Submitted:
04 April 2026
Posted:
08 April 2026
You are already at the latest version
Abstract
Keywords:
1. Introduction: Mechanical Stiffness as a Central Parameter in Cardiac Function
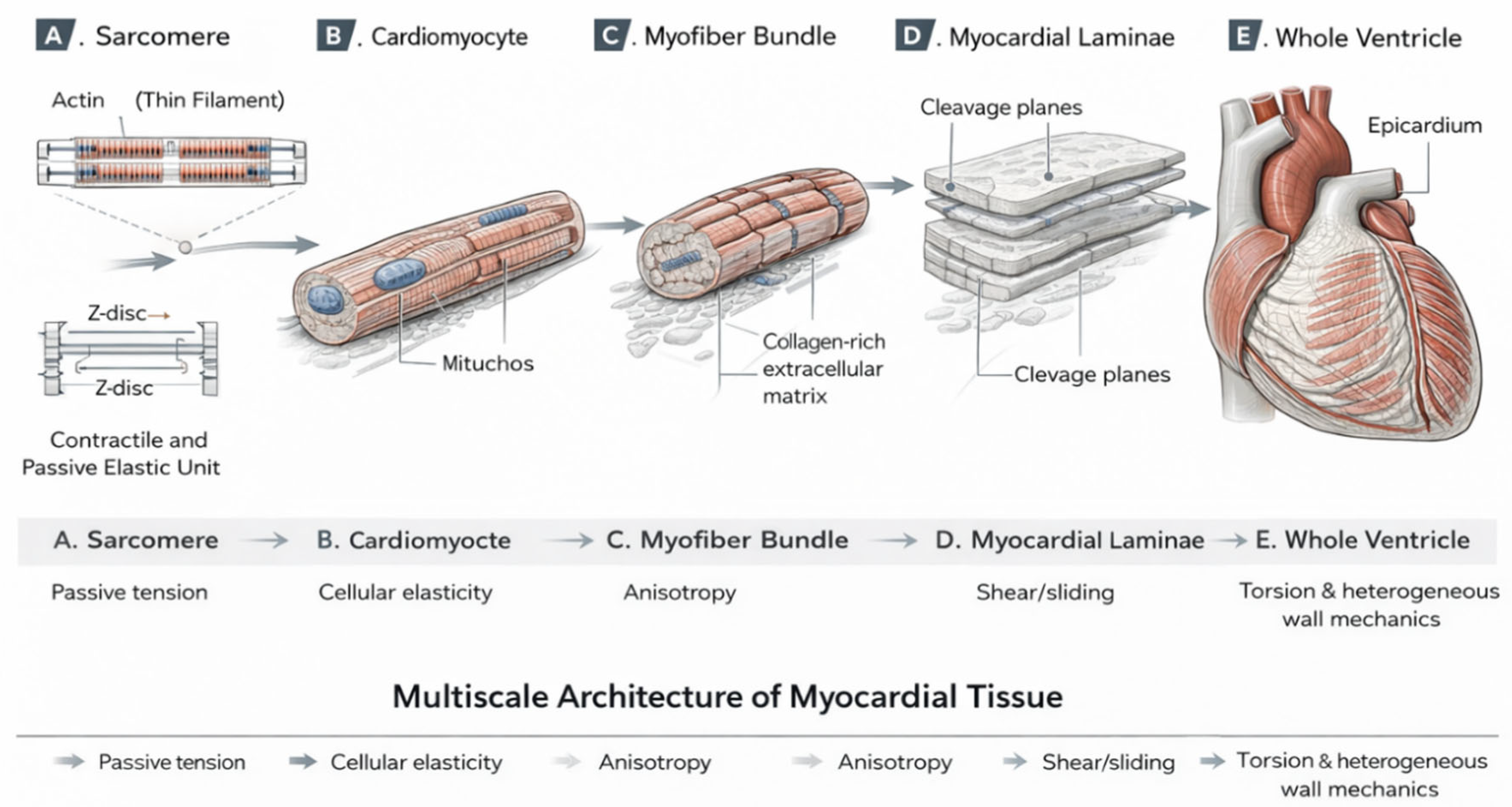
2. Structural Basis of Myocardial Mechanical Behaviour
2.1. Cardiomyocytes and Intracellular Mechanical Elements
2.2. Collagen Fibers and the Extracellular Matrix
2.3. Myocardial Sheets and Laminar Structure
2.4. Fiber Orientation Gradients Across the Ventricular Wall
2.5. Structural Anisotropy and Nonlinear Mechanical Behaviour
3. Experimental Biomechanics of Cardiac Tissue
3.1. Uniaxial Mechanical Testing
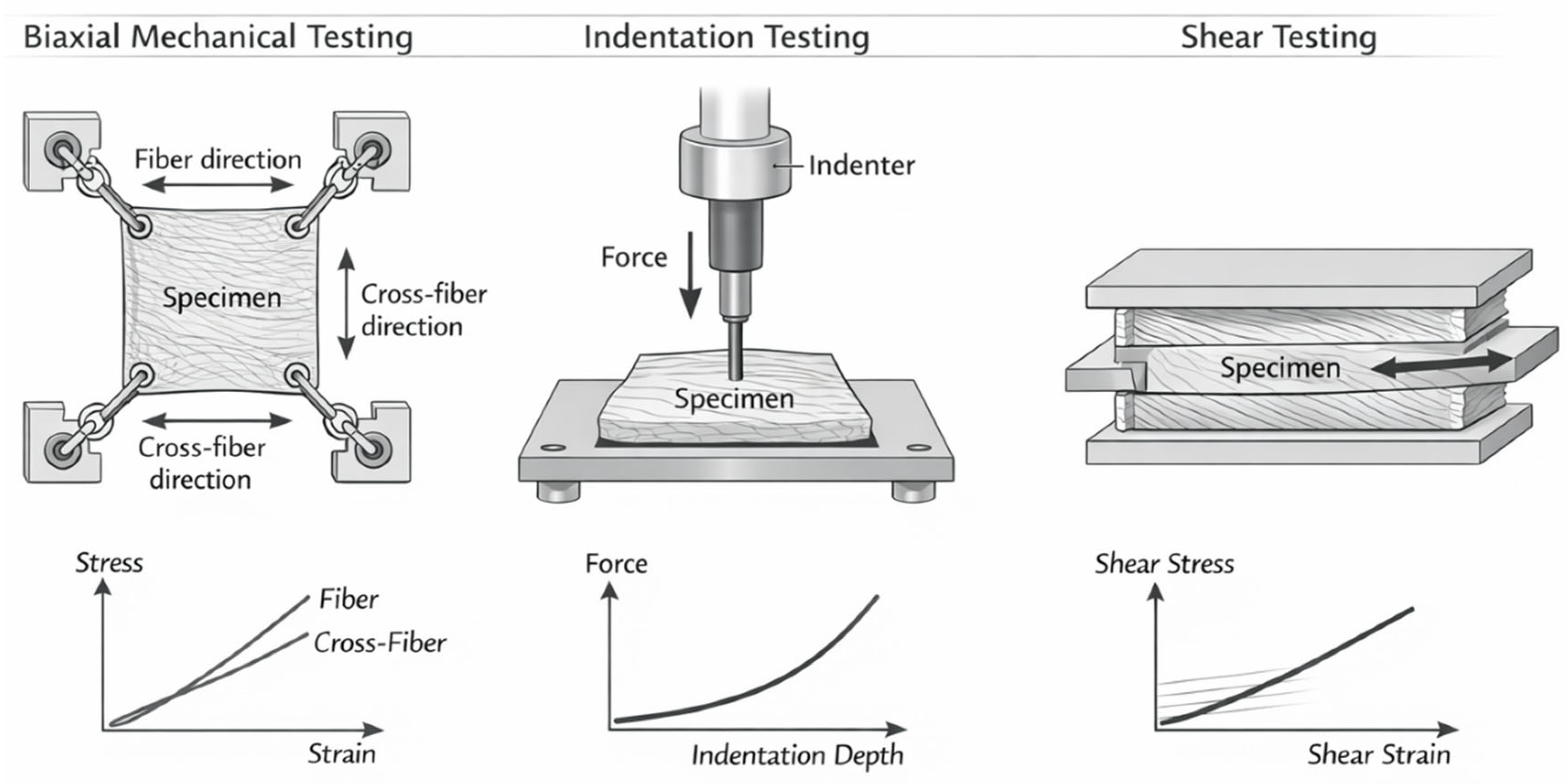
3.2. Biaxial Mechanical Testing
3.3. Shear and Torsion Testing
3.4. Indentation and Micro-Indentation Techniques
3.5. Rheology and Viscoelastic Testing
3.6. Methodological Considerations in Myocardial Mechanical Testing
4. Viscoelastic and Constitutive Modelling of Passive Myocardial Mechanics
4.1. Experimental Basis for time-Dependent Myocardial Behaviour
4.2. Structural Origins of Passive Stiffness and Viscoelasticity
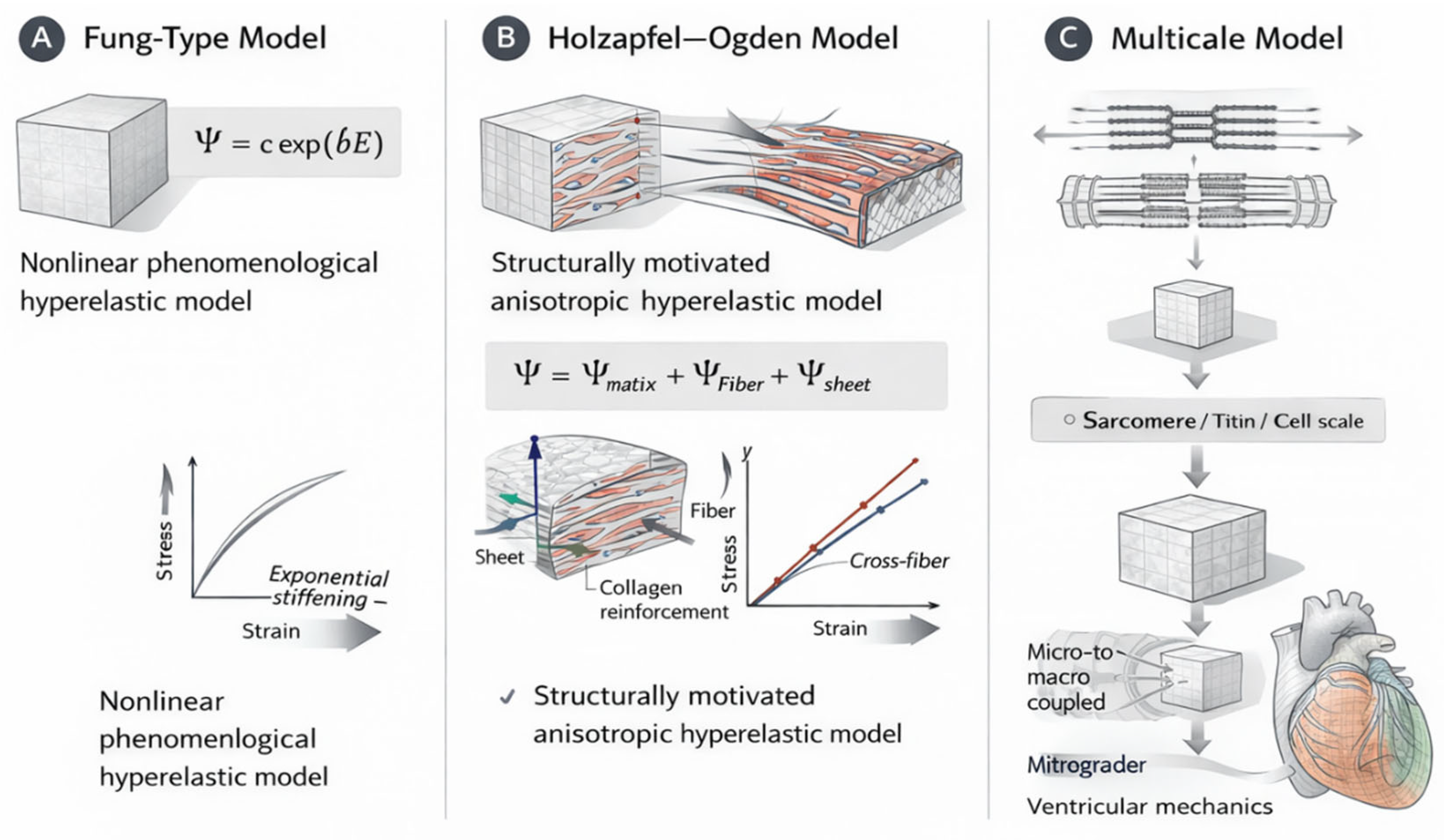
4.3. Hyperelastic Foundations and Structurally Based Constitutive Models
| Constitutive Model | Model Type | Key Features | Advantages | Limitations | Typical Applications |
| Fung-type exponential model | Phenomenological hyperelastic model | Uses exponential strain-energy function to describe nonlinear stress–strain behavior of soft tissues | Simple formulation; widely used in early cardiac biomechanics studies; captures nonlinear stiffening | Limited physiological interpretation; does not explicitly represent myocardial microstructure | Early finite element simulations of ventricular mechanics; basic tissue characterization |
| Guccione transversely isotropic model | Phenomenological anisotropic model | Incorporates preferred fiber direction with transverse isotropy | Captures anisotropic mechanical behavior associated with myocardial fibers; relatively computationally efficient | Does not explicitly include sheet structure or collagen recruitment mechanisms | Ventricular finite element simulations and patient-specific cardiac modeling |
| Holzapfel–Ogden model | Structure-based anisotropic hyperelastic model | Represents myocardium as a fiber-reinforced composite with contributions from matrix, fibers, and sheet structure | Provides physiologically meaningful representation of myocardial architecture; widely used in modern cardiac mechanics models | Requires accurate information on fiber orientation; higher computational cost | Advanced ventricular mechanics simulations; constitutive parameter identification |
| Orthotropic myocardial models | Structure-based anisotropic models | Incorporate fiber, sheet, and sheet-normal directions | More realistic representation of myocardial mechanical anisotropy | Increased number of parameters; parameter identification may be difficult | High-fidelity cardiac finite element models |
| Viscoelastic myocardial models | Time-dependent constitutive models | Incorporate stress relaxation, creep, and strain-rate dependence | Capture experimentally observed viscoelastic behavior of myocardium | Additional parameters increase model complexity | Simulation of time-dependent cardiac tissue behavior |
| Multiscale myocardial models | Multiscale constitutive framework | Integrate cellular mechanics, sarcomere dynamics, and tissue-level deformation | Mechanistically grounded; connects cellular and organ-level mechanics | Computationally intensive; requires detailed parameter calibration | Cardiac digital twins; mechanistic modeling of cardiac disease |
4.4. Viscoelastic Extensions of Myocardial Constitutive Laws
4.5. Disease Remodeling and the Need for Region- and State-Dependent Constitutive Laws
4.6. Implications for Computational Cardiac Mechanics
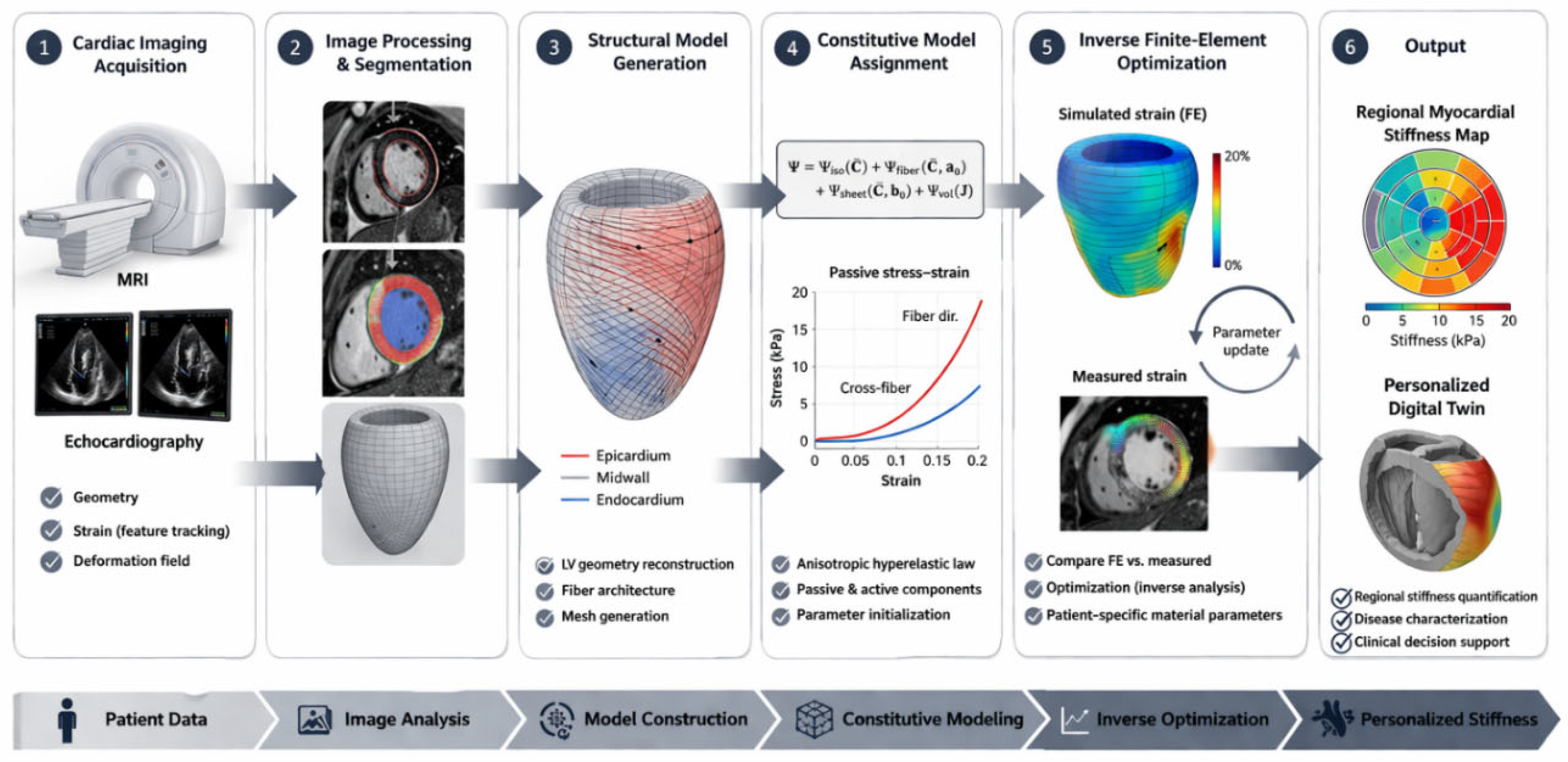
5. Image-Based and Patient-Specific Estimation of Myocardial Stiffness
5.1. MRI-Based Finite Element Inversion
5.2. Magnetic Resonance Elastography
5.3. Echocardiography-Derived Strain Analysis
5.4. Bayesian and Machine Learning Inference Methods
5.5. Toward the Cardiac Digital Twin
6. Disease-Related Remodeling and Clinical Relevance of Myocardial Stiffness
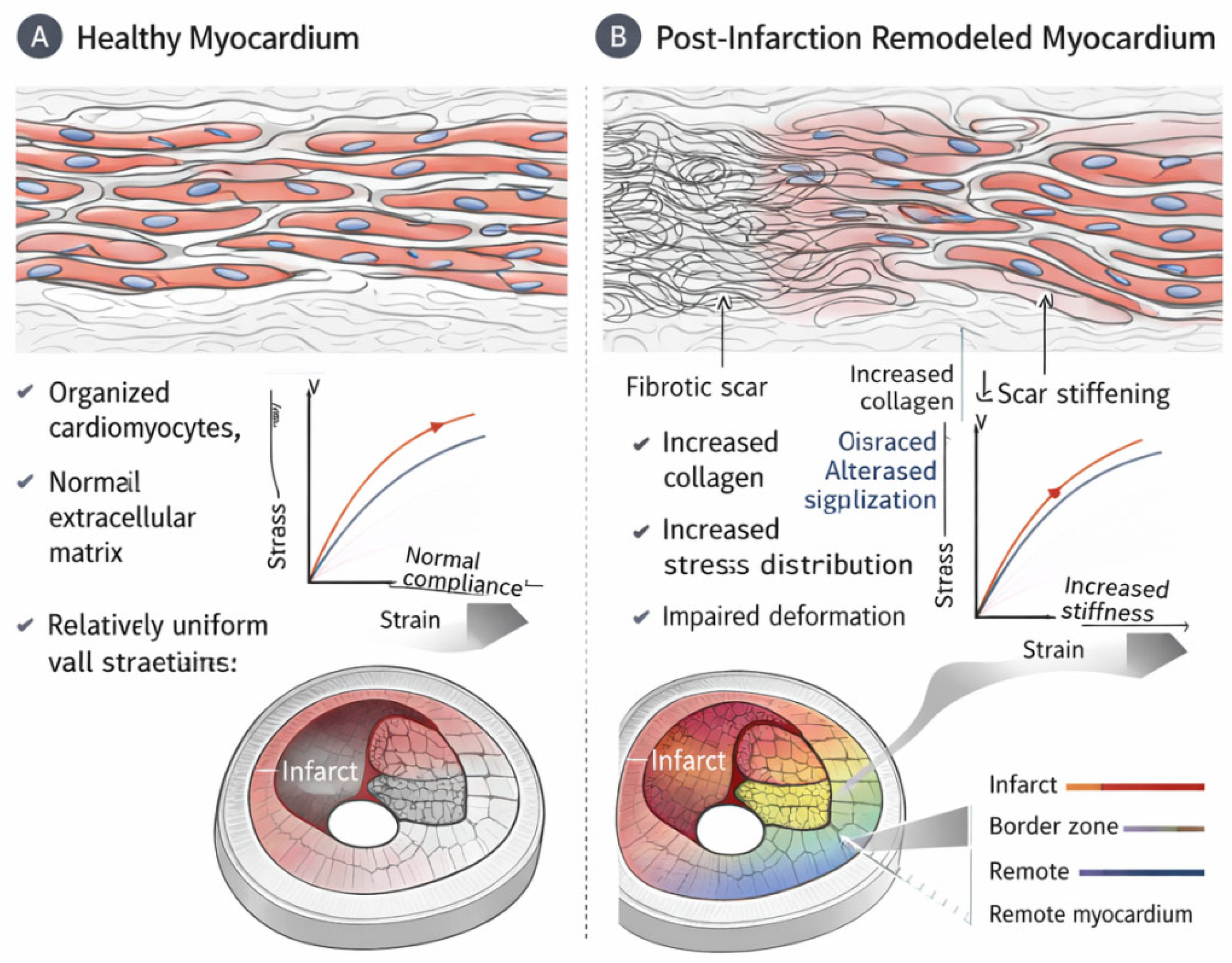
6.1. Post-Myocardial Infarction Remodelling
6.2. Fibrosis and HFpEF
6.3. Hypertrophic Cardiomyopathy as a Representative Cardiomyopathy
6.4. Translational Implication
7. Toward Standardization in Cardiac Tissue Mechanics
7.1. Anatomical Definition of Myocardial Specimens
7.2. Reporting of Myocardial Fiber Orientation
7.3. Standardization of Mechanical Testing Protocols
7.4. Parameter Identifiability in Constitutive Models
7.5. Toward Consensus Guidelines in Cardiac Biomechanics
8. Future Directions and Translational Opportunities
8.1. Personalized Cardiac Digital Twins
8.2. Non-Invasive Biomarkers of Myocardial Stiffness
8.3. Computational Design of Cardiac Patches and Implants
8.4. Integration of Biomechanics with Clinical Cardiology
8.5. Toward Mechanically Informed Cardiovascular Medicine
9. Conclusion
Ethical Approval
Consent to Participate
Consent to Publish
Data Availability Statement
Author Contributions
Funding
Competing Interests
References
- Stroik, D.; Gregorich, Z. R.; Raza, F.; Ge, Y.; Guo, W. Titin: roles in cardiac function and diseases. Frontiers in physiology 2024, vol. 15, 1385821. [Google Scholar] [CrossRef]
- Wagner, F. A. Direction-dependent contributions of cardiac myofilament networks to myocardial passive stiffness reveal a major disparity for titin. Basic Research in Cardiology 2025, vol. 120(no. 4), 761–777. [Google Scholar] [CrossRef]
- Granzier, H. L.; Labeit, S. Discovery of titin and its role in heart function and disease. Circulation Research 2025, vol. 136(no. 1), 135–157. [Google Scholar] [CrossRef]
- Janssens, J. V. The cardiomyocyte origins of diastolic dysfunction: Cellular components of myocardial stiffness. American Journal of Physiology-Heart and Circulatory Physiology 2024, vol. 326(no. 3), H584–H598. [Google Scholar] [CrossRef]
- Janssens, K.; Kraamer, M.; Barbarotta, L.; Bovendeerd, P. H. Post-infarct evolution of ventricular and myocardial function. Biomechanics and Modeling in Mechanobiology 2023, vol. 22(no. 6), 1815–1828. [Google Scholar] [CrossRef]
- Yan, D. The role of myocardial regeneration, cardiomyocyte apoptosis in acute myocardial infarction: A review of current research trends and challenges. Journal of Cardiology 2025, vol. 85(no. 4), 283–292. [Google Scholar] [CrossRef]
- Ebrahimighaei, R. Extracellular matrix stiffness controls cardiac fibroblast proliferation via the nuclear factor-Y (NF-Y) transcription factor. Biochimica et Biophysica Acta (BBA)-Molecular Cell Research 2024, vol. 1871(no. 2), 119640. [Google Scholar] [CrossRef] [PubMed]
- Kolawole, F. O. Characterizing variability in passive myocardial stiffness in healthy human left ventricles using personalized MRI and finite element modeling. Scientific Reports vol. 15(no. 1), 5556, 2025. [CrossRef] [PubMed]
- Yang, H.; Wang, H.; Wang, L.; Jiang, J. The Potential Connection Between Mechanical Stress and Heart Failure with Preserved Ejection Fraction: Mechanistic Insights and Therapeutic Potential. International Journal of Medical Sciences 2026, vol. 23(no. 2), 543. [Google Scholar] [CrossRef] [PubMed]
- Du, X.-J.; She, G.; Wu, W.; Deng, X.-L. Coupling of β-adrenergic and Hippo pathway signaling: Implications for heart failure pathophysiology and metabolic therapy. Mitochondrion 2024, vol. 78, 101941. [Google Scholar] [CrossRef] [PubMed]
- Valente, V.; Beer, B. N.; Savarese, G. Advances in the Pharmacological Treatment of Heart Failure With Preserved Ejection Fraction. International Journal of Heart Failure 2026, vol. 8(no. 1), 24–42. [Google Scholar] [CrossRef] [PubMed]
- Fayyaz, A. U. Pathophysiological insights into HFpEF from studies of human cardiac tissue. Nature Reviews Cardiology 2025, vol. 22(no. 2), 90–104. [Google Scholar] [CrossRef]
- Aradhyula, V. Novel insights into the pathobiology of pulmonary hypertension in heart failure with preserved ejection fraction. American Journal of Physiology-Heart and Circulatory Physiology 2024, vol. 326(no. 6), H1498–H1514. [Google Scholar] [CrossRef] [PubMed]
- Xu, X. Biological aging, left ventricular dysfunction and mortality in patients with heart failure with preserved ejection fraction. npj Aging 2025. [Google Scholar] [CrossRef]
- Zhou, S. The gut microbiota-inflammation-HFpEF axis: deciphering the role of gut microbiota dysregulation in the pathogenesis and management of HFpEF. Frontiers in Cellular and Infection Microbiology vol. 15, 1537576, 2025. [CrossRef]
- Pandelani, T.; Nemavhola, F. Post-MI Remodeling Mechanics: Microstructure-Informed Models, Identifiability, and Uncertainty for Patient-Specific Prediction; 2026. [Google Scholar]
- Holmes, J. W.; Borg, T. K.; Covell, J. W. Structure and mechanics of healing myocardial infarcts. Annu. Rev. Biomed. Eng. 2005, vol. 7(no. 1), 223–253. [Google Scholar] [CrossRef]
- Laita, N.; Martínez, M. Á.; Doblaré, M.; Peña, E. On the myocardium modeling under multimodal deformations: a comparison between costa’s, Holzapfel and Ogden’s formulations. Meccanica 2025, vol. 60(no. 8), 2291–2324. [Google Scholar] [CrossRef]
- Avril, S. Recent Advances in The Virtual Fields Method for Evaluating and Identifying Tissue Biomechanical Properties and Constitutive Parameters. Journal of Biomechanical Engineering 2026, pp. 1–15. [Google Scholar] [CrossRef]
- Masithulela, F. J. Computational biomechanics in the remodelling rat heart post myocardial infarction; 2016. [Google Scholar]
- Masithulela, F. Bi-ventricular finite element model of right ventricle overload in the healthy rat heart. Bio-medical materials and engineering 2016, vol. 27(no. 5), 507–525. [Google Scholar] [CrossRef]
- Masithulela, F. Analysis of passive filling with fibrotic myocardial infarction. In ASME international mechanical engineering congress and exposition; American Society of Mechanical Engineers, 2015; vol. 57380, p. V003T03A004. [Google Scholar]
- Masithulela, F. The effect of over-loaded right ventricle during passive filling in rat heart: A biventricular finite element model. ASME international mechanical engineering congress and exposition, 2015; American Society of Mechanical Engineers; vol. 57380, p. V003T03A005. [Google Scholar]
- Nemavhola, F. Detailed structural assessment of healthy interventricular septum in the presence of remodeling infarct in the free wall–A finite element model. Heliyon 2019, vol. 5(no. 6). [Google Scholar] [CrossRef] [PubMed]
- Celi, S. An image-based approach for the estimation of arterial local stiffness in vivo. Frontiers in Bioengineering and Biotechnology 2023, vol. 11, 1096196. [Google Scholar] [CrossRef] [PubMed]
- Shazly, T. Image-Based Estimation of Left Ventricular Myocardial Stiffness. Journal of Biomechanical Engineering 2025, vol. 147(no. 1), 014501. [Google Scholar] [CrossRef]
- Pandelani, T.; Semakane, L.; Msibi, M.; Kuchumov, A. G.; Nemavhola, F. Passive biaxial mechanical properties of sheep myocardium. Frontiers in Bioengineering and Biotechnology vol. 13, 1549829, 2025. [CrossRef] [PubMed]
- Ngwangwa, H. Determination of cross-directional and cross-wall variations of passive biaxial mechanical properties of rat myocardia. Processes 2022, vol. 10(no. 4), 629. [Google Scholar] [CrossRef]
- Nemavhola, F. Study of biaxial mechanical properties of the passive pig heart: material characterisation and categorisation of regional differences. International Journal of Mechanical and Materials Engineering 2021, vol. 16(no. 1), 6. [Google Scholar] [CrossRef]
- Nemavhola, F.; Ngwangwa, H.; Davies, N.; Franz, T. Passive biaxial tensile dataset of three main rat heart myocardia: left ventricle, mid-wall and right ventricle. 2021. [Google Scholar] [CrossRef]
- Nemavhola, F. Biaxial quantification of passive porcine myocardium elastic properties by region. In Engineering Solid Mechanics; 2017. [Google Scholar]
- Avazmohammadi, R.; Soares, J. S.; Li, D. S.; Raut, S. S.; Gorman, R. C.; Sacks, M. S. A contemporary look at biomechanical models of myocardium. Annual review of biomedical engineering 2019, vol. 21(no. 1), 417–442. [Google Scholar] [CrossRef]
- Ferrario, C.; Padilla, J. R.; Venet, M.; Villemain, O.; Sermesant, M. Myocardial Stiffness Quantification Using Ultrasound Shear Wave Elastography and Reduced Modeling for Subject-Specific Simulations. In in International Conference on Functional Imaging and Modeling of the Heart; Springer, 2025; pp. 41–54. [Google Scholar]
- Ge, Y.; Husmeier, D.; Rabbani, A.; Gao, H. Advanced statistical inference of myocardial stiffness: A time series Gaussian process approach of emulating cardiac mechanics for real-time clinical decision support. Computers in Biology and Medicine 2025, vol. 184, 109381. [Google Scholar] [CrossRef]
- Shi, Q.; Xu, M.; Zhu, C.; Shi, G. Feel the force: Biomechanical homeostasis of the cardiovascular system. Journal of Translational Internal Medicine 2026, vol. 14(no. 1), 6–9. [Google Scholar] [CrossRef]
- Mendiola, E. A.; Mehdi, R. R.; Shah, D. J.; Avazmohammadi, R. On in-silico estimation of left ventricular end-diastolic pressure from cardiac strains. 2024 46th Annual International Conference of the IEEE Engineering in Medicine and Biology Society (EMBC) 2024, 1–4. [Google Scholar]
- Mu, S.; Chan, W. X.; Yap, C. H. A Cycle-Consistent Graph Surrogate for Full-Cycle Left Ventricular Myocardial Biomechanics. arXiv 2026, arXiv:2602.06884. [Google Scholar]
- Nemavhola, F. Fibrotic infarction on the LV free wall may alter the mechanics of healthy septal wall during passive filling. Bio-medical materials and engineering 2017, vol. 28(no. 6), 579–599. [Google Scholar] [CrossRef]
- Nordsletten, D.; Niederer, S.; Nash, M.; Hunter, P.; Smith, N. Coupling multi-physics models to cardiac mechanics. Progress in biophysics and molecular biology 2011, vol. 104(no. 1-3), 77–88. [Google Scholar] [CrossRef]
- Holzapfel, G. A.; Ogden, R. W. Constitutive modelling of passive myocardium: a structurally based framework for material characterization. Philosophical Transactions of the Royal Society A: Mathematical, Physical and Engineering Sciences 2009, vol. 367(no. 1902), 3445–3475. [Google Scholar] [CrossRef]
- Bracamonte, J. H.; Saunders, S. K.; Wilson, J. S.; Truong, U. T.; Soares, J. S. Patient-specific inverse modeling of in vivo cardiovascular mechanics with medical image-derived kinematics as input data: concepts, methods, and applications. Applied Sciences 2022, vol. 12(no. 8), 3954. [Google Scholar] [CrossRef]
- Ramadan, A. Selecting sensitive parameter subsets in dynamical models with application to biomechanical system identification. Journal of biomechanical engineering 2018, vol. 140(no. 7), 074503. [Google Scholar] [CrossRef]
- Linke, W. A.; Krüger, M. The giant protein titin as an integrator of myocyte signaling pathways. Physiology 2010, vol. 25(no. 3), 186–198. [Google Scholar] [CrossRef]
- Crocini, C.; Gotthardt, M. Cardiac sarcomere mechanics in health and disease. Biophysical reviews 2021, vol. 13(no. 5), 637–652. [Google Scholar] [CrossRef] [PubMed]
- Campbell, K. S. Impact of myocyte strain on cardiac myofilament activation. Pflügers Archiv-European Journal of Physiology 2011, vol. 462(no. 1), 3–14. [Google Scholar] [CrossRef] [PubMed]
- Granzier, H. L.; Irving, T. C. Passive tension in cardiac muscle: contribution of collagen, titin, microtubules, and intermediate filaments. Biophysical journal 1995, vol. 68(no. 3), 1027–1044. [Google Scholar] [CrossRef] [PubMed]
- Nagueh, S. F. Altered titin expression, myocardial stiffness, and left ventricular function in patients with dilated cardiomyopathy. Circulation 2004, vol. 110(no. 2), 155–162. [Google Scholar] [CrossRef]
- LeWinter, M. M.; Granzier, H. L. Cardiac titin and heart disease. Journal of cardiovascular pharmacology 2014, vol. 63(no. 3), 207–212. [Google Scholar] [CrossRef]
- Vahle, B. MyoMed205 Counteracts Titin Hyperphosphorylation and the Expression of Contraction-Regulating Proteins in a Rat Model of HFpEF. Journal of Cachexia, Sarcopenia and Muscle vol. 16(no. 3), e13843, 2025. [CrossRef] [PubMed]
- Hamdani, N.; Herwig, M.; Linke, W. A. Tampering with springs: phosphorylation of titin affecting the mechanical function of cardiomyocytes. Biophysical reviews 2017, vol. 9(no. 3), 225–237. [Google Scholar] [CrossRef] [PubMed]
- LeWinter, M. M.; Wu, Y.; Labeit, S.; Granzier, H. Cardiac titin: structure, functions and role in disease. Clinica chimica acta 2007, vol. 375(no. 1-2), 1–9. [Google Scholar] [CrossRef]
- Kleber, A. G.; Saffitz, J. E. Role of the intercalated disc in cardiac propagation and arrhythmogenesis. Frontiers in physiology 2014, vol. 5, 404. [Google Scholar] [CrossRef] [PubMed]
- Lindsey, M. L.; Borg, T. K. Understanding the role of the extracellular matrix in cardiovascular development and disease: where do we go from here? Journal of molecular and cellular cardiology 2009, vol. 48(no. 3), 431. [Google Scholar] [CrossRef]
- Burlew, B. S.; Weber, K. T. Cardiac fibrosis as a cause of diastolic dysfunction. Herz 2002, vol. 27(no. 2), 92–98. [Google Scholar] [CrossRef]
- Fan, D.; Takawale, A.; Lee, J.; Kassiri, Z. Cardiac fibroblasts, fibrosis and extracellular matrix remodeling in heart disease. Fibrogenesis & tissue repair 2012, vol. 5(no. 1), 15. [Google Scholar]
- Robinson, T. F.; Cohen-Gould, L.; Factor, S. M.; Eghbali, M.; Blumenfeld, O. O. Structure and function of connective tissue in cardiac muscle: collagen types I and III in endomysial struts and pericellular fibers. Scanning Microscopy 1988, vol. 2(no. 2), 33. [Google Scholar]
- Holzapfel, G. A.; Gasser, T. C.; Ogden, R. W. A new constitutive framework for arterial wall mechanics and a comparative study of material models. Journal of elasticity and the physical science of solids 2000, vol. 61(no. 1), 1–48. [Google Scholar]
- Fomovsky, G. M.; Thomopoulos, S.; Holmes, J. W. Contribution of extracellular matrix to the mechanical properties of the heart. Journal of molecular and cellular cardiology 2010, vol. 48(no. 3), 490–496. [Google Scholar] [CrossRef]
- Fomovsky, G. M.; Rouillard, A. D.; Holmes, J. W. Regional mechanics determine collagen fiber structure in healing myocardial infarcts. Journal of molecular and cellular cardiology 2012, vol. 52(no. 5), 1083–1090. [Google Scholar] [CrossRef]
- Sommer, G.; Gasser, T. C.; Regitnig, P.; Auer, M.; Holzapfel, G. A. Dissection properties of the human aortic media: an experimental study. Journal of biomechanical engineering 2008, vol. 130(no. 2). [Google Scholar] [CrossRef]
- Gizzi, A.; Pandolfi, A. Visco-hyperelasticity of electro-active soft tissues. Procedia IUTAM 2015, vol. 12, 162–175. [Google Scholar] [CrossRef]
- Niederer, S. A.; Lumens, J.; Trayanova, N. A. Computational models in cardiology. Nature reviews cardiology 2019, vol. 16(no. 2), 100–111. [Google Scholar] [CrossRef] [PubMed]
- Frangogiannis, N. G. The inflammatory response in myocardial injury, repair, and remodelling. Nature Reviews Cardiology 2014, vol. 11(no. 5), 255–265. [Google Scholar] [CrossRef] [PubMed]
- Talman, V.; Ruskoaho, H. Cardiac fibrosis in myocardial infarction—from repair and remodeling to regeneration. Cell and tissue research 2016, vol. 365(no. 3), 563–581. [Google Scholar] [CrossRef]
- Kong, P.; Christia, P.; Frangogiannis, N. G. The pathogenesis of cardiac fibrosis. Cellular and molecular life sciences 2014, vol. 71(no. 4), 549–574. [Google Scholar] [CrossRef]
- LeGrice, J.; Hunter, P. J.; Smaill, B. Laminar structure of the heart: a mathematical model. American Journal of Physiology-Heart and Circulatory Physiology 1997, vol. 272(no. 5), H2466–H2476. [Google Scholar] [CrossRef]
- Costa, D.; Takayama, Y.; McCulloch, A. D.; Covell, J. W. Laminar fiber architecture and three-dimensional systolic mechanics in canine ventricular myocardium. American Journal of Physiology-Heart and Circulatory Physiology 1999, vol. 276(no. 2), H595–H607. [Google Scholar] [CrossRef] [PubMed]
- Costa, D.; Holmes, J. W.; McCulloch, A. D. Modelling cardiac mechanical properties in three dimensions. Philosophical transactions of the Royal Society of London. Series A: Mathematical, physical and engineering sciences 2001, vol. 359(no. 1783), 1233–1250. [Google Scholar] [CrossRef]
- Takayama, Y.; Costa, K. D.; Covell, J. W. Contribution of laminar myofiber architecture to load-dependent changes in mechanics of LV myocardium. American Journal of Physiology-Heart and Circulatory Physiology 2002, vol. 282(no. 4), H1510–H1520. [Google Scholar] [CrossRef]
- Laita, N. On modeling the in vivo ventricular passive mechanical behavior from in vitro experimental properties in porcine hearts. Computers & Structures vol. 292, 107241, 2024.
- Tous, C. Distinguishing shear and tensile myocardial wall stiffness using ex vivo anisotropic Magnetic Resonance Elastography. Acta biomaterialia 2025, vol. 202, 276–291. [Google Scholar] [CrossRef] [PubMed]
- Gültekin, O.; Dal, H. “Computational finite strain orthotropic viscoelasticity of human passive myocardium,” in Solid (Bio) mechanics: Challenges of the Next Decade: A Book Dedicated to Professor Gerhard A; Springer: Holzapfel, 2022; pp. 257–278. [Google Scholar]
- Nash, P.; Hunter, P. J. Computational mechanics of the heart. Journal of elasticity and the physical science of solids 2000, vol. 61(no. 1), 113–141. [Google Scholar]
- Nachlas, L. A multilayered valve leaflet promotes cell-laden collagen type I production and aortic valve hemodynamics. Biomaterials 2020, vol. 240, 119838. [Google Scholar] [CrossRef] [PubMed]
- Helm, P.; Beg, M. F.; Miller, M. I.; Winslow, R. L. Measuring and mapping cardiac fiber and laminar architecture using diffusion tensor MR imaging. Annals of the New York Academy of Sciences 2005, vol. 1047(no. 1), 296–307. [Google Scholar] [CrossRef]
- Gilbert, S. H.; Bernus, O.; Holden, A. V.; Benson, A. P. A quantitative comparison of the myocardial fibre orientation in the rabbit as determined by histology and by diffusion tensor-MRI. in International Conference on Functional Imaging and Modeling of the Heart, 2009; Springer; pp. 49–57. [Google Scholar]
- Buckberg, G.; Hoffman, J. I.; Nanda, N. C.; Coghlan, C.; Saleh, S.; Athanasuleas, C. Ventricular torsion and untwisting: further insights into mechanics and timing interdependence: a viewpoint. Echocardiography 2011, vol. 28(no. 7), 782–804. [Google Scholar] [CrossRef]
- Young, A. A.; Axel, L. Three-dimensional motion and deformation of the heart wall: estimation with spatial modulation of magnetization--a model-based approach. Radiology 1992, vol. 185(no. 1), 241–247. [Google Scholar] [CrossRef]
- Young, A.; Kramer, C. M.; Ferrari, V. A.; Axel, L.; Reichek, N. Three-dimensional left ventricular deformation in hypertrophic cardiomyopathy. Circulation 1994, vol. 90(no. 2), 854–867. [Google Scholar] [CrossRef]
- Scollan, D.; Holmes, A.; Zhang, J.; Winslow, R. Reconstruction of cardiac ventricular geometry and fiber orientation using magnetic resonance imaging. Annals of biomedical engineering 2000, vol. 28(no. 8), 934–944. [Google Scholar] [CrossRef]
- Toussaint; Stoeck, C. T.; Schaeffter, T.; Sermesant, M.; Kozerke, S. Cardiac Laminae Structure Dynamics from In-vivo Diffusion Tensor Imaging. in 21st Annual ISMRM Scientific Meeting and Exhibition 2013: Discovery, Innovation & Application–Advancing MR for Improved Health, Salt Lake City, Utah, USA, 20-26 April 2013, 2013; Curran; pp. 1169–1169. [Google Scholar]
- Mekkaoui. Fiber architecture in remodeled myocardium revealed with a quantitative diffusion CMR tractography framework and histological validation. Journal of Cardiovascular Magnetic Resonance 2012, vol. 14(no. 1), 71. [Google Scholar] [CrossRef]
- Dokos, S.; Smaill, B. H.; Young, A. A.; LeGrice, I. J. Shear properties of passive ventricular myocardium. American Journal of Physiology-Heart and Circulatory Physiology 2002, vol. 283(no. 6), H2650–H2659. [Google Scholar] [CrossRef] [PubMed]
- Guccione, J. M.; McCulloch, A. D.; Waldman, L. Passive material properties of intact ventricular myocardium determined from a cylindrical model; 1991. [Google Scholar]
- Guccione, J. M.; Costa, K. D.; McCulloch, A. D. Finite element stress analysis of left ventricular mechanics in the beating dog heart. Journal of biomechanics 1995, vol. 28(no. 10), 1167–1177. [Google Scholar] [CrossRef] [PubMed]
- Sacks, S.; Sun, W. Multiaxial mechanical behavior of biological materials. Annual review of biomedical engineering 2003, vol. 5(no. 1), 251–284. [Google Scholar] [CrossRef]
- Humphrey, J. D. Cardiovascular solid mechanics: cells, tissues, and organs; Springer Science & Business Media, 2013. [Google Scholar]
- Humphrey, J. D.; Rajagopal, K. A constrained mixture model for growth and remodeling of soft tissues. Mathematical models and methods in applied sciences 2002, vol. 12(no. 03), 407–430. [Google Scholar] [CrossRef]
- Nemavhola, F.; Pandelani, T. Right-Ventricular Remodeling Mechanics: From Fiber Architecture and Constitutive Modeling to Imaging Biomarkers and Clinical Endpoints; 2026. [Google Scholar]
- Nemavhola, F.; Pandelani, T. Infarct Border-Zone Biomechanics After Myocardial Infarction: Linking Mechanotransduction, Fibrosis, and Ventricular Dysfunction; 2026. [Google Scholar]
- Omens, J. H.; Fung, Y.-C. Residual strain in rat left ventricle. Circulation research 1990, vol. 66(no. 1), 37–45. [Google Scholar] [CrossRef]
- Fung, Y. C. What are the residual stresses doing in our blood vessels? Annals of biomedical engineering 1991, vol. 19(no. 3), 237–249. [Google Scholar] [CrossRef] [PubMed]
- Fung, Y.; Liu, S.; Zhou, J. Remodeling of the constitutive equation while a blood vessel remodels itself under stress. 1993. [Google Scholar] [CrossRef]
- Fung, Y.; Liu, S. Determination of the mechanical properties of the different layers of blood vessels in vivo. Proceedings of the National Academy of Sciences 1995, vol. 92(no. 6), 2169–2173. [Google Scholar] [CrossRef] [PubMed]
- Fung, Y.-c. Biomechanics: mechanical properties of living tissues; Springer Science & Business Media, 2013. [Google Scholar]
- Emig, R. Passive myocardial mechanical properties: meaning, measurement, models. Biophysical reviews 2021, vol. 13(no. 5), 587–610. [Google Scholar] [CrossRef] [PubMed]
- Bezy, S. Impact of loading and myocardial mechanical properties on natural shear waves: comparison to pressure-volume loops. Cardiovascular Imaging 2022, vol. 15(no. 12), 2023–2034. [Google Scholar]
- Stella, J. A.; D’Amore, A.; Wagner, W. R.; Sacks, M. S. On the biomechanical function of scaffolds for engineering load-bearing soft tissues. Acta biomaterialia 2010, vol. 6(no. 7), 2365–2381. [Google Scholar] [CrossRef]
- Holzapfel, G. A.; Ogden, R. W. Biomechanics of soft tissue in cardiovascular systems; Springer Science & Business Media, 2003. [Google Scholar]
- Famaey. Community challenge towards consensus on characterization of biological tissue: C4Bio’s first findings. Journal of Biomechanics 2025, 113021. [Google Scholar] [CrossRef]
- Nemavhola, F.; Ngwangwa, H.; Pandelani, T.; Davies, N.; Franz, T. Understanding regional mechanics of rat myocardia by fitting hyperelatsic models. 2021. [Google Scholar] [CrossRef]
- Ndlovu, Z.; Nemavhola, F.; Desai, D. Biaxial mechanical characterization and constitutive modelling of sheep sclera soft tissue. Russian Journal of Biomechanics/Rossijski Zurnal Biomehaniki 2020, vol. 24(no. 1), 84–96. [Google Scholar] [CrossRef]
- Ndlovu, Z.; Desai, D.; Nemavhola, F.; Ngwangwa, H. M. Sheep sclera soft tissue subjected to mechanical equi-biaxial testing. 2021. [Google Scholar] [CrossRef]
- Pandelani, T.; Ngwangwa, H.; Nemavhola, F. Experimental analysis and biaxial biomechanical behaviour of ex-vivo sheep trachea. Frontiers in Materials 2023, vol. 10, 1230789. [Google Scholar] [CrossRef]
- Semakane, L.; Mohan, T. P.; Ngwangwa, H.; Pandelani, T.; Nemavhola, F. Mechanical Behaviour of Breast Tissue: An In-Depth Systematic Review. 2024. [Google Scholar] [CrossRef]
- Avazmohammadi, R. An integrated inverse model-experimental approach to determine soft tissue three-dimensional constitutive parameters: application to post-infarcted myocardium. Biomechanics and modeling in mechanobiology 2018, vol. 17(no. 1), 31–53. [Google Scholar] [CrossRef]
- Hoang; Guccione, J. Finite Element Modeling in Left Ventricular Cardiac Biomechanics: From Computational Tool to Clinical Practice. Bioengineering vol. 12(no. 9), 913, 2025. [CrossRef]
- Mehri, M.; Wenk, J. F. A Unified Finite Element Framework for Cardiac Growth and Remodeling in Mitral Regurgitation Incorporating Fiber Reorientation and Baroreflex. Acta Biomaterialia 2025. [Google Scholar] [CrossRef]
- Mehri, M.; Campbell, K. S.; Lee, L. C.; Wenk, J. F. A multi-scale finite element method for investigating fiber remodeling in hypertrophic cardiomyopathy. Scientific Reports vol. 15(no. 1), 31961, 2025. [CrossRef]
- Salih, A.; Hamandi, F.; Goswami, T. Advancements in finite element modeling for cardiac device leads and 3D heart models. Bioengineering 2024, vol. 11(no. 6), 564. [Google Scholar] [CrossRef]
- Qiu; Weiss, D. Local mechanical characterization of cardiovascular tissues: methods, challenges, and pathways to clinical use. Frontiers in Mechanical Engineering vol. 11, 1703081, 2025. [CrossRef]
- Holzapfel, G. A.; Ogden, R. W. Constitutive Modeling of Soft Biological Tissues. In in Constitutive Modelling of Solid Continua; Springer, 2025; pp. 493–520. [Google Scholar]
- Wilson, A. J.; Sands, G. B.; LeGrice, I. J.; Young, A. A.; Ennis, D. B. Myocardial mesostructure and mesofunction. American Journal of Physiology-Heart and Circulatory Physiology 2022, vol. 323(no. 2), H257–H275. [Google Scholar] [CrossRef] [PubMed]
- Wang, Y.; Chatterjee, E.; Li, G.; Xu, J.; Xiao, J. Force-sensing protein expression in response to cardiovascular mechanotransduction. EBioMedicine 2024, vol. 110. [Google Scholar] [CrossRef]
- Osouli, K.; De Gaetano, F.; Costantino, M. L.; Peirlinck, M. Heart in a knot: unraveling the impact of the nested tori myofiber architecture on ventricular mechanics: K. Osouli et al. Biomechanics and Modeling in Mechanobiology 2025, vol. 24(no. 5), 1815–1835. [Google Scholar] [CrossRef] [PubMed]
- Trifunović-Zamaklar. The basic heart anatomy and physiology from the cardiologist’s perspective: Toward a better understanding of left ventricular mechanics, systolic, and diastolic function. Journal of Clinical Ultrasound 2022, vol. 50(no. 8), 1026–1040. [Google Scholar] [CrossRef]
- Mukherjee, T.; Mendiola, E. A.; Neelakantan, S.; Shah, D. J.; Avazmohammadi, R. Four-dimensional assessment of left ventricular torsion in mitral valve prolapse using CMR. in Medical Imaging 2024: Image Processing 2024, vol. 12926: SPIE, 108–115. [Google Scholar]
- Zhong, Y. Evaluation the relationship between myocardial fibrosis and left ventricular torsion measured by cardiac magnetic resonance feature-tracking in hypertrophic cardiomyopathy patients with preserved ejection fraction. The International Journal of Cardiovascular Imaging 2024, vol. 40(no. 4), 921–930. [Google Scholar] [CrossRef]
- Young, K.; Hooton, C.; Zimmerman, M. B.; Reinking, B.; Gupta, U. Fetal left and right ventricular strain parameters using speckle tracking in congenital heart diseases. The international journal of cardiovascular imaging 2024, vol. 40(no. 6), 1235–1243. [Google Scholar] [CrossRef]
- Muraru. Right ventricular longitudinal strain in the clinical routine: a state-of-the-art review. European Heart Journal-Cardiovascular Imaging 2022, vol. 23(no. 7), 898–912. [Google Scholar] [CrossRef]
- Salte, M. Deep learning for improved precision and reproducibility of left ventricular strain in echocardiography: a test-retest study. Journal of the American Society of Echocardiography 2023, vol. 36(no. 7), 788–799. [Google Scholar] [CrossRef] [PubMed]
- Benech, C.; Romanelli, G. Atomic force microscopy indentation for nanomechanical characterization of live pathological cardiovascular/heart tissue and cells. Micron 2022, vol. 158, 103287. [Google Scholar] [CrossRef]
- Cox, M. A.; Driessen, N. J.; Boerboom, R. A.; Bouten, C. V.; Baaijens, F. P. Mechanical characterization of anisotropic planar biological soft tissues using finite indentation: experimental feasibility. Journal of biomechanics 2008, vol. 41(no. 2), 422–429. [Google Scholar] [CrossRef]
- Codan; Del Favero, G.; Martinelli, V.; Long, C.; Mestroni, L.; Sbaizero, O. Exploring the elasticity and adhesion behavior of cardiac fibroblasts by atomic force microscopy indentation. Materials Science and Engineering: C 2014, vol. 40, 427–434. [Google Scholar] [CrossRef] [PubMed]
- Mathur, A. B.; Collinsworth, A. M.; Reichert, W. M.; Kraus, W. E.; Truskey, G. A. Endothelial, cardiac muscle and skeletal muscle exhibit different viscous and elastic properties as determined by atomic force microscopy. Journal of biomechanics 2001, vol. 34(no. 12), 1545–1553. [Google Scholar] [CrossRef] [PubMed]
- Hiesinger, W. Myocardial tissue elastic properties determined by atomic force microscopy after stromal cell–derived factor 1α angiogenic therapy for acute myocardial infarction in a murine model. The Journal of thoracic and cardiovascular surgery 2012, vol. 143(no. 4), 962–966. [Google Scholar] [CrossRef]
- Gupta, B.; Ratcliffe, M. B.; Fallert, M. A.; Edmunds, L. H., Jr.; Bogen, D. K. Changes in passive mechanical stiffness of myocardial tissue with aneurysm formation. Circulation 1994, vol. 89(no. 5), 2315–2326. [Google Scholar] [CrossRef]
- Holmes, J. W.; Nuñez, J. A.; Covell, J. W. Functional implications of myocardial scar structure. American Journal of Physiology-Heart and Circulatory Physiology 1997, vol. 272(no. 5), H2123–H2130. [Google Scholar] [CrossRef]
- Rausch, K. Growth and remodeling of atrioventricular heart valves: A potential target for pharmacological treatment? Current Opinion in Biomedical Engineering 2020, vol. 15, 10–15. [Google Scholar] [CrossRef]
- Rausch; Karniadakis, G.; Humphrey, J. Modeling soft tissue damage and failure using a combined particle/continuum approach. Biomechanics and modeling in mechanobiology 2017, vol. 16(no. 1), 249–261. [Google Scholar] [CrossRef]
- Mendiola, A. An image-driven micromechanical approach to characterize multiscale remodeling in infarcted myocardium. Acta biomaterialia 2024, vol. 173, 109–122. [Google Scholar] [CrossRef] [PubMed]
- Pitoulis, G. Remodelling of adult cardiac tissue subjected to physiological and pathological mechanical load in vitro. Cardiovascular Research 2022, vol. 118(no. 3), 814–827. [Google Scholar] [CrossRef] [PubMed]
- Frangogiannis, G. Pathophysiology of myocardial infarction. Comprehensive physiology 2015, vol. 5(no. 4), 1841–1875. [Google Scholar] [CrossRef]
- Frangogiannis, G. The mechanistic basis of infarct healing. Antioxidants & redox signaling 2006, vol. 8(no. 11-12), 1907–1939. [Google Scholar]
- Logeart. Patterns of left ventricular remodeling post-myocardial infarction, determinants, and outcome. Clinical Research in Cardiology 2024, vol. 113(no. 12), 1670–1681. [Google Scholar] [CrossRef]
- Gissler, M. C.; Antiochos, P.; Ge, Y.; Heydari, B.; Gräni, C.; Kwong, R. Y. Cardiac magnetic resonance evaluation of LV remodeling post-myocardial infarction: prognosis, monitoring and trial endpoints. Cardiovascular Imaging 2024, vol. 17(no. 11), 1366–1380. [Google Scholar]
- Yong, J.; Tao, J.; Wang, K.; Li, X.; Yang, Y. Post-myocardial infarction cardiac remodeling: multidimensional mechanisms and clinical prospects of stem cell therapy. Stem Cell Reviews and Reports 2025, vol. 21(no. 5), 1369–1427. [Google Scholar] [CrossRef] [PubMed]
- Frantz, S.; Hundertmark, M. J.; Schulz-Menger, J.; Bengel, F. M.; Bauersachs, J. Left ventricular remodelling post-myocardial infarction: pathophysiology, imaging, and novel therapies. European heart journal 2022, vol. 43(no. 27), 2549–2561. [Google Scholar] [CrossRef]
- Peña, B.; Adbel-Hafiz, M.; Cavasin, M.; Mestroni, L.; Sbaizero, O. Atomic force microscopy (AFM) applications in arrhythmogenic cardiomyopathy. International journal of molecular sciences 2022, vol. 23(no. 7), 3700. [Google Scholar] [CrossRef]
- Villarreal, L.; Fontes-Silva, M.; Mendaro, L.; Romanelli, G.; Benech, J. C. Mechanics and disease of heart cells/cardiomyocytes explored through atomic force microscopy: present and future. Biophysical Reviews 2025, vol. 17(no. 2), 347–358. [Google Scholar] [CrossRef]
- Loewe, A.; Hunter, P. J.; Kohl, P. Computational modelling of biological systems now and then: revisiting tools and visions from the beginning of the century. Philosophical Transactions A vol. 383(no. 2296), 20230384, 2025. [CrossRef]
- Lo, S. C. A multi-component, multi-physics computational model for solving coupled cardiac electromechanics and vascular haemodynamics. Computer Methods in Applied Mechanics and Engineering vol. 446, 118185, 2025. [CrossRef]
- Jones, E.; Oomen, P. J. Synergistic biophysics and machine learning modeling to rapidly predict cardiac growth probability. Computers in Biology and Medicine 2025, vol. 184, 109323. [Google Scholar] [CrossRef] [PubMed]
- van Osta, N.; van Loon, T.; Lumens, J. Individual hearts: computational models for improved management of cardiovascular disease. In Heart; 2025. [Google Scholar]
- Familusi, M. A. Computational modeling of the tissue mechanics in rheumatic heart disease patients. 2025. [Google Scholar]
- Faza, N. N.; Harb, S. C.; Wang, D. D.; van den Dorpel, M. M.; Van Mieghem, N.; Little, S. H. Physical and computational modeling for transcatheter structural heart interventions. Cardiovascular Imaging 2024, vol. 17(no. 4), 428–440. [Google Scholar] [CrossRef]
- Hinton, A., Jr. Mitochondrial structure and function in human heart failure. Circulation research 2024, vol. 135(no. 2), 372–396. [Google Scholar] [CrossRef]
- Jiao. Nanopatterned Cell Sheet Assembly of Biomimetic Cardiac Laminae for Modeling Structure–Function Relationships. Biomaterials Research 2026, vol. 30, 0339. [Google Scholar] [CrossRef]
- Rajagopal, V.; Arumugam, S.; Hunter, P. J.; Khadangi, A.; Chung, J.; Pan, M. The cell physiome: what do we need in a computational physiology framework for predicting single-cell biology? Annual Review of Biomedical Data Science 2022, vol. 5(no. 1), 341–366. [Google Scholar] [CrossRef]
- Min, J. Association of pericardial fat with cardiac structure, function, and mechanics: the multi-ethnic study of atherosclerosis. Journal of the American Society of Echocardiography 2022, vol. 35(no. 6), 579–587. e5. [Google Scholar] [CrossRef] [PubMed]
- Tan, S. Differential associations of A-/B-type natriuretic peptides with cardiac structure, function, and prognosis in heart failure. Heart Failure 2024, vol. 12(no. 3), 461–474. [Google Scholar] [CrossRef] [PubMed]
- Lupisella, J. Molecular mechanisms of desensitization underlying the differential effects of formyl peptide receptor 2 agonists on cardiac structure–function post myocardial infarction. ACS Pharmacology & Translational Science 2022, vol. 5(no. 10), 892–906. [Google Scholar] [CrossRef]
- Brown, A. L.; Liu, J.; Ennis, D. B.; Marsden, A. L. Cardiac mechanics modeling: recent developments and current challenges. ArXiv 2509.07971 v1. 2025. [Google Scholar]
- Nordsletten, D. A viscoelastic model for human myocardium. Acta Biomaterialia vol. 135, 441–457, 2021/11/01/ 2021. [CrossRef] [PubMed]
- Zhang, W.; Jadidi, M.; Razian, S. A.; Holzapfel, G. A.; Kamenskiy, A.; Nordsletten, D. A. “A viscoelastic constitutive framework for aging muscular and elastic arteries,” (in eng). Acta Biomater 2024, vol. 188, 223–241. [Google Scholar] [CrossRef]
- Gkouti; Czekanski, A.; AlAttar, A. Simulating and Predicting the Mechanical Behavior of Electrospun Scaffolds for Cardiac Patches Fabrication. Materials 2023, vol. 16(no. 22), 7095. Available online: https://www.mdpi.com/1996-1944/16/22/7095. [CrossRef]
- Sommer. Biomechanical properties and microstructure of human ventricular myocardium. Acta Biomaterialia vol. 24, 172–192, 2015/09/15/ 2015. [CrossRef]
- Richardson, W. J.; Clarke, S. A.; Quinn, T. A.; Holmes, J. W. Physiological implications of myocardial scar structure. Comprehensive Physiology 2015, vol. 5(no. 4), 1877–1909. [Google Scholar] [CrossRef]
- Schwan, J. Anisotropic engineered heart tissue made from laser-cut decellularized myocardium. Scientific Reports 2016/08/30 2016, vol. 6(no. 1), 32068. [Google Scholar] [CrossRef]
- Liang, Z. Structural mechanisms linking muscle fiber alignment to elastic modulus in skeletal muscle. Materials & Design vol. 263, 115528, 2026/03/01/ 2026. [CrossRef]
- Laita, N. A comprehensive experimental analysis of the local passive response across the healthy porcine left ventricle. Acta Biomaterialia vol. 187, 261–277, 2024/10/01/ 2024. [CrossRef] [PubMed]
- Bianco; Levy, A. M.; Grytz, R.; Fazio, M. A. “Effect of different preconditioning protocols on the viscoelastic inflation response of the posterior sclera,” (in eng). Acta Biomater 2021, vol. 128, 332–345. [Google Scholar] [CrossRef]
- Bolli. “Preconditioning: a paradigm shift in the biology of myocardial ischemia,” (in eng). Am J Physiol Heart Circ Physiol 2007, vol. 292(no. 1), H19–27. [Google Scholar] [CrossRef]
- A. C. Silva, C. Pereira, A. C. R. G. Fonseca, P. Pinto-do-Ó, and D. S. Nascimento, “Bearing My Heart: The Role of Extracellular Matrix on Cardiac Development, Homeostasis, and Injury Response,” (in English), Frontiers in Cell and Developmental Biology, Review vol. Volume 8—2020, 2021-January-12 2021. [CrossRef]
- Tran, D. T.; Tsai, L. “Effect of strain rates on the mechanical response of whole muscle bundle,” (in eng). J Biol Phys 2023, vol. 49(no. 2), 257–267. [Google Scholar] [CrossRef] [PubMed]
- Bahadormanesh, N.; Tomka, B.; Kadem, M.; Khodaei, S.; Keshavarz-Motamed, Z. An ultrasound-exclusive non-invasive computational diagnostic framework for personalized cardiology of aortic valve stenosis. Medical Image Analysis 2023, vol. 87, 102795. [Google Scholar] [CrossRef]
- Xia, L.; Zhang, H.; Zheng, D. Multi-Scale Computational Cardiology; Frontiers Media SA, 2022; vol. 13, p. 847118. [Google Scholar]
- Sacks, M.; Chuong, C. Biaxial mechanical properties of passive right ventricular free wall myocardium; 1993. [Google Scholar]
- Krüger, M.; Linke, W. A. Titin-based mechanical signalling in normal and failing myocardium. Journal of molecular and cellular cardiology 2009, vol. 46(no. 4), 490–498. [Google Scholar] [CrossRef] [PubMed]
- Borbély, A. Hypophosphorylation of the Stiff N2B titin isoform raises cardiomyocyte resting tension in failing human myocardium. Circulation research 2009, vol. 104(no. 6), 780–786. [Google Scholar] [CrossRef]
- Hamdani, N. Myocardial titin hypophosphorylation importantly contributes to heart failure with preserved ejection fraction in a rat metabolic risk model. Circulation: Heart Failure 2013, vol. 6(no. 6), 1239–1249. [Google Scholar] [CrossRef]
- Gültekin, O.; Sommer, G.; Holzapfel, G. A. An orthotropic viscoelastic model for the passive myocardium: continuum basis and numerical treatment. Computer methods in biomechanics and biomedical engineering 2016, vol. 19(no. 15), 1647–1664. [Google Scholar] [CrossRef]
- Hussein, T. M.; Criscione, J. C. A new strain energy function representing the passive behavior of the myocardium. Journal of Biomechanical Engineering 2023, vol. 145(no. 11), 111004. [Google Scholar] [CrossRef]
- Namashiri; Allahverdizadeh, A.; Dadashzadeh, B.; Dokos, S. Electromechanical modeling of the left ventricle: considering hyperelastic and viscoelastic properties. Journal of the Brazilian Society of Mechanical Sciences and Engineering 2024, vol. 46(no. 12), 704. [Google Scholar] [CrossRef]
- Pan; Ng, S. M.; Neubauer, S.; Rider, O. J. Phenotyping heart failure by cardiac magnetic resonance imaging of cardiac macro-and microscopic structure: state of the art review. European Heart Journal-Cardiovascular Imaging 2023, vol. 24(no. 10), 1302–1317. [Google Scholar] [CrossRef] [PubMed]
- Sillanmäki, H.-L.; Vainio; Ylä-Herttuala, E.; Husso, M.; Hedman, M. Measuring Cardiac Dyssynchrony with DENSE (Displacement Encoding with Stimulated Echoes)—A Systematic Review. Reviews in Cardiovascular Medicine 2023, vol. 24(no. 9), 261. [Google Scholar] [CrossRef] [PubMed]
- Osman, N. F.; Kerwin, W. S.; McVeigh, E. R.; Prince, J. L. Cardiac motion tracking using CINE harmonic phase (HARP) magnetic resonance imaging. Magnetic Resonance in Medicine: An Official Journal of the International Society for Magnetic Resonance in Medicine 1999, vol. 42(no. 6), 1048–1060. [Google Scholar] [CrossRef]
- Guccione, M.; Waldman, L.; McCulloch, A. D. Mechanics of active contraction in cardiac muscle: Part II—Cylindrical models of the systolic left ventricle; 1993. [Google Scholar]
- Guccione, M.; McCulloch, A. D. Mechanics of active contraction in cardiac muscle: part I—constitutive relations for fiber stress that describe deactivation; 1993. [Google Scholar]
- Wang, V.; Nielsen, P.; Nash, M. Image-based predictive modeling of heart mechanics. Annual review of biomedical engineering 2015, vol. 17(no. 1), 351–383. [Google Scholar] [CrossRef]
- Sacks, S.; Mirnajafi, A.; Sun, W.; Schmidt, P. Bioprosthetic heart valve heterograft biomaterials: structure, mechanical behavior and computational simulation. Expert review of medical devices 2006, vol. 3(no. 6), 817–834. [Google Scholar] [CrossRef]
- Muthupillai; Lomas, D. J.; Rossman, P. J.; Greenleaf, J. F.; Manduca, A.; Ehman, R. L. Magnetic resonance elastography by direct visualization of propagating acoustic strain waves. science 1995, vol. 269(no. 5232), 1854–1857. [Google Scholar] [CrossRef]
- Meyer, K. Free-Breathing, 3D Cardiac Magnetic Resonance Elastography for Myocardial Stiffness Mapping. Magnetic Resonance in Medicine 2026, vol. 95(no. 4), 2356–2369. [Google Scholar] [CrossRef]
- Meyer. Cardiac elastography with external vibration for quantification of diastolic myocardial stiffness. Journal of the American Society of Echocardiography 2025, vol. 38(no. 5), 431–442. [Google Scholar] [CrossRef]
- Sack; Rump, J.; Elgeti, T.; Samani, A.; Braun, J. MR elastography of the human heart: noninvasive assessment of myocardial elasticity changes by shear wave amplitude variations. Magnetic Resonance in Medicine: An Official Journal of the International Society for Magnetic Resonance in Medicine 2009, vol. 61(no. 3), 668–677. [Google Scholar] [CrossRef]
- Kolipaka, A. MR elastography as a method for the assessment of myocardial stiffness: comparison with an established pressure–volume model in a left ventricular model of the heart. Magnetic Resonance in Medicine: An Official Journal of the International Society for Magnetic Resonance in Medicine 2009, vol. 62(no. 1), 135–140. [Google Scholar] [CrossRef]
- Arani. Cardiac MR elastography for quantitative assessment of elevated myocardial stiffness in cardiac amyloidosis. Journal of Magnetic Resonance Imaging 2017, vol. 46(no. 5), 1361–1367. [Google Scholar] [CrossRef] [PubMed]
- Amundsen, H. Noninvasive myocardial strain measurement by speckle tracking echocardiography: validation against sonomicrometry and tagged magnetic resonance imaging. Journal of the American College of Cardiology 2006, vol. 47(no. 4), 789–793. [Google Scholar] [CrossRef] [PubMed]
- Mirea, O. Variability and reproducibility of segmental longitudinal strain measurement: a report from the EACVI-ASE strain standardization task force. JACC: Cardiovascular Imaging 2018, vol. 11(no. 1), 15–24. [Google Scholar]
- Blessberger; Binder, T. Two dimensional speckle tracking echocardiography: basic principles. Heart 2010, vol. 96(no. 9), 716–722. [Google Scholar] [CrossRef]
- Voigt, J.-U. Definitions for a common standard for 2D speckle tracking echocardiography: consensus document of the EACVI/ASE/Industry Task Force to standardize deformation imaging. European Heart Journal-Cardiovascular Imaging 2015, vol. 16(no. 1), 1–11. [Google Scholar] [CrossRef]
- Mor-Avi. Current and evolving echocardiographic techniques for the quantitative evaluation of cardiac mechanics: ASE/EAE consensus statement on methodology and indications endorsed by the Japanese Society of Echocardiography. European Journal of Echocardiography 2011, vol. 12(no. 3), 167–205. [Google Scholar] [CrossRef]
- Gillam, L. D.; Marcoff, L. Echocardiography: past, present, and future. Circulation: Cardiovascular Imaging 2024, vol. 17(no. 4), e016517. [Google Scholar] [CrossRef]
- Al-Assaf, A. M. A. M. An Overview for Echocardiography Techniques-An Updated Review for Modern and Advanced Techniques and Innovations. Journal of Medical and Life Science 2024, vol. 6(no. 4), 611–632. [Google Scholar] [CrossRef]
- Chakraborti, S. Jana; Dey, S. The advancement of echocardiography through evolutionary development. J. Community Health Management 2023, vol. 9(no. 4), 123–131. [Google Scholar]
- Kaipio, P.; Somersalo, E. Statistical and computational inverse problems; Springer, 2005. [Google Scholar]
- Borowska; Gao, H.; Lazarus, A.; Husmeier, D. Bayesian optimisation for efficient parameter inference in a cardiac mechanics model of the left ventricle. International Journal for Numerical Methods in Biomedical Engineering 2022, vol. 38(no. 5), e3593. [Google Scholar] [CrossRef] [PubMed]
- Sundnes; Rodríguez-Cantano, R. A Bayesian approach to parameter estimation in cardiac mechanics. In in Solid (Bio) mechanics: Challenges of the Next Decade: A Book Dedicated to Professor Gerhard A. Holzapfel; Springer, 2022; pp. 245–256. [Google Scholar]
- Doraiswamy, S.; Criscione, J. C.; Srinivasa, A. R. A technique for the classification of tissues by combining mechanics based models with Bayesian inference. International Journal of Engineering Science 2016, vol. 106, 95–109. [Google Scholar] [CrossRef]
- Wang, S. Bayesian inference-based estimation of normal aortic, aneurysmal and atherosclerotic tissue mechanical properties: From material testing, modeling and histology. IEEE Transactions on Biomedical Engineering 2019, vol. 66(no. 8), 2269–2278. [Google Scholar] [CrossRef] [PubMed]
- Ngwangwa, H.; Modungwa, D.; Pandelani, T.; Nemavhola, F. Estimation of the biaxial tensile behavior of ovine esophageal tissue using artificial neural networks. BioMedical Engineering OnLine 2024, vol. 23(no. 1), 1–44. [Google Scholar] [CrossRef]
- Ngwangwa, H. M.; Nemavhola, F. Evaluating computational performances of hyperelastic models on supraspinatus tendon uniaxial tensile test data. Journal of Computational Applied Mechanics 2021, vol. 52(no. 1), 27–43. [Google Scholar]
- Shrestha, S.; Sengupta, P. P. The mechanics of machine learning: from a concept to value. Journal of the American Society of Echocardiography 2018, vol. 31(no. 12), 1285–1287. [Google Scholar] [CrossRef]
- Arzani, A.; Wang, J.-X.; Sacks, M. S.; Shadden, S. C. Machine learning for cardiovascular biomechanics modeling: challenges and beyond. Annals of Biomedical Engineering 2022, vol. 50(no. 6), 615–627. [Google Scholar] [CrossRef] [PubMed]
- Dabiri, Y.; Van der Velden, A.; Sack, K. L.; Choy, J. S.; Kassab, G. S.; Guccione, J. M. Prediction of left ventricular mechanics using machine learning. Frontiers in physics 2019, vol. 7, 117. [Google Scholar] [CrossRef] [PubMed]
- Francesco. Mathematical modeling and Machine Learning for the numerical simulation of cardiac electromechanics; 2020. [Google Scholar]
- Karniadakis, G. E.; Kevrekidis, I. G.; Lu, L.; Perdikaris, P.; Wang, S.; Yang, L. Physics-informed machine learning. Nature Reviews Physics 2021, vol. 3(no. 6), 422–440. [Google Scholar] [CrossRef]
- Corral-Acero, J. The ‘Digital Twin’to enable the vision of precision cardiology. European heart journal 2020, vol. 41(no. 48), 4556–4564. [Google Scholar] [CrossRef]
- Morotti. Precision cardiac electrophysiology: Toward digital twins and beyond. The Journal of Precision Medicine: Health and Disease 2025, vol. 2. [Google Scholar] [CrossRef]
- Thangaraj, M.; Benson, S. H.; Oikonomou, E. K.; Asselbergs, F. W.; Khera, R. Cardiovascular care with digital twin technology in the era of generative artificial intelligence. European Heart Journal 2024, vol. 45(no. 45), 4808–4821. [Google Scholar] [CrossRef]
- Clarke, A.; Richardson, W. J.; Holmes, J. W. Modifying the mechanics of healing infarcts: Is better the enemy of good? Journal of molecular and cellular cardiology 2016, vol. 93, 115–124. [Google Scholar] [CrossRef]
- Zile, R.; Baicu, C. F.; Gaasch, W. H. Diastolic heart failure—abnormalities in active relaxation and passive stiffness of the left ventricle. New England Journal of Medicine 2004, vol. 350(no. 19), 1953–1959. [Google Scholar] [CrossRef]
- Chiorescu, R. M. Current insights and future directions in the treatment of heart failure with preserved ejection fraction. International Journal of Molecular Sciences 2023, vol. 25(no. 1), 440. [Google Scholar] [CrossRef]
- Paulus, J.; Tschöpe, C. A novel paradigm for heart failure with preserved ejection fraction: comorbidities drive myocardial dysfunction and remodeling through coronary microvascular endothelial inflammation. Journal of the American college of cardiology 2013, vol. 62(no. 4), 263–271. [Google Scholar] [CrossRef]
- Westermann. Role of left ventricular stiffness in heart failure with normal ejection fraction. Circulation 2008, vol. 117(no. 16), 2051–2060. [Google Scholar] [CrossRef]
- Ho, Y. Myocardial fibrosis as an early manifestation of hypertrophic cardiomyopathy. New England Journal of Medicine 2010, vol. 363(no. 6), 552–563. [Google Scholar] [CrossRef] [PubMed]
- Marian, A. J.; Braunwald, E. Hypertrophic cardiomyopathy: genetics, pathogenesis, clinical manifestations, diagnosis, and therapy. Circulation research 2017, vol. 121(no. 7), 749–770. [Google Scholar] [CrossRef]
- Humphrey, J. D.; Tellides, G. Central artery stiffness and thoracic aortopathy. American Journal of Physiology-Heart and Circulatory Physiology 2019, vol. 316(no. 1), H169–H182. [Google Scholar] [CrossRef]
- Humphrey, J. D.; Schwartz, M. A.; Tellides, G.; Milewicz, D. M. Role of mechanotransduction in vascular biology: focus on thoracic aortic aneurysms and dissections. Circulation research 2015, vol. 116(no. 8), 1448–1461. [Google Scholar] [CrossRef]
- Humphrey, J. D. Mechanisms of arterial remodeling in hypertension: coupled roles of wall shear and intramural stress. Hypertension 2008, vol. 52(no. 2), 195–200. [Google Scholar] [CrossRef]
- Dell’Italia, J.; Kane, M. S.; Zheng, J.; Huang, S. W.; Pat, B.; Denney, T. S. Persistent Shift in Principal Strain Angles and Postsurgical Decline in Left Ventricular Ejection Fraction in Patients With Primary Mitral Regurgitation. Journal of the American Heart Association 2025, vol. 14(no. 22), e043558. [Google Scholar] [CrossRef] [PubMed]
- Rodríguez-Padilla, J. Impact of intraventricular septal fiber orientation on cardiac electromechanical function. American Journal of Physiology-Heart and Circulatory Physiology 2022, vol. 322(no. 6), H936–H952. [Google Scholar] [CrossRef] [PubMed]
- Scollan, F.; Holmes, A.; Winslow, R.; Forder, J. Histological validation of myocardial microstructure obtained from diffusion tensor magnetic resonance imaging. American Journal of Physiology-Heart and Circulatory Physiology 1998, vol. 275(no. 6), H2308–H2318. [Google Scholar] [CrossRef]
- Cook, P.; Dhahri, W.; Laflamme, M. A.; Ghugre, N. R.; Wright, G. A. Using diffusion tensor imaging to depict myocardial changes after matured pluripotent stem cell-derived cardiomyocyte transplantation. Journal of Cardiovascular Magnetic Resonance 2024, vol. 26(no. 2), 101045. [Google Scholar] [CrossRef]
- Lohr, D. Assessment of myocardial microstructure in a murine model of obesity-related cardiac dysfunction by diffusion tensor magnetic resonance imaging at 7T. Frontiers in Cardiovascular Medicine 2022, vol. 9, 839714. [Google Scholar] [CrossRef]
- Ambiga, R.; Verma, S. Morphology, microstructure and biomechanical properties of tendinous cords of heart–a systematic review of cadaveric studies. Anatomy 2025, vol. 19(no. 1), 30–40. [Google Scholar] [CrossRef]
- Baghersad; Sathish Kumar, A.; Kipper, M. J.; Popat, K.; Wang, Z. Recent advances in tissue-engineered cardiac scaffolds—the progress and gap in mimicking native myocardium mechanical behaviors. Journal of Functional Biomaterials 2023, vol. 14(no. 5), 269. [Google Scholar] [CrossRef]
- Eastep, R. A Comparative Biomechanical Investigation of Porcine Neonatal and Adult Left Ventricular Tissues; The University of Texas at Arlington, 2023. [Google Scholar]
- Sultan, A. A. Image-Based Models for Left Heart Flow Morphodynamics in Hypertrophic Obstructive Cardiomyopathy; The American University in Cairo (Egypt), 2025. [Google Scholar]
- Kakaletsis, S. Histomechanical characterization and microstructure-based modeling of right ventricular myocardium. 2023. [Google Scholar]
- Ghafarinatanzi, M. Identification of Mechanical Properties for Patient-Specific Left Ventricle Models from CMR Using Virtual Fields Method; Ecole Polytechnique: Montreal (Canada), 2023. [Google Scholar]
- Bragard, J. R. Cardiac computational modelling. Revista Española de Cardiología (English Edition) 2021, vol. 74(no. 1), 65–71. [Google Scholar] [CrossRef]
- Trayanova, A. Computational cardiology: the heart of the matter. International Scholarly Research Notices 2012, vol. 2012(no. 1), 269680. [Google Scholar] [CrossRef]
- Petrescu, A. Ultrasound shear wave elastography for detection of myocardial fibrosis. European Heart Journal-Cardiovascular Imaging 2025, vol. 26(no. 9), 1537–1545. [Google Scholar] [CrossRef]
- Benz, D. C. Echocardiographic characterization of myocardial stiffness in healthy volunteers, cardiac amyloidosis, and hypertrophic cardiomyopathy: a case-control study using multimodality imaging. Circulation: Cardiovascular Imaging vol. 18(no. 3), e017475, 2025. [CrossRef]
- Anders, M. S. Time-resolved mapping of myocardial stiffness using 2D multifrequency spiral MR elastography with and without external vibration. Magnetic Resonance in Medicine vol. 94(no. 6), 2421–2434, 2025. [CrossRef]
- Castelein, J. Reproducibility of cardiac multifrequency MR elastography in assessing left ventricular stiffness and viscosity. Journal of Magnetic Resonance Imaging vol. 61(no. 5), 2146–2154, 2025. [CrossRef]
- González, A.; Schelbert, E. B.; Díez, J.; Butler, J. Myocardial interstitial fibrosis in heart failure: biological and translational perspectives. Journal of the American College of Cardiology 2018, vol. 71(no. 15), 1696–1706. [Google Scholar] [CrossRef]
- Schotten, A. Goette; Verheule, S. Translation of pathophysiological mechanisms of atrial fibrosis into new diagnostic and therapeutic approaches. Nature Reviews Cardiology 2025, vol. 22(no. 4), 225–240. [Google Scholar] [CrossRef] [PubMed]
- Ghazal, R.; Wang, M.; Liu, D.; Tschumperlin, D. J.; Pereira, N. L. Cardiac fibrosis in the multi-omics era: implications for heart failure. Circulation research 2025, vol. 136(no. 7), 773–802. [Google Scholar] [CrossRef]
- Tocchetti, G. How to facilitate seamless translation from basic concepts to new heart failure drugs. A scientific statement of the Heart Failure Association of the ESC. European journal of heart failure 2025, vol. 27(no. 8), 1379–1392. [Google Scholar] [CrossRef]
- Zivalj, L. M. S.; Schleicher; Krsek, A.; Diamee, H. J. F.; Raljevic, D.; Baticic, L. MicroRNAs in Heart Failure Pathogenesis and Progression: Mechanistic Control, Biomarker Potential, and Translational Perspectives. Life 2026, vol. 16(no. 3), 400. [Google Scholar] [CrossRef]
- Imran, T. F. Translating Mechanistic Insights Into Action and Revealing New Pathways: Machine Learning Approaches in Heart Failure With Preserved Ejection Fraction. Journal of the American Heart Association 2026, e043617. [Google Scholar] [CrossRef]
- Paillard, M. Mitochondrial targets in ischaemic heart disease and heart failure, and their potential for a more efficient clinical translation. A scientific statement of the ESC Working Group on Cellular Biology of the Heart and the ESC Working Group on Myocardial Function. European journal of heart failure 2025, vol. 27(no. 9), 1720–1736. [Google Scholar]
- Janssens, K. L.; Bovendeerd, P. H. The Role of Infarct Stiffness in Cardiac Patch Therapy: A Computational Study Using an Idealized Left Ventricular Geometry. Journal of biomechanical engineering 2025, vol. 147(no. 8), 081005. [Google Scholar] [CrossRef] [PubMed]
- Neelakantan, S.; Pinto, D.; Mehdi, R. R.; Peighambari, S. B.; Myers, K. J.; Avazmohammadi, R. Intracardiac Hemodynamics Alteration in Myocardial Infarction. In in International Conference on Functional Imaging and Modeling of the Heart; Springer, 2025; pp. 430–441. [Google Scholar]
- Awadi, R. An Improved Computational Model Based on Cardiac Imaging Data for the Quantitative and Qualitative Assessment of the Mechanical Properties of the Human Left Ventricle. Journal of Computational and Cognitive Engineering 2026, vol. 5(no. 1), 62–73. [Google Scholar] [CrossRef]
- Magaña; Pezzuto, S.; Costabal, F. Sahli. Ensemble learning of the atrial fibre orientation with physics-informed neural networks. The Journal of Physiology 2025. [Google Scholar] [CrossRef] [PubMed]
- Gültekin, O.; Moeineddin, A.; Cansız, B.; Sveric, K.; Linke, A.; Kaliske, M. A Physics-Informed Neural Network Model for the Anisotropic Hyperelasticity of the Human Passive Myocardium. International Journal for Numerical Methods in Engineering vol. 126(no. 14), e70067, 2025. [CrossRef]
- Yalcinkaya, M. Physics-informed deep learning model selection for robust segmentation of multi-center stress perfusion datasets: results from the SCMR registry. Journal of Cardiovascular Magnetic Resonance vol. 27, 2025. [CrossRef]
- Höfler, M. Physics-informed neural network estimation of active material properties in time-dependent cardiac biomechanical models. arXiv 2025, arXiv:2505.03382. [Google Scholar] [CrossRef]
| Experimental Method | Loading Mode | Mechanical Properties Measured | Advantages | Limitations | Typical Applications |
| Uniaxial tensile testing | Tensile loading along a single direction (fiber or cross-fiber) | Stress–strain relationship, elastic modulus, nonlinear stiffness | Simple experimental setup; historically widely used; useful for directional stiffness estimation | Does not capture anisotropic coupling between directions; boundary effects can influence results | Early studies of myocardial stiffness; characterization of directional mechanics |
| Biaxial mechanical testing | Simultaneous loading along two orthogonal directions | Anisotropic stress–strain behavior; fiber vs cross-fiber stiffness; constitutive parameter identification | Considered the gold standard for soft tissue mechanics; captures anisotropic coupling | Requires complex specimen preparation; experimental setup more demanding | Parameter identification for constitutive models; finite element simulations of cardiac mechanics |
| Shear testing | Tangential deformation under controlled shear strain | Shear modulus; laminar sheet mechanics; inter-sheet sliding | Provides insight into sheet architecture and ventricular wall thickening mechanisms | Difficult specimen preparation; shear boundary conditions can be challenging | Investigation of myocardial sheet mechanics and laminar deformation |
| Torsion testing | Rotational deformation applied to tissue samples | Torsional stiffness; fiber–sheet coupling behavior | Mimics physiological torsional deformation of the ventricle | Experimental implementation is complex; rarely applied to small samples | Study of ventricular torsion and fiber orientation effects |
| Indentation / micro-indentation testing | Localized compressive loading using rigid indenter | Local tissue stiffness; spatial heterogeneity; regional mechanical properties | Enables mapping of stiffness across tissue surfaces; useful for infarct and scar regions | Interpretation requires contact mechanics models; sensitive to boundary effects | Characterization of infarct stiffness; mapping mechanical heterogeneity |
| Atomic force microscopy (AFM) | Nanoscale indentation using cantilever probe | Microscale elastic modulus; cellular and extracellular matrix stiffness | High spatial resolution; useful for cellular-scale mechanics | Limited penetration depth; sensitive to surface conditions | Measurement of stiffness in cardiomyocytes and extracellular matrix |
| Rheological testing | Oscillatory shear loading | Viscoelastic properties; storage and loss moduli; frequency-dependent stiffness | Quantifies time-dependent mechanical behavior | Often requires homogenized or modified samples | Study of myocardial viscoelasticity and damping behavior |
| Inflation testing (ventricular pressurization) | Pressure-driven deformation of intact ventricular wall | Global ventricular stiffness; pressure–volume relationships | Closely mimics physiological loading conditions | Requires intact specimens; difficult parameter identification | Whole-heart mechanical characterization and model validation |
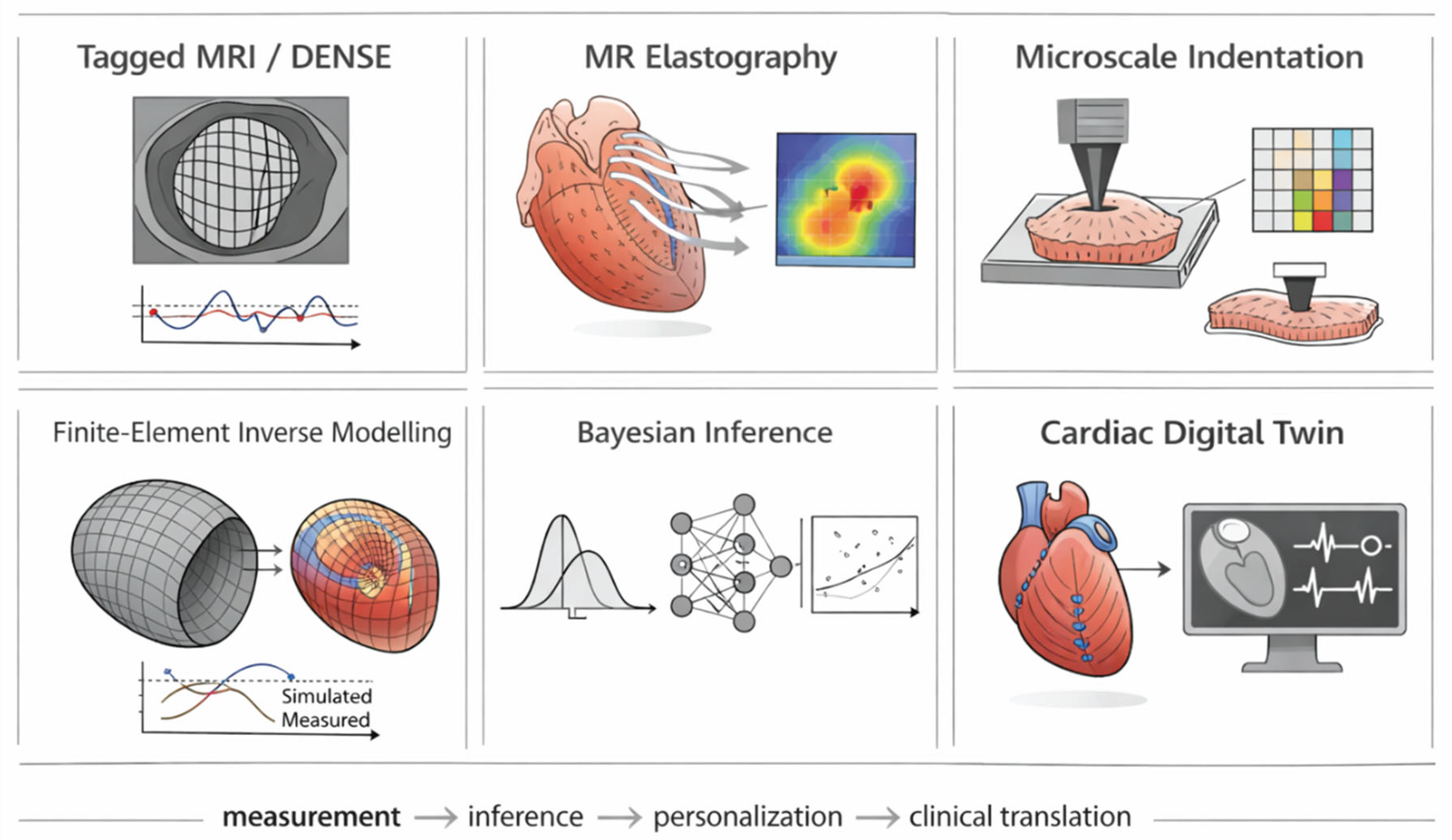
| Imaging Modality | Measurement Principle | Mechanical Parameters Estimated | Advantages | Limitations | Typical Applications |
| Tagged Magnetic Resonance Imaging (Tagged MRI) | Spatial modulation of magnetization creates tag lines that deform with myocardial motion | Myocardial strain, regional deformation patterns | High spatial resolution; well-established technique for cardiac strain analysis | Requires specialized pulse sequences; time-consuming image processing | Quantification of ventricular deformation; validation of computational models |
| Displacement Encoding with Stimulated Echoes (DENSE MRI) | Direct encoding of tissue displacement within the MRI signal phase | Myocardial displacement fields; strain tensors | High accuracy in displacement measurements; suitable for detailed strain mapping | Technically complex acquisition; sensitive to motion artifacts | Quantitative assessment of myocardial mechanics and ventricular deformation |
| Feature Tracking MRI | Post-processing tracking of anatomical features across cine MRI frames | Global and regional myocardial strain | Does not require specialized imaging sequences; compatible with standard cine MRI | Lower spatial resolution compared with tagged MRI; dependent on image quality | Clinical assessment of ventricular mechanics |
| Magnetic Resonance Elastography (MRE) | Mechanical shear waves propagated through tissue and measured with MRI | Shear modulus; regional myocardial stiffness | Direct estimation of tissue stiffness; spatial mapping of mechanical properties | Technically challenging due to cardiac motion; limited clinical availability | Quantification of myocardial stiffness in fibrosis and heart failure |
| Speckle Tracking Echocardiography | Tracking of natural acoustic speckle patterns in ultrasound images | Longitudinal, circumferential, and radial strain | Widely available; non-invasive; real-time imaging | Dependent on image quality and acoustic window; lower spatial resolution | Clinical evaluation of myocardial function and early detection of dysfunction |
| Ultrasound Shear Wave Elastography | Ultrasound-generated shear waves used to estimate tissue elasticity | Shear modulus and stiffness distribution | Rapid acquisition; non-invasive stiffness estimation | Limited penetration depth; sensitive to motion artifacts | Experimental assessment of myocardial stiffness |
| Diffusion Tensor MRI (DT-MRI) | Measurement of water diffusion anisotropy within myocardial tissue | Fiber orientation and structural anisotropy | Provides detailed myocardial microstructure; useful for modeling fiber architecture | Requires long acquisition times; mainly used in research settings | Reconstruction of myocardial fiber architecture for computational models |
| Computed Tomography (CT)-based motion analysis | High-resolution imaging combined with motion tracking algorithms | Ventricular deformation and strain estimates | High spatial resolution; useful when MRI is contraindicated | Radiation exposure; limited soft tissue contrast | Structural and functional cardiac imaging |
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2026 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).




