Submitted:
07 April 2026
Posted:
08 April 2026
You are already at the latest version
Abstract
Keywords:
1. Introduction
2. The Roles of Microbiomes Play in CRC Formation and Progression
2.1. Metabolism
2.2. Immune
2.3. Inflammation
2.4. Proliferation and Apoptosis
2.5. Genotoxin and Mucosal Barrier
3. Contemporary Applications of Microbiological Markers in CRC Diagnosis or Prognosis
3.1. Genetic Detective Methods and Carriers
3.2. Microbial Biomarkers
4. Discussion
Institutional Review Board Statement
Informed Consent Statement
Data Availability
Conflicts of Interest
Abbreviations
References
- Siegel, R.L.; Giaquinto, A.N.; Jemal, A. Cancer statistics, 2024. Ca A Cancer J. Clin. 2024, 74, 12–49. [Google Scholar] [CrossRef]
- Santucci, C.; Mignozzi, S.; Malvezzi, M.; Boffetta, P.; Collatuzzo, G.; Levi, F.; La Vecchia, C.; Negri, E. European cancer mortality predictions for the year 2024 with focus on colorectal cancer. Ann. Oncol. 2024, 35, 308–316. [Google Scholar] [CrossRef]
- Helsingen, L.M.; Kalager, M. Colorectal Cancer Screening—Approach, Evidence, and Future Directions. NEJM Evid. 2022, 1, EVIDra2100035. [Google Scholar] [CrossRef]
- England, N. Clinical Advice for the Commissioning of the Whole Bowel Cancer Pathway. Available online: https://www.england.nhs.uk/south/wp-content/uploads/sites/6/2018/10/Clinical-Advice-for-the-Provision-of-Bowel-Cancer-Services-Jan-2018.pdf.
- Liu, Z.; Xu, Y.; Xu, G.J.; Baklaushev, V.P.; Chekhonin, V.P.; Peltzer, K.; Ma, W.J.; Wang, X.; Wang, G.W.; Zhang, C. Nomogram for predicting overall survival in colorectal cancer with distant metastasis. BMC Gastroenterol. 2021, 21, 103. [Google Scholar] [CrossRef]
- Robinson, J.R.; Newcomb, P.A.; Hardikar, S.; Cohen, S.A.; Phipps, A.I. Stage IV colorectal cancer primary site and patterns of distant metastasis. Cancer Epidemiol. 2017, 48, 92–95. [Google Scholar] [CrossRef]
- Joachim, C.; Macni, J.; Drame, M.; Pomier, A.; Escarmant, P.; Veronique-Baudin, J.; Vinh-Hung, V. Overall survival of colorectal cancer by stage at diagnosis Data from the Martinique Cancer Registry. Medicine 2019, 98, e16941. [Google Scholar] [CrossRef]
- Robertson, D.J.; Rex, D.K.; Ciani, O.; Drummond, M.F. Colonoscopy vs the Fecal Immunochemical Test: Which is Best? Gastroenterology 2024, 166, 758–771. [Google Scholar] [CrossRef]
- Castells, A.; Quintero, E.; Bujanda, L.; Castán-Cameo, S.; Cubiella, J.; Díaz-Tasende, J.; Lanas, A.; Ono, A.; Serra-Burriel, M.; Frías-Arrocha, E.; et al. Effect of invitation to colonoscopy versus faecal immunochemical test screening on colorectal cancer mortality (COLONPREV): A pragmatic, randomised, controlled, non-inferiority trial. Lancet 2025, 405, 1231–1239. [Google Scholar] [CrossRef]
- Laszlo, H.E.; Seward, E.; Ayling, R.M.; Lake, J.; Malhi, A.; Stephens, C.; Pritchard-Jones, K.; Chung, D.; Hackshaw, A.; Machesney, M. Faecal immunochemical test for patients with ‘high-risk’ bowel symptoms: A large prospective cohort study and updated literature review. Br. J. Cancer 2022, 126, 736–743. [Google Scholar] [CrossRef]
- Borrego-Ruiz, A.; Borrego, J.J. The Gut Microbiome in Human Obesity: A Comprehensive Review. Biomedicines 2025, 13, 2173. [Google Scholar] [CrossRef]
- Rinninella, E.; Tohumcu, E.; Raoul, P.; Fiorani, M.; Cintoni, M.; Mele, M.C.; Cammarota, G.; Gasbarrini, A.; Ianiro, G. The role of diet in shaping human gut microbiota. Best Pract. Res. Clin. Gastroenterol. 2023, 62–63, 101828. [Google Scholar] [CrossRef]
- Bohm, M.S.; Joseph, S.C.; Sipe, L.M.; Kim, M.; Leathem, C.T.; Mims, T.S.; Willis, N.B.; Tanveer, U.A.; Elasy, J.H.; Grey, E.W.; et al. The gut microbiome enhances breast cancer immunotherapy following bariatric surgery. JCI Insight 2025, 10, 187683. [Google Scholar] [CrossRef]
- Limeta, A.; Ji, B.Y.; Levin, M.; Gatto, F.; Nielsen, J. Meta-analysis of the gut microbiota in predicting response to cancer immunotherapy in metastatic melanoma. JCI Insight 2020, 5, 140940. [Google Scholar] [CrossRef]
- Mandal, D.P.; Mohanty, N.; Behera, P.K.; Gopinath, D.; Panda, S.; Al-Kheraif, A.A.; Divakar, D.D.; Anil, S.; Panda, S. A Plausible Proposition of CCL20-Related Mechanism in Fusobacterium nucleatum-Associated Oral Carcinogenesis. Life 2021, 11, 1218. [Google Scholar] [CrossRef]
- Pignatelli, P.; Nuccio, F.; Piattelli, A.; Curia, M.C. The Role of Fusobacterium nucleatum in Oral and Colorectal Carcinogenesis. Microorganisms 2023, 11, 2358. [Google Scholar] [CrossRef]
- Yu, L.L.; Zhao, G.; Wang, L.J.; Zhou, X.; Sun, J.; Li, X.X.; Zhu, Y.S.; He, Y.Z.; Kofonikolas, K.; Bogaert, D.; et al. A systematic review of microbial markers for risk prediction of colorectal neoplasia. Br. J. Cancer 2022, 126, 1318–1328. [Google Scholar] [CrossRef]
- Li, Z.R.; Liu, J.; Li, J.; Zhou, Z.K.; Huang, X.F.; Gopinath, D.; Luo, P.; Wang, Q.; Shan, D. Fusobacterium in the microbiome: From health to disease across the oral-gut axis and beyond. npj Biofilms Microbiomes 2025, 11, 200. [Google Scholar] [CrossRef]
- Myszkowska, J.; Derevenkov, I.; Makarov, S.V.; Spiekerkoetter, U.; Hannibal, L. Biosynthesis, Quantification and Genetic Diseases of the Smallest Signaling Thiol Metabolite: Hydrogen Sulfide. Antioxidants 2021, 10, 1065. [Google Scholar] [CrossRef]
- Birg, A.; Lin, H.C. The Role of Bacteria-Derived Hydrogen Sulfide in Multiple Axes of Disease. Int. J. Mol. Sci. 2025, 26, 3340. [Google Scholar] [CrossRef]
- Módis, K.; Coletta, C.; Erdélyi, K.; Papapetropoulos, A.; Szabo, C. Intramitochondrial hydrogen sulfide production by 3-mercaptopyruvate sulfurtransferase maintains mitochondrial electron flow and supports cellular bioenergetics. FASEB J. 2013, 27, 601–611. [Google Scholar] [CrossRef]
- Hill, B.C.; Woon, T.C.; Nicholls, P.; Peterson, J.; Greenwood, C.; Thomson, A.J. INTERACTIONS OF SULFIDE AND OTHER LIGANDS WITH CYTOCHROME-C OXIDASE—AN ELECTRON-PARAMAGNETIC-RESONANCE STUDY. Biochem. J. 1984, 224, 591–600. [Google Scholar] [CrossRef]
- Attene-Ramos, M.S.; Wagner, E.D.; Plewa, M.J.; Gaskins, H.R. Evidence that hydrogen sulfide is a genotoxic agent. Mol. Cancer Res. 2006, 4, 9–14. [Google Scholar] [CrossRef]
- Jiang, J.J.; Chan, A.; Ali, S.; Saha, A.; Haushalter, K.J.; Lam, W.L.M.; Glasheen, M.; Parker, J.; Brenner, M.; Mahon, S.B.; et al. Hydrogen Sulfide-Mechanisms of Toxicity and Development of an Antidote. Sci. Rep. 2016, 6, 20831. [Google Scholar] [CrossRef]
- Caro, A.A.; Thompson, S.; Tackett, J. Increased oxidative stress and cytotoxicity by hydrogen sulfide in HepG2 cells overexpressing cytochrome P450 2E1. Cell Biol. Toxicol. 2011, 27, 439–453. [Google Scholar] [CrossRef]
- Thompson, R.W.; Valentine, H.L.; Valentine, W.M. Cytotoxic mechanisms of hydrosulfide anion and cyanide anion in primary rat hepatocyte cultures. Toxicology 2003, 188, 149–159. [Google Scholar] [CrossRef]
- Mallardi, D.; Chimienti, G.; Maqoud, F.; Orlando, A.; Drago, S.; Malerba, E.; De Virgilio, C.; Akbarali, H.I.; Russo, F. The Dual Role of Exogenous Hydrogen Sulfide (H2S) in Intestinal Barrier Mitochondrial Function: Insights into Cytoprotection and Cytotoxicity Under Non-Stressed Conditions. Antioxidants 2025, 14, 384. [Google Scholar] [CrossRef]
- Pieper, R.; Tudela, C.V.; Taciak, M.; Bindelle, J.; Pérez, J.F.; Zentek, J. Health relevance of intestinal protein fermentation in young pigs. Anim. Health Res. Rev. 2016, 17, 137–147. [Google Scholar] [CrossRef]
- Steuber, J.; Cypionka, H.; Kroneck, P.M.H. MECHANISM OF DISSIMILATORY SULFITE REDUCTION BY DESULFOVIBRIO-DESULFURICANS—PURIFICATION OF A MEMBRANE-BOUND SULFITE REDUCTASE AND COUPLING WITH CYTOCHROME C(3) AND HYDROGENASE. Arch. Microbiol. 1994, 162, 255–260. [Google Scholar]
- Singh, S.B.; Carroll-Portillo, A.; Lin, H.C. Desulfovibrio in the Gut: The Enemy within? Microorganisms 2023, 11, 1772. [Google Scholar] [CrossRef]
- Szabo, C. Gasotransmitters in cancer: From pathophysiology to experimental therapy. Nat. Rev. Drug Discov. 2016, 15, 185–203. [Google Scholar] [CrossRef]
- Singh, S.B.; Coffman, C.N.; Varga, M.G.; Carroll-Portillo, A.; Braun, C.A.; Lin, H.C. Intestinal Alkaline Phosphatase Prevents Sulfate Reducing Bacteria-Induced Increased Tight Junction Permeability by Inhibiting Snail Pathway. Front. Cell. Infect. Microbiol. 2022, 12, 882498. [Google Scholar] [CrossRef]
- Fleishman, J.S.; Kumar, S. Bile acid metabolism and signaling in health and disease: Molecular mechanisms and therapeutic targets. Signal Transduct. Target. Ther. 2024, 9, 97. [Google Scholar] [CrossRef]
- Zhang, Y.X.; Zhou, Z.; Li, Y.S.; Xu, M.Y.; Li, Y.; Yang, Y.Q.; Hu, X.F.; Wang, X.N.; Yi, H.W.; Wu, X.D. Clostridial-dominated gut microbiota promotes 7α-dehydroxylation of obeticholic acid to lithocholic acid in cholestasis treatment. Biochem. Biophys. Res. Commun. 2025, 782, 152565. [Google Scholar] [CrossRef]
- Liu, Y.; Xu, J.; Ren, X.H.; Zhang, Y.; Ke, Z.L.; Zhou, J.H.; Wang, Y.; Zhang, Y.F.; Liu, Y.L. Cholecystectomy-induced secondary bile acids accumulation ameliorates colitis through inhibiting monocyte/macrophage recruitment. Gut Microbes 2022, 14, 2107387. [Google Scholar] [CrossRef]
- Cheng, K.R.; Raufman, J.P. Bile acid-induced proliferation of a human colon cancer cell line is mediated by transactivation of epidermal growth factor receptors. Biochem. Pharmacol. 2005, 70, 1035–1047. [Google Scholar] [CrossRef]
- Cao, H.L.; Xu, M.Q.; Dong, W.X.; Deng, B.R.; Wang, S.N.; Zhang, Y.J.; Wang, S.; Luo, S.H.; Wang, W.Q.; Qi, Y.R.; et al. Secondary bile acid-induced dysbiosis promotes intestinal carcinogenesis. Int. J. Cancer 2017, 140, 2545–2556. [Google Scholar] [CrossRef]
- Fu, T.; Coulter, S.; Yoshihara, E.; Oh, T.G.; Fang, S.; Cayabyab, F.; Zhu, Q.Y.; Zhang, T.; Leblanc, M.; Liu, S.H.; et al. FXR Regulates Intestinal Cancer Stem Cell Proliferation. Cell 2019, 176, 1098–1112.e18. [Google Scholar] [CrossRef]
- Yoshimoto, S.; Loo, T.M.; Atarashi, K.; Kanda, H.; Sato, S.; Oyadomari, S.; Iwakura, Y.; Oshima, K.; Morita, H.; Hattori, M.; et al. Obesity-induced gut microbial metabolite promotes liver cancer through senescence secretome. Nature 2013, 499, 97–101. [Google Scholar] [CrossRef]
- Wirbel, J.; Pyl, P.T.; Kartal, E.; Zych, K.; Kashani, A.; Milanese, A.; Fleck, J.S.; Voigt, A.Y.; Palleja, A.; Ponnudurai, R.; et al. Meta-analysis of fecal metagenomes reveals global microbial signatures that are specific for colorectal cancer. Nat. Med. 2019, 25, 679–689. [Google Scholar] [CrossRef]
- Yachida, S.; Mizutani, S.; Shiroma, H.; Shiba, S.; Nakajima, T.; Sakamoto, T.; Watanabe, H.; Masuda, K.; Nishimoto, Y.; Kubo, M.; et al. Metagenomic and metabolomic analyses reveal distinct stage-specific phenotypes of the gut microbiota in colorectal cancer. Nat. Med. 2019, 25, 968–976. [Google Scholar] [CrossRef]
- Ridlon, J.M.; Wolf, P.G.; Gaskins, H.R. Taurocholic acid metabolism by gut microbes and colon cancer. Gut Microbes 2016, 7, 201–215. [Google Scholar] [CrossRef]
- Shao, J.W.; Ge, T.T.; Tang, C.L.; Wang, G.; Pang, L.T.; Chen, Z. Synergistic anti-inflammatory effect of gut microbiota and lithocholic acid on liver fibrosis. Inflamm. Res. 2022, 71, 1389–1401. [Google Scholar] [CrossRef]
- Liu, L.; Dong, W.X.; Wang, S.N.; Zhang, Y.J.; Liu, T.Y.; Xie, R.X.; Wang, B.M.; Cao, H.L. Deoxycholic acid disrupts the intestinal mucosal barrier and promotes intestinal tumorigenesis. Food Funct. 2018, 9, 5588–5597. [Google Scholar] [CrossRef]
- Guo, S.H.; Peng, Y.; Lou, Y.; Cao, L.J.; Liu, J.Q.; Lin, N.M.; Cai, S.; Kang, Y.; Zeng, S.; Yu, L.S. Downregulation of the farnesoid X receptor promotes colorectal tumorigenesis by facilitating enterotoxigenic Bacteroides fragilis colonization. Pharmacol. Res. 2022, 177, 106101. [Google Scholar] [CrossRef]
- Byndloss, M.X.; Olsan, E.E.; Rivera-Chávez, F.; Tiffany, C.R.; Cevallos, S.A.; Lokken, K.L.; Torres, T.P.; Byndloss, A.J.; Faber, F.; Gao, Y.D.; et al. Microbiota-activated PPAR-γ signaling inhibits dysbiotic Enterobacteriaceae expansion. Science 2017, 357, 570–575. [Google Scholar] [CrossRef]
- Ma, H.; Yu, Y.; Wang, M.M.; Li, Z.X.; Xu, H.S.; Tian, C.; Zhang, J.; Ye, X.L.; Li, X.G. Correlation between microbes and colorectal cancer: Tumor apoptosis is induced by sitosterols through promoting gut microbiota to produce short-chain fatty acids. Apoptosis 2019, 24, 168–183. [Google Scholar] [CrossRef]
- Deleu, S.; Machiels, K.; Raes, J.; Verbeke, K.; Vermeire, S. Short chain fatty acids and its producing organisms: An overlooked therapy for IBD? eBioMedicine 2021, 66, 103293. [Google Scholar] [CrossRef]
- Louis, P.; Hold, G.L.; Flint, H.J. The gut microbiota, bacterial metabolites and colorectal cancer. Nat. Rev. Microbiol. 2014, 12, 661–672. [Google Scholar] [CrossRef]
- Lahouty, M.; Fadaee, M.; Aghaei, R.; Alizadeh, F.; Jafari, A.; Sharifi, Y. Gut microbiome and colorectal cancer: From pathogenesis to treatment. Pathol. Res. Pract. 2025, 271, 156034. [Google Scholar] [CrossRef]
- Liu, T.F.; Li, J.; Liu, Y.X.; Xiao, N.; Suo, H.T.; Xie, K.; Yang, C.L.; Wu, C. Short-Chain Fatty Acids Suppress Lipopolysaccharide-Induced Production of Nitric Oxide and Proinflammatory Cytokines Through Inhibition of NF-κB Pathway in RAW264.7 Cells. Inflammation 2012, 35, 1676–1684. [Google Scholar] [CrossRef]
- Shao, X.Y.; Sun, S.S.; Zhou, Y.Q.; Wang, H.Y.; Yu, Y.; Hu, T.; Yao, Y.Z.; Zhou, C.L. Bacteroides fragilis restricts colitis-associated cancer via negative regulation of the NLRP3 axis. Cancer Lett. 2021, 523, 170–181. [Google Scholar] [CrossRef] [PubMed]
- Ji, J.; Shu, D.M.; Zheng, M.Z.; Wang, J.; Luo, C.L.; Wang, Y.; Guo, F.Y.; Zou, X.; Lv, X.H.; Li, Y.; et al. Microbial metabolite butyrate facilitates M2 macrophage polarization and function. Sci. Rep. 2016, 6, 24838. [Google Scholar] [CrossRef]
- Coker, O.O.; Liu, C.; Wu, W.K.K.; Wong, S.H.; Jia, W.; Sung, J.J.Y.; Yu, J. Altered gut metabolites and microbiota interactions are implicated in colorectal carcinogenesis and can be non-invasive diagnostic biomarkers. Microbiome 2022, 10, 35. [Google Scholar] [CrossRef] [PubMed]
- Zhang, A.K.; Xu, Y.Z.; Xu, H.S.; Ren, J.; Meng, T.; Ni, Y.J.; Zhu, Q.W.; Zhang, W.B.; Pan, Y.B.; Jin, J.L.; et al. Lactate-induced M2 polarization of tumor-associated macrophages promotes the invasion of pituitary adenoma by secreting CCL17. Theranostics 2021, 11, 3839–3852. [Google Scholar] [CrossRef]
- Khalyfa, A.; Qiao, Z.H.; Raju, M.; Shyu, C.R.; Coghill, L.; Ericsson, A.; Gozal, D. Monocarboxylate Transporter-2 Expression Restricts Tumor Growth in a Murine Model of Lung Cancer: A Multi-Omic Analysis. Int. J. Mol. Sci. 2021, 22, 10616. [Google Scholar] [CrossRef]
- Gur, C.; Ibrahim, Y.; Isaacson, B.; Yamin, R.; Abed, J.; Gamliel, M.; Enk, J.; Bar-On, Y.; Stanietsky-Kaynan, N.; Coppenhagen-Glazer, S.; et al. Binding of the Fap2 Protein of Fusobacterium nucleatum to Human Inhibitory Receptor TIGIT Protects Tumors from Immune Cell Attack. Immunity 2015, 42, 344–355. [Google Scholar] [CrossRef]
- Abed, J.; Emgård, J.E.M.; Zamir, G.; Faroja, M.; Almogy, G.; Grenov, A.; Sol, A.; Naor, R.; Pikarsky, E.; Atlan, K.A.; et al. Fap2 Mediates Fusobacterium nucleatum Colorectal Adenocarcinoma Enrichment by Binding to Tumor-Expressed Gal-GalNAc. Cell Host Microbe 2016, 20, 215–225. [Google Scholar] [CrossRef] [PubMed]
- Rubinstein, M.R.; Wang, X.W.; Liu, W.D.; Hao, Y.J.; Cai, G.F.; Han, Y.P.W. Fusobacterium nucleatum Promotes Colorectal Carcinogenesis by Modulating E-Cadherin/β-Catenin Signaling via its FadA Adhesin. Cell Host Microbe 2013, 14, 195–206. [Google Scholar] [CrossRef]
- Dong, H.D.; Strome, S.E.; Salomao, D.R.; Tamura, H.; Hirano, F.; Flies, D.B.; Roche, P.C.; Lu, J.; Zhu, G.F.; Tamada, K.; et al. Tumor-associated B7-H1 promotes T-cell apoptosis: A potential mechanism of immune evasion. Nat. Med. 2002, 8, 793–800. [Google Scholar] [CrossRef]
- Coussens, L.M.; Pollard, J.W. Leukocytes in Mammary Development and Cancer. Cold Spring Harb. Perspect. Biol. 2011, 3, a003285. [Google Scholar] [CrossRef]
- Qian, B.Z.; Pollard, J.W. Macrophage Diversity Enhances Tumor Progression and Metastasis. Cell 2010, 141, 39–51. [Google Scholar] [CrossRef]
- Kostic, A.D.; Chun, E.Y.; Robertson, L.; Glickman, J.N.; Gallini, C.A.; Michaud, M.; Clancy, T.E.; Chung, D.C.; Lochhead, P.; Hold, G.L.; et al. Fusobacterium nucleatum Potentiates Intestinal Tumorigenesis and Modulates the Tumor-Immune Microenvironment. Cell Host Microbe 2013, 14, 207–215. [Google Scholar] [CrossRef]
- Xu, C.C.; Fan, L.N.; Lin, Y.F.; Shen, W.Y.; Qi, Y.D.; Zhang, Y.; Chen, Z.H.; Wang, L.; Long, Y.Q.; Hou, T.Y.; et al. Fusobacterium nucleatum promotes colorectal cancer metastasis through miR-1322/CCL20 axis and M2 polarization. Gut Microbes 2021, 13, 1980347. [Google Scholar] [CrossRef] [PubMed]
- Sears, C.L.; Geis, A.L.; Housseau, F. Bacteroides fragilis subverts mucosal biology: From symbiont to colon carcinogenesis. J. Clin. Investig. 2014, 124, 4166–4172. [Google Scholar] [CrossRef] [PubMed]
- Rhee, K.J.; Wu, S.G.; Wu, X.Q.; Huso, D.L.; Karim, B.; Franco, A.A.; Rabizadeh, S.; Golub, J.E.; Mathews, L.E.; Shin, J.; et al. Induction of Persistent Colitis by a Human Commensal, Enterotoxigenic Bacteroides fragilis, in Wild-Type C57BL/6 Mice. Infect. Immun. 2009, 77, 1708–1718. [Google Scholar] [CrossRef] [PubMed]
- Wu, S.G.; Rhee, K.J.; Albesiano, E.; Rabizadeh, S.; Wu, X.Q.; Yen, H.R.; Huso, D.L.; Brancati, F.L.; Wick, E.; McAllister, F.; et al. A human colonic commensal promotes colon tumorigenesis via activation of T helper type 17 T cell responses. Nat. Med. 2009, 15, 1016–1022. [Google Scholar] [CrossRef]
- Geis, A.L.; Fan, H.N.; Wu, X.Q.; Wu, S.G.; Huso, D.L.; Wolfe, J.L.; Sears, C.L.; Pardoll, D.M.; Housseau, F. Regulatory T-cell Response to Enterotoxigenic Bacteroides fragilis Colonization Triggers IL17-Dependent Colon Carcinogenesis. Cancer Discov. 2015, 5, 1098–1109. [Google Scholar] [CrossRef]
- Orberg, E.T.; Fan, H.; Tam, A.J.; Dejea, C.M.; Shields, C.E.D.; Wu, S.; Chung, L.; Finard, B.B.; Wu, X.; Fathi, P.; et al. The myeloid immune signature of enterotoxigenic Bacteroides fragilis-induced murine colon tumorigenesis. Mucosal Immunol. 2017, 10, 421–433. [Google Scholar] [CrossRef]
- Chung, L.; Orberg, E.T.; Geis, A.L.; Chan, J.L.; Fu, K.; Shields, C.E.D.; Dejea, C.M.; Fathi, P.; Chen, J.; Finard, B.B.; et al. Bacteroides fragilis Toxin Coordinates a Pro-carcinogenic Inflammatory Cascade via Targeting of Colonic Epithelial Cells. Cell Host Microbe 2018, 23, 203–214.e5. [Google Scholar] [CrossRef]
- Kikuchi, T.; Mimura, K.; Ashizawa, M.; Okayama, H.; Endo, E.; Saito, K.; Sakamoto, W.; Fujita, S.; Endo, H.; Saito, M.; et al. Characterization of tumor-infiltrating immune cells in relation to microbiota in colorectal cancers. Cancer Immunol. Immunother. 2020, 69, 23–32. [Google Scholar] [CrossRef]
- Lopès, A.; Billard, E.; Casse, A.; Villéger, R.; Veziant, J.; Roche, G.; Carrier, G.; Sauvanet, P.; Briat, A.; Pagès, F.; et al. Colibactin-positive Escherichia coli induce a procarcinogenic immune environment leading to immunotherapy resistance in colorectal cancer. Int. J. Cancer 2020, 146, 3147–3159. [Google Scholar] [CrossRef]
- Zhang, Y.C.; Weng, Y.J.; Gan, H.Y.; Zhao, X.M.; Zhi, F.C. Streptococcus gallolyticus conspires myeloid cells to promote tumorigenesis of inflammatory bowel disease. Biochem. Biophys. Res. Commun. 2018, 506, 907–911. [Google Scholar] [CrossRef]
- Furusawa, Y.; Obata, Y.; Fukuda, S.; Endo, T.A.; Nakato, G.; Takahashi, D.; Nakanishi, Y.; Uetake, C.; Kato, K.; Kato, T.; et al. Commensal microbe-derived butyrate induces the differentiation of colonic regulatory T cells. Nature 2013, 504, 446–450. [Google Scholar] [CrossRef]
- Singh, N.K.; Beckett, J.M.; Kalpurath, K.; Ishaq, M.; Ahmad, T.; Eri, R.D. Synbiotics as Supplemental Therapy for the Alleviation of Chemotherapy-Associated Symptoms in Patients with Solid Tumours. Nutrients 2023, 15, 1759. [Google Scholar] [CrossRef] [PubMed]
- Chen, S.J.; Fan, L.A.; Lin, Y.F.; Qi, Y.D.; Xu, C.C.; Ge, Q.W.; Zhang, Y.; Wang, Q.W.; Jia, D.J.C.; Wang, L.; et al. Bifidobacterium adolescentis orchestrates CD143+ cancer-associated fibroblasts to suppress colorectal tumorigenesis byWnt signaling-regulated GAS1. Cancer Commun. 2023, 43, 1027–1047. [Google Scholar] [CrossRef] [PubMed]
- Yoon, Y.; Kim, G.; Jeon, B.N.; Fang, S.; Park, H. Bifidobacterium Strain-Specific Enhances the Efficacy of Cancer Therapeutics in Tumor-Bearing Mice. Cancers 2021, 13, 957. [Google Scholar] [CrossRef] [PubMed]
- Sivaprakasam, S.; Prasad, P.D.; Singh, N. Benefits of short-chain fatty acids and their receptors in inflammation and carcinogenesis. Pharmacol. Ther. 2016, 164, 144–151. [Google Scholar] [CrossRef]
- Chang, P.V.; Hao, L.M.; Offermanns, S.; Medzhitov, R. The microbial metabolite butyrate regulates intestinal macrophage function via histone deacetylase inhibition. Proc. Natl. Acad. Sci. USA 2014, 111, 2247–2252. [Google Scholar] [CrossRef]
- Greten, F.R.; Grivennikov, S. Inflammation and Cancer: Triggers, Mechanisms, and Consequences. Immunity 2019, 51, 27–41. [Google Scholar] [CrossRef]
- Goodwin, A.C.; Shields, C.E.D.; Wu, S.G.; Huso, D.L.; Wu, X.Q.; Murray-Stewart, T.R.; Hacker-Prietz, A.; Rabizadeh, S.; Woster, P.M.; Sears, C.L.; et al. Polyamine catabolism contributes to enterotoxigenic Bacteroides fragilis-induced colon tumorigenesis. Proc. Natl. Acad. Sci. USA 2011, 108, 15354–15359. [Google Scholar] [CrossRef]
- Abdulamir, A.S.; Hafidh, R.R.; Abu Bakar, F. Molecular detection, quantification, and isolation of Streptococcus gallolyticus bacteria colonizing colorectal tumors: Inflammation-driven potential of carcinogenesis via IL-1, COX-2, and IL-8. Mol. Cancer 2010, 9, 249. [Google Scholar] [CrossRef] [PubMed]
- Miquel, S.; Martín, R.; Rossi, O.; Bermúdez-Humarán, L.G.; Chatel, J.M.; Sokol, H.; Thomas, M.; Wells, J.M.; Langella, P. Faecalibacterium prausnitzii and human intestinal health. Curr. Opin. Microbiol. 2013, 16, 255–261. [Google Scholar] [CrossRef]
- Atarashi, K.; Tanoue, T.; Shima, T.; Imaoka, A.; Kuwahara, T.; Momose, Y.; Cheng, G.H.; Yamasaki, S.; Saito, T.; Ohba, Y.; et al. Induction of Colonic Regulatory T Cells by Indigenous Clostridium Species. Science 2011, 331, 337–341. [Google Scholar] [CrossRef]
- Malik, A.; Sharma, D.; Malireddi, R.K.S.; Guy, C.S.; Chang, T.C.; Olsen, S.R.; Neale, G.; Vogel, P.; Kanneganti, T.D. SYK-CARD9 Signaling Axis Promotes Gut Fungi-Mediated Inflammasome Activation to Restrict Colitis and Colon Cancer. Immunity 2018, 49, 515–530.e5. [Google Scholar] [CrossRef]
- Alipourgivi, F.; Lu, T. Celebrating the 40-year milestone: NF-KB in oncoimmunity. Cancer Lett. 2026, 636, 218087. [Google Scholar] [CrossRef] [PubMed]
- Mao, H.M.; Zhao, X.C.; Sun, S.C. NF-κB in inflammation and cancer. Cell. Mol. Immunol. 2025, 22, 811–839. [Google Scholar] [CrossRef]
- Kim, B.Y.; Gaynor, R.B.; Song, K.; Dritschilo, A.; Jung, M. Constitutive activation of NF-κB in Ki-ras-transformed prostate epithelial cells. Oncogene 2002, 21, 4490–4497. [Google Scholar] [CrossRef]
- Zhang, Y.C.; Huo, F.C.; Wei, L.L.; Gong, C.C.; Pan, Y.J.; Mou, J.; Pei, D.S. PAK5-mediated phosphorylation and nuclear translocation of NF-κB-p65 promotes breast cancer cell proliferation in vitro and in vivo. J. Exp. Clin. Cancer Res. 2017, 36, 146. [Google Scholar] [CrossRef]
- Li, R.; Zhou, R.; Wang, H.; Li, W.D.; Pan, M.X.; Yao, X.Q.; Zhan, W.Q.; Yang, S.B.; Xu, L.J.; Ding, Y.Q.; et al. Gut microbiota-stimulated cathepsin K secretion mediates TLR4-dependent M2 macrophage polarization and promotes tumor metastasis in colorectal cancer. Cell Death Differ. 2019, 26, 2447–2463. [Google Scholar] [CrossRef]
- Sears, C.L.; Pardoll, D.M. Perspective: Alpha-Bugs, Their Microbial Partners, and the Link to Colon Cancer. J. Infect. Dis. 2011, 203, 306–311. [Google Scholar] [CrossRef] [PubMed]
- Kim, J.M.; Jung, H.Y.; Lee, J.Y.; Youn, J.; Lee, C.H.; Kim, K.H. Mitogen-activated protein kinase and activator protein-1 dependent signals are essential for Bacteroides fragilis enterotoxin-induced enteritis. Eur. J. Immunol. 2005, 35, 2648–2657. [Google Scholar] [CrossRef]
- Long, X.H.; Wong, C.C.; Tong, L.; Chu, E.S.H.; Szeto, C.H.; Go, M.Y.Y.; Coker, O.O.; Chan, A.W.H.; Chan, F.K.L.; Sung, J.J.Y.; et al. Peptostreptococcus anaerobius promotes colorectal carcinogenesis and modulates tumour immunity. Nat. Microbiol. 2019, 4, 2319–2330. [Google Scholar] [CrossRef]
- Xue, C.; Chu, Q.F.; Shi, Q.M.; Zeng, Y.F.; Lu, J.; Li, L.J. Wnt signaling pathways in biology and disease: Mechanisms and therapeutic advances. Signal Transduct. Target. Ther. 2025, 10, 106. [Google Scholar] [CrossRef]
- Pelka, K.; Hofree, M.; Chen, J.H.; Sarkizova, S.; Pirl, J.D.; Jorgji, V.; Bejnood, A.; Dionne, D.; Ge, W.H.; Xu, K.H.; et al. Spatially organized multicellular immune hubs in human colorectal cancer. Cell 2021, 184, 4734–4752.e20. [Google Scholar] [CrossRef]
- Chu, H.T.; Khosravi, A.; Kusumawardhani, I.P.; Kwon, A.H.K.; Vasconcelos, A.C.; Cunha, L.D.; Mayer, A.E.; Shen, Y.; Wu, W.L.; Kambal, A.; et al. Gene-microbiota interactions contribute to the pathogenesis of inflammatory bowel disease. Science 2016, 352, 1116–1120. [Google Scholar] [CrossRef]
- Ha, J.; Kim, D.; Yeom, J.; Kim, Y.; Yoo, S.M.; Yoon, S.H. Identification of a gene cluster for D-tagatose utilization in Escherichia coli B2 phylogroup. Iscience 2022, 25, 105655. [Google Scholar] [CrossRef] [PubMed]
- Tripathi, P.; Bruner, S.D. Structural Basis for the Interactions of the Colibactin Resistance Gene Product ClbS with DNA. Biochemistry 2021, 60, 1619–1625. [Google Scholar] [CrossRef]
- Buc, E.; Dubois, D.; Sauvanet, P.; Raisch, J.; Delmas, J.; Darfeuille-Michaud, A.; Pezet, D.; Bonnet, R. High Prevalence of Mucosa-Associated E-coli Producing Cyclomodulin and Genotoxin in Colon Cancer. PLoS ONE 2013, 8, e56964. [Google Scholar] [CrossRef] [PubMed]
- Condamine, B.; Morel-Journel, T.; Tesson, F.; Royer, G.; Magnan, M.; Bernheim, A.; Denamur, E.; Blanquart, F.; Clermont, O. Strain phylogroup and environmental constraints shape Escherichia coli dynamics and diversity over a 20-year human gut time series. ISME J. 2025, 19, wrae245. [Google Scholar] [CrossRef]
- Qiu, L.; Chirman, D.; Clark, J.R.; Xing, Y.K.; Santos, H.H.; Vaughan, E.E.; Maresso, A.W. Vaccines against extraintestinal pathogenic Escherichia coli (ExPEC): Progress and challenges. Gut Microbes 2024, 16, 2359691. [Google Scholar] [CrossRef] [PubMed]
- Cuevas-Ramos, G.; Petit, C.R.; Marcq, I.; Boury, M.; Oswald, E.; Nougayrède, J.P. Escherichia coli induces DNA damage in vivo and triggers genomic instability in mammalian cells. Proc. Natl. Acad. Sci. USA 2010, 107, 11537–11542. [Google Scholar] [CrossRef]
- Cevallos, S.A.; Lee, J.Y.; Tiffany, C.R.; Byndloss, A.J.; Johnston, L.; Byndloss, M.X.; Bäumler, A.J. Increased Epithelial Oxygenation Links Colitis to an Expansion of Tumorigenic Bacteria. mBio 2019, 10, e02244-19. [Google Scholar] [CrossRef]
- Dziubanska-Kusibab, P.J.; Berger, H.; Battistini, F.; Bouwman, B.A.M.; Iftekhar, A.; Katainen, R.; Cajuso, T.; Crosetto, N.; Orozco, M.; Aaltonen, L.A.; et al. Colibactin DNA-damage signature indicates mutational impact in colorectal cancer. Nat. Med. 2020, 26, 1063–1069. [Google Scholar] [CrossRef] [PubMed]
- Lara-Tejero, M.; Galán, J.E. A bacterial toxin that controls cell cycle progression as a deoxyribonuclease I-like protein. Science 2000, 290, 354–357. [Google Scholar] [CrossRef]
- Overstreet, A.M.C.; Burge, M.; Anderson, B.; Zhu, X.R.; Tao, Y.; Cham, C.M.; Michaud, B.; Horam, S.; Sangwan, N.; Dwidar, M.; et al. HMGB1 functions as a critical mediator of host defense at the gut mucosal barrier. Cell Host Microbe 2026, 34, 230–244.e13. [Google Scholar] [CrossRef]
- Qiao, Y.R.; He, C.E.; Xia, Y.X.; Ocansey, D.K.W.; Mao, F. Intestinal mucus barrier: A potential therapeutic target for IBD. Autoimmun. Rev. 2025, 24, 103717. [Google Scholar] [CrossRef]
- Ko, S.H.; Jeon, J.I.; Woo, H.A.; Kim, J.M. Bacteroides fragilis enterotoxin upregulates heme oxygenase-1 in dendritic cells via reactive oxygen species-, mitogen-activated protein kinase-, and Nrf2-dependent pathway. World J. Gastroenterol. 2020, 26, 291–306. [Google Scholar] [CrossRef] [PubMed]
- Kim, W.S.; Hwang, S.; Gwon, S.Y.; Jo, M.; Yoo, S.H.; Hong, J.; Jang, H.; Hong, J.E.; Kang, D.H.; Yun, M.; et al. Bacteroides fragilis Toxin Induces Sequential Proteolysis of E-Cadherin and Inflammatory Response in Mouse Intestinal Epithelial Cell Line. Microorganisms 2025, 13, 781. [Google Scholar] [CrossRef] [PubMed]
- Patterson, L.; Allen, J.; Posey, I.; Shaw, J.J.P.; Costa-Pinheiro, P.; Walker, S.J.; Gademsey, A.; Wu, X.N.; Wu, S.G.; Zachos, N.C.; et al. Glucosylceramide production maintains colon integrity in response toBacteroides fragilistoxin-induced colon epithelial cell signaling. FASEB J. 2020, 34, 15922–15945. [Google Scholar] [CrossRef]
- Prorok-Hamon, M.; Friswell, M.K.; Alswied, A.; Roberts, C.L.; Song, F.; Flanagan, P.K.; Knight, P.; Codling, C.; Marchesi, J.R.; Winstanley, C.; et al. Colonic mucosa-associated diffusely adherent afaC plus Escherichia coli expressing lpfA and pks are increased in inflammatory bowel disease and colon cancer. Gut 2014, 63, 761–770. [Google Scholar] [CrossRef]
- Liu, H.; Hong, X.L.; Sun, T.T.; Huang, X.W.; Wang, J.L.; Xiong, H. Fusobacterium nucleatumexacerbates colitis by damaging epithelial barriers and inducing aberrant inflammation. J. Dig. Dis. 2020, 21, 385–398. [Google Scholar] [CrossRef]
- Sillanpää, J.; Nallapareddy, S.R.; Singh, K.V.; Ferraro, M.J.; Murray, B.E. Adherence characteristics of endocarditis-derived Streptococcus gallolyticus ssp gallolyticus (Streptococcus bovis biotype I) isolates to host extracellular matrix proteins. Fems Microbiol. Lett. 2008, 289, 104–109. [Google Scholar] [CrossRef] [PubMed]
- Intarajak, T.; Udomchaiprasertkul, W.; Khoiri, A.N.; Sutheeworapong, S.; Kusonmano, K.; Kittichotirat, W.; Thammarongtham, C.; Cheevadhanarak, S. Distinct gut microbiomes in Thai patients with colorectal polyps. World J. Gastroenterol. 2024, 30, 3336–3355. [Google Scholar] [CrossRef] [PubMed]
- Kumar, R.; Taylor, J.C.; Jain, A.; Jung, S.Y.; Garza, V.; Xu, Y. Modulation of the extracellular matrix by Streptococcus gallolyticus subsp. gallolyticus and importance in cell proliferation. PLoS Pathog. 2022, 18, e1010894. [Google Scholar] [CrossRef] [PubMed]
- Pasquereau-Kotula, E.; Martins, M.; Aymeric, L.; Dramsi, S. Significance of Streptococcus gallolyticus subsp gallolyticus Association With Colorectal Cancer. Front. Microbiol. 2018, 9, 614. [Google Scholar] [CrossRef]
- Villéger, R.; Lopès, A.; Carrier, G.; Veziant, J.; Billard, E.; Barnich, N.; Gagnière, J.; Vazeille, E.; Bonnet, M. Intestinal Microbiota: A Novel Target to Improve Anti-Tumor Treatment? Int. J. Mol. Sci. 2019, 20, 4584. [Google Scholar] [CrossRef]
- Wang, Q.; He, M.X.; Liang, J.; Tan, X.H.; Wu, Q.M.; Wang, J.; Li, X.; Qiao, M.Q.; Huang, Z.M.; Xie, Q.; et al. Chinese guidelines for integrated diagnosis and treatment of intestinal microecology technologies in tumor application (2024 Edition). J. Cancer Res. Ther. 2024, 20, 1130–1140. [Google Scholar] [CrossRef]
- López-Aladid, R.; Fernández-Barat, L.; Alcaraz-Serrano, V.; Bueno-Freire, L.; Vázquez, N.; Pastor-Ibáñez, R.; Palomeque, A.; Oscanoa, P.; Torres, A. Determining the most accurate 16S rRNA hypervariable region for taxonomic identification from respiratory samples. Sci. Rep. 2023, 13, 3974. [Google Scholar] [CrossRef]
- Clarridge, J.E. Impact of 16S rRNA gene sequence analysis for identification of bacteria on clinical microbiology and infectious diseases. Clin. Microbiol. Rev. 2004, 17, 840–862. [Google Scholar] [CrossRef]
- Imperiale, T.F.; Gruber, R.N.; Stump, T.E.; Emmett, T.W.; Monahan, P.O. Performance Characteristics of Fecal Immunochemical Tests for Colorectal Cancer and Advanced Adenomatous Polyps A Systematic Review and Meta-analysis. Ann. Intern. Med. 2019, 170, 319–329. [Google Scholar] [CrossRef]
- Kisiel, J.B.; Limburg, P.J. Colorectal Cancer Screening With the Multitarget Stool DNA Test. Am. J. Gastroenterol. 2020, 115, 1737–1740. [Google Scholar] [CrossRef]
- Liu, Y.X.; Qin, Y.; Chen, T.; Lu, M.P.; Qian, X.B.; Guo, X.X.; Bai, Y. A practical guide to amplicon and metagenomic analysis of microbiome data. Protein Cell 2021, 12, 315–330. [Google Scholar] [CrossRef]
- Quince, C.; Walker, A.W.; Simpson, J.T.; Loman, N.J.; Segata, N. Shotgun metagenomics, from sampling to analysis. Nat. Biotechnol. 2017, 35, 833–844. [Google Scholar] [CrossRef]
- Chen, X.; Yin, X.L.; Xu, X.Q.; Zhang, T. Species-resolved profiling of antibiotic resistance genes in complex metagenomes through long-read overlapping with Argo. Nat. Commun. 2025, 16, 1744. [Google Scholar] [CrossRef]
- Feng, Q.; Liang, S.S.; Jia, H.J.; Stadlmayr, A.; Tang, L.Q.; Lan, Z.; Zhang, D.Y.; Xia, H.H.; Xu, X.Y.; Jie, Z.Y.; et al. Gut microbiome development along the colorectal adenoma-carcinoma sequence. Nat. Commun. 2015, 6, 6528. [Google Scholar] [CrossRef]
- Zeller, G.; Tap, J.; Voigt, A.Y.; Sunagawa, S.; Kultima, J.R.; Costea, P.I.; Amiot, A.; Böohm, J.; Brunetti, F.; Habermann, N.; et al. Potential of fecal microbiota for early-stage detection of colorectal cancer. Mol. Syst. Biol. 2014, 10, 766. [Google Scholar] [CrossRef]
- Pleguezuelos-Manzano, C.; Puschhof, J.; Huber, A.R.; van Hoeck, A.; Wood, H.M.; Nomburg, J.; Gurjao, C.; Manders, F.; Dalmasso, G.; Stege, P.B.; et al. Mutational signature in colorectal cancer caused by genotoxic pks+E. coli. Nature 2020, 580, 269–273. [Google Scholar] [CrossRef] [PubMed]
- Butt, J.; Blot, W.J.; Teras, L.R.; Visvanathan, K.; Le Marchand, L.; Haiman, C.A.; Chen, Y.; Bao, Y.; Sesso, H.D.; Wassertheil-Smoller, S.; et al. Antibody Responses to Streptococcus Gallolyticus Subspecies Gallolyticus Proteins in a Large Prospective Colorectal Cancer Cohort Consortium. Cancer Epidemiol. Biomark. Prev. 2018, 27, 1186–1194. [Google Scholar] [CrossRef] [PubMed]
- Wang, H.F.; Li, L.F.; Guo, S.H.; Zeng, Q.Y.; Ning, F.; Liu, W.L.; Zhang, G. Evaluation of antibody level against Fusobacterium nucleatum in the serological diagnosis of colorectal cancer. Sci. Rep. 2016, 6, 33440. [Google Scholar] [CrossRef] [PubMed]
- Flint, H.J.; Scott, K.P.; Louis, P.; Duncan, S.H. The role of the gut microbiota in nutrition and health. Nat. Rev. Gastroenterol. Hepatol. 2012, 9, 577–589. [Google Scholar] [CrossRef]
- Magne, F.; Gotteland, M.; Gauthier, L.; Zazueta, A.; Pesoa, S.; Navarrete, P.; Balamurugan, R. The Firmicutes/Bacteroidetes Ratio: A Relevant Marker of Gut Dysbiosis in Obese Patients? Nutrients 2020, 12, 1474. [Google Scholar] [CrossRef]
- Azad, M.B.; Konya, T.; Guttman, D.S.; Field, C.J.; Sears, M.R.; HayGlass, K.T.; Mandhane, P.J.; Turvey, S.E.; Subbarao, P.; Becker, A.B.; et al. Infant gut microbiota and food sensitization: Associations in the first year of life. Clin. Exp. Allergy 2015, 45, 632–643. [Google Scholar] [CrossRef]
- Xu, K.H.; Jiang, B. Analysis of Mucosa-Associated Microbiota in Colorectal Cancer. Med. Sci. Monit. 2017, 23, 4422–4430. [Google Scholar] [CrossRef] [PubMed]
- Yang, Y.Z.; Misra, B.B.; Liang, L.; Bi, D.X.; Weng, W.H.; Wu, W.; Cai, S.J.; Qin, H.L.; Goel, A.; Li, X.X.; et al. Integrated microbiome and metabolome analysis reveals a novel interplay between commensal bacteria and metabolites in colorectal cancer. Theranostics 2019, 9, 4101–4114. [Google Scholar] [CrossRef]
- de Carvalho, A.C.; Pereira, L.D.; Datorre, J.G.; dos Santos, W.; Berardinelli, G.N.; Matsushita, M.D.; Oliveira, M.A.; Duraes, R.O.; Guimaraes, D.P.; Reis, R.M. Microbiota Profile and Impact of Fusobacterium nucleatum in Colorectal Cancer Patients of Barretos Cancer Hospital. Front. Oncol. 2019, 9, 813. [Google Scholar] [CrossRef] [PubMed]
- Yu, J.; Feng, Q.; Wong, S.H.; Zhang, D.; Liang, Q.Y.; Qin, Y.W.; Tang, L.Q.; Zhao, H.; Stenvang, J.; Li, Y.L.; et al. Metagenomic analysis of faecal microbiome as a tool towards targeted non-invasive biomarkers for colorectal cancer. Gut 2017, 66, 70–78. [Google Scholar] [CrossRef]
- Guo, S.H.; Li, L.F.; Xu, B.L.; Li, M.H.; Zeng, Q.Y.; Xiao, H.; Xue, Y.; Wu, Y.X.; Wang, Y.D.; Liu, W.L.; et al. A Simple and Novel Fecal Biomarker for Colorectal Cancer: Ratio of Fusobacterium Nucleatum to Probiotics Populations, Based on Their Antagonistic Effect. Clin. Chem. 2018, 64, 1327–1337. [Google Scholar] [CrossRef]
- Wong, S.H.; Kwong, T.N.Y.; Chow, T.C.; Luk, A.K.C.; Dai, R.Z.W.; Nakatsu, G.; Lam, T.Y.T.; Zhang, L.; Wu, J.C.Y.; Chan, F.K.L.; et al. Quantitation of faecal Fusobacterium improves faecal immunochemical test in detecting advanced colorectal neoplasia. Gut 2017, 66, 1441–1448. [Google Scholar] [CrossRef]
- Zhang, X.; Zhang, Y.P.; Gui, X.R.; Zhang, Y.L.; Zhang, Z.H.; Chen, W.D.; Zhang, X.W.; Wang, Y.X.; Zhang, M.J.; Shang, Z.Q.; et al. Salivary Fusobacterium nucleatum serves as a potential biomarker for colorectal cancer. Iscience 2022, 25, 104203. [Google Scholar] [CrossRef] [PubMed]
- Wassenaar, T.M. E-coli and colorectal cancer: A complex relationship that deserves a critical mindset. Crit. Rev. Microbiol. 2018, 44, 619–632. [Google Scholar] [CrossRef]
- Eklöf, V.; Löfgren-Burström, A.; Zingmark, C.; Edin, S.; Larsson, P.; Karling, P.; Alexeyev, O.; Rutegård, J.; Wikberg, M.L.; Palmqvist, R. Cancer-associated fecal microbial markers in colorectal cancer detection. Int. J. Cancer 2017, 141, 2528–2536. [Google Scholar] [CrossRef]
- Piawah, S.; Venook, A.P. Targeted therapy for colorectal cancer metastases: A review of current methods of molecularly targeted therapy and the use of tumor biomarkers in the treatment of metastatic colorectal cancer. Cancer 2019, 125, 4139–4147. [Google Scholar] [CrossRef]
- Xie, Y.H.; Gao, Q.Y.; Cai, G.X.; Sun, X.M.; Zou, T.H.; Chen, H.M.; Yu, S.Y.; Qiu, Y.W.; Gu, W.Q.; Chen, X.Y.; et al. Fecal Clostridium symbiosum for Noninvasive Detection of Early and Advanced Colorectal Cancer: Test and Validation Studies. eBioMedicine 2017, 25, 32–40. [Google Scholar] [CrossRef] [PubMed]
- He, Z.; Yu, J.; Gong, J.L.; Wu, J.J.; Zong, X.; Luo, Z.H.; He, X.W.; Cheng, W.M.; Liu, Y.G.; Liu, C.; et al. Campylobacter jejuni-derived cytolethal distending toxin colorectal cancer metastasis. Cell Host Microbe 2024, 32, 2080–2091.e6. [Google Scholar] [CrossRef]
- Xu, S.Y.; Yin, W.; Zhang, Y.L.; Lv, Q.M.M.; Yang, Y.J.; He, J. Foes or Friends? Bacteria Enriched in the Tumor Microenvironment of Colorectal Cancer. Cancers 2020, 12, 372. [Google Scholar] [CrossRef]
- Lin, Y.F.; Lau, H.C.H.; Liu, C.F.; Ding, X.; Sun, Y.; Rong, J.M.; Zhang, X.; Wang, L.Y.; Yuan, K.; Miao, Y.L.; et al. Multi-cohort analysis reveals colorectal cancer tumor location-associated fecal microbiota and their clinical impact. Cell Host Microbe 2025, 33, 589–601. [Google Scholar] [CrossRef]
- Li, J.Y.; Pan, J.S.; Wang, L.S.; Ji, G.; Dang, Y.Q. Colorectal Cancer: Pathogenesis and Targeted Therapy. Medcomm 2025, 6, e70127. [Google Scholar] [CrossRef] [PubMed]
- Zhang, R.L.; Perekatt, A.; Chen, L. Metabolic regulation of intestinal homeostasis: Molecular and cellular mechanisms and diseases. Medcomm 2024, 5, e776. [Google Scholar] [CrossRef] [PubMed]
- Monleón, D.; Morales, J.M.; Barrasa, A.; López, J.A.; Vázquez, C.; Celda, B. Metabolite profiling of fecal water extracts from human colorectal cancer. Nmr Biomed. 2009, 22, 342–348. [Google Scholar] [CrossRef]
- Lin, Y.; Ma, C.C.; Liu, C.K.; Wang, Z.N.; Yang, J.R.; Liu, X.M.; Shen, Z.W.; Wu, R.H. NMR-based fecal metabolomics fingerprinting as predictors of earlier diagnosis in patients with colorectal cancer. Oncotarget 2016, 7, 29454–29464. [Google Scholar] [CrossRef]
- Xie, Y.L.; Chen, H.M.; Fang, J.Y. Amino acid metabolism-based molecular classification of colon adenocarcinomavia in silico analysis. Front. Immunol. 2022, 13, 1018334. [Google Scholar] [CrossRef]
- Amelio, I.; Cutruzzolá, F.; Antonov, A.; Agostini, M.; Melino, G. Serine and glycine metabolism in cancer. Trends Biochem. Sci. 2014, 39, 191–198. [Google Scholar] [CrossRef] [PubMed]
- Dadgar-Zankbar, L.; Elahi, Z.; Shariati, A.; Khaledi, A.; Razavi, S.; Khoshbayan, A. Exploring the role of Fusobacterium nucleatum in colorectal cancer: Implications for tumor proliferation and chemoresistance. Cell Commun. Signal. 2024, 22, 547. [Google Scholar] [CrossRef] [PubMed]
- Morgan, E.; Arnold, M.; Gini, A.; Lorenzoni; Cabasag, C.J.; Laversanne, M.; Vignat, J.; Ferlay, J.; Murphy, N.; Bray, F. Global burden of colorectal cancer in 2020 and 2040: Incidence and mortality estimates from GLOBOCAN. Gut 2023, 72, 338–344. [Google Scholar] [CrossRef] [PubMed]
- Tan, E.; Gouvas, N.; Nicholls, R.J.; Ziprin, P.; Xynos, E.; Tekkis, P.P. Diagnostic precision of carcinoembryonic antigen in the detection of recurrence of colorectal cancer. Surg. Oncol.-Oxf. 2009, 18, 15–24. [Google Scholar] [CrossRef]
| Substances and Toxins | Microbiomes | Effects and Mechanisms on CRC |
|---|---|---|
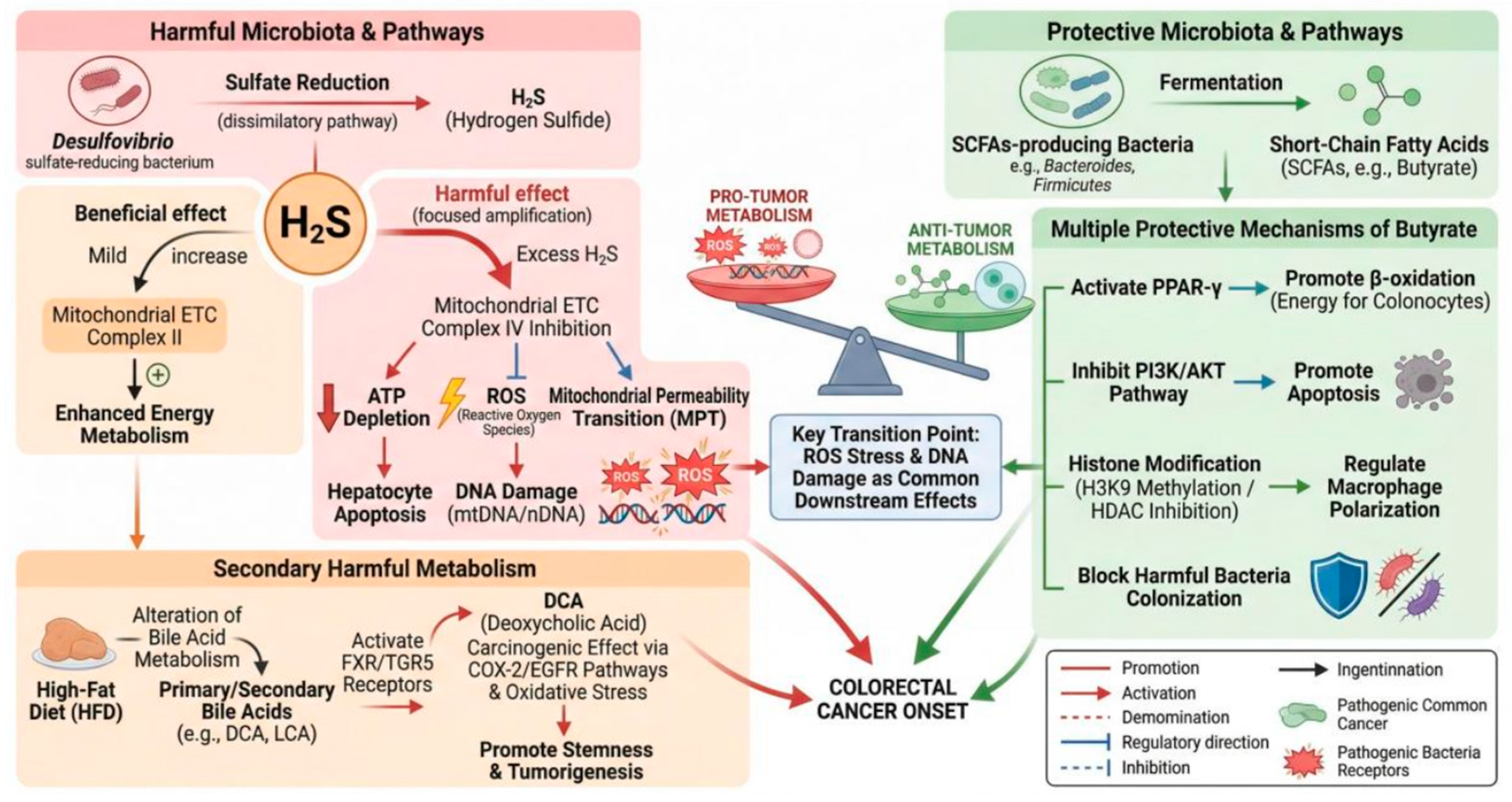
| Hydrogen sulfide | Sulfur-producing microorganisms: Bilophila wadsworthia, Desulfovibrio, Pyramidobacter etc. | 1. Genotoxicity: Direct DNA chains break and ROS elevation induced indirect damages. 2. Energy cutoff: Suppression on mitochondrial respiratory chain complex IV and butyrate application in energy supply of crypt cells. 3. Proliferation: Crypt epithelial cells proliferation ratio rise at about 50 percents level. |
| Deoxycholic acids | 7α-Decarboxylating bacteria: Clostridium scindens etc. | 1. Angiogenesis: Promotion on colonic cytomembraneous arachidonic acids excretion and COX-2 stimulation, facilitating neovascularization. 2. Propelling stem and invasiveness of CRC: FXR and TGR5 upregulation to endow stem. Deteriorating β-catenin pathway and adding aggressiveness to CRC. 3. DNA injures: ROS accumulation and DNA damages. 4. Cancer suppressor protein interruption: Proteasome activation and p53 protein degradation, selecting DNA injured cells survival to retain genetic instability. 5. Dysbiosis: Pathogen abundance augment and probiotics slash. |
| Lactate | General species | 1. Macrophages repolarization: Reshaping TAMs into M2 phenotype. 2. Lift on tumor invasiveness: Inducing TAMs to secrete CCL17 and MDSCs infiltration. |
| Amino acid metabolites (alanine, glycine etc.) | General species | 1. Nutrition supply: Supporting biosynthetic precursors and energetic sources for exuberant tumor cells. 2. Synergism: Glycine cooperating with P. anaerobius on proliferation acceleration. |
| Butyrate | Fiber-fermenting bacteria including Faecalibacterium prausnitzii, Roseburia, Clostridium butyricum, Eubacterium etc. | 1. Predominant energetic origin of colonic epithelium. 2. Epigenetic regulation: Effectuating histone hyperacetylation as a histone deacetylase inhibitor. 3. Differentiation and apoptosis expedition 4. Immune mediated inflammation management: Hastening Tregs differentiation to restrain inflammation. 5. Barrier restoration: Mucin synthetic enhancement and reconstruct damaged mucosal barrier. |
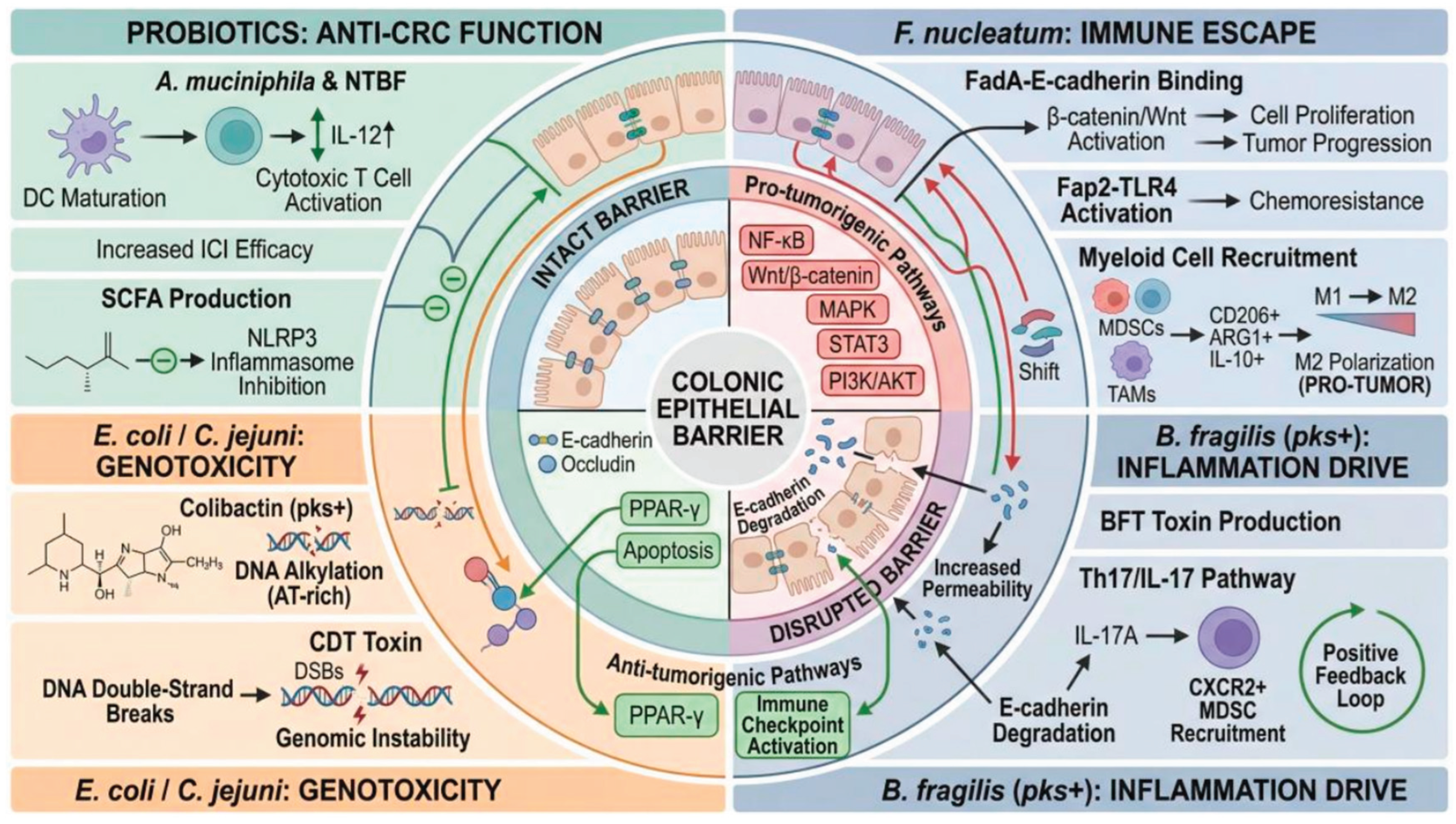
| Colibactin | pks+ E. coli | 1. Genotoxin: Direct cleavage on host DNA double chains. 2. Genetic mutation: Leading to genetic instability and chromosomal aberration and leaving mutational signature within adenine-rich regions. 3. Cell aging: Urging cell aging and following secretion of HGF to spur CRC growth. |
| BFT | ETBF | 1. Debonding epithelial barrier: Decomposition of E-cadherin, intra-cellular adhesion loss and permeability increase. 2. Proliferation: Secreting β-catenin to activate Wnt and MAPK signal pathways, raising c-Myc expression. 3. Immune suppression: Arousing IL-8 and TNF-α via NF-κB pathway, triggering Th17 and IL-17 mediated inflammation. Reinforcing immunosuppressive microenvironment by MDSCs recruitment. 4. Oxidative stress: stimulating SPO in ROS production. |
| FadA and LPS | F. nucleatum | 1. Proliferation: β-catenin amplification by binding with E-cadherin and prompting carcinogenesis. 2. Chronic inflammation: Inflammation cascade initiation from combination of LPS and T cell expressed TLR4. 3. Chemotherapeutic tolerance: 5-Fu therapeutic efficacy elimination through autophage control. |
| Signal Pathways or Targets | Microbiomes | Launcher and Procedures |
|---|---|---|
| Wnt/β-catenin signal pathway | F. Nucleatum, ETBF, S. gallolyticus, P. micra etc. | 1. F. nucleatum: Combination of FadA and E-cadherin actuates β-catenin secretion and activation. 2. ETBF: Cleavage of E-cadherin from BFT enable β-catenin entrance into nucleus. 3. S. Gallolyticus: Nuclear β-catenin level elevation. |
| NF-κB signal pathway | F. nucleatum, P. anaerobius, ETBF, E. faecalis | 1. F. nucleatum: TLR4 activation by LPS. 2. P. anaerobius: Membraneous protein PCWBR2 adhesion with integrin. 3. ETBF: BFT expels inflammation by COX-2 and PGE2 in the downstream of NF-κB, facilitating CRC immune escape. |
| PI3K/Akt pathway | P. anaerobius | 1. P. anaerobius: PCWBR2 binding with integrin α2/β1 starts PI3K/Akt pathway and promotes tumorigenesis. 2. SCFAs: Preventing PI3K/Akt pathway from excessive expression. |
| STAT3 pathway | ETBF, F. nucleatum, Prevotella copri | 1. F. nucleatum: MDSCs infiltration through STAT3 pathway. 2. ETBF and P. copri: Selectively triggering STAT3 and Th17 cells differentiation, contributing to tumorigenesis. 3. STAT3 maintains survival and proliferative status in CRC cells. |
| TIGIT | F. nucleatum | F. nucleatum escapes from immune elimination under the recognition and binding of surface protein Fap2 and TIGIT which is expressed at T cells and NK cells. |
| FXR/TGR5 | Bacteroides, Clostridium etc. | Bacteroides, Clostridium etc. transfer PCAs into SCAs. Later activation of TGR5 and FXR suppression consolidate stem and cause DNA damages. |
| Approaches | Principles | Indicators | Characteristics | Advantages | Weaknesses | Significance |
|---|---|---|---|---|---|---|
| Metagenomic Sequencing (Shotgun Sequencing) | Random dissection on all genomes within feces and indiscriminate sequencing. | Specified bacteria abundance and functional genes. | 1. High discrimination: Accurate quantification at bacterial strain level. 2. Functional analysis: Discovering genetic metabolic pathway alteration. 3. Position recognition: Distinguishing microbial proportions at different sites of colon and rectum. |
1. Distinctions at bacterial strain level. 2. Genetic information of various functional or metabolic pathways. 3. High diagnostic precision for CRC with 0.84 AUC value. 4. Exemption from tumor stage leaded effects. |
1. High volume of automatic analysis occupies huge hashrate. 2. Tissue sample is apt to lose efficacy when interrupted with host DNAs. |
1. Decent identification on adenoma and early CRC. 2. Improved sensitivity as united with FIT. |
| 16S rRNA Sequencing | Conserved and hypervariable tRNA domains amplification and sequencing dependent of Fluorescence in Situ Hybridization (FISH). | Microorganic composition and class abundance. | 1. Extensive classification: Flora diversity and community emerging. 2. Acceptable economical burden: Lower cost permits large-scale screening of 16S rRNA rather than metagenomic methods. |
Mature test technology with existing substantial databases. | 1. Distinguishing efficiency just arrive genus specificity. 2. No cellular function representation supported. |
1. CRC risk: Diversity attenuation and F. nucleatum enrichment indicates incremental CRC possibility. 2. Survival predication: Excessive colonization of B. fragilis and F. nucleatum are independent predictive factors for curtailed survival rate in CRC patients. |
| PCR-based Virulence Gene Detection | Specified amplification of function gene related to toxicity or invasiveness. | Selected virulence factors like Colibactin and BFT. | 1. Explicit target: Concentrating on pathogenic genes while not flora itself. 2. Timesaving: Standard procedure and rapid automatic process facilitate its utilization in practice. |
1. High efficacy and convenience. 2. Thorough translation into clinical medicine. |
Constricted test fields with some potentially significant and undiscovered pathogens. | 1. Early prediction: C. symbiosum and Colibactin detection serve higher sensitivity in earlier CRC development. 2. Invasiveness prejudging: Clarifying highly invasive microbiomic subtypes. 3. United test scheme: Combination together with FIT attaches improved detective rate in CRC screening stage. |
| Metabolomic Analysis | Identification and quantification of microbial products within circulation, feces and other biological samples. NMR and mass spectrum technology are common measures. | Metabolites, such as DCA, sulfuretted hydrogen and butyrate. | 1. Most direct and visualized function measurement. 2. Suitable for extracting nutritional and dietary clues on CRC. |
1. Functional regulation realization upon genomic alterations is dependent on downstream substances. Metabolomics offered closest information of this part. 2. Instantaneity: Other than subscribing clues among longer interval, metabolomics transmits instant circumstances within lower gastrointestinal tract. |
1. Timely instability: Daily dietary partterns may got a sudden change in a short time. Some temporary confounding factors cast unstable but obvious differences. 2. Material fluctuation: Some products degrade or transform in poor control conditions. |
1. Whole stage evaluation: DCA, butyrate and alanine assist in judgement of distinct stage lesion from adenoma to cancer. 2. Barrier status assessment: SCFAs test for curative effects from probiotics or dietary administration. Butyrate level reflects epithelial barrier integrity. |
| Metatranscriptomics | Probe on transcription levels and directions from intestinal or fecal samples. | RNAs | 1. Representation of virulent activity, not only existence. 2. Emphasis on more temporary effects than persistent ones. |
1. Concentrating on functions ongoing, closer than DNAs. 2. Containing more messages of recent coming adjustment than merely substances. |
RNAs high instability calls for delicate preservation and brings about indeterminacy. | Not yet systematic put into clinical practice to date. |
| Microbiomes | Indicative Fields | Mechanisms | Applications |
|---|---|---|---|
| F. nucleatum | Diagnostics, prognostics and therapeutic effect predication | 1. Surface FadA ignite β-catenin signal pathway to accelerate proliferation. 2. Inhibition on autophage pathways significantly reduce chemotherapeutic response. |
1. Adenoma and tumor diagnostic marker. 2. High abundance indicates shorten survival, cancer relapse and lymph node metastasis. 3. Reminding 5-Fu resistance. |
| pks+ E. coli | Diagnostics, classification and stages indication | Colibactin induces DNA double chains break and chromosome instability. | 1. CoPEC is an early actuator of carcinogenesis. 2. Higher concentration in TNM III/IV stages mucosal tissue than earlier periods. 3. Tending to be positive in MSS CRC. 4. Enrichment in familial adenomatous polyposis (FAP). |
| ETBF | Diagnostics, classification and prognostics | 1. Triggering STAT3 signal and lay pro-tumor inflammatory microenvironment by BFT release. 2. Cleavaging E-cadherin and initializing Wnt pathway and Th17 intermediated inflammation. |
1. ETBF enrichment is an independent risk factor of CRC overall survival. 2. Remarkable relevance to CIMP, CpG gene island mythelated phenotype. 3. Significant in early CRC screen. 4. Effective indicator of PD-1/PD-L1 immunotherapy. |
| P. anaerobius | Diagnostics and position marker | Combining membraneous PCWBR2 with integrin. Instigating PI3K-Akt signal to expedite uncontrollable proliferation. | 1. Climbing from adenoma to CRC as a potential early set biomarker. 2. Characteristic signature of rectal cancer. |
| C. symbiosum | Diagnostics | - | 1. AUC value of advanced adenoma or early stage CRC surpasses that of F. nucleatum. 2. Multistep uprising from healthy state, adenoma, early cancer to advanced CRC. |
| Faecalibacterium prausnitzii | Diagnostics and prognostics | A source of butyrates. | 1. Dramatic reduction along tumorigenesis. 2. Positively correlated with long-term cancer related and overall survival. 3. Postoperative abundance restoration prompts intestinal functional recovery. |
| A. muciniphila | Curative effect predication | Enhancing Th1 cells and renovating bowel mucosal integrity. | Ample colonization of A. muciniphila is positively relevant to treatment efficacy. |
| S. gallolyticus | Diagnostics | Upregulating IL-1, IL-8 and other inflammatory cytokines to build tumor microenviroment. | S. gallolyticus caused bacteremia declares latent CRC. |
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2026 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).




