Submitted:
10 April 2026
Posted:
13 April 2026
You are already at the latest version
Abstract
Pet birds are increasingly recognized as reservoirs of zoonotic and antimicrobial-resistant bacteria, raising concerns within the One Health framework. However, data on bacterial diversity and resistance profiles in clinically affected ornamental birds remain limited. This study, conducted over three years (November 2022–March 2026), included 198 pet birds presenting with digestive and respiratory disorders. Samples were analyzed bacteriologically, with isolates identified by MALDI-TOF mass spectrometry, and antimicrobial susceptibility assessed using the Kirby-Bauer disk diffusion method according to EUCAST and CLSI guidelines. Bacterial growth was detected in 87.9% of cases, yielding 249 isolates across 26 genera. Gram-positive cocci predominated (62.3%), particularly Staphylococcus spp. (33.3%) and Enterococcus spp. (9.6%), while Escherichia coli (9.2%) was the most common Gram-negative species. Polymicrobial infections occurred in 39.1% of cases. High resistance rates were observed for tylosin (71.6%), oxytetracycline (68.4%), and trimethoprim–sulfamethoxazole (63.4%), whereas amikacin showed the highest susceptibility (58%). Overall, 57% of isolates were multidrug-resistant, with Staphylococcus spp. contributing most to this burden. These findings highlight pet birds as important reservoirs of antimicrobial-resistant bacteria and highlight the need for routine diagnostics and improved antimicrobial stewardship in avian medicine.
Keywords:
1. Introduction
2. Results
2.1. Clinical Presentation
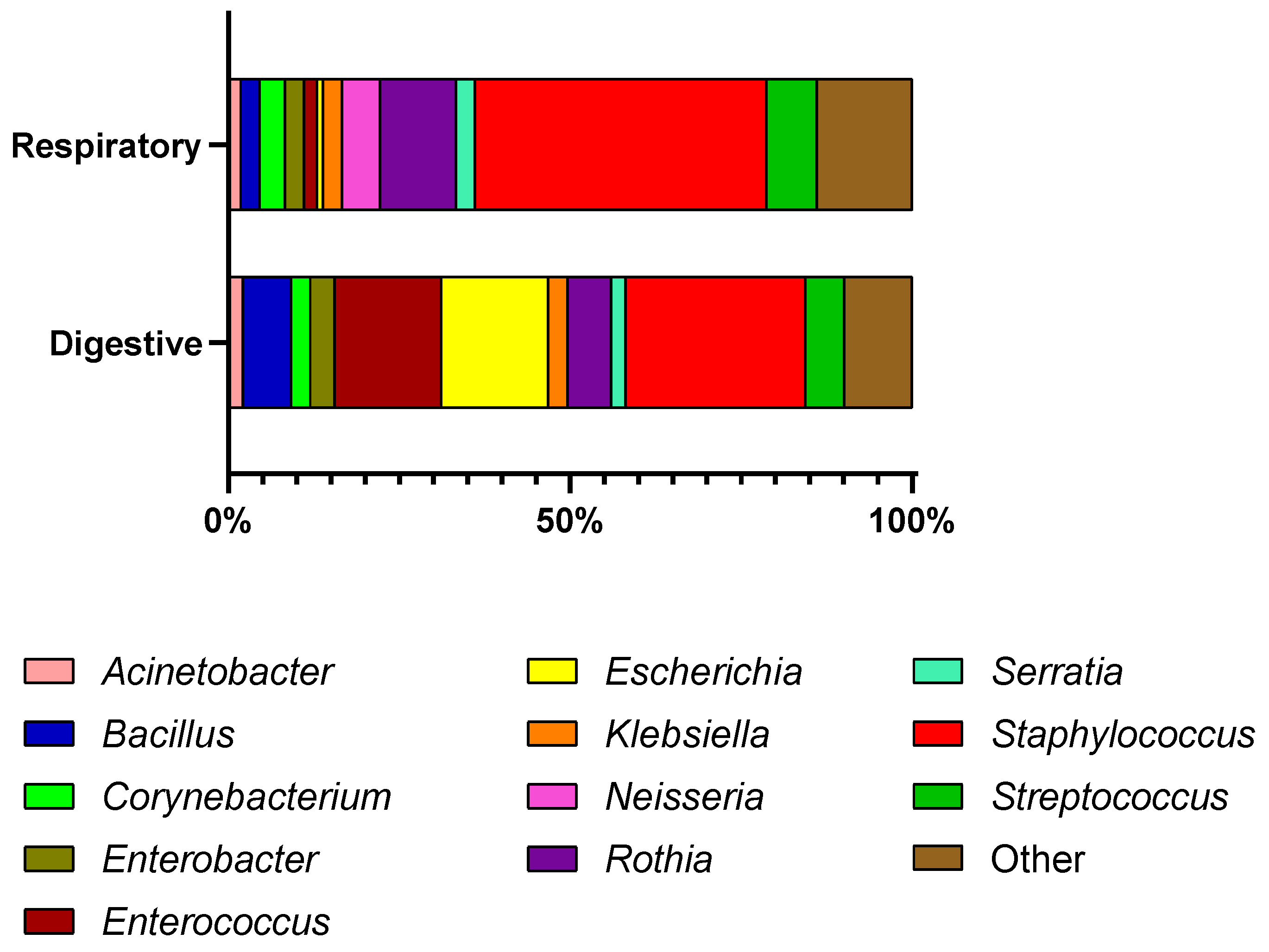
2.2. Bacterial Identification
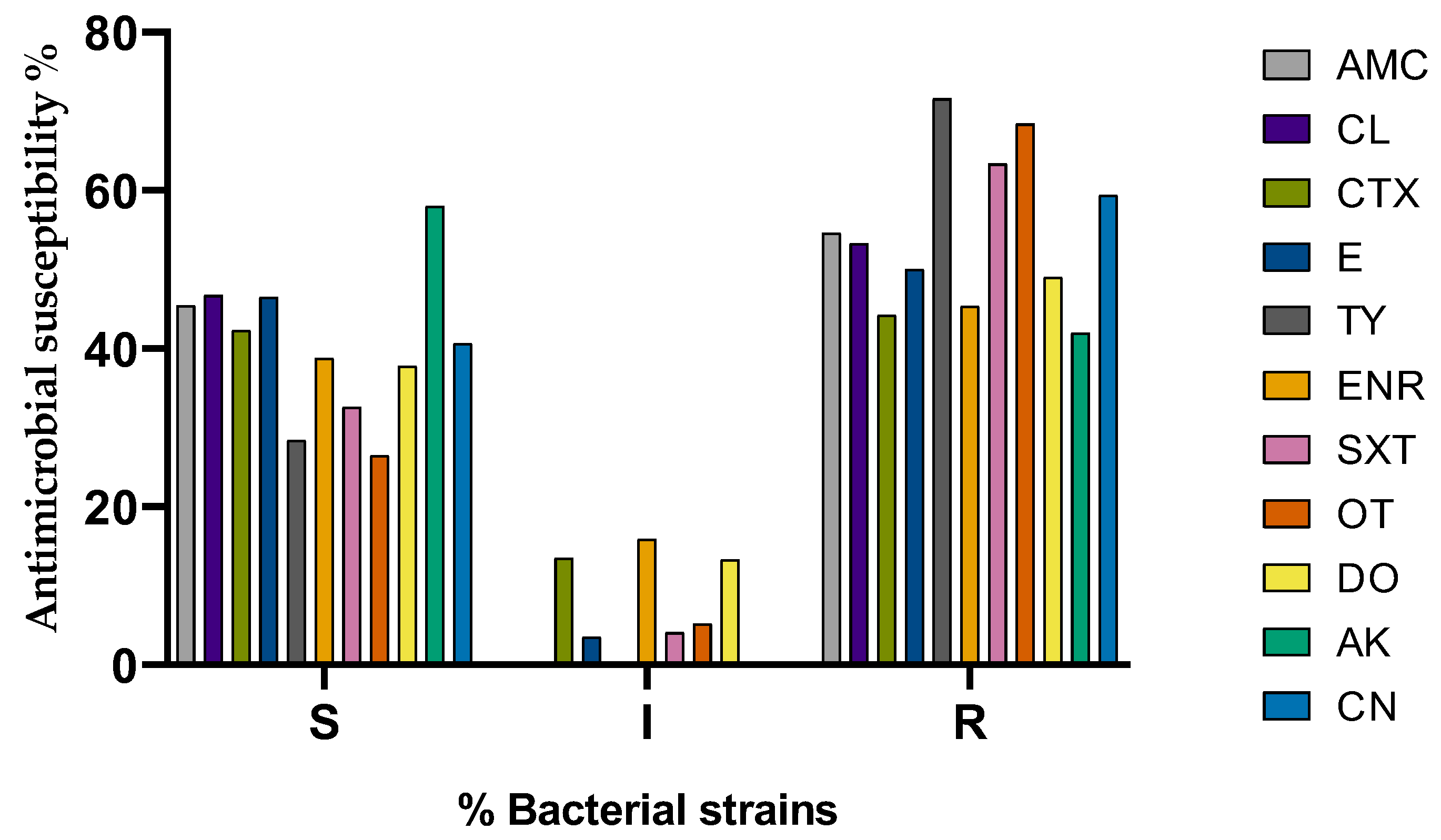
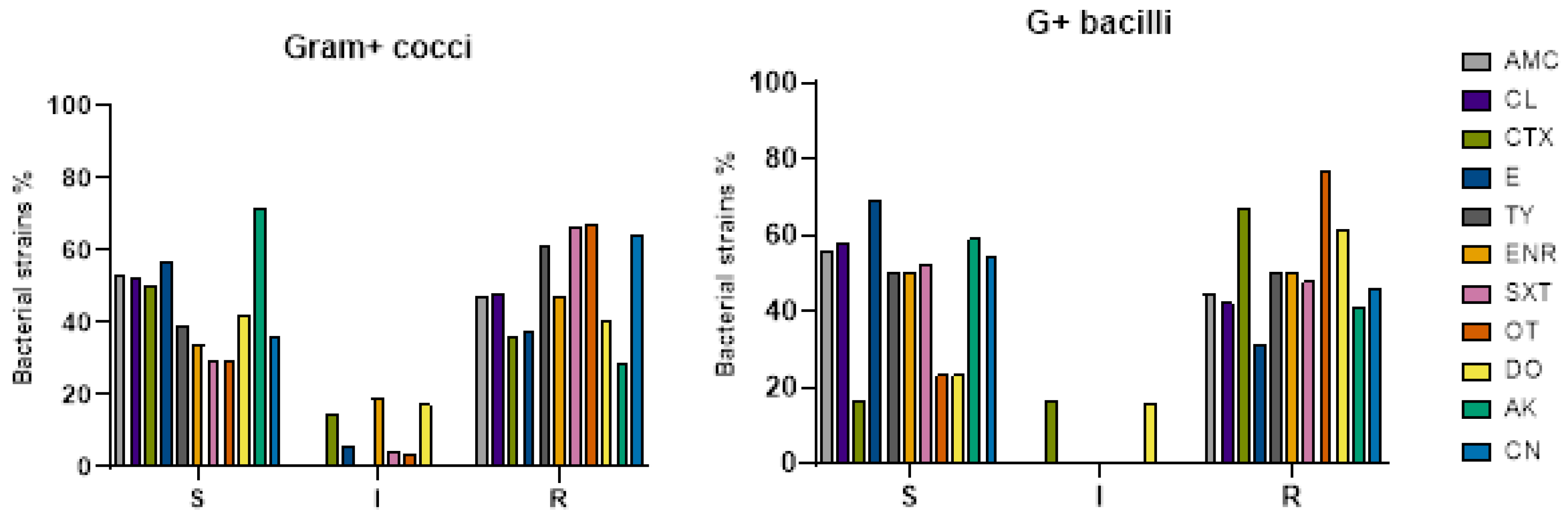
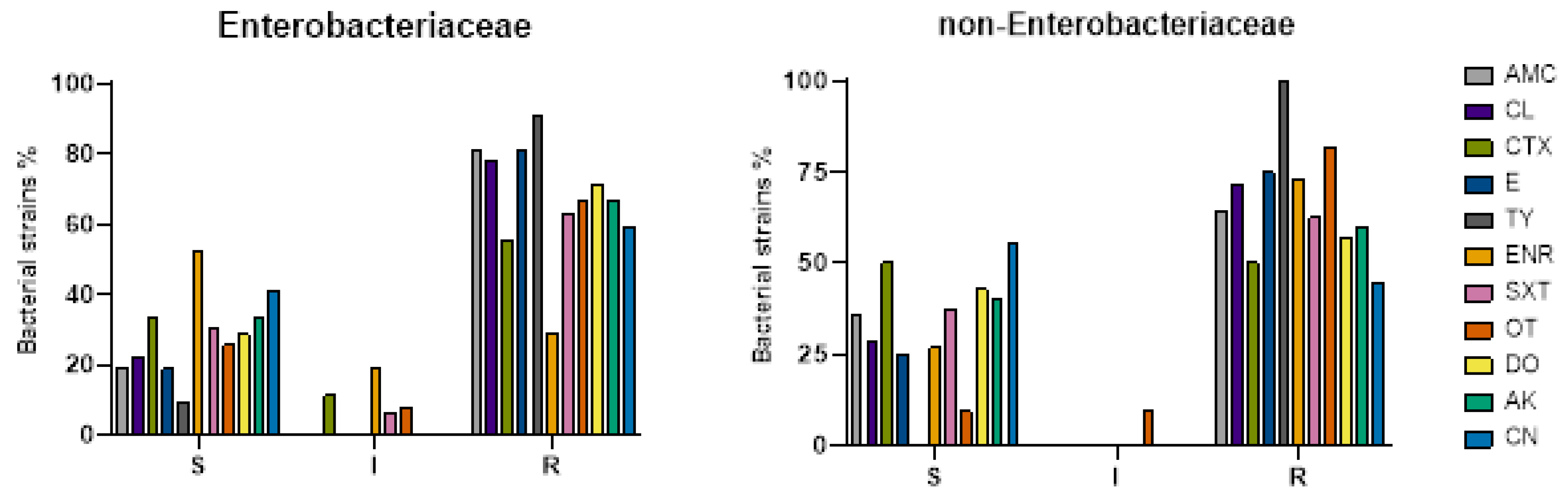
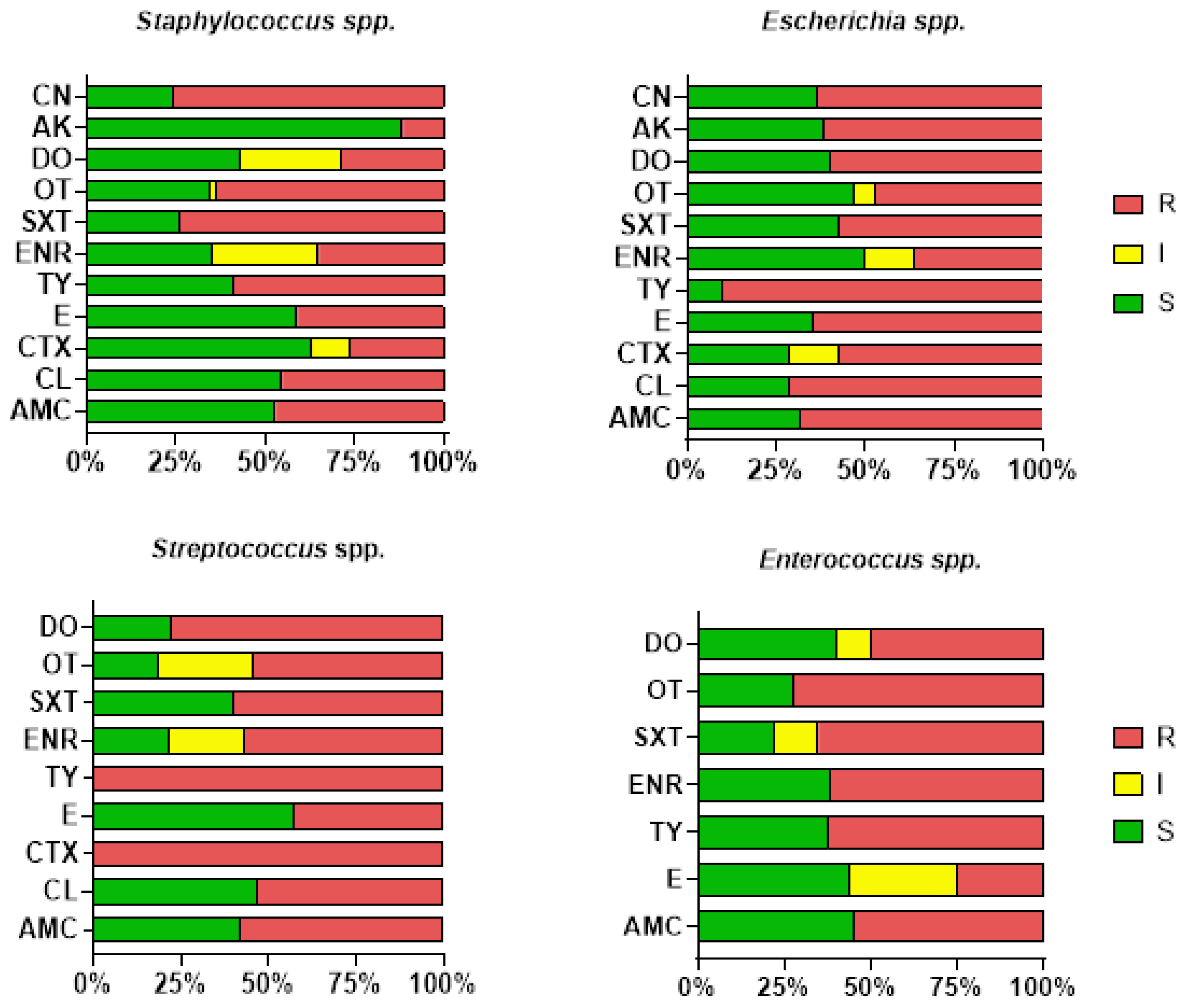
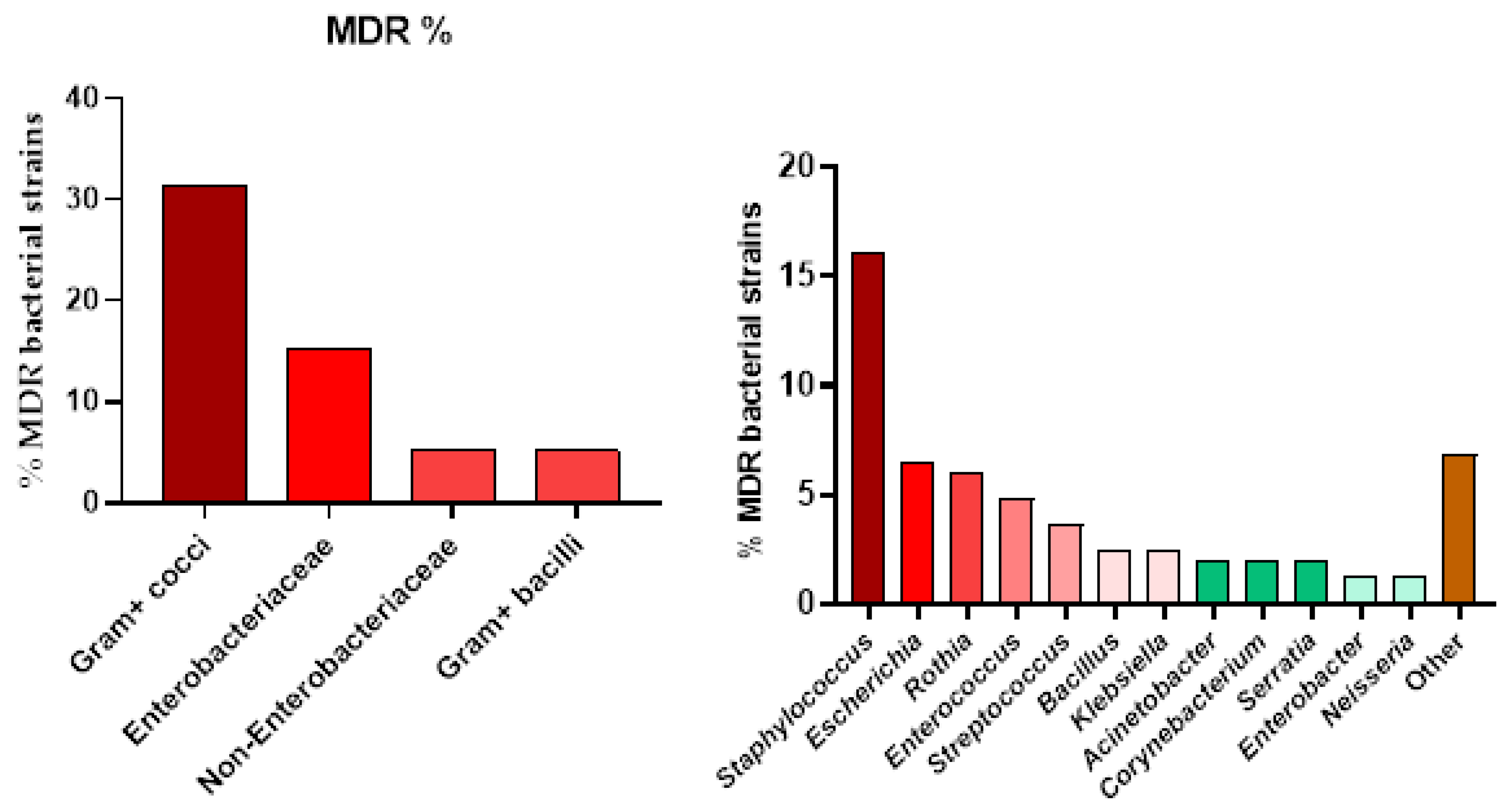
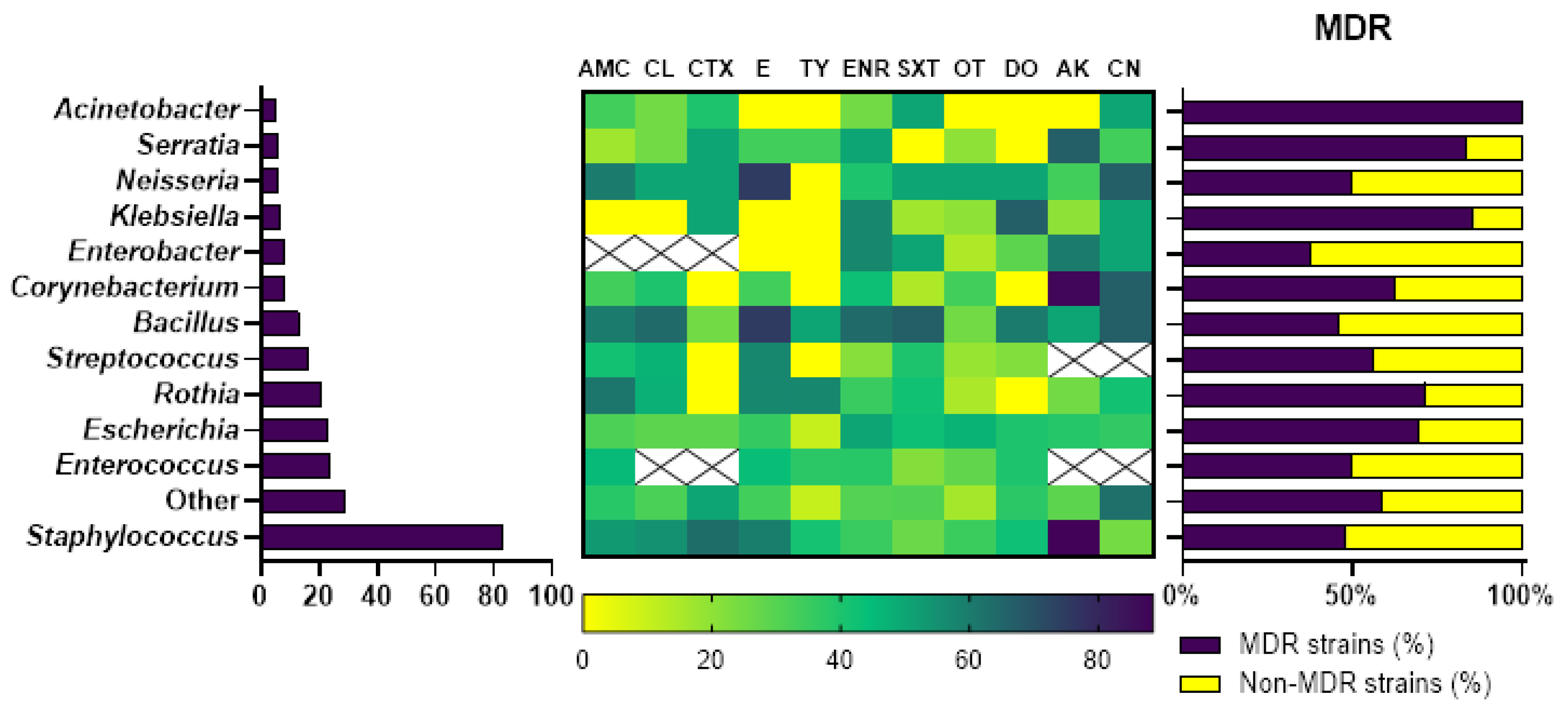
2.3. Antibiotic Susceptibility Testing

3. Discussion
4. Materials and Methods
4.1. Animals and Study Design
4.2. Microbial Identification
4.3. Antimicrobial Susceptibility Testing
4.4. Statistical Analysis
5. Conclusions
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Conflicts of Interest
References
- Cardoso-Freitas, F.; Mota, S.M.; Silva, V.; Martinez-Silvestre, A.; Martins, Â.; Poeta, P. A One Health perspective on multidrug resistance amongst Iberian exotic pet owners. Vet. Sci. 2025, 12, 64. [Google Scholar] [CrossRef]
- Espinosa García-San Román, J.; Quesada-Canales, Ó.; Arbelo Hernández, M.; Déniz Suárez, S.; Castro-Alonso, A. Veterinary education and training on non-traditional companion animals, exotic, zoo, and wild animals: concepts review and challenging perspective on zoological medicine. Vet. Sci. 2023, 10, 357. [Google Scholar] [CrossRef] [PubMed]
- Ahmed, H.A.; Awad, N.F.; Abd El-Hamid, M.I.; Shaker, A.; Mohamed, R.E.; Elsohaby, I. Pet birds as potential reservoirs of virulent and antibiotic resistant zoonotic bacteria. Comp. Immunol. Microbiol. Infect. Dis. 2021, 75, 101606. [Google Scholar] [CrossRef]
- Okunlade, A.O.; Esan, O.O.; Ogunleye, A.O. Multidrug resistant E. coli recovered from household reared female budgerigar pet bird (Melopsittacus undulatus) in Ibadan, Oyo State Nigeria: a case report. Bull. Natl. Res. Cent. 2021, 45, 155. [Google Scholar] [CrossRef]
- Baldrey, V. Guide to using antibiotics in pet birds. In Pract. 2020, 42, 394–404. [Google Scholar] [CrossRef]
- Royal, S.S.; Sigma, S.H.; Ashraf, M.N.; Rimi, S.S.; Affroze, S.; Yousup, S.; et al. Isolation and molecular detection of antibiotic resistant Staphylococcus aureus from pet birds of Mymensigh City Corporation areas, Bangladesh. Am. J. Microbiol. Res. 2024, 12, 1–6. [Google Scholar]
- Durge, A.; Dhaigude, V.; Baby, B.M.; Gujjalkar, P.; Chouraddi, R.; Yadav, S.; et al. Public health threats from pet bird zoonoses. J. Sci. Res. Rep. 2022, 28, 10–20. [Google Scholar] [CrossRef]
- Hasib, F.M.Y.; Bayzid, M.; Das, T.; Rana, E.A.; Kamal, T.; Hossain, M.A.; et al. Multidrug-resistant extended-spectrum beta-lactamase (ESBL) producing Escherichia coli in pet birds of Bangladesh. Vet. Med. Sci. 2025, 11, e70509. [Google Scholar] [CrossRef]
- da Silva, B.D.C.T.; de Carvalho, D.U.O.G.; Sakauchi, V.T.S.; Ferreira, J.S.; Cortez, A.; Heinemann, M.B.; et al. Investigating antimicrobial-resistant bacteria from exotic domestic birds–a One Health concern. Braz. J. Vet. Med. 2024, 46, e001624. [Google Scholar] [CrossRef]
- Monteiro, H.I.G.; Silva, V.; de Sousa, T.; Calouro, R.; Saraiva, S.; Igrejas, G.; et al. Antimicrobial resistance in European companion animals practice: a One Health approach. Animals 2025, 15, 1708. [Google Scholar] [CrossRef]
- Song, Y.; An, Q.; Chen, S.; Dai, H.; Ma, S.; Wu, C.; et al. Antimicrobial resistance of pet-derived bacteria in China, 2000–2020. Antimicrob. Agents Chemother. 2025, 69, e01657-24. [Google Scholar] [CrossRef] [PubMed]
- Mazari, Y.L.; Mouslim, B.A.R.A. Preliminary report on antibiotic-resistant bacteria isolated from select domestic and exotic pets in Guelma, northeast Algeria. Next Res. 2025, 101251. [Google Scholar] [CrossRef]
- Flammer, K. Antibiotic drug selection in companion birds. J. Exotic Pet Med. 2006, 15, 166–176. [Google Scholar] [CrossRef]
- Davies, Y.M.; Cunha, M.P.; Oliveira, M.G.; Oliveira, M.C.; Philadelpho, N.; Romero, D.C.; et al. Virulence and antimicrobial resistance of Klebsiella pneumoniae isolated from passerine and psittacine birds. Avian Pathol. 2016, 45, 194–201. [Google Scholar] [CrossRef]
- Nworie, A.; Onyema, S.I.; Okekpa, S.I.; Elom, M.O.; Umoh, N.O.; Usanga, V.U.; et al. A novel methicillin-resistant Staphylococcus aureus t11469 and a poultry endemic strain t002 (ST5) are present in chicken in Ebonyi State, Nigeria. Biomed Res. Int. 2017, 2017. [Google Scholar] [CrossRef]
- Masud, R.I.; Jahan, R.; Bakhtiyar, Z.; Antor, T.H.; Jany, D.A.; Fahim, N.A.I.; et al. Role of ornamental birds in the transmission of zoonotic pathogen and AMR: a growing public health concern. Ger. J. Microbiol. 2024, 3, 0042. [Google Scholar]
- Soh, H.Y.; Tan, P.X.Y.; Ng, T.T.M.; Chng, H.T.; Xie, S. A critical review of the pharmacokinetics, pharmacodynamics, and safety data of antibiotics in avian species. Antibiotics 2022, 11, 741. [Google Scholar] [CrossRef]
- Muñoz-Ibarra, E.; Molina-López, R.A.; Durán, I.; Garcias, B.; Martín, M.; Darwich, L. Antimicrobial resistance in bacteria isolated from exotic pets: the situation in the Iberian Peninsula. Animals 2022, 12, 1912. [Google Scholar] [CrossRef] [PubMed]
- Caneschi, A.; Bardhi, A.; Barbarossa, A.; Zaghini, A. The use of antibiotics and antimicrobial resistance in veterinary medicine, a complex phenomenon: a narrative review. Antibiotics 2023, 12, 487. [Google Scholar] [CrossRef]
- Crăciun, S.; Novac, C.Ş.; Fiţ, N.I.; Bouari, C.M.; Bel, L.V.; Nadăş, G.C. Bacterial diversity in pet rabbits: implications for public health, zoonotic risks, and antimicrobial resistance. Microorganisms 2025, 13, 653. [Google Scholar] [CrossRef]
- Cardoso, S.; Loc’h, A.L.; Marques, I.; Almeida, A.; Sousa, S.; Saavedra, M.J.; et al. Unveiling the emergence of multidrug-resistant pathogens in exotic pets from France: a comprehensive study (2017-2019). One Health Implement. Res. 2023, 3, 161–176. [Google Scholar] [CrossRef]
- Barbosa, C.K.; Teixeira, V.N.; Pimpão, C.T. Antibiotic usage patterns in exotic pets: a study in Curitiba, Paraná, Brazil. Open Vet. J. 2023, 13, 1543–1553. [Google Scholar] [CrossRef] [PubMed]
- Prestinaci, F.; Pezzotti, P.; Pantosti, A. Antimicrobial resistance: a global multifaceted phenomenon. Pathog. Glob. Health 2015, 109, 309–318. [Google Scholar] [CrossRef]
- Bhat, A.H. Bacterial zoonoses transmitted by household pets and as reservoirs of antimicrobial resistant bacteria. Microb. Pathog. 2021, 155, 104891. [Google Scholar] [CrossRef]
- Ponzo, E.; De Gaetano, S.; Midiri, A.; Mancuso, G.; Giovanna, P.; Giuliana, D.; et al. The antimicrobial resistance pandemic is here: implementation challenges and the need for the One Health approach. Hygiene 2024, 4, 297–316. [Google Scholar] [CrossRef]
- Horvat, O.; Kovačević, Z. Human and veterinary medicine collaboration: synergistic approach to address antimicrobial resistance through the lens of planetary health. Antibiotics 2025, 14, 38. [Google Scholar] [CrossRef]
- Magiorakos, A.P.; Srinivasan, A.; Carey, R.B.; Carmeli, Y.; Falagas, M.E.; Giske, C.G.; et al. Multidrug-resistant, extensively drug-resistant and pandrug-resistant bacteria: an international expert proposal for interim standard definitions for acquired resistance. Clin. Microbiol. Infect. 2012, 18, 268–281. [Google Scholar] [CrossRef]
- Dunn, J.C.; Clegg, S.R. High prevalence of multi-drug-resistant bacteria in faecal samples from UK passerine birds. Sci. Rep. 2025, 15, 28130. [Google Scholar] [CrossRef]
- De Oliveira, D.M.P.; Forde, B.M.; Kidd, T.J.; Harris, P.N.A.; Schembri, M.A.; Beatson, S.A.; et al. Antimicrobial resistance in ESKAPE pathogens. Clin. Microbiol. Rev. 2020, 33, e00181-19. [Google Scholar] [CrossRef]
- Doneley, B. Avian Medicine and Surgery in Practice: Companion and Aviary Birds; CRC Press: Boca Raton, FL, USA, 2018. [Google Scholar]
- Fit, N.I.; Rapuntean, G.; Rapuntean, S.; Chirila, F.; Nadas, G.C. Antibacterial effect of essential vegetal extracts on Staphylococcus aureus compared to antibiotics. Not. Bot. Horti Agrobot. Cluj-Napoca 2009, 37, 117–123. [Google Scholar] [CrossRef]
- Nadăş, G.C.; Manchon, A.M.; Bouari, C.M.; Fiț, N.I. Veterinary clinics as reservoirs for Pseudomonas aeruginosa: a neglected pathway in One Health surveillance. Antibiotics 2025, 14, 720. [Google Scholar] [CrossRef]
- Kim, D.H.; Kim, J.H. Efficacy of tigecycline and linezolid against pan-drug-resistant bacteria isolated from companion dogs in South Korea. Front. Vet. Sci. 2021, 8, 723232. [Google Scholar] [CrossRef]
- Ballash, G.A.; Parker, E.M.; Mollenkopf, D.F.; Wittum, T.E. The One Health dissemination of antimicrobial resistance occurs in both natural and clinical environments. J. Am. Vet. Med. Assoc. 2024, 262, 451–458. [Google Scholar] [CrossRef]
- Nofouzi, K.; Shakeri, S.; Nikkhah, S.; Hosseini, R.H.; Khordadmehr, M.; Madadi, M.; et al. Gallibacterium anatis as an emerging pathogen in pet birds: biofilm formation contributes to treatment challenges and persistence. BMC Microbiol. 2025, 25, 518. [Google Scholar] [CrossRef]
- Masudi, B.; Litvinchuk, T.; Byrd, J. Lactococcus garvieae in rural Alabama: a case report. Cureus 2023, 15, e39560. [Google Scholar] [CrossRef]
- Sirakov, I.; Strateva, T.V.; Boyanov, V.S.; Orozova, P.; Yordanov, D.; Rusenova, N.; et al. Identification, characterization, and epidemiological analysis of Lactococcus garvieae fish isolates obtained in a period of eighteen years. Microorganisms 2025, 13, 436. [Google Scholar] [CrossRef]
- Zayet, S.; Lang, S.; Garnier, P.; Pierron, A.; Plantin, J.; Toko, L.; et al. Leclercia adecarboxylata as emerging pathogen in human infections: clinical features and antimicrobial susceptibility testing. Pathogens 2021, 10, 1399. [Google Scholar] [CrossRef] [PubMed]
- Di Gregorio, M.; Tiecco, G.; Colangelo, C.; Corbellini, S.; Caccuri, F. Leclercia adecarboxylata. An Emerging Pathogen: a Narrative Review. J. Infect. Dis. Ther. 2023, 11, 2. [Google Scholar]
- Aurori, M.; Novac, C.Ș.; Nadăş, G.C.; Crăciun, S.; Fiţ, N.; Andrei, S. The antimicrobial effect of Cornus mas L. and Sorbus aucuparia L. fruit extracts against resistant uropathogens in correlation with the prevalence of urinary tract infections in companion animals. Pharmaceuticals 2024, 17, 814. [Google Scholar] [CrossRef] [PubMed]
- Nadăş, G.C.; Novac, C.Ș.; Matei, I.A.; Bouari, C.M.; Gal, Z.M.; Tamas-Krumpe, O.M.; et al. Prevalence of antimicrobial resistant bacteria from conjunctival flora in an eye infection prone breed (Saint Bernard). Molecules 2021, 26*, 2219. [Google Scholar] [CrossRef]
- Qureshi, M.H.F.; Azam, F.; Shafique, M.; Aslam, B.; Farooq, M.; Rafique, M.K.; et al. A One Health perspective of pet birds bacterial zoonosis and prevention. Pak. Vet. J. 2024, 44, 1. [Google Scholar]
- Cabral, B.G.; Davies, Y.M.; Menão, M.C.; Saidenberg, A.B.; Gomes, V.T.M.; Moreno, L.Z.; et al. Companion psittacine birds as reservoir of gentamicin and vancomycin-resistant Enterococcus spp. Pesq. Vet. Bras. 2020, 40, 129–133. [Google Scholar] [CrossRef]
- Punom, S.A.; Islam, M.S.; Fahim, N.A.I.; Hasan, M.A.E.; Antor, M.T.H.; Saha, S.; et al. Ornamental birds: hidden carriers of potentially virulent and antimicrobial-resistant Enterococcus faecalis in Bangladesh. Microbiol. Spectr. 2025, 13, e01974-25. [Google Scholar] [CrossRef]
- Damborg, P.; Broens, E.M.; Chomel, B.B.; Guenther, S.; Pasmans, F.; Wagenaar, J.A.; et al. Bacterial zoonoses transmitted by household pets: state-of-the-art and future perspectives for targeted research and policy actions. J. Comp. Pathol. 2016, 155, S27–S40. [Google Scholar] [CrossRef] [PubMed]
- Matuschek, E.; Brown, D.F.; Kahlmeter, G. Development of the EUCAST disk diffusion antimicrobial susceptibility testing method and its implementation in routine microbiology laboratories. Clin. Microbiol. Infect. 2014, 20, O255–O266. [Google Scholar] [CrossRef] [PubMed]
- EUCAST. 2026. Available online: http://www.eucast.org.
- Performance Standards for Antimicrobial Disk and Dilution Susceptibility Tests for Bacteria Isolated from Animals, 7th ed.; CLSI Supplement VET01S; Clinical and Laboratory Standards Institute: Wayne, PA, USA, 2024.
| Birds % | Total | ||||
|---|---|---|---|---|---|
| Gender | Male 59 (n=117) | Female 41 (n=81) | 100 (n=198) |
||
| Age | Juvenile 30.3 (n=60) | Adult 69.7 (n=138) | 100 (n=198) |
||
| Season | Spring 22.7 (n=45) | Summer 27.3 (n=54) | Autumn 23.7 (n=47) | Winter 26.3 (n=52) | 100 (n=198) |
| Pathology |
Digestive 62.1 (n=123) |
Respiratory 37.9 (n=75) |
100 (n=198) |
||
| Antibiotic usage | Yes 21.7 (n=43) | No 78.3 (n=155) | 100 (n=198) |
||
| Digestive | Respiratory | Total | ||
|---|---|---|---|---|
| Gender (%) | Male | 59.3% (n=73) | 58.7% (n=44) | 100 (n=198) |
| Female | 40.7% (n=50) | 41.3% (n=31) | ||
| Age (%) | Young | 31.7% (n=39) | 28% (n=21) | 100 (n=198) |
| Adult | 68.3% (n=84) | 72% (n=54) | ||
| Season (%) | Spring | 24.4% (n=30) | 20% (n=15) | 100 (n=198) |
| Summer | 30% (n=37) | 22.7% (n=17) | ||
| Autumn | 23.6% (n=29) | 24% (n=18) | ||
| Winter | 22% (n=27) | 33.3% (n=25) | ||
| Total (n) | 62.1 (n=123) | 37.9 (n75) | 100 (n=198) | |
| Group | Genus | Species | Digestive | Respiratory | Total |
|---|---|---|---|---|---|
| Gram + cocci | Dermacoccus | Dermacoccus nishinomiyaensis | - | 1.21% (n=3) | 1.21% (n=3) |
| Enterococcus | Enterococcus faecalis | 7.23% (n=18) | - | 7.23% (n=18) | |
| Enterococcus faecium | 1.61% (n=4) | 0.81% (n=2) | 2.42% (n=6) | ||
| Kocuria | Kocuria kristinae | - | 0.81% (n=2) | 0.81% (n=2) | |
| Kocuria rhizophila | - | 0.41% (n=1) | 0.41% (n=1) | ||
| Kocuria rosea | - | 0.41% (n=1) | 0.41% (n=1) | ||
| Lactococcus | Lactococcus garvieae | - | 0.41% (n=1) | 0.41% (n=1) | |
| Micrococcus | Micrococcus luteus | - | 1.21% (n=3) | 1.21% (n=3) | |
| Rothia | Rothia dentocariosa | 0.41% (n=1) | 0.81% (n=2) | 1.22% (n=3) | |
| Rothia kristinae | 2.01% (n=5) | 0.41% (n=1) | 2.42% (n=6) | ||
| Rothia nasimurium | 1.21% (n=3) | 3.62% (n=9) | 4.83% (n=12) | ||
| Staphylococcus | Staphylococcus aureus | 0.41% (n=1) | 2.01% (n=5) | 2.42% (n=6) | |
| Staphylococcus borealis | - | 1.21% (n=3) | 1.21% (n=3) | ||
| Staphylococcus delphini | 0.81% (n=2) | - | 0.81% (n=2) | ||
| Staphylococcus epidermidis | 0.41% (n=1) | 1.21% (n=3) | 1.62% (n=4) | ||
| Staphylococcus equorum | 0.41% (n=1) | - | 0.41% (n=1) | ||
| Staphylococcus gallinarum | 0.41% (n=1) | 1.21% (n=3) | 1.62% (n=4) | ||
| Staphylococcus haemolyticus | 2.41% (n=6) | 8.04% (n=20) | 10.45% (n=26) | ||
| Staphylococcus hominis | - | 0.41% (n=1) | 0.41% (n=1) | ||
| Staphylococcus hyicus | - | 0.41% (n=1) | 0.41% (n=1) | ||
| Staphylococcus intermedius | - | 0.41% (n=1) | 0.41% (n=1) | ||
| Staphylococcus kloosii | 0.41% (n=1) | - | 0.41% (n=1) | ||
| Staphylococcus saprophyticus | 0.81% (n=2) | - | 0.81% (n=2) | ||
| Staphylococcus sciuri | 2.41% (n=6) | 2.01% (n=5) | 4.42% (n=11) | ||
| Staphylococcus simulans | - | 0.41% (n=1) | 0.41% (n=1) | ||
| Staphylococcus vitulinus | 0.81% (n=2) | - | 0.81% (n=2) | ||
| Staphylococcus warneri | 93.62% (n=9) | 1.21% (n=3) | 4.83% (n=12) | ||
| Staphylococcus xylosus | 2.01% (n=5) | - | 2.01% (n=5) | ||
| Streptococcus | Streptococcus gallolyticus | - | 0.81% (n=2) | 0.81% (n=2) | |
| Streptococcus mitis | - | 1.61% (n=4) | 1.61% (n=4) | ||
| Streptococcus pluranimalium | 0.41% (n=1) | - | 0.41% (n=1) | ||
| Streptococcus salivarius | 2.01% (n=5) | 0.41% (n=1) | 2.42% (n=6) | ||
| Streptococcus sanguinus | 0.41% (n=1) | - | 0.41% (n=1) | ||
| Streptococcus suis | 0.41% (n=1) | 0.41% (n=1) | 0.82% (n=2) | ||
| G + bacilli | Bacillus | Bacillus cereus | 1.21% (n=3) | - | 1.21% (n=3) |
| Bacillus circulans | 0.41% (n=1) | - | 0.41% (n=1) | ||
| Bacillus licheniformis | - | 0.41% (n=1) | 0.41% (n=1) | ||
| Bacillus pumilus | 0.41% (n=1) | - | 0.41% (n=1) | ||
| Bacillus spp. | 1.61% (n=4) | 2 (0.81%) (n=2) | 2.42% (n=6) | ||
| Bacillus velezensis | 1 (0.41%) (n=1) | - | 0.41% (n=1) | ||
| Corynebacterium | Corynebacterium amycolatum | 0.41% (n=1) | - | 0.41% (n=1) | |
| Corynebacterium durum | - | 0.41% (n=1) | 0.41% (n=1) | ||
| Corynebacterium falsenii | - | 0.41% (n=1) | 0.41% (n=1) | ||
| Corynebacterium spp. | 1.21% (n=3) | 0.81% (n=2) | 2.02% (n=5) | ||
| Lactobacillus | Lactobacillus crispatus | 0.41% (n=1) | - | 0.41% (n=1) | |
| Lactobacillus johnsonii | 0.41% (n=1) | - | 0.41% (n=1) | ||
| Lactobacillus salivarius | 0.41% (n=1) | - | 0.41% (n=1) | ||
| Microbacterium | Microbacterium aerolatum | 0.41% (n=1) | - | 0.41% (n=1) | |
| Enterobacteriaceae | Enterobacter | Enterobacter asburiae | - | 0.41% (n=1) | 0.41% (n=1) |
| Enterobacter cloacae | 1.21% (n=3) | - | 1.21% (n=3) | ||
| Enterobacter hormaechei | 0.41% (n=1) | 0.81% (n=2) | 1.22% (n=3) | ||
| Enterobacter kobei | 0.41% (n=1) | - | 0.41% (n=1) | ||
| Escherichia | Escherichia coli | 8.84% (n=22) | 0.41% (n=1) | 9.24% (n=23) | |
| Klebsiella | Klebsiella oxytoca | 0.41% (n=1) | 0.41% (n=1) | 0.82% (n=2) | |
| Klebsiella pneumoniae | 1.21% (n=3) | 0.81% (n=2) | 2.02% (n=5) | ||
| Kosakonia | Kosakonia cowanii | 0.41% (n=1) | - | 0.41% (n=1) | |
| Leclercia | Leclercia adecarboxylata | 0.41% (n=1) | - | 0.41% (n=1) | |
| Mixta | Mixta gaviniae | 0.41% (n=1) | - | 0.41% (n=1) | |
| Proteus | Proteus mirabilis | - | 1.21% (n=3) | 1.21% (n=3) | |
| Proteus vulgaris | 0.41% (n=1) | - | 0.41% (n=1) | ||
| Raoultella | Raoultella terrigena | 0.41% (n=1) | - | 0.41% (n=1) | |
| Serratia | Serratia marcescens | 1.21% (n=3) | 1.21% (n=3) |
2.42% (n=6) | |
| Non-Enterobacteriaceae | Acinetobacter | Acinetobacter baumannii | 0.41%) (n=1) | - | 0.41% (n=1) |
| Acinetobacter pittii | 0.41% (n=1) | 0.41% (n=1) | 0.82% (n=2) | ||
| Acinetobacter seifertii | 0.41% (n=1) | - | 0.41% (n=1) | ||
| Acinetobacter ursingii | - | 0.41% (n=1) | 0.41% (n=1) | ||
| Aeromonas | Aeromonas caviae | 0.41% (n=1) | - | 0.41% (n=1) | |
| Aeromonas salmonicida | 0.41% (n=1) | 0.41% (n=1) | 0.82% (n=2) | ||
| Gallibacterium | Gallibacterium anatis | 0.81% (n=2) | - | 0.81% (n=2) | |
| Neisseria | Neisseria denitrificans | - | 0.41% (n=1) | 0.41% (n=1) | |
| Neisseria sicca | - | 2.01% (n=1) | 2.01% (n=1) | ||
| Pseudomonas | Pseudomonas monteilii | 0.41% (n=1) | - | 0.41% (n=1) | |
| Total | 56.63% (n=141) | 43.37% (n=108) | 100% (n=249) | ||
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2026 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).




