Submitted:
09 April 2026
Posted:
13 April 2026
You are already at the latest version
Abstract
Keywords:
1. Introduction
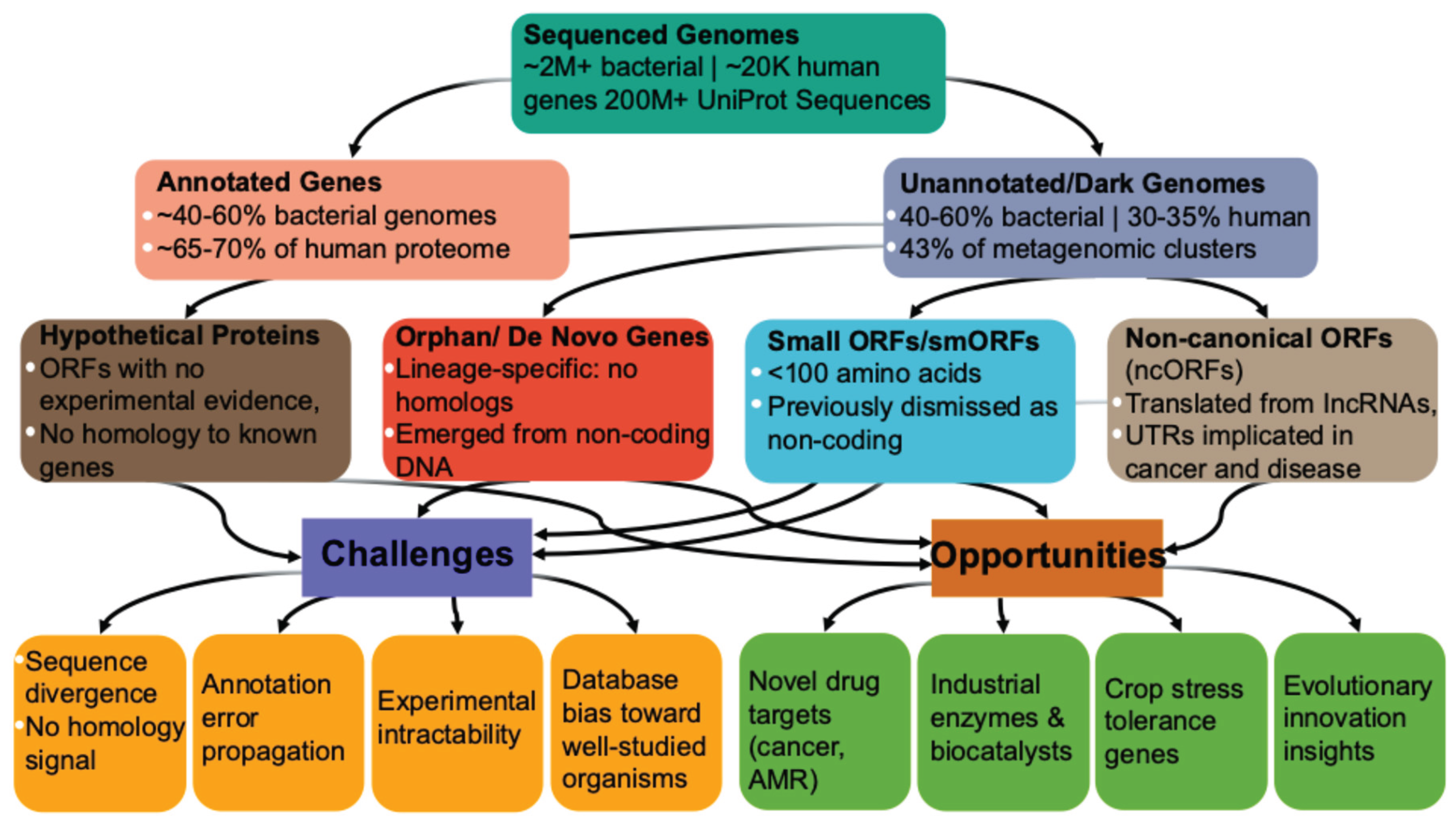
2. The Landscape of Unannotated Genes: Categories and Origins
2.1. Hypothetical Proteins (HPs)
2.2. Orphan and De Novo Genes
2.3. Small Open Reading Frames (smORFs) and Micropeptides
2.4. Non-Canonical ORFs (ncORFs)
3. Challenges in Genome Annotation
3.1. The Fundamental Limitations of Homology-Based Inference
3.2. Annotation Error Propagation and Database Contamination
3.3. Experimental Intractability and the “Streetlight Effect”
3.4. Biological Complexity: Multifunctionality and Context-Dependence
4. Biotechnological Opportunities in the Dark Genome
4.1. Plant Genetics, Biotechnology, and Agricultural Applications
4.2. Animal Genetics and Biotechnology
4.3. Medical Biotechnology and Genomic Medicine
4.4. Microbial, Industrial, and Environmental Biotechnology
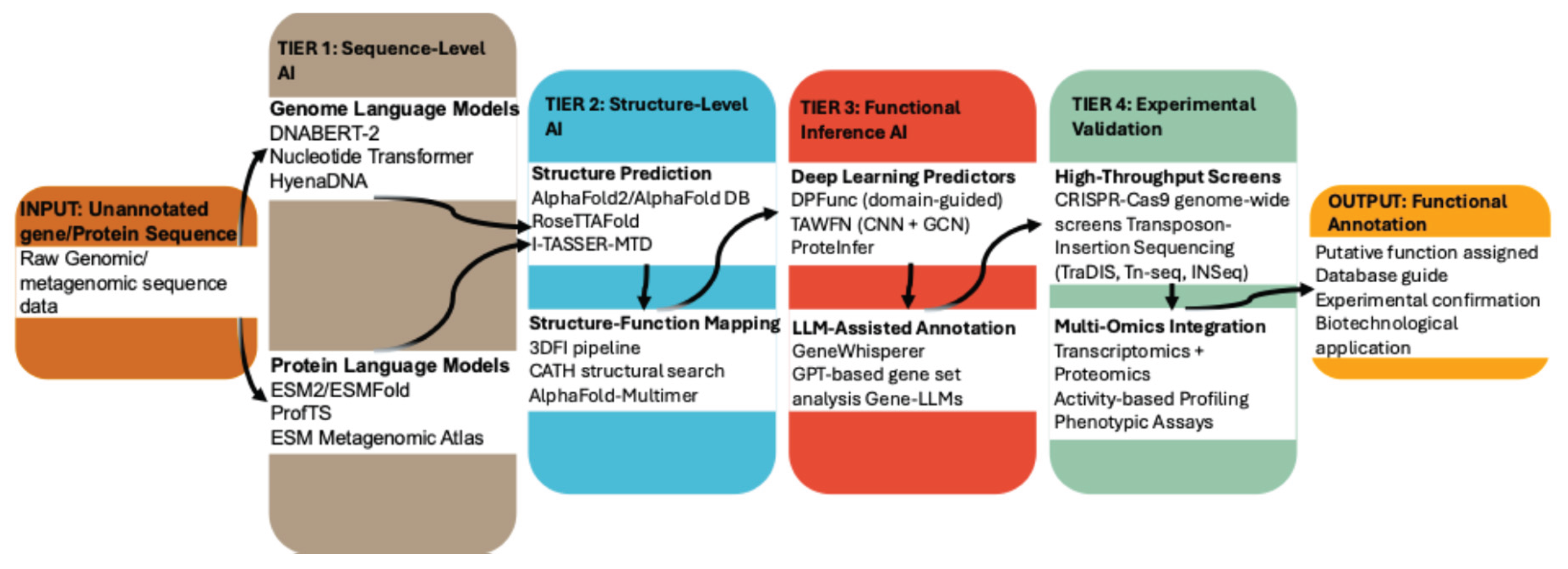
5. AI Solutions for Functional Annotation: A Multi-Tiered Framework
5.1. Tier 1: Sequence Level AI Using Genome and Protein Language Models
5.2. Tier 2: Structure Level AI for Protein Structure Prediction
5.3. Tier 3: Functional Inference AI Using Deep Learning and Large Language Models
5.4. Tier 4: Experimental Validation Using High-Throughput Functional Genomics
6. Integrative Strategies and Future Perspectives
7. Conclusions
Supplementary Materials
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Acknowledgments
Conflicts of Interest
Abbreviations
| AI | Artificial Intelligence |
| AMR | Antimicrobial Resistance |
| CNN | Convolutional Neural Network |
| dORF | Downstream Open Reading Frame |
| EC | Enzyme Commission |
| GCN | Graph Convolutional Network |
| GO | Gene Ontology |
| HITS | High-throughput Insertion Tracking by Deep Sequencing |
| HP | Hypothetical Protein |
| HPP | Human Proteome Project |
| INSeq | Insertion Sequencing |
| LLM | Large Language Model |
| lncRNA | Long Non-coding RNA |
| MAG | Metagenome-Assembled Genome |
| ML | Machine Learning |
| ncORF | Non-canonical Open Reading Frame |
| NGS | Next-Generation Sequencing |
| NR | Non-Redundant |
| nuORF | Novel Unannotated Open Reading Frame |
| ORF | Open Reading Frame |
| PFAS | Per- and Polyfluoroalkyl Substances |
| PLM | Protein Language Model |
| PUF | Protein of Unknown Function |
| Ribo-seq | Ribosome Profiling |
| SmORF | Small Open Reading Frame |
| TAWFN | Two-model Adaptive Weight Fusion Network |
| TIS | Transposon-Insertion Sequencing |
| Tn-seq | Transposon Sequencing |
| TraDIS | Transposon-Directed Insertion-Site Sequencing |
| uORF | Upstream Open Reading Frame |
| UTR | Untranslated Region |
| U/U | Unknown/Uncharacterized |
References
- Fleischmann, R. D.; et al. (1995). Whole-genome random sequencing and assembly of Haemophilus influenzae Rd. Science, 269(5223), 496-512. [CrossRef]
- UniProt Consortium. (2025). UniProt: The Universal Protein Knowledgebase in 2025. Nucleic Acids Research, 53(D1), D609-D617. https://academic.oup.com/nar/article/53/D1/D609/7902999.
- Nolan, L. M., Webber, M. A., & Filloux, A. (2025). Throwing a spotlight on genomic dark matter: The power and potential of transposon-insertion sequencing. Journal of Biological Chemistry, 301(6), 110231. [CrossRef]
- Moitra, T., & Larrouy-Maumus, G. (2026). Integrated approaches for discovery and functional annotation of proteins of unknown function. Trends in Biochemical Sciences, 51(1), 80-92. [CrossRef]
- Vanni, C.; et al. (2022). Unifying the known and unknown microbial coding sequence space. eLife, 11, e67667. [CrossRef]
- Rocha, J. J.; et al. (2023). Functional unknomics: Systematic screening of conserved genes of unknown function. PLoS Biology, 21(8), e3002222. [CrossRef]
- Rodríguez del Río, Á.; et al. (2024). Functional and evolutionary significance of unknown genes. Nature, 626, 104-111. [CrossRef]
- Wang, X., Wang, B., & Yuan, F. (2023). Deciphering the roles of unknown/uncharacterized genes in plant development and stress responses. Frontiers in Plant Science, 14, 1276559. [CrossRef]
- Ge, A., Chan, C., & Yang, X. (2024). Exploring the dark matter of human proteome: The emerging role of non-canonical open reading frame (ncORF) in cancer diagnosis, biology, and therapy. Cancers, 16(15), 2660. [CrossRef]
- Vincent, A. T. (2024). Bacterial hypothetical proteins may be of functional interest. Frontiers in Bacteriology, 3, 1334712. [CrossRef]
- Jumper, J.; et al. (2021). Highly accurate protein structure prediction with AlphaFold. Nature, 596, 583-589. [CrossRef]
- Zhang, Y.; et al. (2025). Predicting functions of uncharacterized gene products from microbial communities. Nature Biotechnology. [CrossRef]
- Lobb, B.; et al. (2020). An assessment of genome annotation coverage across the bacterial tree of life. Microbial Genomics, 6(5), mgen000341. [CrossRef]
- Casola, C. (2025). De Novo Genes: Current Status and Future Goals. Genome Biology and Evolution, 17(12), evaf230. [CrossRef]
- Grandchamp, A.; et al. (2025). De Novo Gene Emergence: Summary, Classification, and Challenges of Current Methods. Genome Biology and Evolution, 17(11), evaf197. [CrossRef]
- Luo, M.; et al. (2025). Rethinking de novo genes in plants: Mechanisms, methodological progress, and future prospects. Frontiers in Plant Science, 16, 1724832. [CrossRef]
- Baena-Angulo, C.; et al. (2025). Cis to trans: Small ORF functions emerging through evolution. Trends in Genetics, 41(2), 119-131. [CrossRef]
- Ruiz-Orera, J.; et al. (2025). The non-canonical proteome: A novel contributor to cancer biology. Nature Reviews Cancer. https://pmc.ncbi.nlm.nih.gov/articles/PMC11909265/.
- Schnoes, A. M.; et al. (2009). Annotation Error in Public Databases: Misannotation of Molecular Function in Enzyme Superfamilies. PLoS Computational Biology, 5(12), e1000605. [CrossRef]
- Goudey, B.; et al. (2022). Propagation, detection and correction of errors using the sequence database network. Briefings in Bioinformatics, 23(6), bbac416. [CrossRef]
- Yao, C.; et al. (2022). Overexpression of a Malus baccata MYB Transcription Factor Gene MbMYB4 Increases Cold and Drought Tolerance in Arabidopsis thaliana. International Journal of Molecular Sciences, 23(3):1794. [CrossRef]
- Kumar, B.; et al. (2023). Orphan crops: A genetic treasure trove for hunting stress tolerance genes. Food and Energy Security, 12(1), e436. [CrossRef]
- Fierro-Monti, I. (2025). Tiny proteins, great impacts: Non canonical ORFs in cancer. Academia Molecular Biology and Genomics, 2(2). [CrossRef]
- Park, B. S.; et al. (2025). Perturbomics: CRISPR–Cas screening-based functional genomics approach for drug target discovery. Experimental & Molecular Medicine, 57, 1-12. [CrossRef]
- Saggu, S. K., Kumar, M., & Kumar, S. (2026). Metagenomics and its impact on environmental and therapeutic microbiology. Archives of Microbiology, 208, 1-18. [CrossRef]
- Rennie, M. L., & Oliver, M. R. (2025). Emerging frontiers in protein structure prediction following the AlphaFold revolution. Journal of the Royal Society Interface, 22(225), 20240886. [CrossRef]
- Varadi, M.; et al. (2022). AlphaFold Protein Structure Database: Massively expanding the structural coverage of protein-sequence space with high-accuracy models. Nucleic Acids Research, 50(D1), D439-D444. [CrossRef]
- Sousounis, K.; et al. (2012). Protein function and structure: A systems biology perspective. Briefings in Bioinformatics, 13(5), 527-538.
- Julian, T.; et al. (2021). 3DFI: A pipeline for structural-based functional annotation of proteins. Bioinformatics, 37(18), 3028-3030.
- Lin, Z.; et al. (2023). Evolutionary-scale prediction of atomic-level protein structure with a language model. Science, 379(6637), 1123-1130. [CrossRef]
- Dalla-Torre, H.; et al. (2024). Nucleotide Transformer: Building and evaluating robust foundation models for human genomics. Nature Methods, 21, 1-10. [CrossRef]
- Zhou, Z.; et al. (2024). DNABERT-2: Efficient Foundation Model and Benchmark For Multi-Species Genome. arXiv preprint, arXiv:2306.15006. https://arxiv.org/abs/2306.15006.
- Wang, W.; et al. (2025). DPFunc: Accurately predicting protein function via deep learning with domain-guided structure information. Nature Communications, 16, 1-15. [CrossRef]
- Meng, L.; et al. (2024). TAWFN: A deep learning framework for protein function prediction. Bioinformatics, 40(10), btae571. [CrossRef]
- Balakrishnan, P.; et al. (2025). Gene-LLMs: A comprehensive survey of transformer-based genomic language models for regulatory and clinical genomics. Frontiers in Genetics, 16, 1634882. [CrossRef]
- Przybyla, L., & Gilbert, L. A. (2022). A new era in functional genomics screens. Nature Reviews Genetics, 23(2), 89-103. [CrossRef]
| Application Domain | Opportunity | Key Example | Reference |
|---|---|---|---|
| Plant & Agricultural Biotechnology | Novel stress tolerance genes in dispensable genome | MbMYBC1 enhancing drought resistance | [8,21] |
| Medical Biotechnology | ncORFs as cancer biomarkers and drug targets | C9orf50 in colorectal cancer | [9,24] |
| Microbial Biotechnology | Novel biocatalysts from environmental metagenomes | Extremophilic enzymes for precision fermentation | [7,25] |
| Industrial Biotechnology | Undiscovered metabolic pathways for bioproduction | Novel biosynthetic gene clusters | [7] |
| Environmental Biotechnology | Novel degradative enzymes for bioremediation | PFAS and microplastic degradation | [25] |
| Animal Genomics | Lineage-specific genes in livestock adaptation | Selective sweeps in cattle populations | [14] |
| AI Tool/Method | Category | Key Capability | Application to Dark Genome | Reference |
|---|---|---|---|---|
| AlphaFold2 | Structure Prediction | Atomic-level 3D structure from sequence | Structural annotation of HPs | [12] |
| ESMFold / ESM2 | Protein Language Model | Structure & function from sequence | Metagenomic protein annotation | [30] |
| DNABERT-2 | Genome Language Model | Regulatory element & ORF annotation | smORF and ncORF discovery | [32] |
| Nucleotide Transformer | Genome Language Model | Multi-species genomic task performance | Cross-species annotation | [31] |
| DPFunc | Deep Learning Predictor | GO term prediction with domain guidance | HP function inference | [33] |
| TAWFN | Deep Learning Predictor | CNN + GCN fusion for function prediction | Multi-scale functional annotation | [34] |
| GeneWhisperer | LLM-Assisted Annotation | Literature-guided gene curation | Automated HP annotation | [35] |
| TraDIS / Tn-seq (TIS) | Experimental Validation | Genome-wide fitness profiling in bacteria | Bacterial HP characterization | [6] |
| CRISPR Perturbomics | Experimental Validation | Genome-wide knockout screens | Eukaryotic dark proteome discovery | [36] |
| 3DFI Pipeline | Structure-Function Mapping | Automated structural comparison | HP function inference via CATH | [29] |
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2026 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).




