Submitted:
03 April 2026
Posted:
07 April 2026
You are already at the latest version
Abstract
Keywords:
1. Introduction
2. Related Work
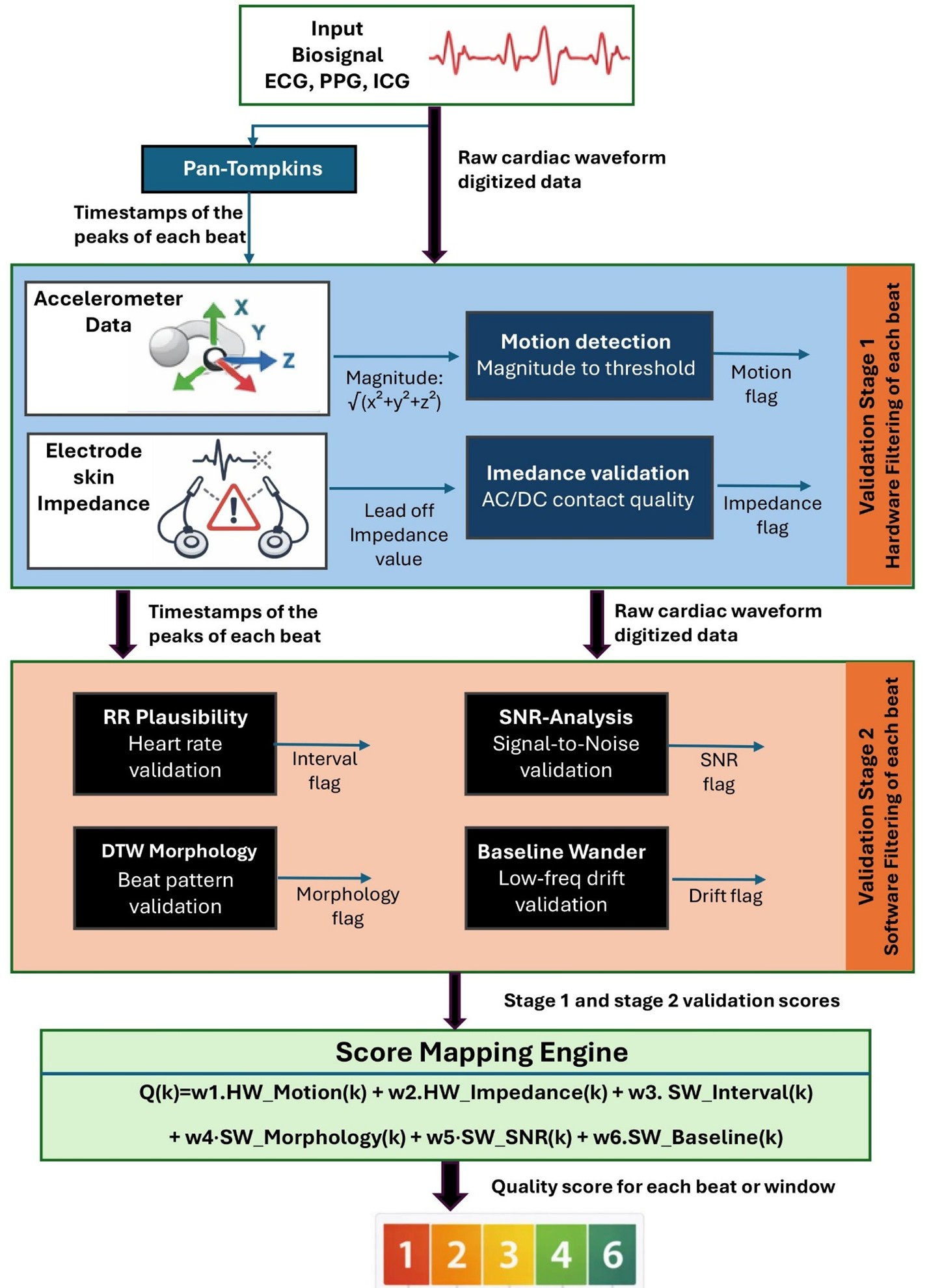
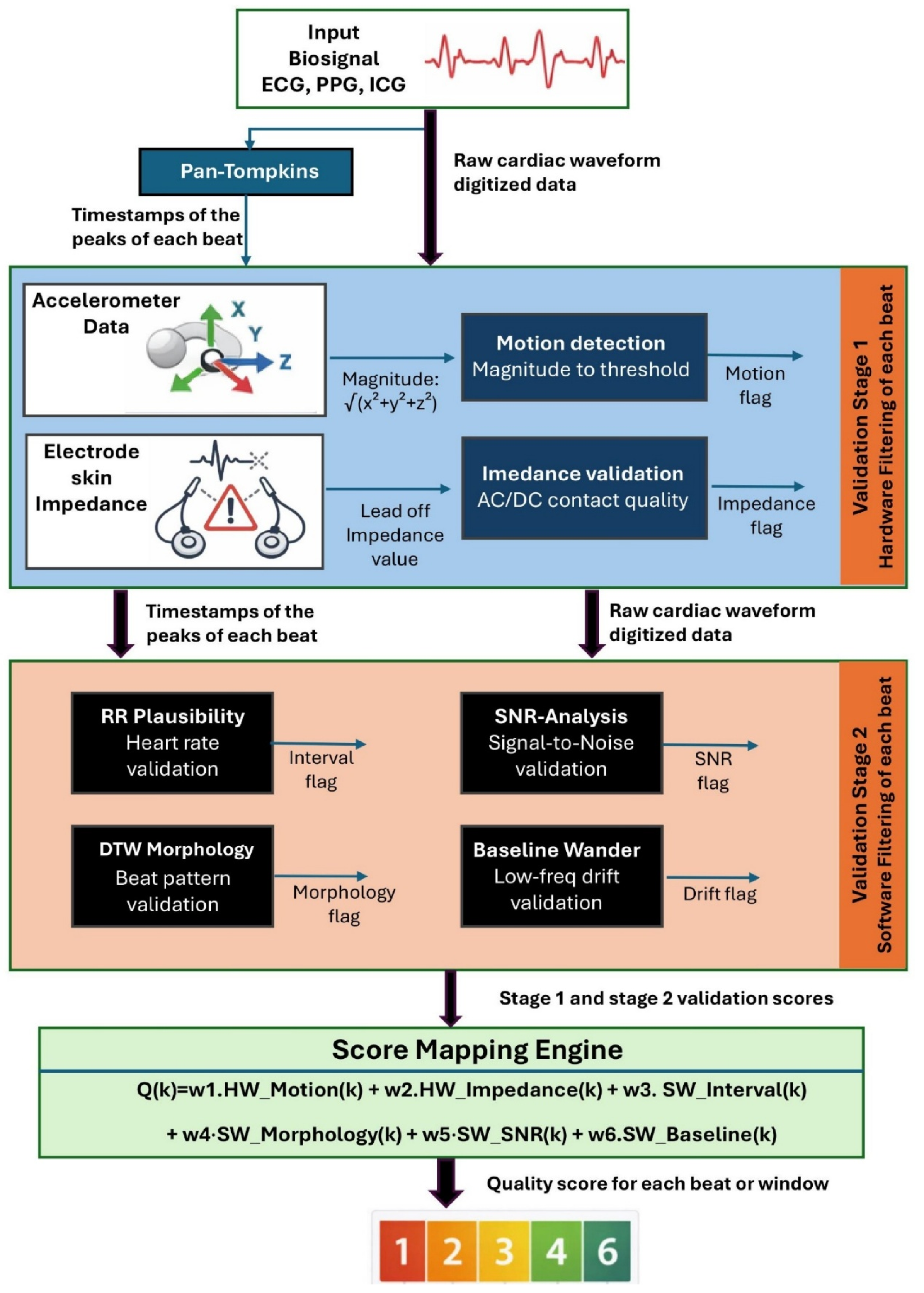
3. Methodology
3.1. Initial Preprocessing
3.2. Stage I: Hardware Based Sensor Integrity Filtering
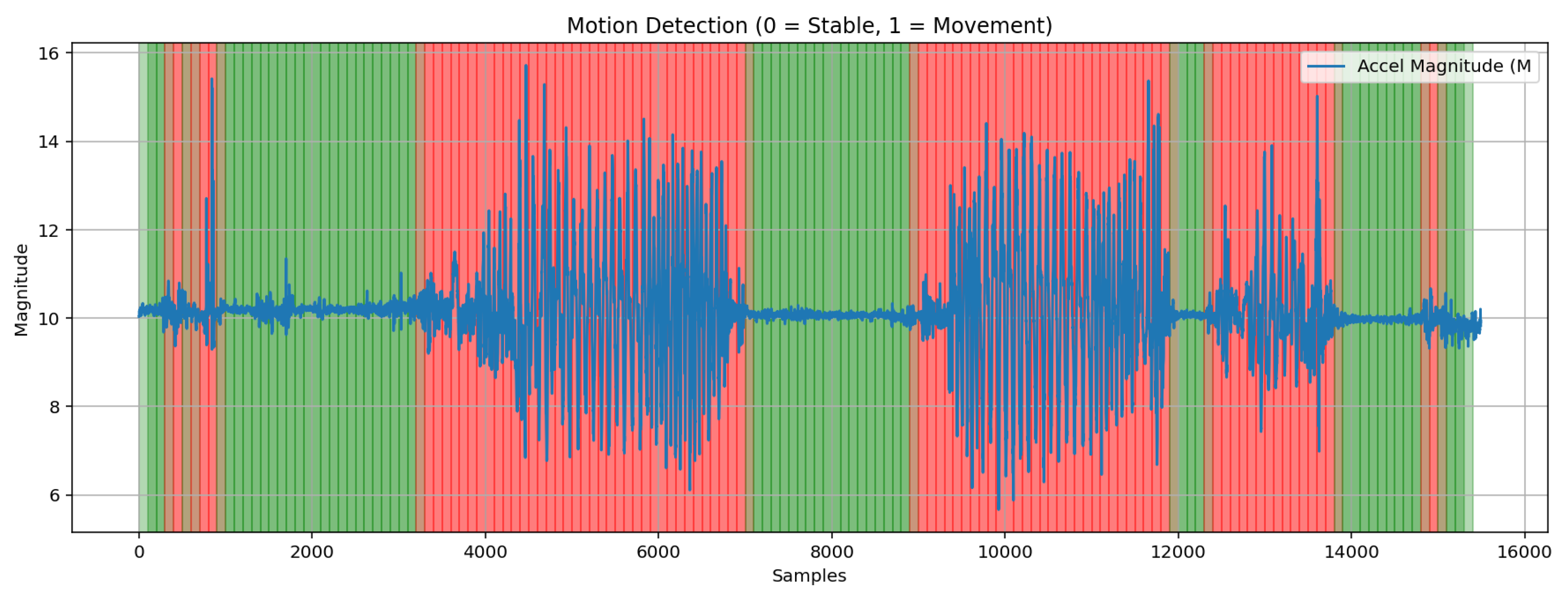
3.2.1. Motion Artifact Detection
3.2.2. Electrode Lead-Off Impedance Verification
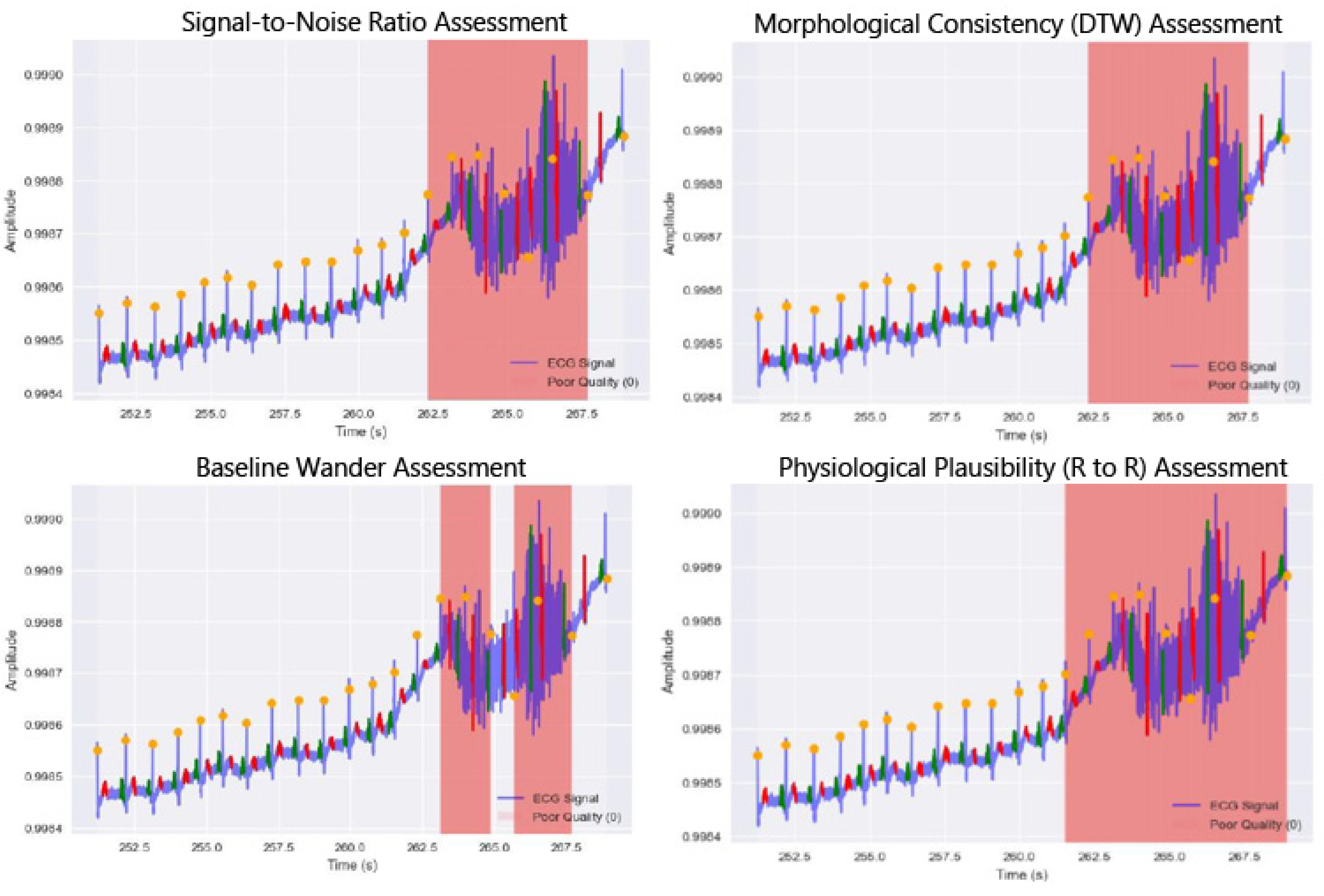
3.3. Stage II: Software-Based Signal Quality Index Evaluation
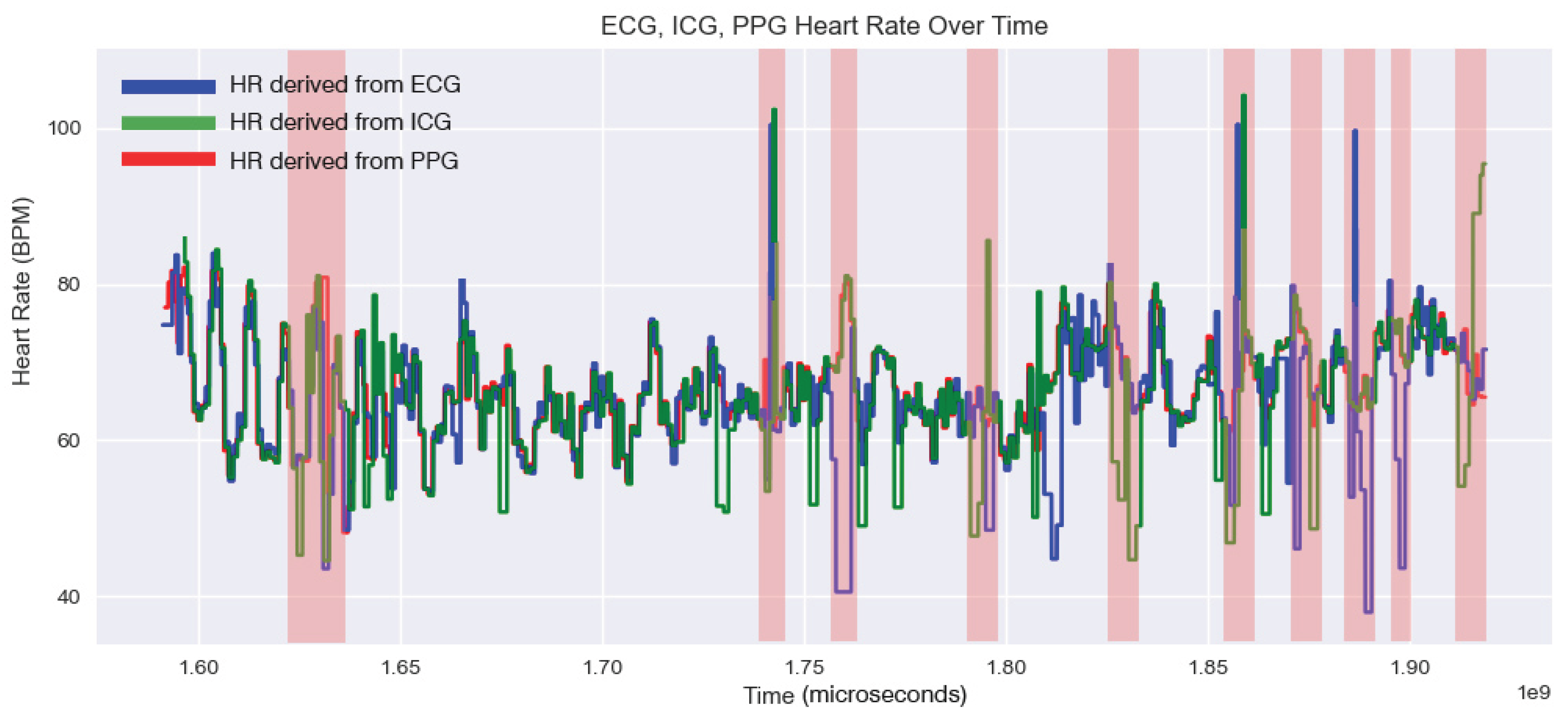
3.3.1. Physiological Plausibility (RR Interval Analysis)
R-Peak Period Stability Assessment
- the RR interval falls outside predefined physiological bounds (e.g., 50bpm to 250bpm), or
- the deviation from the recent R-R median exceeds a relative tolerance threshold.
R-Peak Amplitude Stability Assessment
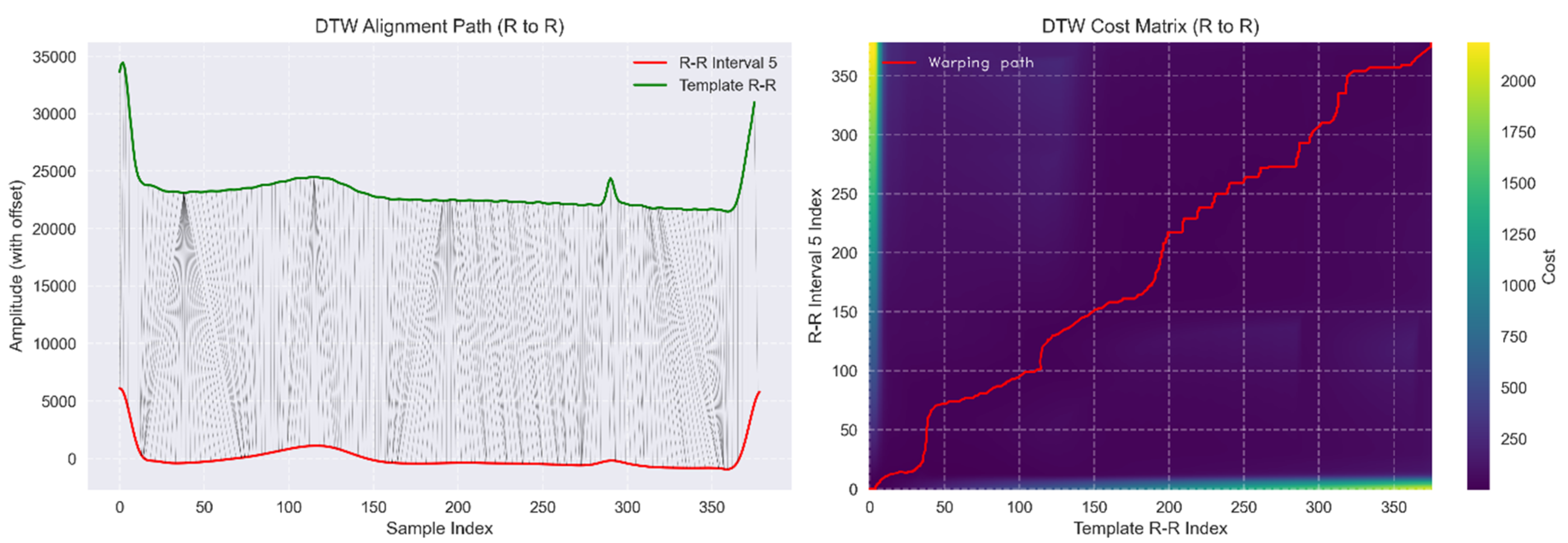
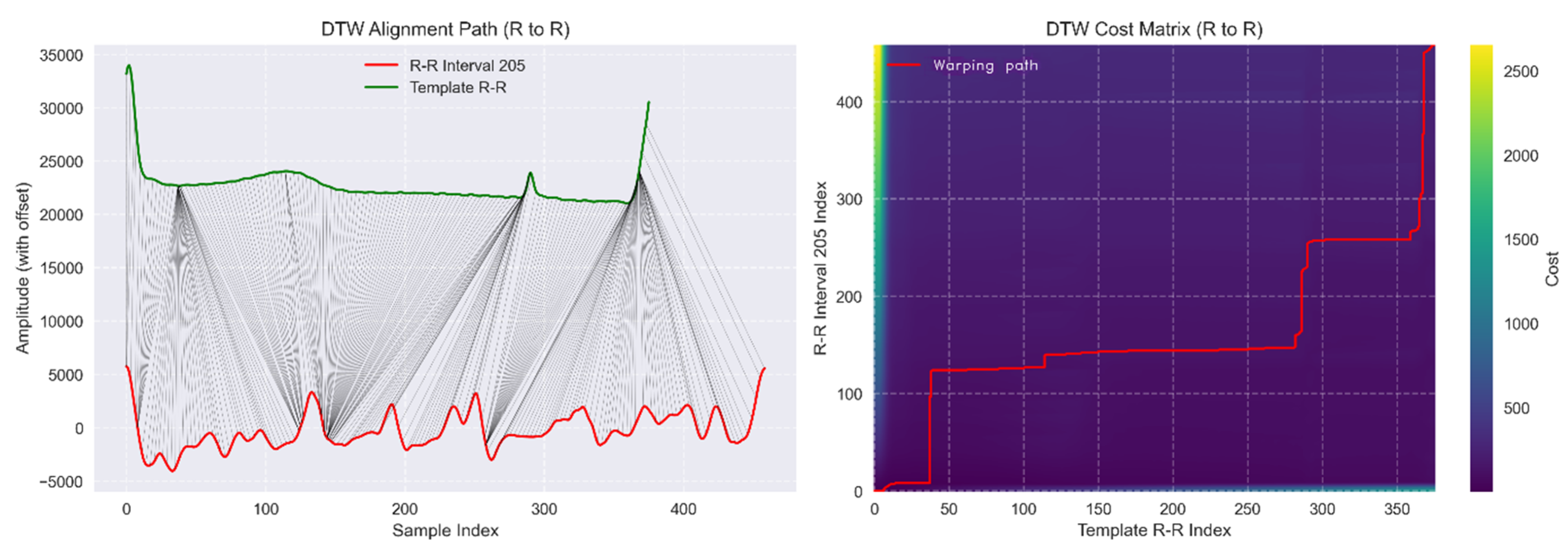
3.3.2. Morphological Consistency Using Multi Directional DTW Pattern Matching
3.3.3. Signal-to-Noise Ratio (SNR)
- 20–50 dB: clinical-grade, high-quality signal
- 10–20 dB: acceptable quality for monitoring
- 0–10 dB: noisy waveform with reduced diagnostic reliability
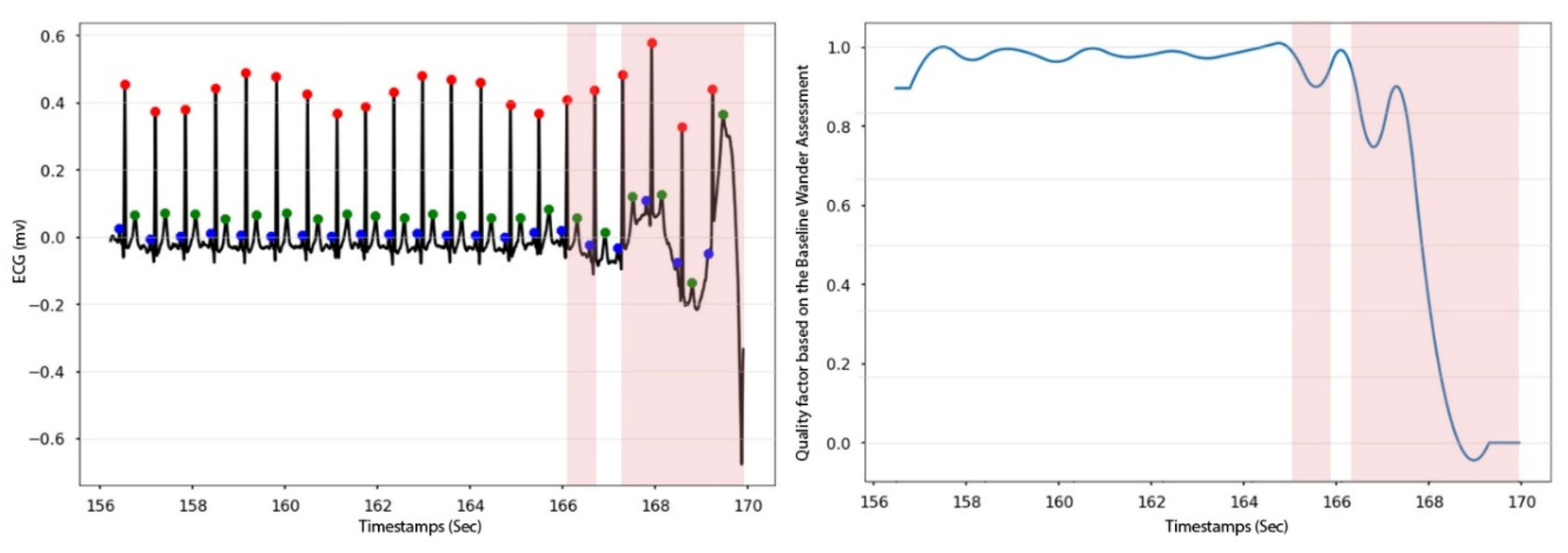
3.3.4. Baseline Wander Assessment
3.4. Global Quality Factor Computation
4. Data Acquisition and Experimental Protocol
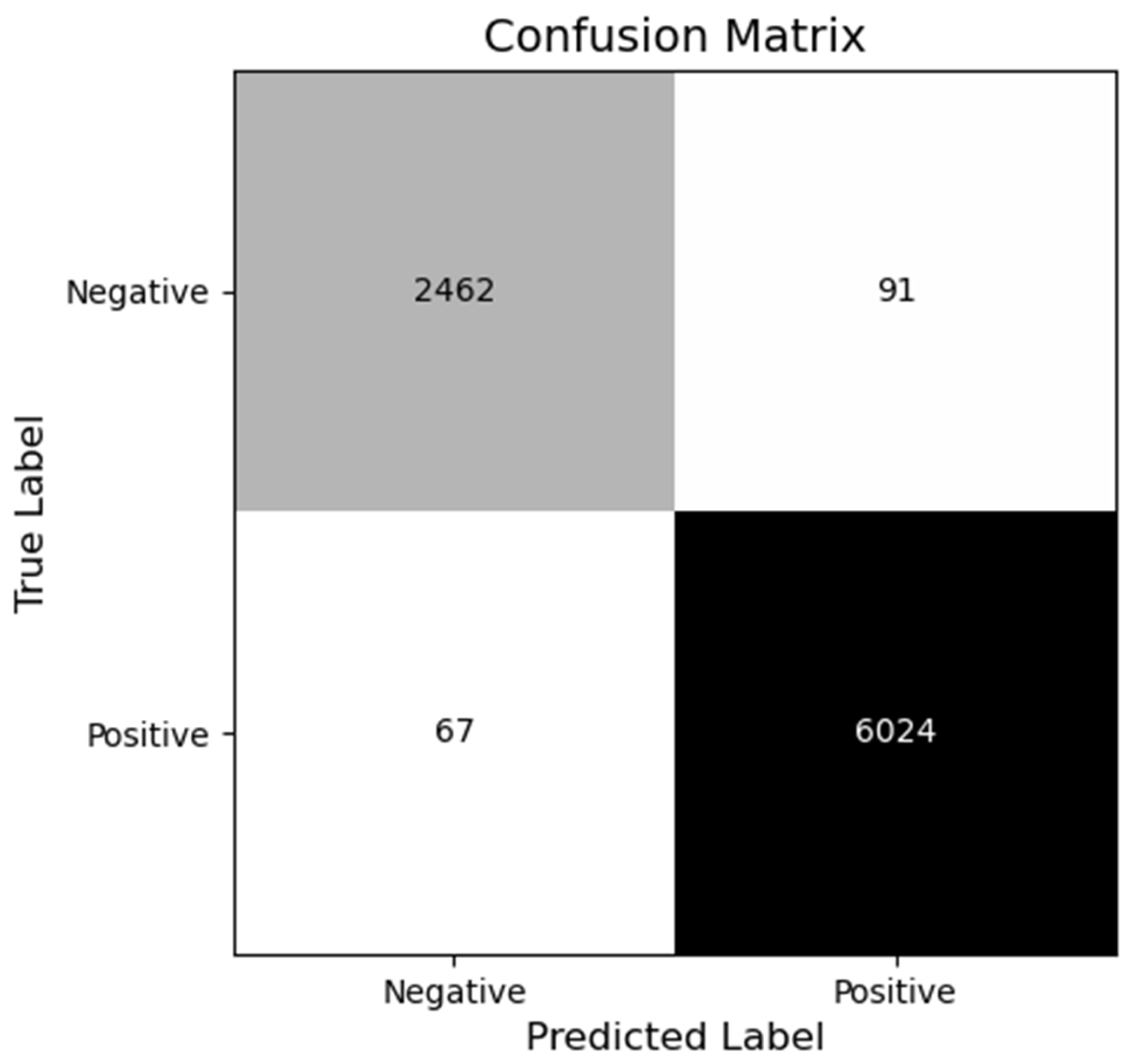
5. Results
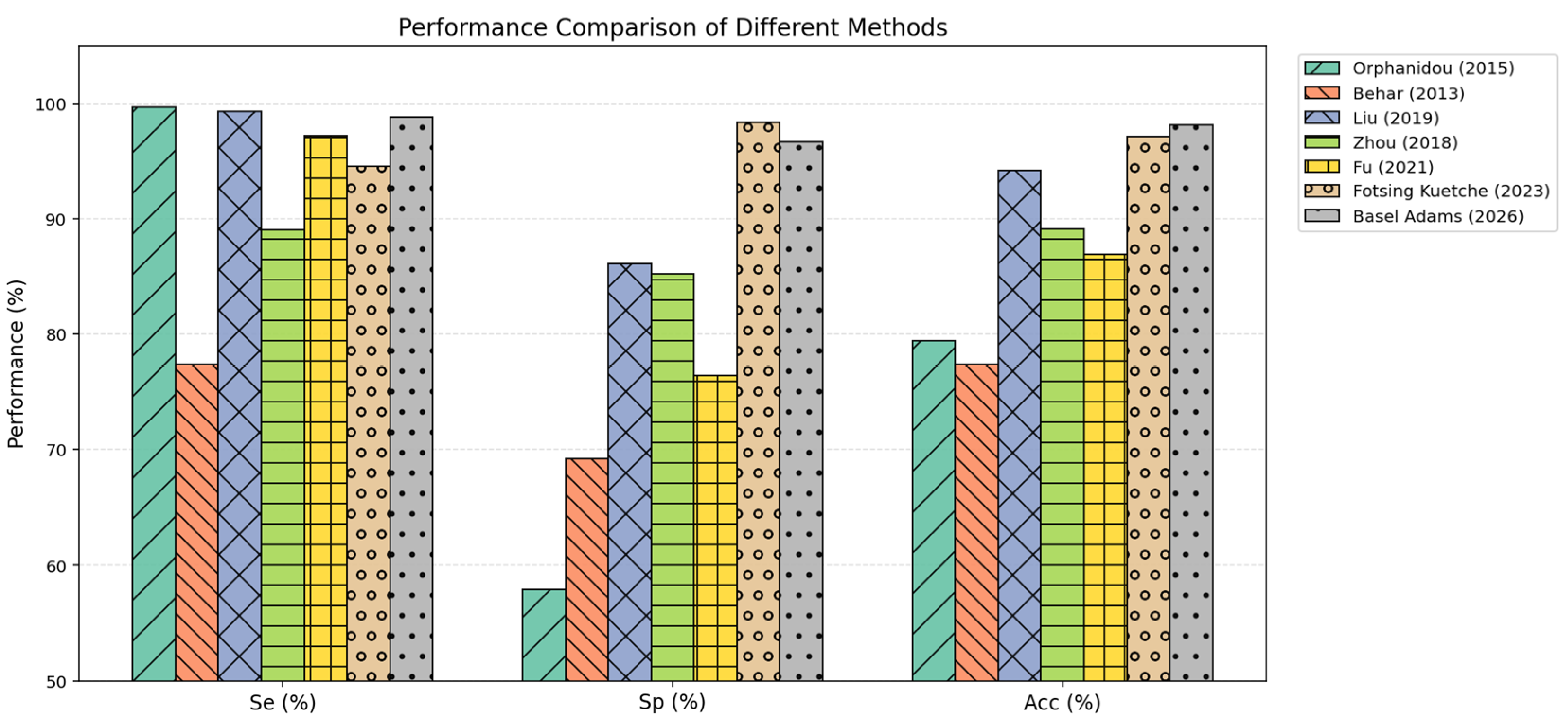
6. Performance Comparison with State-of-the-Art Methods
| Method | Evaluation | Sensitivity (%) | Specificity (%) | Accuracy (%) |
| Orphanidou (2015) | Set of Rules +average template matching | 99.70 | 57.94 | 79.41 |
| Behar (2013) | SVM | 77.34 | 69.20 | 77.37 |
| Liu (2019) | SVM | 99.36 | 86.10 | 94.17 |
| Zhou (2018) | CNN | 89.02 | 85.23 | 89.12 |
| Fu (2021) | LSTM | 97.19 | 76.42 | 86.90 |
| Fotsing Kuetche (2023) | Set of Rules + beats correlation and clustering | 94.59 | 98.38 | 97.10 |
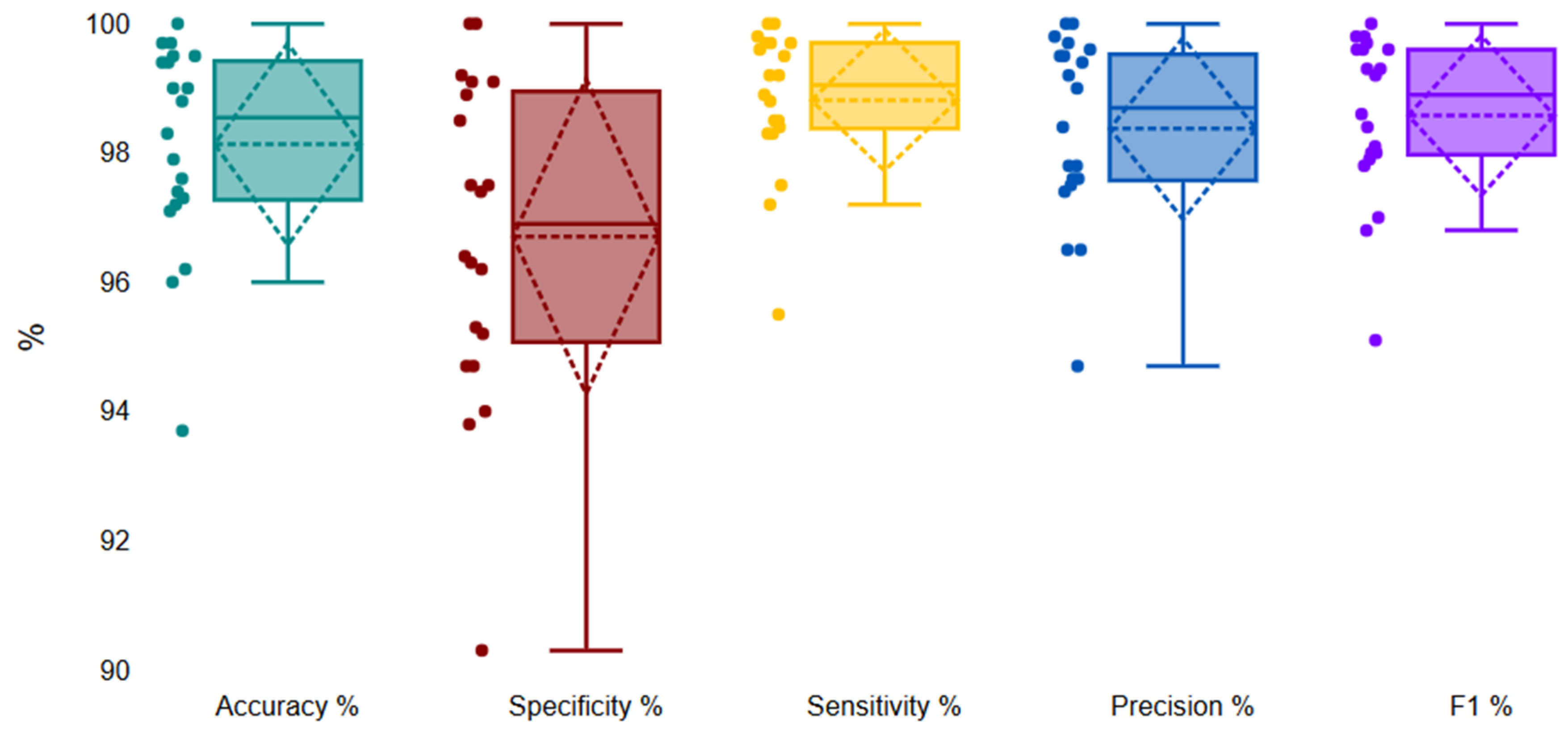
| Basel Adams (2026) | Sensor Fusion and Software Indices | 98.81 | 96.70 | 98.13 |
7. Discussion
8. Conclusions
Institutional Review Board Statement
Acknowledgments
Conflicts of Interest
References
- Adams, B; Dils, C.L. AI-Enhanced Smart Textile System for the Monitoring of Cardiovascular Insufficiency. 2024 IEEE 10th Electronics System-Integration Technology Conference, 2025. [Google Scholar] [CrossRef]
- Poon, C. C.; Zhang, Y. T. Cuff-less and noninvasive measurements of arterial blood pressure by pulse transit time. IEEE Engineering in Medicine and Biology Magazine 2005, 24(5), 20–27. [Google Scholar]
- Tong, D. A.; Bartels, K. A.; Honeyager, K. S. Adaptive reduction of motion artifact in the electrocardiogram. IEEE Transactions on Biomedical Engineering 2002, 49(9), 1085–1088. [Google Scholar]
- Liu, S. H.; Cheng, D. C.; Lin, C. M. Arrhythmia identification with two-lead electrocardiograms using artificial neural networks and support vector machines for a portable ECG monitor system. Sensors 2013, 13(1), 813–828. [Google Scholar] [CrossRef] [PubMed]
- Medical Instrumentation: Application and Design, 4th ed.; Webster, J. G., Ed.; John Wiley & Sons, 2009. [Google Scholar]
- Meziane, N.; Webster, J. G.; Attari, M.; Nimunkar, A. J. Dry electrodes for electrocardiography. Physiological Measurement 2013, 34(9), R47–R69. [Google Scholar] [CrossRef]
- Pan, J.; Tompkins, W. J. A real-time QRS detection algorithm. IEEE Transactions on Biomedical Engineering 1985, BME-32(3), 230–236. [Google Scholar] [CrossRef] [PubMed]
- Clifford, G. D.; Behar, J.; Li, Q.; Rezek, I. Signal quality indices and data fusion for determining clinical acceptability of electrocardiograms. Physiological Measurement 2012, 33(9), 1419–1433. [Google Scholar] [CrossRef] [PubMed]
- Moody, G. B.; Mark, R. G. QRS morphology representation and noise estimation using the Karhunen-Loeve transform. In Computers in Cardiology; 1989; pp. 269–272. [Google Scholar]
- Hayn, D.; Kollmann, A.; Schreier, G. Automated quality evaluation of ECG using morphology-based features. In Computers in Cardiology; 2008; pp. 661–664. [Google Scholar]
- Krasteva, V.; Jekova, I.; Leber, R.; Schmid, R.; Abächerli, R. Real-time arrhythmia detection with supplementary ECG quality and pulse wave monitoring for the reduction of false alarms in ICUs. Physiological Measurement 2016, 37(8), 1273–1297. [Google Scholar] [CrossRef]
- Zhao, Z.; Zhang, Y. SQI quality evaluation mechanism of single-lead ECG signal based on simple statistics. Computer Methods and Programs in Biomedicine 2005, 125, 135–145. [Google Scholar]
- Spicher, N.; Kukuk, M. Delineation of electrocardiograms using multiscale parameter estimation. IEEE Journal of Biomedical and Health Informatics 2015, 19(4), 1224–1234. [Google Scholar] [CrossRef]
- Li, Q.; Mark, R. G.; Clifford, G. D. Robust heart rate estimation from multiple asynchronous noisy sources using signal quality indices and a Kalman filter. Physiological Measurement 2008, 29(1), 15–32. [Google Scholar] [CrossRef] [PubMed]
- Johannesen, L.; Vicente, J.; Gray, R. A.; Galeotti, L.; Loring, Z.; Garnett, C. E.; Strauss, D. G. Improving the assessment of heart rate variability in drug safety studies. Clinical Pharmacology & Therapeutics 2014, 96(3), 372–374. [Google Scholar]
- Li, Q.; Clifford, G. D. Dynamic time warping and machine learning for signal quality assessment of pulsatile signals. Physiological Measurement 2012, 33(9), 1491–1501. [Google Scholar] [CrossRef]
- Orphanidou, C.; Bonnici, T.; Charlton, P.; Clifton, D.; Vallance, D.; Tarassenko, L. Signal-quality indices for the electrocardiogram and photoplethysmogram: Derivation and applications to wireless monitoring. IEEE Journal of Biomedical and Health Informatics 2015, 19(3), 832–838. [Google Scholar] [CrossRef]
- Choi, A.; Shin, H.; Kim, J. Quantifying the physiological response during rest-to-exercise transition using time-varying heart rate variability indices. In Computing in Cardiology; 2011; pp. 613–616. [Google Scholar]
- Gradl, S.; Kugler, P.; Lohmann, C.; Eskofier, B. M. Real-time ECG monitoring and arrhythmia detection using Android-based mobile devices. Annual International Conference of the IEEE Engineering in Medicine and Biology Society, 2012; pp. 2452–2455. [Google Scholar]
- Smital, L.; Vítek, M.; Kozumplík, J.; Provazník, I. Adaptive wavelet Wiener filtering of ECG signals. IEEE Transactions on Biomedical Engineering 2013, 60(2), 437–445. [Google Scholar] [CrossRef]
- Satija, U.; Ramkumar, B.; Manikandan, M. S. A review of signal processing techniques for electrocardiogram signal quality assessment. IEEE Reviews in Biomedical Engineering 2018, 11, 36–52. [Google Scholar] [CrossRef] [PubMed]
- Elgendi, M. Optimal signal quality index for photoplethysmogram signals. Bioengineering 2016, 3(4), 21. [Google Scholar] [CrossRef]
- Sukor, J. A.; Redmond, S. J.; Lovell, N. H. Signal quality measures for pulse oximetry through waveform morphology analysis. Physiological Measurement 2011, 32(3), 369–384. [Google Scholar] [CrossRef] [PubMed]
- Sherwood, A.; Allen, M. T.; Fahrenberg, J.; Kelsey, R. M.; Lovallo, W. R.; Van Doornen, L. J. Methodological guidelines for impedance cardiography. Psychophysiology 1990, 27(1), 1–23. [Google Scholar] [CrossRef]
- Springer, D. B.; Tarassenko, L.; Clifford, G. D. Logistic regression-HSMM-based heart sound segmentation. IEEE Transactions on Biomedical Engineering 2016, 63(4), 822–832. [Google Scholar] [CrossRef] [PubMed]
- Clancy, E. A.; Morin, E. L.; Merletti, R. Sampling, noise-reduction and amplitude estimation issues in surface electromyography. Journal of Electromyography and Kinesiology 2002, 12(1), 1–16. [Google Scholar] [CrossRef]
- Li, Q.; Clifford, G. D. Dynamic time warping and machine learning for signal quality assessment of pulsatile signals. Physiological Measurement 2012, vol. 33(no. 9), 1491–1501. [Google Scholar] [CrossRef] [PubMed]
|
Accuracy Metric |
Mean (%) |
± SD (%) |
Min (%) |
Max (%) |
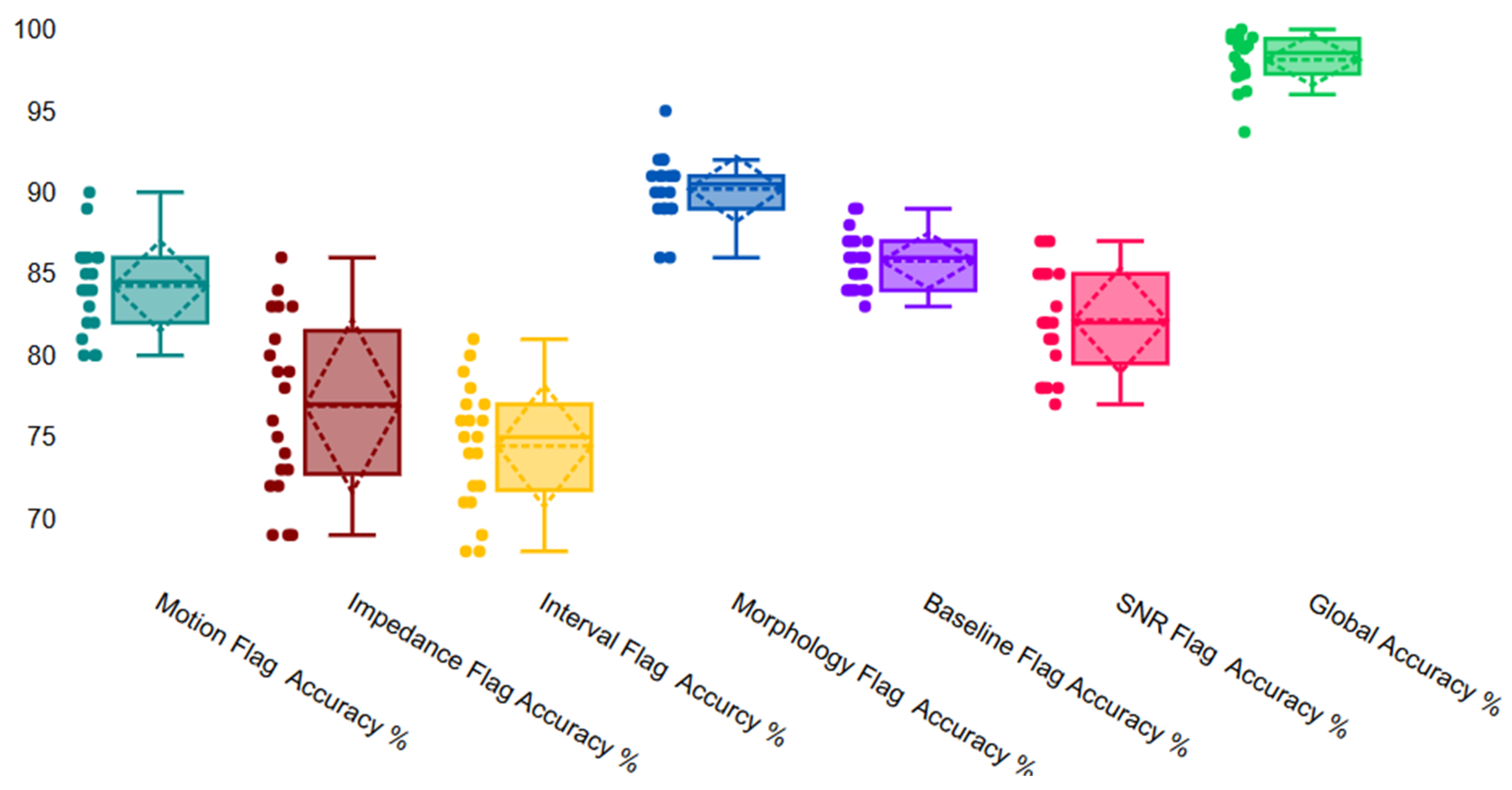
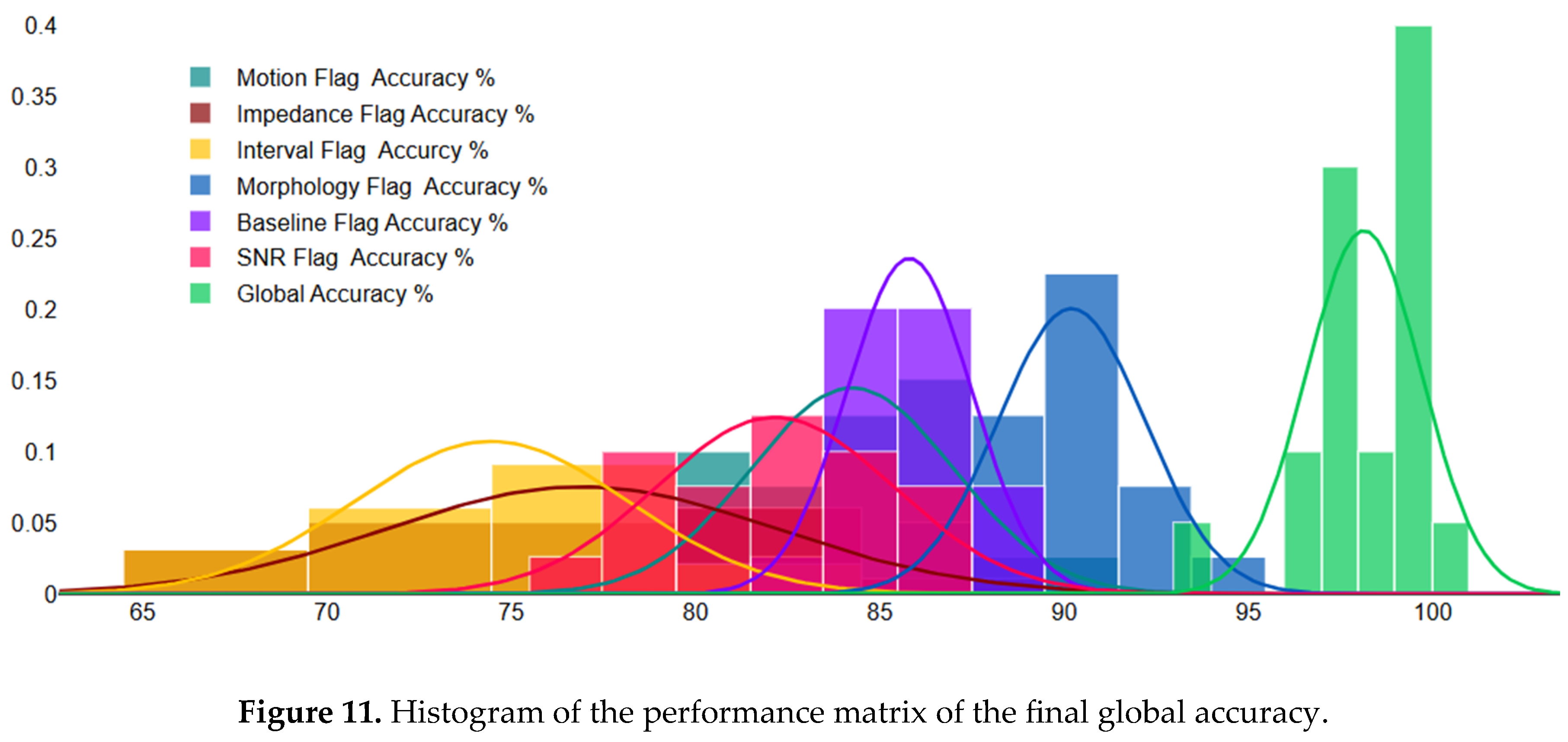
| Motion Flag | 84.4 | 2.7 | 80 | 90 |
| Impedance Flag | 77.7 | 5.3 | 69 | 86 |
| Interval Flag | 74.8 | 3.8 | 68 | 81 |
| Morphology Flag | 90.4 | 2.2 | 86 | 95 |
| SNR Flag | 82.2 | 3.2 | 77 | 87 |
| Baseline Flag | 85.9 | 1.9 | 83 | 89 |
| Global Validation | 98.1 | 1.5 | 93.6 | 100 |
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2026 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).




