Submitted:
08 April 2026
Posted:
13 April 2026
You are already at the latest version
Abstract
Keywords:
Introduction
Methods
Open-Source Code
AI-assisted Pipeline
Shape Dataset
Braitenberg Vehicle Dataset
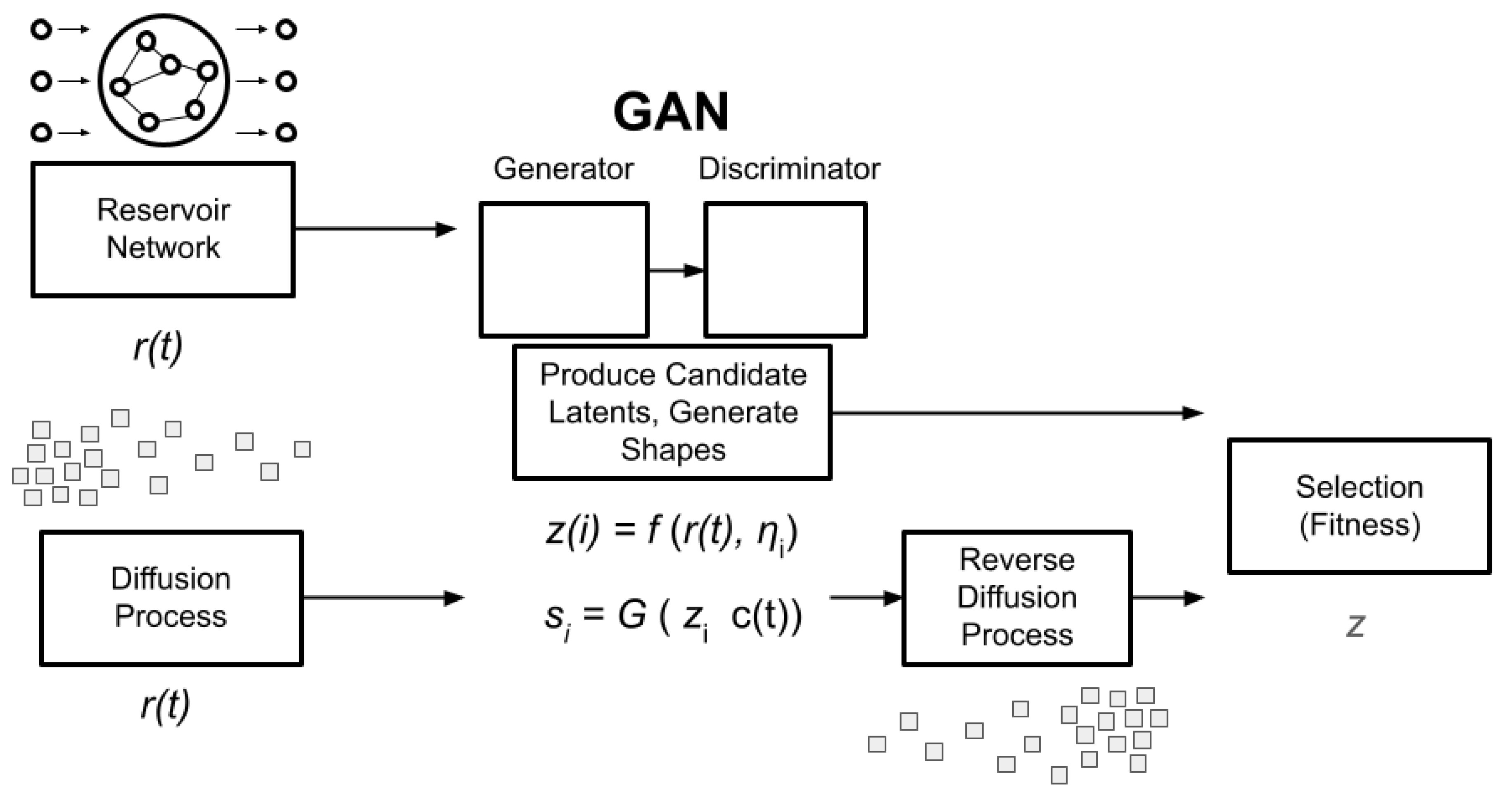
Reservoir Network
Diffusion Process
GAN Implementation
Evolutionary Algorithm
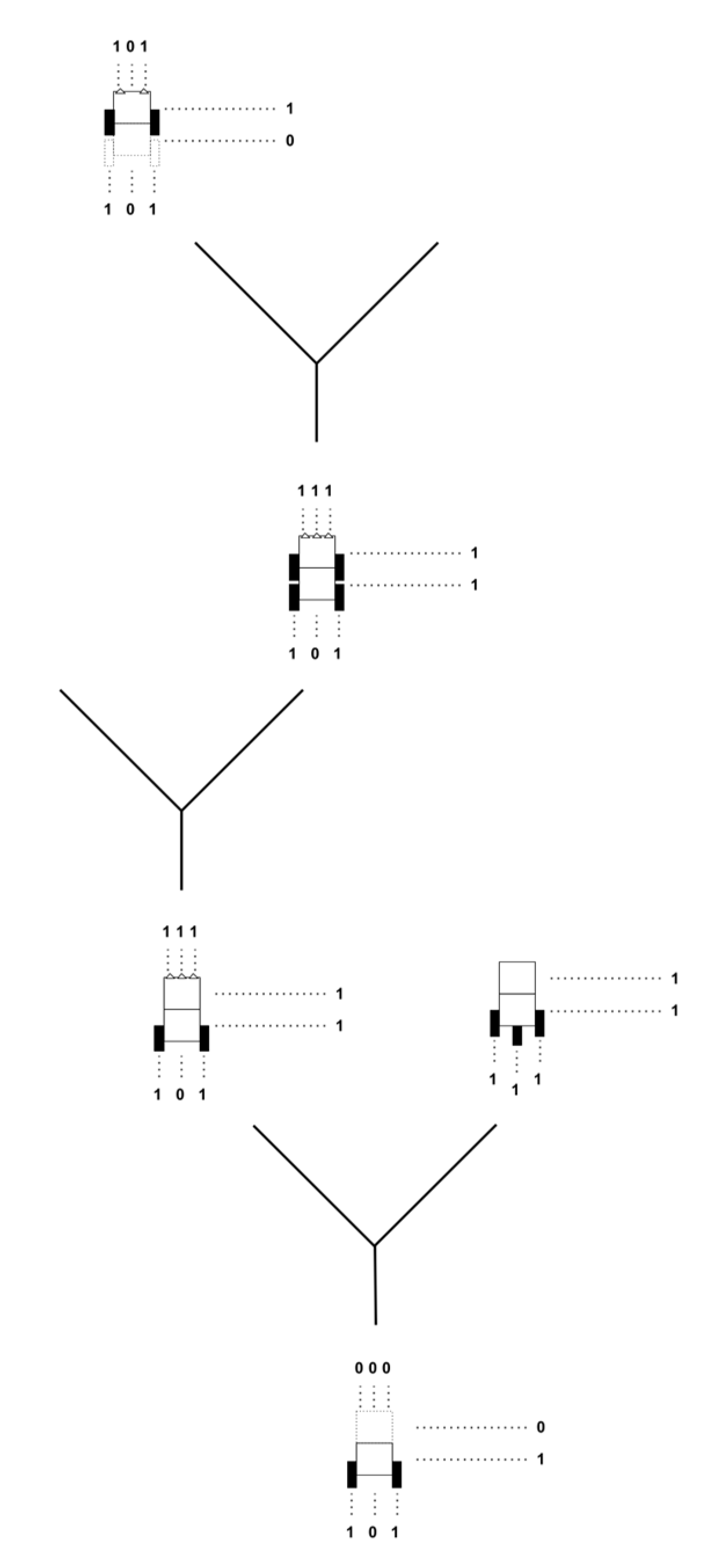
Evolutionary Trees
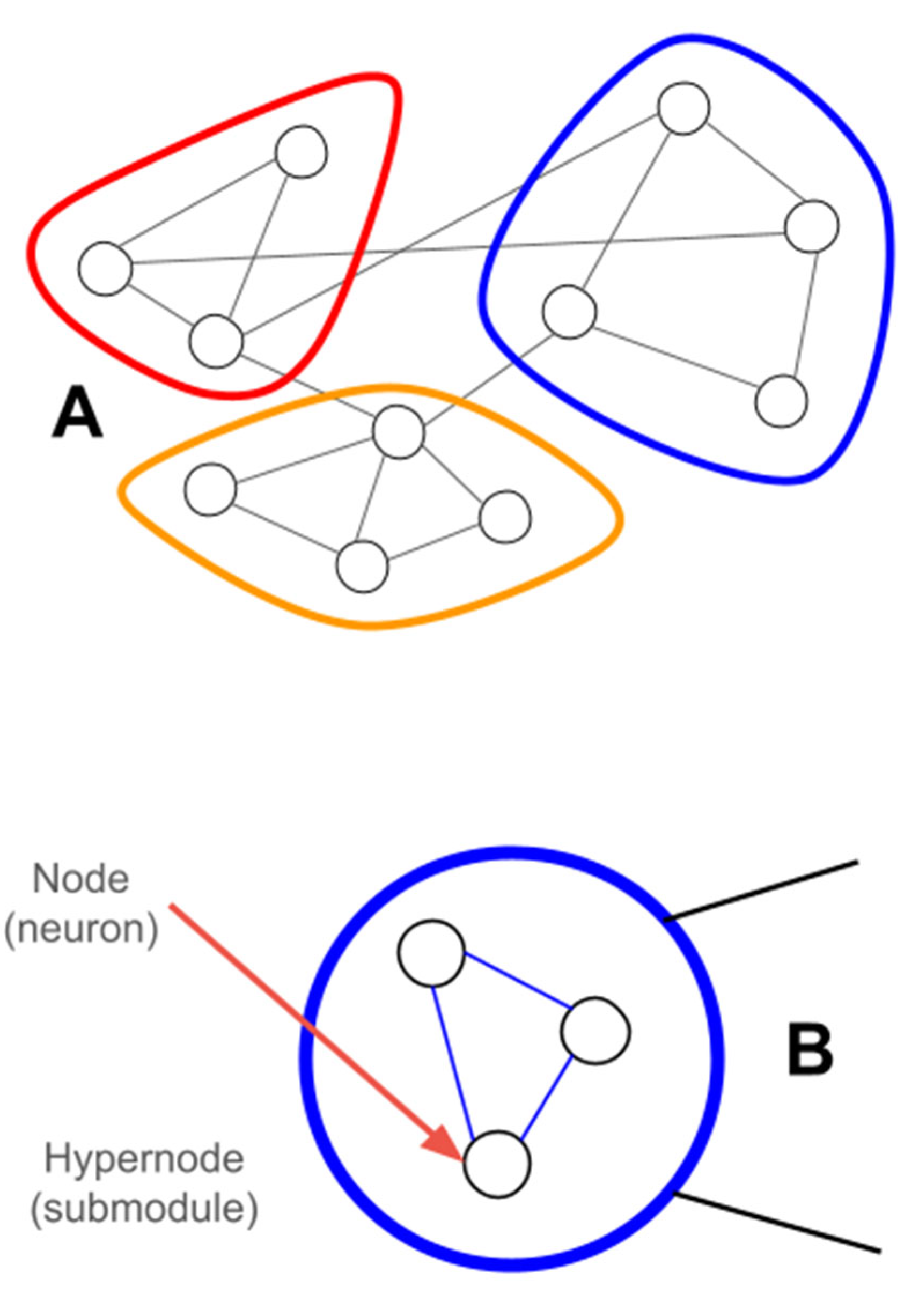
Hypergraphs
Quasi-Experimental Approach
Core Architectural Pipeline
Complexity Bands
Fitness Function
Results
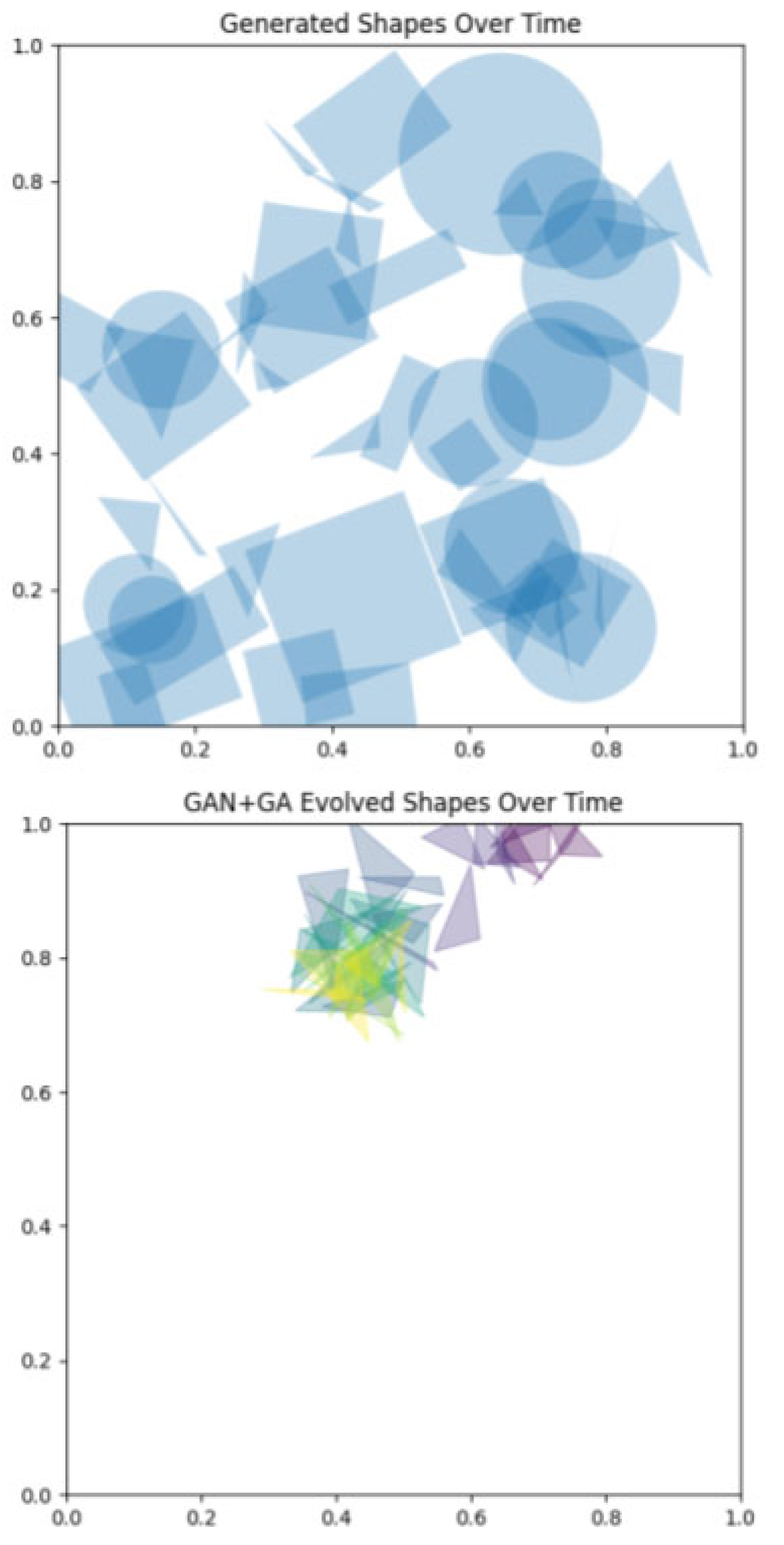
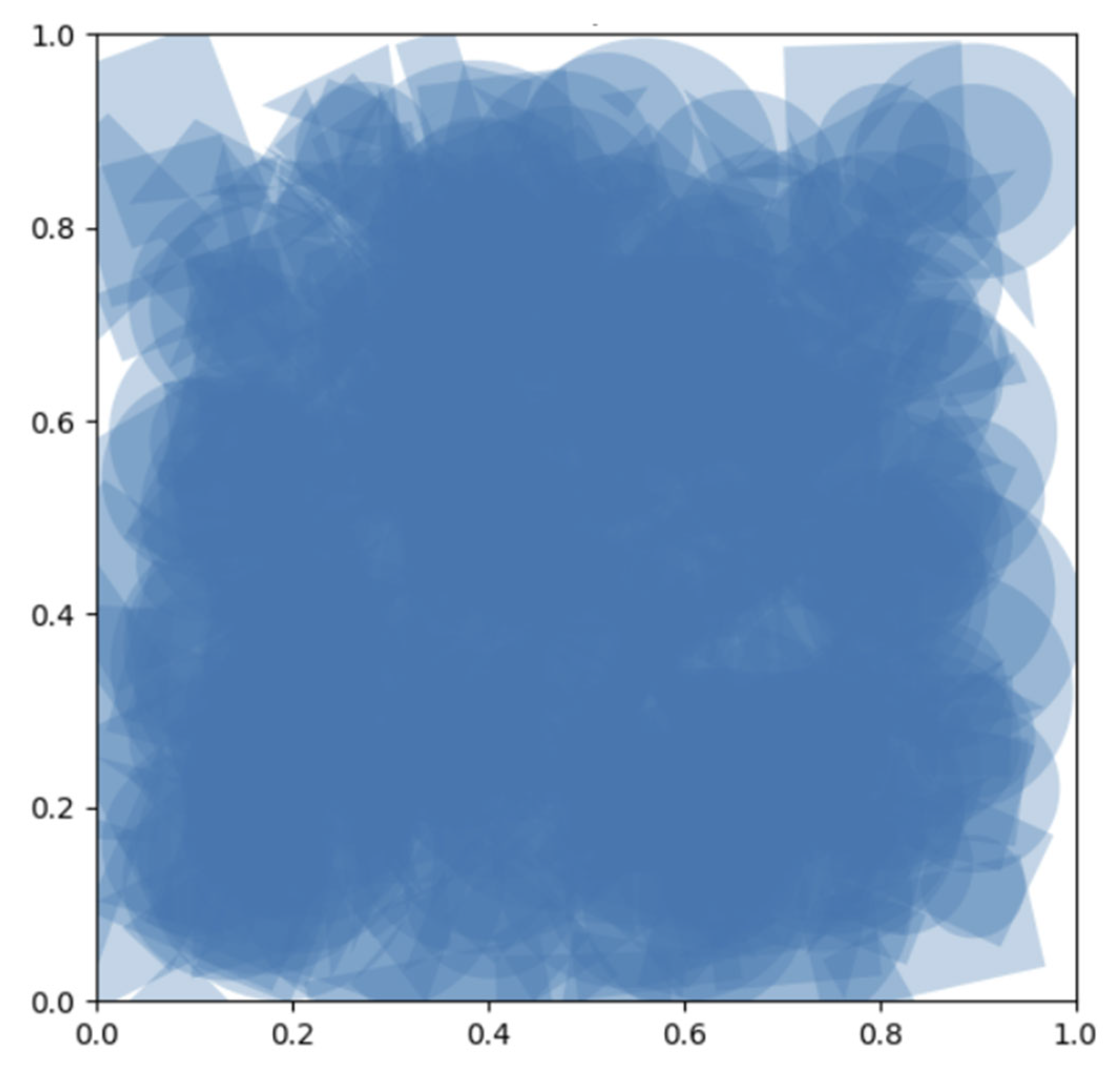
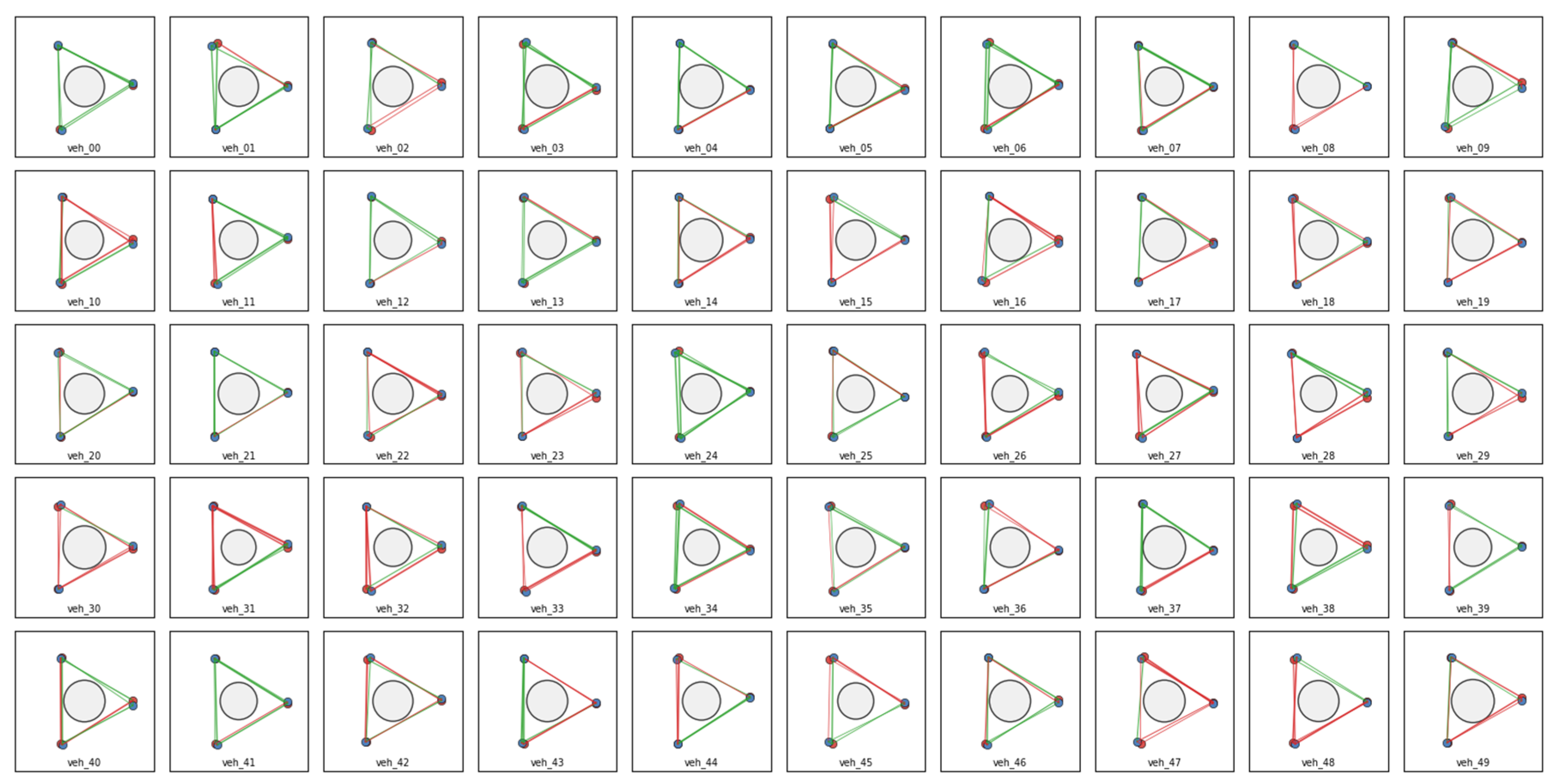
Initialization of Populations
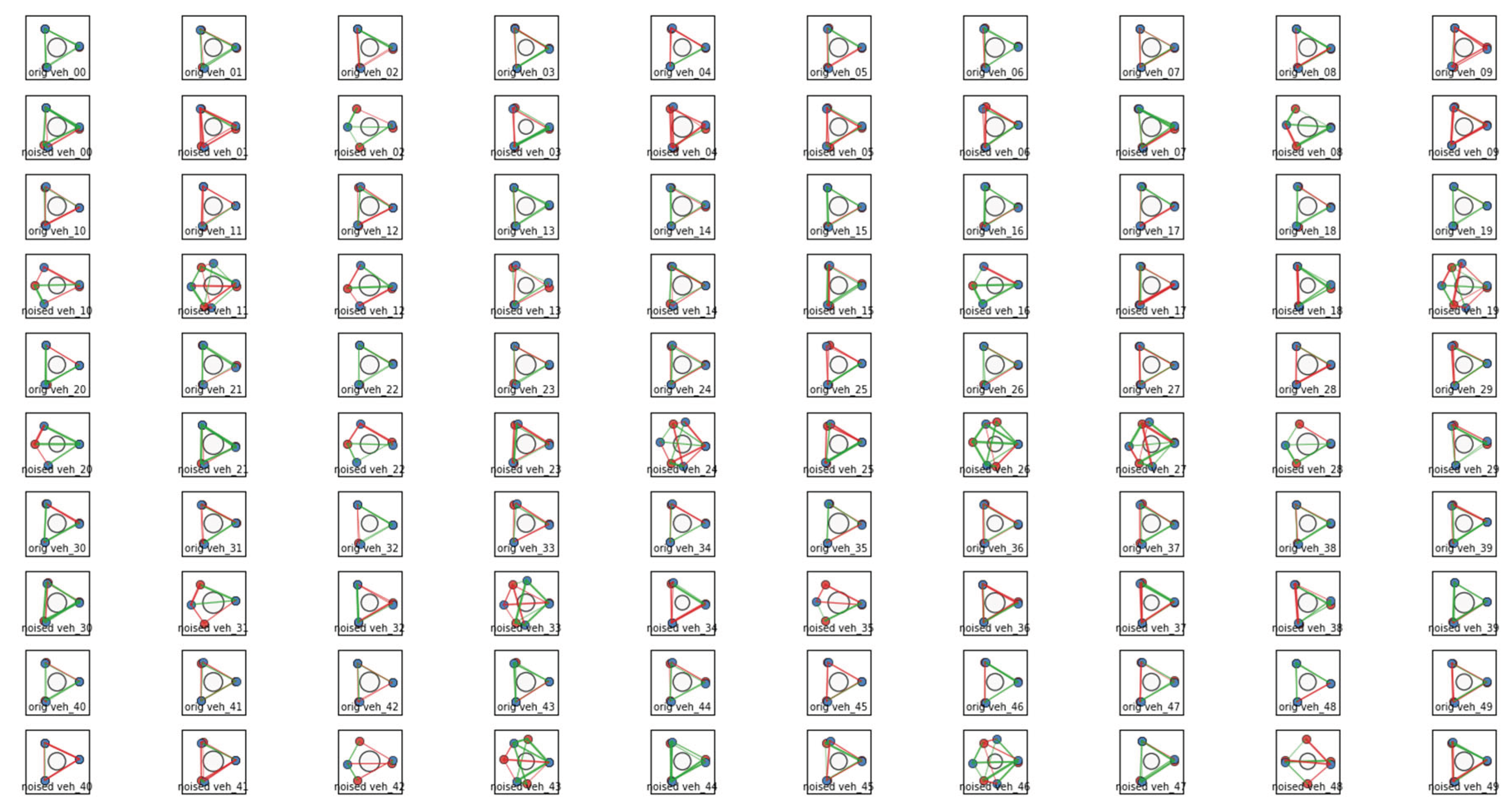
Diffusion Noise and Evolvability
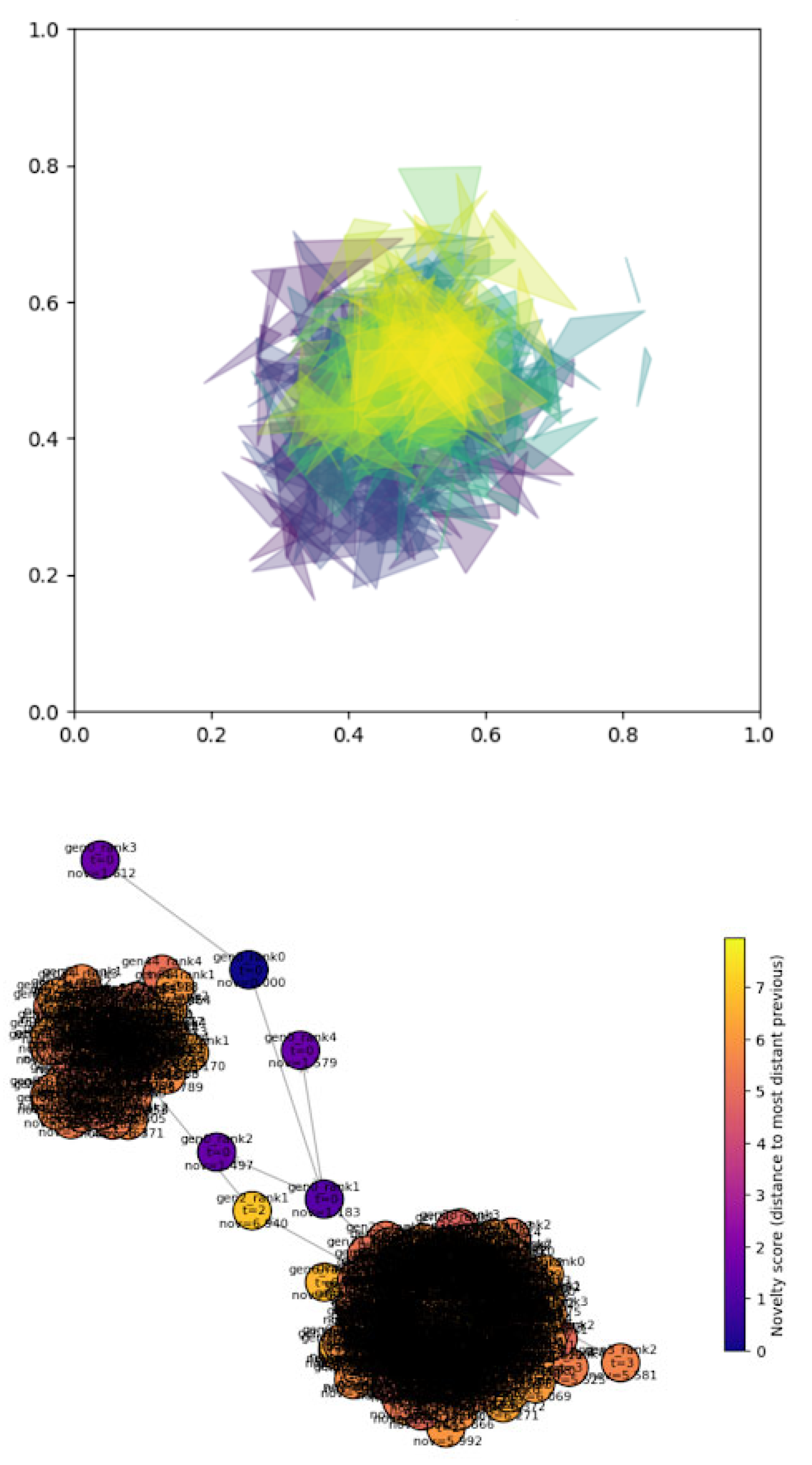
Evolution and Selection
Hypergraph Representations of Variation
Analysis with Embodied Agents
Discussion
References
- Alicea, B. Phylogenetic Models of Embodied Agents: an eco-evo-devo approach. IOP Conference Series: Materials Science and Engineering; 2026; 1343, p. 012006. [Google Scholar]
- Alicea, B.; Chakrabarty, R.; Dvoretskii, S.; Gopiiswaminathan, A.V.; Lim, A.; Parent, J. Continual Developmental Neurosimulation Using Embodied Computational Agents. IOP Conference Series: Materials Science and Engineering; 2024; 1321, p. 012013. [Google Scholar]
- Alicea, B.; Gordon, R.; Parent, J. Embodied Cognitive Morphogenesis as a Route to Intelligent Systems. Royal Society Interface Focus 2023, 13(3), 20220067. [Google Scholar] [CrossRef] [PubMed]
- Braitenberg, V. Vehicles: experiments in synthetic psychology; MIT Press; Cambridge, MA, 1984. [Google Scholar]
- Brooks, R. A robust layered control system for a mobile robot. IEEE Journal of Robotics and Automation 1986, 2(1), 14–23. [Google Scholar] [CrossRef]
- Chauhan, V.K.; Zhou, J.; Lu, P.; Molaei, S.; Clifton, D.A. A brief review of hypernetworks in deep learning. arXiv, 2024; arXiv:2306.06955. [Google Scholar]
- Cisek, P. Resynthesizing behavior through phylogenetic refinement. Attention, Perception, and Psychophysics 2009, 81(7), 2265–2287. [Google Scholar] [CrossRef] [PubMed]
- Ehlers, P.J.; Nurdin, H.L.; Soh, D. Stochastic reservoir computers. Nature Communications 2025, 16, 3070. [Google Scholar] [CrossRef] [PubMed]
- Eppe, M.; Oudeyer, P.Y. Intelligent behavior depends on the ecological niche. KI-Künstliche Intelligenz 2021, 35(1), 103–108. [Google Scholar] [CrossRef]
- Fodor, J.A. (1983). The Modularity of Mind, MIT Press, Cambridge, MA. Ghiselin, M.T. (2016). Homology, convergence and parallelism. Philosophical Transactions of the Royal Society B, 371 (1685), 20150035.
- Goodfellow, I.; Pouget-Abadie, J.; Mirza, M.; Xu, B.; Warde-Farley, D.; Ozair, S.; Courville, A.; Bengio, Y. Generative Adversarial Nets. Proceedings of Neural Information Processing Systems 2014, 27, 2672–2680. [Google Scholar]
- Lavanchy, G.; Schwander, T. Hybridogenesis. Current Biology 2019, 29(3), 3p539. [Google Scholar] [CrossRef] [PubMed]
- Levin, M. Technological Approach to Mind Everywhere: An Experimentally-Grounded Framework for Understanding Diverse Bodies and Minds. Frontiers in Systems Neuroscience 2022, 16, 768201. [Google Scholar] [CrossRef] [PubMed]
- MacLean, E.L. Unraveling the evolution of uniquely human cognition. PNAS 2016, 113(23), 6348–6354. [Google Scholar] [CrossRef] [PubMed]
- Margolis, E.; Laurence, S. Making sense of domain specificity. Cognition 2023, 240, 105583. [Google Scholar] [CrossRef] [PubMed]
- Marshall, P.J.; Houser, T.M.; Weiss, S.M. The Shared Origins of Embodiment and Development. Frontiers in Systems Neuroscience 2021, 15, 726403. [Google Scholar] [CrossRef] [PubMed]
- Miikkulainen, R. Neuroevolution insights into biological neural computation. Science 2025, 387(6735). [Google Scholar] [CrossRef] [PubMed]
- Moczek, A.P. When the end modifies its means: the origins of novelty and the evolution of innovation. Biological Journal of the Linnean Society 2023, 139, 433–440. [Google Scholar] [CrossRef]
- Nobukawa, S.; Bhattacharya, A.K.; Hirose, A. Editorial: Deep neural network architectures and reservoir computing. Frontiers in Artificial Intelligence 2025, 8, 1676744. [Google Scholar] [CrossRef] [PubMed]
- Pedersen, J.W.; Plantec, E.; Nisioti, E.; Barylli, M.; Montero, M.; Korte, K.; Risi, S. Hypernetworks That Evolve Themselves. arXiv, 2025; arXiv:2512.16406. [Google Scholar]
- Sohl-Dickstein, J.; Weiss, E.; Maheswaranathan, N.; Ganguli, S. Deep unsupervised learning using nonequilibrium thermodynamics. In Proceedings of the ICLR; 2015; pp. 2256–2265. [Google Scholar]
- Tanaka, G.; Yamane, T.; Heroux, J.B.; Nakane, R.; Kanazawa, N.; Takeda, S.; Numata, H.; Nakano, D.; Hirose, A. Recent advances in physical reservoir computing: a review. Neural Networks 2019, 115, 100–123. [Google Scholar] [CrossRef] [PubMed]
- te Vrugt, M. An introduction to reservoir computing. arXiv, 2024.
- Thomas, M.S C.; McClelland, J.L. Sun, R., Ed.; Connectionist models of cognition. In Cambridge handbook of computational psychology; Cambridge University Press; Cambridge, UK, 2008; pp. pgs. 23–58. [Google Scholar]
- Valle-Lisboa, J.C.; Pomi, A.; Mizraji, E. Multiplicative processing in the modeling of cognitive activities in large neural networks. Biophysical Reviews 2023, 15(4), 767–785. [Google Scholar] [CrossRef] [PubMed]
- van Hemmen, J.L.; Schuz, A.; Aertsen, A. Structural aspects of biological cybernetics: Valentino Braitenberg, neuroanatomy, and brain function. Biological Cybernetics 2014, 108, 517–525. [Google Scholar] [CrossRef] [PubMed]
- Verstraeten, D.; Schrauwen, B.; d’Haene, M.; Stroobandt, D. An experimental unification of reservoir computing methods. Neural Networks 2007, 20(3), 391–403. [Google Scholar] [CrossRef] [PubMed]
- Willmore, K.E. The Body Plan Concept and Its Centrality in Evo-Devo. Evolution: Education and Outreach 2012, 5, 219–230. [Google Scholar] [CrossRef]
| Reservoir Networks | Diffusion Models | |
|---|---|---|
| Primary output | High-dimensional state vectors used by a trained linear readout. | Denoised samples produced by iterative reverse diffusion. |
| Inference cost and latency | Low per step; single forward pass per timestep. | High: iterative sampling (many steps) unless using accelerated samplers. |
| Interpretability | Moderate: readout weights interpretable; reservoir dynamics opaque. | Low: deep denoisers are black boxes; intermediate noisy states are uninterpretable. |
| Robustness and stability | Good for stable temporal embeddings; sensitive to hyperparameters (spectral radius). | Sensitive to noise schedule and model capacity; sampling stability improved by recent methods. |
| Ideal Tasks | Real-time control, low compute budgets, small datasets. | High-quality generative tasks, complex data distributions, conditional synthesis. |
| Band | Description |
|---|---|
| 1 | Single sensor-effector connection |
| 2 | Simple sensor-motor couplings that perform simple behaviors (phototaxis) |
| 3 | Added selection or preference mechanisms |
| 4 | Concept-like or multi-stage behaviors |
| 5 | Chaining, rule use, or internal state dynamics |
| 6 | Foresight, planning, or complex internal models |
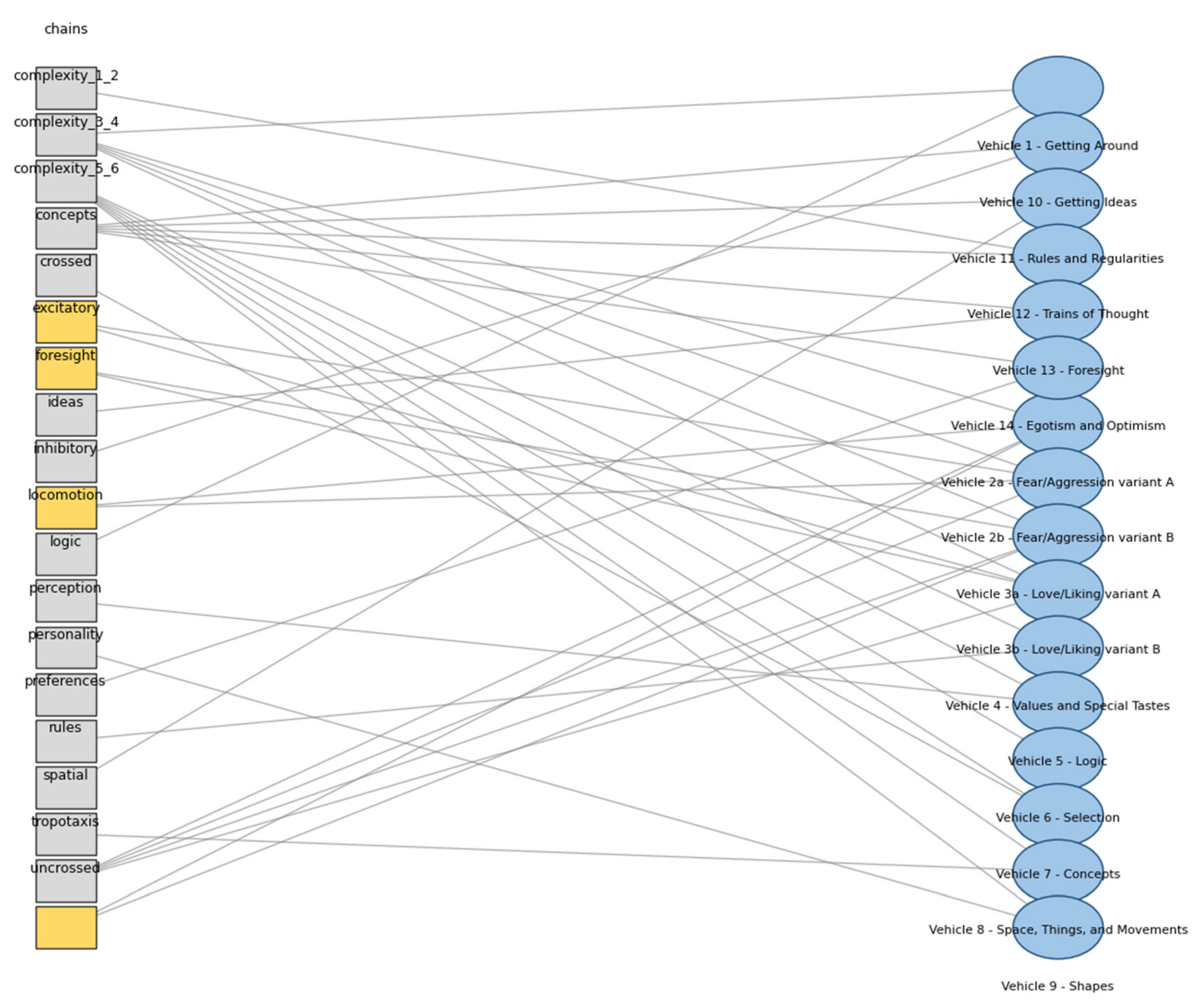
| Vehicle | Description | Theme | Wiring | Sign | Complexity |
|---|---|---|---|---|---|
| 1 | Getting Around | Locomotion | Uncrossed | Inhibitory | 2 |
| 2a | Fear/Aggression variant A | Tropotaxis | Crossed | Inhibitory | 2 |
| 2b | Fear/Aggression variant B | Tropotaxis | Uncrossed | Excitatory | 2 |
| 3a | Love/Liking variant A | Tropotaxis | Uncrossed | Excitatory | 2 |
| 3b | Love/Liking variant B | Tropotaxis | Crossed | Excitatory | 2 |
| 4 | Values and Special Tastes | Preferences | 3 | ||
| 5 | Logic | Logic | 3 | ||
| 6 | Selection | Selection | 3 | ||
| 7 | Concepts | Concepts | 4 | ||
| 8 | Space, Things, and Movements | Spatial | 4 | ||
| 9 | Shapes | Perception | 4 | ||
| 10 | Getting Ideas | Ideas | 5 | ||
| 11 | Rules and Regularities | Rules | 5 | ||
| 12 | Trains of Thought | Chains | 5 | ||
| 13 | Foresight | Foresight | 6 | ||
| 14 | Egotism and Optimism | Personality | 6 |
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2026 by the author. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license.




