Submitted:
09 April 2026
Posted:
10 April 2026
You are already at the latest version
Abstract
Keywords:
1. Introduction
2. Materials and Methods
2.1. Study Setting
2.2. Sample Size
2.3. Residual Clinical Specimens
2.4. Reference Material
2.5. SARS-CoV-2 Viral Culture Samples
2.6. FluoroType® SARS-CoV-2 varID Q Assay
2.7. Nucleic Acid Extraction
2.8. Nucleic Acid Amplification
2.9. Statistical Analysis
3. Results
3.1. Overall Results
3.2. Accuracy and Agreement (Cohen’s Kappa Coefficient)
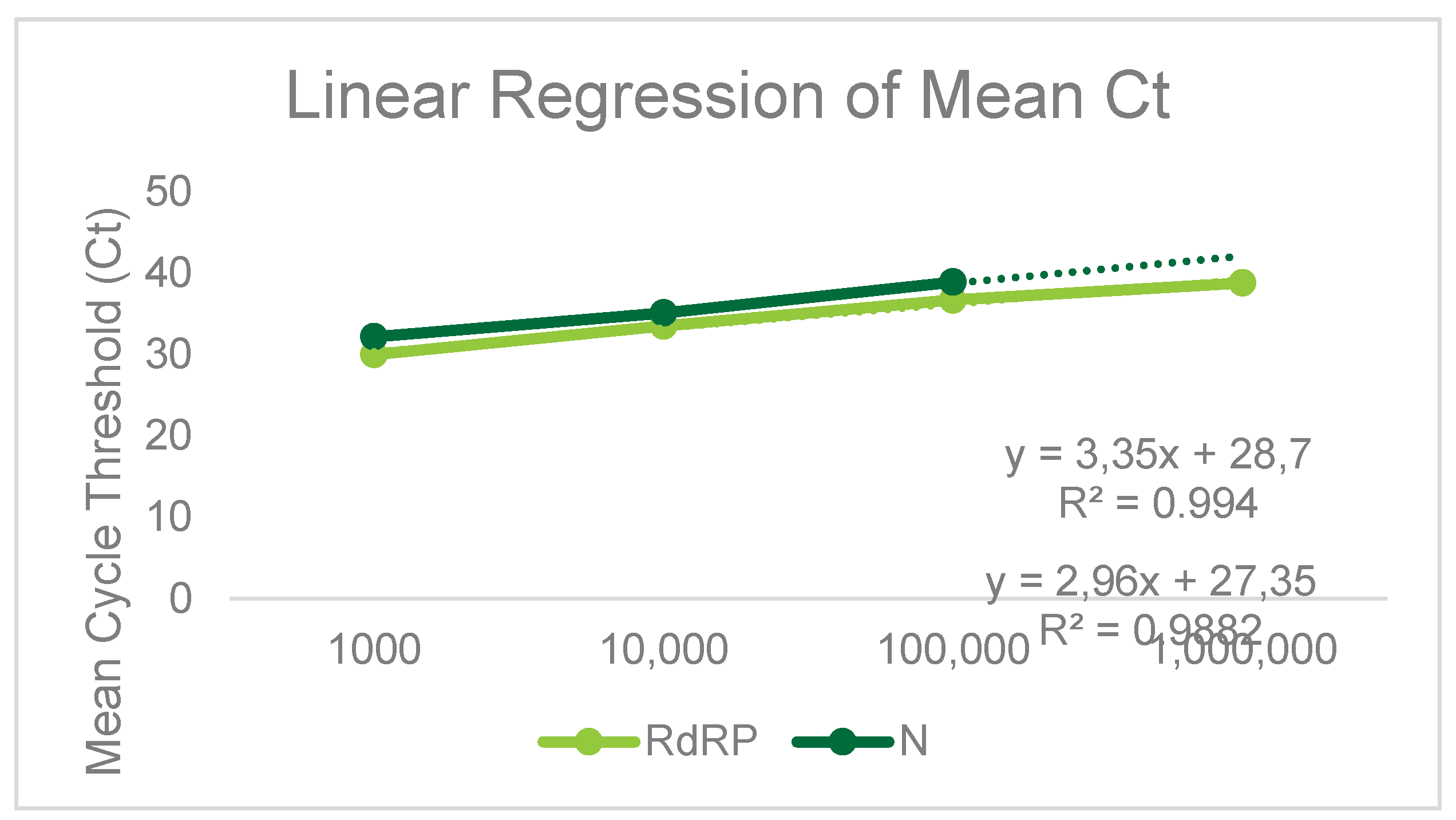
3.3. Precision Analysis (Reproducibility) and Linearity
3.4. Limit of Detection (LOD)
3.5. Variant Detection
4. Discussion
Author Contributions
Funding
Institutional Review Board Statement
Data Availability Statement
Acknowledgments
Conflicts of Interest
References
- Jin, Y; Yang, H; Ji, W; Wu, W; Chen, S; Zhang, W; et al. Virology, epidemiology, pathogenesis, and control of COVID-19. Viruses 2020, 12(4), 372. [Google Scholar] [CrossRef]
- Hu, B; Guo, H; Zhou, P; Shi, Z-L. Characteristics of SARS-CoV-2 and COVID-19. Nature reviews microbiology 2021, 19(3), 141–54. [Google Scholar] [CrossRef]
- Shereen, MA; Khan, S; Kazmi, A; Bashir, N; Siddique, R. COVID-19 infection: Emergence, transmission, and characteristics of human coronaviruses. Journal of advanced research 2020, 24, 91–8. [Google Scholar] [CrossRef] [PubMed]
- Feng, W; Newbigging, AM; Le, C; Pang, B; Peng, H; Cao, Y; et al. Molecular diagnosis of COVID-19: challenges and research needs. Analytical chemistry 2020, 92(15), 10196–209. [Google Scholar] [CrossRef] [PubMed]
- Choi, JY; Smith, DM. SARS-CoV-2 variants of concern. Yonsei medical journal 2021, 62(11), 961. [Google Scholar] [CrossRef] [PubMed]
- Sanyaolu, A; Okorie, C; Marinkovic, A; Haider, N; Abbasi, AF; Jaferi, U; et al. The emerging SARS-CoV-2 variants of concern. Therapeutic advances in infectious disease 2021, 8, 20499361211024372. [Google Scholar] [CrossRef]
- Lin, L; Liu, Y; Tang, X; He, D. The disease severity and clinical outcomes of the SARS-CoV-2 variants of concern. Frontiers in public health 2021, 9, 775224. [Google Scholar] [CrossRef]
- Gupta, P; Gupta, V; Singh, CM; Singhal, L. Emergence of COVID-19 variants: an update. Cureus 2023, 15(7). [Google Scholar] [CrossRef]
- Farahat, RA; Abdelaal, A; Umar, TP; El-Sakka, AA; Benmelouka, AY; Albakri, K; et al. The emergence of SARS-CoV-2 Omicron subvariants: current situation and future trends. Le infezioni in medicina 2022, 30(4), 480. [Google Scholar]
- Viana, R; Moyo, S; Amoako, DG; Tegally, H; Scheepers, C; Althaus, CL; et al. Rapid epidemic expansion of the SARS-CoV-2 Omicron variant in southern Africa. Nature 2022, 603(7902), 679–86. [Google Scholar] [CrossRef] [PubMed]
- Xu, A; Hong, B; Lou, F; Wang, S; Li, W; Shafqat, A; et al. Sub-lineages of the SARS-CoV-2 Omicron variants: Characteristics and prevention. MedComm 2022, 3(3), e172. [Google Scholar] [CrossRef]
- Kevadiya, BD; Machhi, J; Herskovitz, J; Oleynikov, MD; Blomberg, WR; Bajwa, N; et al. Diagnostics for SARS-CoV-2 infections. Nature materials 2021, 20(5), 593–605. [Google Scholar] [CrossRef]
- Afzal, A. Molecular diagnostic technologies for COVID-19: Limitations and challenges. Journal of advanced research 2020, 26, 149–59. [Google Scholar] [CrossRef] [PubMed]
- Rai, P; Kumar, BK; Deekshit, VK; Karunasagar, I; Karunasagar, I. Detection technologies and recent developments in the diagnosis of COVID-19 infection. Applied microbiology and biotechnology 2021, 105(2), 441–55. [Google Scholar] [CrossRef]
- Boan, P; Jardine, A; Pryce, TM. Clinical associations of SARS-CoV-2 viral load using the first WHO international standard for SARS-CoV-2 RNA. Pathology 2022, 54(3), 344–50. [Google Scholar] [CrossRef] [PubMed]
- Dadras, O; Afsahi, AM; Pashaei, Z; Mojdeganlou, H; Karimi, A; Habibi, P; et al. The relationship between COVID-19 viral load and disease severity: A systematic review. Immunity, inflammation and disease 2022, 10(3), e580. [Google Scholar] [CrossRef]
- Specification criteria for COVID-19 molecular Meril COVID-19 One-step RT-PCR Kits MD018: SAHPRA; SAHPRA: South Africa, 2025; Available online: https://www.sahpra.org.za/medical-devices/medical-devices-and-in-vitro-diagnostics-test-kits/.
- Poljak, M; Korva, M; Knap Gašper, N; Fujs Komloš, K; Sagadin, M; Uršič, T; et al. Clinical evaluation of the cobas SARS-CoV-2 test and a diagnostic platform switch during 48 hours in the midst of the COVID-19 pandemic. Journal of Clinical Microbiology 2020, 58(6). [Google Scholar] [CrossRef]
- Perchetti, GA; Pepper, G; Shrestha, L; LaTurner, K; Kim, DY; Huang, M-L; et al. Performance characteristics of the Abbott Alinity m SARS-CoV-2 assay. Journal of Clinical Virology 2021, 140, 104869. [Google Scholar] [CrossRef] [PubMed]
- Newman, H; Maritz, J. Basic overview of method validation in the clinical virology laboratory. Reviews in Medical Virology 2017, 27(5), e1940. [Google Scholar] [CrossRef] [PubMed]
- Pum, J. A practical guide to validation and verification of analytical methods in the clinical laboratory. Advances in clinical chemistry 2019, 90, 215–81. [Google Scholar]
- Subramoney, K; Mtileni, N; Giandhari, J; Naidoo, Y; Ramphal, Y; Pillay, S; et al. Molecular epidemiology of SARS-CoV-2 during five COVID-19 waves and the significance of low-frequency lineages. Viruses 2023, 15(5), 1194. [Google Scholar] [CrossRef] [PubMed]
- Furusawa, Y; Kiso, M; Iida, S; Uraki, R; Hirata, Y; Imai, M; et al. SARS-CoV-2 delta variants, Spike-P681R and D950N promote membrane fusion, Spike-P681R enhances spike cleavage, but neither substitution affects pathogenicity in hamsters. EBioMedicine 2023, 91. [Google Scholar] [CrossRef] [PubMed]
- Gangavarapu, K; Latif, AA; Mullen, JL; Alkuzweny, M; Hufbauer, E; Tsueng, G; et al. Outbreak. info genomic reports: scalable and dynamic surveillance of SARS-CoV-2 variants and mutations. Nature Methods 2023, 20(4), 512–22. [Google Scholar] [CrossRef] [PubMed]
| Type | n | Sensitivity/Specificity (CI) | PPV/NPP (CI) | Cohen Kappa (CI) |
|---|---|---|---|---|
| As described above | 220 | 98.4% (94.2 – 99.8)/ 100% (95.9 – 100.0) |
100%/ 97.8% (91.8 – 99.4) |
0.981 (0.954 – 1.000) |
| Viral Culture (run in quadruplicate) | RdRP gene (mean, SD, CV%) | N gene (mean, SD, CV%) |
|---|---|---|
| 1:1000 | 30.0, 0.24, 0.96 | 32.2, 0.45, 1.4 |
| 1:10000 | 33.5, 0.15, 0.46 | 35.1, 0.47, 1.35 |
| 1:100000 | 36.7, 0.82, 2.23 | 38.9, 0.95, 2.45 |
| 1:1000000 | 38.75, 1.49, 3.83 | - |
| VOC | del69-70 | E484K | N501Y | D80A |
|---|---|---|---|---|
| Alpha | ||||
| Beta | ||||
| Gamma | ||||
| Delta | ||||
| Omicron |
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2026 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).




