Submitted:
04 April 2026
Posted:
06 April 2026
You are already at the latest version
Abstract
Keywords:
1. Introduction
1.1. Limitations of Conventional Modeling in Heterogeneous Populations
1.2. Physiological Senescence and Pharmacokinetic Deviations
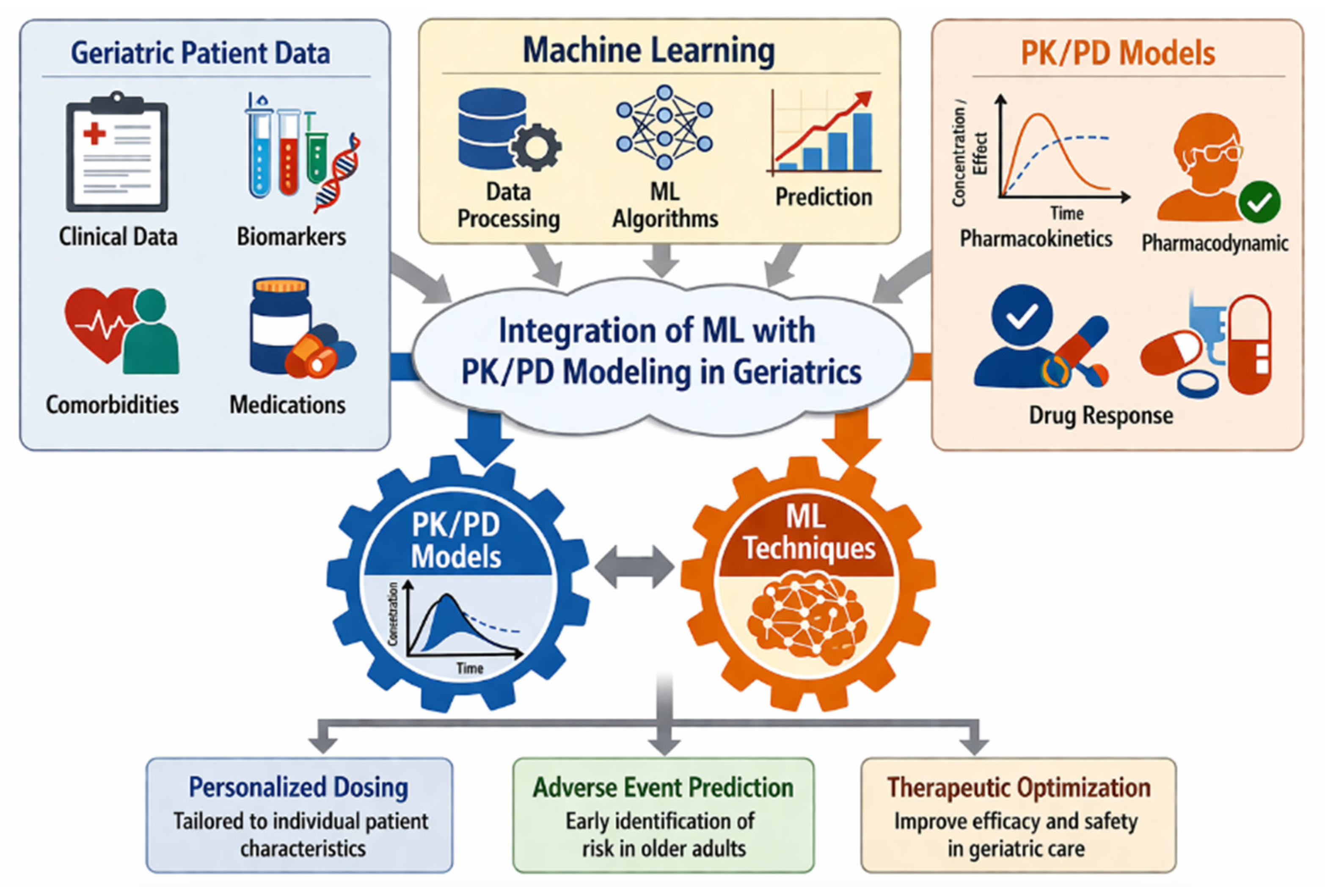
1.3. AI Role in Improving Precision Therapy
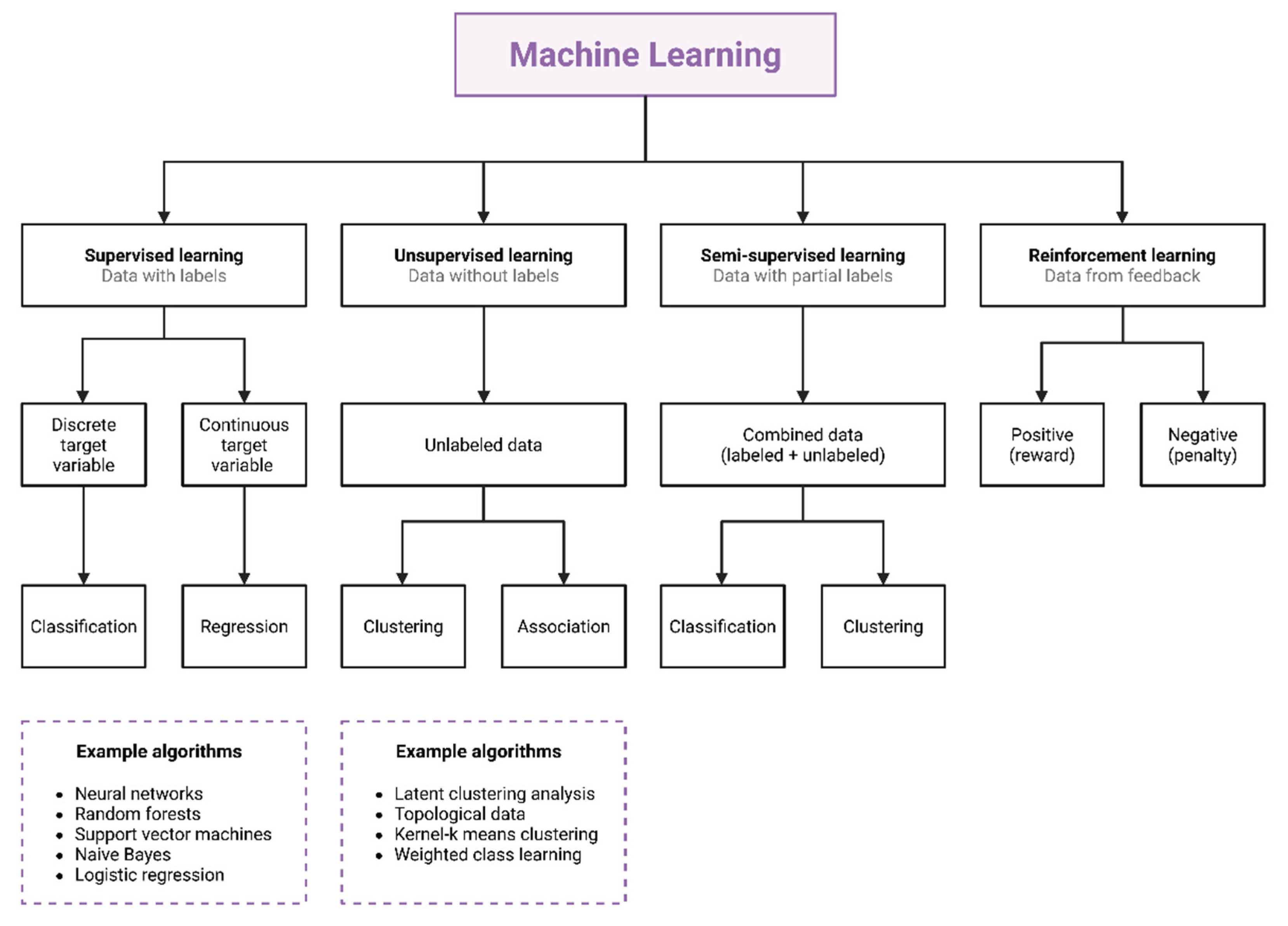
1.4. The Machine Learning Paradigm Shift
1.5. Aims of the Review
1.6. Methods
2. Age-Related PK/PD Challenges in Elderly Patients
2.1. Physiological Senescence and Organ Functional Decline
2.2. Pharmacokinetic Variability in Geriatric Populations
2.3. Pharmacodynamic Sensitivity and Immuno-Senescence
2.4. The Burden of Multimorbidity and Polypharmacy
2.5. Geriatric-Specific Determinants
3. Traditional PK/PD Modelling Approaches and Limitations
3.1. Mechanistic Basis and Compartmental Modeling
3.2. Nonlinear Mixed-Effects (NLME)
- Fixed effects
- 2.
- Random effects
3.3. Bayesian Forecasting and a Priori Dose Optimization
3.4. Limitations and Challenges of Traditional PK/PD Modelling
- Time and labor demands: Developing population PK models involves a progressive, fundamentally manual process that requires technical expertise. Evaluating different structural models and examining covariate effects is time-consuming, and the overall process of model development and validation can take weeks to months [16,34].
- Physiological oversimplification: To maintain computational tractability, conventional models often rely on one- or two-compartment structures. Despite their practicality, such formulations may induce model misspecification. when representing complex pharmacokinetic behavior, particularly for drugs exhibiting nonlinear clearance or heterogeneous tissue distribution [33].
- Limitations in covariate selection: Covariate selection procedures become inefficient because They are poorly suited to high-dimensional datasets, as a result, they tend to perform poorly when there is strong collinearity among predictors [33].
- Limited generalizability and risk of bias: Models developed and evaluated in highly selected populations exhibit limited generalizability to other clinical groups. For example, several meropenem population models have shown limited predictive accuracy in patients undergoing continuous renal replacement therapy (CRRT) due to the absence of covariates that effectively reflect physiological alteration. Although mechanistic statements provide a useful structural framework, they can lead to systematic bias if the biological mechanisms are inaccurate, incomplete, or not applicable to the target population [16,35].
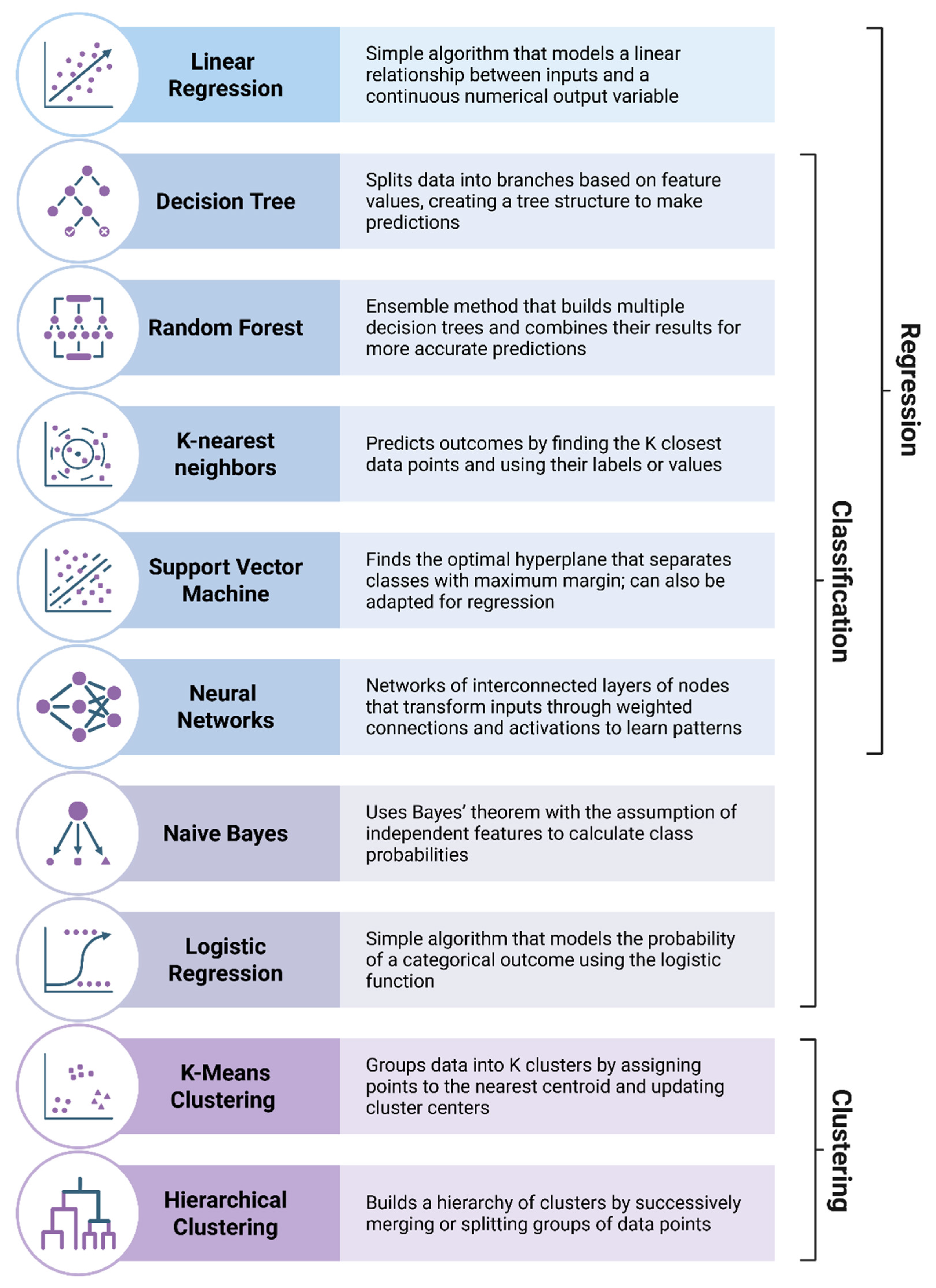
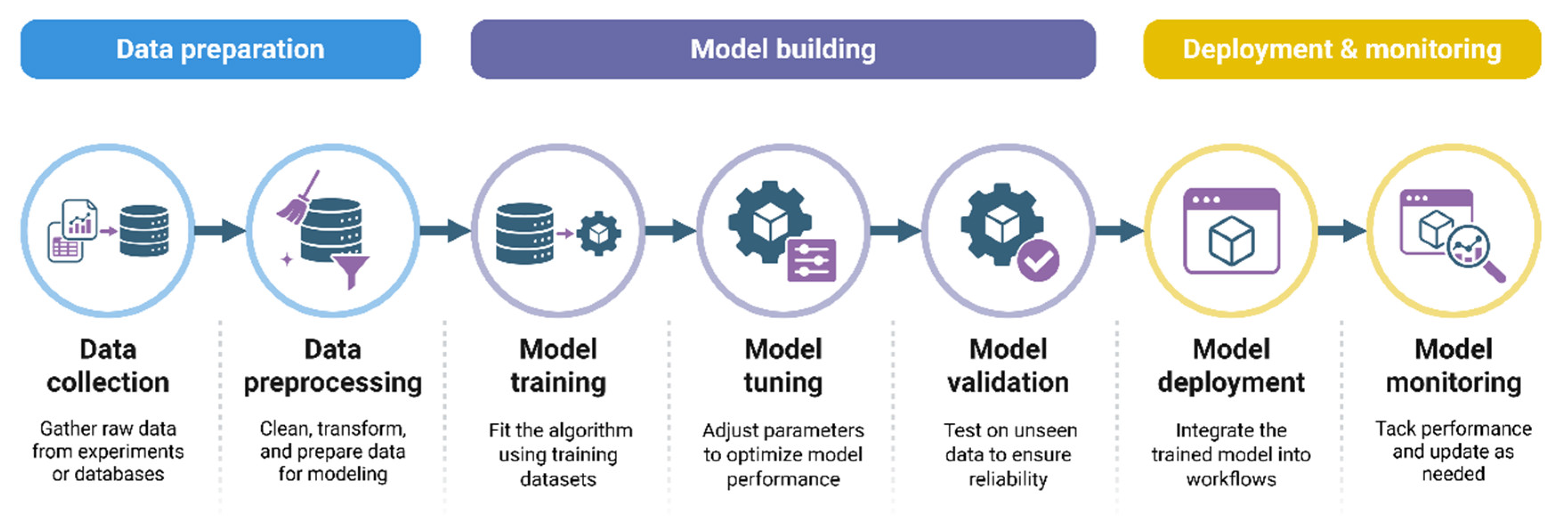
4. Machine Learning Methods Applied to PK/PD Modelling
5. Data Requirements and Feature Engineering in Geriatric Populations
6. Performance of ML-Enhanced PK/PD Models
6.1. Superiority in Predictive Accuracy and Bias Reduction
6.2. Uncovering Non-Linear Covariate Interactions
6.3. ML-Guided Model Selection and Ensembling
6.4. Performance Variability Across Clinical Data Availability
7. Impact of ML on the Accuracy of Medication Regimen Design
7.1. Superiority of Nonlinear ML Models over Traditional Methods
7.2. Enhancing Accuracy Through ML-PopPK Hybridization
7.3. Feature Importance and Clinical Interpretability
7.4. Efficiency in Computational Workflows and Run-Times
8. Clinical Implications for Personalized Dosing in the Elderly
8.1. Transitioning to Individual-Level Therapeutic Targets
8.2. Adaptive Dosing via Reinforcement Learning (RL)
8.3. Decision Support Tools and Point-of-Care Integration
8.4. Reducing Adverse Drug Reactions and Enhancing Safety
8.5. Geriatric-Specific Implementation Challenges: Limitations of Clinical Trials in the Elderly
9. Current Limitations and Ethical Considerations
9.1. Challenges in Generalizability and Site-Specific Biases
9.2. Algorithmic Bias and Demographic Disparities
- Gender and race bias: [88] found that both standard clinical tools (like Pooled Cohort Risk Equations) and ML models were biased against women, resulting in lower true positive rates and lower positive prediction rates compared to men.
- Underdiagnosis bias: In radiological deep learning models, under-served patient populations (including female, Black, Hispanic, and lower socioeconomic status patients (proxied by Medicaid insurance)) are consistently and selectively underdiagnosed. Which, in turn, is particularly harmful because underdiagnosis falsely labels a sick individual as healthy, potentially delaying or denying access to critical life-saving medication adjustments or triage priority [87].
- Intersectional vulnerabilities: These biases are often found in intersectional subgroups. For example, Hispanic female patients may face higher rates of algorithmic underdiagnosis bias than white female patients, which would in turn conclude that simple demographic checks are insufficient to capture the depth of algorithmic unfairness [87].
9.3. Label Selection and Proxy Bias
9.4. Explainability and Clinician Trust
9.5. Data Requirements and Lack of Standardized Guidelines
- 68% of developers claimed that a lack of fair data as a primary reason for algorithmic bias.
- 49% referred to a lack of guidelines or recommendations for technically implementing fairness in clinical applications
- 50% of projects rely on data from only one center, limiting the model's exposure to diverse patient populations and acquisition protocols, which would in return contribute to the site-specific bias mentioned before.
10. Future Directions
10.1. Incorporating Multi-Modal and High-Resolution Data Sources
10.2. Real-Time Dynamic Decision Support and Bedside Integration
10.3. Methodological Advancements in Algorithmic Robustness
10.4. Transition to Prospective Clinical Validation
11. Conclusions
Abbreviations
| PK | Pharmacokinetic |
| PD | Pharmacodynamic |
| AI | Artificial Intelligence |
| Vd | Volume of Distribution |
| GFR | Glomerular Filtration Rate |
| CYP450 | Cytochrome P450 |
| CNS | Central Nervous System |
| SSRI | Selective Serotonin Reuptake Inhibitors |
| GI | Gastro-Intestinal |
| ADEs | Adverse Drug Events |
| DDIs | Drug-Drug Interactions |
| ADRs | Adverse Drug Reactions |
| ODEs | Ordinary differential equations |
| ADME | Absorption, distribution, metabolism, and elimination |
| CL | Clearance |
| E-max | maximum drug effect |
| NLME | Nonlinear Mixed-Effects |
| PopPK | Population PK |
| IIV | Inter-individual variability |
| IOV | Inter-occasion variability |
| RUV | Residual unexplained variability |
| MIPD | Model-Informed Precision Dosing |
| TDM | Therapeutic drug monitoring |
| CRRT | Continuous renal replacement therapy |
| ML | machine learning |
| RF | Random Forest |
| GDA | General discriminant analysis |
| TPR | true positive rate |
| INR | International Normalized Ratio |
| aPTT | activated partial thromboplastin time |
| SNN | Shallow Neural Network |
| XGBoost | Extreme Gradient Boosting |
| SVM | Support Vector Machine |
| AdaBoost | Adaptive Boosting |
| MLP | Multilayer Perceptron |
| EHR | Electronic health records |
| DT | Decision Tree |
| MAPE | Mean absolute percentage error |
| RMSE | Root Mean Squared Error |
| CrCl | Creatinine Clearance |
| BNP | B-type Natriuretic Peptide |
| CRP | C-reactive protein |
| MAP | Maximum a posteriori |
| ANN | Artificial Neural Networks |
| SHAP | Shapley Additive exPlanations |
| RL | Reinforcement Learning |
| MIRL | Model-Informed RL |
| FGFR | Fibroblast Growth Factor Receptor |
| CHR | Complete Hematologic Response |
| DOACs | Direct Oral Anti-Coagulants |
| CDSS | Clinical Decision Support System |
| PACS | Picture Archiving and Communication System |
| ICU | Intensive Care Unit |
| FHIR | Fast Healthcare Interoperability Resources |
| SNOMED CT | Systematized Nomenclature of Medicine Clinical Terms |
| FTL | Federated Transfer Learning |
| GDPR | General Data Protection Regulation |
References
- Gettman, D. Integrating Pharmacokinetics/Pharmacodynamics (PK/PD) with Artificial Intelligence to Advance Predictive Modeling of Therapeutic Windows. 2023. [Google Scholar]
- Mangoni, A.A.; Jackson, S.H.D. Age-Related Changes in Pharmacokinetics and Pharmacodynamics: Basic Principles and Practical Applications. Br. J. Clin. Pharmacol. 2004, 57, 6–14. [Google Scholar] [CrossRef]
- Yang, K.; Gonzalez, D.; Woodhead, J.L.; Bhargava, P.; Ramanathan, M. Leveraging In Silico and Artificial Intelligence Models to Advance Drug Disposition and Response Predictions Across the Lifespan. Clin. Transl. Sci. 2025, 18, e70272. [Google Scholar] [CrossRef]
- Papadopoulos, D.; Magliozzi, R.; Mitsikostas, D.D.; Gorgoulis, V.G.; Nicholas, R.S. Aging, Cellular Senescence, and Progressive Multiple Sclerosis. Front. Cell. Neurosci. 2020, 14, 178. [Google Scholar] [CrossRef] [PubMed]
- Klotz, U. Pharmacokinetics and Drug Metabolism in the Elderly. Drug Metab. Rev. 2009, 41, 67–76. [Google Scholar] [CrossRef] [PubMed]
- Yang, K.; Gonzalez, D.; Woodhead, J.L.; Bhargava, P.; Ramanathan, M. Leveraging in Silico and Artificial Intelligence Models to Advance Drug Disposition and Response Predictions across the Lifespan. Clin. Transl. Sci. 2025, 18, e70272. [Google Scholar] [CrossRef] [PubMed]
- Keutzer, L.; You, H.; Farnoud, A.; Nyberg, J.; Wicha, S.G.; Maher-Edwards, G.; Vlasakakis, G.; Moghaddam, G.K.; Svensson, E.M.; Menden, M.P. Machine Learning and Pharmacometrics for Prediction of Pharmacokinetic Data: Differences, Similarities and Challenges Illustrated with Rifampicin. Pharmaceutics 2022, 14, 1530. [Google Scholar] [CrossRef] [PubMed]
- Stankevičiūtė, K.; Woillard, J.-B.; Peck, R.W.; Marquet, P.; van der Schaar, M. Bridging the Worlds of Pharmacometrics and Machine Learning. Clin. Pharmacokinet. 2023, 62, 1551–1565. [Google Scholar] [CrossRef]
- Mangoni, A.A.; Jackson, S.H. Age-related Changes in Pharmacokinetics and Pharmacodynamics: Basic Principles and Practical Applications. Br. J. Clin. Pharmacol. 2004, 57, 6–14. [Google Scholar] [CrossRef]
- Zhu, X.; Huang, W.; Lu, H.; Wang, Z.; Ni, X.; Hu, J.; Deng, S.; Tan, Y.; Li, L.; Zhang, M. A Machine Learning Approach to Personalized Dose Adjustment of Lamotrigine Using Noninvasive Clinical Parameters. Sci. Rep. 2021, 11, 5568. [Google Scholar] [CrossRef]
- Wicha, S.G.; Märtson, A.; Nielsen, E.I.; Koch, B.C.; Friberg, L.E.; Alffenaar, J.; Minichmayr, I.K. International Society of Anti-Infective Pharmacology (ISAP), the P. study group of the E.S. of C.M., Infectious Diseases (EPASG) From Therapeutic Drug Monitoring to Model-informed Precision Dosing for Antibiotics. Clin. Pharmacol. Ther. 2021, 109, 928–941. [Google Scholar] [CrossRef]
- Dudarski, L. Harnessing Machine Learning for Predictive Pharmacokinetics:Revolutionizing Drug Development and Personalized Medicine. Curr. Issues Pharm. Med. Sci. 2025, 38, 125–131. [Google Scholar] [CrossRef]
- Parvathaneni, M.; Awol, A.K.; Kumari, M.; Lan, K.; Lingam, M. Application of Artificial Intelligence and Machine Learning in Drug Discovery and Development. J. Drug Deliv. Ther. 2023, 13, 151–158. [Google Scholar] [CrossRef]
- Mao, J.; Chen, Y.; Xu, L.; Chen, W.; Chen, B.; Fang, Z.; Qin, W.; Zhong, M. Applying Machine Learning to the Pharmacokinetic Modeling of Cyclosporine in Adult Renal Transplant Recipients: A Multi-Method Comparison. Front. Pharmacol. 2022, 13, 1016399. [Google Scholar] [CrossRef] [PubMed]
- Chen, Y.; Liu, R.; Ma, P.; Shang, S.; Yu, M.; Xie, L.; Zhao, L.; Huang, Y. A Real-Time Plasma Concentration Prediction Model for Voriconazole in Elderly Patients via Machine Learning Combined with Population Pharmacokinetics 2024.
- Keutzer, L.; You, H.; Farnoud, A.; Nyberg, J.; Wicha, S.G.; Maher-Edwards, G.; Vlasakakis, G.; Moghaddam, G.K.; Svensson, E.M.; Menden, M.P.; et al. Machine Learning and Pharmacometrics for Prediction of Pharmacokinetic Data: Differences, Similarities and Challenges Illustrated with Rifampicin. Pharmaceutics 2022, 14, 1530. [Google Scholar] [CrossRef]
- Ma, P.; Ma, H.; Liu, R.; Wen, H.; Li, H.; Huang, Y.; Li, Y.; Xiong, L.; Xie, L.; Wang, Q. Prediction of Vancomycin Plasma Concentration in Elderly Patients Based on Multi-Algorithm Mining Combined with Population Pharmacokinetics. Sci. Rep. 2024, 14, 27165. [Google Scholar] [CrossRef]
- Sarker, I.H. Machine Learning: Algorithms, Real-World Applications and Research Directions. SN Comput. Sci. 2021, 2, 1–21. [Google Scholar] [CrossRef] [PubMed]
- Stankevičiūtė, K.; Woillard, J.-B.; Peck, R.W.; Marquet, P.; van der Schaar, M. Bridging the Worlds of Pharmacometrics and Machine Learning. Clin. Pharmacokinet. 2023, 62, 1551–1565. [Google Scholar] [CrossRef] [PubMed]
- Kneller, L.A.; Hempel, G. Modelling Age-Related Changes in the Pharmacokinetics of Risperidone and 9-Hydroxyrisperidone in Different CYP2D6 Phenotypes Using a Physiologically Based Pharmacokinetic Approach. Pharm. Res. 2020, 37, 110. [Google Scholar] [CrossRef]
- Błeszyńska, E.; Wierucki, Ł.; Zdrojewski, T.; Renke, M.; Błeszyńska, E.; Wierucki, Ł.; Zdrojewski, T.; Renke, M. Pharmacological Interactions in the Elderly. Medicina (Mex.) 2020, 56. [Google Scholar] [CrossRef]
- Zubair, H. Personalized Medicine in Aging: Strategies and Specific Needs of Older Adults. 2025, 7. [Google Scholar]
- Woodford, H.J. Polypharmacy in Older Patients. Br. J. Hosp. Med. 2024, 85, 1–12. [Google Scholar] [CrossRef]
- Calderón-Larrañaga, A.; Vetrano, D.L.; Onder, G.; Gimeno-Feliu, L.A.; Coscollar-Santaliestra, C.; Carfí, A.; Pisciotta, M.S.; Angleman, S.; Melis, R.J.F.; Santoni, G.; et al. Assessing and Measuring Chronic Multimorbidity in the Older Population: A Proposal for Its Operationalization. J. Gerontol. Ser. A 2017, 72, 1417–1423. [Google Scholar] [CrossRef] [PubMed]
- Michael, A.E.; Michael, N.; Erfurth, A.; Kujovic, M. Clozapine for the Treatment of Aggressiveness and Agitation in Advanced Dementia. Psychogeriatrics 2023, 23, 963–972. [Google Scholar] [CrossRef]
- Clegg, A.; Young, J.; Iliffe, S.; Rikkert, M.O.; Rockwood, K. Frailty in Elderly People. The lancet 2013, 381, 752–762. [Google Scholar] [CrossRef] [PubMed]
- Fried, L.P.; Tangen, C.M.; Walston, J.; Newman, A.B.; Hirsch, C.; Gottdiener, J.; Seeman, T.; Tracy, R.; Kop, W.J.; Burke, G. Frailty in Older Adults: Evidence for a Phenotype. J. Gerontol. A. Biol. Sci. Med. Sci. 2001, 56, M146–M157. [Google Scholar] [CrossRef]
- Hilmer, S.N.; McLachlan, A.J.; Le Couteur, D.G. Clinical Pharmacology in the Geriatric Patient. Fundam. Clin. Pharmacol. 2007, 21, 217–230. [Google Scholar] [CrossRef]
- Turnheim, K. When Drug Therapy Gets Old: Pharmacokinetics and Pharmacodynamics in the Elderly. Exp. Gerontol. 2003, 38, 843–853. [Google Scholar] [CrossRef]
- Hilmer, S.N.; McLachlan, A.J.; Le Couteur, D.G. Clinical Pharmacology in the Geriatric Patient. Fundam. Clin. Pharmacol. 2007, 21, 217–230. [Google Scholar] [CrossRef] [PubMed]
- Kneller, L.A.; Hempel, G. Modelling Age-Related Changes in the Pharmacokinetics of Risperidone and 9-Hydroxyrisperidone in Different CYP2D6 Phenotypes Using a Physiologically Based Pharmacokinetic Approach: Kneller and Hempel. Pharm. Res. 2020, 37, 110. [Google Scholar] [CrossRef]
- Ho, R.H.; Kim, R.B. Transporters and Drug Therapy: Implications for Drug Disposition and Disease. Clin. Pharmacol. Ther. 2005, 78, 260–277. [Google Scholar] [CrossRef]
- Stankevičiūtė, K.; Woillard, J.-B.; Peck, R.W.; Marquet, P.; van der Schaar, M. Bridging the Worlds of Pharmacometrics and Machine Learning. Clin. Pharmacokinet. 2023, 62, 1551–1565. [Google Scholar] [CrossRef] [PubMed]
- Huang, Z.; Denti, P.; Mistry, H.; Kloprogge, F. Machine Learning and Artificial Intelligence in PK-PD Modeling: Fad, Friend, or Foe? Clin. Pharmacol. Ther. 2024, 115, 652–654. [Google Scholar] [CrossRef]
- Yang, N.; Wang, J.; Xie, Y.; Ding, J.; Wu, C.; Liu, J.; Pei, Q. External Evaluation of Population Pharmacokinetic Models to Inform Precision Dosing of Meropenem in Critically Ill Patients. Front. Pharmacol. 2022, 13. [Google Scholar] [CrossRef]
- Mtshali, S.; Jacobs, B. Machine Learning-Based Prediction of Pharmacokinetic Parameters for Individualized Drug Dosage Optimization. Int. J. Inf. Technol. 2024, 17. [Google Scholar] [CrossRef]
- Xue, L.; He, S.; Singla, R.K.; Qin, Q.; Ding, Y.; Liu, L.; Ding, X.; Bediaga-Bañeres, H.; Arrasate, S.; Durado-Sanchez, A.; et al. Machine Learning Guided Prediction of Warfarin Blood Levels for Personalized Medicine Based on Clinical Longitudinal Data from Cardiac Surgery Patients: A Prospective Observational Study. Int. J. Surg. Lond. Engl. 2024, 110, 6528–6540. [Google Scholar] [CrossRef]
- Su, L.; Liu, C.; Li, D.; He, J.; Zheng, F.; Jiang, H.; Wang, H.; Gong, M.; Hong, N.; Zhu, W.; et al. Toward Optimal Heparin Dosing by Comparing Multiple Machine Learning Methods: Retrospective Study. JMIR Med. Inform. 2020, 8, e17648. [Google Scholar] [CrossRef] [PubMed]
- Hernández-Mustieles, M.A.; Lima-Carmona, Y.E.; Pacheco-Ramírez, M.A.; Mendoza-Armenta, A.A.; Romero-Gómez, J.E.; Cruz-Gómez, C.F.; Rodríguez-Alvarado, D.C.; Arceo, A.; Cruz-Garza, J.G.; Ramírez-Moreno, M.A. Wearable Biosensor Technology in Education: A Systematic Review. Sensors 2024, 24, 2437. [Google Scholar] [CrossRef] [PubMed]
- Landi, I.; Glicksberg, B.S.; Lee, H.-C.; Cherng, S.; Landi, G.; Danieletto, M.; Dudley, J.T.; Furlanello, C.; Miotto, R. Deep Representation Learning of Electronic Health Records to Unlock Patient Stratification at Scale. NPJ Digit. Med. 2020, 3, 96. [Google Scholar] [CrossRef]
- Wimmer, B.C.; Dent, E.; Visvanathan, R.; Wiese, M.D.; Johnell, K.; Chapman, I.; Bell, J.S. Polypharmacy and Medication Regimen Complexity as Factors Associated with Hospital Discharge Destination among Older People: A Prospective Cohort Study. Drugs Aging 2014, 31, 623–630. [Google Scholar] [CrossRef]
- Syed, M.; Syed, S.; Sexton, K.; Syeda, H.B.; Garza, M.; Zozus, M.; Syed, F.; Begum, S.; Syed, A.U.; Sanford, J. Application of Machine Learning in Intensive Care Unit (ICU) Settings Using MIMIC Dataset: Systematic Review; MDPI, 2021; Vol. 8, p. 16. [Google Scholar] [CrossRef]
- Rind, S.; Zhang, N.; Khan, W.U.; Zhang, Q. Sweat-Based Wearable Biosensors: A New Era of Continuous, Noninvasive Health Monitoring and Diagnostics. Wearable Electron. 2025, 2, 323–343. [Google Scholar] [CrossRef]
- Zolnoori, M.; Barrón, Y.; Song, J.; Noble, J.; Burgdorf, J.; Ryvicker, M.; Topaz, M. HomeADScreen: Developing Alzheimer’s Disease and Related Dementia Risk Identification Model in Home Healthcare. Int. J. Med. Inf. 2023, 177, 105146. [Google Scholar] [CrossRef] [PubMed]
- Rajkomar, A.; Oren, E.; Chen, K.; Dai, A.M.; Hajaj, N.; Hardt, M.; Liu, P.J.; Liu, X.; Marcus, J.; Sun, M.; et al. Scalable and Accurate Deep Learning with Electronic Health Records. NPJ Digit. Med. 2018, 1, 18. [Google Scholar] [CrossRef]
- Gibson, L.L.; Mueller, C.; Stewart, R.; Aarsland, D. Characteristics Associated with Progression to Probable Dementia with Lewy Bodies in a Cohort with Very Late-Onset Psychosis. Psychol. Med. 2024, 54, 3479–3488. [Google Scholar] [CrossRef]
- Mataraso, S.J.; Espinosa, C.A.; Seong, D.; Reincke, S.M.; Berson, E.; Reiss, J.D.; Kim, Y.; Ghanem, M.; Shu, C.-H.; James, T.; et al. A Machine Learning Approach to Leveraging Electronic Health Records for Enhanced Omics Analysis. Nat. Mach. Intell. 2025, 7, 293–306. [Google Scholar] [CrossRef]
- Bräm, D.S.; Nahum, U.; Atkinson, A.; Koch, G.; Pfister, M. Evaluation of Machine Learning Methods for Covariate Data Imputation in Pharmacometrics. CPT Pharmacomet. Syst. Pharmacol. 2022, 11, 1638–1648. [Google Scholar] [CrossRef]
- Bräm, D.S.; Nahum, U.; Atkinson, A.; Koch, G.; Pfister, M. Evaluation of Machine Learning Methods for Covariate Data Imputation in Pharmacometrics. CPT Pharmacomet. Syst. Pharmacol. 2022, 11, 1638–1648. [Google Scholar] [CrossRef]
- Ali, A.; Alsayed, A.R.; Seder, N.; Jarrar, Y.; Altabanjeh, R.H.; Zihlif, M.; Ata, O.A.; Samara, A.; Zihlif, M. Unveiling Etiology and Mortality Risks in Community-Acquired Pneumonia: A Machine Learning Approach. Biomol. Biomed. 2025, 26, 333. [Google Scholar] [CrossRef] [PubMed]
- Al-Shajlawi, M.; Alsayed, A.R.; Abazid, H.; Awajan, D.; Al-Imam, A.; Basheti, I. Using Laboratory Parameters as Predictors for the Severity and Mortality of COVID-19 in Hospitalized Patients. Pharm. Pract. 2022, 20, 2721. [Google Scholar] [CrossRef]
- Alsayed, A.R.; Alsayed, Y. Integrating Telemonitoring in COPD Exacerbations Care: A Multinational Study Using the Alsayed System for Applied Medical-Care Improvement (ASAMI) Database. OBM Transplant. 2025, 9, 1–20. [Google Scholar] [CrossRef]
- Rajkomar, A.; Oren, E.; Chen, K.; Dai, A.M.; Hajaj, N.; Hardt, M.; Liu, P.J.; Liu, X.; Marcus, J.; Sun, M.; et al. Scalable and Accurate Deep Learning With Electronic Health Records. NPJ Digit. Med. 2018, 1. [Google Scholar] [CrossRef] [PubMed]
- Stankevičiūtė, K.; Woillard, J.-B.; Peck, R.W.; Marquet, P.; van der Schaar, M. Bridging the Worlds of Pharmacometrics and Machine Learning. Clin. Pharmacokinet. 2023, 62, 1551–1565. [Google Scholar] [CrossRef]
- Keutzer, L.; You, H.; Farnoud, A.; Nyberg, J.; Wicha, S.G.; Maher-Edwards, G.; Vlasakakis, G.; Moghaddam, G.K.; Svensson, E.M.; Menden, M.P. Machine Learning and Pharmacometrics for Prediction of Pharmacokinetic Data: Differences, Similarities and Challenges Illustrated with Rifampicin. Pharmaceutics 2022, 14, 1530. [Google Scholar] [CrossRef] [PubMed]
- Alsayed, A.R. Illustrating How to Use the Validated Alsayed_v1 Tools to Improve Medical Care: A Particular Reference to the Global Initiative for Asthma 2022 Recommendations. Patient Prefer. Adherence 2023, 1161–1179. [Google Scholar] [CrossRef]
- Bräm, D.S.; Nahum, U.; Atkinson, A.; Koch, G.; Pfister, M. Evaluation of Machine Learning Methods for Covariate Data Imputation in Pharmacometrics. CPT Pharmacomet. Syst. Pharmacol. 2022, 11, 1638–1648. [Google Scholar] [CrossRef] [PubMed]
- Mansoor, A.R.; Abed, A.; Alqudah, A.; Alsayed, A.R. Assessment of Medical Care Strategies for Primary Hypertension in Iraqi Adults: A Hospital-Based Problem-Oriented Plan. Patient Prefer. Adherence 2025, 1317–1335. [Google Scholar] [CrossRef]
- Alsayed, A.R.; Ahmed, S.I.; Shweiki, A.O.A.; Al-Shajlawi, M.; Hakooz, N. The Laboratory Parameters in Predicting the Severity and Death of COVID-19 Patients: Future Pandemic Readiness Strategies. Biomol. Biomed. 2024, 24, 238. [Google Scholar] [CrossRef]
- Asai, Y.; Tashiro, T.; Kondo, Y.; Hayashi, M.; Arihara, H.; Omote, S.; Tanio, E.; Yamashita, S.; Higuchi, T.; Hashimoto, E.; et al. Machine Learning-Based Prediction of Digoxin Toxicity in Heart Failure: A Multicenter Retrospective Study. Biol. Pharm. Bull. 2023, 46, 614–620. [Google Scholar] [CrossRef]
- Hu, T.; Ding, X.; Han, F.; An, Z. Machine Learning Approach for Personalized Vancomycin Steady-State Trough Concentration Prediction: A Superior Approach over Bayesian Population Pharmacokinetic Model. Front. Pharmacol. 2025, 16, 1549500. [Google Scholar] [CrossRef]
- van Os, W.; O’Jeanson, A.; Troisi, C.; Liu, C.; Brooks, J.T.; Hughes, J.H.; Tong, D.M.H.; Keizer, R.J. Machine Learning-Based Model Selection and Averaging Outperform Single-Model Approaches for a Priori Vancomycin Precision Dosing. CPT Pharmacomet. Syst. Pharmacol. 2025, 14, 1650–1660. [Google Scholar] [CrossRef]
- Chen, K.; Wang, C.; Wei, Y.; Ma, S.; Huang, W.; Dong, Y.; Wang, Y. Machine Learning and Population Pharmacokinetics: A Hybrid Approach for Optimizing Vancomycin Therapy in Sepsis Patients. Microbiol. Spectr. 2025, 13, e00499-25. [Google Scholar] [CrossRef]
- Hu, T.; Ding, X.; Han, F.; An, Z. Machine Learning Approach for Personalized Vancomycin Steady-State Trough Concentration Prediction: A Superior Approach over Bayesian Population Pharmacokinetic Model. Front. Pharmacol. 2025, 16, 1549500. [Google Scholar] [CrossRef]
- Zhu, X.; Huang, W.; Lu, H.; Wang, Z.; Ni, X.; Hu, J.; Deng, S.; Tan, Y.; Li, L.; Zhang, M.; et al. A Machine Learning Approach to Personalized Dose Adjustment of Lamotrigine Using Noninvasive Clinical Parameters. Sci. Rep. 2021, 11, 5568. [Google Scholar] [CrossRef] [PubMed]
- Kamiya, Y.; Handa, K.; Miura, T.; Ohori, J.; Kato, A.; Shimizu, M.; Kitajima, M.; Yamazaki, H. Simplified Physiologically Based Pharmacokinetic Model Subsequently. Biol Pharm Bull.
- Tang, J.; Liu, R.; Zhang, Y.-L.; Liu, M.-Z.; Hu, Y.-F.; Shao, M.-J.; Zhu, L.-J.; Xin, H.-W.; Feng, G.-W.; Shang, W.-J.; et al. Application of Machine-Learning Models to Predict Tacrolimus Stable Dose in Renal Transplant Recipients. Sci. Rep. 2017, 7, 42192. [Google Scholar] [CrossRef]
- Tosca, E.M.; De Carlo, A.; Ronchi, D.; Magni, P. Model-Informed Reinforcement Learning for Enabling Precision Dosing Via Adaptive Dosing. Clin. Pharmacol. Ther. 2024, 116, 619–636. [Google Scholar] [CrossRef]
- Kaul, R.; Ossai, C.; Forkan, A.R.M.; Jayaraman, P.P.; Zelcer, J.; Vaughan, S.; Wickramasinghe, N. The Role of AI for Developing Digital Twins in Healthcare: The Case of Cancer Care. WIREs Data Min. Knowl. Discov. 2023, 13, e1480. [Google Scholar] [CrossRef]
- Srivastava, D.S. Pharmacokinetic–Pharmacodynamic (PK–PD) Modeling in Personalized Medicine: Optimizing Dosage for High-Risk Patient Populations. Int. J. 2025, 01. [Google Scholar] [CrossRef]
- Zhu, X.; Huang, W.; Lu, H.; Wang, Z.; Ni, X.; Hu, J.; Deng, S.; Tan, Y.; Li, L.; Zhang, M. A Machine Learning Approach to Personalized Dose Adjustment of Lamotrigine Using Noninvasive Clinical Parameters. Sci. Rep. 2021, 11, 5568. [Google Scholar] [CrossRef] [PubMed]
- Wicha, S.G.; Märtson, A.; Nielsen, E.I.; Koch, B.C.; Friberg, L.E.; Alffenaar, J.; Minichmayr, I.K. International Society of Anti-Infective Pharmacology (ISAP), the P. study group of the E.S. of C.M., Infectious Diseases (EPASG) From Therapeutic Drug Monitoring to Model-informed Precision Dosing for Antibiotics. Clin. Pharmacol. Ther. 2021, 109, 928–941. [Google Scholar] [CrossRef] [PubMed]
- Spanakis, M.; Ioannou, P.; Tzalis, S.; Chouzouri, F.; Patelarou, E.; Kofteridis, D.P.; Antoniou, K.M.; Schiza, S.E.; Patelarou, A.; Tzanakis, N. Evaluation of Drug Interactions in Hospitalized Patients with Respiratory Disorders in Greece. Adv. Respir. Med. 2023, 91, 74–92. [Google Scholar] [CrossRef]
- Peruzzi, E.; Roncato, R.; De Mattia, E.; Bignucolo, A.; Swen, J.J.; Guchelaar, H.; Toffoli, G.; Cecchin, E. Implementation of Pre-emptive Testing of a Pharmacogenomic Panel in Clinical Practice: Where Do We Stand? Br. J. Clin. Pharmacol. 2025, 91, 270–282. [Google Scholar] [CrossRef]
- Dr. P. Devi Guna Sireesha; Shaik Ayesha Sidhikha; Tokala Divya; Inapanurthy kavya; Gorlollapalli Thanveer Apsha AI PERSONALIZED DRUG DOSING FOR RESPIRATORY MEDICATIONS. 2025. [CrossRef]
- Van Dijkman, S.C.; Wicha, S.G.; Danhof, M.; Della Pasqua, O.E. Individualized Dosing Algorithms and Therapeutic Monitoring for Antiepileptic Drugs. Clin. Pharmacol. Ther. 2018, 103, 663–673. [Google Scholar] [CrossRef]
- Uhlig, K.; Leff, B.; Kent, D.; Dy, S.; Brunnhuber, K.; Burgers, J.S.; Greenfield, S.; Guyatt, G.; High, K.; Leipzig, R. A Framework for Crafting Clinical Practice Guidelines That Are Relevant to the Care and Management of People with Multimorbidity. J. Gen. Intern. Med. 2014, 29, 670–679. [Google Scholar] [CrossRef]
- Cherubini, A.; Oristrell, J.; Pla, X.; Ruggiero, C.; Ferretti, R.; Diestre, G.; Clarfield, A.M.; Crome, P.; Hertogh, C.; Lesauskaite, V. The Persistent Exclusion of Older Patients from Ongoing Clinical Trials Regarding Heart Failure. Arch. Intern. Med. 2011, 171, 550–556. [Google Scholar] [CrossRef] [PubMed]
- Al Maqbali, M.; Alsayed, A.; Bashayreh, I. Quality of Life and Psychological Impact among Chronic Disease Patients during the COVID-19 Pandemic. J. Integr. Nurs. 2022, 4, 217–223. [Google Scholar] [CrossRef]
- Khader, H.; Hasoun, L.Z.; Alsayed, A.; Abu-Samak, M. Potentially Inappropriate Medications Use and Its Associated Factors among Geriatric Patients: A Cross-Sectional Study Based on 2019 Beers Criteria. Pharmacia 2021, 68, 789–795. [Google Scholar] [CrossRef]
- Alimam, S.M.; Alhmoud, J.F.; Khader, H.A.; Alsayed, A.R.; Abusamak, M.; Mohammad, B.A.; Mosleh, I.; Khadra, K.A.; Aljaberi, A.; Habash, M. Effect of Weekly High-Dose Vitamin D3 Supplementation on the Association between Circulatory FGF-23 and A1c Levels in People with Vitamin D Deficiency: A Randomized Controlled 10-Week Follow-up Trial. Pharm. Pract. 2024, 22, 1–8. [Google Scholar] [CrossRef]
- Khaled, R.A.; Alhmoud, J.F.; Issa, R.; Khader, H.A.; Mohammad, B.A.; Alsayed, A.R.; Khadra, K.A.; Habash, M.; Aljaberi, A.; Hasoun, L. The Variations of Selected Serum Cytokines Involved in Cytokine Storm after Omega-3 Daily Supplements: A Randomized Clinical Trial in Jordanians with Vitamin D Deficiency. Pharm. Pract. 2024, 22, 1–10. [Google Scholar] [CrossRef]
- Daboul, S.M.; Abusamak, M.; Mohammad, B.A.; Alsayed, A.R.; Habash, M.; Mosleh, I.; Al-Shakhshir, S.; Issa, R.; Abu-Samak, M. The Effect of Omega-3 Supplements on the Serum Levels of ACE/ACE2 Ratio as a Potential Key in Cardiovascular Disease: A Randomized Clinical Trial in Participants with Vitamin D Deficiency. Pharm. Pract. 2022, 21, 2761. [Google Scholar] [CrossRef]
- Vorisek, C.N.; Stellmach, C.; Mayer, P.J.; Klopfenstein, S.A.I.; Bures, D.M.; Diehl, A.; Henningsen, M.; Ritter, K.; Thun, S. Artificial Intelligence Bias in Health Care: Web-Based Survey. J. Med. Internet Res. 2023, 25, e41089. [Google Scholar] [CrossRef] [PubMed]
- Zech, J.R.; Badgeley, M.A.; Liu, M.; Costa, A.B.; Titano, J.J.; Oermann, E.K. Variable Generalization Performance of a Deep Learning Model to Detect Pneumonia in Chest Radiographs: A Cross-Sectional Study. PLOS Med. 2018, 15, e1002683. [Google Scholar] [CrossRef]
- Obermeyer, Z.; Powers, B.; Vogeli, C.; Mullainathan, S. Dissecting Racial Bias in an Algorithm Used to Manage the Health of Populations. Science 2019, 366, 447–453. [Google Scholar] [CrossRef]
- Seyyed-Kalantari, L.; Zhang, H.; McDermott, M.B.A.; Chen, I.Y.; Ghassemi, M. Underdiagnosis Bias of Artificial Intelligence Algorithms Applied to Chest Radiographs in Under-Served Patient Populations. Nat. Med. 2021, 27, 2176–2182. [Google Scholar] [CrossRef]
- Li, F.; Wu, P.; Ong, H.H.; Peterson, J.F.; Wei, W.-Q.; Zhao, J. Evaluating and Mitigating Bias in Machine Learning Models for Cardiovascular Disease Prediction. J. Biomed. Inform. 2023, 138, 104294. [Google Scholar] [CrossRef]
- Tonekaboni, S.; Joshi, S.; McCradden, M.D.; Goldenberg, A. What Clinicians Want: Contextualizing Explainable Machine Learning for Clinical End Use.
- Reich, C.; Meder, B. The Heart and Artificial Intelligence—How Can We Improve Medicine Without Causing Harm. Curr. Heart Fail. Rep. 2023, 20, 271–279. [Google Scholar] [CrossRef]
- Norori, N.; Hu, Q.; Aellen, F.M.; Faraci, F.D.; Tzovara, A. Addressing Bias in Big Data and AI for Health Care: A Call for Open Science. Patterns 2021, 2. [Google Scholar] [CrossRef]
- Topol, E.J. Bias in Healthcare Algorithms. Nat. Med. 2019, 25, 44–56. [Google Scholar] [CrossRef] [PubMed]
- Nemati, S.; Ghassemi, M.M.; Clifford, G.D. Optimal Medication Dosing from Suboptimal Clinical Examples: A Deep Reinforcement Learning Approach. Annu. Int. Conf. IEEE Eng. Med. Biol. Soc. IEEE Eng. Med. Biol. Soc. Annu. Int. Conf. 2016, 2016, 2978–2981. [Google Scholar] [CrossRef]
- Komorowski, M.; Celi, L.A.; Badawi, O.; Gordon, A.C.; Faisal, A.A. The Artificial Intelligence Clinician Learns Optimal Treatment Strategies for Sepsis in Intensive Care. Nat. Med. 2018, 24, 1716–1720. [Google Scholar] [CrossRef] [PubMed]
- Cristea, A.-M.; Dobre, C.; Cristea, A.-M.; Dobre, C. Federated Transfer Learning for Tomato Leaf Disease Detection Using Neuro-Graph Hybrid Model. AgriEngineering 2025, 7. [Google Scholar] [CrossRef]
- Luck, M.; Sylvain, T.; Cardinal, H.; Lodi, A.; Bengio, Y. Deep Learning for Patient-Specific Kidney Graft Survival Analysis 2017.
| Reason | Underlying Change | Clinical Comment/Impact |
|---|---|---|
| Changes in body composition | ↑ Proportion of body fat | Lipophilic drugs (e.g., benzodiazepines) have ↑ volume of distribution → prolonged half-life and accumulation |
| ↓ Proportion of total body water | Hydrophilic drugs (e.g., digoxin, lithium) reach higher plasma concentrations | |
| ↓ Muscle mass | Reduced creatinine production → renal function may be overestimated | |
| Altered drug clearance | ↓ Hepatic metabolism | Reduced phase I metabolism → ↑ bioavailability and toxicity of many drugs |
| ↓ Renal elimination | Accumulation of renally cleared drugs → ↑ risk of adverse drug reactions | |
| Increased vulnerability to toxicity | ↑ Blood–brain barrier permeability | Greater Central Nervous System (CNS) effects (confusion, sedation, delirium) |
| Atypical presentation of adverse effects | Non-specific symptoms | Falls or delirium may not be immediately recognized as medication-related |
| Polypharmacy | Multiple concurrent medications | ↑ Risk of drug–drug interactions (e.g., aspirin + Selective Serotonin Reuptake Inhibitors (SSRI) → ↑ Gastro-Intestinal (GI) bleeding risk) |
| Multimorbidity | Multiple coexisting diseases | ↑ Risk of drug–disease interactions (e.g., opioids worsening constipation) |
| Pharmacodynamic Factor | Impact in Geriatrics |
|---|---|
| Altered receptor sensitivity | Changes in receptor number or responsiveness may reduce or enhance drug effects, sometimes necessitating dose adjustments |
| Drug metabolism (PD-related response) | Slower metabolic processes can prolong drug action and increase the risk of accumulation and adverse effects |
| Physiological changes | Age-related alterations in organ function influence drug distribution, elimination, and tissue responsiveness |
| Increased susceptibility to Adverse Drug Events (ADEs) | Older adults have greater vulnerability to ADEs, requiring cautious dosing and close monitoring |
| Pharmacodynamic interactions | Coexisting diseases and polypharmacy increase the likelihood of clinically significant drug–drug and drug–disease interactions |
| Algorithm | Clinical Application | Output Type | Performance & Key Findings | Source |
|---|---|---|---|---|
| RF | Warfarin blood levels (International Normalized Ratio (INR)) | Continuous | Optimal for raw data/59.0% ideal prediction percentage and 0.595 correlation in validation. | [37] |
| GDA | Warfarin blood levels (INR) | Categorical | Best for categorical output/total True Positive Rate (TPR) of 95.6%. Provides an explicit equation. | [37] |
| Shallow Neural Network (SNN) | Heparin dosing (activated partial thromboplastin time (aPTT)) | Categorical | Performed best across 3 datasets (F1 scores: 85.98%–87.55%). Outperformed ensemble models. | [38] |
| L-BFGS (Optimizer) | 2-Compartment PK parameters | Continuous | Significantly faster training (~3.4s) compared to Adam (>2400s) for small-scale datasets. | [36] |
| Adam (Optimizer) | 2-Compartment PK parameters | Continuous | Used in a hybrid approach; effective for solving inverse problems but slower in small datasets. | [36] |
| Extreme Gradient Boosting (XGBoost) | Heparin dosing (aPTT) | Categorical | Achieved second-best F1 scores (73.94%–78.85%). More robust than traditional RF for heparin data. | [38] |
| Support Vector Machine (SVM) | Heparin dosing (aPTT) | Categorical | Achieved 100% precision for normal and supratherapeutic classes but had lower recall than SNN. | [38] |
| Adaptive Boosting (AdaBoost) | Heparin dosing (aPTT) | Categorical | Performance was competitive with XGBoost, reaching up to 81.67% F1 score in specific datasets. | [38] |
| Multilayer Perceptron (MLP) | Warfarin blood levels (INR) | Continuous/Categorical | Demonstrated lower correlation (0.475) and higher error rates compared to RF or GDA. | [37] |
| Drug | ML Model | ML Performance (R²) | Population PK Performance (R²) | Accuracy Improvement (F30%) | Comments |
|---|---|---|---|---|---|
| Cyclosporine | ANN | 0.75 | 0.68 | 56.46% (ML) vs 51.22% (PopPK) | ML showed modest but clinically relevant improvement over PopPK |
| Vancomycin | ML Ensemble (SVR, LightGBM, CatBoost) | 0.656 | 0.218 | 76.62% (ML) vs 53.75% (PopPK) | Marked superiority of ML in exposure prediction |
| Voriconazole | ML Ensemble | 0.828 | Not specified | Minimal MAPE achieved (0.772) | High predictive accuracy despite lack of PopPK comparator |
| Tacrolimus | Regression Tree | 0.73 | 0.71 (Multiple Linear Regression) | 56.1% ideal rate | ML marginally outperformed traditional regression |
| Clinical Condition | Primary Biomarker(s) | Individual-Level Target | Clinical Goal | Clinical Rationale | Source |
|---|---|---|---|---|---|
| Metastatic Cancer (Erdafitinib) | Serum phosphate ([PO₄³⁻]) | 5.5–7.0 mg/dL | Balance antitumor efficacy while limiting hyperphosphatemia | Serum phosphate serves as a pharmacodynamic biomarker reflecting Fibroblast Growth Factor Receptor (FGFR) inhibition intensity | [68,69] |
| Polycythemia Vera (Givinostat) | Platelets, WBC, Hematocrit | Complete Hematologic Response (CHR) | Simultaneous normalization of all three blood cell lines | Multi-parameter control is required to reduce thrombotic risk and disease progression | [68,69] |
| Renal Impairment (Direct Oral Anti-Coagulants (DOACs)) | Plasma drug concentration / renal function | 25–30% dose reduction | Maintain therapeutic anticoagulation while minimizing bleeding risk | Age-related renal decline significantly increases DOAC exposure | [70] |
| Critical Care (Vancomycin) | Serum concentration (or AUC-guided exposure) | 15–25 mg/L (or AUC/MIC 400–600) | Achieve stable exposure while preventing nephrotoxicity | Narrow therapeutic index; elderly patients are highly susceptible to Acute Kidney Injury (AKI) | [69] |
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2026 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).




