Submitted:
10 April 2026
Posted:
13 April 2026
You are already at the latest version
Abstract
Keywords:
1. Introduction
2. Methods
3. Results
3.1. Dataset Construction
3.2. Scoring Results
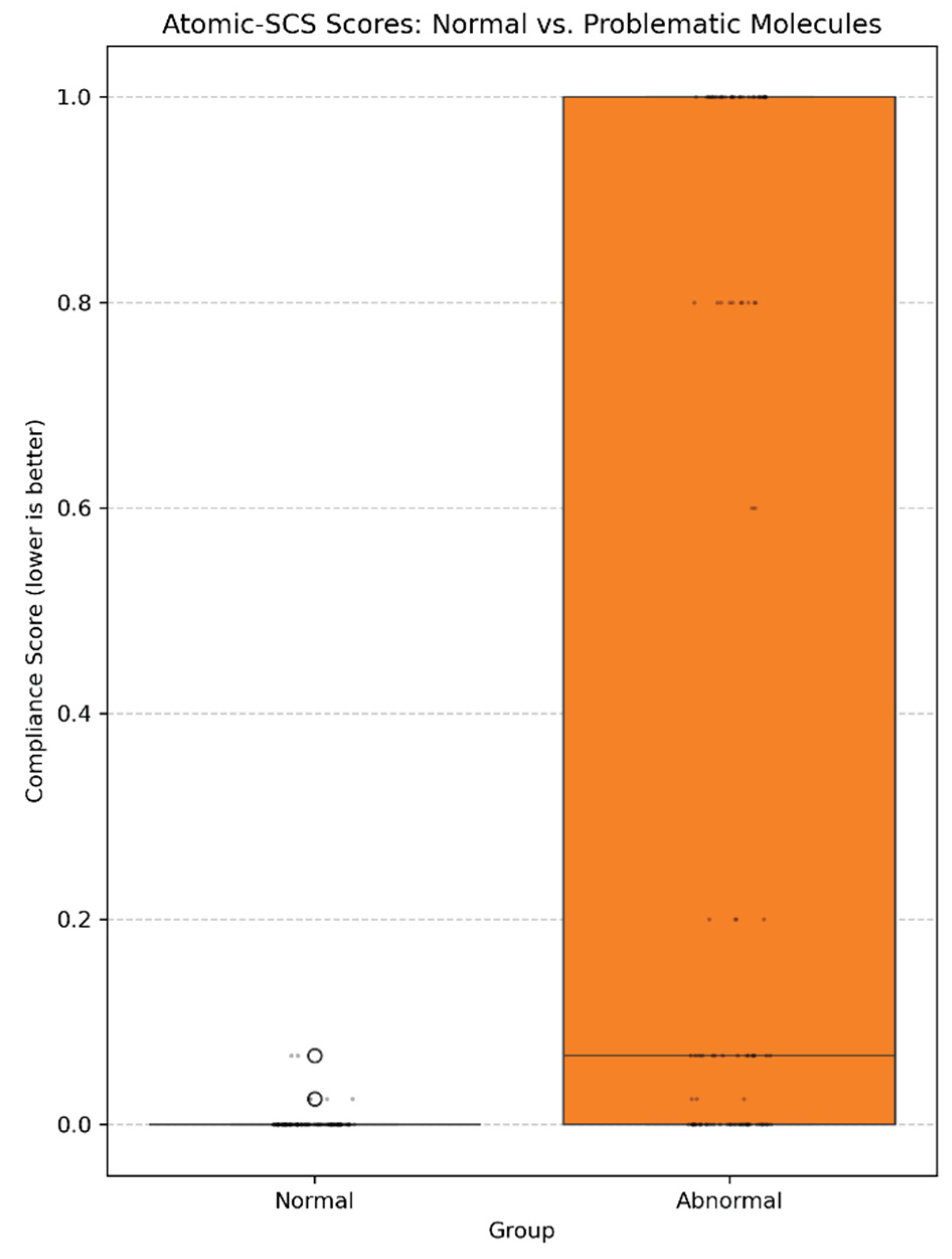
3.3. Performance
4. Discussion and Limitations
Limitations
Future Work
Data Availability Statement
Acknowledgments
References
- Olivecrona, M.; Blaschke, T.; Engkvist, O.; Chen, H. Molecular De Novo Design through Deep Reinforcement Learning. J. Cheminform. 2017, 9(1), 48. [Google Scholar] [CrossRef] [PubMed]
- Gilmer, J.; Schoenholz, S.S.; Riley, P.F.; Vinyals, O.; Dahl, G.E. Neural message passing for quantum chemistry. In Proceedings of the 34th International Conference on Machine Learning (ICML), 2017; pp. pp 1263–1272. [Google Scholar]
- RDKit: Open-source cheminformatics. Available online: https://www.rdkit.org (accessed on 7 April 2026).
- ChemAudit: An Open-Source Chemical Structure Validation Suite. NFDI4Chem. 2026. Available online: https://nfdi4chem.de/introducing-chemaudit/ (accessed on 5 April 2026).
- Geng, C.; Zhang, L.; Zhang, M.; Ye, H.; Zhao, Z.; Si, X. Neural proposals, symbolic guarantees: Neuro-symbolic graph generation with hard constraints. arXiv 2026, arXiv:2602.16954. [Google Scholar] [CrossRef]
- Christopher, J.K.; Caldei, M.; Liang, J.; Fioretto, F. Neuro-symbolic generative diffusion models for physically grounded, robust, and safe generation. In Proceedings of the Second International Conference on Neuro-Symbolic Systems (NeuS; 2025. [Google Scholar]
- Landrum, G. RDKit: A software suite for cheminformatics, computational chemistry, and predictive modeling. GitHub. 2016. Available online: https://github.com/rdkit/rdkit.
- Kusner, M.J.; Paige, B.; Hernández-Lobato, J.M. Grammar variational autoencoder. In Proceedings of the 34th International Conference on Machine Learning (ICML), 2017; pp. pp 1945–1954. [Google Scholar]
- Jin, W.; Barzilay, R.; Jaakkola, T. Junction tree variational autoencoder for molecular graph generation. In Proceedings of the 35th International Conference on Machine Learning (ICML), 2018; pp. pp 2323–2332. [Google Scholar]
- Elton, D.C.; Boukouvalas, Z.; Fuge, M.D.; Chung, P.W. Deep learning for molecular design—a review of the state of the art. Mol. Syst. Des. Eng. 2019, 4, 828–849. [Google Scholar] [CrossRef]
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2026 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license.



