Submitted:
09 April 2026
Posted:
10 April 2026
You are already at the latest version
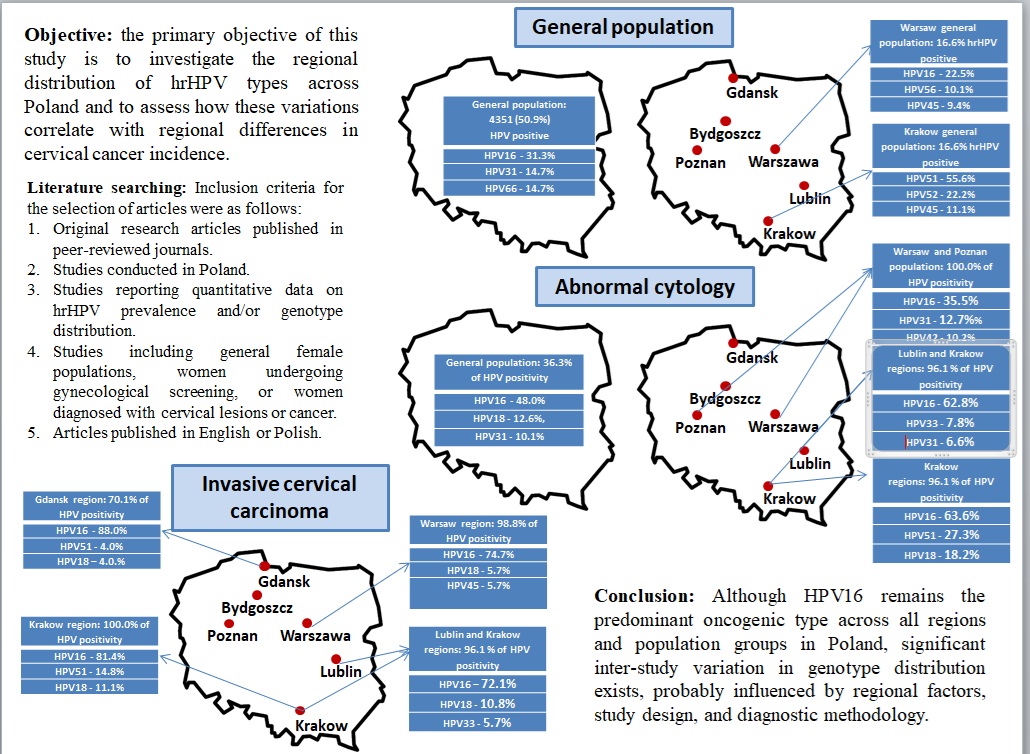
Abstract
Keywords:
1. Introduction
2. Literature Search
- Original research articles published in peer-reviewed journals.
- Studies conducted in Poland.
- Studies reporting quantitative data on hrHPV prevalence and/or genotype distribution.
- Studies including general female populations, women undergoing gynecological screening, or women diagnosed with cervical lesions or cancer.
- Articles published in English or Polish.
- Review articles, conference abstracts, case reports, and editorials.
- Studies with insufficient data on HPV genotype distribution.
- Studies using outdated or non-specific HPV detection methods without genotype resolution.
3. Results
3.1. Human Papillomavirus Prevalence and Genotype Distribution in the General Female Population Across Different Regions of Poland
3.2. Human Papillomavirus Prevalence and Genotype Distribution Among Women with Abnormal Cytology Across Different Regions of Poland
3.3. The prevalence and Genotype Distribution of Human Papillomavirus in Women with Invasive Cervical Cancer (ICC) Across Different Regions of Poland
4. Interpretation of Findings
5. Conclusions
Author Contributions
Funding
Conflicts of Interest
References
- Bray, F.; Laversanne, M.; Sung, H.; Ferlay, J.; Siegel, R.L.; Soerjomataram, I.; Jemal, A. Global cancer statistics 2022: GLOBOCAN estimates of incidence and mortality worldwide for 36 cancers in 185 countries. CA Cancer J Clin. 2024, 74(3), 229–263. [Google Scholar] [CrossRef]
- Didkowska, J.; Barańska, K.; Miklewska, M.J.; Wojciechowska, U. Cancer incidence and mortality in Poland in 2023. Nowotwory. Journal of Oncology 2024, 74(2), 75–93. [Google Scholar] [CrossRef]
- IARC Working Group on the Evaluation of Carcinogenic Risks to Humans. Biological agents. IARC Monogr Eval Carcinog Risks Hum. 2012, 100 Pt B, 1–441.
- Human Papillomaviruses. IARC Monographs on the Evaluation of Carcinogenic Risks to Humans, IARC. 2007, ISBN-13, 978-92-832-1290-4.
- Target product profiles for human papillomavirus screening tests to detect cervical precancer and cancer. Geneva: World Health Organization 2024, Licence: CC BY-NC-SA 3.0 IGO.
- Osmani, V.; Hörner, L.; Nkurunziza, T.; Rank, S.; Tanaka, L.F.; Klug, S.J. Global prevalence of cervical human papillomavirus in women aged 50 years and older with normal cytology: a systematic review and meta-analysis. Lancet Microbe 2025, 6(1), 100955. [Google Scholar] [CrossRef]
- Bruni, L.; Diaz, M.; Castellsagué, X.; Ferrer, E.; Bosch, F.X.; de Sanjosé, S. Cervical human papillomavirus prevalence in 5 continents: meta-analysis of 1 million women with normal cytological findings. J Infect Dis. 2010, 15;202(12), 1789–99. [Google Scholar] [CrossRef]
- de Sanjosé, S.; Diaz, M.; Castellsagué, X.; Clifford, G.; Bruni, L.; Muñoz, N.; Bosch, FX. Worldwide prevalence and genotype distribution of cervical human papillomavirus DNA in women with normal cytology: a meta-analysis. Lancet Infect Dis. 2007, 7(7), 453–9. [Google Scholar] [CrossRef]
- Clifford, G.M.; Gallus, S.; Herrero, R.; Muñoz, N.; Snijders, P.J.; Vaccarella, S.; Anh, P.T.; Ferreccio, C.; Hieu, N.T.; Matos, E.; Molano, M.; Rajkumar, R.; Ronco, G.; de Sanjosé, S.; Shin, H.R.; Sukvirach, S.; Thomas, J.O.; Tunsakul, S.; Meijer, C.J.; Franceschi, S. Worldwide distribution of human papillomavirus types in cytologically normal women in the International Agency for Research on Cancer HPV prevalence surveys: a pooled analysis. Lancet 2005, 366, 991–98. [Google Scholar] [CrossRef]
- Clifford, G.M.; Rana, R.K.; Franceschi, S.; Smith, J.S.; Gough, G.; Pimenta, J.M. Human papillomavirus genotype distribution in low-grade cervical lesions: comparison by geographic region and with cervical cancer. Cancer Epidemiol Biomarkers Prev. 2005, 14(5), 1157–64. [Google Scholar] [CrossRef] [PubMed]
- Wei, F.; Georges, D.; Man, I.; Baussano, I.; Clifford, G.M. Causal attribution of human papillomavirus genotypes to invasive cervical cancer worldwide: a systematic analysis of the global literature. Lancet 2024, 404(10451), 43. [Google Scholar] [CrossRef] [PubMed]
- Guan, P.; Howell-Jones, R.; Li, N.; Bruni, L.; de Sanjosé, S.; Franceschi, S.; Franceschi, S.; Clifford, G.M. Human Papillomavirus Types in 115,789 HPV-Positive Women: A Meta-Analysis from Cervical Infection to Cancer. Int J Cancer 2012, 131, 2349–2359. [Google Scholar] [CrossRef]
- Li, N.; Franceschi, S.; Howell-Jones, R.; Snijders, P.J.; Clifford, G.M. Human papillomavirus type distribution in 30,848 invasive cervical cancers worldwide: Variation by geographical region, histological type and year of publication. Int J Cancer 2011, 128(4), 927–35. [Google Scholar] [CrossRef]
- de Sanjose, S.; Quint, W.G.; Alemany, L.; Geraets, D.T.; Klaustermeier, J.E.; Lloveras, B.; Tous, S.; Felix, A.; Bravo, L.E.; Shin, H.R.; Vallejos, C.S.; de Ruiz, P.A.; Lima, M.A.; Guimera, N.; Clavero, O.; Alejo, M.; Llombart-Bosch, A.; Cheng-Yang, C.; Tatti, S.A.; Kasamatsu, E.; Iljazovic, E.; Odida, M.; Prado, R.; Seoud, M.; Grce, M.; Usubutun, A.; Jain, A.; Suarez, G.A.; Lombardi, L.E.; Banjo, A.; Menéndez, C.; Domingo, E.J.; Velasco, J.; Nessa, A.; Chichareon, S.C.; Qiao, Y.L.; Lerma, E.; Garland, S.M.; Sasagawa, T.; Ferrera, A.; Hammouda, D.; Mariani, L.; Pelayo, A.; Steiner, I.; Oliva, E.; Meijer, C.J.; Al-Jassar, W.F.; Cruz, E.; Wright, T.C.; Puras, A.; Llave, C.L.; Tzardi, M.; Agorastos, T.; Garcia-Barriola, V.; Clavel, C.; Ordi, J.; Andújar, M.; Castellsagué, X.; Sánchez, G.I.; Nowakowski, A.M.; Bornstein, J.; Muñoz, N.; Bosch, F.X. Retrospective International Survey and HPV Time Trends Study Group. Human papillomavirus genotype attribution in invasive cervical cancer: a retrospective cross-sectional worldwide study. Lancet Oncol. 2010, 11(11), 1048–56. [Google Scholar] [CrossRef] [PubMed]
- Smith, J.S.; Lindsay, L.; Hoots, B.; Keys, J.; Franceschi, S.; Winer, R.; Clifford, G.M. Human papillomavirus type distribution in invasive cervical cancer and high-grade cervical lesions: a meta-analysis update. Int J Cancer 2007, 121(3), 621–32. [Google Scholar] [CrossRef]
- Clifford, G.M.; Smith, J.S.; Aguado, T.; Franceschi, S. Comparison of HPV type distribution in high-grade cervical lesions and cervical cancer: a meta-analysis. Br J Cancer 2003, 89, 101–05. [Google Scholar] [CrossRef]
- Pinheiro, M.; Gage, J.C.; Clifford, G.M.; Demarco, M.; Cheung, L.C.; Chen, Z.; Yeager, M.; Cullen, M.; Boland, J.F.; Chen, X.; Raine-Bennett, T.; Steinberg, M.; Bass, S.; Befano, B.; Xiao, Y.; Tenet, V.; Walker, J.; Zuna, R.; Poitras, N.E.; Gold, M.A.; Dunn, T.; Yu, K.; \; Zhu, B.; Burdett, L.; Turan, S.; Lorey, T.; Castle, P.; Wentzensen, N.; Burk, R.; Schiffman, M.; Mirabello, L. Association of HPV35 with cervical carcinogenesis among women of African ancestry: Evidence of viral-host interaction with implications for disease intervention. Int J Cancer 2020, 147(10), 2677–2686. [Google Scholar] [CrossRef]
- Poljak, M.; Seme, K.; Maver, P.J.; Kocjan, B.; Cuschieri, K.S.; Rogovskaya, S.I.; Arbyn, M.; Syrjänen, S. Human papillomavirus prevalence and type-distribution, cervical cancer screening practices and current status of vaccination implementation in Central and Eastern Europe. Vaccine 2013, 31 Suppl 7, H59–70. [Google Scholar] [CrossRef]
- Reuschenbach, M.; Valente, S.; Takyar, J.; Dhawan, A.; Hall, A.; Agrawal, N.; Ghelardi, A.; Del Pino, M.; Nowakowski, A.; Sabale, U. Treatment characteristics, HPV genotype distribution and risk of subsequent disease among women with high-grade cervical intraepithelial neoplasia in Europe: A systematic literature review. Eur J Obstet Gynecol Reprod Biol. 2024, 300, 129–140. [Google Scholar] [CrossRef]
- Tjalma, W.A.; Fiander, A.; Reich, O.; Powell, N.; Nowakowski, A.M.; Kirschner, B.; Koiss, R.; O'Leary, J.; Joura, E.A.; Rosenlund, M.; Colau, B.; Schledermann, D.; Kukk, K.; Damaskou, V.; Repanti, M.; Vladareanu, R.; Kolomiets, L.; Savicheva, A.; Shipitsyna, E.; Ordi, J.; Molijn, A.; Quint, W.; Raillard, A.; Rosillon, D.; De Souza, S.C.; Jenkins, D.; Holl, K. HERACLES/SCALE Study Group. Differences in human papillomavirus type distribution in high-grade cervical intraepithelial neoplasia and invasive cervical cancer in Europe. Int J Cancer 2013, 132(4), 854–67. [Google Scholar] [CrossRef] [PubMed]
- Liang, L.A.; Tanaka, L.F.; Radde, K.; Bussas, U.; Ikenberg, H.; Heideman, D.A.M.; Meijer, C.J.L.M.; Blettner, M.; Klug, S.J. Population-based age- and type-specific prevalence of human papillomavirus among non-vaccinated women aged 30 years and above in Germany. BMC Infect Dis. 2024, 24(1), 1008. [Google Scholar] [CrossRef] [PubMed]
- Cenci, M.; Rossi, F.; Pisani, T. Detection of 14 High-risk Human Papillomavirus (HPV) Genotypes Within the Italian Cervical Cancer Screening. In Vivo 2023, 37(5), 2161–2165. [Google Scholar] [CrossRef]
- López, N.; Torné, A.; Franco, A.; San-Martin, M.; Viayna, E.; Barrull, C.; Perulero, N. Epidemiologic and economic burden of HPV diseases in Spain: implication of additional 5 types from the 9-valent vaccine. Infect Agent Cancer 2018, 13, 15. [Google Scholar] [CrossRef] [PubMed]
- Glińska, P.; Macios, A.; Jaworski, R.; Bobinski, M.; Pruski, D.; Przybylski, M.; Zielinska, A.; Sawicki, W.; Nowakowski, A. Baseline data on distribution of human papillomavirus (HPV) genotypes in cervical samples of gynecological patients before implementation of population-based HPV vaccination program in Poland. Ginekol Pol. 2024, 95(11), 870–878. [Google Scholar] [CrossRef]
- Bardin, A.; Vaccarella, S.; Clifford, G.M.; Lissowska, J.; Rekosz, M.; Bobkiewicz, P.; Kupryjańczyk, J.; Krynicki, R.; Jonska-Gmyrek, J.; Danska-Bidzinska, A.; Snijders, P.J.; Meijer, C.J.; Zatonski, W.; Franceschi, S. Human papillomavirus infection in women with and without cervical cancer in Warsaw, Poland. Eur J Cancer 2008, 44(4), 557–64. [Google Scholar] [CrossRef]
- Szostek, S.; Klimek, M.; Zawilinska, B.; Kosz-Vnenchak, M. Genotype-specific human papillomavirus detection in cervical smears. Acta Biochim Pol. 2008, 55(4), 687–92. [Google Scholar] [CrossRef]
- Bebyn, M.G.; Śledzińska, P.; Wojtysiak, J.; Jóźwicki, W.; Mierzwa, T.; Dziedzic, J.; Kowalewski, J.; Lewandowska, M. HPV RNA and DNA testing in Polish women screened for cervical cancer - A single oncological center study. Eur J Obstet Gynecol Reprod Biol. 2022, 268, 129–134. [Google Scholar] [CrossRef]
- Kiwerska, K.; Jozefiak, A.; Markowska, J.; Kedzia, W.; Jackowska, J.; Wierzbicka, M. Oral-genital human papillomavirus infection in Polish couples: frequent detection of HPV 42. BMC Infect Dis. 2019, 9(1), 122. [Google Scholar] [CrossRef] [PubMed]
- Nowakowski, A.; de Souza, S.C.; Jach, R.; Rosillon, D.; Książek, A.; Holl, K. HPV-type distribution and reproducibility of histological diagnosis in cervical neoplasia in Poland. Pathol Oncol Res. 2015, 21(3), 703–11. [Google Scholar] [CrossRef]
- Liss, J.; Łukaszuk, K.; Gulczyński, J.; Zwaliński, M.; Woźniak, I.; Emerich, J.; Wójcikowski, Cz. Występowanie DNA wirusa HPV u pacjentek z rakiem szyjki macicy w regionie gdańskim. Ginekol Pol. 2002, 73(9), 740–745. [Google Scholar] [PubMed]
| Authors [year of publication] | Cohort | Geagraphical region | Material |
Method of HPV detection |
hrHPV frequency |
|---|---|---|---|---|---|
| Glińska et al. [2024] [24] |
11 151 woman gynecological tested between 2018 - 2021 |
556 sites across Poland located all over the country in all 16 voivodeshipes | Cervical cytology material |
Amplisens HPV HCR genotype –titre FRT PCR KITand Alinity m HR HPV Assay/Abbot, BD OncoclarityLinear Array HPV Genotyping Test, Roche; Ampliquality HPV-type express v. 3.0, AB Analitica; Inno-LIPA Genotyping Extra II, Fuji Rebio |
General population: 4351 (50.9%) HPV positive: HPV16 - 31.3% HPV31 - 14.7% HPV66 - 14.7% HPV51 – 13.0%, HPV56 - 11.0% HPV52 - 10.1% HPV39 - 8.3% HPV18 - 7.4% HPV18 -7.4%, HPV58 - 7.2%, HPV59 - 6.8% HPV45 - 6.4% HPV33 - 6.0% HPV68 - 4.6% HPV35 - 2.6% |
| Baradin et al. [2008] [25] | 834 woman from general population |
Warsaw region |
Exfoliated cervical cells and cervical cancer biopsy |
HPV DNA Immunoassay with oligoprobe cocktails, detection of 44 HPV genotypes |
General population: 138 (16.6%) hrHPV positive HPV16 - 22.5% HPV56 - 10.1% HPV45 - 9.4% HPV31 - 8.7%, HPV52 - 8.7% HPV33 - 6.5% HPV58 - 5.1% HPV18 - 4.3%, HPV73 - 4.3%, HPV35 - 2.2% HPV39 - 2.2% HPV59 - 2.2% HPV68 - 2.2% HPV82 - 2.2% |
| Szostek et al. [2008] [26] | 42woman without abnormality in cytology |
Krakow region |
Cervical cytology material | INNO-LiPA HPV Genotyping |
Normal cytology population: 9 (21.0%) hrHPV: HPV51 - 55.6% HPV52 - 22.2% HPV45 - 11.1% HPV56 - 11.1% HPV66 - 11.1% |
|
Authors [year of publication] |
Cohort | Geagraphical region | Material |
Method of HPV detection |
hrHPV frequency |
|---|---|---|---|---|---|
| Bebyn et al. (2022) [27] |
1840 most with abnormal cytology LSIL, HSIL,ASC-US, ASC-H, AGC |
Bydgoszcz region |
Cervical cytology material |
mRNA analysis NucliSENS EasyQ HPV (detection of five HPV genotypes) |
General population: 198 (36.3%) HPV RNA positive HPV16 - 48.0% HPV18 - 12.6%, HPV31 - 10.1%, HPV33 - 8.6%, HPV45 - 4.5% More than one HPV type – 16.2% |
| Kiwerska et al. (2019) [28] |
197 females with cervical pathology, including cancer 2014-2016 |
Warsaw and Poznan regions |
Cervical cytology material | AnyplexTM IIHPV28 Detection system, Segene |
General population: 197 (100.0%) HPV positive: HPV16 - 35.5% HPV31 - 12.7% HPV42 – 10.2% HPV39 - 9.1% HPV54 – 8.1%, HPV18 - 7.1% HPV33 - 6.6% HPV53 - 6.6% HPV35 - 6.1% HPV66 - 5.1%, HPV56 - 4.7% HPV51 - 4.1% HPV68 - 4.1% HPV58 - 2.5% HPV59 - 2.5% HPV45 - 1.5% HPV52 – 0.5% |
| Nowakowski et al. (2014) [29] | 205 woman with HG-CIN and 193 ICC between 2001 - 2008 |
Lublin and Krakow regions |
FFPE | SPF10-DEIAliPA25-polymerase chain reaction (PCR) system |
HG-CIN poplutaion: : 197 (96.1%) hrHPV positive: HPV16 - 62.8% HPV33 - 7.8% HPV31 - 6.6% HPV52 - 3.7% HPV45 - 2.6% HPV58 - 2.6% |
| Szostek et al. (2008) [26] | 125 woman with LSIL (n=44), HSIL (n=12) |
Krakow region |
Cervical cytology material | INNO-LiPA HPV Genotyping |
L-SIL population: 43 (98.0%) hrHPV: HPV51 - 30.2% HPV52 - 25.6% HPV18 - 11.6% HPV31 - 11.6% HPV16 - 9.3% HPV33 – 7.0% HPV58 - 7.0% HPV66 – 7.0% HPV56 - 4.6% HPV35 - 2.3% HPV39 - 2.3% H-SIL population: 11 (92.0%) hrHPV: HPV16 - 63.6% HPV51 – 27.3% HPV18 - 18.2% HPV31 – 9.1% |
| Authors [year of publication] | Cohort | Geagraphical region | Material | Method of HPV detection | hrHPV frequency |
|---|---|---|---|---|---|
| Nowakowski et al. (2014) [29] |
193 pts with ICC 2001 - 2008 |
Lublin and Krakow regions |
FFPE | SPF10-DEIAliPA25-polymerase chain reaction (PCR) system |
176 (91.2%) hrHPV positive pts: HPV16 - 72.1% HPV18 - 10.8% HPV33 - 5.7% HPV45 - 3.4% |
| Baradin et al. [2008] [25] |
88 pts with ICC |
Warsaw region |
Exfoliated cervical cells and cervical cancer biopsy |
HPV DNA Immunoassay with oligoprobe cocktails, detection of 44 HPV genotypes |
87 (98.8%) hrHPV positive pts : HPV16 - 74.7% HPV18 - 5.7%, HPV45 - 5.7% HPV31 - 3.4%, HPV52 – 3.4% HPV56 – 3.4% |
| Szostek et al. [2008] [26] |
27 pts with ICC |
Southern Poland |
Cervical cytology material | INNO-LiPA HPV Genotyping |
27 (100.0%) hrHPV positive pts HPV16 - 81.4% HPV51 - 14.8% HPV18 - 11.1% HPV66 – 11.1% HPV45 – 2.2% HPV68 – 2.2% |
| Liss et al. (2002) [30] |
107 pts with ICC 1997 - 1998 |
Gdańsk region |
Cervical cytology material | DNA PCR |
75 (70.1%) hrHPV positive pts: HPV16 - 88.0% HPV31 - 4.0%, HPV33 - 4.0% HPV45 - 1.3%, HPV52 – 1.3% |
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2026 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).




