Submitted:
20 March 2026
Posted:
10 April 2026
You are already at the latest version
Abstract
Keywords:
1. Introduction
2. Materials and Methods
2.1. Data Sources and Gene Set Curation
2.2. Genomic Coordinate Processing
2.3. LINE-1 Annotation and Intersection Analysis
2.4. Statistical Analysis
3. Results and Discussion
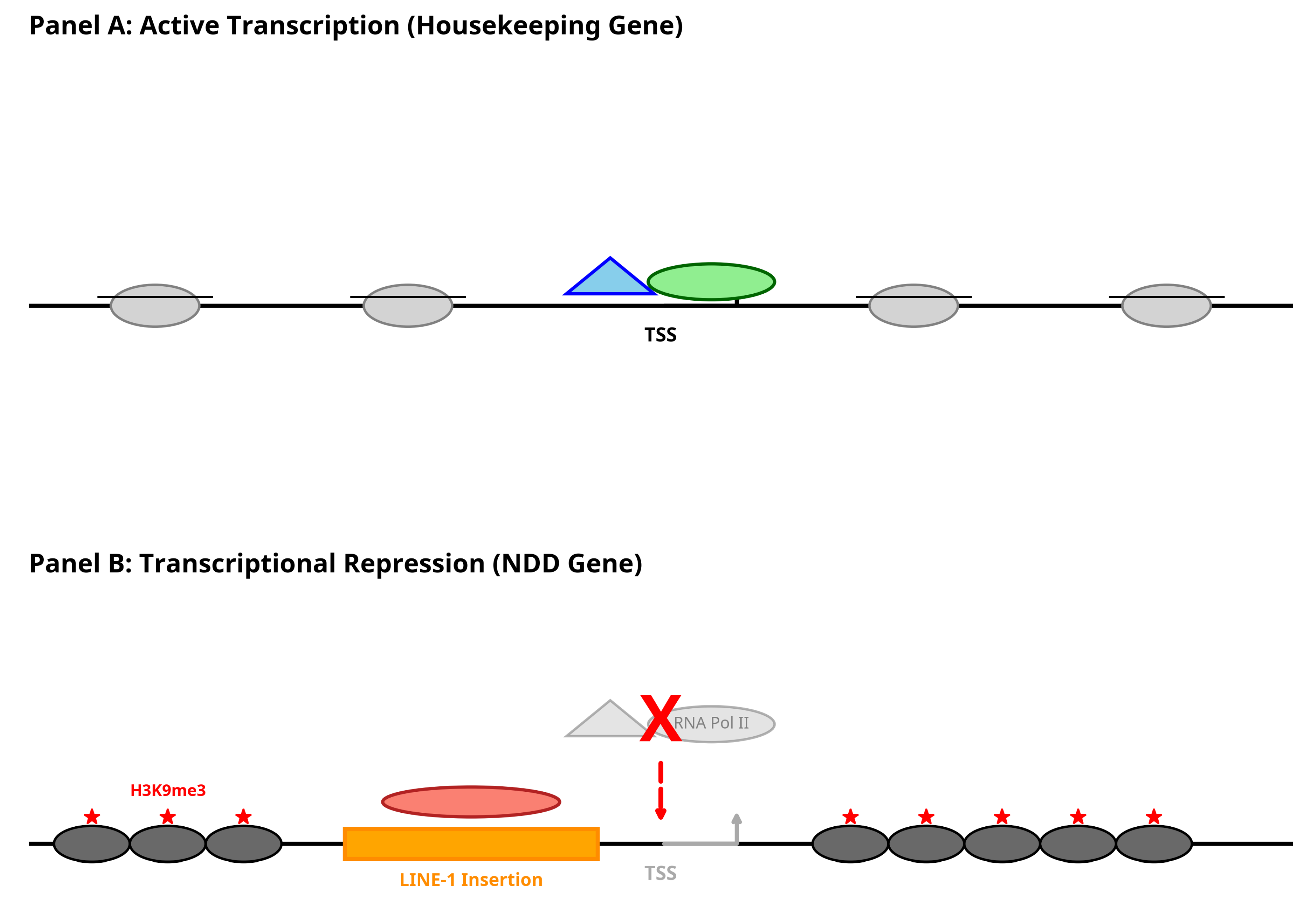
3.1. LINE-1 Occupancy Is Reduced in NDD Gene Promoters
3.2. A Gradient of LINE-1 Depletion Across NDD Subgroups
3.3. Intronic LINE-1 Enrichment Is Primarily Driven by Gene Length Confounding
3.4. Promoter GC Content and CpG Density Do Not Explain LINE-1 Depletion
3.5. Interpretation of Main Findings
3.6. Limitations
4. Conclusions
Funding
Author Contributions: Can Sevilmiş
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Use of Artificial Intelligence
Acknowledgments
Conflicts of Interest
References
- Lander, E.S.; et al. Initial sequencing and analysis of the human genome. Nature 2001, 409, 860–921. [Google Scholar] [CrossRef]
- ENCODE Project Consortium. An integrated encyclopedia of DNA elements in the human genome. Nature 2012, 489, 57–74. [Google Scholar] [CrossRef]
- Ardeljan, D.; Taylor, M.S.; Ting, D.T.; Burns, K.H. The human LINE-1 retrotransposon: an emerging biomarker of neoplasia. Clinical Chemistry 2017, 63, 816–822. [Google Scholar] [CrossRef]
- Beck, C.R.; Collier, P.; Macfarlane, C.; Malig, M.; Kidd, J.M.; Eichler, E.E.; Badge, R.M.; Moran, J.V. LINE-1 retrotransposition activity in human genomes. Cell 2010, 141, 1159–1170. [Google Scholar] [CrossRef]
- Lavie, L.; Maldener, E.; Brouha, B.; Meese, E.U.; Mayer, J. The human L1 promoter: variable transcription initiation sites and a major impact of upstream flanking sequence on promoter activity. Genome Research 2004, 14, 2253–2260. [Google Scholar] [CrossRef]
- Honda, T.; Nishikawa, Y.; Nishimura, K.; Teng, D.; Takemoto, K.; Ueda, K. Effects of activation of the LINE-1 antisense promoter on the growth of cultured cells. Scientific Reports 2020, 10, 22136. [Google Scholar] [CrossRef] [PubMed]
- Jönsson, M.E.; et al. Activation of neuronal genes via LINE-1 elements upon global DNA demethylation in human neural progenitors. Nature Communications 2019, 10, 3182. [Google Scholar] [CrossRef]
- Muotri, A.R.; Chu, V.T.; Marchetto, M.C.; Deng, W.; Moran, J.V.; Gage, F.H. Somatic mosaicism in neuronal precursor cells mediated by L1 retrotransposition. Nature 2005, 435, 903–910. [Google Scholar] [CrossRef] [PubMed]
- Coufal, N.G.; et al. L1 retrotransposition in human neural progenitor cells. Nature 2009, 460, 1127–1131. [Google Scholar] [CrossRef]
- Singer, T.; McConnell, M.J.; Marchetto, M.C.; Coufal, N.G.; Gage, F.H. LINE-1 retrotransposons: mediators of somatic variation in neuronal genomes? Trends in Neurosciences 2010, 33, 345–354. [Google Scholar] [CrossRef]
- Abrahams, B.S.; et al. SFARI Gene 2.0: a community-driven knowledgebase for the autism spectrum disorders. Molecular Autism 2013, 4, 36. [Google Scholar] [CrossRef] [PubMed]
- Köhler, S.; et al. The Human Phenotype Ontology in 2021. Nucleic Acids Research 2021, 49, D1207–D1217. [Google Scholar] [CrossRef]
- Fan, H.C.; Chiang, K.L.; Chang, K.H.; Chen, C.M.; Tsai, J.D. Epilepsy and attention deficit hyperactivity disorder: connection, chance, and challenges. International Journal of Molecular Sciences 2023, 24, 5270. [Google Scholar] [CrossRef] [PubMed]
- Hounkpe, B.W.; Chenou, F.; de Lima, F.; De Paula, E.V. HRT Atlas v1.0 database: redefining human and mouse housekeeping genes and candidate reference transcripts by mining massive RNA-seq datasets. Nucleic Acids Research 2021, 49, D947–D955. [Google Scholar] [CrossRef]
- Frankish, A.; et al. GENCODE: reference annotation for the human and mouse genomes in 2023. Nucleic Acids Research 2023, 51, D942–D949. [Google Scholar] [CrossRef]
- Smit, A.; Hubley, R.; Green, P. RepeatMasker Open-4.0, 2013. Available online: http://www.repeatmasker.org.
- Quinlan, A.R.; Hall, I.M. BEDTools: a flexible suite of utilities for comparing genomic features. Bioinformatics 2010, 26, 841–842. [Google Scholar] [CrossRef]
- Simonti, C.N.; Pavličev, M.; Capra, J.A. Transposable Element Exaptation into Regulatory Regions Is Rare, Influenced by Evolutionary Age, and Subject to Pleiotropic Constraints. Molecular Biology and Evolution 2017, 34, 2856–2869. [Google Scholar] [CrossRef]
- You, J.S.; Pierce, S.; Liang, G.; Jones, P.A. Roles of transposable elements and DNA methylation in the formation of CpG islands and CpG-depleted regulatory elements. Proceedings of the National Academy of Sciences 2025, 122, e2502963122. [Google Scholar] [CrossRef] [PubMed]
- Zug, R. Developmental disorders caused by haploinsufficiency of transcriptional regulators: a perspective based on cell fate determination. Biology Open 2022, 11, bio058896. [Google Scholar] [CrossRef]
- Darbandi, S.F.; An, J.Y.; Lim, K.; Page, N.F.; Liang, L.; Young, D.M.; Ypsilanti, A.R.; State, M.W.; Nord, A.S.; Sanders, S.J.; et al. Five autism-associated transcriptional regulators target shared loci proximal to brain-expressed genes. Cell Reports 2024, 43, 114329. [Google Scholar] [CrossRef]
- Koesterich, J.; An, J.Y.; Inoue, F.; Sohota, A.; Ahituv, N.; Sanders, S.J.; Kreimer, A. Characterization of De Novo Promoter Variants in Autism Spectrum Disorder with Massively Parallel Reporter Assays. International Journal of Molecular Sciences 2023, 24, 3509. [Google Scholar] [CrossRef]
- Liang, K.C.; Tseng, J.T.; Tsai, S.J.; Sun, H.S. Characterization and distribution of repetitive elements in association with genes in the human genome. Computational Biology and Chemistry 2015, 57, 29–38. [Google Scholar] [CrossRef]
- Chuong, E.B.; Elde, N.C.; Feschotte, C. Regulatory activities of transposable elements: from conflicts to benefits. Nature Reviews Genetics 2016, 18, 71–86. [Google Scholar] [CrossRef]
- Lu, J.Y.; Shao, W.; Chang, L.; Yin, Y.; Li, T.; Zhang, H.; Hong, Y.; Percharde, M.; Guo, L.; Wu, Z.; et al. Genomic Repeats Categorize Genes with Distinct Functions for Orchestrated Regulation. Cell Reports 2020, 30, 3296–3311. [Google Scholar] [CrossRef] [PubMed]
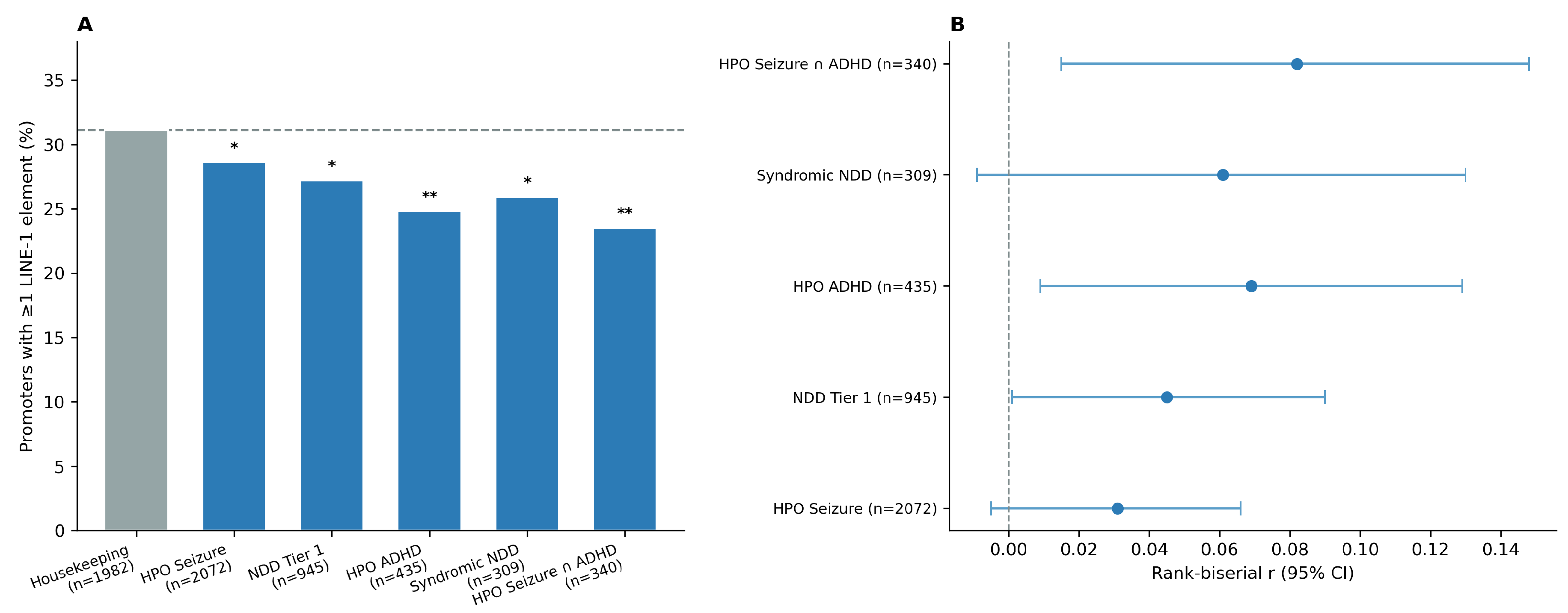
| Gene Set | n | LINE-1 (%) | p-value | r (95% CI) |
|---|---|---|---|---|
| Housekeeping | 1,982 | 31.1 | — | — |
| HPO Seizure | 2,072 | 28.6 | 0.0360 | 0.031 (−0.00, 0.07) |
| NDD Tier 1 (SFARI score 1–2) | 945 | 27.2 | 0.0138 | 0.045 (0.00, 0.09) |
| HPO ADHD | 435 | 24.8 | 0.0051 | 0.069 (0.01, 0.13) |
| Syndromic NDD | 309 | 25.9 | 0.0344 | 0.061 (−0.01, 0.13) |
| HPO Seizure ∩ ADHD | 340 | 23.5 | 0.0029 | 0.082 (0.02, 0.15) |
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2026 by the author. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).




