Submitted:
10 April 2026
Posted:
14 April 2026
You are already at the latest version
Abstract
Keywords:
Introduction
Materials and Methods
Cell lines and culture conditions
Plasmid construction for site-directed mutagenesis
Plasmid construction for CRISPR/Cas9 mediated gene knockout
CRISPR/Cas9 Prime Editing system construction
Genotyping
Whole transcriptome sequencing
CYP17A1, CYP19A1, CYP21A2 Enzyme Activity Assays
Western Blotting
Steroid Profiling
Statistical analysis
Results
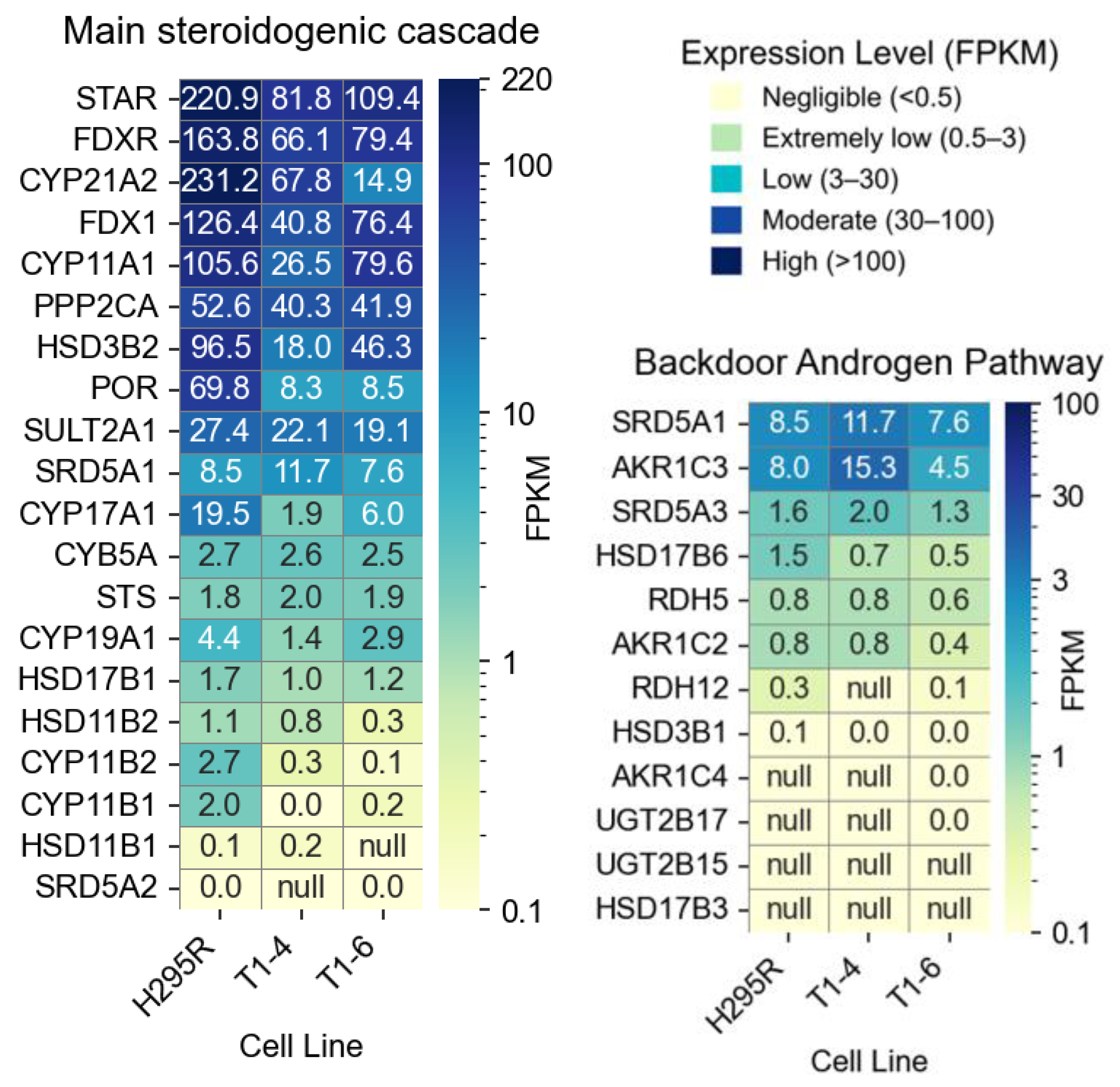
Whole transcriptome sequencing of the H295R and HEK293T POR knockout cell lines.
Steroid profile in POR knockout H295R cells.
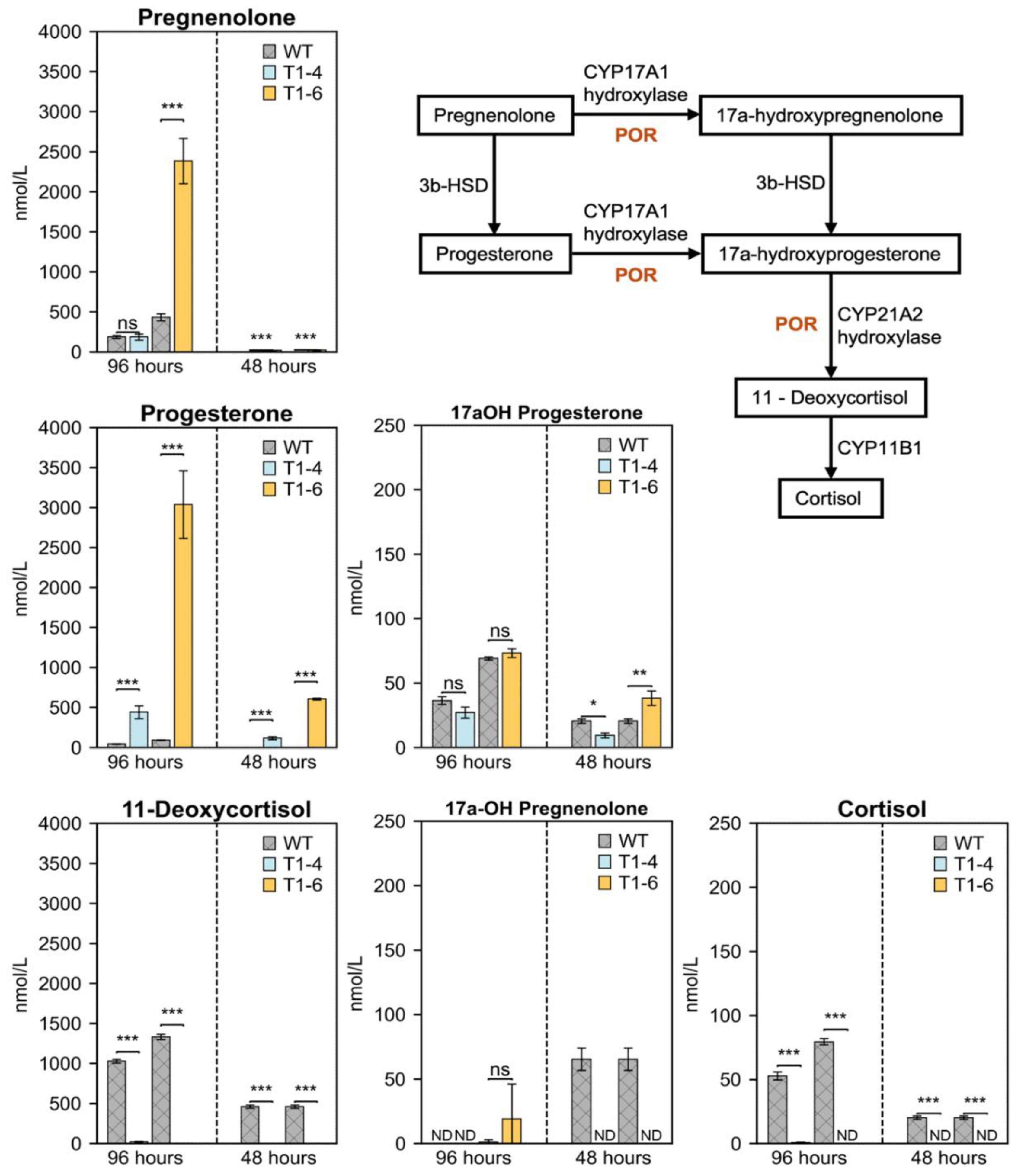
POR Deficiency Disrupts Glucocorticoid Pathway Intermediates in H295R Cells
POR Knockout Disrupts Aldosterone Pathway Intermediates in H295R Cells
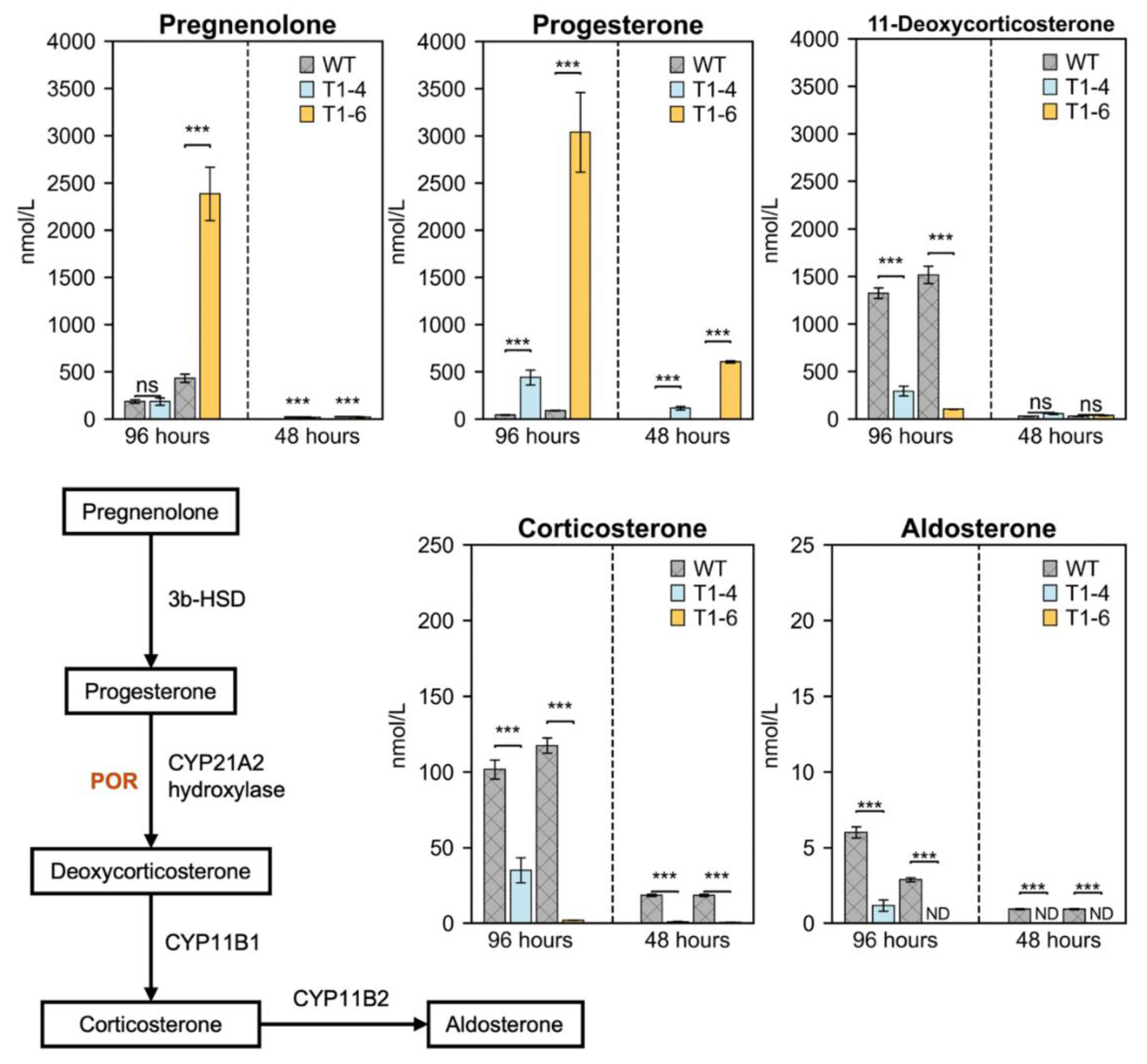
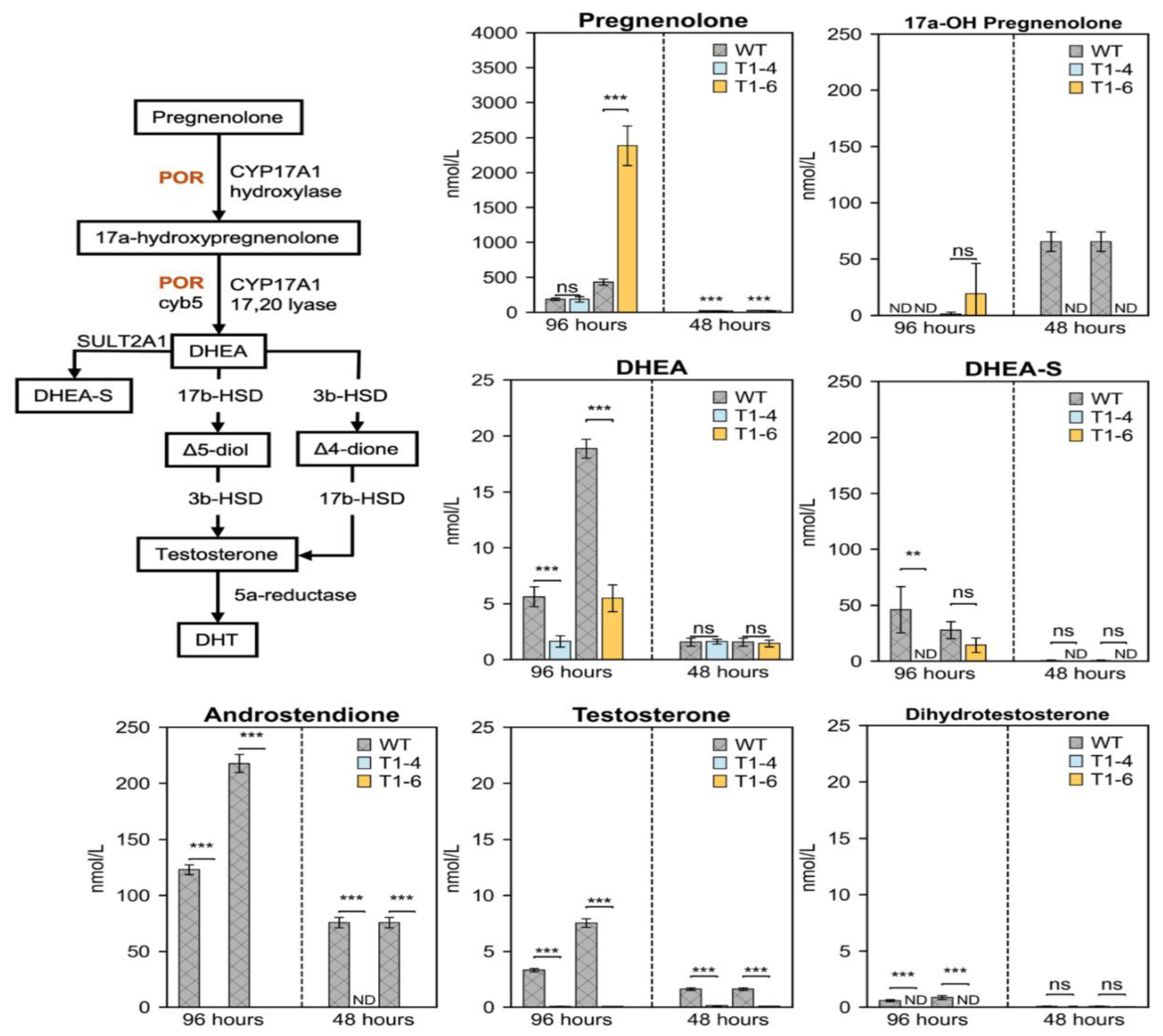
Metabolic Shunting of Progesterone in POR-Knockout Cells
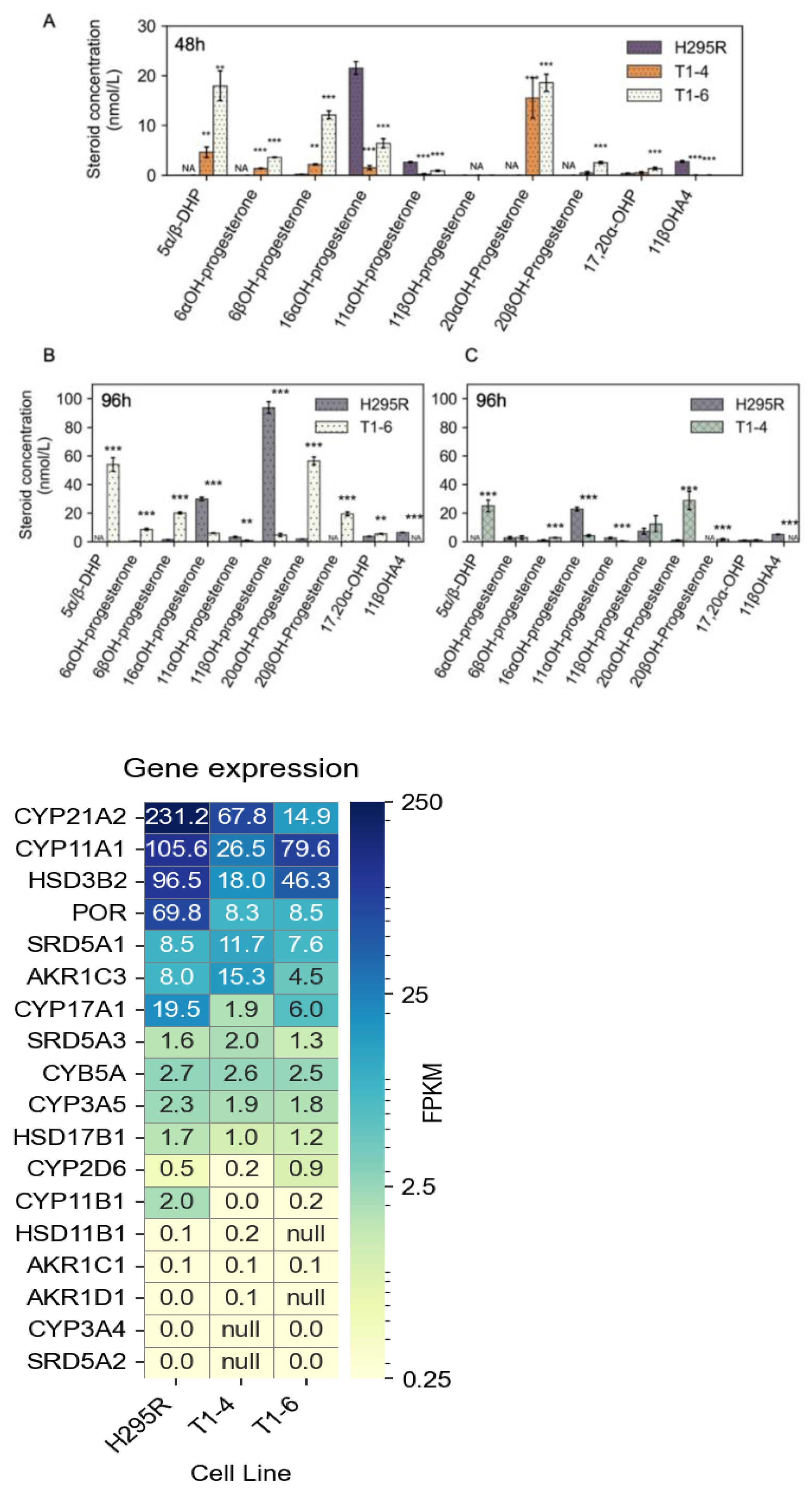
Steroid-metabolizing enzyme activities
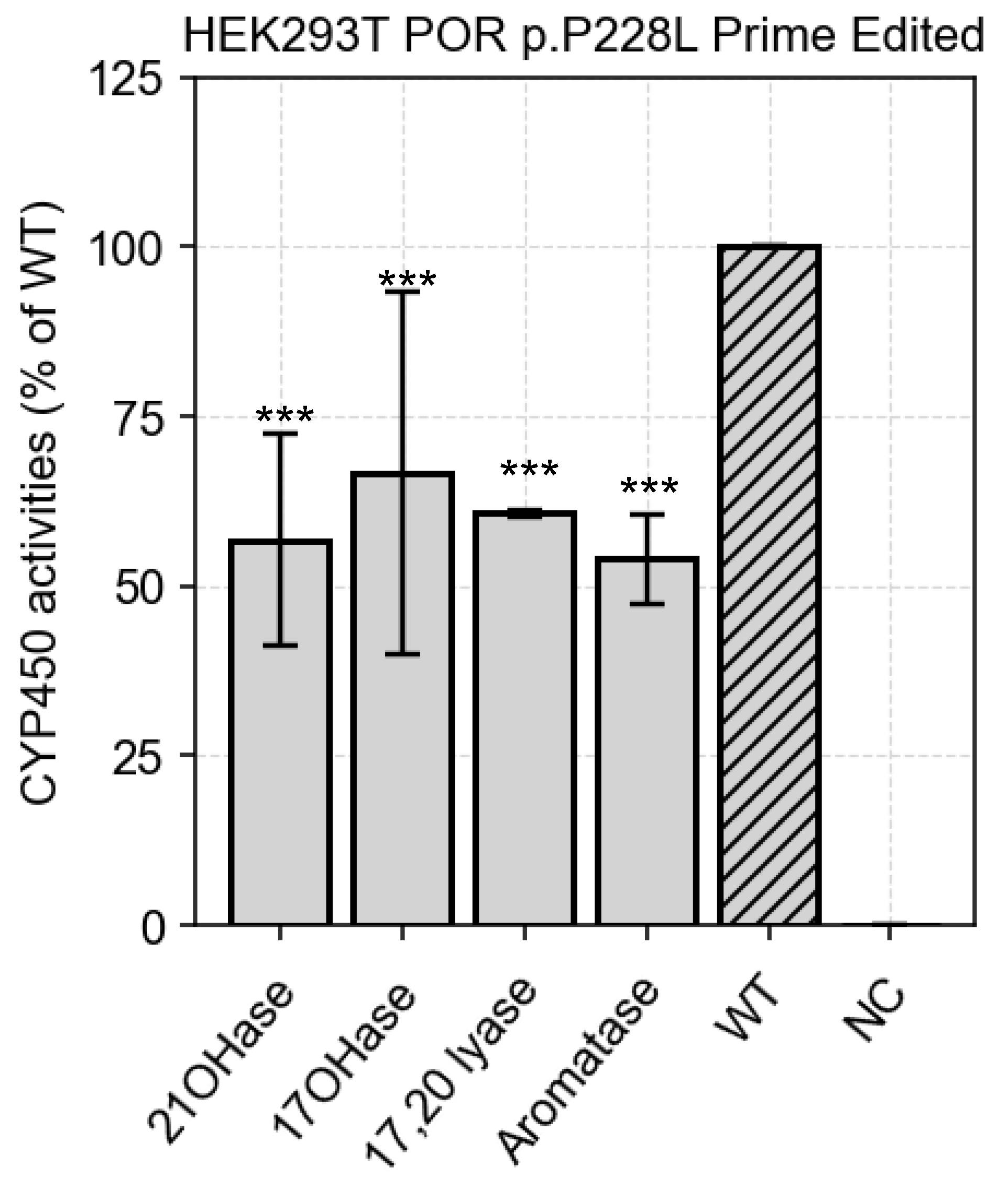
Prime edited cells
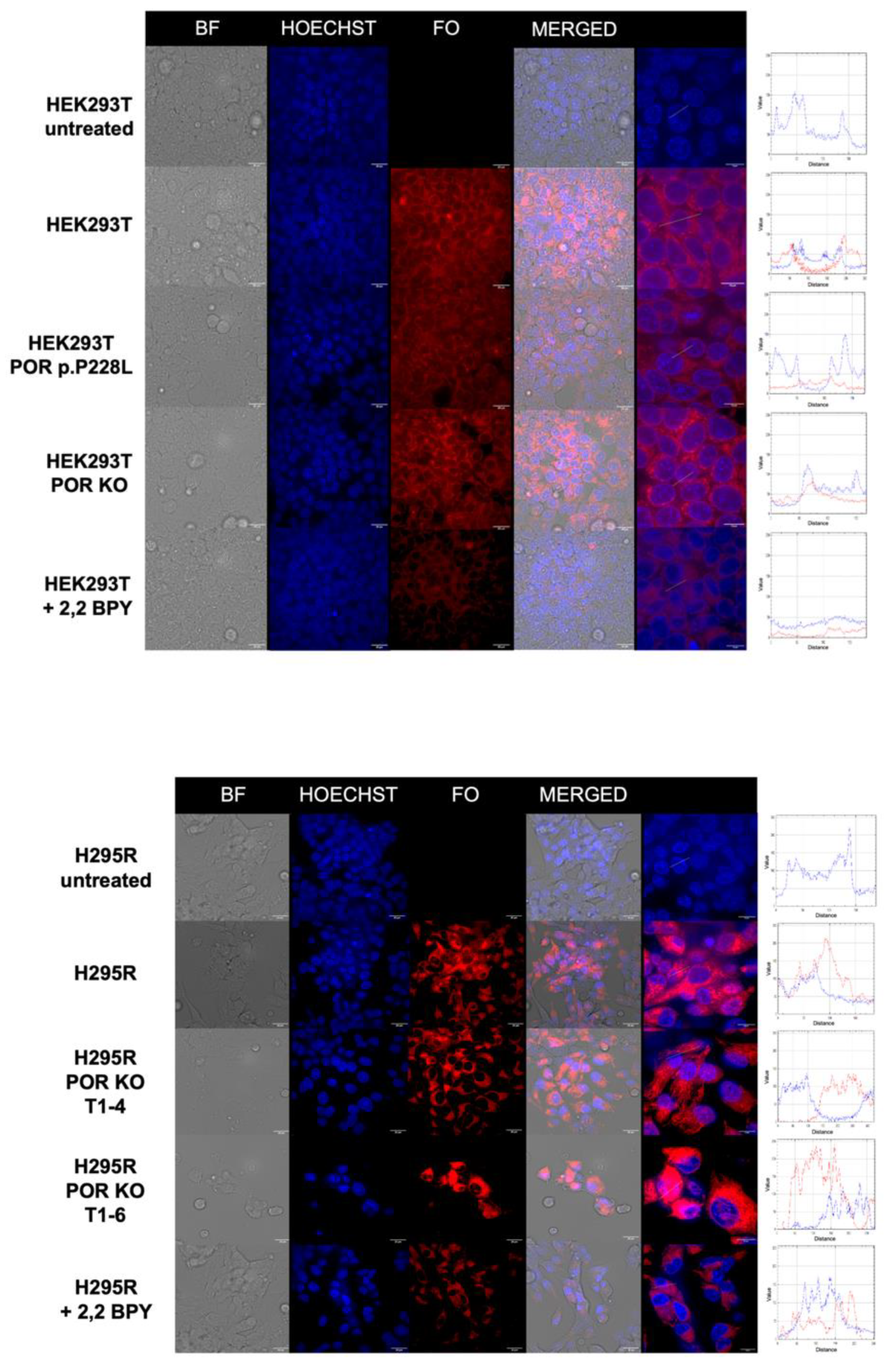
Confocal imaging of Fe(II) storage in POR knockout and prime-edited HEK293T and H295R cells.
Discussion
Funding
Acknowledgments
Disclosure Statement
References
- Pandey, V.; Flück, C. E. NADPH P450 oxidoreductase: Structure, function, and pathology of diseases. Pharmacology & Therapeutics 2013, 138, 229–254. [Google Scholar] [CrossRef]
- Riddick, D. S. NADPH–Cytochrome P450 Oxidoreductase: Roles in Physiology, Pharmacology, and Toxicology. Drug Metabolism and Disposition 2013, 41, 12–23. [Google Scholar] [CrossRef]
- Miller, W. L. Minireview: Regulation of Steroidogenesis by Electron Transfer. Endocrinology 2005, 146, 2544–2550. [Google Scholar] [CrossRef]
- Pandey, A.V.; Sproll, P. Pharmacogenomics of human P450 oxidoreductase. Frontiers in Pharmacology 2014, 5. [Google Scholar] [CrossRef] [PubMed]
- Miller, W. L.; Auchus, R. J. The Molecular Biology, Biochemistry, and Physiology of Human Steroidogenesis and Its Disorders. Endocrine Reviews 2011, 32, 81–151. [Google Scholar] [CrossRef] [PubMed]
- Xia et al., Structural basis for human NADPH-cytochrome P450 oxidoreductase deficiency. Proceedings of the National Academy of Sciences 2011, 108, 13486–13491. [CrossRef]
- Zhang, T. Clinical and genetic analysis of cytochrome P450 oxidoreductase (POR) deficiency in a female and the analysis of a novel POR intron mutation causing alternative mRNA splicing: Overall analysis of a female with POR deficiency. Journal of Assisted Reproduction and Genetics 2020, 37, 2503–2511. [Google Scholar] [CrossRef]
- Pandey, A.V.; Flück Ce Fau - Mullis, P. E.; Mullis, P. E. Altered heme catabolism by heme oxygenase-1 caused by mutations in human NADPH cytochrome P450 reductase. [CrossRef] [PubMed]
- Dean, B. P450 Oxidoreductase Deficiency: A Systematic Review and Meta-analysis of Genotypes, Phenotypes, and Their Relationships. The Journal of Clinical Endocrinology & Metabolism 2020, 105, e42–e52. [Google Scholar]
- Burkhard, F. Z.; Parween, S.; Udhane, S. S.; Flück, C. E.; Pandey, A. V. P450 Oxidoreductase deficiency: Analysis of mutations and polymorphisms. The Journal of Steroid Biochemistry and Molecular Biology 2017, 165, 38–50. [Google Scholar] [CrossRef]
- Krone, N. Genotype-Phenotype Analysis in Congenital Adrenal Hyperplasia due to P450 Oxidoreductase Deficiency. The Journal of Clinical Endocrinology & Metabolism 2012, 97, E257–E267. [Google Scholar] [CrossRef]
- Idkowiak, J. Pubertal Presentation in Seven Patients with Congenital Adrenal Hyperplasia due to P450 Oxidoreductase Deficiency. The Journal of Clinical Endocrinology & Metabolism 2011, 96, E453–E462. [Google Scholar] [CrossRef]
- Fukami, M. Cytochrome P450 Oxidoreductase Deficiency: Identification and Characterization of Biallelic Mutations and Genotype-Phenotype Correlations in 35 Japanese Patients. The Journal of Clinical Endocrinology & Metabolism 2009, 94, 1723–1731. [Google Scholar] [CrossRef]
- Miller, W. L.; Huang, N.; Pandey, A. V.; Flück, C. E.; Agrawal, V. P450 Oxidoreductase Deficiency: A New Disorder of Steroidogenesis. Annals of the New York Academy of Sciences 2005, 1061, 100–108. [Google Scholar] [CrossRef]
- Bai, Y.; Li, J.; Wang, X. Cytochrome P450 oxidoreductase deficiency caused by R457H mutation in POR gene in Chinese: case report and literature review. Journal of Ovarian Research 2017, 10, 16. [Google Scholar] [CrossRef]
- Williamson, L.; Arlt, W.; Shackleton, C.; Kelley, R. I.; Braddock, S. R. Linking Antley–Bixler syndrome and congenital adrenal hyperplasia: A novel case of P450 oxidoreductase deficiency. American Journal of Medical Genetics Part A 2006, 140A, 1797–1803. [Google Scholar] [CrossRef] [PubMed]
- Huang, N. Diversity and Function of Mutations in P450 Oxidoreductase in Patients with Antley-Bixler Syndrome and Disordered Steroidogenesis. The American Journal of Human Genetics 2005, 76, 729–749. [Google Scholar] [CrossRef] [PubMed]
- Arlt, W. Arlt, P450 oxidoreductase deficiency and Antley–Bixler syndrome. Reviews in Endocrine and Metabolic Disorders 2007, 8, 301–307. [Google Scholar] [CrossRef] [PubMed]
- Gusmano et al., Congenital adrenal hyperplasia, disorders of sex development, and infertility in patients with POR gene pathogenic variants: a systematic review of the literature. Journal of Endocrinological Investigation 2022, 46, 1–14.
- Kelley, R. I. Abnormal sterol metabolism in a patient with Antley-Bixler syndrome and ambiguous genitalia. American Journal of Medical Genetics 2002, 110, 95–102. [Google Scholar] [CrossRef]
- Shackleton et al., Biochemical diagnosis of Antley–Bixler syndrome by steroid analysis. American Journal of Medical Genetics Part A 2004, 128A, 223–231. [CrossRef]
- Nakanishi, K. P450 oxidoreductase deficiency with maternal virilization during pregnancy. Clinical and Experimental Obstetrics & Gynecology 2016, 43, 902–904. [Google Scholar] [CrossRef]
- Puiu, M. A Case of Antley Bixler Syndrome: Diagnosis and Outcome. Acta Endocrinologica (Bucharest) 2012, 8, 479–484. [Google Scholar] [CrossRef]
- Flück, C. E.; Nicolo, C.; Pandey, A. V. Clinical, structural and functional implications of mutations and polymorphisms in human NADPH P450 oxidoreductase. Fundamental & Clinical Pharmacology 2007, 21, 399–410. [Google Scholar] [CrossRef]
- Homma, K. Urine Steroid Hormone Profile Analysis in Cytochrome P450 Oxidoreductase Deficiency: Implication for the Backdoor Pathway to Dihydrotestosterone. The Journal of Clinical Endocrinology & Metabolism 2006, 91, 2643–2649. [Google Scholar] [CrossRef]
- Khadilkar, K.; Jagtap, V.; Lila, A.; Bandgar, T.; Shah, N. Cytochrome P450 oxidoreductase deficiency: Novel cause of ambiguity with primary amenorrhea. Indian Journal of Endocrinology and Metabolism 2017, 21, 360. [Google Scholar] [CrossRef] [PubMed]
- Pan, P. Successful live birth in a Chinese woman with P450 oxidoreductase deficiency through frozen-thawed embryo transfer: a case report with review of the literature. Journal of Ovarian Research 2021, 14, 22. [Google Scholar] [CrossRef]
- Zhang; Ding, L.; Deng, S.; Tian, Q. Clinical Characteristics and Molecular Aetiology of Cytochrome P450 Oxidoreductase Deficiency Diagnosed in 46,XX Patients. Reproductive Sciences 2025, 32, 2474–2483. [Google Scholar] [CrossRef]
- Cragun, D. L. Undetectable maternal serum uE3 and postnatal abnormal sterol and steroid metabolism in Antley–Bixler syndrome. American Journal of Medical Genetics Part A 2004, 129A, 1–7. [Google Scholar] [CrossRef]
- Sahakitrungruang, T. Clinical, Genetic, and Enzymatic Characterization of P450 Oxidoreductase Deficiency in Four Patients. The Journal of Clinical Endocrinology & Metabolism 2009, 94, 4992–5000. [Google Scholar] [CrossRef]
- Adachi, M. Compound heterozygous mutations of cytochrome P450 oxidoreductase gene ( POR ) in two patients with Antley–Bixler syndrome. American Journal of Medical Genetics Part A 2004, 128A, 333–339. [Google Scholar] [CrossRef]
- Reisch, N. Prenatal Diagnosis of Congenital Adrenal Hyperplasia Caused by P450 Oxidoreductase Deficiency. The Journal of Clinical Endocrinology & Metabolism 2013, 98, E528–E536. [Google Scholar] [CrossRef]
- Gomes, A.M. Pharmacogenomics of Human Liver Cytochrome P450 Oxidoreductase: Multifactorial Analysis and Impact on Microsomal Drug Oxidation. Pharmacogenomics 2009, 10, 579–599. [Google Scholar] [CrossRef]
- Tomková, M. Genetic Variations in NADPH-CYP450 Oxidoreductase in a Czech Slavic Cohort. Pharmacogenomics 2015, 16, 205–215. [Google Scholar] [CrossRef]
- Saito, Y. Genetic Polymorphisms and Haplotypes of POR, Encoding Cytochrome P450 Oxidoreductase, in a Japanese Population. Drug Metabolism and Pharmacokinetics 2011, 26, 107–116. [Google Scholar] [CrossRef]
- Chen, S. A genomic mutational constraint map using variation in 76,156 human genomes. Nature 2024, 625, 92–100. [Google Scholar] [CrossRef]
- Byrska-Bishop, M. High-coverage whole-genome sequencing of the expanded 1000 Genomes Project cohort including 602 trios. Cell 2022, 185, 3426–3440.e3419. [Google Scholar] [CrossRef]
- Iyanagi, T.; Makino, R.; Anan, F. K. Studies on the microsomal mixed-function oxidase system: mechanism of action of hepatic NADPH-cytochrome P-450 reductase. Biochemistry 1981, 20, 1722–1730. [Google Scholar] [CrossRef] [PubMed]
- Wang, M. Three-dimensional structure of NADPH–cytochrome P450 reductase: Prototype for FMN- and FAD-containing enzymes. Proceedings of the National Academy of Sciences 1997, 94, 8411–8416. [Google Scholar] [CrossRef] [PubMed]
- Xia, C. Conformational Changes of NADPH-Cytochrome P450 Oxidoreductase Are Essential for Catalysis and Cofactor Binding. Journal of Biological Chemistry 2011, 286, 16246–16260. [Google Scholar] [CrossRef] [PubMed]
- Guengerich, P. Mechanisms of cytochrome P450 substrate oxidation: MiniReview. Journal of Biochemical and Molecular Toxicology 2007, 21, 163–168. [Google Scholar] [CrossRef] [PubMed]
- Behrendorff, J. B. Y. H. Reductive Cytochrome P450 Reactions and Their Potential Role in Bioremediation. Frontiers in Microbiology 2021, 12, 649273. [Google Scholar] [CrossRef]
- Rojas Velazquez, M. N.; Noebauer, M.; Pandey, A. V. Loss of Protein Stability and Function Caused by P228L Variation in NADPH-Cytochrome P450 Reductase Linked to Lower Testosterone Levels. International Journal of Molecular Sciences 2022, 23, 10141. [Google Scholar] [CrossRef]
- Flück, C. E. Deletion of P399_E401 in NADPH cytochrome P450 oxidoreductase results in partial mixed oxidase deficiency. Biochemical and Biophysical Research Communications 2011, 412, 572–577. [Google Scholar] [CrossRef]
- Kumar, R.; Jayaraman, M.; Ramadas, K.; Chandrasekaran, A. Computational identification and analysis of deleterious non-synonymous single nucleotide polymorphisms (nsSNPs) in the human POR gene: a structural and functional impact. Journal of Biomolecular Structure and Dynamics 2024, 42, 1518–1532. [Google Scholar] [CrossRef]
- Flück, C. E.; Mullis, P. E.; Pandey, A. V. Modeling of human P450 oxidoreductase structure by in silico mutagenesis and MD simulation. Molecular and Cellular Endocrinology 2009, 313, 17–22. [Google Scholar] [CrossRef] [PubMed]
- Nurhafizuddin, M.; Azizi, A.; Ming, L. C.; Shafqat, N. In Silico Analysis of PORD Mutations on the 3D Structure of P450 Oxidoreductase. Molecules 2022, 27, 4646. [Google Scholar] [CrossRef]
- McCammon, K. M. Instability of the Human Cytochrome P450 Reductase A287P Variant Is the Major Contributor to Its Antley-Bixler Syndrome-like Phenotype. Journal of Biological Chemistry 2016, 291, 20487–20502. [Google Scholar] [CrossRef]
- Pandey, A. V.; Kempná, P.; Hofer, G.; Mullis, P. E.; Flück, C. E. Modulation of Human CYP19A1 Activity by Mutant NADPH P450 Oxidoreductase. Molecular Endocrinology 2007, 21, 2579–2595. [Google Scholar] [CrossRef]
- Agrawal, V.; Huang, N.; Miller, W. L. Pharmacogenetics of P450 oxidoreductase: effect of sequence variants on activities of CYP1A2 and CYP2C19. Pharmacogenetics and Genomics 2008, 18, 569–576. [Google Scholar] [CrossRef] [PubMed]
- Shen, A.L.; O’Leary, K. A.; Kasper, C. B. Association of Multiple Developmental Defects and Embryonic Lethality with Loss of Microsomal NADPH-Cytochrome P450 Oxidoreductase. Journal of Biological Chemistry 2002, 277, 6536–6541. [Google Scholar] [CrossRef]
- Uehara, S. Humanized liver TK-NOG mice with functional deletion of hepatic murine cytochrome P450s as a model for studying human drug metabolism. Scientific Reports 2022, 12, 14907. [Google Scholar] [CrossRef] [PubMed]
- Heintze, T.; Klein, K.; Hofmann, U.; Zanger, U. M. Differential effects on human cytochromes P450 by CRISPR/Cas9-induced genetic knockout of cytochrome P450 reductase and cytochrome b5 in HepaRG cells. Scientific Reports 2021, 11, 1000. [Google Scholar] [CrossRef] [PubMed]
- Reed, L.; Jarvis, I. W. H.; Phillips, D. H.; Arlt, V. M. Enhanced DNA adduct formation by benzo(a)pyrene in human liver cells lacking cytochrome P450 oxidoreductase. Mutation Research/Genetic Toxicology and Environmental Mutagenesis 2020, 852, 503162. [Google Scholar] [CrossRef]
- Concordet, J.-P.; Haeussler, M. CRISPOR: intuitive guide selection for CRISPR/Cas9 genome editing experiments and screens. Nucleic Acids Research 2018, 46, W242–W245. [Google Scholar] [CrossRef]
- Labun, K. CHOPCHOP v3: expanding the CRISPR web toolbox beyond genome editing. Nucleic Acids Research 2019, 47, W171–W174. [Google Scholar] [CrossRef]
- Ran, A. Genome engineering using the CRISPR-Cas9 system. Nature Protocols 2013, 8, 2281–2308. [Google Scholar] [CrossRef]
- Menzorov, A.G.; Lukyanchikova, V. A.; Korablev, A. N.; Serova, I. A.; Fishman, V. S. Genome editing using CRISPR/ Cas9 system: a practical guide. Vavilov Journal of Genetics and Breeding 2016, 20, 930–944. [Google Scholar] [CrossRef]
- Brinkman, E. K.; Chen, T.; Amendola, M.; van Steensel, B. Easy quantitative assessment of genome editing by sequence trace decomposition. Nucleic Acids Research 2014, 42, e168–e168. [Google Scholar] [CrossRef]
- Happi Mbakam, C.; Rousseau, J.; Tremblay, G.; Yameogo, P.; Tremblay, J. P. Prime Editing Permits the Introduction of Specific Mutations in the Gene Responsible for Duchenne Muscular Dystrophy. International Journal of Molecular Sciences 2022, 23, 6160. [Google Scholar] [CrossRef]
- Mathis, N. Machine learning prediction of prime editing efficiency across diverse chromatin contexts. Nature Biotechnology 2025, 43, 712–719. [Google Scholar] [CrossRef]
- Paquet, D. Efficient introduction of specific homozygous and heterozygous mutations using CRISPR/Cas9. Nature 2016, 533, 125–129. [Google Scholar] [CrossRef]
- Anzalone, A. V. Search-and-replace genome editing without double-strand breaks or donor DNA. Nature 2019, 576, 149–157. [Google Scholar] [CrossRef]
- Lephart, E. D.; Simpson, E. R. “(45) Assay of aromatase activity” in Methods in enzymology; Elsevier, 1991; vol. 206, pp. 477–483. [Google Scholar]
- Prado, M. J. Characterization of Mutations Causing CYP21A2 Deficiency in Brazilian and Portuguese Populations. International Journal of Molecular Sciences 2021, 23, 296. [Google Scholar] [CrossRef]
- Sharma, K. Effect of Essential Oil Components on the Activity of Steroidogenic Cytochrome P450. Biomolecules 2024, 14, 203. [Google Scholar] [CrossRef]
- Cohen, M. In Silico Structural and Biochemical Functional Analysis of a Novel CYP21A2 Pathogenic Variant. International Journal of Molecular Sciences 2020, 21, 5857. [Google Scholar] [CrossRef]
- Andrieu, T.; Du Toit, T.; Vogt, B.; Mueller, M. D.; Groessl, M. Parallel targeted and non-targeted quantitative analysis of steroids in human serum and peritoneal fluid by liquid chromatography high-resolution mass spectrometry. Analytical and Bioanalytical Chemistry 2022, 414, 7461–7472. [Google Scholar] [CrossRef]
- Kempná, P.; Marti, N.; Udhane, S.; Flück, C. E. Regulation of androgen biosynthesis – A short review and preliminary results from the hyperandrogenic starvation NCI-H295R cell model. Molecular and Cellular Endocrinology 2015, 408, 124–132. [Google Scholar] [CrossRef] [PubMed]
- Duranova et al., Human adrenocortical carcinoma cell line (NCI-H295R): An in vitro screening model for the assessment of endocrine disruptors’ actions on steroidogenesis with an emphasis on cell ultrastructural features. Acta Histochemica 2022, 124, 151912. [CrossRef] [PubMed]
- du Toit, T.; Groessl, M.; Pignatti, E.; Swart, A. C.; Flück, C. E. Characterization of Steroid Metabolic Pathways in Established Human and Mouse Cell Models. International Journal of Molecular Sciences 2025, 26, 9721. [Google Scholar] [CrossRef] [PubMed]
- Kolatorova, L.; Vitku, J.; Suchopar, J.; Hill, M.; Parizek, A. Progesterone: A Steroid with Wide Range of Effects in Physiology as Well as Human Medicine. International Journal of Molecular Sciences 2022, 23, 7989. [Google Scholar] [CrossRef]
- Niwa, T. Effect of genetic polymorphism on the metabolism of endogenous neuroactive substances, progesterone and p-tyramine, catalyzed by CYP2D6. Molecular Brain Research 2004, 129, 117–123. [Google Scholar] [CrossRef] [PubMed]
- Quinney, S. K.; Benjamin, T.; Zheng, X.; Patil, A. S. Characterization of Maternal and Fetal CYP3A-Mediated Progesterone Metabolism. Fetal and Pediatric Pathology 2017, 36, 400–411. [Google Scholar] [CrossRef]
- Penning, T. M.; Wangtrakuldee, P.; Auchus, R. J. Structural and Functional Biology of Aldo-Keto Reductase Steroid-Transforming Enzymes. Endocrine Reviews 2019, 40, 447–475. [Google Scholar] [CrossRef]
- Storbeck, K.-H.; O’Reilly, M. W. The clinical and biochemical significance of 11-oxygenated androgens in human health and disease. European Journal of Endocrinology 2023, 188, R98–R109. [Google Scholar] [CrossRef]
- Bruque, C. D. Structure-based activity prediction of CYP21A2 stability variants: A survey of available gene variations. Scientific Reports 2016, 6, 39082. [Google Scholar] [CrossRef] [PubMed]
- Zou, Y. Cytochrome P450 oxidoreductase contributes tophospholipid peroxidation in ferroptosis. Nature Chemical Biology 2020, 16, 302–309. [Google Scholar] [CrossRef]
- Yan, B. Membrane Damage during Ferroptosis Is Caused by Oxidation of Phospholipids Catalyzed by the Oxidoreductases POR and CYB5R1. Molecular Cell 2021, 81, 355–369.e310. [Google Scholar] [CrossRef]
- Constable, E. C.; Housecroft, C. E. The Early Years of 2,2′-Bipyridine—A Ligand in Its Own Lifetime. Molecules 2019, 24, 3951. [Google Scholar] [CrossRef]
- Wang, C.; Tian, Q. Diagnostic challenges and management advances in cytochrome P450 oxidoreductase deficiency, a rare form of congenital adrenal hyperplasia, with 46, XX karyotype. Frontiers in Endocrinology 2023, 14, 1226387. [Google Scholar] [CrossRef] [PubMed]
- Papadakis, G. E. Non-classic cytochrome P450 oxidoreductase deficiency strongly linked with menstrual cycle disorders and female infertility as primary manifestations. Human Reproduction 2020, 35, 939–949. [Google Scholar] [CrossRef] [PubMed]
- Bayrak Demirel et al., Hemodynamic and microvascular abnormalities in P450 oxidoreductase deficiency: evidence for COX-dependent dysfunction. European Journal of Endocrinology 2026, 194, 264–274. [CrossRef] [PubMed]
- Thibeault, A.-A. H.; Deroy, K.; Vaillancourt, C.; Sanderson, J. T. A Unique Co-culture Model for Fundamental and Applied Studies of Human Fetoplacental Steroidogenesis and Interference by Environmental Chemicals. Environmental Health Perspectives 2014, 122, 371–377. [Google Scholar] [CrossRef]
- Strajhar, P. Steroid profiling in H295R cells to identify chemicals potentially disrupting the production of adrenal steroids. Toxicology 2017, 381, 51–63. [Google Scholar] [CrossRef]
- Roth, M. Y. Serum insulin-like factor 3 is highly correlated with intratesticular testosterone in normal men with acute, experimental gonadotropin deficiency stimulated with low-dose human chorionic gonadotropin: a randomized, controlled trial. Fertility and Sterility 2013, 99, 132–139. [Google Scholar] [CrossRef]
- Zheng, M. Progesterone and 17-hydroxy-progesterone concentrations in follicular fluid and serum reflect their production in granulosa and theca cells. Reproductive BioMedicine Online 2024, 49, 103853. [Google Scholar] [CrossRef]
- Miller, Walter L. Congenital Adrenal Hyperplasia: Time to Replace 17OHP with 21-Deoxycortisol. Hormone Research in Paediatrics 2019, 91, 416–420. [Google Scholar] [CrossRef]
- L. Claahsen - Van Der Grinten et al., Congenital Adrenal Hyperplasia—Current Insights in Pathophysiology, Diagnostics, and Management. Endocrine Reviews 2022, 43, 91–159. [CrossRef]
- Auchus, R. J. Overview of Dehydroepiandrosterone Biosynthesis. Seminars in Reproductive Medicine 2004, 22, 281–288. [Google Scholar] [CrossRef]
- Auchus, R. J. Steroid 17-hydroxylase and 17,20-lyase deficiencies, genetic and pharmacologic. The Journal of Steroid Biochemistry and Molecular Biology 2017, 165, 71–78. [Google Scholar] [CrossRef] [PubMed]
- Miller, W. L. Androgen biosynthesis from cholesterol to DHEA. Molecular and Cellular Endocrinology 2002, 198, 7–14. [Google Scholar] [CrossRef] [PubMed]
- Byrne, G. C.; Perry, Y. S.; Winter, J. S. D. Kinetic Analysis of Adrenal 3β-Hydroxysteroid Dehydrogenase Activity During Human Development*. The Journal of Clinical Endocrinology & Metabolism 1985, 60, 934–939. [Google Scholar] [CrossRef]
- Rhéaume, E. Structure and Expression of a New Complementary DNA Encoding the almost Exclusive 3β-Hydroxysteroid Dehydrogenase/Δ5 -Δ4 -lsomerase in Human Adrenals and Gonads. Molecular Endocrinology 1991, 5, 1147–1157. [Google Scholar] [CrossRef]
- Cascio, C.; Prasad, V. V. K.; Lin, Y. Y.; Lieberman, S.; Papadopoulos, V. Detection of P450c17-independent pathways for dehydroepiandrosterone (DHEA) biosynthesis in brain glial tumor cells. Proceedings of the National Academy of Sciences 1998, 95, 2862–2867. [Google Scholar] [CrossRef]
- Brown, R. C.; Cascio, C.; Papadopoulos, V. Pathways of Neurosteroid Biosynthesis in Cell Lines from Human Brain: Regulation of Dehydroepiandrosterone Formation by Oxidative Stress and β-Amyloid Peptide. Journal of Neurochemistry 2000, 74, 847–859. [Google Scholar] [CrossRef]
- Rammouz, G.; Lecanu, L.; Aisen, P.; Papadopoulos, V. A Lead Study on Oxidative Stress-Mediated Dehydroepiandrosterone Formation in Serum: The Biochemical Basis for a Diagnosis of Alzheimer’s Disease. Journal of Alzheimer’s Disease 2011, 24, 5–16. [Google Scholar] [CrossRef]
- Rammouz, G.; Lecanu, L.; Papadopoulos, V. Oxidative Stress-Mediated Brain Dehydroepiandrosterone (DHEA) Formation in Alzheimer’s Disease Diagnosis. Frontiers in Endocrinology 2011, 2. [Google Scholar] [CrossRef] [PubMed]
- Liere, P. Analysis of pregnenolone and dehydroepiandrosterone in rodent brain: cholesterol autoxidation is the key. Journal of Lipid Research 2009, 50, 2430–2444. [Google Scholar] [CrossRef] [PubMed]
- Zhu, Z. A bypass gateway from cholesterol to sex steroid biosynthesis circumnavigates CYP17A1. Nature Communications 2025, 16, 10611. [Google Scholar] [CrossRef]
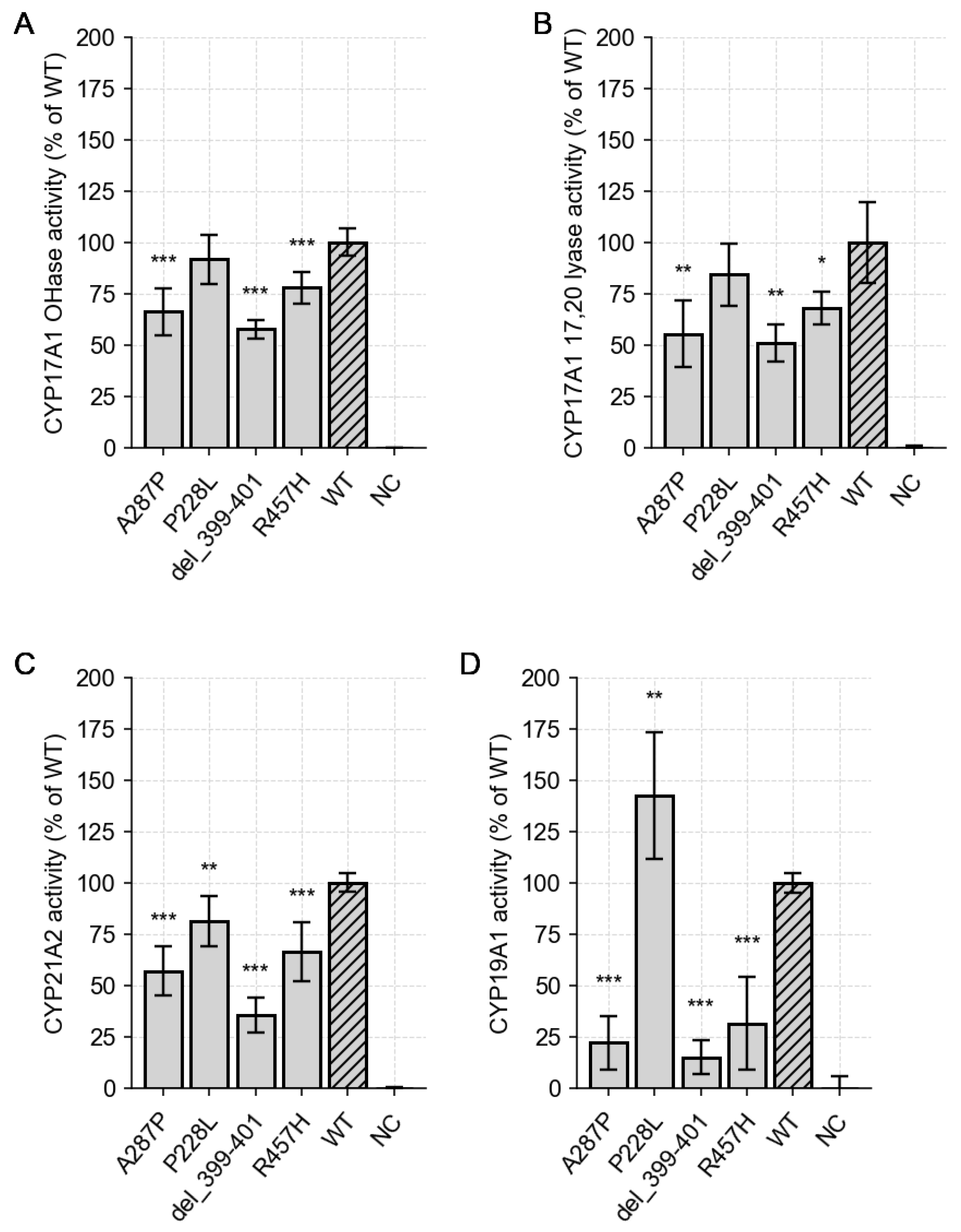
| CYP17A1 hydroxylase activity | ||
| Mutant | Mean % of CYP17A1 activity (±SD) | p-value (T-test) |
| A287P | 66.1 (± 11.4) | 9 x 10-5 (***) |
| P228L | 91.7 (± 12.0) | 0.13 (ns) |
| del399-401 | 57.7 (± 4.5) | 6 x 10-8 (***) |
| R457H | 78.0 (± 7.7) | 1 x 10-4 (***) |
| CYP17A1 17,20-lyase activity | ||
| Mutant | Mean % of CYP17A1 activity (±SD) | p-value (T-test) |
| A287P | 55.5 (± 16.4) | 0.008 (**) |
| P228L | 84.3 (± 15.1) | 0.194 (ns) |
| del399-401 | 51.1 (± 9.0) | 0.006 (**) |
| R457H | 68.1 (± 8.0) | 0.022 (*) |
| CYP21A2 hydroxylase activity | ||
| Mutant | Mean % of CYP21A2 activity (±SD) | p-value (T-test) |
| A287P | 57.1(± 11.9) | 4 x 10-5 (***) |
| P228L | 81.2 (± 12.3) | 0.006 (**) |
| del399-401 | 35.7 (± 8.4) | 5 x 10-8 (***) |
| R457H | 66.4 (± 14.4) | 7 x 10-4 (***) |
| CYP19A1 aromatase activity | ||
| Mutant | Mean % of CYP19A1 activity (±SD) | p-value (T-test) |
| A287P | 22.1 (± 12.9) | 2 x 10-9 (***) |
| P228L | 142.5 (± 30.8) | 0.001 (**) |
| del399-401 | 14.9 (± 8.2) | 1 x 10-13 (***) |
| R457H | 31.4 (± 22.5) | 4 x 10-6 (***) |
| POR c.P228L activities towards metabolic partners | ||
|---|---|---|
| Metabolic partner | Mean % of WT activity (±SD) | p-value (T-test) |
| CYP21A2 | 53.0 (± 12.3) | 0.001 (***) |
| CYP17A1 - OHase | 58.4 (± 21.9) | 0.013 (***) |
| CYP17A1 - 17, 20 lyase | 60.6 (± 0.5) | 5 x 10-5 (***) |
| CYP19A1 aromatase | 53.9 (± 6.7) | 0.007 (***) |
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2026 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).




