Submitted:
03 April 2026
Posted:
07 April 2026
You are already at the latest version
Abstract
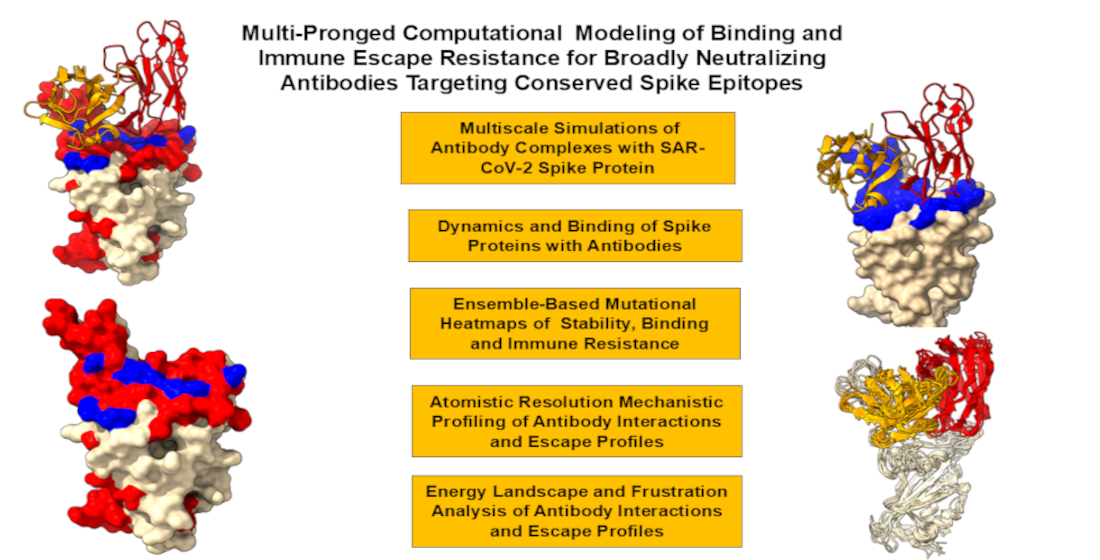
Keywords:
1. Introduction
2. Results
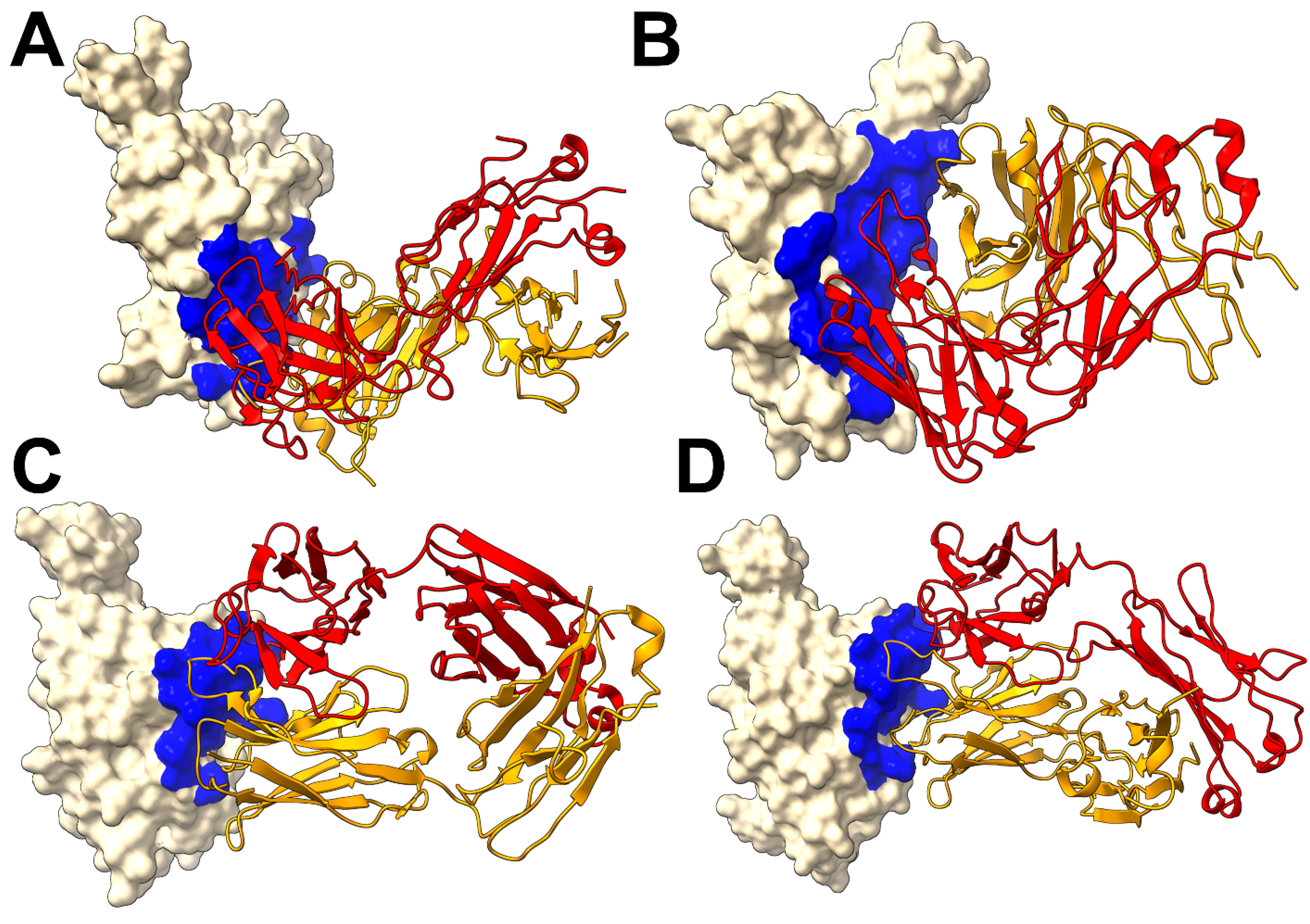
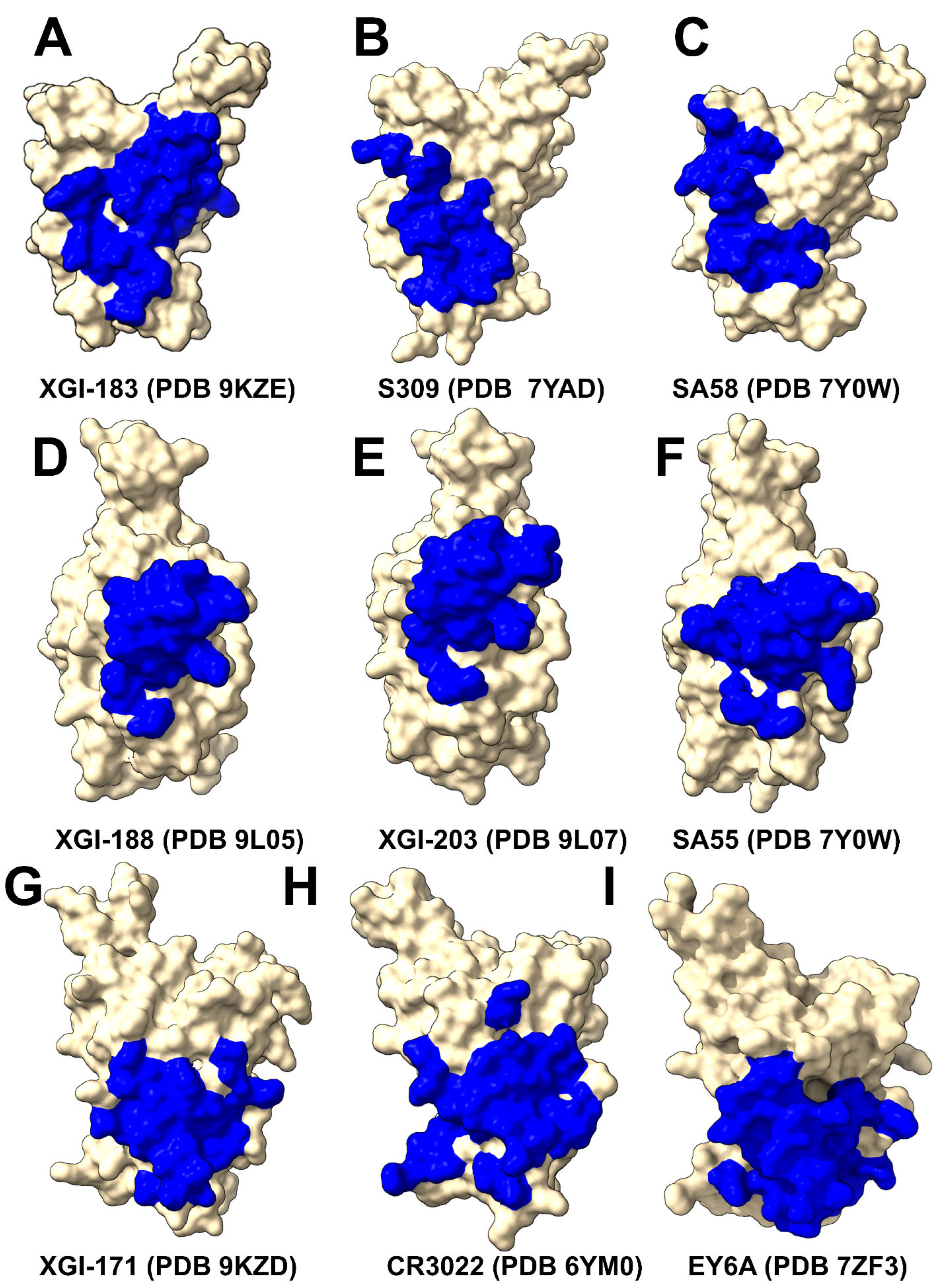
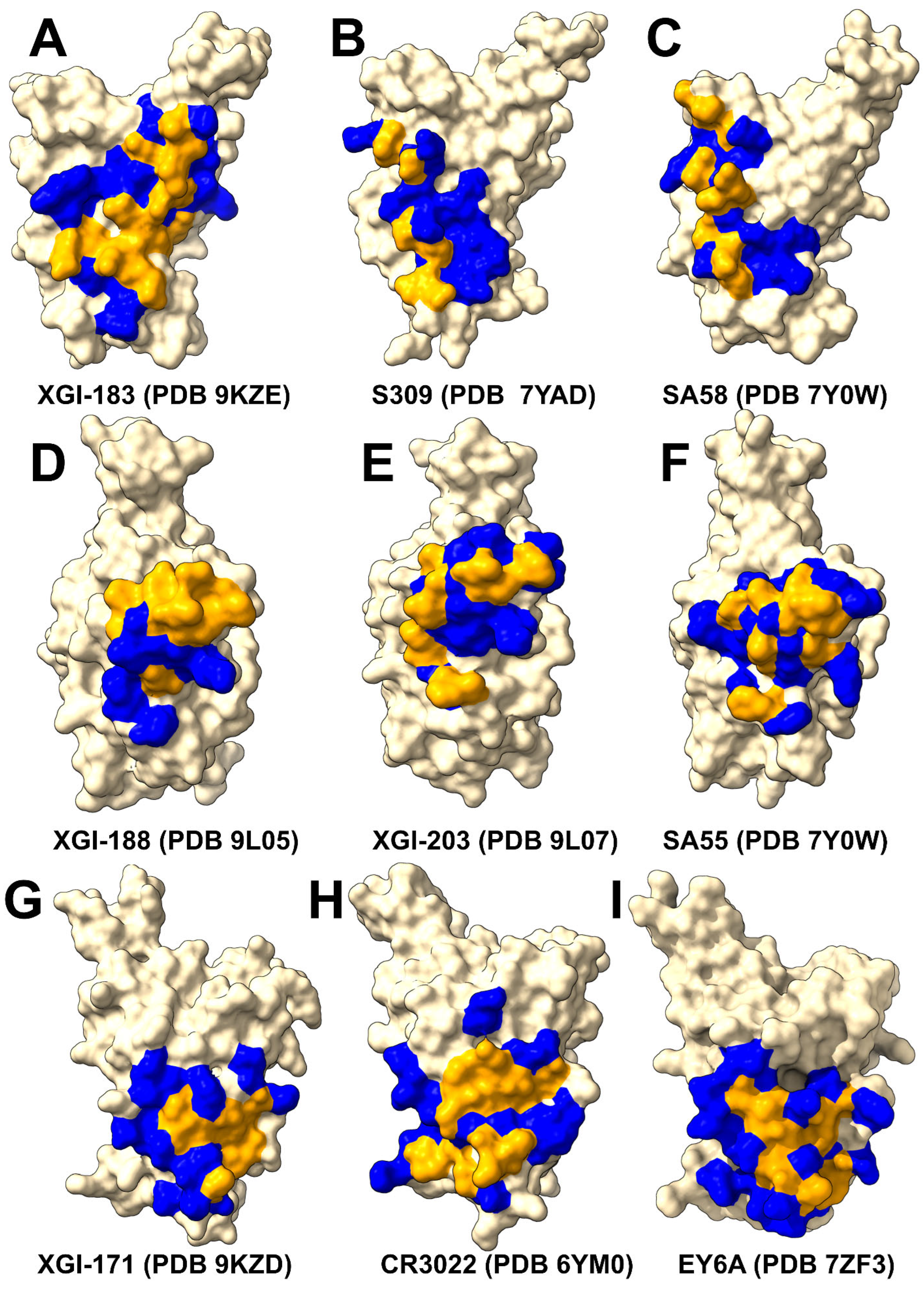
2.1. Structural Characterization of the Conserved RBD Epitopes Targeted by XGI Antibodies
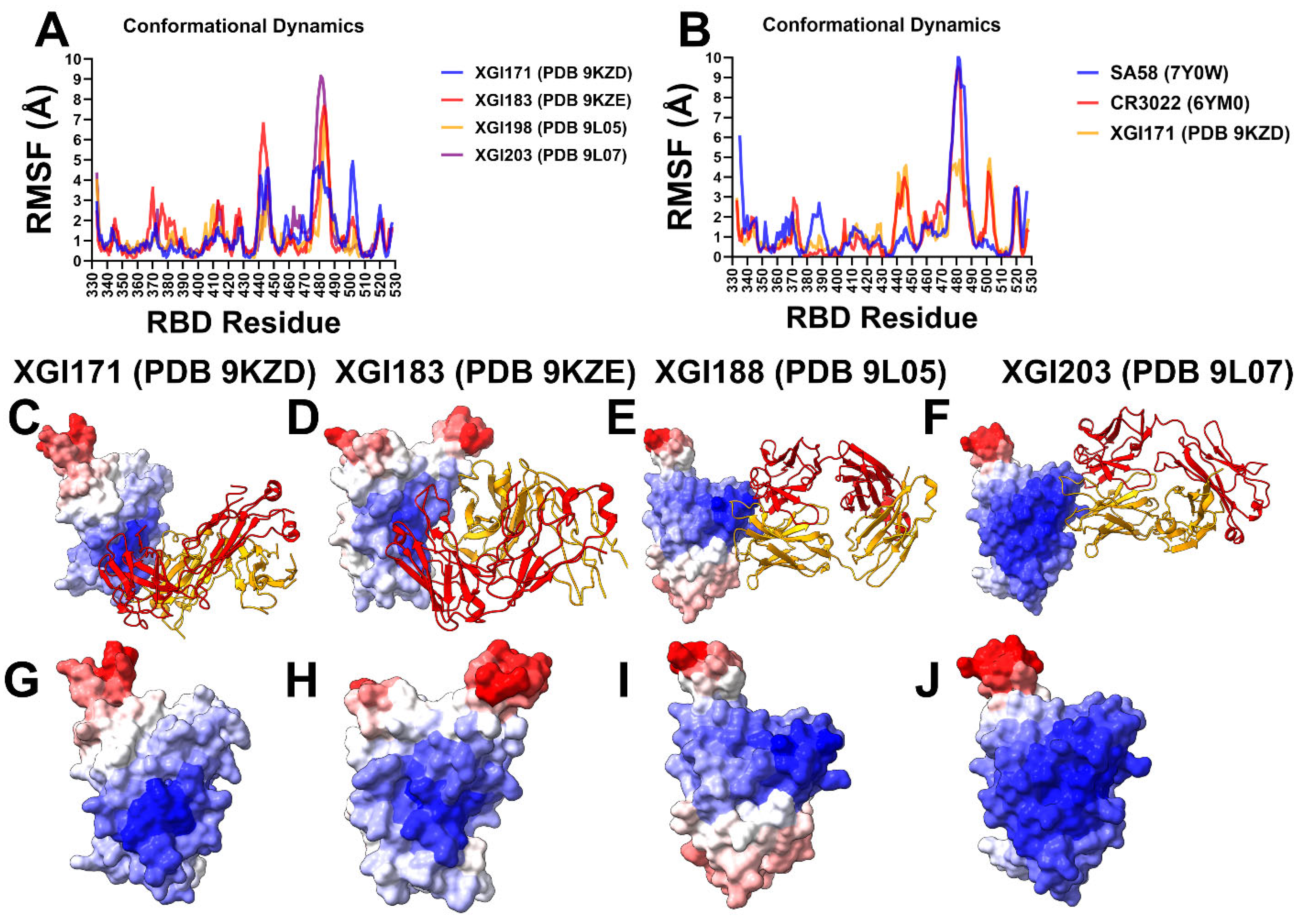
2.2. Conformational Dynamics of XGI Antibody–RBD Complexes Reveal Distinct Flexibility Signatures Linked to Escape Vulnerability
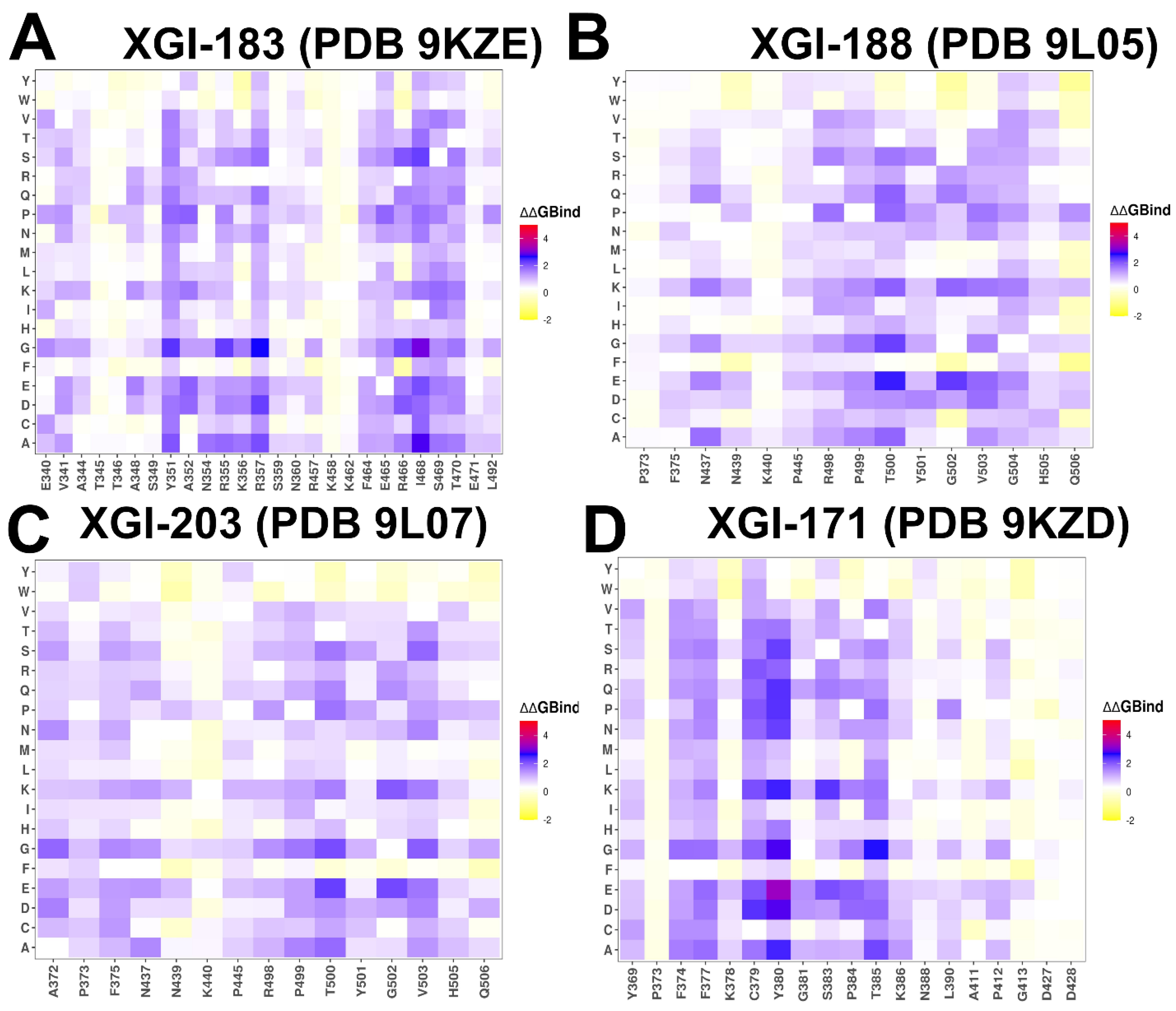
2.3. Mutational Profiling of Antibody-RBD Binding Interactions Interfaces Reveals Molecular Determinants of Immune Sensitivity
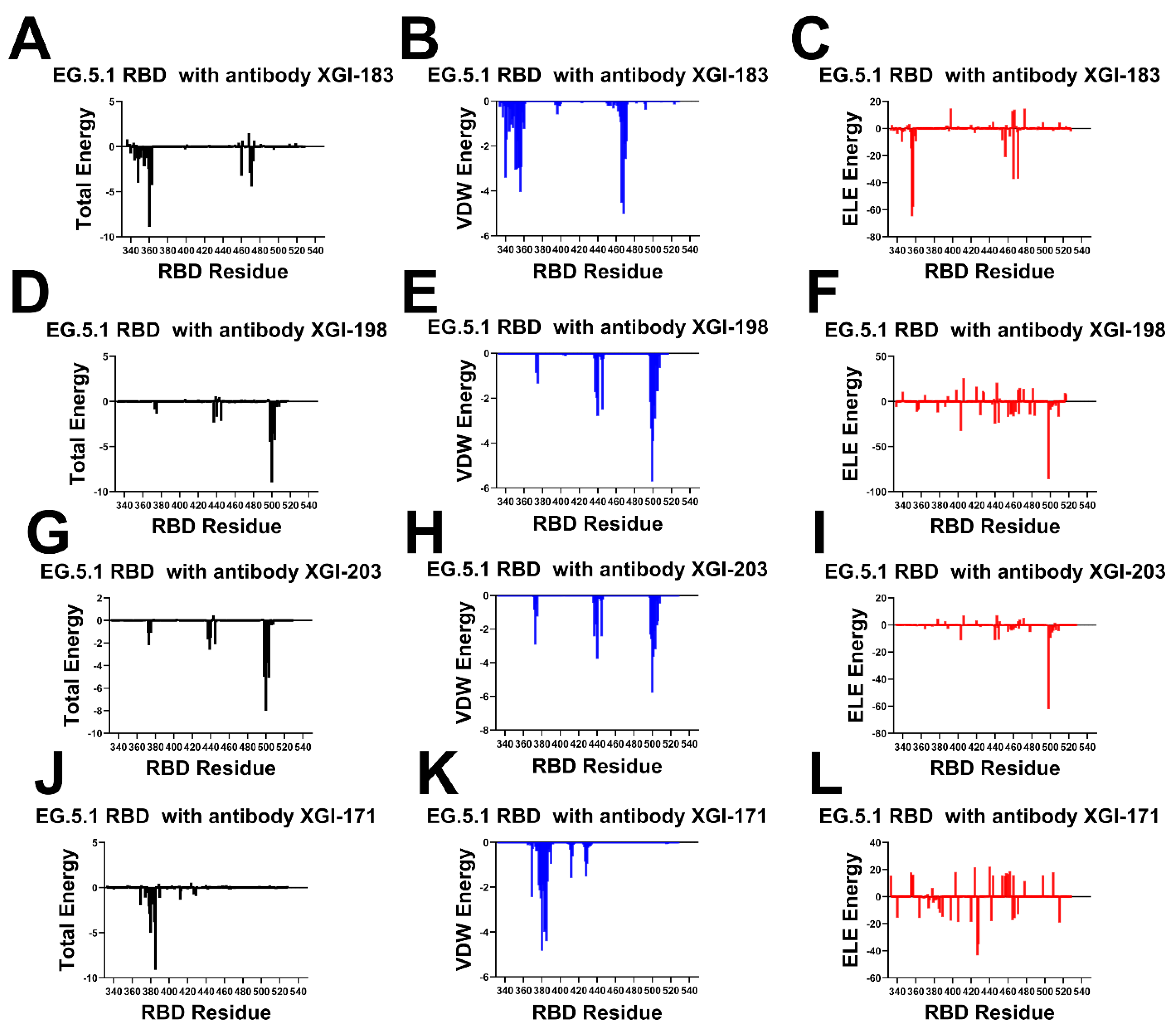
2.4. Energetic Architecture of XGI Antibody–RBD Interfaces: Common Principles Across Three Conserved Epitopes
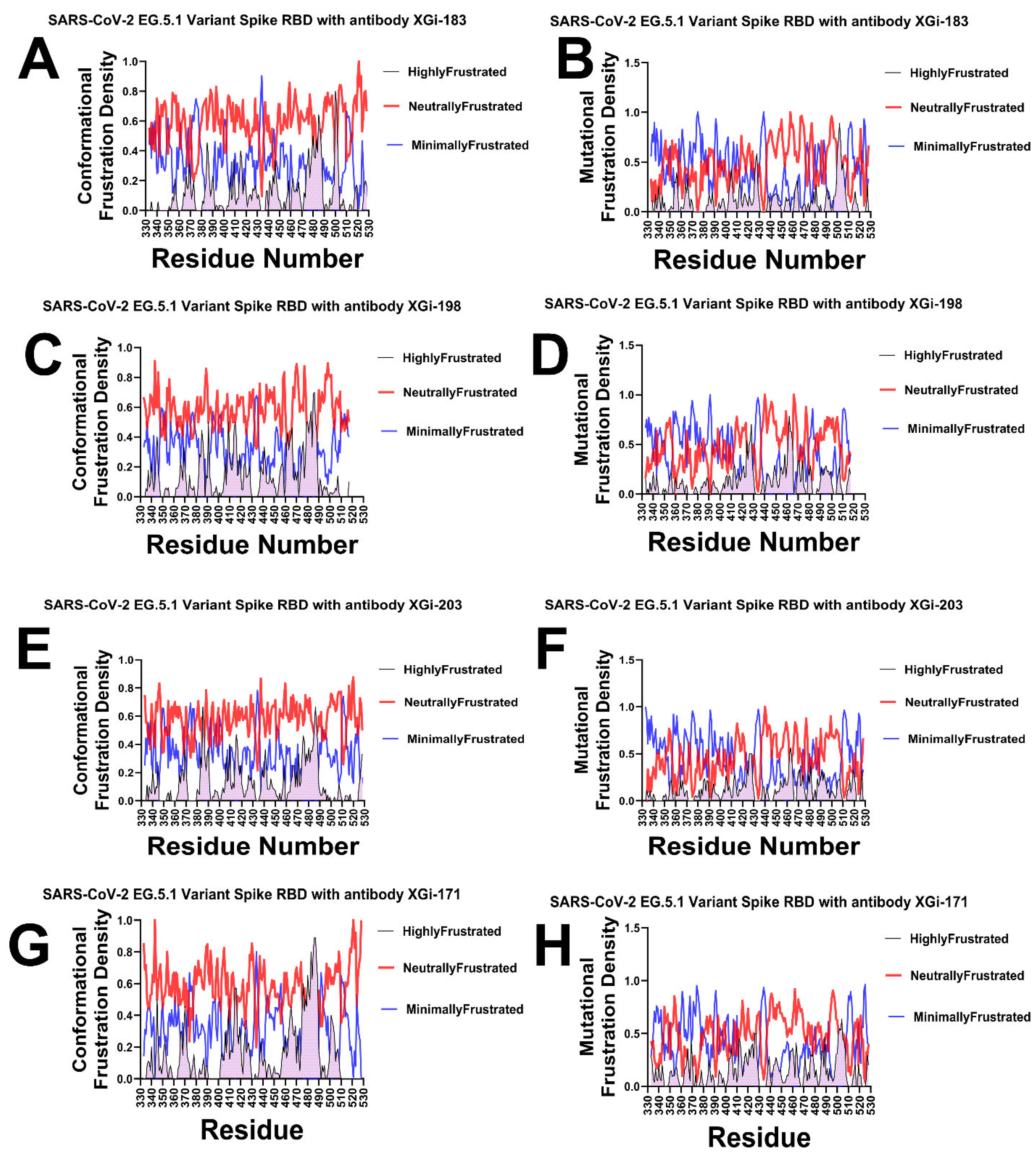
2.5. Frustration Landscape Analysis of Antibody–RBD Interfaces Reveals Energetic Signatures of Binding and Resistance to Immune Escape
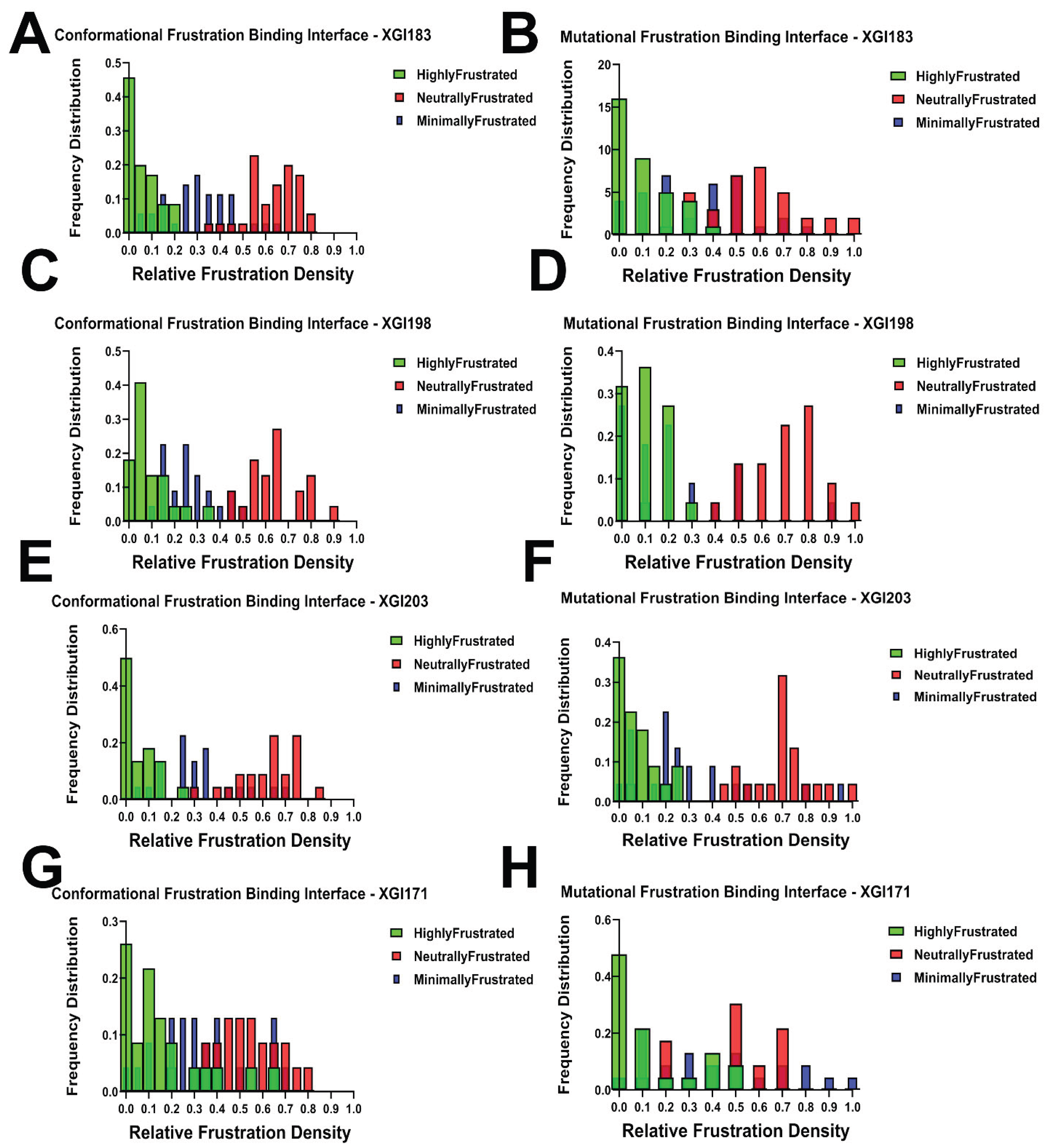
2.6. Comparative Interface-Specific Frustration Distributions: Conformational Plasticity Versus Mutational Constraint
3. Discussion
3. Materials and Methods
3.1. Structure Preparation and Analysis
3.2. Coarse-Grained Simulations
3.3. All-Atom Molecular Dynamics Simulations
3.4. Mutational Scanning of the RBD-Antibody Binding Interfaces
3.5. Binding Free Energy Computations of the RBD Complexes with Antibodies
3.6. Local Frustration Analysis of Conformational Ensembles
5. Conclusions
Supplementary Materials
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Acknowledgments
Conflicts of Interest
References
- Tai, W.; He, L.; Zhang, X.; Pu, J.; Voronin, D.; Jiang, S.; Zhou, Y.; Du, L. Characterization of the receptor-binding domain (RBD) of 2019 novel coronavirus: implication for development of RBD protein as a viral attachment inhibitor and vaccine. Cell. Mol. Immunol. 2020, 17, 613–620. [Google Scholar] [CrossRef]
- Wang, Q.; Zhang, Y.; Wu, L.; Niu, S.; Song, C.; Zhang, Z.; Lu, G.; Qiao, C.; Hu, Y.; Yuen, K. Y.; Wang, Q.; Zhou, H.; Yan, J.; Qi, J. Structural and functional basis of SARS-CoV-2 entry by using human ACE2. Cell 2020, 181, 894–904.e9. [Google Scholar] [CrossRef]
- Walls, A. C.; Park, Y. J.; Tortorici, M. A.; Wall, A.; McGuire, A. T.; Veesler, D. Structure, Function, and Antigenicity of the SARS-CoV-2 Spike Glycoprotein. Cell 2020, 181, 281–292.e6. [Google Scholar] [CrossRef] [PubMed]
- Wrapp, D.; Wang, N.; Corbett, K. S.; Goldsmith, J. A.; Hsieh, C. L.; Abiona, O.; Graham, B. S.; McLellan, J. S. Cryo-EM structure of the 2019-nCoV spike in the prefusion conformation. Science 2020, 367, 1260–1263. [Google Scholar] [CrossRef] [PubMed]
- Cai, Y.; Zhang, J.; Xiao, T.; Peng, H.; Sterling, S. M.; Walsh, R. M., Jr.; Rawson, S.; Rits-Volloch, S.; Chen, B. Distinct conformational states of SARS-CoV-2 spike protein. Science 2020, 369, 1586–1592. [Google Scholar] [CrossRef]
- Hsieh, C. L.; Goldsmith, J. A.; Schaub, J. M.; DiVenere, A. M.; Kuo, H. C.; Javanmardi, K.; Le, K. C.; Wrapp, D.; Lee, A. G.; Liu, Y.; Chou, C.W.; Byrne, P.O.; Hjorth, C.K.; Johnson, N.V.; Ludes-Meyers, J.; Nguyen, A.W.; Park, J.; Wang, N.; Amengor, D.; Lavinder, J.J.; Ippolito, G.C.; Maynard, J.A.; Finkelstein, I.J.; McLellan, J.S. Structure-based design of prefusion-stabilized SARS-CoV-2 spikes. Science 2020, 369, 1501–1505. [Google Scholar] [CrossRef]
- Henderson, R.; Edwards, R. J.; Mansouri, K.; Janowska, K.; Stalls, V.; Gobeil, S. M. C.; Kopp, M.; Li, D.; Parks, R.; Hsu, A. L.; Borgnia, M.J.; Haynes, B.F.; Acharya, P. Controlling the SARS-CoV-2 spike glycoprotein conformation. Nat. Struct. Mol. Biol. 2020, 27, 925–933. [Google Scholar] [CrossRef]
- McCallum, M.; Walls, A. C.; Bowen, J. E.; Corti, D.; Veesler, D. Structure-guided covalent stabilization of coronavirus spike glycoprotein trimers in the closed conformation. Nat. Struct. Mol. Biol. 2020, 27, 942–949. [Google Scholar] [CrossRef]
- Xiong, X.; Qu, K.; Ciazynska, K. A.; Hosmillo, M.; Carter, A. P.; Ebrahimi, S.; Ke, Z.; Scheres, S. H. W.; Bergamaschi, L.; Grice, G. L.; Zhang, Y.; CITIID-NIHR COVID-19 BioResource Collaboration; Nathan, J.A.; Baker, S.; James, L.C.; Baxendale, H.E.; Goodfellow, I.; Doffinger, R.; Briggs, J.A.G. A thermostable, closed SARS-CoV-2 spike protein trimer. Nat. Struct. Mol. Biol. 2020, 27, 934–941. [Google Scholar] [CrossRef] [PubMed]
- Costello, S.M.; Shoemaker, S.R.; Hobbs, H.T.; Nguyen, A.W.; Hsieh, C.L.; Maynard, J.A.; McLellan, J.S.; Pak, J.E.; Marqusee, S. The SARS-CoV-2 spike reversibly samples an open-trimer conformation exposing novel epitopes. Nat. Struct. Mol. Biol. 2022, 27, 229–238. [Google Scholar] [CrossRef]
- McCormick, K.D.; Jacobs, J.L.; Mellors, J.W. The emerging plasticity of SARS-CoV-2. Science 2021, 371, 1306–1308. [Google Scholar] [CrossRef]
- Ghimire, D.; Han, Y.; Lu, M. Structural Plasticity and Immune Evasion of SARS-CoV-2 Spike Variants. Viruses 2022, 14, 1255. [Google Scholar] [CrossRef]
- Xu, C.; Wang, Y.; Liu, C.; Zhang, C.; Han, W.; Hong, X.; Wang, Y.; Hong, Q.; Wang, S.; Zhao, Q.; Wang, Y.; Yang, Y.; Chen, K.; Zheng, W.; Kong, L.; Wang, F.; Zuo, Q.; Huang, Z.; Cong, Y. Conformational dynamics of SARS-CoV-2 trimeric spike glycoprotein in complex with receptor ACE2 revealed by cryo-EM. Sci. Adv. 2021, 7, eabe5575. [Google Scholar] [CrossRef] [PubMed]
- Benton, D. J.; Wrobel, A. G.; Xu, P.; Roustan, C.; Martin, S. R.; Rosenthal, P. B.; Skehel, J. J.; Gamblin, S. J. Receptor binding and priming of the spike protein of SARS-CoV-2 for membrane fusion. Nature 2020, 588, 327–330. [Google Scholar] [CrossRef]
- Turoňová, B.; Sikora, M.; Schuerman, C.; Hagen, W. J. H.; Welsch, S.; Blanc, F. E. C.; von Bülow, S.; Gecht, M.; Bagola, K.; Hörner, C.; van Zandbergen, G.; Landry, J.; de Azevedo, N. T. D.; Mosalaganti, S.; Schwarz, A.; Covino, R.; Mühlebach, M. D.; Hummer, G.; Krijnse Locker, J.; Beck, M. In situ structural analysis of SARS-CoV-2 spike reveals flexibility mediated by three hinges. Science 2020, 370, 203–208. [Google Scholar] [CrossRef]
- Lu, M.; Uchil, P. D.; Li, W.; Zheng, D.; Terry, D. S.; Gorman, J.; Shi, W.; Zhang, B.; Zhou, T.; Ding, S.; Gasser, R.; Prevost, J.; Beaudoin-Bussieres, G.; Anand, S. P.; Laumaea, A.; Grover, J. R.; Lihong, L.; Ho, D. D.; Mascola, J.R.; Finzi, A.; Kwong, P. D.; Blanchard, S. C.; Mothes, W. Real-time conformational dynamics of SARS-CoV-2 spikes on virus particles. Cell Host Microbe 2020, 28, 880–891.e8. [Google Scholar] [CrossRef]
- Yang, Z.; Han, Y.; Ding, S.; Shi, W.; Zhou, T.; Finzi, A.; Kwong, P.D.; Mothes, W.; Lu, M. SARS-CoV-2 Variants Increase Kinetic Stability of Open Spike Conformations as an Evolutionary Strategy. mBio 2022, 13, e0322721. [Google Scholar] [CrossRef] [PubMed]
- Díaz-Salinas, M.A.; Li, Q.; Ejemel, M.; Yurkovetskiy, L.; Luban, J.; Shen, K.; Wang, Y.; Munro, J.B. Conformational dynamics and allosteric modulation of the SARS-CoV-2 spike. Elife 2022, 11, e75433. [Google Scholar] [CrossRef] [PubMed]
- Wang, Y.; Liu, C.; Zhang, C.; Wang, Y.; Hong, Q.; Xu, S.; Li, Z.; Yang, Y.; Huang, Z.; Cong, Y. Structural Basis for SARS-CoV-2 Delta Variant Recognition of ACE2 Receptor and Broadly Neutralizing Antibodies. Nat. Commun. 2022, 13, 871. [Google Scholar] [CrossRef]
- Mannar, D.; Saville, J.W.; Zhu, X.; Srivastava, S.S.; Berezuk, A.M.; Tuttle, K.S.; Marquez, A.C.; Sekirov, I.; Subramaniam, S. SARS-CoV-2 Omicron Variant: Ab Evasion and Cryo-EM Structure of Spike Protein–ACE2 Complex. Science 2022, 375, 760–764. [Google Scholar] [CrossRef]
- Hong, Q.; Han, W.; Li, J.; Xu, S.; Wang, Y.; Xu, C.; Li, Z.; Wang, Y.; Zhang, C.; Huang, Z.; Cong, Y. Molecular Basis of Receptor Binding and Ab Neutralization of Omicron. Nature 2022. [Google Scholar] [CrossRef] [PubMed]
- McCallum, M.; Czudnochowski, N.; Rosen, L.E.; Zepeda, S.K.; Bowen, J.E.; Walls, A.C.; Hauser, K.; Joshi, A.; Stewart, C.; Dillen, J.R.; Powell, A.E.; Croll, T.I.; Nix, J.; Virgin, H.W.; Corti, D.; Snell, G.; Veesler, D. Structural Basis of SARS-CoV-2 Omicron Immune Evasion and Receptor Engagement. Science 2022, 375, 864–868. [Google Scholar] [CrossRef]
- Yin, W.; Xu, Y.; Xu, P.; Cao, X.; Wu, C.; Gu, C.; He, X.; Wang, X.; Huang, S.; Yuan, Q.; Wu, K.; Hu, W.; Huang, Z.; Liu, J.; Wang, Z.; Jia, F.; Xia, K.; Liu, P.; Wang, X.; Song, B.; Zheng, J.; Jiang, H.; Cheng, X.; Jiang, Y.; Deng, S.J.; Xu, H.E. Structures of the Omicron Spike Trimer with ACE2 and an Anti-Omicron Ab. Science 2022, 375, 1048–1053. [Google Scholar] [CrossRef] [PubMed]
- Gobeil, S. M.-C.; Henderson, R.; Stalls, V.; Janowska, K.; Huang, X.; May, A.; Speakman, M.; Beaudoin, E.; Manne, K.; Li, D.; Parks, R.; Barr, M.; Deyton, M.; Martin, M.; Mansouri, K.; Edwards, R. J.; Eaton, A.; Montefiori, D. C.; Sempowski, G. D.; Saunders, K. O.; Wiehe, K.; Williams, W.; Korber, B.; Haynes, B. F.; Acharya, P. Structural Diversity of the SARS-CoV-2 Omicron Spike. Mol Cell. 2022, 82, 2050–2068.e6. [Google Scholar] [CrossRef]
- Cui, Z.; Liu, P.; Wang, N.; Wang, L.; Fan, K.; Zhu, Q.; Wang, K.; Chen, R.; Feng, R.; Jia, Z.; Yang, M.; Xu, G.; Zhu, B.; Fu, W.; Chu, T.; Feng, L.; Wang, Y.; Pei, X.; Yang, P.; Xie, X.S.; Cao, L.; Cao, Y.; Wang, X. Structural and Functional Characterizations of Infectivity and Immune Evasion of SARS-CoV-2 Omicron. Cell 2022, 185, 860–871.e13. [Google Scholar] [CrossRef]
- Wang, Q.; Mellis, I. A.; Ho, J.; Bowen, A.; Kowalski-Dobson, T.; Valdez, R.; Katsamba, P. S.; Wu, M.; Lee, C.; Shapiro, L.; Gordon, A.; Guo, Y.; Ho, D. D.; Liu, L. Recurrent SARS-CoV-2 Spike Mutations Confer Growth Advantages to Select JN.1 Sublineages. Emerg Microbes Infect. 2024, 13, 2402880. [Google Scholar] [CrossRef]
- Ito, J.; Suzuki, R.; Uriu, K.; Itakura, Y.; Zahradnik, J.; Kimura, K. T.; Deguchi, S.; Wang, L.; Lytras, S.; Tamura, T.; Kida, I.; Nasser, H.; Shofa, M.; Begum, M. M.; Tsuda, M.; Oda, Y.; Suzuki, T.; Sasaki, J.; Sasaki-Tabata, K.; Fujita, S.; Yoshimatsu, K.; Ito, H.; Nao, N.; Asakura, H.; Nagashima, M.; Sadamasu, K.; Yoshimura, K.; Yamamoto, Y.; Nagamoto, T.; Kuramochi, J.; Schreiber, G.; Suzuki, S.; Kato, M.; Ferdous, Z.; Mouri, H.; Shishido, K.; Misawa, N.; Kimura, I.; Kosugi, Y.; Lin, P.; Suganami, M.; Chiba, M.; Yoshimura, R.; Yasuda, K.; Iida, K.; Ohsumi, N.; Strange, A. P.; Sauter, D.; Nakagawa, S.; Wu, J.; Watanabe, Y.; Sakamoto, A.; Yasuhara, N.; Nakajima, Y.; Yajima, H.; Shirakawa, K.; Takaori-Kondo, A.; Nagata, K.; Kazuma, Y.; Nomura, R.; Horisawa, Y.; Tashiro, Y.; Kawa, Y.; Irie, T.; Kawabata, R.; Shimizu, R.; Takahashi, O.; Ichihara, K.; Motozono, C.; Toyoda, M.; Ueno, T.; Shibatani, Y.; Nishiuchi, T.; Saito, A.; Matsuno, K.; Takayama, K.; Hashiguchi, T.; Tanaka, S.; Fukuhara, T.; Ikeda, T.; Sato, K. Convergent Evolution of SARS-CoV-2 Omicron Subvariants Leading to the Emergence of BQ.1.1 Variant. Nat Commun. 2023, 14, 2671. [Google Scholar] [CrossRef]
- Focosi, D.; Quiroga, R.; McConnell, S.; Johnson, M.C.; Casadevall, A. Convergent Evolution in SARS-CoV-2 Spike Creates a Variant Soup from Which New COVID-19 Waves Emerge. Int. J. Mol. Sci. 2023, 24, 2264. [Google Scholar] [CrossRef]
- Jian, F.; Wang, J.; Yisimayi, A.; Song, W.; Xu, Y.; Chen, X.; Niu, X.; Yang, S.; Yu, Y.; Wang, P.; Sun, H.; Yu, L.; Wang, J.; Wang, Y.; An, R.; Wang, W.; Ma, M.; Xiao, T.; Gu, Q.; Shao, F.; Wang, Y.; Shen, Z.; Jin, R.; Cao, Y. Evolving Antibody Response to SARS-CoV-2 Antigenic Shift from XBB to JN.1. Nature 2024. [Google Scholar] [CrossRef]
- Taylor, A. L.; Starr, T. N. Deep Mutational Scanning of SARS-CoV-2 Omicron BA.2.86 and Epistatic Emergence of the KP.3 Variant. Virus Evol. 2024, 10, veae067. [Google Scholar] [CrossRef] [PubMed]
- Feng, L.; Sun, Z.; Zhang, Y.; Jian, F.; Yang, S.; Xia, K.; Yu, L.; Wang, J.; Shao, F.; Wang, X.; Cao, Y. Structural and Molecular Basis of the Epistasis Effect in Enhanced Affinity between SARS-CoV-2 KP.3 and ACE2. Cell Discov. 2024, 10, 123. [Google Scholar] [CrossRef] [PubMed]
- Kaku, Y.; Okumura, K.; Kawakubo, S.; Uriu, K.; Chen, L.; Kosugi, Y.; Uwamino, Y.; Begum, M. M.; Leong, S.; Ikeda, T.; Sadamasu, K.; Asakura, H.; Nagashima, M.; Yoshimura, K.; Ito, J.; Sato, K. Virological Characteristics of the SARS-CoV-2 XEC Variant. Lancet Infect Dis. 2024, S1473-3099(24), 00731-X. [Google Scholar] [CrossRef] [PubMed]
- Wang, Q.; Guo, Y.; Mellis, I. A.; Wu, M.; Mohri, H.; Gherasim, C.; Valdez, R.; Purpura, L. J.; Yin, M. T.; Gordon, A.; Ho, D. D. Antibody Evasiveness of SARS-CoV-2 Subvariants KP.3.1.1 and XEC. Cell Rep. 2025, 44, 115543. [Google Scholar] [CrossRef]
- Kaku, Y.; Uriu, K.; Okumura, K.; Ito, J.; Sato, K. Virological Characteristics of the SARS-CoV-2 KP.3.1.1 Variant. Lancet Infect Dis. 2024, 24, e609. [Google Scholar] [CrossRef]
- Liu, J.; Yu, Y.; Yang, S.; Jian, F.; Song, W.; Yu, L.; Shao, F.; Cao, Y. Virological and Antigenic Characteristics of SARS-CoV-2 Variants LF.7.2.1, NP.1, and LP.8.1. Lancet Infect Dis. 2025, 25, e128–e130. [Google Scholar] [CrossRef]
- Guo, C.; Yu, Y.; Liu, J.; Jian, F.; Yang, S.; Song, W.; Yu, L.; Shao, F.; Cao, Y. Antigenic and Virological Characteristics of SARS-CoV-2 Variants BA.3.2, XFG, and NB.1.8.1. Lancet Infect Dis. 2025, 25, e374–e377. [Google Scholar] [CrossRef]
- Mellis, I. A.; Wu, M.; Hong, H.; Tzang, C.-C.; Bowen, A.; Wang, Q.; Gherasim, C.; Pierce, V. M.; Shah, J. G.; Purpura, L. J.; Yin, M. T.; Gordon, A.; Guo, Y.; Ho, D. D. Antibody Evasion and Receptor Binding of SARS-CoV-2 LP.8.1.1, NB.1.8.1, XFG, and Related Subvariants. Cell Rep. 2025, 44, 116440. [Google Scholar] [CrossRef]
- Uriu, K.; Kaku, Y.; Kosugi, Y.; Chen, L.; Itoh, N.; Uwamino, Y.; Fujiwara, H.; Satoh, H.; Sato, K. Humoral Immunity Induced by XEC Monovalent Vaccines against SARS-CoV-2 Variants Including XEC, LP.8.1, NB.1.8.1, XFG, and BA.3.2. Vaccine 2026, 28311. [Google Scholar] [CrossRef]
- Wang, Y.; Hu, Y.; Chen, Z.; Zou, J.; Zhang, K.; Ren, P.; Shi, P.-Y.; Liang, B.; Xie, X. Functional and Structural Basis of Omicron BA.3.2.1 Spike. bioRxiv 2026, 2026.02.02.702852. [Google Scholar] [CrossRef]
- Barnes, C. O.; Jette, C. A.; Abernathy, M. E.; Dam, K.-M. A.; Esswein, S. R.; Gristick, H. B.; Malyutin, A. G.; Sharaf, N. G.; Huey-Tubman, K. E.; Lee, Y. E.; Robbiani, D. F.; Nussenzweig, M. C.; West, A. P., Jr.; Bjorkman, P. J. SARS-CoV-2 neutralizing antibody structures inform therapeutic strategies. Nature 2020, 588, 682–687. [Google Scholar] [CrossRef] [PubMed]
- Greaney, A. J.; Starr, T. N.; Barnes, C. O.; Weisblum, Y.; Schmidt, F.; Caskey, M.; Gaebler, C.; Cho, A.; Agudelo, M.; Finkin, S.; Wang, Z.; Poston, D.; Muecksch, F.; Hatziioannou, T.; Bieniasz, P. D.; Robbiani, D. F.; Nussenzweig, M. C.; Bjorkman, P. J.; Bloom, J. D. Mapping mutations to the SARS-CoV-2 RBD that escape binding by different classes of antibodies. Nat Commun 2021, 12, 4196. [Google Scholar] [CrossRef]
- Cao, Y.; Wang, J.; Jian, F.; Xiao, T.; Song, W.; Yisimayi, A.; Huang, W.; Li, Q.; Wang, P.; An, R.; Wang, J.; Wang, Y.; Niu, X.; Yang, S.; Liang, H.; Sun, H.; Li, T.; Yu, Y.; Cui, Q.; Liu, S.; Yang, X.; Du, S.; Zhang, Z.; Hao, X.; Shao, F.; Jin, R.; Wang, X.; Xiao, J.; Wang, Y.; Xie, X. S. Omicron Escapes the Majority of Existing SARS-CoV-2 Neutralizing Antibodies. Nature 2022, 602, 657–663. [Google Scholar] [CrossRef]
- Cao, Y.; Yisimayi, A.; Jian, F.; Song, W.; Xiao, T.; Wang, L.; Du, S.; Wang, J.; Li, Q.; Chen, X.; Yu, Y.; Wang, P.; Zhang, Z.; Liu, P.; An, R.; Hao, X.; Wang, Y.; Wang, J.; Feng, R.; Sun, H.; Zhao, L.; Zhang, W.; Zhao, D.; Zheng, J.; Yu, L.; Li, C.; Zhang, N.; Wang, R.; Niu, X.; Yang, S.; Song, X.; Chai, Y.; Hu, Y.; Shi, Y.; Zheng, L.; Li, Z.; Gu, Q.; Shao, F.; Huang, W.; Jin, R.; Shen, Z.; Wang, Y.; Wang, X.; Xiao, J.; Xie, X. S. BA.2.12.1, BA.4 and BA.5 Escape Antibodies Elicited by Omicron Infection. Nature 2022, 608, 593–602. [Google Scholar] [CrossRef]
- Cao, Y.; Jian, F.; Wang, J.; Yu, Y.; Song, W.; Yisimayi, A.; Wang, J.; An, R.; Chen, X.; Zhang, N.; Wang, Y.; Wang, P.; Zhao, L.; Sun, H.; Yu, L.; Yang, S.; Niu, X.; Xiao, T.; Gu, Q.; Shao, F.; Hao, X.; Xu, Y.; Jin, R.; Shen, Z.; Wang, Y.; Xie, X. S. Imprinted SARS-CoV-2 Humoral Immunity Induces Convergent Omicron RBD Evolution. Nature 2023, 614, 521–529. [Google Scholar] [CrossRef]
- Jian, F.; Wec, A. Z.; Feng, L.; Yu, Y.; Wang, L.; Wang, P.; Yu, L.; Wang, J.; Hou, J.; Berrueta, D. M.; Lee, D.; Speidel, T.; Ma, L.; Kim, T.; Yisimayi, A.; Song, W.; Wang, J.; Liu, L.; Yang, S.; Niu, X.; Xiao, T.; An, R.; Wang, Y.; Shao, F.; Wang, Y.; Pecetta, S.; Wang, X.; Walker, L. M.; Cao, Y. Viral Evolution Prediction Identifies Broadly Neutralizing Antibodies to Existing and Prospective SARS-CoV-2 Variants. Nat Microbiol 2025. [Google Scholar] [CrossRef] [PubMed]
- Schendel, S. L.; Yu, X.; Halfmann, P. J.; Mahita, J.; Ha, B.; Hastie, K. M.; Li, H.; Bedinger, D.; Troup, C.; Li, K.; Kuzmina, N.; Torrelles, J. B.; Munt, J. E.; Mattocks, M.; Osei-Twum, M.; Callaway, H. M.; Reece, S.; Palser, A.; Kellam, P.; Dennison, S. M.; Huntwork, R. H. C.; Horn, G. Q.; Abraha, M.; Feeney, E.; Martinez-Sobrido, L.; Pino, P. A.; Hicks, A.; Ye, C.; Park, J.-G.; Maingot, B.; Periasamy, S.; Mallory, M.; Scobey, T.; Lepage, M.-N.; St-Amant, N.; Khan, S.; Gambiez, A.; Baric, R. S.; Bukreyev, A.; Gagnon, L.; Germann, T.; Kawaoka, Y.; Tomaras, G. D.; Peters, B.; Saphire, E. O. A Global Collaboration for Systematic Analysis of Broad-Ranging Antibodies against the SARS-CoV-2 Spike Protein. Cell Rep. 2025, 44, 115499. [Google Scholar] [CrossRef] [PubMed]
- Jiang, J.; Boughter, C. T.; Ahmad, J.; Natarajan, K.; Boyd, L. F.; Meier-Schellersheim, M.; Margulies, D. H. SARS-CoV-2 Antibodies Recognize 23 Distinct Epitopic Sites on the Receptor Binding Domain. Commun Biol. 2023, 6, 953. [Google Scholar] [CrossRef]
- Feng, Z.; Sang, Z.; Xiang, Y.; Escalera, A.; Weshler, A.; Schneidman-Duhovny, D.; García-Sastre, A.; Shi, Y. One Thousand SARS-CoV-2 Antibody Structures Reveal Convergent Binding and near-Universal Immune Escape. Cell Syst. 2026, 17, 101452. [Google Scholar] [CrossRef] [PubMed]
- Rosen, L. E.; Tortorici, M. A.; De Marco, A.; Pinto, D.; Foreman, W. B.; Taylor, A. L.; Park, Y.-J.; Bohan, D.; Rietz, T.; Errico, J. M.; Hauser, K.; Dang, H. V.; Chartron, J. W.; Giurdanella, M.; Cusumano, G.; Saliba, C.; Zatta, F.; Sprouse, K. R.; Addetia, A.; Zepeda, S. K.; Brown, J.; Lee, J.; Dellota, E., Jr.; Rajesh, A.; Noack, J.; Tao, Q.; DaCosta, Y.; Tsu, B.; Acosta, R.; Subramanian, S.; de Melo, G. D.; Kergoat, L.; Zhang, I.; Liu, Z.; Guarino, B.; Schmid, M. A.; Schnell, G.; Miller, J. L.; Lempp, F. A.; Czudnochowski, N.; Cameroni, E.; Whelan, S. P. J.; Bourhy, H.; Purcell, L. A.; Benigni, F.; di Iulio, J.; Pizzuto, M. S.; Lanzavecchia, A.; Telenti, A.; Snell, G.; Corti, D.; Veesler, D.; Starr, T. N. A Potent Pan-Sarbecovirus Neutralizing Antibody Resilient to Epitope Diversification. Cell 2024, 187, 7196–7213.e26. [CrossRef]
- Wang, Q.; Guo, Y.; Casner, R. G.; Yu, J.; Nair, M. S.; Ho, J.; Reddem, E. R.; Mellis, I. A.; Wu, M.; Tzang, C.-C.; Hong, H.; Huang, Y.; Shapiro, L.; Liu, L.; Ho, D. D. Optimizing a Human Monoclonal Antibody for Better Neutralization of SARS-CoV-2. Nat Commun. 2025, 16, 6195. [Google Scholar] [CrossRef]
- Luo, M.; Zhou, R.; Tang, B.; Liu, H.; Chen, B.; Liu, N.; Mo, Y.; Zhang, P.; Lee, Y. L.; Ip, J. D.; Wing-Ho Chu, A.; Chan, W.-M.; Man, H.-O.; Chen, Y.; To, K. K.-W.; Yuen, K.-Y.; Dang, S.; Chen, Z. Ultrapotent Class I Neutralizing Antibodies Post Omicron Breakthrough Infection Overcome Broad SARS-CoV-2 Escape Variants. EBioMedicine. 2024, 108, 105354. [Google Scholar] [CrossRef]
- Schmidt, P.; Li, Y.; Popejoy, M. Immunobridging for Pemivibart, a Monoclonal Antibody for Prevention of Covid-19. N Engl J Med. 2024, 391, 1860–1862. [Google Scholar] [CrossRef] [PubMed]
- Cao, Y.; Jian, F.; Zhang, Z.; Yisimayi, A.; Hao, X.; Bao, L.; Yuan, F.; Yu, Y.; Du, S.; Wang, J.; Xiao, T.; Song, W.; Zhang, Y.; Liu, P.; An, R.; Wang, P.; Wang, Y.; Yang, S.; Niu, X.; Zhang, Y.; Gu, Q.; Shao, F.; Hu, Y.; Yin, W.; Zheng, A.; Wang, Y.; Qin, C.; Jin, R.; Xiao, J.; Xie, X. S. Rational Identification of Potent and Broad Sarbecovirus-Neutralizing Antibody Cocktails from SARS Convalescents. Cell Rep. 2022, 41, 111845. [Google Scholar] [CrossRef]
- Xie, M.; Qiu, Y.; Zhao, X.; Shi, J.; Liu, Y.; Zhang, Q.; He, J.; Li, J.; Liu, L.; Sun, S.; Zhu, Y.; Mao, Q.; Long, Y.; Oliveira, T. Y.; Wang, Z.; Zhou, Y.; Yan, Y.; Xia, A.; Zai, W.; Mayer, C. T.; Xie, Y.; Jiang, S.; Lu, L.; Xia, R.; Wu, F.; Sun, L.; Wang, P.; Chu, H.; Wang, Q. Orphan Broadly RBD-Binding Antibodies Annotate Three Remaining Conserved RBD Epitopes along SARS-CoV-2 Evolution. Nat Commun. 2025, 16, 10566. [Google Scholar] [CrossRef] [PubMed]
- Song, G.; Yuan, M.; Liu, H.; Capozzola, T.; Lin, R. N.; Torres, J. L.; He, W.; Musharrafieh, R.; Dueker, K.; Zhou, P.; Callaghan, S.; Mishra, N.; Yong, P.; Anzanello, F.; Avillion, G.; Vo, A. L.; Li, X.; Zhang, Y.; Makhdoomi, M.; Feng, Z.; Zhu, X.; Peng, L.; Nemazee, D.; Safonova, Y.; Briney, B.; Ward, A. B.; Burton, D. R.; Wilson, I. A.; Andrabi, R. Broadly Neutralizing Antibodies Targeting a Conserved Silent Face of Spike RBD Resist Extreme SARS-CoV-2 Antigenic Drift. Cell Rep. 2025, 44, 115948. [Google Scholar] [CrossRef]
- Starr, T. N.; Czudnochowski, N.; Liu, Z.; Zatta, F.; Park, Y.-J.; Addetia, A.; Pinto, D.; Beltramello, M.; Hernandez, P.; Greaney, A. J.; Marzi, R.; Glass, W. G.; Zhang, I.; Dingens, A. S.; Bowen, J. E.; Tortorici, M. A.; Walls, A. C.; Wojcechowskyj, J. A.; De Marco, A.; Rosen, L. E.; Zhou, J.; Montiel-Ruiz, M.; Kaiser, H.; Dillen, J. R.; Tucker, H.; Bassi, J.; Silacci-Fregni, C.; Housley, M. P.; di Iulio, J.; Lombardo, G.; Agostini, M.; Sprugasci, N.; Culap, K.; Jaconi, S.; Meury, M.; Dellota, E., Jr.; Abdelnabi, R.; Foo, S.-Y. C.; Cameroni, E.; Stumpf, S.; Croll, T. I.; Nix, J. C.; Havenar-Daughton, C.; Piccoli, L.; Benigni, F.; Neyts, J.; Telenti, A.; Lempp, F. A.; Pizzuto, M. S.; Chodera, J. D.; Hebner, C. M.; Virgin, H. W.; Whelan, S. P. J.; Veesler, D.; Corti, D.; Bloom, J. D.; Snell, G. SARS-CoV-2 RBD Antibodies That Maximize Breadth and Resistance to Escape. Nature 2021, 597, 97–102. [Google Scholar] [CrossRef]
- Dejnirattisai, W.; Zhou, D.; Ginn, H. M.; Duyvesteyn, H. M. E.; Supasa, P.; Case, J. B.; Zhao, Y.; Walter, T. S.; Mentzer, A. J.; Liu, C.; Wang, B.; Paesen, G. C.; Slon-Campos, J.; López-Camacho, C.; Kafai, N. M.; Bailey, A. L.; Chen, R. E.; Ying, B.; Thompson, C.; Bolton, J.; Fyfe, A.; Gupta, S.; Tan, T. K.; Gilbert-Jaramillo, J.; James, W.; Knight, M.; Carroll, M. W.; Skelly, D.; Dold, C.; Peng, Y.; Levin, R.; Dong, T.; Pollard, A. J.; Knight, J. C.; Klenerman, P.; Temperton, N.; Hall, D. R.; Williams, M. A.; Paterson, N. G.; Bertram, F. K. R.; Siebert, C. A.; Clare, D. K.; Howe, A.; Radecke, J.; Song, Y.; Townsend, A. R.; Huang, K.-Y. A.; Fry, E. E.; Mongkolsapaya, J.; Diamond, M. S.; Ren, J.; Stuart, D. I.; Screaton, G. R. The Antigenic Anatomy of SARS-CoV-2 Receptor Binding Domain. Cell 2021, 184, 2183–2200.e22. [Google Scholar] [CrossRef] [PubMed]
- Zhan, W.; Tian, X.; Zhang, X.; Xing, S.; Song, W.; Liu, Q.; Hao, A.; Hu, Y.; Zhang, M.; Ying, T.; Chen, Z.; Lan, F.; Sun, L. Structural Study of SARS-CoV-2 Antibodies Identifies a Broad-Spectrum Antibody That Neutralizes the Omicron Variant by Disassembling the Spike Trimer. J Virol. 2022, 96, e0048022. [Google Scholar] [CrossRef]
- Wang, S.; Sun, H.; Zhang, Y.; Yuan, L.; Wang, Y.; Zhang, T.; Wang, S.; Zhang, J.; Yu, H.; Xiong, H.; Tang, Z.; Liu, L.; Huang, Y.; Chen, X.; Li, T.; Ying, D.; Liu, C.; Chen, Z.; Yuan, Q.; Zhang, J.; Cheng, T.; Li, S.; Guan, Y.; Zheng, Q.; Zheng, Z.; Xia, N. Three SARS-CoV-2 Antibodies Provide Broad and Synergistic Neutralization against Variants of Concern, Including Omicron. Cell Rep. 2022, 39, 110862. [Google Scholar] [CrossRef]
- Zimmerman, M. I.; Porter, J. R.; Ward, M. D.; Singh, S.; Vithani, N.; Meller, A.; Mallimadugula, U. L.; Kuhn, C. E.; Borowsky, J. H.; Wiewiora, R. P.; Hurley, M.F.D.; Harbison, A.M.; Fogarty, C.A.; Coffland, J.E.; Fadda, E.; Voelz, V.A.; Chodera, J.D.; Bowman, G.R. SARS-CoV-2 simulations go exascale to predict dramatic spike opening and cryptic pockets across the proteome. Nat. Chem. 2021, 13, 651–659. [Google Scholar] [CrossRef]
- Verkhivker, G.M.; Agajanian, S.; Oztas, D.Y.; Gupta, G. Comparative Perturbation-Based Modeling of the SARS-CoV-2 Spike Protein Binding with Host Receptor and Neutralizing Antibodies: Structurally Adaptable Allosteric Communication Hotspots Define Spike Sites Targeted by Global Circulating Mutations. Biochemistry 2021, 60, 1459–1484. [Google Scholar] [CrossRef] [PubMed]
- Verkhivker, G.M.; Agajanian, S.; Oztas, D.Y.; Gupta, G. Dynamic Profiling of Binding and Allosteric Propensities of the SARS-CoV-2 Spike Protein with Different Classes of Antibodies: Mutational and Perturbation-Based Scanning Reveals the Allosteric Duality of Functionally Adaptable Hotspots. J. Chem. Theory Comput. 2021, 17, 4578–4598. [Google Scholar] [CrossRef]
- Xiao, S.; Alshahrani, M.; Gupta, G.; Tao, P.; Verkhivker, G. Markov State Models and Perturbation-Based Approaches Reveal Distinct Dynamic Signatures and Hidden Allosteric Pockets in the Emerging SARS-Cov-2 Spike Omicron Variant Complexes with the Host Receptor: The Interplay of Dynamics and Convergent Evolution Modulates Allostery and Functional Mechanisms. J. Chem. Inf. Model. 2023, 63, 5272–5296. [Google Scholar] [CrossRef]
- Raisinghani, N.; Alshahrani, M.; Gupta, G.; Xiao, S.; Tao, P.; Verkhivker, G. AlphaFold2 Predictions of Conformational Ensembles and Atomistic Simulations of the SARS-CoV-2 Spike XBB Lineages Reveal Epistatic Couplings between Convergent Mutational Hotspots That Control ACE2 Affinity. J. Phys. Chem. B. 2024, 128, 4696–4715. [Google Scholar] [CrossRef]
- Tulsian, N. K.; Palur, R. V.; Qian, X.; Gu, Y.; D/O Shunmuganathan, B.; Samsudin, F.; Wong, Y. H.; Lin, J.; Purushotorman, K.; Kozma, M. M.; Wang, B.; Lescar, J.; Wang, C.-I.; Gupta, R. K.; Bond, P. J.; MacAry, P. A. Defining Neutralization and Allostery by Antibodies against COVID-19 Variants. Nat Commun. 2023, 14, 6967. [Google Scholar] [CrossRef]
- Li, W.; Xu, Z.; Niu, T.; Xie, Y.; Zhao, Z.; Li, D.; He, Q.; Sun, W.; Shi, K.; Guo, W.; Chang, Z.; Liu, K.; Fan, Z.; Qi, J.; Gao, G. F. Key Mechanistic Features of the Trade-off between Antibody Escape and Host Cell Binding in the SARS-CoV-2 Omicron Variant Spike Proteins. EMBO J. 2024, 43, 1484–1498. [Google Scholar] [CrossRef] [PubMed]
- Alshahrani, M.; Parikh, V.; Foley, B.; Verkhivker, G. Dynamic Mutational Profiling of Binding Interactions and Allosteric Networks in Conformational Ensembles of the SARS-CoV-2 Spike Protein Complexes with Classes of Antibodies Targeting Cryptic Binding Sites: Confluence of Binding and Allostery Determines Molecular Mechanisms and Hotspots of Immune Escape. Phys. Chem. Chem. Phys. 2025, 27, 18263–18287. [Google Scholar] [CrossRef]
- Parra, R. G.; Schafer, N. P.; Radusky, L. G.; Tsai, M. Y.; Guzovsky, A. B.; Wolynes, P. G.; Ferreiro, D. U. Protein Frustratometer 2: A Tool to Localize Energetic Frustration in Protein Molecules, Now With Electrostatics. Nucleic Acids Res. 2016, 44(W1), W356–360. [Google Scholar] [CrossRef]
- Chen, M.; Chen, X.; SSchafer, N.P.; Clementi, C.; Komives, E.A. Ferreiro, D.U.; Wolynes, P.G. Surveying biomolecular frustration at atomic resolution. Nat. Commun. 2020, 11, 5944. [CrossRef] [PubMed]
- Parra, R. G.; Komives, E. A.; Wolynes, P. G.; Ferreiro, D. U. Frustration in Physiology and Molecular Medicine. Mol Aspects Med. 2025, 103, 101362. [Google Scholar] [CrossRef] [PubMed]
- Ma, N.; Bhattacharya, S.; Muk, S.; Jandova, Z.; Schmalhorst, P. S.; Ghosh, S.; Le, K.; Diers, E.; Trainor, N.; Farnaby, W.; Roy, M. J.; Kofink, C.; Greb, P.; Weinstabl, H.; Ciulli, A.; Bader, G.; Sankar, K.; Bergner, A.; Vaidehi, N. Frustration in the Protein-Protein Interface Plays a Central Role in the Cooperativity of PROTAC Ternary Complexes. Nat Commun. 2025, 16, 8595. [Google Scholar] [CrossRef]
- Alshahrani, M.; Parikh, V.; Foley, B.; Verkhivker, G. Multiscale Modeling and Dynamic Mutational Profiling of Binding Energetics and Immune Escape for Class I Antibodies with SARS-CoV-2 Spike Protein: Dissecting Mechanisms of High Resistance to Viral Escape Against Emerging Variants. Viruses 2025, 17, 1029. [Google Scholar] [CrossRef]
- Alshahrani, M.; Parikh, V.; Foley, B.; Verkhivker, G. Dissecting Binding and Immune Evasion Mechanisms for Ultrapotent Class I and Class 4/1 Neutralizing Antibodies of SARS-CoV-2 Spike Protein Using a Multi-Pronged Computational Approach: Neutral Frustration Architecture of Binding Interfaces and Immune Escape Hotspots Drives Adaptive Evolution. Phys. Chem. Chem. Phys. 2026, 28, 4518–4543. [Google Scholar] [CrossRef]
- Alshahrani, M.; Parikh, V.; Foley, B.; Verkhivker, G. Dynamic Mutational Profiling of Binding Interactions and Allosteric Networks in Conformational Ensembles of the SARS-CoV-2 Spike Protein Complexes with Classes of Antibodies Targeting Cryptic Binding Sites: Confluence of Binding and Allostery Determines Molecular Mechanisms and Hotspots of Immune Escape. Phys. Chem. Chem. Phys. 2025, 27, 18263–18287. [Google Scholar] [CrossRef]
- Cui, L.; Li, T.; Xue, W.; Zhang, S.; Wang, H.; Liu, H.; Gu, Y.; Xia, N.; Li, S. Comprehensive Overview of Broadly Neutralizing Antibodies against SARS-CoV-2 Variants. Viruses 2024, 16, 900. [Google Scholar] [CrossRef] [PubMed]
- Piccoli, L.; Park, Y.-J.; Tortorici, M. A.; Czudnochowski, N.; Walls, A. C.; Beltramello, M.; Silacci-Fregni, C.; Pinto, D.; Rosen, L. E.; Bowen, J. E.; Acton, O. J.; Jaconi, S.; Guarino, B.; Minola, A.; Zatta, F.; Sprugasci, N.; Bassi, J.; Peter, A.; De Marco, A.; Nix, J. C.; Mele, F.; Jovic, S.; Rodriguez, B. F.; Gupta, S. V.; Jin, F.; Piumatti, G.; Lo Presti, G.; Pellanda, A. F.; Biggiogero, M.; Tarkowski, M.; Pizzuto, M. S.; Cameroni, E.; Havenar-Daughton, C.; Smithey, M.; Hong, D.; Lepori, V.; Albanese, E.; Ceschi, A.; Bernasconi, E.; Elzi, L.; Ferrari, P.; Garzoni, C.; Riva, A.; Snell, G.; Sallusto, F.; Fink, K.; Virgin, H. W.; Lanzavecchia, A.; Corti, D.; Veesler, D. Mapping Neutralizing and Immunodominant Sites on the SARS-CoV-2 Spike Receptor-Binding Domain by Structure-Guided High-Resolution Serology. Cell 2020, 183, 1024–1042.e21. [Google Scholar] [CrossRef]
- Zhao, Z.; Zhou, J.; Tian, M.; Huang, M.; Liu, S.; Xie, Y.; Han, P.; Bai, C.; Han, P.; Zheng, A.; Fu, L.; Gao, Y.; Peng, Q.; Li, Y.; Chai, Y.; Zhang, Z.; Zhao, X.; Song, H.; Qi, J.; Wang, Q.; Wang, P.; Gao, G. F. Omicron SARS-CoV-2 Mutations Stabilize Spike up-RBD Conformation and Lead to a Non-RBM-Binding Monoclonal Antibody Escape. Nat Commun. 2022, 13, 4958. [Google Scholar] [CrossRef]
- Nutalai, R.; Zhou, D.; Tuekprakhon, A.; Ginn, H. M.; Supasa, P.; Liu, C.; Huo, J.; Mentzer, A. J.; Duyvesteyn, H. M. E.; Dijokaite-Guraliuc, A.; Skelly, D.; Ritter, T. G.; Amini, A.; Bibi, S.; Adele, S.; Johnson, S. A.; Constantinides, B.; Webster, H.; Temperton, N.; Klenerman, P.; Barnes, E.; Dunachie, S. J.; Crook, D.; Pollard, A. J.; Lambe, T.; Goulder, P.; Paterson, N. G.; Williams, M. A.; Hall, D. R.; Mongkolsapaya, J.; Fry, E. E.; Dejnirattisai, W.; Ren, J.; Stuart, D. I.; Screaton, G. R. Potent Cross-Reactive Antibodies Following Omicron Breakthrough in Vaccinees. Cell 2022, 185, 2116–2131.e18. [Google Scholar] [CrossRef] [PubMed]
- Nithin, C.; Fornari, R. P.; Pilla, S. P.; Wroblewski, K.; Zalewski, M.; Madaj, R.; Kolinski, A.; Macnar, J. M.; Kmiecik, S. Exploring Protein Functions from Structural Flexibility Using CABS-Flex Modeling. Protein Sci. 2024, 33, e5090. [Google Scholar] [CrossRef] [PubMed]
- Rose, P. W.; Prlic, A.; Altunkaya, A.; Bi, C.; Bradley, A. R.; Christie, C. H.; Costanzo, L. D.; Duarte, J. M.; Dutta, S.; Feng, Z.; Green, R. K.; Goodsell, D. S.; Hudson, B.; Kalro, T.; Lowe, R.; Peisach, E.; Randle, C.; Rose, A. S.; Shao, C.; Tao, Y. P.; Valasatava, Y.; Voigt, M.; Westbrook, J. D.; Woo, J.; Yang, H.; Young, J. Y.; Zardecki, C.; Berman, H. M.; Burley, S. K. The RCSB protein data bank: integrative view of protein, gene and 3D structural information. Nucleic Acids Res. 2017, 45, D271–D281. [Google Scholar]
- Hooft, R. W.; Sander, C.; Vriend, G. Positioning hydrogen atoms by optimizing hydrogen-bond networks in protein structures. Proteins 1996, 26, 363–376. [Google Scholar] [CrossRef]
- Hekkelman, M. L.; Te Beek, T. A.; Pettifer, S. R.; Thorne, D.; Attwood, T. K.; Vriend, G. WIWS: A protein structure bioinformatics web service collection. Nucleic Acids Res. 2010, 38, W719–W723. [Google Scholar] [CrossRef] [PubMed]
- Fiser, A.; Sali, A. ModLoop: Automated modeling of loops in protein structures. Bioinformatics 2003, 19, 2500–25001. [Google Scholar] [CrossRef] [PubMed]
- Fernandez-Fuentes, N.; Zhai, J.; Fiser, A. ArchPRED: A template based loop structure prediction server. Nucleic Acids Res. 2006, 34, W173–W176. [Google Scholar] [CrossRef] [PubMed]
- Ko, J.; Lee, D.; Park, H.; Coutsias, E. A.; Lee, J.; Seok, C. The FALC-Loop web server for protein loop modeling. Nucleic Acids Res. 2011, 39, W210–W214. [Google Scholar] [CrossRef]
- Krivov, G. G.; Shapovalov, M. V.; Dunbrack, R. L., Jr. Improved prediction of protein side-chain conformations with SCWRL4. Proteins 2009, 77, 778–795. [Google Scholar] [CrossRef]
- Bhattacharya, D.; Nowotny, J.; Cao, R.; Cheng, J. 3Drefine: an interactive web server for efficient protein structure refinement. Nucleic Acids Res. 2016, 44, W406–W409. [Google Scholar] [CrossRef]
- Kmiecik, S.; Kolinski, A. Characterization of protein-folding pathways by reduced-space modeling. Proc. Natl. Acad. Sci. U. S. A 2007, 104, 12330–12335. [Google Scholar] [CrossRef]
- Kmiecik, S.; Gront, D.; Kolinski, M.; Wieteska, L.; Dawid, A.E.; Kolinski, A. Coarse-grained protein models and their applications. Chem. Rev. 2016, 116, 7898–7936. [Google Scholar] [CrossRef]
- Kmiecik, S.; Kouza, M.; Badaczewska-Dawid, A.E.; Kloczkowski, A.; Kolinski, A. Modeling of protein structural flexibility and large-scale dynamics: Coarse-grained simulations and elastic network models. Int. J. Mol. Sci. 2018, 19, 3496. [Google Scholar] [CrossRef]
- Ciemny, M.P.; Badaczewska-Dawid, A.E.; Pikuzinska, M.; Kolinski, A.; Kmiecik, S. Modeling of disordered protein structures using monte carlo simulations and knowledge-based statistical force fields. Int. J. Mol. Sci. 2019, 20, 606. [Google Scholar] [CrossRef] [PubMed]
- Kurcinski, M.; Oleniecki, T.; Ciemny, M.P.; Kuriata, A.; Kolinski, A.; Kmiecik, S. CABS-flex standalone: A simulation environment for fast modeling of protein flexibility. Bioinformatics 2019, 35, 694–695. [Google Scholar] [CrossRef]
- Badaczewska-Dawid, A. E.; Kolinski, A.; Kmiecik, S. Protocols for fast simulations of protein structure flexibility using CABS-Flex and SURPASS. Methods Mol. Biol. 2020, 2165, 337–353. [Google Scholar] [CrossRef]
- Marti-Renom, M. A.; Stuart, A. C.; Fiser, A.; Sanchez, R.; Melo, F.; Sali, A. Comparative protein structure modeling of genes and genomes. Annu. Rev. Biophys. Biomol. Struct. 2000, 29, 291–325. [Google Scholar] [CrossRef]
- Søndergaard, C. R.; Olsson, M. H.; Rostkowski, M.; Jensen, J. H. Improved treatment of ligands and coupling effects in empirical calculation and rationalization of pKa values. J. Chem. Theory Comput. 2011, 7, 2284–2295. [Google Scholar] [CrossRef]
- Olsson, M. H.; Søndergaard, C. R.; Rostkowski, M.; Jensen, J. H. PROPKA3: consistent treatment of internal and surface residues in empirical pKa predictions. J. Chem. Theory Comput. 2011, 7, 525–537. [Google Scholar] [CrossRef]
- Lee, J.; Cheng, X.; Swails, J. M.; Yeom, M. S.; Eastman, P. K.; Lemkul, J. A.; Wei, S.; Buckner, J.; Jeong, J. C.; Qi, Y.; Jo, S.; Pande, V. S.; Case, D. A.; Brooks, C. L., III; MacKerell, A. D., Jr.; Klauda, J. B.; Im, W. CHARMM-GUI Input Generator for NAMD, GROMACS, AMBER, OpenMM, and CHARMM/OpenMM Simulations Using the CHARMM36 Additive Force Field. J Chem Theory Comput. 2016, 12, 405–413. [Google Scholar] [CrossRef] [PubMed]
- Park, S.-J.; Lee, J.; Qi, Y.; Kern, N. R.; Lee, H. S.; Jo, S.; Joung, I.; Joo, K.; Lee, J.; Im, W. CHARMM-GUI Glycan Modeler for Modeling and Simulation of Carbohydrates and Glycoconjugates. Glycobiology 2019, 29, 320–331. [Google Scholar] [CrossRef] [PubMed]
- Phillips, J.C.; Hardy, D.J.; Maia, J.D.C.; Stone, J.E.; Ribeiro, J.V.; Bernardi, R.C.; Buch, R.; Fiorin, G.; Hénin, J.; Jiang, W.; et al. Scalable Molecular Dynamics on CPU and GPU Architectures with NAMD. J. Chem. Phys. 2020, 153, 044130. [Google Scholar] [CrossRef]
- Huang, J.; Rauscher, S.; Nawrocki, G.; Ran, T.; Feig, M.; de Groot, B.L.; Grubmüller, H.; MacKerell, A.D., Jr. CHARMM36m: An improved force field for folded and intrinsically disordered proteins. Nat. Methods 2017, 14, 71–73. [Google Scholar] [CrossRef]
- Jorgensen, W.L.; Chandrasekhar, J.; Madura, J.D.; Impey, R.W.; Klein, M.L. Comparison of Simple Potential Functions for Simulating Liquid Water. J. Chem. Phys. 1983, 79, 926–935. [Google Scholar] [CrossRef]
- Hoover, W. G. Canonical Dynamics: Equilibrium Phase-Space Distributions. Phys. Rev. A 1985, 31, 1695–1697. [Google Scholar] [CrossRef]
- Parrinello, M.; Rahman, A. Polymorphic transitions in single crystals: a new molecular dynamics method. J. Appl. Phys. 1981, 52, 7182–7190. [Google Scholar] [CrossRef]
- Di Pierro, M.; Elber, R.; Leimkuhler, B. A Stochastic Algorithm for the Isobaric-Isothermal Ensemble with Ewald Summations for All Long Range Forces. J. Chem. Theory Comput. 2015, 11, 5624–5637. [Google Scholar] [CrossRef]
- Martyna, G.J.; Tobias, D.J.; Klein, M.L. Constant pressure molecular dynamics algorithms. J. Chem. Phys. 1994, 101, 4177–4189. [Google Scholar] [CrossRef]
- Feller, S.E.; Zhang, Y.; Pastor, R.W.; Brooks, B.R. Constant pressure molecular dynamics simulation: The Langevin piston method. J. Chem. Phys. 1995, 103, 4613–4621. [Google Scholar] [CrossRef]
- Davidchack, R.L.; Handel, R.; Tretyakov, M.V. Langevin thermostat for rigid body dynamics. J. Chem. Phys. 2009, 130, 234101. [Google Scholar] [CrossRef]
- Dehouck, Y.; Kwasigroch, J.M.; Rooman, M.; Gilis, D. BeAtMuSiC: Prediction of changes in protein-protein binding affinity on mutations. Nucleic Acids Res. 2013, 41, W333–W339. [Google Scholar] [CrossRef] [PubMed]
- Dehouck, Y.; Gilis, D.; Rooman, M. A new generation of statistical potentials for proteins. Biophys. J. 2006, 90, 4010–4017. [Google Scholar] [CrossRef] [PubMed]
- Dehouck, Y.; Grosfils, A.; Folch, B.; Gilis, D.; Bogaerts, P.; Rooman, M. Fast and accurate predictions of protein stability changes upon mutations using statistical potentials and neural networks:PoPMuSiC-2.0. Bioinformatics 2009, 25, 2537–2543. [Google Scholar] [CrossRef] [PubMed]
- Srinivasan, J.; Cheatham, T.E.; Cieplak, P.; Kollman, P.A.; Case, D.A. Continuum Solvent Studies of the Stability of DNA, RNA, and Phosphoramidate−DNA Helices. J. Amer. Chem. Soc. 1998, 120, 9401–9409. [Google Scholar] [CrossRef]
- Kollman, P.A.; Massova, I.; Reyes, C.; Kuhn, B.; Huo, S.; Chong, L.; Lee, M.; Lee, T.; Duan, Y.; Wang, W.; et al. Calculating Structures and Free Energies of Complex Molecules: Combining Molecular Mechanics and Continuum Models. Acc. Chem. Res. 2000, 33, 889–897. [Google Scholar] [CrossRef]
- Hou, T.; Wang, J.; Li, Y.; Wang, W. Assessing the Performance of the MM/PBSA and MM/GBSA Methods. 1. The Accuracy of Binding Free Energy Calculations Based on Molecular Dynamics Simulations. J. Chem. Inf. Model. 2011, 51, 69–82. [Google Scholar] [CrossRef]
- Weng, G.; Wang, E.; Wang, Z.; Liu, H.; Zhu, F.; Li, D.; Hou, T. HawkDock: A Web Server to Predict and Analyze the Protein–Protein Complex Based on Computational Docking and MM/GBSA. Nucleic Acids Res. 2019, 47, W322–W330. [Google Scholar] [CrossRef] [PubMed]
- Mongan, J.; Simmerling, C.; McCammon, J.A.; Case, D.A.; Onufriev, A. Generalized Born Model with a Simple, Robust Molecular Volume Correction. J. Chem. Theory Comput. 2007, 3, 156–169. [Google Scholar] [CrossRef]
- Williams, A.H.; Zhan, C.-G. Generalized Methodology for the Quick Prediction of Variant SARS-CoV-2 Spike Binding Affinities with Human Angiotensin-Converting Enzyme II. J. Phys. Chem. B 2022, 126, 2353–2360. [Google Scholar] [CrossRef]
- Sun, H.; Duan, L.; Chen, F.; Liu, H.; Wang, Z.; Pan, P.; Zhu, F.; Zhang, J.Z.H.; Hou, T. Assessing the Performance of MM/PBSA and MM/GBSA Methods. 7. Entropy Effects on the Performance of End-Point Binding Free Energy Calculation Approaches. Phys. Chem. Chem. Phys. 2018, 20, 14450–14460. [Google Scholar] [CrossRef]
- Case, D. A.; Aktulga, H. M.; Belfon, K.; Cerutti, D. S.; Cisneros, G. A.; Cruzeiro, V. W. D.; Forouzesh, N.; Giese, T. J.; Götz, A. W.; Gohlke, H.; Izadi, S.; Kasavajhala, K.; Kaymak, M. C.; King, E.; Kurtzman, T.; Lee, T.-S.; Li, P.; Liu, J.; Luchko, T.; Luo, R.; Manathunga, M.; Machado, M. R.; Nguyen, H. M.; O’Hearn, K. A.; Onufriev, A. V.; Pan, F.; Pantano, S.; Qi, R.; Rahnamoun, A.; Risheh, A.; Schott-Verdugo, S.; Shajan, A.; Swails, J.; Wang, J.; Wei, H.; Wu, X.; Wu, Y.; Zhang, S.; Zhao, S.; Zhu, Q.; Cheatham, T. E., III; Roe, D. R.; Roitberg, A.; Simmerling, C.; York, D. M.; Nagan, M. C.; Merz, K. M., Jr. AmberTools. J. Chem. Inf. Model. 2023, 63, 6183–6191. [Google Scholar] [CrossRef]
- Miller, B. R., III; McGee, T. D., Jr.; Swails, J. M.; Homeyer, N.; Gohlke, H.; Roitberg, A. E. MMPBSA.Py: An Efficient Program for End-State Free Energy Calculations. J Chem Theory Comput. 2012, 8, 3314–3321. [Google Scholar] [CrossRef] [PubMed]
- Valdés-Tresanco, M. S.; Valdés-Tresanco, M. E.; Valiente, P. A.; Moreno, E. gmx_MMPBSA: A New Tool to Perform End-State Free Energy Calculations with GROMACS. J Chem Theory Comput. 2021, 17, 6281–6291. [Google Scholar] [CrossRef]
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2026 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).



