Turnover in Podocytes Exposed to High Glucose. Cells 2022, 11. [Google Scholar] [CrossRef] [PubMed]
- Lewko, B.; Wodzinska, M.; Daca, A.; Ploska, A.; Obremska, K.; Kalinowski, L. Urolithin A Ameliorates the TGF Beta-Dependent Impairment of Podocytes Exposed to High Glucose. J Pers Med 2024, 14. [Google Scholar] [CrossRef]
- Wilson, P. C.; Wu, H.; Kirita, Y.; Uchimura, K.; Ledru, N.; Rennke, H. G.; Welling, P. A.; Waikar, S. S.; Humphreys, B. D. The single-cell transcriptomic landscape of early human diabetic nephropathy. Proc Natl Acad Sci U S A 2019, 116, 19619–19625. [Google Scholar] [CrossRef]
- Wang, Y.; Jin, M.; Cheng, C. K.; Li, Q. Tubular injury in diabetic kidney disease: molecular mechanisms and potential therapeutic perspectives. Front Endocrinol (Lausanne) 2023, 14, 1238927. [Google Scholar] [CrossRef] [PubMed]
- Janosevic, D.; De Luca, T.; Kidney Precision Medicine, P.; Eadon, M. T. The Kidney Precision Medicine Project and Single-Cell Biology of the Injured Proximal Tubule. Am J Pathol 2025, 195, 7–22. [Google Scholar] [CrossRef]
- Rego, S. M.; Snyder, M. P. High Throughput Sequencing and Assessing Disease Risk. Cold Spring Harb Perspect Med 2019, 9. [Google Scholar] [CrossRef]
- Berger, S. I.; Iyengar, R. Network analyses in systems pharmacology. Bioinformatics 2009, 25, 2466–72. [Google Scholar] [CrossRef] [PubMed]
- Kitchen, D. B.; Decornez, H.; Furr, J. R.; Bajorath, J. Docking and scoring in virtual screening for drug discovery: methods and applications. Nat Rev Drug Discov 2004, 3, 935–49. [Google Scholar] [CrossRef] [PubMed]
- Gao, Z.; S, A.; Li, X. M.; Li, X. L.; Sui, L. N. Identification of Key Candidate Genes and Chemical Perturbagens in Diabetic Kidney Disease Using Integrated Bioinformatics Analysis. Front Endocrinol (Lausanne) 2021, 12, 721202. [Google Scholar] [CrossRef]
- Ye, S.; Cheng, Z.; Zhuo, D.; Liu, S. Different Types of Cell Death in Diabetic Neuropathy: A Focus on Mechanisms and Therapeutic Strategies. Int J Mol Sci 2024, 25. [Google Scholar] [CrossRef]
- Chen, H.; Zhang, H.; Li, A. M.; Liu, Y. T.; Liu, Y.; Zhang, W.; Yang, C.; Song, N.; Zhan, M.; Yang, S. VDR regulates mitochondrial function as a protective mechanism against renal tubular cell injury in diabetic rats. Redox Biol 2024, 70, 103062. [Google Scholar] [CrossRef]
- Lanfranchi, M.; Yandiev, S.; Meyer-Dilhet, G.; Ellouze, S.; Kerkhofs, M.; Dos Reis, R.; Garcia, A.; Blondet, C.; Amar, A.; Kneppers, A.; Polveche, H.; Plassard, D.; Foretz, M.; Viollet, B.; Sakamoto, K.; Mounier, R.; Bourgeois, C. F.; Raineteau, O.; Goillot, E.; Courchet, J. The AMPK-related kinase NUAK1 controls cortical axons branching by locally modulating mitochondrial metabolic functions. Nat Commun 2024, 15, 2487. [Google Scholar] [CrossRef]
- Poullennec, K. G.; Jnoff, E.; Abendroth, J.; Bhuma, N.; Calmiano, M.; Calmus, L.; Cardenas, A.; Courade, J. P.; Delatour, C.; Hall, A.; de Haro, T.; Delker, S. L.; Demaude, T.; Gaikwad, N.; Ghavate, D.; Gholap, A. R.; Kierkowicz, M.; Le Mestre, R.; Van Hijfte, N.; Verheijden, S.; Vernerova, K.; De Wever, V.; Waghmode, N. Discovery of UCB9386: A Potent, Selective, and Brain-Penetrant Nuak1 Inhibitor Suitable for In Vivo Pharmacological Studies. J Med Chem 2024, 67, 20879–20910. [Google Scholar] [CrossRef]
- Seo, M. S.; Jung, K. H.; Kim, K.; Lee, J. E.; Han, B. S.; Ko, S.; Kim, J. H.; Hong, S.; Lee, S. H.; Hong, S. S. Discovery of a novel NUAK1 inhibitor against pancreatic cancer. Biomed Pharmacother 2022, 152, 113241. [Google Scholar] [CrossRef]
- Zhang, T.; He, X.; Caldwell, L.; Goru, S. K.; Ulloa Severino, L.; Tolosa, M. F.; Misra, P. S.; McEvoy, C. M.; Christova, T.; Liu, Y.; Atin, C.; Zhang, J.; Hu, C.; Vukosa, N.; Chen, X.; Krizova, A.; Kirpalani, A.; Gregorieff, A.; Ni, R.; Chan, K.; Gill, M. K.; Attisano, L.; Wrana, J. L.; Yuen, D. A. NUAK1 promotes organ fibrosis via YAP and TGF-beta/SMAD signaling. Sci Transl Med 2022, 14, eaaz4028. [Google Scholar] [CrossRef]
- Buensuceso, A.; Fritz, J. L.; Collins, O.; Valdes, Y. R.; Borrelli, M. J.; DiMattia, G. E.; Shepherd, T. G. Loss of LKB1-NUAK1 signalling enhances NF-kappaB activity in a spheroid model of high-grade serous ovarian cancer. Sci Rep 2022, 12, 3011. [Google Scholar] [CrossRef]
- Jin, Q.; Liu, T.; Qiao, Y.; Liu, D.; Yang, L.; Mao, H.; Ma, F.; Wang, Y.; Peng, L.; Zhan, Y. Oxidative stress and inflammation in diabetic nephropathy: role of polyphenols. Front Immunol 2023, 14, 1185317. [Google Scholar] [CrossRef] [PubMed]
- Guo, L.; Wu, P.; Li, Q.; Feng, Q.; Lin, X.; Luo, Y.; Wang, Y.; Wu, M.; Cai, F.; Zhang, J.; Hu, Y.; Wang, H.; Wang, Y.; Luo, S.; Tian, L.; Fan, X.; Wang, L.; Xue, Y.; Guan, M. NUAK1 Promotes Diabetic Kidney Disease by Accelerating Renal Tubular Senescence via the ROS/P53 Axis. Diabetes 2025, 74, 2405–2417. [Google Scholar] [CrossRef]
- Marable, S. S.; Chung, E.; Park, J. S. Hnf4a Is Required for the Development of Cdh6-Expressing Progenitors into Proximal Tubules in the Mouse Kidney. J Am Soc Nephrol 2020, 31, 2543–2558. [Google Scholar] [CrossRef]
- Iliaki, S.; Beyaert, R.; Afonina, I. S. Polo-like kinase 1 (PLK1) signaling in cancer and beyond. Biochem Pharmacol 2021, 193, 114747. [Google Scholar] [CrossRef] [PubMed]
- Su, S.; Chhabra, G.; Singh, C. K.; Ndiaye, M. A.; Ahmad, N. PLK1 inhibition-based combination therapies for cancer management. Transl Oncol 2022, 16, 101332. [Google Scholar] [CrossRef] [PubMed]
- Kalous, J.; Aleshkina, D. Multiple Roles of PLK1 in Mitosis and Meiosis. Cells 2023, 12. [Google Scholar] [CrossRef]
- Du, Y.; Shang, Y.; Qian, Y.; Guo, Y.; Chen, S.; Lin, X.; Cao, W.; Tang, X.; Zhou, A.; Huang, S.; Zhang, A.; Jia, Z.; Zhang, Y. Plk1 promotes renal tubulointerstitial fibrosis by targeting autophagy/lysosome axis. Cell Death Dis 2023, 14, 571. [Google Scholar] [CrossRef] [PubMed]
- Jiang, A.; Chen, S.; Yu, X.; Jia, Y.; Sun, J.; Bian, Y.; Du, X.; Gu, X. Polo-like kinase 1 drives hypoxia-induced renal fibrosis via PTEN/PGK1-mediated glycolytic activation. Int J Biol Macromol 2025, 319, 145305. [Google Scholar] [CrossRef] [PubMed]
- Wang, M.; Huang, Z.; Li, X.; He, P.; Sun, H.; Peng, Y.; Fan, Q. Apabetalone, a BET protein inhibitor, inhibits kidney damage in diabetes by preventing pyroptosis via modulating the P300/H3K27ac/PLK1 axis. Pharmacol Res 2024, 207, 107306. [Google Scholar] [CrossRef] [PubMed]
- Zhu, B.; Hu, Y.; Wu, R.; Yu, Q.; Wen, W. FBXO45 levels regulated ferroptosis renal tubular epithelial cells in a model of diabetic nephropathy by PLK1. Open Med (Wars) 2024, 19, 20240971. [Google Scholar] [CrossRef]
- Uehara-Watanabe, N.; Okuno-Ozeki, N.; Minamida, A.; Nakamura, I.; Nakata, T.; Nakai, K.; Yagi-Tomita, A.; Ida, T.; Ikeda, K.; Kitani, T.; Yamashita, N.; Kamezaki, M.; Kirita, Y.; Matoba, S.; Tamagaki, K.; Kusaba, T. Direct evidence of proximal tubular proliferation in early diabetic nephropathy. Sci Rep 2022, 12, 778. [Google Scholar] [CrossRef]
- Rosenzweig, R.; Nillegoda, N. B.; Mayer, M. P.; Bukau, B. The Hsp70 chaperone network. Nat Rev Mol Cell Biol 2019, 20, 665–680. [Google Scholar] [CrossRef]
- Liu, Y.; Zhou, L.; Xu, Y.; Li, K.; Zhao, Y.; Qiao, H.; Xu, Q.; Zhao, J. Heat Shock Proteins and Ferroptosis. Front Cell Dev Biol 2022, 10, 864635. [Google Scholar] [CrossRef] [PubMed]
- Wang, Q.; Ke, S.; Liu, Z.; Shao, H.; He, M.; Guo, J. HSPA5 Promotes the Proliferation, Metastasis and Regulates Ferroptosis of Bladder Cancer. Int J Mol Sci 2023, 24. [Google Scholar] [CrossRef]
- Lv, M.; Cai, Y.; Hou, W.; Peng, K.; Xu, K.; Lu, C.; Yu, W.; Zhang, W.; Liu, L. The RNA-binding protein SND1 promotes the degradation of GPX4 by destabilizing the HSPA5 mRNA and suppressing HSPA5 expression, promoting ferroptosis in osteoarthritis chondrocytes. Inflamm Res 2022, 71, 461–472. [Google Scholar] [CrossRef]
- Nargesi, A. A.; Shalchi, M.; Nargesi, R. A.; Sadeghpour, N.; Zarifkar, M.; Mozaffari, M.; Imani, M.; Esteghamati, A.; Nakhjavani, M. The lost correlation between heat shock protein 70 (HSPA1A) and plasminogen activator inhibitor-1 in patients with type 2 diabetes and albuminuria. Cell Stress Chaperones 2016, 21, 361–5. [Google Scholar] [CrossRef] [PubMed]
- Barutta, F.; Pinach, S.; Giunti, S.; Vittone, F.; Forbes, J. M.; Chiarle, R.; Arnstein, M.; Perin, P. C.; Camussi, G.; Cooper, M. E.; Gruden, G. Heat shock protein expression in diabetic nephropathy. Am J Physiol Renal Physiol 2008, 295, F1817–24. [Google Scholar] [CrossRef]
- Zhang, J.; Cai, Y.; Qin, Y.; Liu, J.; Ding, J.; Xu, M.; Yang, L.; Zheng, Y.; Zhang, X. Heat shock protein 70 promotes the progression of type 2 diabetic nephropathy by inhibiting T-cell immunoglobulin and mucin domain-3 and thereby promoting Th17/Treg imbalance. Nephrology (Carlton) 2024, 29, 806–814. [Google Scholar] [CrossRef]
- Wang, Y.; Chen, Z.; Li, J.; Wen, Y.; Li, J.; Lv, Y.; Pei, Z.; Pei, Y. A Paramagnetic Metal-Organic Framework Enhances Mild Magnetic Hyperthermia Therapy by Downregulating Heat Shock Proteins and Promoting Ferroptosis via Aggravation of Two-Way Regulated Redox Dyshomeostasis. Adv Sci (Weinh) 2024, 11, e2306178. [Google Scholar] [CrossRef]
- Wang, C. H.; Chou, P. C.; Chung, F. T.; Lin, H. C.; Huang, K. H.; Kuo, H. P. Heat shock protein70 is implicated in modulating NF-kappaB activation in alveolar macrophages of patients with active pulmonary tuberculosis. Sci Rep 2017, 7, 1214. [Google Scholar] [CrossRef]
- Hulina, A.; Grdic Rajkovic, M.; Jaksic Despot, D.; Jelic, D.; Dojder, A.; Cepelak, I.; Rumora, L. Extracellular Hsp70 induces inflammation and modulates LPS/LTA-stimulated inflammatory response in THP-1 cells. Cell Stress Chaperones 2018, 23, 373–384. [Google Scholar] [CrossRef]
- Tukaj, S. Heat Shock Protein 70 as a Double Agent Acting Inside and Outside the Cell: Insights into Autoimmunity. Int J Mol Sci 2020, 21. [Google Scholar] [CrossRef]
- Arao, Y.; Korach, K. S. The physiological role of estrogen receptor functional domains. Essays Biochem 2021, 65, 867–875. [Google Scholar] [CrossRef]
- Gui, Y.; Palanza, Z.; Gupta, P.; Li, H.; Pan, Y.; Wang, Y.; Hargis, G.; Kreutzer, D. L.; Wang, Y.; Bastacky, S. I.; Liu, Y.; Liu, S.; Zhou, D. Calponin 2 regulates ketogenesis to mitigate acute kidney injury. JCI Insight 2023, 8. [Google Scholar] [CrossRef] [PubMed]
- Wells, C. C.; Riazi, S.; Mankhey, R. W.; Bhatti, F.; Ecelbarger, C.; Maric, C. Diabetic nephropathy is associated with decreased circulating estradiol levels and imbalance in the expression of renal estrogen receptors. Gend Med 2005, 2, 227–37. [Google Scholar] [CrossRef] [PubMed]
- Neugarten, J.; Acharya, A.; Lei, J.; Silbiger, S. Selective estrogen receptor modulators suppress mesangial cell collagen synthesis. Am J Physiol Renal Physiol 2000, 279, F309–18. [Google Scholar] [CrossRef]
- Lu, K.; Wang, L.; Fu, Y.; Li, G.; Zhang, X.; Cao, M. Bioinformatics analysis identifies immune-related gene signatures and subtypes in diabetic nephropathy. Front Endocrinol (Lausanne) 2022, 13, 1048139. [Google Scholar] [CrossRef]
- Zhang, F.; Wang, C.; Wen, X.; Chen, Y.; Mao, R.; Cui, D.; Li, L.; Liu, J.; Chen, Y.; Cheng, J.; Lu, Y. Mesenchymal stem cells alleviate rat diabetic nephropathy by suppressing CD103(+) DCs-mediated CD8(+) T cell responses. J Cell Mol Med 2020, 24, 5817–5831. [Google Scholar] [CrossRef]
- Zheng, J. M.; Yao, G. H.; Cheng, Z.; Wang, R.; Liu, Z. H. Pathogenic role of mast cells in the development of diabetic nephropathy: a study of patients at different stages of the disease. Diabetologia 2012, 55, 801–11. [Google Scholar] [CrossRef] [PubMed]
- Moon, J. Y.; Jeong, K. H.; Lee, T. W.; Ihm, C. G.; Lim, S. J.; Lee, S. H. Aberrant recruitment and activation of T cells in diabetic nephropathy. Am J Nephrol 2012, 35, 164–74. [Google Scholar] [CrossRef] [PubMed]
- Chen, J.; Liu, Q.; He, J.; Li, Y. Immune responses in diabetic nephropathy: Pathogenic mechanisms and therapeutic target. Front Immunol 2022, 13, 958790. [Google Scholar] [CrossRef] [PubMed]
- Romero, M.; Ortega, A.; Olea, N.; Arenas, M. I.; Izquierdo, A.; Bover, J.; Esbrit, P.; Bosch, R. J. Novel role of parathyroid hormone-related protein in the pathophysiology of the diabetic kidney: evidence from experimental and human diabetic nephropathy. J Diabetes Res 2013, 2013, 162846. [Google Scholar] [CrossRef]
- Lu, Y. A.; Liao, C. T.; Raybould, R.; Talabani, B.; Grigorieva, I.; Szomolay, B.; Bowen, T.; Andrews, R.; Taylor, P. R.; Fraser, D. Single-Nucleus RNA Sequencing Identifies New Classes of Proximal Tubular Epithelial Cells in Kidney Fibrosis. J Am Soc Nephrol 2021, 32, 2501–2516. [Google Scholar] [CrossRef]
- Kirita, Y.; Wu, H.; Uchimura, K.; Wilson, P. C.; Humphreys, B. D. Cell profiling of mouse acute kidney injury reveals conserved cellular responses to injury. Proc Natl Acad Sci U S A 2020, 117, 15874–15883. [Google Scholar] [CrossRef]
- Li, L.; Tao, M.; Gao, X.; Cao, Q.; Liao, Z.; Chen, F.; Yusufu, A.; Nie, H.; Zeng, Z.; Huang, K.; Deng, X.; Gao, P.; Wu, X. Uncovering key markers and therapeutic targets for renal fibrosis in diabetic kidney disease through bulk and single-cell RNA sequencing. J Transl Med 2025, 23, 742. [Google Scholar] [CrossRef]
- Wu, J.; Sun, Z.; Yang, S.; Fu, J.; Fan, Y.; Wang, N.; Hu, J.; Ma, L.; Peng, C.; Wang, Z.; Lee, K.; He, J. C.; Li, Q. Kidney single-cell transcriptome profile reveals distinct response of proximal tubule cells to SGLT2i and ARB treatment in diabetic mice. Mol Ther 2022, 30, 1741–1753. [Google Scholar] [CrossRef] [PubMed]
- Tan, W.; Chen, J.; Wang, Y.; Xiang, K.; Lu, X.; Han, Q.; Hou, M.; Yang, J. Single-cell RNA sequencing in diabetic kidney disease: a literature review. Ren Fail 2024, 46, 2387428. [Google Scholar] [CrossRef] [PubMed]
- Erekat, N. S. Programmed Cell Death in Diabetic Nephropathy: A Review of Apoptosis, Autophagy, and Necroptosis. Med Sci Monit 2022, 28, e937766. [Google Scholar] [CrossRef]
- Love, M. I.; Huber, W.; Anders, S. Moderated estimation of fold change and dispersion for RNA-seq data with DESeq2. Genome Biol 2014, 15, 550. [Google Scholar] [CrossRef]
- Gustavsson, E. K.; Zhang, D.; Reynolds, R. H.; Garcia-Ruiz, S.; Ryten, M. ggtranscript: an R package for the visualization and interpretation of transcript isoforms using ggplot2. Bioinformatics (Oxford, England) 2022, 38, 3844–3846. [Google Scholar]
- Gu, Z.; Eils, R.; Schlesner, M. Complex heatmaps reveal patterns and correlations in multidimensional genomic data. Bioinformatics 2016, 32, 2847–9. [Google Scholar] [CrossRef]
- Shannon, P.; Markiel, A.; Ozier, O.; Baliga, N. S.; Wang, J. T.; Ramage, D.; Amin, N.; Schwikowski, B.; Ideker, T. Cytoscape: a software environment for integrated models of biomolecular interaction networks. Genome Res 2003, 13, 2498–504. [Google Scholar] [CrossRef]
- Chen, H.; Boutros, P. C. VennDiagram: a package for the generation of highly-customizable Venn and Euler diagrams in R. BMC bioinformatics 2011, 12, 35. [Google Scholar] [CrossRef]
- Wu, T.; Hu, E.; Xu, S.; Chen, M.; Guo, P.; Dai, Z.; Feng, T.; Zhou, L.; Tang, W.; Zhan, L.; Fu, X.; Liu, S.; Bo, X.; Yu, G. clusterProfiler 4.0: A universal enrichment tool for interpreting omics data. Innovation (Camb) 2021, 2, 100141. [Google Scholar] [CrossRef]
- Li, Y.; Lu, F.; Yin, Y. Applying logistic LASSO regression for the diagnosis of atypical Crohn’s disease. Sci Rep 2022, 12, 11340. [Google Scholar] [CrossRef] [PubMed]
- Long, G.; Yang, C. A six-gene support vector machine classifier contributes to the diagnosis of pediatric septic shock. Mol Med Rep 2020, 21, 1561–1571. [Google Scholar] [CrossRef] [PubMed]
- Robin, X.; Turck, N.; Hainard, A.; Tiberti, N.; Lisacek, F.; Sanchez, J. C.; Müller, M. pROC: an open-source package for R and S+ to analyze and compare ROC curves. BMC bioinformatics 2011, 12, 77. [Google Scholar] [CrossRef]
- Robles-Jimenez, L. E.; Aranda-Aguirre, E.; Castelan-Ortega, O. A.; Shettino-Bermudez, B. S.; Ortiz-Salinas, R.; Miranda, M.; Li, X.; Angeles-Hernandez, J. C.; Vargas-Bello-Pérez, E.; Gonzalez-Ronquillo, M. Worldwide Traceability of Antibiotic Residues from Livestock in Wastewater and Soil: A Systematic Review. Animals (Basel) 2021, 12. [Google Scholar] [CrossRef] [PubMed]
- Craven, K. E.; Gökmen-Polar, Y.; Badve, S. S. CIBERSORT analysis of TCGA and METABRIC identifies subgroups with better outcomes in triple negative breast cancer. Sci Rep 2021, 11, 4691. [Google Scholar] [CrossRef]
- Liu, W.; Han, F.; Wan, M.; Yang, X. Z. Integrated bioinformatics analysis identifies shared immune changes between ischemic stroke and COVID 19. Front Immunol 2023, 14, 1102281. [Google Scholar] [CrossRef]
- Pereira, W. J.; Almeida, F. M.; Conde, D.; Balmant, K. M.; Triozzi, P. M.; Schmidt, H. W.; Dervinis, C.; Pappas, G. J., Jr.; Kirst, M. Asc-Seurat: analytical single-cell Seurat-based web application. BMC Bioinformatics 2021, 22, 556. [Google Scholar] [CrossRef]
- Griss, J.; Viteri, G.; Sidiropoulos, K.; Nguyen, V.; Fabregat, A.; Hermjakob, H. ReactomeGSA - Efficient Multi-Omics Comparative Pathway Analysis. Molecular & cellular proteomics: MCP 2020, 19, 2115–2125. [Google Scholar]
- Trapnell, C.; Cacchiarelli, D.; Grimsby, J.; Pokharel, P.; Li, S.; Morse, M.; Lennon, N. J.; Livak, K. J.; Mikkelsen, T. S.; Rinn, J. L. The dynamics and regulators of cell fate decisions are revealed by pseudotemporal ordering of single cells. Nat Biotechnol 2014, 32, 381–386. [Google Scholar] [CrossRef]
- Cheng, Q.; Chen, X.; Wu, H.; Du, Y. Three hematologic/immune system-specific expressed genes are considered as the potential biomarkers for the diagnosis of early rheumatoid arthritis through bioinformatics analysis. J Transl Med 2021, 19, 18. [Google Scholar] [CrossRef] [PubMed]
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2026 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).



