Submitted:
04 April 2026
Posted:
07 April 2026
You are already at the latest version
Abstract
Keywords:
1. Introduction: Mitochondrial Function and Health
2. Skeletal Muscle and Mitochondrial Quality Control
3. A Brief History of Mitochondrial Stress Response
4. The Integrated Stress Response (ISR): A Conserved Cellular Sentinel
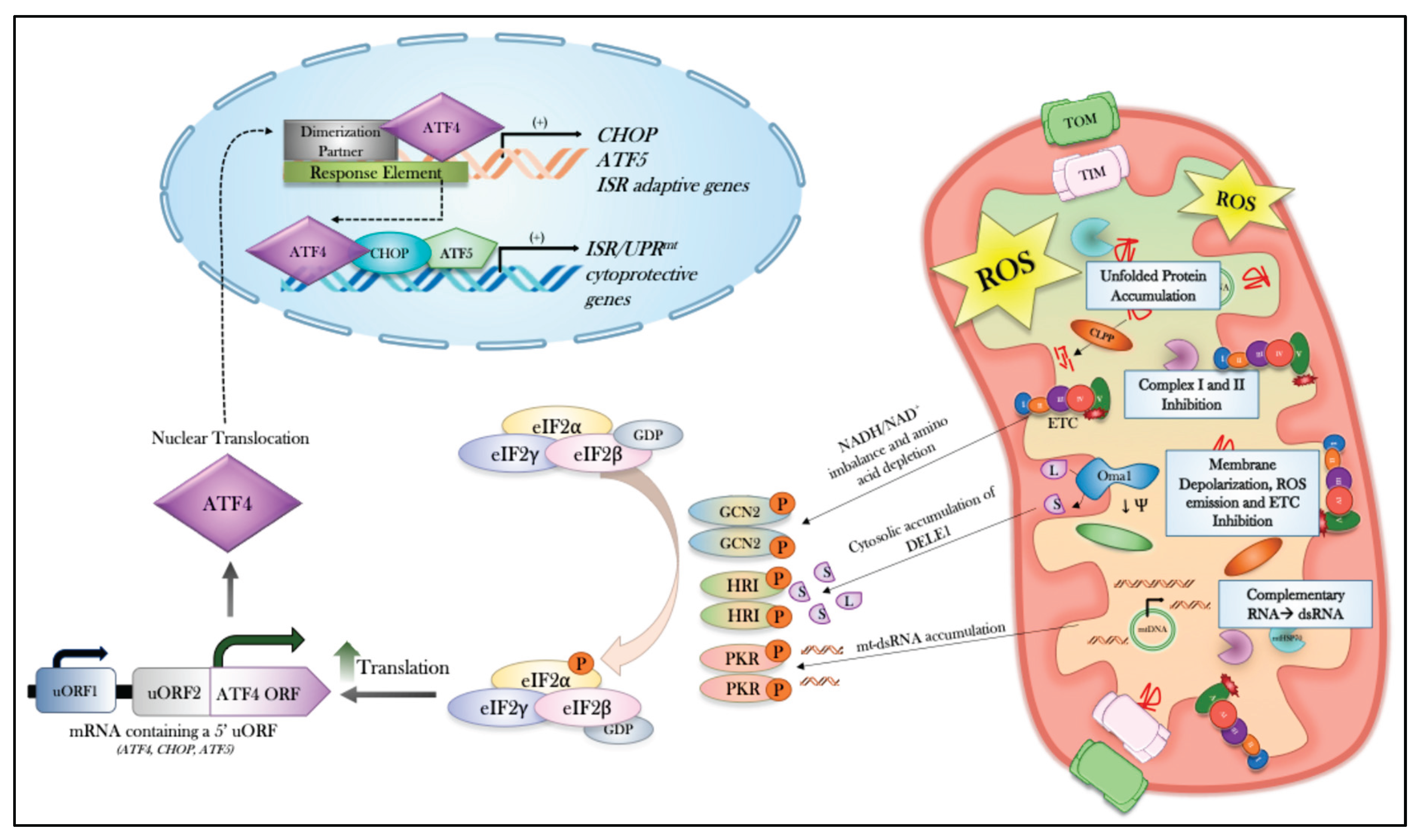
4.1. General Activation and Regulation of the ISR
4.2. Mitochondrial Stress-Specific Activation of the ISR
4.2.1. HRI and Mitochondrial Stress
4.2.2. GCN2 and Mitochondrial Stress
4.2.3. PKR and Mitochondrial Stress
4.2.4. Kinase-Independent Activation of ISR
4.3. Downstream Consequences of ISR Activation: Mitochondrial Stress Resolution
4.4. Cell Guardian or Executioner? The ISR as a Double-Edge Sword
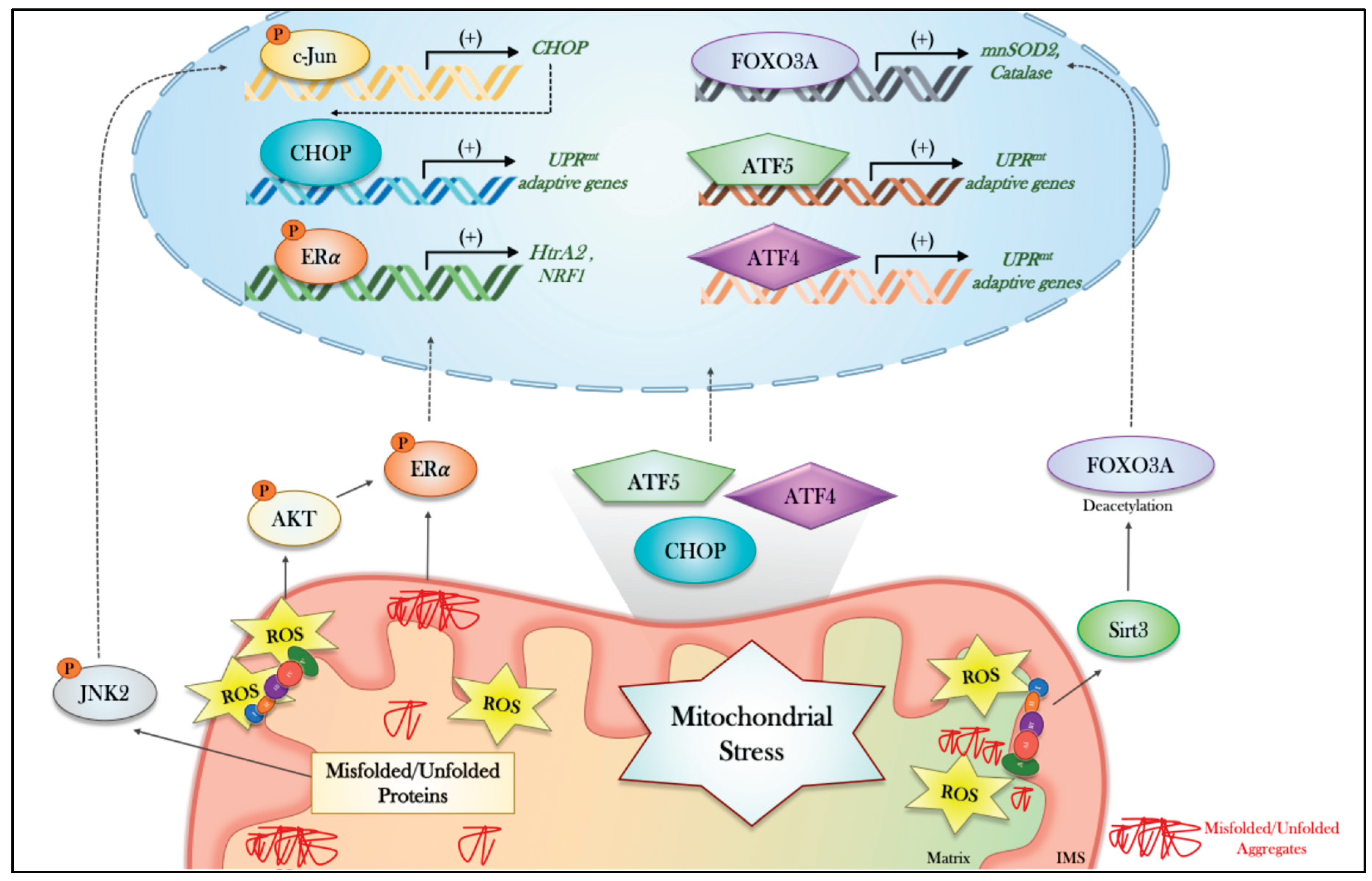
4.5. The UPRᵐᵗ in Mammalian Cells
4.6. ISR and UPRmt Interconnection
5. The Mitochondrial Stress Response in Skeletal Muscle
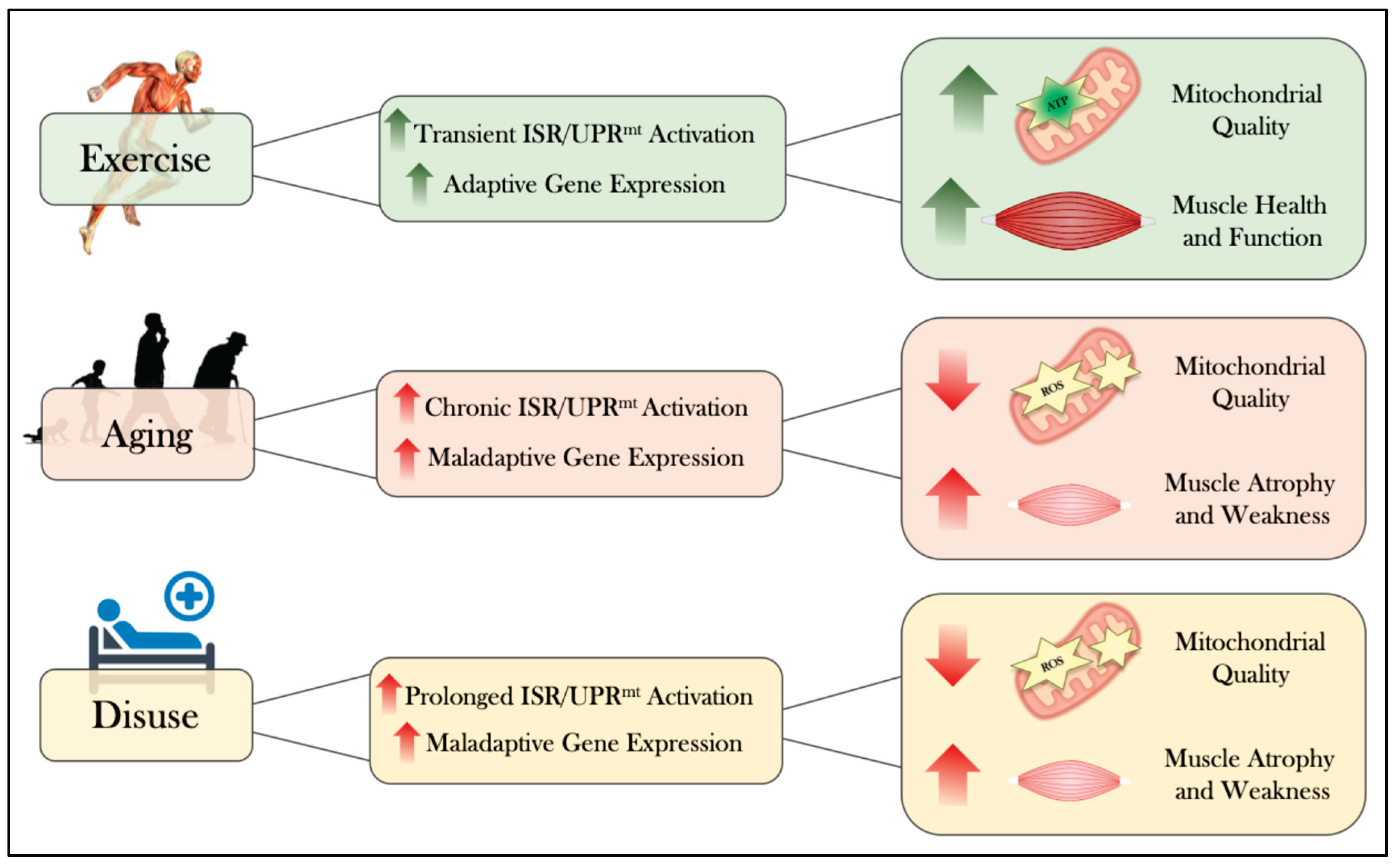
5.1. Activation of the ISR/UPRmt During Acute Exercise
5.2. ISR/UPRmt Mediated Adaptations Following Chronic Exercise
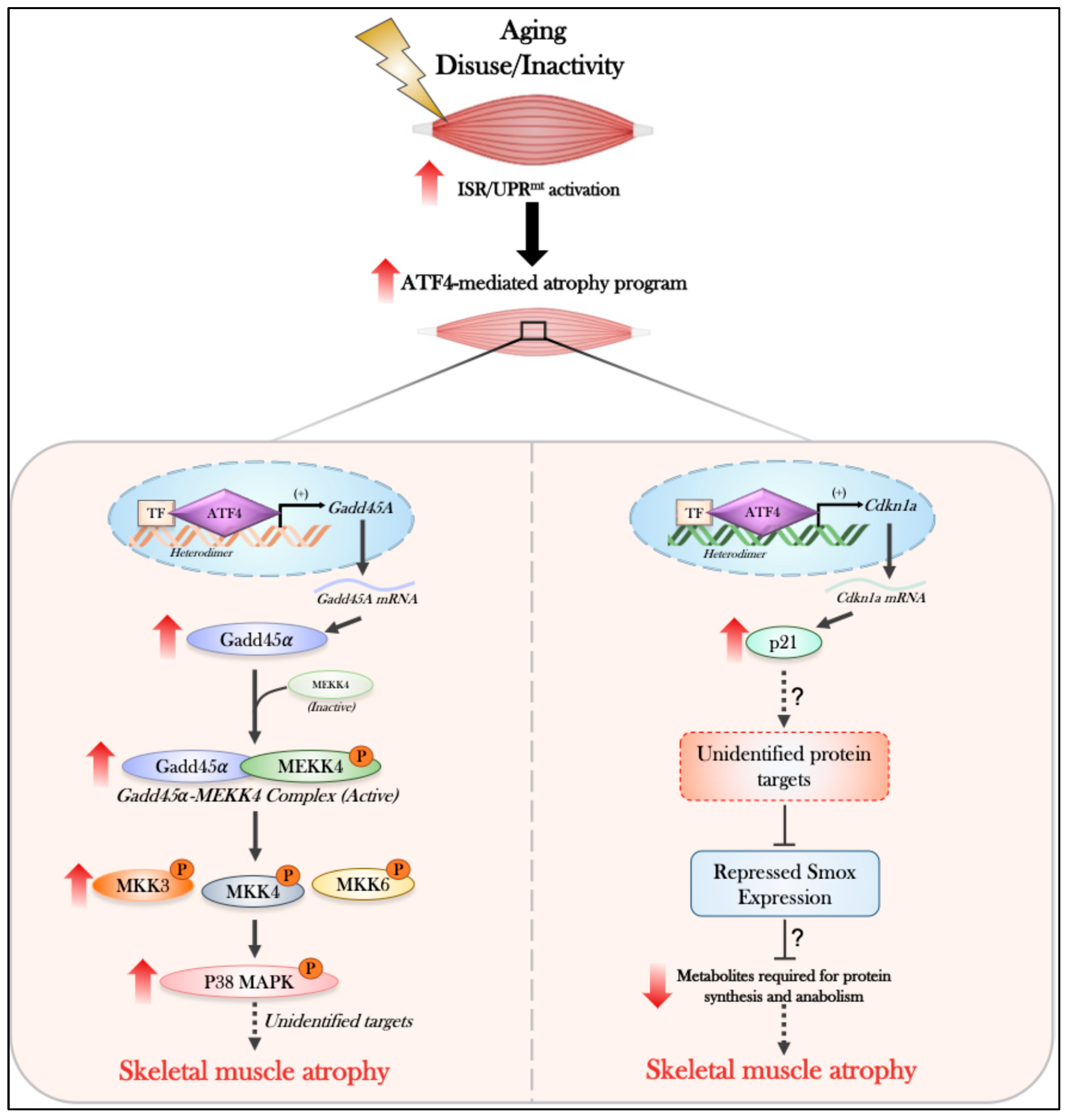
6. The Intersection of Aging and the ISR/UPRmt
7. The Activation of the ISR/UPRmt During Muscle Inactivity
8. Modulation of the Mitochondrial Stress Responses to Improve Muscle Health
8.1. Direct Pharmacological Targeting of the ISR/UPRmt
8.2. Exercise as Medicine
9. Conclusion and Future Perspectives
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Conflicts of Interest
References
- Martin, W.F.; Garg, S.; Zimorski, V. Endosymbiotic Theories for Eukaryote Origin. Philos. Trans. R. Soc. B Biol. Sci. 2015, 370, 20140330. [Google Scholar] [CrossRef]
- Roger, A.J.; Muñoz-Gómez, S.A.; Kamikawa, R. The Origin and Diversification of Mitochondria. Curr. Biol. 2017, 27, R1177–R1192. [Google Scholar] [CrossRef]
- Ernster, L.; Schatz, G. Mitochondria: A Historical Review. J. Cell Biol. 1981, 91, 227s–255s. [Google Scholar] [CrossRef]
- Greaves, L.C.; Reeve, A.K.; Taylor, R.W.; Turnbull, D.M. Mitochondrial DNA and Disease. J. Pathol. 2012, 226, 274–286. [Google Scholar] [CrossRef] [PubMed]
- Osellame, L.D.; Blacker, T.S.; Duchen, M.R. Cellular and Molecular Mechanisms of Mitochondrial Function. Best Pract. Res. Clin. Endocrinol. Metab. 2012, 26, 711–723. [Google Scholar] [CrossRef]
- Quirós, P.M.; Mottis, A.; Auwerx, J. Mitonuclear Communication in Homeostasis and Stress. Nat. Rev. Mol. Cell Biol. 2016, 17, 213–226. [Google Scholar] [CrossRef]
- Frontera, W.R.; Ochala, J. Skeletal Muscle: A Brief Review of Structure and Function. Calcif. Tissue Int. 2015, 96, 183–195. [Google Scholar] [CrossRef] [PubMed]
- Hood, D.A.; Irrcher, I.; Ljubicic, V.; Joseph, A.-M. Coordination of Metabolic Plasticity in Skeletal Muscle. J. Exp. Biol. 2006, 209, 2265–2275. [Google Scholar] [CrossRef] [PubMed]
- Bo, H.; Zhang, Y.; Ji, L.L. Redefining the Role of Mitochondria in Exercise: A Dynamic Remodeling. Ann. N. Y. Acad. Sci. 2010, 1201, 121–128. [Google Scholar] [CrossRef] [PubMed]
- Drake, J.C.; Wilson, R.J.; Yan, Z. Molecular Mechanisms for Mitochondrial Adaptation to Exercise Training in Skeletal Muscle. FASEB J. 2016, 30, 13–22. [Google Scholar] [CrossRef]
- Kotiadis, V.N.; Duchen, M.R.; Osellame, L.D. Mitochondrial Quality Control and Communications with the Nucleus Are Important in Maintaining Mitochondrial Function and Cell Health. Biochim. Biophys. Acta BBA - Gen. Subj. 2014, 1840, 1254–1265. [Google Scholar] [CrossRef]
- Roca-Portoles, A.; Tait, S.W.G. Mitochondrial Quality Control: From Molecule to Organelle. Cell. Mol. Life Sci. 2021, 78, 3853–3866. [Google Scholar] [CrossRef]
- Eckl, E.-M.; Ziegemann, O.; Krumwiede, L.; Fessler, E.; Jae, L.T. Sensing, Signaling and Surviving Mitochondrial Stress. Cell. Mol. Life Sci. 2021, 78, 5925–5951. [Google Scholar] [CrossRef] [PubMed]
- Pakos-Zebrucka, K.; Koryga, I.; Mnich, K.; Ljujic, M.; Samali, A.; Gorman, A.M. The Integrated Stress Response. EMBO Rep. 2016, 17, 1374–1395. [Google Scholar] [CrossRef]
- Anderson, N.S.; Haynes, C.M. Folding the Mitochondrial UPR into the Integrated Stress Response. Trends Cell Biol. 2020, 30, 428–439. [Google Scholar] [CrossRef]
- Anderson, N.S.; Haynes, C.M. Folding the Mitochondrial UPR into the Integrated Stress Response. Trends Cell Biol. 2020, 30, 428–439. [Google Scholar] [CrossRef]
- Hood, D.A.; Memme, J.M.; Oliveira, A.N.; Triolo, M. Maintenance of Skeletal Muscle Mitochondria in Health, Exercise, and Aging. Annu. Rev. Physiol. 2019, 81, 19–41. [Google Scholar] [CrossRef] [PubMed]
- Joseph, A.; Adhihetty, P.J.; Buford, T.W.; Wohlgemuth, S.E.; Lees, H.A.; Nguyen, L.M. -D.; Aranda, J.M.; Sandesara, B.D.; Pahor, M.; Manini, T.M.; et al. The Impact of Aging on Mitochondrial Function and Biogenesis Pathways in Skeletal Muscle of Sedentary High- and Low-functioning Elderly Individuals. Aging Cell 2012, 11, 801–809. [Google Scholar] [CrossRef] [PubMed]
- Parikh, V.S.; Morgan, M.M.; Scott, R.; Clements, L.S.; Butow, R.A. The Mitochondrial Genotype Can Influence Nuclear Gene Expression in Yeast. Science 1987, 235, 576–580. [Google Scholar] [CrossRef] [PubMed]
- Liu, Z.; Butow, R.A. Mitochondrial Retrograde Signaling. Annu. Rev. Genet. 2006, 40, 159–185. [Google Scholar] [CrossRef]
- Martinus, R.D.; Garth, G.P.; Webster, T.L.; Cartwright, P.; Naylor, D.J.; Høj, P.B.; Hoogenraad, N.J. Selective Induction of Mitochondrial Chaperones in Response to Loss of the Mitochondrial Genome. Eur. J. Biochem. 1996, 240, 98–103. [Google Scholar] [CrossRef]
- Zhao, Q. A Mitochondrial Specific Stress Response in Mammalian Cells. EMBO J. 2002, 21, 4411–4419. [Google Scholar] [CrossRef]
- Cortopassi, G.; Danielson, S.; Alemi, M.; Zhan, S.S.; Tong, W.; Carelli, V.; Martinuzzi, A.; Marzuki, S.; Majamaa, K.; Wong, A. Mitochondrial Disease Activates Transcripts of the Unfolded Protein Response and Cell Cycle and Inhibits Vesicular Secretion and Oligodendrocyte-Specific Transcripts. Mitochondrion 2006, 6, 161–175. [Google Scholar] [CrossRef]
- Costa-Mattioli, M.; Walter, P. The Integrated Stress Response: From Mechanism to Disease. Science 2020, 368, eaat5314. [Google Scholar] [CrossRef] [PubMed]
- Mick, E.; Titov, D.V.; Skinner, O.S.; Sharma, R.; Jourdain, A.A.; Mootha, V.K. Distinct Mitochondrial Defects Trigger the Integrated Stress Response Depending on the Metabolic State of the Cell. eLife 2020, 9, e49178. [Google Scholar] [CrossRef] [PubMed]
- Donnelly, N.; Gorman, A.M.; Gupta, S.; Samali, A. The eIF2α Kinases: Their Structures and Functions. Cell. Mol. Life Sci. 2013, 70, 3493–3511. [Google Scholar] [CrossRef]
- Kimball, S.R. Eukaryotic Initiation Factor eIF2. Int. J. Biochem. Cell Biol. 1999, 31, 25–29. [Google Scholar] [CrossRef]
- Dever, T.E. Gene-Specific Regulation by General Translation Factors. Cell 2002, 108, 545–556. [Google Scholar] [CrossRef] [PubMed]
- Wek, R.C. Role of eIF2α Kinases in Translational Control and Adaptation to Cellular Stress. Cold Spring Harb. Perspect. Biol. 2018, 10, a032870. [Google Scholar] [CrossRef] [PubMed]
- Harding, H.P.; Novoa, I.; Zhang, Y.; Zeng, H.; Wek, R.; Schapira, M.; Ron, D. Regulated Translation Initiation Controls Stress-Induced Gene Expression in Mammalian Cells. Mol. Cell 2000, 6, 1099–1108. [Google Scholar] [CrossRef]
- Algire, M.A.; Maag, D.; Lorsch, J.R. Pi Release from eIF2, Not GTP Hydrolysis, Is the Step Controlled by Start-Site Selection during Eukaryotic Translation Initiation. Mol. Cell 2005, 20, 251–262. [Google Scholar] [CrossRef] [PubMed]
- Jackson, R.J.; Hellen, C.U.T.; Pestova, T.V. The Mechanism of Eukaryotic Translation Initiation and Principles of Its Regulation. Nat. Rev. Mol. Cell Biol. 2010, 11, 113–127. [Google Scholar] [CrossRef]
- Sonenberg, N.; Hinnebusch, A.G. Regulation of Translation Initiation in Eukaryotes: Mechanisms and Biological Targets. Cell 2009, 136, 731–745. [Google Scholar] [CrossRef]
- Pavitt, G.D. Regulation of Translation Initiation Factor eIF2B at the Hub of the Integrated Stress Response. WIREs RNA 2018, 9, e1491. [Google Scholar] [CrossRef] [PubMed]
- Sidrauski, C.; Tsai, J.C.; Kampmann, M.; Hearn, B.R.; Vedantham, P.; Jaishankar, P.; Sokabe, M.; Mendez, A.S.; Newton, B.W.; Tang, E.L.; et al. Pharmacological Dimerization and Activation of the Exchange Factor eIF2B Antagonizes the Integrated Stress Response. eLife 2015, 4, e07314. [Google Scholar] [CrossRef]
- Vattem, K.M.; Wek, R.C. Reinitiation Involving Upstream ORFs Regulates ATF4 mRNA Translation in Mammalian Cells. Proc. Natl. Acad. Sci. 2004, 101, 11269–11274. [Google Scholar] [CrossRef]
- Jousse, C. Inhibition of CHOP Translation by a Peptide Encoded by an Open Reading Frame Localized in the Chop 5’UTR. Nucleic Acids Res. 2001, 29, 4341–4351. [Google Scholar] [CrossRef]
- Watatani, Y.; Ichikawa, K.; Nakanishi, N.; Fujimoto, M.; Takeda, H.; Kimura, N.; Hirose, H.; Takahashi, S.; Takahashi, Y. Stress-Induced Translation of ATF5 mRNA Is Regulated by the 5′-Untranslated Region. J. Biol. Chem. 2008, 283, 2543–2553. [Google Scholar] [CrossRef]
- Young, S.K.; Wek, R.C. Upstream Open Reading Frames Differentially Regulate Gene-Specific Translation in the Integrated Stress Response. J. Biol. Chem. 2016, 291, 16927–16935. [Google Scholar] [CrossRef] [PubMed]
- Koncha, R.R.; Ramachandran, G.; Sepuri, N.B.V.; Ramaiah, K.V.A. CCCP-induced Mitochondrial Dysfunction – Characterization and Analysis of Integrated Stress Response to Cellular Signaling and Homeostasis. FEBS J. 2021, 288, 5737–5754. [Google Scholar] [CrossRef]
- Cohen, P.T.W. Protein Phosphatase 1 – Targeted in Many Directions. J. Cell Sci. 2002, 115, 241–256. [Google Scholar] [CrossRef]
- Novoa, I.; Zeng, H.; Harding, H.P.; Ron, D. Feedback Inhibition of the Unfolded Protein Response by GADD34 -Mediated Dephosphorylation of eIF2α. J. Cell Biol. 2001, 153, 1011–1022. [Google Scholar] [CrossRef]
- Jousse, C.; Oyadomari, S.; Novoa, I.; Lu, P.; Zhang, Y.; Harding, H.P.; Ron, D. Inhibition of a Constitutive Translation Initiation Factor 2α Phosphatase, CReP, Promotes Survival of Stressed Cells. J. Cell Biol. 2003, 163, 767–775. [Google Scholar] [CrossRef] [PubMed]
- Chung, H.K.; Ryu, D.; Kim, K.S.; Chang, J.Y.; Kim, Y.K.; Yi, H.-S.; Kang, S.G.; Choi, M.J.; Lee, S.E.; Jung, S.-B.; et al. Growth Differentiation Factor 15 Is a Myomitokine Governing Systemic Energy Homeostasis. J. Cell Biol. 2017, 216, 149–165. [Google Scholar] [CrossRef]
- Durieux, J.; Wolff, S.; Dillin, A. The Cell-Non-Autonomous Nature of Electron Transport Chain-Mediated Longevity. Cell 2011, 144, 79–91. [Google Scholar] [CrossRef] [PubMed]
- Forsström, S.; Jackson, C.B.; Carroll, C.J.; Kuronen, M.; Pirinen, E.; Pradhan, S.; Marmyleva, A.; Auranen, M.; Kleine, I.-M.; Khan, N.A.; et al. Fibroblast Growth Factor 21 Drives Dynamics of Local and Systemic Stress Responses in Mitochondrial Myopathy with mtDNA Deletions. Cell Metab. 2019, 30, 1040–1054.e7. [Google Scholar] [CrossRef]
- Khan, N.A.; Nikkanen, J.; Yatsuga, S.; Jackson, C.; Wang, L.; Pradhan, S.; Kivelä, R.; Pessia, A.; Velagapudi, V.; Suomalainen, A. mTORC1 Regulates Mitochondrial Integrated Stress Response and Mitochondrial Myopathy Progression. Cell Metab. 2017, 26, 419–428.e5. [Google Scholar] [CrossRef] [PubMed]
- Quirós, P.M.; Prado, M.A.; Zamboni, N.; D’Amico, D.; Williams, R.W.; Finley, D.; Gygi, S.P.; Auwerx, J. Multi-Omics Analysis Identifies ATF4 as a Key Regulator of the Mitochondrial Stress Response in Mammals. J. Cell Biol. 2017, 216, 2027–2045. [Google Scholar] [CrossRef]
- Baron, K.R.; Oviedo, S.; Krasny, S.; Zaman, M.; Aldakhlallah, R.; Bora, P.; Mathur, P.; Pfeffer, G.; Bollong, M.J.; Shutt, T.E.; et al. Pharmacologic Activation of Integrated Stress Response Kinases Inhibits Pathologic Mitochondrial Fragmentation 2025.
- Sánchez-Vera, I.; Núñez-Vázquez, S.; Saura-Esteller, J.; Cosialls, A.M.; Heib, J.; Nadal Rodríguez, P.; Ghashghaei, O.; Lavilla, R.; Pons, G.; Gil, J.; et al. The Prohibitin-Binding Compound Fluorizoline Activates the Integrated Stress Response through the eIF2α Kinase HRI. Int. J. Mol. Sci. 2023, 24, 8064. [Google Scholar] [CrossRef]
- Zhu, S.; Nguyen, A.; Pang, J.; Zhao, J.; Chen, Z.; Liang, Z.; Gu, Y.; Huynh, H.; Bao, Y.; Lee, S.; et al. Mitochondrial Stress Induces an HRI-eIF2α Pathway Protective for Cardiomyopathy. Circulation 2022, 146, 1028–1031. [Google Scholar] [CrossRef]
- Bora, P.; Zaman, M.; Oviedo, S.; Kutseikin, S.; Madrazo, N.; Mathur, P.; Pannikkat, M.; Krasny, S.; Aldakhlallah, R.; Chu, A.; et al. Drug Repurposing Screen Identifies an HRI Activating Compound That Promotes Adaptive Mitochondrial Remodeling in MFN2-Deficient Cells. Proc. Natl. Acad. Sci. 2025, 122, e2517552122. [Google Scholar] [CrossRef]
- Sánchez-Vera, I.; Cosialls, A.M.; Maritorena-Hualde, N.; Schuler, M.-H.; Lavilla, R.; Pons, G.; Jae, L.T.; Iglesias-Serret, D.; Gil, J. Targeting Prohibitins Activates the ISR through DELE1-HRI by Impairing Protein Import into the Mitochondrial Matrix. Cell Death Differ. 2025. [Google Scholar] [CrossRef]
- Rivera-Mejías, P.; Narbona-Pérez, Á.J.; Hasberg, L.; Kroczek, L.; Bahat, A.; Lawo, S.; Folz-Donahue, K.; Schumacher, A.-L.; Ahola, S.; Mayer, F.C.; et al. The Mitochondrial Protease OMA1 Acts as a Metabolic Safeguard upon Nuclear DNA Damage. Cell Rep. 2023, 42, 112332. [Google Scholar] [CrossRef]
- Guo, X.; Aviles, G.; Liu, Y.; Tian, R.; Unger, B.A.; Lin, Y.-H.T.; Wiita, A.P.; Xu, K.; Correia, M.A.; Kampmann, M. Mitochondrial Stress Is Relayed to the Cytosol by an OMA1–DELE1–HRI Pathway. Nature 2020, 579, 427–432. [Google Scholar] [CrossRef] [PubMed]
- Fessler, E.; Eckl, E.-M.; Schmitt, S.; Mancilla, I.A.; Meyer-Bender, M.F.; Hanf, M.; Philippou-Massier, J.; Krebs, S.; Zischka, H.; Jae, L.T. A Pathway Coordinated by DELE1 Relays Mitochondrial Stress to the Cytosol. Nature 2020, 579, 433–437. [Google Scholar] [CrossRef] [PubMed]
- Alavi, M.V. OMA1—An Integral Membrane Protease? Biochim. Biophys. Acta BBA - Proteins Proteomics 2021, 1869, 140558. [Google Scholar] [CrossRef] [PubMed]
- Baker, M.J.; Lampe, P.A.; Stojanovski, D.; Korwitz, A.; Anand, R.; Tatsuta, T.; Langer, T. Stress-Induced OMA1 Activation and Autocatalytic Turnover Regulate OPA1-Dependent Mitochondrial Dynamics. EMBO J. 2014, 33, 578–593. [Google Scholar] [CrossRef]
- Fessler, E.; Krumwiede, L.; Jae, L.T. DELE1 Tracks Perturbed Protein Import and Processing in Human Mitochondria. Nat. Commun. 2022, 13, 1853. [Google Scholar] [CrossRef]
- Schmidt, O.; Pfanner, N.; Meisinger, C. Mitochondrial Protein Import: From Proteomics to Functional Mechanisms. Nat. Rev. Mol. Cell Biol. 2010, 11, 655–667. [Google Scholar] [CrossRef]
- Fessler, E.; Eckl, E.-M.; Schmitt, S.; Mancilla, I.A.; Meyer-Bender, M.F.; Hanf, M.; Philippou-Massier, J.; Krebs, S.; Zischka, H.; Jae, L.T. A Pathway Coordinated by DELE1 Relays Mitochondrial Stress to the Cytosol. Nature 2020, 579, 433–437. [Google Scholar] [CrossRef]
- Sekine, Y.; Houston, R.; Eckl, E.-M.; Fessler, E.; Narendra, D.P.; Jae, L.T.; Sekine, S. A Mitochondrial Iron-Responsive Pathway Regulated by DELE1. Mol. Cell 2023, 83, 2059–2076.e6. [Google Scholar] [CrossRef] [PubMed]
- Chaveroux, C.; Lambert-Langlais, S.; Parry, L.; Carraro, V.; Jousse, C.; Maurin, A.-C.; Bruhat, A.; Marceau, G.; Sapin, V.; Averous, J.; et al. Identification of GCN2 as New Redox Regulator for Oxidative Stress Prevention in Vivo. Biochem. Biophys. Res. Commun. 2011, 415, 120–124. [Google Scholar] [CrossRef] [PubMed]
- De Vito, A.; Lazzaro, M.; Palmisano, I.; Cittaro, D.; Riba, M.; Lazarevic, D.; Bannai, M.; Gabellini, D.; Schiaffino, M.V. Amino Acid Deprivation Triggers a Novel GCN2-Independent Response Leading to the Transcriptional Reactivation of Non-Native DNA Sequences. PLOS ONE 2018, 13, e0200783. [Google Scholar] [CrossRef] [PubMed]
- Darawshi, O.; Yassin, O.; Shmuel, M.; Wek, R.C.; Mahdizadeh, S.J.; Eriksson, L.A.; Hatzoglou, M.; Tirosh, B. Phosphorylation of GCN2 by mTOR Confers Adaptation to Conditions of Hyper-mTOR Activation under Stress. J. Biol. Chem. 2024, 300, 107575. [Google Scholar] [CrossRef] [PubMed]
- Li, J.; Gao, L.; Chen, J.; Zhang, W.-W.; Zhang, X.-Y.; Wang, B.; Zhang, C.; Wang, Y.; Huang, Y.-C.; Wang, H.; et al. Mitochondrial ROS-Mediated Ribosome Stalling and GCN2 Activation Are Partially Involved in 1-Nitropyrene-Induced Steroidogenic Inhibition in Testes. Environ. Int. 2022, 167, 107393. [Google Scholar] [CrossRef] [PubMed]
- Baker, B.M.; Nargund, A.M.; Sun, T.; Haynes, C.M. Protective Coupling of Mitochondrial Function and Protein Synthesis via the eIF2α Kinase GCN-2. PLoS Genet. 2012, 8, e1002760. [Google Scholar] [CrossRef] [PubMed]
- Falcón, P.; Escandón, M.; Brito, Á.; Matus, S. Nutrient Sensing and Redox Balance: GCN2 as a New Integrator in Aging. Oxid. Med. Cell. Longev. 2019, 2019, 1–9. [Google Scholar] [CrossRef]
- Li, J.; Gao, L.; Chen, J.; Zhang, W.-W.; Zhang, X.-Y.; Wang, B.; Zhang, C.; Wang, Y.; Huang, Y.-C.; Wang, H.; et al. Mitochondrial ROS-Mediated Ribosome Stalling and GCN2 Activation Are Partially Involved in 1-Nitropyrene-Induced Steroidogenic Inhibition in Testes. Environ. Int. 2022, 167, 107393. [Google Scholar] [CrossRef]
- Takase, S.; Kurokawa, R.; Kondoh, Y.; Honda, K.; Suzuki, T.; Kawahara, T.; Ikeda, H.; Dohmae, N.; Osada, H.; Shin-ya, K.; et al. Mechanism of Action of Prethioviridamide, an Anticancer Ribosomally Synthesized and Post-Translationally Modified Peptide with a Polythioamide Structure. ACS Chem. Biol. 2019, 14, 1819–1828. [Google Scholar] [CrossRef] [PubMed]
- Luengo, A.; Li, Z.; Gui, D.Y.; Sullivan, L.B.; Zagorulya, M.; Do, B.T.; Ferreira, R.; Naamati, A.; Ali, A.; Lewis, C.A.; et al. Increased Demand for NAD+ Relative to ATP Drives Aerobic Glycolysis. Mol. Cell 2021, 81, 691–707.e6. [Google Scholar] [CrossRef]
- Ye, J.; Palm, W.; Peng, M.; King, B.; Lindsten, T.; Li, M.O.; Koumenis, C.; Thompson, C.B. GCN2 Sustains mTORC1 Suppression upon Amino Acid Deprivation by Inducing Sestrin2. Genes Dev. 2015, 29, 2331–2336. [Google Scholar] [CrossRef]
- Rath, E.; Berger, E.; Messlik, A.; Nunes, T.; Liu, B.; Kim, S.C.; Hoogenraad, N.; Sans, M.; Sartor, R.B.; Haller, D. Induction of dsRNA-Activated Protein Kinase Links Mitochondrial Unfolded Protein Response to the Pathogenesis of Intestinal Inflammation. Gut 2012, 61, 1269–1278. [Google Scholar] [CrossRef] [PubMed]
- Borowski, L.S.; Dziembowski, A.; Hejnowicz, M.S.; Stepien, P.P.; Szczesny, R.J. Human Mitochondrial RNA Decay Mediated by PNPase–hSuv3 Complex Takes Place in Distinct Foci. Nucleic Acids Res. 2013, 41, 1223–1240. [Google Scholar] [CrossRef]
- Wang, D.D.-H.; Shu, Z.; Lieser, S.A.; Chen, P.-L.; Lee, W.-H. Human Mitochondrial SUV3 and Polynucleotide Phosphorylase Form a 330-kDa Heteropentamer to Cooperatively Degrade Double-Stranded RNA with a 3′-to-5′ Directionality. J. Biol. Chem. 2009, 284, 20812–20821. [Google Scholar] [CrossRef] [PubMed]
- Murphy, W.I.; Attardi, B.; Tu, C.; Attardi, G. Evidence for Complete Symmetrical Transcription in Vivo of Mitochondrial DNA in HeLa Cells. J. Mol. Biol. 1975, 99, 809–814. [Google Scholar] [CrossRef] [PubMed]
- Young, P.G.; Attardi, G. Characterization of Double-Stranded RNA from HeLa Cell Mitochondria. Biochem. Biophys. Res. Commun. 1975, 65, 1201–1207. [Google Scholar] [CrossRef]
- Kim, Y.; Park, J.; Kim, S.; Kim, M.; Kang, M.-G.; Kwak, C.; Kang, M.; Kim, B.; Rhee, H.-W.; Kim, V.N. PKR Senses Nuclear and Mitochondrial Signals by Interacting with Endogenous Double-Stranded RNAs. Mol. Cell 2018, 71, 1051–1063.e6. [Google Scholar] [CrossRef] [PubMed]
- Kusuma, F.; Park, S.; Nguyen, K.A.; Elvira, R.; Lee, D.; Han, J. PKR Mediates the Mitochondrial Unfolded Protein Response through Double-Stranded RNA Accumulation under Mitochondrial Stress. Int. J. Mol. Sci. 2024, 25, 7738. [Google Scholar] [CrossRef] [PubMed]
- Kim, S.; Lee, K.; Choi, Y.S.; Ku, J.; Kim, H.; Kharbash, R.; Yoon, J.; Lee, Y.S.; Kim, J.-H.; Lee, Y.J.; et al. Mitochondrial Double-Stranded RNAs Govern the Stress Response in Chondrocytes to Promote Osteoarthritis Development. Cell Rep. 2022, 40, 111178. [Google Scholar] [CrossRef]
- Yoon, J.; Lee, M.; Ali, A.A.; Oh, Y.R.; Choi, Y.S.; Kim, S.; Lee, N.; Jang, S.G.; Park, S.; Chung, J.-H.; et al. Mitochondrial Double-Stranded RNAs as a Pivotal Mediator in the Pathogenesis of Sjӧgren’s Syndrome. Mol. Ther. - Nucleic Acids 2022, 30, 257–269. [Google Scholar] [CrossRef]
- Fu, Y.; Sacco, O.; DeBitetto, E.; Kanshin, E.; Ueberheide, B.; Sfeir, A. Mitochondrial DNA Breaks Activate an Integrated Stress Response to Reestablish Homeostasis. Mol. Cell 2023, 83, 3740–3753.e9. [Google Scholar] [CrossRef]
- Ben-Sahra, I.; Hoxhaj, G.; Ricoult, S.J.H.; Asara, J.M.; Manning, B.D. mTORC1 Induces Purine Synthesis through Control of the Mitochondrial Tetrahydrofolate Cycle. Science 2016, 351, 728–733. [Google Scholar] [CrossRef] [PubMed]
- Li, T.Y.; Wang, Q.; Gao, A.W.; Li, X.; Sun, Y.; Mottis, A.; Shong, M.; Auwerx, J. Lysosomes Mediate the Mitochondrial UPR via mTORC1-Dependent ATF4 Phosphorylation. Cell Discov. 2023, 9, 92. [Google Scholar] [CrossRef]
- Quirós, P.M.; Prado, M.A.; Zamboni, N.; D’Amico, D.; Williams, R.W.; Finley, D.; Gygi, S.P.; Auwerx, J. Multi-Omics Analysis Identifies ATF4 as a Key Regulator of the Mitochondrial Stress Response in Mammals. J. Cell Biol. 2017, 216, 2027–2045. [Google Scholar] [CrossRef]
- Sasaki, K.; Uchiumi, T.; Toshima, T.; Yagi, M.; Do, Y.; Hirai, H.; Igami, K.; Gotoh, K.; Kang, D. Mitochondrial Translation Inhibition Triggers ATF4 Activation, Leading to Integrated Stress Response but Not to Mitochondrial Unfolded Protein Response. Biosci. Rep. 2020, 40, BSR20201289. [Google Scholar] [CrossRef]
- Mahor, D.; Pandey, R.; Bulusu, V. TCA Cycle off, ATF4 on for Metabolic Homeostasis. Trends Biochem. Sci. 2022, 47, 558–560. [Google Scholar] [CrossRef]
- Kasai, S.; Shimizu, S.; Tatara, Y.; Mimura, J.; Itoh, K. Regulation of Nrf2 by Mitochondrial Reactive Oxygen Species in Physiology and Pathology. Biomolecules 2020, 10, 320. [Google Scholar] [CrossRef]
- Harding, H.P.; Zhang, Y.; Zeng, H.; Novoa, I.; Lu, P.D.; Calfon, M.; Sadri, N.; Yun, C.; Popko, B.; Paules, R.; et al. An Integrated Stress Response Regulates Amino Acid Metabolism and Resistance to Oxidative Stress. Mol. Cell 2003, 11, 619–633. [Google Scholar] [CrossRef] [PubMed]
- Fusakio, M.E.; Willy, J.A.; Wang, Y.; Mirek, E.T.; Al Baghdadi, R.J.T.; Adams, C.M.; Anthony, T.G.; Wek, R.C. Transcription Factor ATF4 Directs Basal and Stress-Induced Gene Expression in the Unfolded Protein Response and Cholesterol Metabolism in the Liver. Mol. Biol. Cell 2016, 27, 1536–1551. [Google Scholar] [CrossRef] [PubMed]
- Madrazo, N.; Khattar, Z.; Powers, E.T.; Rosarda, J.D.; Wiseman, R.L. Mapping Stress-Responsive Signaling Pathways Induced by Mitochondrial Proteostasis Perturbations. Mol. Biol. Cell 2024, 35, ar74. [Google Scholar] [CrossRef]
- Uoselis, L.; Lindblom, R.; Lam, W.K.; Küng, C.J.; Skulsuppaisarn, M.; Khuu, G.; Nguyen, T.N.; Rudler, D.L.; Filipovska, A.; Schittenhelm, R.B.; et al. Temporal Landscape of Mitochondrial Proteostasis Governed by the UPRmt. Sci. Adv. 2023, 9, eadh8228. [Google Scholar] [CrossRef]
- Kühl, I.; Miranda, M.; Atanassov, I.; Kuznetsova, I.; Hinze, Y.; Mourier, A.; Filipovska, A.; Larsson, N.-G. Transcriptomic and Proteomic Landscape of Mitochondrial Dysfunction Reveals Secondary Coenzyme Q Deficiency in Mammals. eLife 2017, 6, e30952. [Google Scholar] [CrossRef]
- Bond, S.T.; King, E.J.; Walker, S.M.; Yang, C.; Liu, Y.; Liu, K.H.; Zhuang, A.; Jurrjens, A.W.; Fang, H.A.; Formosa, L.E.; et al. Mitochondrial Damage in Muscle Specific PolG Mutant Mice Activates the Integrated Stress Response and Disrupts the Mitochondrial Folate Cycle. Nat. Commun. 2025, 16, 2338. [Google Scholar] [CrossRef]
- Nikkanen, J.; Forsström, S.; Euro, L.; Paetau, I.; Kohnz, R.A.; Wang, L.; Chilov, D.; Viinamäki, J.; Roivainen, A.; Marjamäki, P.; et al. Mitochondrial DNA Replication Defects Disturb Cellular dNTP Pools and Remodel One-Carbon Metabolism. Cell Metab. 2016, 23, 635–648. [Google Scholar] [CrossRef] [PubMed]
- Rezaei Adariani, S.; Agne, D.; Koska, S.; Burhop, A.; Seitz, C.; Warmers, J.; Janning, P.; Metz, M.; Pahl, A.; Sievers, S.; et al. Detection of a Mitochondrial Fragmentation and Integrated Stress Response Using the Cell Painting Assay. J. Med. Chem. 2024, 67, 13252–13270. [Google Scholar] [CrossRef]
- Keerthiga, R.; Pei, D.-S.; Fu, A. Mitochondrial Dysfunction, UPRmt Signaling, and Targeted Therapy in Metastasis Tumor. Cell Biosci. 2021, 11, 186. [Google Scholar] [CrossRef] [PubMed]
- B’chir, W.; Maurin, A.-C.; Carraro, V.; Averous, J.; Jousse, C.; Muranishi, Y.; Parry, L.; Stepien, G.; Fafournoux, P.; Bruhat, A. The eIF2α/ATF4 Pathway Is Essential for Stress-Induced Autophagy Gene Expression. Nucleic Acids Res. 2013, 41, 7683–7699. [Google Scholar] [CrossRef] [PubMed]
- Örd, T.; Örd, D.; Adler, P.; Örd, T. Genome-Wide Census of ATF4 Binding Sites and Functional Profiling of Trait-Associated Genetic Variants Overlapping ATF4 Binding Motifs. PLoS Genet. 2023, 19, e1011014. [Google Scholar] [CrossRef] [PubMed]
- Neill, G.; Masson, G.R. A Stay of Execution: ATF4 Regulation and Potential Outcomes for the Integrated Stress Response. Front. Mol. Neurosci. 2023, 16, 1112253. [Google Scholar] [CrossRef]
- Clemens, M.J. Initiation Factor eIF2α Phosphorylation in Stress Responses and Apoptosis. In Signaling Pathways for Translation; Progress in Molecular and Subcellular Biology; Rhoads, R.E., Ed.; Springer Berlin Heidelberg: Berlin, Heidelberg, 2001; Vol. 27, pp. 57–89. ISBN 978-3-642-07505-6. [Google Scholar]
- Hu, H.; Tian, M.; Ding, C.; Yu, S. The C/EBP Homologous Protein (CHOP) Transcription Factor Functions in Endoplasmic Reticulum Stress-Induced Apoptosis and Microbial Infection. Front. Immunol. 2018, 9, 3083. [Google Scholar] [CrossRef] [PubMed]
- Tian, X.; El-Deiry, W.S. Integrated Stress Response (ISR) Activation and Apoptosis through HRI Kinase by PG3 and Other P53 Pathway-Restoring Cancer Therapeutics. Oncotarget 2024, 15, 614–633. [Google Scholar] [CrossRef]
- Fritsch, R.M.; Schneider, G.; Saur, D.; Scheibel, M.; Schmid, R.M. Translational Repression of MCL-1 Couples Stress-Induced eIF2α Phosphorylation to Mitochondrial Apoptosis Initiation. J. Biol. Chem. 2007, 282, 22551–22562. [Google Scholar] [CrossRef] [PubMed]
- Núñez-Vázquez, S.; Sánchez-Vera, I.; Saura-Esteller, J.; Cosialls, A.M.; Noisier, A.F.M.; Albericio, F.; Lavilla, R.; Pons, G.; Iglesias-Serret, D.; Gil, J. NOXA Upregulation by the Prohibitin-binding Compound Fluorizoline Is Transcriptionally Regulated by Integrated Stress Response-induced ATF3 and ATF4. FEBS J. 2021, 288, 1271–1285. [Google Scholar] [CrossRef]
- Kalkavan, H.; Chen, M.J.; Crawford, J.C.; Quarato, G.; Fitzgerald, P.; Tait, S.W.G.; Goding, C.R.; Green, D.R. Sublethal Cytochrome c Release Generates Drug-Tolerant Persister Cells. Cell 2022, 185, 3356–3374.e22. [Google Scholar] [CrossRef]
- Li, T.; Su, L.; Lei, Y.; Liu, X.; Zhang, Y.; Liu, X. DDIT3 and KAT2A Proteins Regulate TNFRSF10A and TNFRSF10B Expression in Endoplasmic Reticulum Stress-Mediated Apoptosis in Human Lung Cancer Cells. J. Biol. Chem. 2015, 290, 11108–11118. [Google Scholar] [CrossRef]
- Liu, K.; Shi, Y.; Guo, X.; Wang, S.; Ouyang, Y.; Hao, M.; Liu, D.; Qiao, L.; Li, N.; Zheng, J.; et al. CHOP Mediates ASPP2-Induced Autophagic Apoptosis in Hepatoma Cells by Releasing Beclin-1 from Bcl-2 and Inducing Nuclear Translocation of Bcl-2. Cell Death Dis. 2014, 5, e1323–e1323. [Google Scholar] [CrossRef]
- Ghosh, A.P.; Klocke, B.J.; Ballestas, M.E.; Roth, K.A. CHOP Potentially Co-Operates with FOXO3a in Neuronal Cells to Regulate PUMA and BIM Expression in Response to ER Stress. PLoS ONE 2012, 7, e39586. [Google Scholar] [CrossRef]
- Tsukano, H.; Gotoh, T.; Endo, M.; Miyata, K.; Tazume, H.; Kadomatsu, T.; Yano, M.; Iwawaki, T.; Kohno, K.; Araki, K.; et al. The Endoplasmic Reticulum Stress-C/EBP Homologous Protein Pathway-Mediated Apoptosis in Macrophages Contributes to the Instability of Atherosclerotic Plaques. Arterioscler. Thromb. Vasc. Biol. 2010, 30, 1925–1932. [Google Scholar] [CrossRef] [PubMed]
- Sora, V.; Papaleo, E. Structural Details of BH3 Motifs and BH3-Mediated Interactions: An Updated Perspective. Front. Mol. Biosci. 2022, 9, 864874. [Google Scholar] [CrossRef] [PubMed]
- Maillet, L.; Fétiveau, A.; Lalier, L.; Martin, N.; Barillé-Nion, S.; Guette, C.; Gautier, F.; Téletchéa, S.; Juin, P.P. Allosteric Regulation of BH3-in-Groove Interactions by Tail Anchors of BCL-xL Complexes Limits BH3 Mimetic Antagonism. Nat. Commun. 2025, 16, 10621. [Google Scholar] [CrossRef]
- Cheng, E.H.-Y.; Sheiko, T.V.; Fisher, J.K.; Craigen, W.J.; Korsmeyer, S.J. VDAC2 Inhibits BAK Activation and Mitochondrial Apoptosis. Science 2003, 301, 513–517. [Google Scholar] [CrossRef]
- Hohorst, L.; Ros, U.; Garcia-Saez, A.J. Mitochondrial Dynamics and Pore Formation in Regulated Cell Death Pathways. Trends Biochem. Sci. 2025, 50, 1001–1014. [Google Scholar] [CrossRef]
- Nguyen, T.T.; Wei, S.; Nguyen, T.H.; Jo, Y.; Zhang, Y.; Park, W.; Gariani, K.; Oh, C.-M.; Kim, H.H.; Ha, K.-T.; et al. Mitochondria-Associated Programmed Cell Death as a Therapeutic Target for Age-Related Disease. Exp. Mol. Med. 2023, 55, 1595–1619. [Google Scholar] [CrossRef] [PubMed]
- Kim, H.; Tu, H.-C.; Ren, D.; Takeuchi, O.; Jeffers, J.R.; Zambetti, G.P.; Hsieh, J.J.-D.; Cheng, E.H.-Y. Stepwise Activation of BAX and BAK by tBID, BIM, and PUMA Initiates Mitochondrial Apoptosis. Mol. Cell 2009, 36, 487–499. [Google Scholar] [CrossRef]
- Sarosiek, K.A.; Chi, X.; Bachman, J.A.; Sims, J.J.; Montero, J.; Patel, L.; Flanagan, A.; Andrews, D.W.; Sorger, P.; Letai, A. BID Preferentially Activates BAK While BIM Preferentially Activates BAX, Affecting Chemotherapy Response. Mol. Cell 2013, 51, 751–765. [Google Scholar] [CrossRef]
- Puthalakath, H.; O’Reilly, L.A.; Gunn, P.; Lee, L.; Kelly, P.N.; Huntington, N.D.; Hughes, P.D.; Michalak, E.M.; McKimm-Breschkin, J.; Motoyama, N.; et al. ER Stress Triggers Apoptosis by Activating BH3-Only Protein Bim. Cell 2007, 129, 1337–1349. [Google Scholar] [CrossRef]
- Galehdar, Z.; Swan, P.; Fuerth, B.; Callaghan, S.M.; Park, D.S.; Cregan, S.P. Neuronal Apoptosis Induced by Endoplasmic Reticulum Stress Is Regulated by ATF4–CHOP-Mediated Induction of the Bcl-2 Homology 3-Only Member PUMA. J. Neurosci. 2010, 30, 16938–16948. [Google Scholar] [CrossRef] [PubMed]
- Zappa, F.; Muniozguren, N.L.; Conrad, J.E.; Acosta-Alvear, D. The Integrated Stress Response Engages a Cell-Autonomous, Ligand-Independent, DR5-Driven Apoptosis Switch. Cell Death Dis. 2025, 16, 101. [Google Scholar] [CrossRef] [PubMed]
- Chan, Z.C.K.; Leong, K.H.; Kareem, H.S.; Norazit, A.; Noor, S.M.; Ariffin, A. Activation of Death Receptor, DR5 and Mitochondria-Mediated Apoptosis by a 3,4,5-Trimethoxybenzyloxy Derivative in Wild-Type and P53 Mutant Colorectal Cancer Cell Lines. Naunyn. Schmiedebergs Arch. Pharmacol. 2020, 393, 405–417. [Google Scholar] [CrossRef] [PubMed]
- Lu, M.; Lawrence, D.A.; Marsters, S.; Acosta-Alvear, D.; Kimmig, P.; Mendez, A.S.; Paton, A.W.; Paton, J.C.; Walter, P.; Ashkenazi, A. Opposing Unfolded-Protein-Response Signals Converge on Death Receptor 5 to Control Apoptosis. Science 2014, 345, 98–101. [Google Scholar] [CrossRef] [PubMed]
- Iurlaro, R.; Püschel, F.; León-Annicchiarico, C.L.; O’Connor, H.; Martin, S.J.; Palou-Gramón, D.; Lucendo, E.; Muñoz-Pinedo, C. Glucose Deprivation Induces ATF4-Mediated Apoptosis through TRAIL Death Receptors. Mol. Cell. Biol. 2017, 37, e00479-16. [Google Scholar] [CrossRef]
- Muniozguren, N.L.; Zappa, F.; Acosta-Alvear, D. The Integrated Stress Response Induces a Common Cell-Autonomous Death Receptor 5-Dependent Apoptosis Switch 2022.
- Kischkel, F.C.; Lawrence, D.A.; Chuntharapai, A.; Schow, P.; Kim, K.J.; Ashkenazi, A. Apo2L/TRAIL-Dependent Recruitment of Endogenous FADD and Caspase-8 to Death Receptors 4 and 5. Immunity 2000, 12, 611–620. [Google Scholar] [CrossRef] [PubMed]
- Taketani, K.; Kawauchi, J.; Tanaka-Okamoto, M.; Ishizaki, H.; Tanaka, Y.; Sakai, T.; Miyoshi, J.; Maehara, Y.; Kitajima, S. Key Role of ATF3 in P53-Dependent DR5 Induction upon DNA Damage of Human Colon Cancer Cells. Oncogene 2012, 31, 2210–2221. [Google Scholar] [CrossRef]
- Nargund, A.M.; Pellegrino, M.W.; Fiorese, C.J.; Baker, B.M.; Haynes, C.M. Mitochondrial Import Efficiency of ATFS-1 Regulates Mitochondrial UPR Activation. Science 2012, 337, 587–590. [Google Scholar] [CrossRef] [PubMed]
- Nargund, A.M.; Fiorese, C.J.; Pellegrino, M.W.; Deng, P.; Haynes, C.M. Mitochondrial and Nuclear Accumulation of the Transcription Factor ATFS-1 Promotes OXPHOS Recovery during the UPRmt. Mol. Cell 2015, 58, 123–133. [Google Scholar] [CrossRef]
- Naresh, N.U.; Haynes, C.M. Signaling and Regulation of the Mitochondrial Unfolded Protein Response. Cold Spring Harb. Perspect. Biol. 2019, 11, a033944. [Google Scholar] [CrossRef]
- Lu, H.; Wang, X.; Li, M.; Ji, D.; Liang, D.; Liang, C.; Liu, Y.; Zhang, Z.; Cao, Y.; Zou, W. Mitochondrial Unfolded Protein Response and Integrated Stress Response as Promising Therapeutic Targets for Mitochondrial Diseases. Cells 2022, 12, 20. [Google Scholar] [CrossRef]
- Shpilka, T.; Haynes, C.M. The Mitochondrial UPR: Mechanisms, Physiological Functions and Implications in Ageing. Nat. Rev. Mol. Cell Biol. 2018, 19, 109–120. [Google Scholar] [CrossRef]
- Horibe, T.; Hoogenraad, N.J. The Chop Gene Contains an Element for the Positive Regulation of the Mitochondrial Unfolded Protein Response. PLoS ONE 2007, 2, e835. [Google Scholar] [CrossRef]
- Aldridge, J.E.; Horibe, T.; Hoogenraad, N.J. Discovery of Genes Activated by the Mitochondrial Unfolded Protein Response (mtUPR) and Cognate Promoter Elements. PLoS ONE 2007, 2, e874. [Google Scholar] [CrossRef]
- Fiorese, C.J.; Schulz, A.M.; Lin, Y.-F.; Rosin, N.; Pellegrino, M.W.; Haynes, C.M. The Transcription Factor ATF5 Mediates a Mammalian Mitochondrial UPR. Curr. Biol. 2016, 26, 2037–2043. [Google Scholar] [CrossRef]
- Teske, B.F.; Fusakio, M.E.; Zhou, D.; Shan, J.; McClintick, J.N.; Kilberg, M.S.; Wek, R.C. CHOP Induces Activating Transcription Factor 5 (ATF5) to Trigger Apoptosis in Response to Perturbations in Protein Homeostasis. Mol. Biol. Cell 2013, 24, 2477–2490. [Google Scholar] [CrossRef]
- Fiorese, C.J.; Schulz, A.M.; Lin, Y.-F.; Rosin, N.; Pellegrino, M.W.; Haynes, C.M. The Transcription Factor ATF5 Mediates a Mammalian Mitochondrial UPR. Curr. Biol. 2016, 26, 2037–2043. [Google Scholar] [CrossRef] [PubMed]
- Jiang, D.; Cui, H.; Xie, N.; Banerjee, S.; Liu, R.-M.; Dai, H.; Thannickal, V.J.; Liu, G. ATF4 Mediates Mitochondrial Unfolded Protein Response in Alveolar Epithelial Cells. Am. J. Respir. Cell Mol. Biol. 2020, 63, 478–489. [Google Scholar] [CrossRef]
- Papa, L.; Germain, D. SirT3 Regulates the Mitochondrial Unfolded Protein Response. Mol. Cell. Biol. 2014, 34, 699–710. [Google Scholar] [CrossRef] [PubMed]
- Kenny, T.C.; Hart, P.; Ragazzi, M.; Sersinghe, M.; Chipuk, J.; Sagar, M.A.K.; Eliceiri, K.W.; LaFramboise, T.; Grandhi, S.; Santos, J.; et al. Selected Mitochondrial DNA Landscapes Activate the SIRT3 Axis of the UPRmt to Promote Metastasis. Oncogene 2017, 36, 4393–4404. [Google Scholar] [CrossRef]
- Kenny, T.C.; Germain, D. From Discovery of the CHOP Axis and Targeting ClpP to the Identification of Additional Axes of the UPRmt Driven by the Estrogen Receptor and SIRT3. J. Bioenerg. Biomembr. 2017, 49, 297–305. [Google Scholar] [CrossRef]
- Papa, L.; Germain, D. Estrogen Receptor Mediates a Distinct Mitochondrial Unfolded Protein Response. J. Cell Sci. 2011, 124, 1396–1402. [Google Scholar] [CrossRef] [PubMed]
- Zhou, D.; Palam, L.R.; Jiang, L.; Narasimhan, J.; Staschke, K.A.; Wek, R.C. Phosphorylation of eIF2 Directs ATF5 Translational Control in Response to Diverse Stress Conditions. J. Biol. Chem. 2008, 283, 7064–7073. [Google Scholar] [CrossRef]
- Hinnebusch, A.G.; Ivanov, I.P.; Sonenberg, N. Translational Control by 5′-Untranslated Regions of Eukaryotic mRNAs. Science 2016, 352, 1413–1416. [Google Scholar] [CrossRef] [PubMed]
- Watatani, Y.; Ichikawa, K.; Nakanishi, N.; Fujimoto, M.; Takeda, H.; Kimura, N.; Hirose, H.; Takahashi, S.; Takahashi, Y. Stress-Induced Translation of ATF5 mRNA Is Regulated by the 5′-Untranslated Region. J. Biol. Chem. 2008, 283, 2543–2553. [Google Scholar] [CrossRef] [PubMed]
- Ma, Z.; Horrocks, J.; Mir, D.A.; Cox, M.; Ruzga, M.; Rollins, J.; Rogers, A.N. The Integrated Stress Response Protects against ER Stress but Is Not Required for Altered Translation and Lifespan from Dietary Restriction in Caenorhabditis Elegans. Front. Cell Dev. Biol. 2023, 11, 1263344. [Google Scholar] [CrossRef]
- Fusakio, M.E.; Willy, J.A.; Wang, Y.; Mirek, E.T.; Al Baghdadi, R.J.T.; Adams, C.M.; Anthony, T.G.; Wek, R.C. Transcription Factor ATF4 Directs Basal and Stress-Induced Gene Expression in the Unfolded Protein Response and Cholesterol Metabolism in the Liver. Mol. Biol. Cell 2016, 27, 1536–1551. [Google Scholar] [CrossRef]
- Liu, K.; Zhao, C.; Adajar, R.C.; DeZwaan-McCabe, D.; Rutkowski, D.T. A Beneficial Adaptive Role for CHOP in Driving Cell Fate Selection during ER Stress 2023.
- Ma, Y.; Brewer, J.W.; Alan Diehl, J.; Hendershot, L.M. Two Distinct Stress Signaling Pathways Converge Upon the CHOP Promoter During the Mammalian Unfolded Protein Response. J. Mol. Biol. 2002, 318, 1351–1365. [Google Scholar] [CrossRef] [PubMed]
- Su, N.; Kilberg, M.S. C/EBP Homology Protein (CHOP) Interacts with Activating Transcription Factor 4 (ATF4) and Negatively Regulates the Stress-Dependent Induction of the Asparagine Synthetase Gene. J. Biol. Chem. 2008, 283, 35106–35117. [Google Scholar] [CrossRef] [PubMed]
- Vincent, A.E.; White, K.; Davey, T.; Philips, J.; Ogden, R.T.; Lawless, C.; Warren, C.; Hall, M.G.; Ng, Y.S.; Falkous, G.; et al. Quantitative 3D Mapping of the Human Skeletal Muscle Mitochondrial Network. Cell Rep. 2019, 26, 996–1009.e4. [Google Scholar] [CrossRef]
- Chen, C.C.W.; Erlich, A.T.; Crilly, M.J.; Hood, D.A. Parkin Is Required for Exercise-Induced Mitophagy in Muscle: Impact of Aging. Am. J. Physiol.-Endocrinol. Metab. 2018, 315, E404–E415. [Google Scholar] [CrossRef]
- Oliveira, A.N.; Hood, D.A. Effect of Tim23 Knockdown in Vivo on Mitochondrial Protein Import and Retrograde Signaling to the UPRmt in Muscle. Am. J. Physiol.-Cell Physiol. 2018, 315, C516–C526. [Google Scholar] [CrossRef] [PubMed]
- Pöverlein, M.C.; Jussupow, A.; Kim, H.; Kaila, V.R. Protein-Induced Membrane Strain Drives Supercomplex Formation 2025.
- Kayvanpour, E.; Wisdom, M.; Lackner, M.K.; Sedaghat-Hamedani, F.; Boeckel, J.-N.; Müller, M.; Eghbalian, R.; Dudek, J.; Doroudgar, S.; Maack, C.; et al. VARS2 Depletion Leads to Activation of the Integrated Stress Response and Disruptions in Mitochondrial Fatty Acid Oxidation. Int. J. Mol. Sci. 2022, 23, 7327. [Google Scholar] [CrossRef] [PubMed]
- Touvier, T.; De Palma, C.; Rigamonti, E.; Scagliola, A.; Incerti, E.; Mazelin, L.; Thomas, J.-L.; D’Antonio, M.; Politi, L.; Schaeffer, L.; et al. Muscle-Specific Drp1 Overexpression Impairs Skeletal Muscle Growth via Translational Attenuation. Cell Death Dis. 2015, 6, e1663–e1663. [Google Scholar] [CrossRef]
- Jackson, C.B.; Marmyleva, A.; Monteuuis, G.; Awadhpersad, R.; Mito, T.; Zamboni, N.; Tatsuta, T.; Vincent, A.E.; Wang, L.; Khan, N.A.; et al. De Novo Serine Biosynthesis Is Protective in Mitochondrial Disease. Cell Rep. 2025, 44, 115710. [Google Scholar] [CrossRef] [PubMed]
- Agnew, T.; Goldsworthy, M.; Aguilar, C.; Morgan, A.; Simon, M.; Hilton, H.; Esapa, C.; Wu, Y.; Cater, H.; Bentley, L.; et al. A Wars2 Mutant Mouse Model Displays OXPHOS Deficiencies and Activation of Tissue-Specific Stress Response Pathways. Cell Rep. 2018, 25, 3315–3328.e6. [Google Scholar] [CrossRef]
- Streeter, J.; Persaud, L.; Gao, J.; Manika, D.; Fairman, W.; García-Peña, L.M.; Marti, A.; Manika, C.; Gaddi, S.; Schickling, B.; et al. ATF4-Dependent and Independent Mitokine Secretion from OPA1 Deficient Skeletal Muscle in Mice Is Sexually Dimorphic. Front. Endocrinol. 2024, 15, 1325286. [Google Scholar] [CrossRef]
- Lin, S.; Wu, J.; Lian, G.; Wu, W.; Chen, W.; Chen, A.; Luo, L.; Xie, L. Protective Effects of Salubrinal against H2O2-Induced Muscle Wasting via eIF2α/ATF4 Signaling Pathway. Front. Pharmacol. 2025, 16, 1607606. [Google Scholar] [CrossRef]
- Guo, Q.; Xu, Z.; Zhou, D.; Fu, T.; Wang, W.; Sun, W.; Xiao, L.; Liu, L.; Ding, C.; Yin, Y.; et al. Mitochondrial Proteostasis Stress in Muscle Drives a Long-Range Protective Response to Alleviate Dietary Obesity Independently of ATF4. Sci. Adv. 2022, 8, eabo0340. [Google Scholar] [CrossRef] [PubMed]
- Keipert, S.; Ost, M.; Johann, K.; Imber, F.; Jastroch, M.; Van Schothorst, E.M.; Keijer, J.; Klaus, S. Skeletal Muscle Mitochondrial Uncoupling Drives Endocrine Cross-Talk through the Induction of FGF21 as a Myokine. Am. J. Physiol.-Endocrinol. Metab. 2014, 306, E469–E482. [Google Scholar] [CrossRef] [PubMed]
- Lin, H.-P.; Petersen, J.D.; Gilsrud, A.J.; Madruga, A.; D’Silva, T.M.; Huang, X.; Shammas, M.K.; Randolph, N.P.; Johnson, K.R.; Li, Y.; et al. DELE1 Maintains Muscle Proteostasis to Promote Growth and Survival in Mitochondrial Myopathy. EMBO J. 2024, 43, 5548–5585. [Google Scholar] [CrossRef] [PubMed]
- Igual Gil, C.; Löser, A.; Lossow, K.; Schwarz, M.; Weber, D.; Grune, T.; Kipp, A.P.; Klaus, S.; Ost, M. Temporal Dynamics of Muscle Mitochondrial Uncoupling-Induced Integrated Stress Response and Ferroptosis Defense. Front. Endocrinol. 2023, 14, 1277866. [Google Scholar] [CrossRef]
- Vásquez-Trincado, C.; Dunn, J.; Han, J.I.; Hymms, B.; Tamaroff, J.; Patel, M.; Nguyen, S.; Dedio, A.; Wade, K.; Enigwe, C.; et al. Frataxin Deficiency Lowers Lean Mass and Triggers the Integrated Stress Response in Skeletal Muscle. JCI Insight 2022, 7, e155201. [Google Scholar] [CrossRef]
- Mick, E.; Titov, D.V.; Skinner, O.S.; Sharma, R.; Jourdain, A.A.; Mootha, V.K. Distinct Mitochondrial Defects Trigger the Integrated Stress Response Depending on the Metabolic State of the Cell. eLife 2020, 9, e49178. [Google Scholar] [CrossRef]
- Memme, J.M.; Sanfrancesco, V.C.; Hood, D.A. Activating Transcription Factor 4 Regulates Mitochondrial Content, Morphology, and Function in Differentiating Skeletal Muscle Myotubes. Am. J. Physiol.-Cell Physiol. 2023, 325, C224–C242. [Google Scholar] [CrossRef] [PubMed]
- Slavin, M.B.; Kumari, R.; Hood, D.A. ATF5 Is a Regulator of Exercise-Induced Mitochondrial Quality Control in Skeletal Muscle. Mol. Metab. 2022, 66, 101623. [Google Scholar] [CrossRef] [PubMed]
- Kuthe, J.; Hood, D.A. ATF5 Is Required for the Normal Stress Response during Skeletal Muscle Denervation. Sports Med. Health Sci. 2026, S2666337626000132. [Google Scholar] [CrossRef]
- Layec, G.; Blain, G.M.; Rossman, M.J.; Park, S.Y.; Hart, C.R.; Trinity, J.D.; Gifford, J.R.; Sidhu, S.K.; Weavil, J.C.; Hureau, T.J.; et al. Acute High-Intensity Exercise Impairs Skeletal Muscle Respiratory Capacity. Med. Sci. Sports Exerc. 2018, 50, 2409–2417. [Google Scholar] [CrossRef] [PubMed]
- Yoo, S.-Z.; No, M.-H.; Heo, J.-W.; Park, D.-H.; Kang, J.-H.; Kim, J.-H.; Seo, D.-Y.; Han, J.; Jung, S.-J.; Kwak, H.-B. Effects of Acute Exercise on Mitochondrial Function, Dynamics, and Mitophagy in Rat Cardiac and Skeletal Muscles. Int. Neurourol. J. 2019, 23, S22–31. [Google Scholar] [CrossRef]
- Vargas-Mendoza, N.; Angeles-Valencia, M.; Morales-González, Á.; Madrigal-Santillán, E.O.; Morales-Martínez, M.; Madrigal-Bujaidar, E.; Álvarez-González, I.; Gutiérrez-Salinas, J.; Esquivel-Chirino, C.; Chamorro-Cevallos, G.; et al. Oxidative Stress, Mitochondrial Function and Adaptation to Exercise: New Perspectives in Nutrition. Life 2021, 11, 1269. [Google Scholar] [CrossRef] [PubMed]
- Musci, R.; Hamilton, K.; Linden, M. Exercise-Induced Mitohormesis for the Maintenance of Skeletal Muscle and Healthspan Extension. Sports 2019, 7, 170. [Google Scholar] [CrossRef] [PubMed]
- Wang, F.; Wang, X.; Liu, Y.; Zhang, Z. Effects of Exercise-Induced ROS on the Pathophysiological Functions of Skeletal Muscle. Oxid. Med. Cell. Longev. 2021, 2021, 3846122. [Google Scholar] [CrossRef] [PubMed]
- Bouviere, J.; Fortunato, R.S.; Dupuy, C.; Werneck-de-Castro, J.P.; Carvalho, D.P.; Louzada, R.A. Exercise-Stimulated ROS Sensitive Signaling Pathways in Skeletal Muscle. Antioxidants 2021, 10, 537. [Google Scholar] [CrossRef]
- Lessard, S.J.; MacDonald, T.L.; Pathak, P.; Han, M.S.; Coffey, V.G.; Edge, J.; Rivas, D.A.; Hirshman, M.F.; Davis, R.J.; Goodyear, L.J. JNK Regulates Muscle Remodeling via Myostatin/SMAD Inhibition. Nat. Commun. 2018, 9, 3030. [Google Scholar] [CrossRef] [PubMed]
- Memme, J.M.; Oliveira, A.N.; Hood, D.A. Chronology of UPR Activation in Skeletal Muscle Adaptations to Chronic Contractile Activity. Am. J. Physiol.-Cell Physiol. 2016, 310, C1024–C1036. [Google Scholar] [CrossRef]
- Sanfrancesco, V.C.; Hood, D.A. Acute Contractile Activity Induces the Activation of the Mitochondrial Integrated Stress Response and the Transcription Factor ATF4. J. Appl. Physiol. 2025, 138, 857–871. [Google Scholar] [CrossRef]
- Gaspar, R.S.; Katashima, C.K.; Crisol, B.M.; Carneiro, F.S.; Sampaio, I.; Silveira, L.D.R.; Silva, A.S.R.D.; Cintra, D.E.; Pauli, J.R.; Ropelle, E.R. Physical Exercise Elicits UPRmt in the Skeletal Muscle: The Role of c-Jun N-Terminal Kinase. Mol. Metab. 2023, 78, 101816. [Google Scholar] [CrossRef]
- Zhang, P.; Langland, J.O.; Jacobs, B.L.; Samuel, C.E. Protein Kinase PKR-Dependent Activation of Mitogen-Activated Protein Kinases Occurs through Mitochondrial Adapter IPS-1 and Is Antagonized by Vaccinia Virus E3L. J. Virol. 2009, 83, 5718–5725. [Google Scholar] [CrossRef]
- Holloszy, J.O. Biochemical Adaptations in Muscle. J. Biol. Chem. 1967, 242, 2278–2282. [Google Scholar] [CrossRef] [PubMed]
- Ornatsky, O.I.; Connor, M.K.; Hood, D.A. Expression of Stress Proteins and Mitochondrial Chaperonins in Chronically Stimulated Skeletal Muscle. Biochem. J. 1995, 311, 119–123. [Google Scholar] [CrossRef]
- Mesbah Moosavi, Z.S.; Hood, D.A. The Unfolded Protein Response in Relation to Mitochondrial Biogenesis in Skeletal Muscle Cells. Am. J. Physiol.-Cell Physiol. 2017, 312, C583–C594. [Google Scholar] [CrossRef]
- Janssen, I.; Heymsfield, S.B.; Wang, Z.; Ross, R. Skeletal Muscle Mass and Distribution in 468 Men and Women Aged 18–88 Yr. J. Appl. Physiol. 2000, 89, 81–88. [Google Scholar] [CrossRef]
- Koltai, E.; Hart, N.; Taylor, A.W.; Goto, S.; Ngo, J.K.; Davies, K.J.A.; Radak, Z. Age-Associated Declines in Mitochondrial Biogenesis and Protein Quality Control Factors Are Minimized by Exercise Training. Am. J. Physiol.-Regul. Integr. Comp. Physiol. 2012, 303, R127–R134. [Google Scholar] [CrossRef]
- Marzetti, E.; Calvani, R.; Coelho-Júnior, H.J.; Landi, F.; Picca, A. Mitochondrial Quantity and Quality in Age-Related Sarcopenia. Int. J. Mol. Sci. 2024, 25, 2052. [Google Scholar] [CrossRef]
- Ladiges, W.; Morton, J.; Blakely, C.; Gale, M. Tissue Specific Expression of PKR Protein Kinase in Aging B6D2F1 Mice. Mech. Ageing Dev. 2000, 114, 123–132. [Google Scholar] [CrossRef]
- Ubaida-Mohien, C.; Lyashkov, A.; Gonzalez-Freire, M.; Tharakan, R.; Shardell, M.; Moaddel, R.; Semba, R.D.; Chia, C.W.; Gorospe, M.; Sen, R.; et al. Discovery Proteomics in Aging Human Skeletal Muscle Finds Change in Spliceosome, Immunity, Proteostasis and Mitochondria. eLife 2019, 8, e49874. [Google Scholar] [CrossRef]
- Michie, K.L.; Kunz, H.E.; Dasari, S.; Lanza, I.R. The Influence of Aging on the Unfolded Protein Response in Human Skeletal Muscle at Rest and after Acute Exercise. Med. Sci. Sports Exerc. 2024, 56, 2135–2145. [Google Scholar] [CrossRef] [PubMed]
- Michel, J.M.; Godwin, J.S.; Kerr, N.R.; Childs, T.E.; Booth, F.W.; Mobley, C.B.; Hughes, D.C.; Roberts, M.D. Skeletal Muscle Atrophy Induced by Aging and Disuse Atrophy Are Strongly Associated with the Upregulation of the Endoplasmic Stress Protein CHOP in Rats. Mol. Biol. Rep. 2025, 52, 322. [Google Scholar] [CrossRef] [PubMed]
- Urbina-Varela, R.; Castillo, N.; Videla, L.A.; Del Campo, A. Impact of Mitophagy and Mitochondrial Unfolded Protein Response as New Adaptive Mechanisms Underlying Old Pathologies: Sarcopenia and Non-Alcoholic Fatty Liver Disease. Int. J. Mol. Sci. 2020, 21, 7704. [Google Scholar] [CrossRef]
- Chen, J.; Kastroll, J.; Bello, F.M.; Pangburn, M.M.; Murali, A.; Smith, P.M.; Rychcik, K.; Loughridge, K.E.; Vandevender, A.M.; Dedousis, N.; et al. Skeletal Muscle Mitochondrial Dysfunction Is Associated with Increased Gdf15 Expression and Circulating GDF15 Levels in Aged Mice. Sci. Rep. 2025, 15, 8101. [Google Scholar] [CrossRef] [PubMed]
- Ebert, S.M.; Dyle, M.C.; Bullard, S.A.; Dierdorff, J.M.; Murry, D.J.; Fox, D.K.; Bongers, K.S.; Lira, V.A.; Meyerholz, D.K.; Talley, J.J.; et al. Identification and Small Molecule Inhibition of an Activating Transcription Factor 4 (ATF4)-Dependent Pathway to Age-Related Skeletal Muscle Weakness and Atrophy. J. Biol. Chem. 2015, 290, 25497–25511. [Google Scholar] [CrossRef] [PubMed]
- Miller, M.J.; Marcotte, G.R.; Basisty, N.; Wehrfritz, C.; Ryan, Z.C.; Strub, M.D.; McKeen, A.T.; Stern, J.I.; Nath, K.A.; Rasmussen, B.B.; et al. The Transcription Regulator ATF4 Is a Mediator of Skeletal Muscle Aging. GeroScience 2023, 45, 2525–2543. [Google Scholar] [CrossRef]
- Von Ruff, Z.D.; Miller, M.J.; Moro, T.; Reidy, P.T.; Ebert, S.M.; Volpi, E.; Adams, C.M.; Rasmussen, B.B. Resistance Exercise Training in Older Men Reduces ATF4-Activated and Senescence-Associated mRNAs in Skeletal Muscle. GeroScience 2025, 47, 4601–4622. [Google Scholar] [CrossRef]
- Hwee, D.T.; Baehr, L.M.; Philp, A.; Baar, K.; Bodine, S.C. Maintenance of Muscle Mass and Load-induced Growth in Muscle RING Finger 1 Null Mice with Age. Aging Cell 2014, 13, 92–101. [Google Scholar] [CrossRef]
- Cordeiro, A.V.; Peruca, G.F.; Braga, R.R.; Brícola, R.S.; Lenhare, L.; Silva, V.R.R.; Anaruma, C.P.; Katashima, C.K.; Crisol, B.M.; Barbosa, L.T.; et al. High-Intensity Exercise Training Induces Mitonuclear Imbalance and Activates the Mitochondrial Unfolded Protein Response in the Skeletal Muscle of Aged Mice. GeroScience 2021, 43, 1513–1518. [Google Scholar] [CrossRef]
- Adams, C.M.; Ebert, S.M.; Dyle, M.C. Role of ATF4 in Skeletal Muscle Atrophy. Curr. Opin. Clin. Nutr. Metab. Care 2017, 20, 164–168. [Google Scholar] [CrossRef]
- Ebert, S.M.; Rasmussen, B.B.; Judge, A.R.; Judge, S.M.; Larsson, L.; Wek, R.C.; Anthony, T.G.; Marcotte, G.R.; Miller, M.J.; Yorek, M.A.; et al. Biology of Activating Transcription Factor 4 (ATF4) and Its Role in Skeletal Muscle Atrophy. J. Nutr. 2022, 152, 926–938. [Google Scholar] [CrossRef]
- Ebert, S.M.; Bullard, S.A.; Basisty, N.; Marcotte, G.R.; Skopec, Z.P.; Dierdorff, J.M.; Al-Zougbi, A.; Tomcheck, K.C.; DeLau, A.D.; Rathmacher, J.A.; et al. Activating Transcription Factor 4 (ATF4) Promotes Skeletal Muscle Atrophy by Forming a Heterodimer with the Transcriptional Regulator C/EBPβ. J. Biol. Chem. 2020, 295, 2787–2803. [Google Scholar] [CrossRef] [PubMed]
- Ebert, S.M.; Dyle, M.C.; Kunkel, S.D.; Bullard, S.A.; Bongers, K.S.; Fox, D.K.; Dierdorff, J.M.; Foster, E.D.; Adams, C.M. Stress-Induced Skeletal Muscle Gadd45a Expression Reprograms Myonuclei and Causes Muscle Atrophy. J. Biol. Chem. 2012, 287, 27290–27301. [Google Scholar] [CrossRef] [PubMed]
- Bullard, S.A.; Seo, S.; Schilling, B.; Dyle, M.C.; Dierdorff, J.M.; Ebert, S.M.; DeLau, A.D.; Gibson, B.W.; Adams, C.M. Gadd45a Protein Promotes Skeletal Muscle Atrophy by Forming a Complex with the Protein Kinase MEKK4. J. Biol. Chem. 2016, 291, 17496–17509. [Google Scholar] [CrossRef]
- Marcotte, G.R.; Miller, M.J.; Kunz, H.E.; Ryan, Z.C.; Strub, M.D.; Vanderboom, P.M.; Heppelmann, C.J.; Chau, S.; Von Ruff, Z.D.; Kilroe, S.P.; et al. GADD45A Is a Mediator of Mitochondrial Loss, Atrophy, and Weakness in Skeletal Muscle. JCI Insight 2023, 8, e171772. [Google Scholar] [CrossRef]
- Glass, D.J. Skeletal Muscle Hypertrophy and Atrophy Signaling Pathways. Int. J. Biochem. Cell Biol. 2005, 37, 1974–1984. [Google Scholar] [CrossRef]
- Miyake, Z.; Takekawa, M.; Ge, Q.; Saito, H. Activation of MTK1/MEKK4 by GADD45 through Induced N-C Dissociation and Dimerization-Mediated Trans Autophosphorylation of the MTK1 Kinase Domain. Mol. Cell. Biol. 2007, 27, 2765–2776. [Google Scholar] [CrossRef]
- Inoue, Y.; Kawachi, S.; Ohkubo, T.; Nagasaka, M.; Ito, S.; Fukuura, K.; Itoh, Y.; Ohoka, N.; Morishita, D.; Hayashi, H. The CDK Inhibitor P21 Is a Novel Target Gene of ATF 4 and Contributes to Cell Survival under ER Stress. FEBS Lett. 2017, 591, 3682–3691. [Google Scholar] [CrossRef] [PubMed]
- Fox, D.K.; Ebert, S.M.; Bongers, K.S.; Dyle, M.C.; Bullard, S.A.; Dierdorff, J.M.; Kunkel, S.D.; Adams, C.M. P53 and ATF4 Mediate Distinct and Additive Pathways to Skeletal Muscle Atrophy during Limb Immobilization. Am. J. Physiol.-Endocrinol. Metab. 2014, 307, E245–E261. [Google Scholar] [CrossRef] [PubMed]
- Bongers, K.S.; Fox, D.K.; Kunkel, S.D.; Stebounova, L.V.; Murry, D.J.; Pufall, M.A.; Ebert, S.M.; Dyle, M.C.; Bullard, S.A.; Dierdorff, J.M.; et al. Spermine Oxidase Maintains Basal Skeletal Muscle Gene Expression and Fiber Size and Is Strongly Repressed by Conditions That Cause Skeletal Muscle Atrophy. Am. J. Physiol.-Endocrinol. Metab. 2015, 308, E144–E158. [Google Scholar] [CrossRef] [PubMed]
- Abbas, T.; Dutta, A. P21 in Cancer: Intricate Networks and Multiple Activities. Nat. Rev. Cancer 2009, 9, 400–414. [Google Scholar] [CrossRef] [PubMed]
- Besson, A.; Dowdy, S.F.; Roberts, J.M. CDK Inhibitors: Cell Cycle Regulators and Beyond. Dev. Cell 2008, 14, 159–169. [Google Scholar] [CrossRef]
- Casero, R.A.; Pegg, A.E. Polyamine Catabolism and Disease. Biochem. J. 2009, 421, 323–338. [Google Scholar] [CrossRef] [PubMed]
- Cervelli, M.; Bellini, A.; Bianchi, M.; Marcocci, L.; Nocera, S.; Polticelli, F.; Federico, R.; Amendola, R.; Mariottini, P. Mouse Spermine Oxidase Gene Splice Variants: Nuclear Subcellular Localization of a Novel Active Isoform. Eur. J. Biochem. 2004, 271, 760–770. [Google Scholar] [CrossRef]
- Fan, J.; Yang, X.; Li, J.; Shu, Z.; Dai, J.; Liu, X.; Li, B.; Jia, S.; Kou, X.; Yang, Y.; et al. Spermidine Coupled with Exercise Rescues Skeletal Muscle Atrophy from D-Gal-Induced Aging Rats through Enhanced Autophagy and Reduced Apoptosis via AMPK-FOXO3a Signal Pathway. Oncotarget 2017, 8, 17475–17490. [Google Scholar] [CrossRef]
- Edgerton, V.R.; Roy, R.R.; Allen, D.L.; Monti, R.J. Adaptations in Skeletal Muscle Disuse or Decreased-Use Atrophy. Am. J. Phys. Med. Rehabil. 2002, 81, S127–S147. [Google Scholar] [CrossRef]
- Nunes, E.A.; Stokes, T.; McKendry, J.; Currier, B.S.; Phillips, S.M. Disuse-Induced Skeletal Muscle Atrophy in Disease and Nondisease States in Humans: Mechanisms, Prevention, and Recovery Strategies. Am. J. Physiol.-Cell Physiol. 2022, 322, C1068–C1084. [Google Scholar] [CrossRef]
- Memme, J.M.; Oliveira, A.N.; Hood, D.A. P53 Regulates Skeletal Muscle Mitophagy and Mitochondrial Quality Control Following Denervation-Induced Muscle Disuse. J. Biol. Chem. 2022, 298, 101540. [Google Scholar] [CrossRef] [PubMed]
- Wang, L.; Pang, X.; Li, S.; Li, W.; Chen, X.; Zhang, P. ER Stress Is Activated and Involved in Disuse-Induced Muscle Atrophy. Front. Biosci.-Landmark 2023, 28, 136. [Google Scholar] [CrossRef] [PubMed]
- Gallot, Y.S.; Bohnert, K.R. Confounding Roles of ER Stress and the Unfolded Protein Response in Skeletal Muscle Atrophy. Int. J. Mol. Sci. 2021, 22, 2567. [Google Scholar] [CrossRef]
- Ibrahim, Z.; Ramachandran, G.; El-Huneidi, W.; Elmoselhi, A.; Qaisar, R. Suppression of Endoplasmic Reticulum Stress Prevents Disuse Muscle Atrophy in a Mouse Model of Microgravity. Life Sci. Space Res. 2022, 34, 45–52. [Google Scholar] [CrossRef] [PubMed]
- Fox, D.K.; Ebert, S.M.; Bongers, K.S.; Dyle, M.C.; Bullard, S.A.; Dierdorff, J.M.; Kunkel, S.D.; Adams, C.M. P53 and ATF4 Mediate Distinct and Additive Pathways to Skeletal Muscle Atrophy during Limb Immobilization. Am. J. Physiol.-Endocrinol. Metab. 2014, 307, E245–E261. [Google Scholar] [CrossRef]
- Wang, L.; Pang, X.; Li, S.; Li, W.; Chen, X.; Zhang, P. ER Stress Is Activated and Involved in Disuse-Induced Muscle Atrophy. Front. Biosci.-Landmark 2023, 28, 136. [Google Scholar] [CrossRef]
- Mcglory, C.; Gorissen, S.H.M.; Kamal, M.; Bahniwal, R.; Hector, A.J.; Baker, S.K.; Chabowski, A.; Phillips, S.M. Omega-3 Fatty Acid Supplementation Attenuates Skeletal Muscle Disuse Atrophy during Two Weeks of Unilateral Leg Immobilization in Healthy Young Women. FASEB J. 2019, 33, 4586–4597. [Google Scholar] [CrossRef]
- Stevenson, E.J.; Giresi, P.G.; Koncarevic, A.; Kandarian, S.C. Global Analysis of Gene Expression Patterns during Disuse Atrophy in Rat Skeletal Muscle. J. Physiol. 2003, 551, 33–48. [Google Scholar] [CrossRef]
- Cussonneau, L.; Coudy-Gandilhon, C.; Deval, C.; Chaouki, G.; Djelloul-Mazouz, M.; Delorme, Y.; Hermet, J.; Gauquelin-Koch, G.; Polge, C.; Taillandier, D.; et al. Induction of ATF4-Regulated Atrogenes Is Uncoupled from Muscle Atrophy during Disuse in Halofuginone-Treated Mice and in Hibernating Brown Bears. Int. J. Mol. Sci. 2022, 24, 621. [Google Scholar] [CrossRef] [PubMed]
- Bongers, K.S.; Fox, D.K.; Ebert, S.M.; Kunkel, S.D.; Dyle, M.C.; Bullard, S.A.; Dierdorff, J.M.; Adams, C.M. Skeletal Muscle Denervation Causes Skeletal Muscle Atrophy through a Pathway That Involves Both Gadd45a and HDAC4. Am. J. Physiol.-Endocrinol. Metab. 2013, 305, E907–E915. [Google Scholar] [CrossRef]
- Zhang, G.; Wang, X.; Li, C.; Li, Q.; An, Y.A.; Luo, X.; Deng, Y.; Gillette, T.G.; Scherer, P.E.; Wang, Z.V. Integrated Stress Response Couples Mitochondrial Protein Translation With Oxidative Stress Control. Circulation 2021, 144, 1500–1515. [Google Scholar] [CrossRef] [PubMed]
- Miyake, M.; Nomura, A.; Ogura, A.; Takehana, K.; Kitahara, Y.; Takahara, K.; Tsugawa, K.; Miyamoto, C.; Miura, N.; Sato, R.; et al. Skeletal Muscle-specific Eukaryotic Translation Initiation Factor 2α Phosphorylation Controls Amino Acid Metabolism and Fibroblast Growth Factor 21-mediated Non-cell-autonomous Energy Metabolism. FASEB J. 2016, 30, 798–812. [Google Scholar] [CrossRef]
- Kunkel, S.D.; Suneja, M.; Ebert, S.M.; Bongers, K.S.; Fox, D.K.; Malmberg, S.E.; Alipour, F.; Shields, R.K.; Adams, C.M. mRNA Expression Signatures of Human Skeletal Muscle Atrophy Identify a Natural Compound That Increases Muscle Mass. Cell Metab. 2011, 13, 627–638. [Google Scholar] [CrossRef]
- Cordeiro, A.V.; Brícola, R.S.; Braga, R.R.; Lenhare, L.; Silva, V.R.R.; Anaruma, C.P.; Katashima, C.K.; Crisol, B.M.; Simabuco, F.M.; Silva, A.S.R.; et al. Aerobic Exercise Training Induces the Mitonuclear Imbalance and UPRmt in the Skeletal Muscle of Aged Mice. J. Gerontol. Ser. A 2020, 75, 2258–2261. [Google Scholar] [CrossRef] [PubMed]
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2026 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).



