Submitted:
03 April 2026
Posted:
07 April 2026
You are already at the latest version
Abstract
Keywords:
1. Introduction
- What is the optimal percentage of synthetic data beyond which performance gains cease or begin to decline?
- How does synthetic data affect different neural network architectures?
- How does targeted augmentation of minority classes affect model behavior?
- How sensitive are models to the choice of augmentation strategy?
- In which task scenarios is this approach trustworthy?
2. Materials and Methods
2.1. Datasets
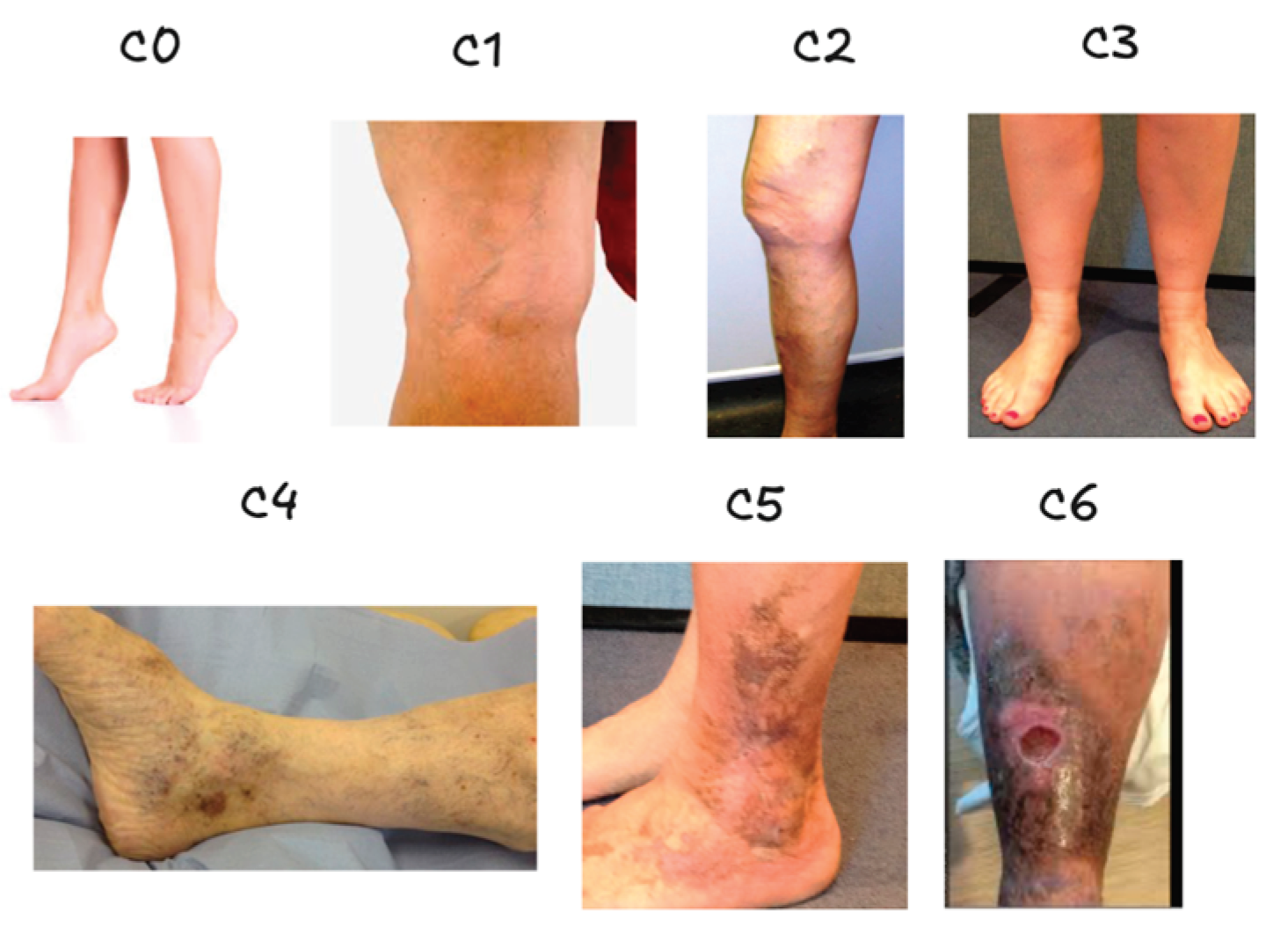
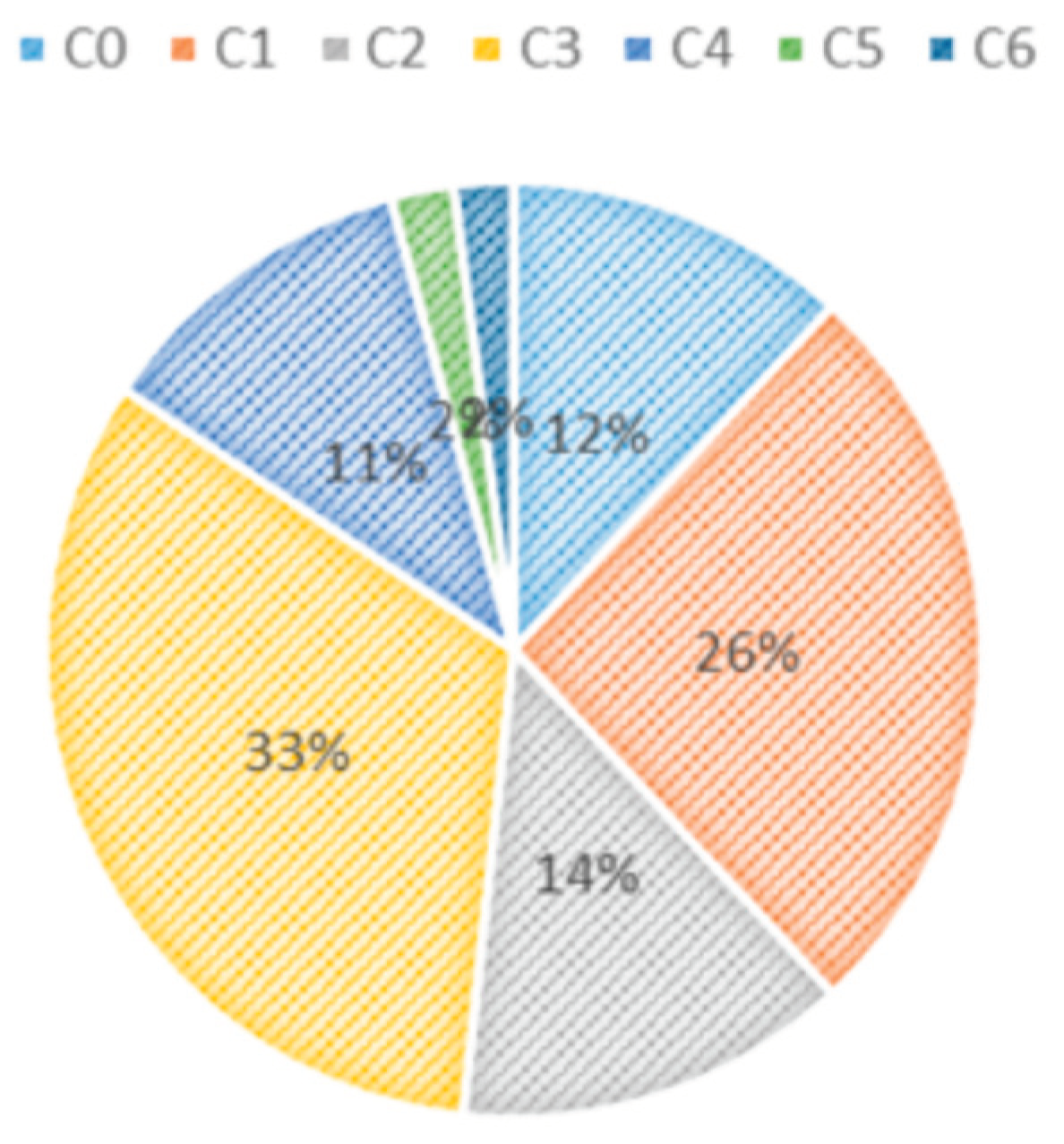
| Class | C0 | C1 | C2 | C3 | C4 | C5 | C6 |
| Number of images | 2494 | 5495 | 2861 | 6850 | 2386 | 458 | 427 |
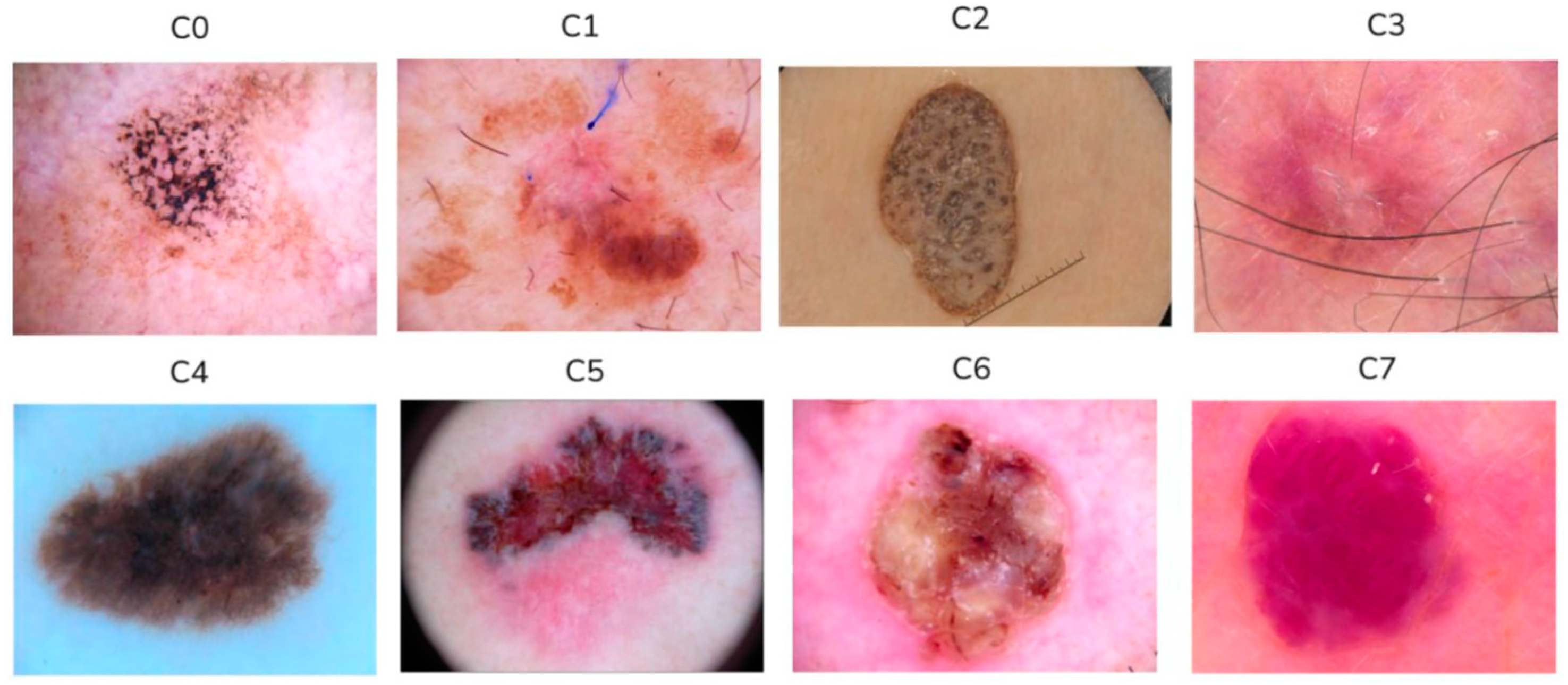
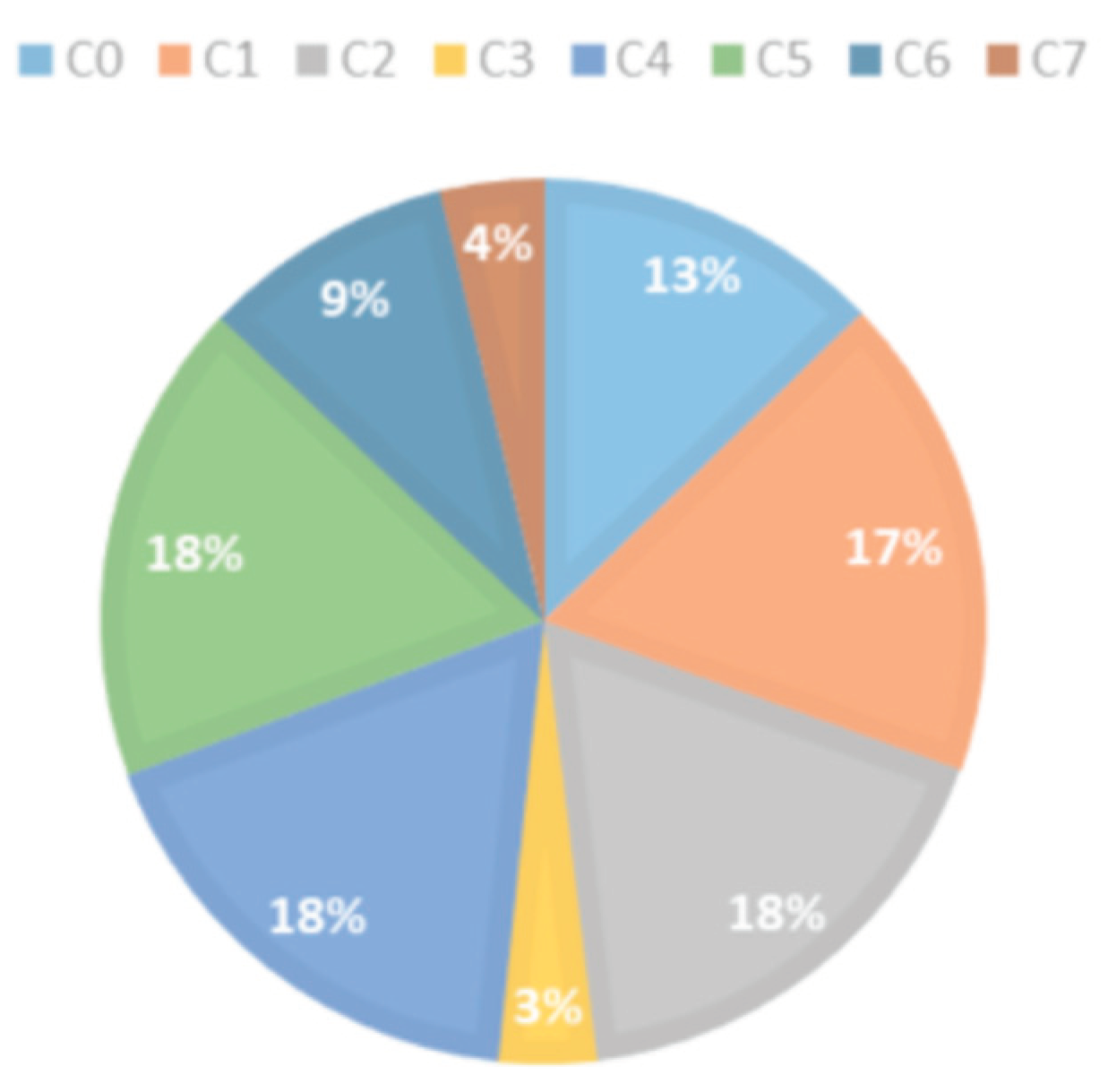
| Class | C0 | C1 | C2 | C3 | C4 | C5 | C6 | C7 |
| Number of images | 882 | 1154 | 1221 | 204 | 1222 | 1221 | 611 | 271 |
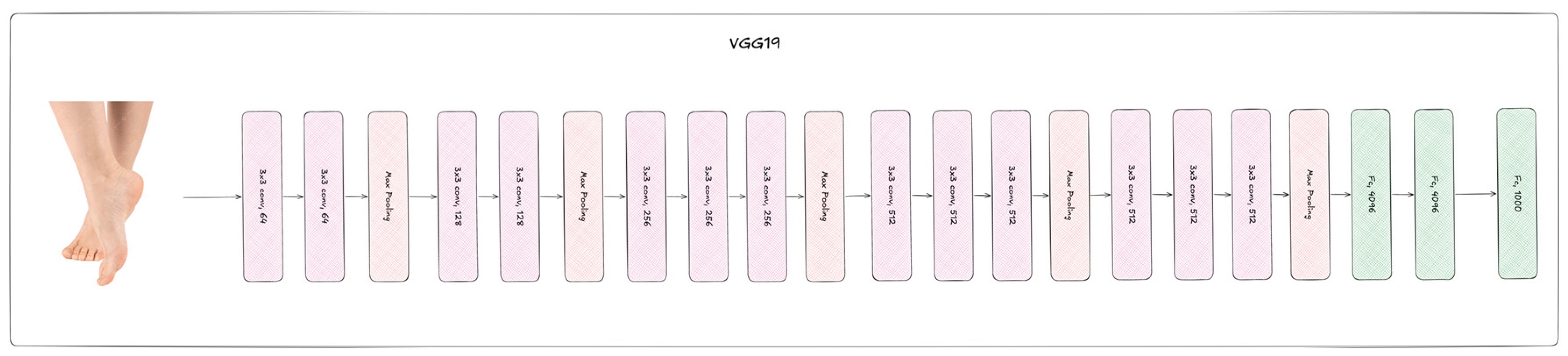
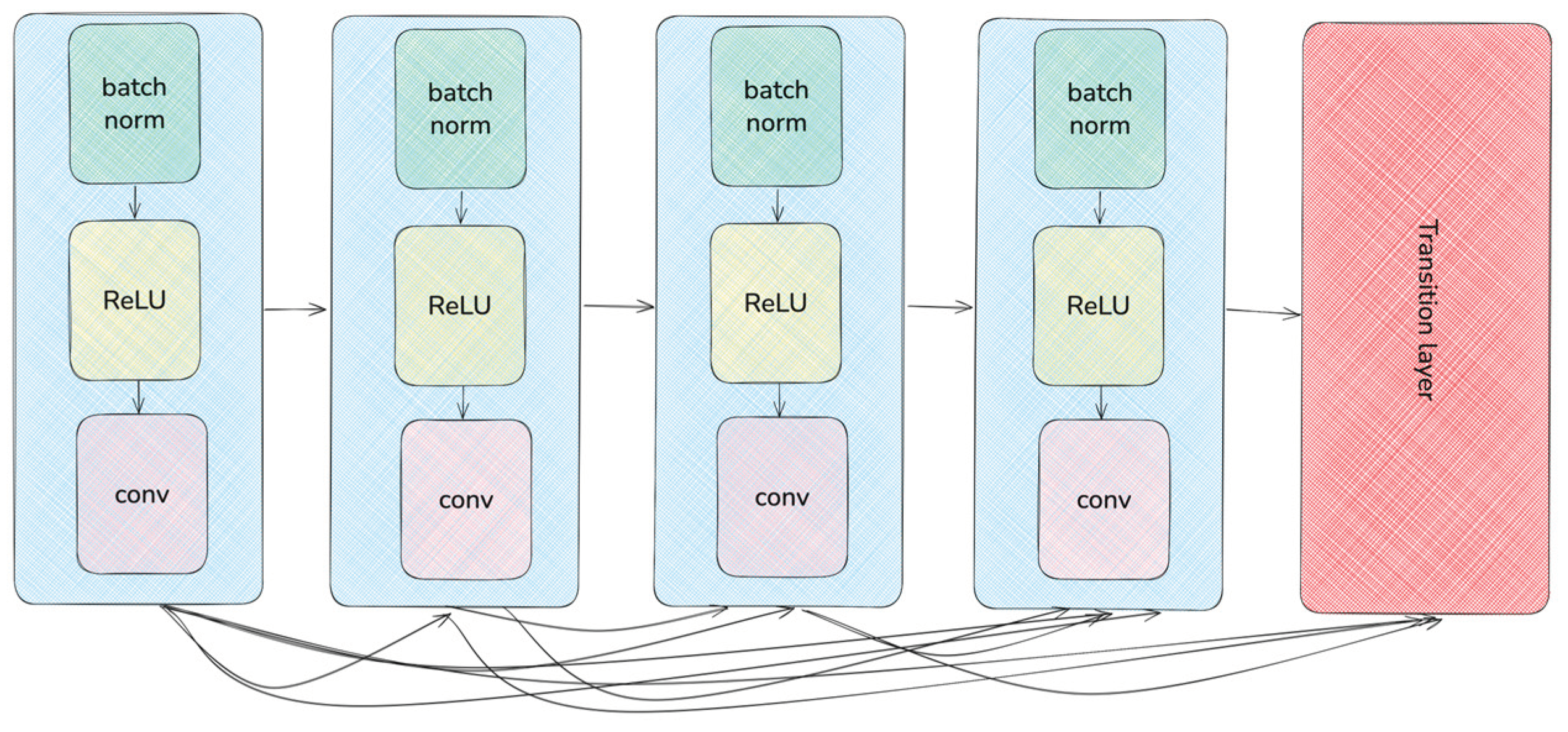
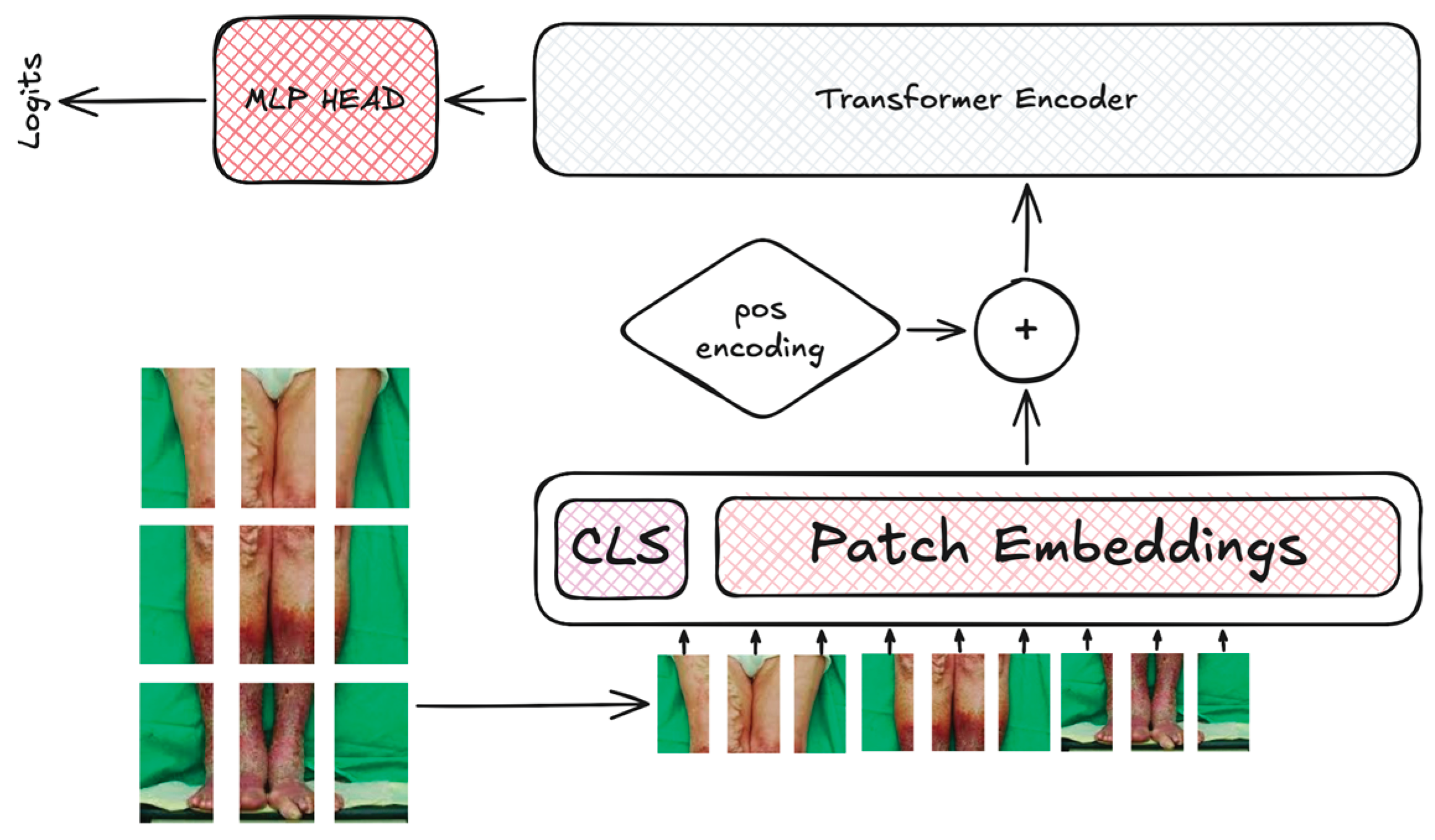
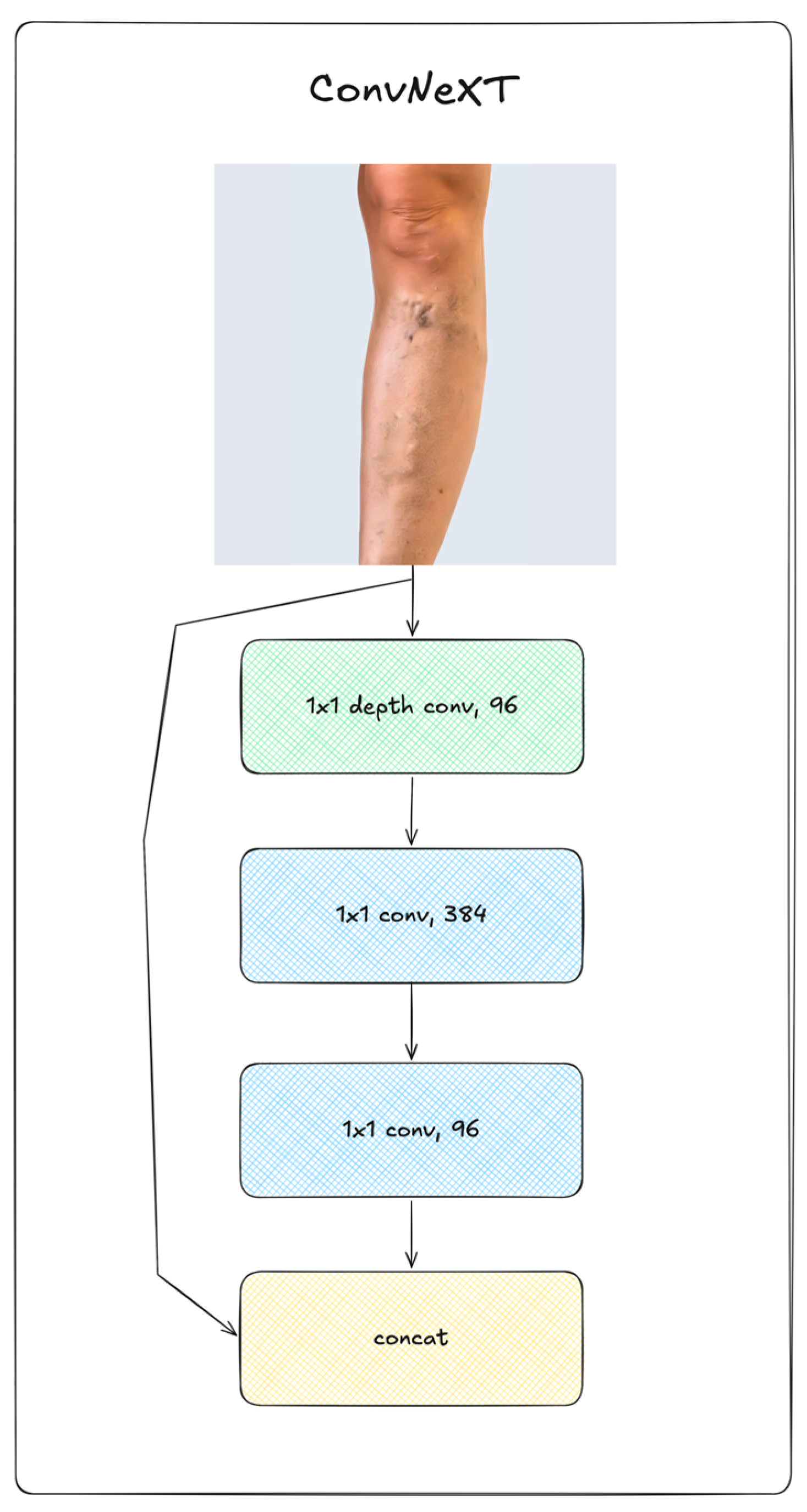
2.2. Deep Learning Neural Networks
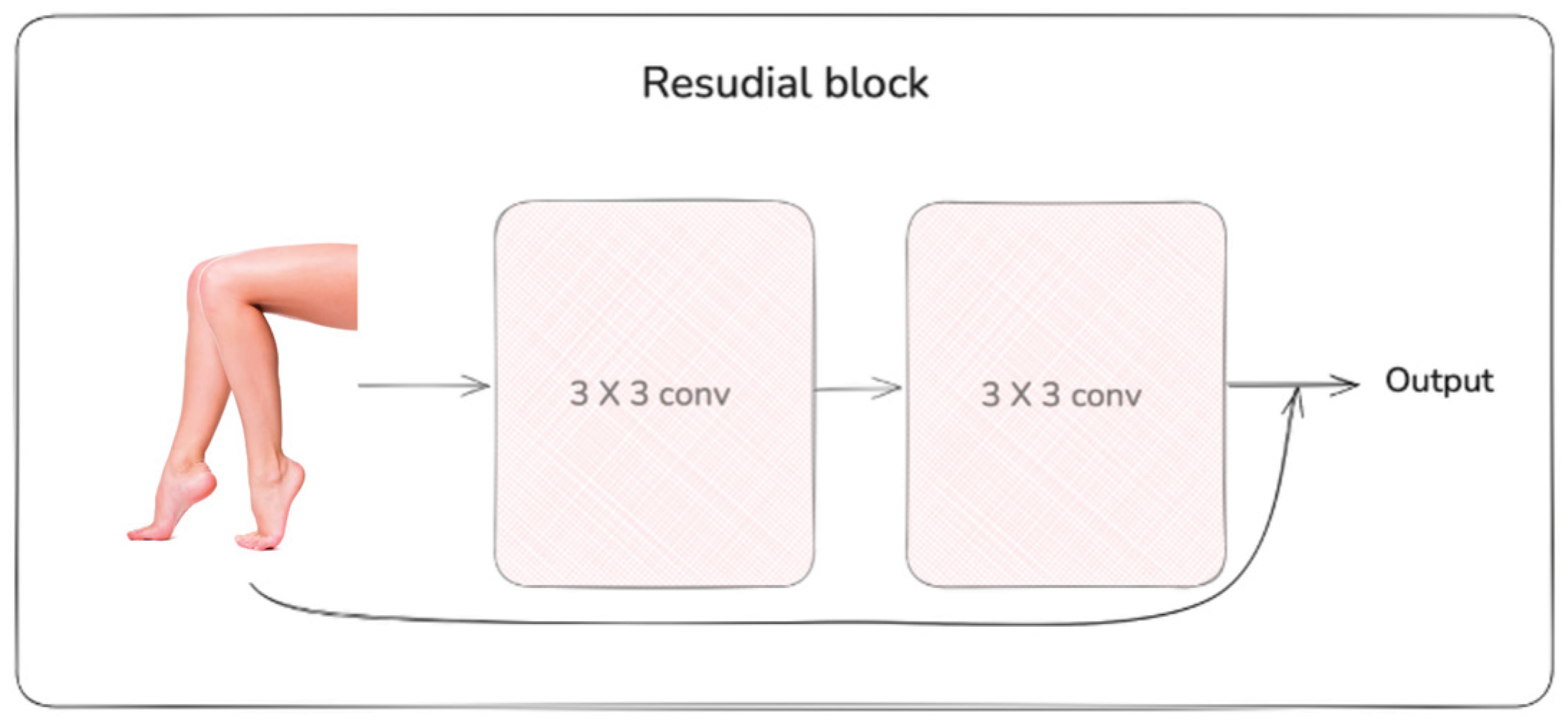
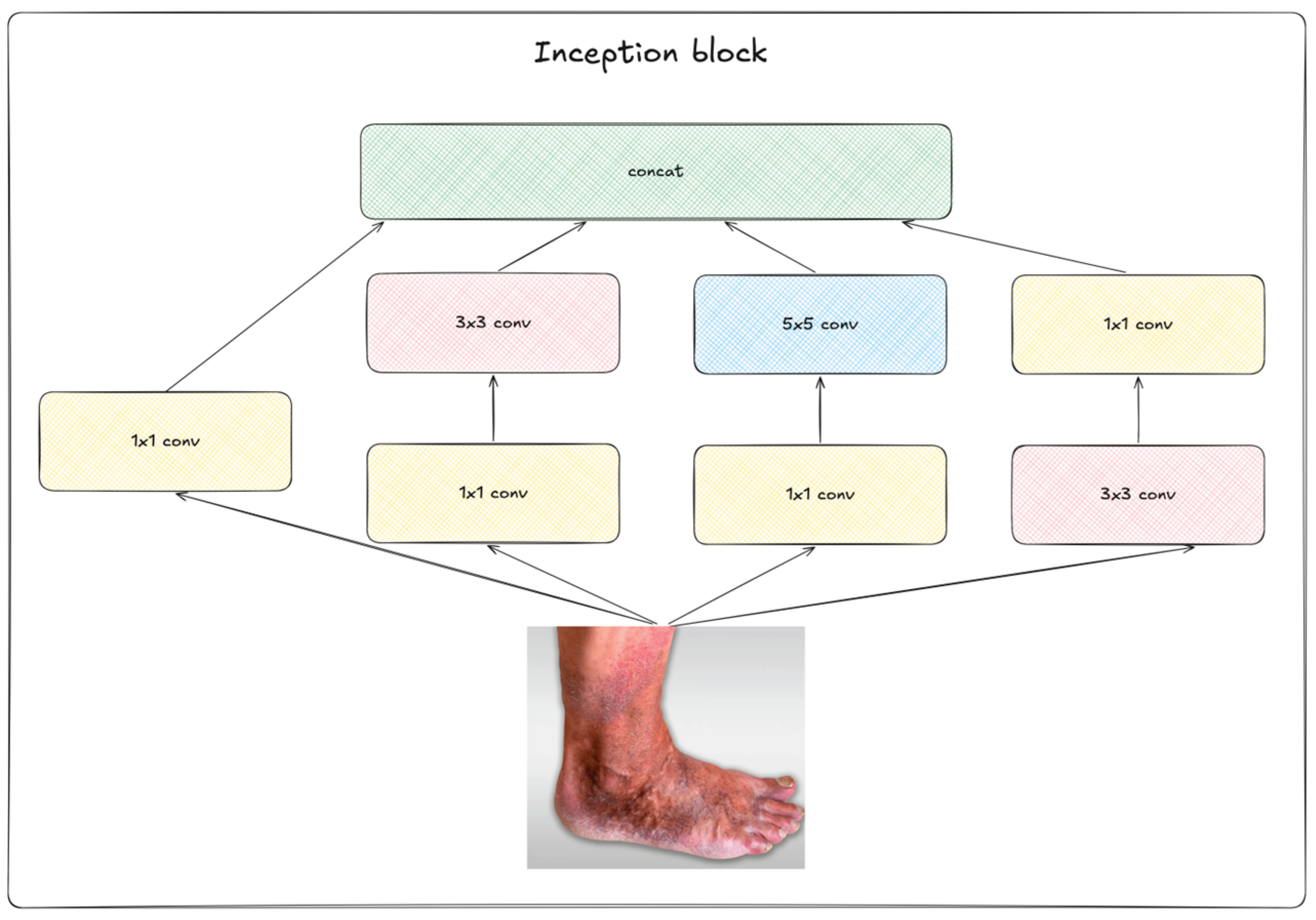
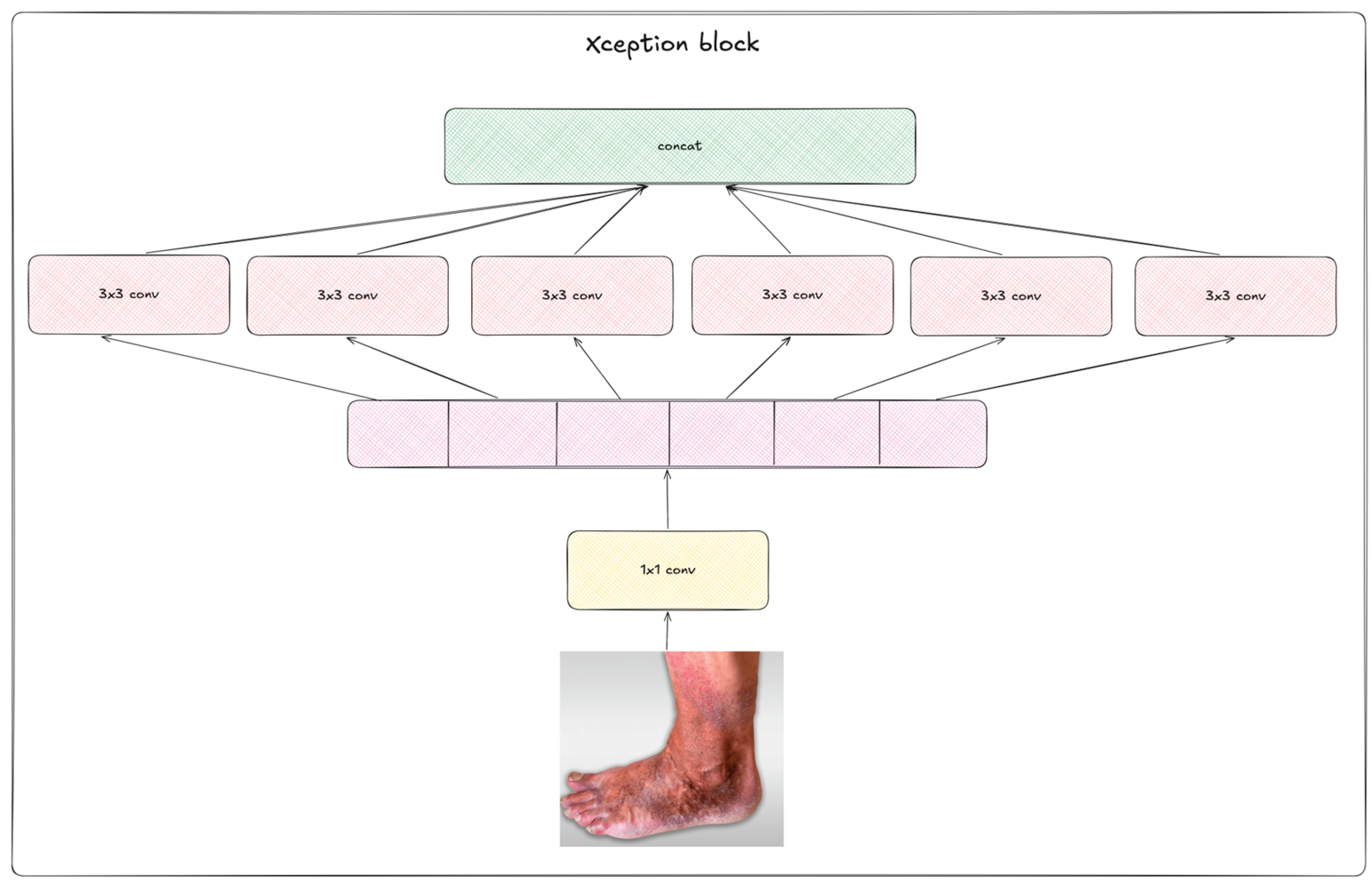
2.3. Augmentation Methods
2.4. Training Procedure
2.5. Metrics
3. Results
4. Discussion
5. Conclusions
Author Contributions
Funding
Data Availability Statement
Acknowledgments
Conflicts of Interest
Appendix A
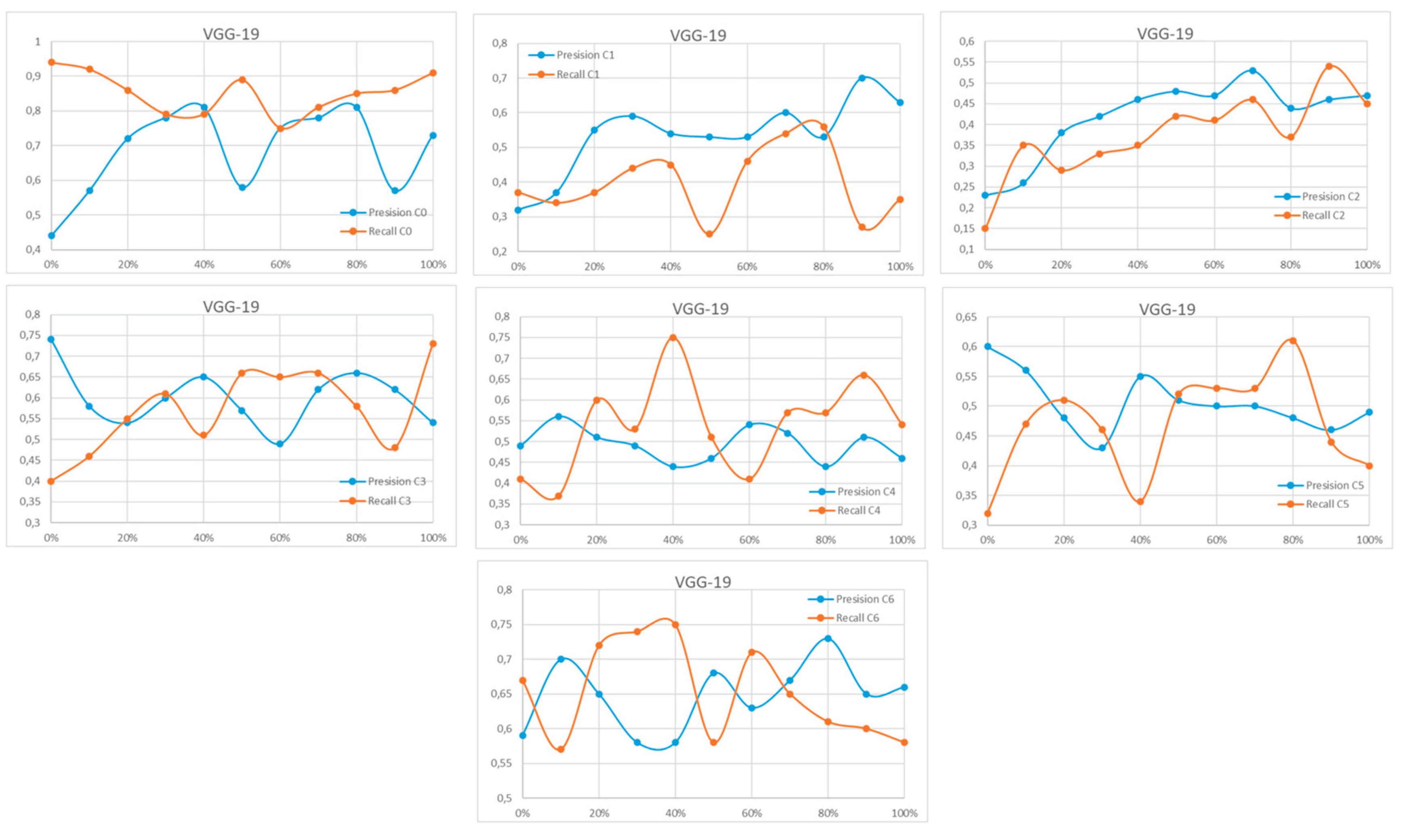
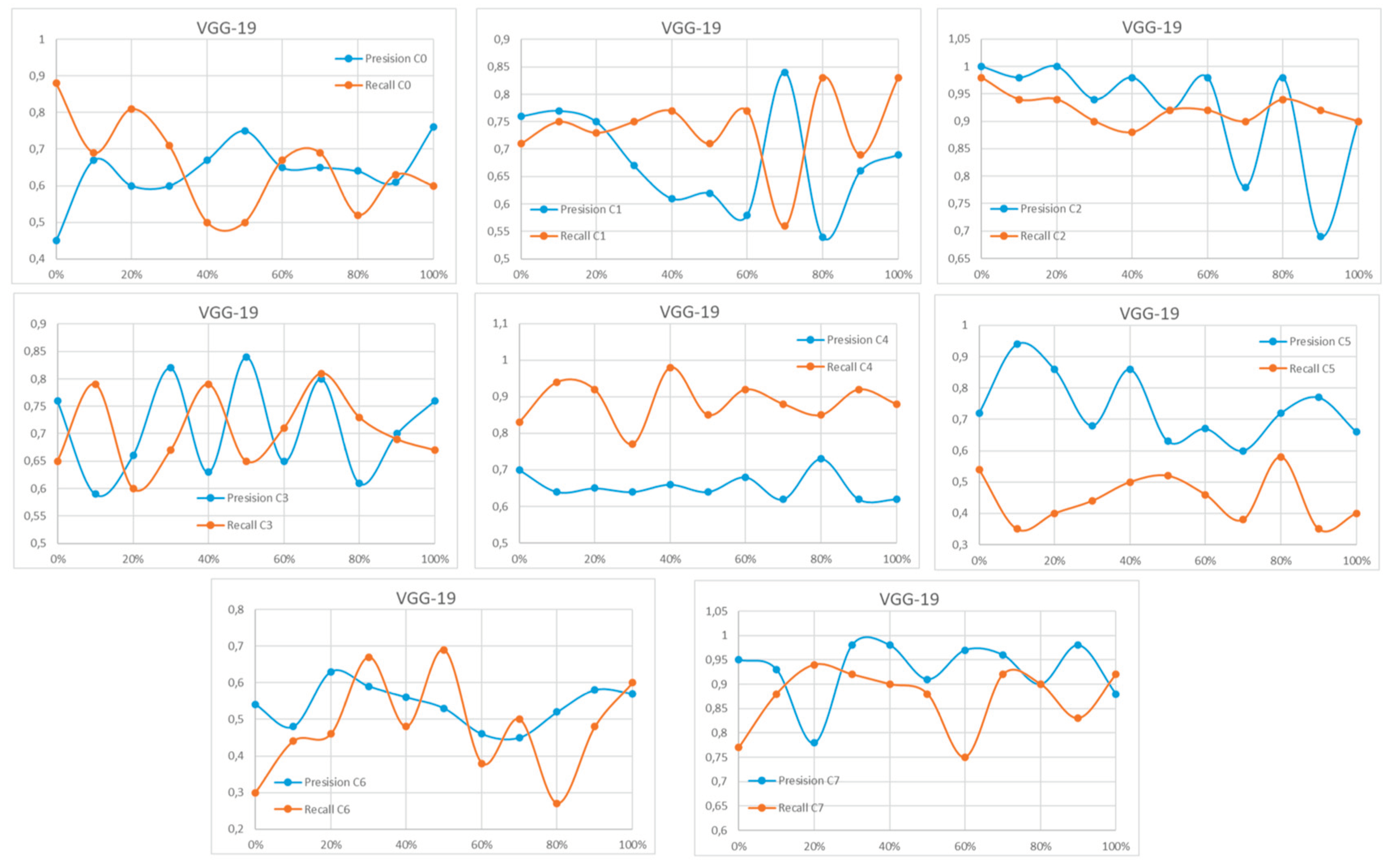
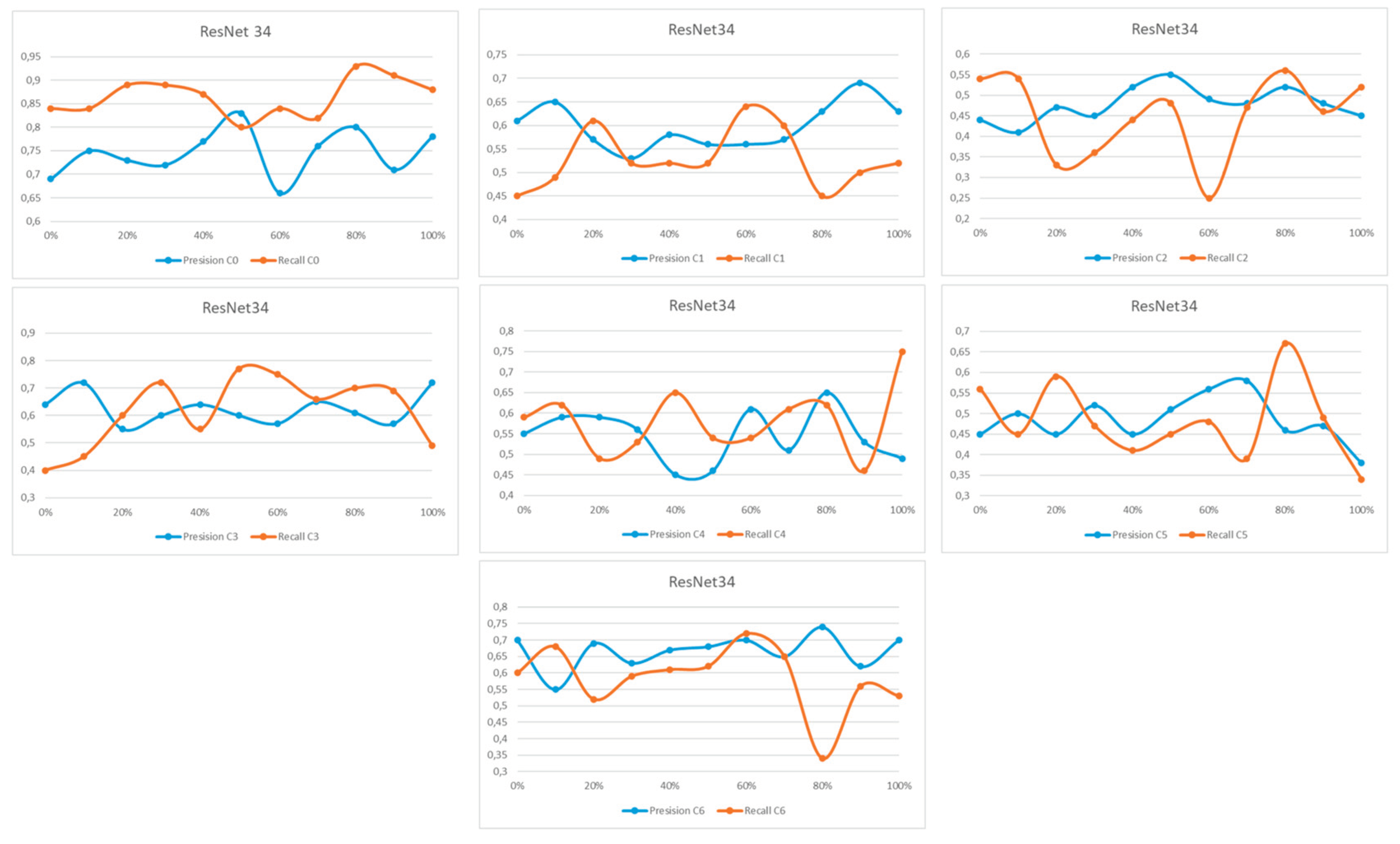
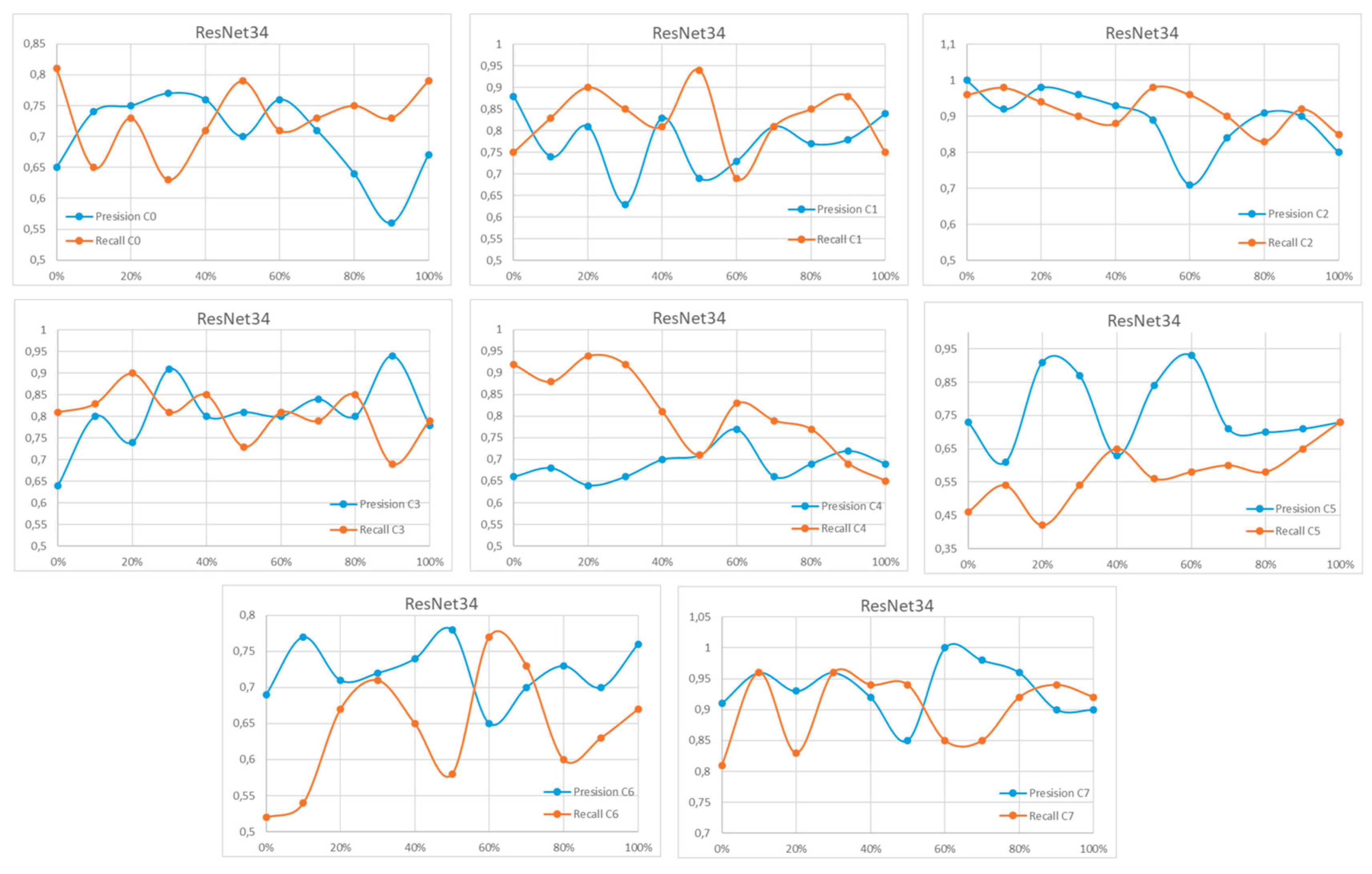
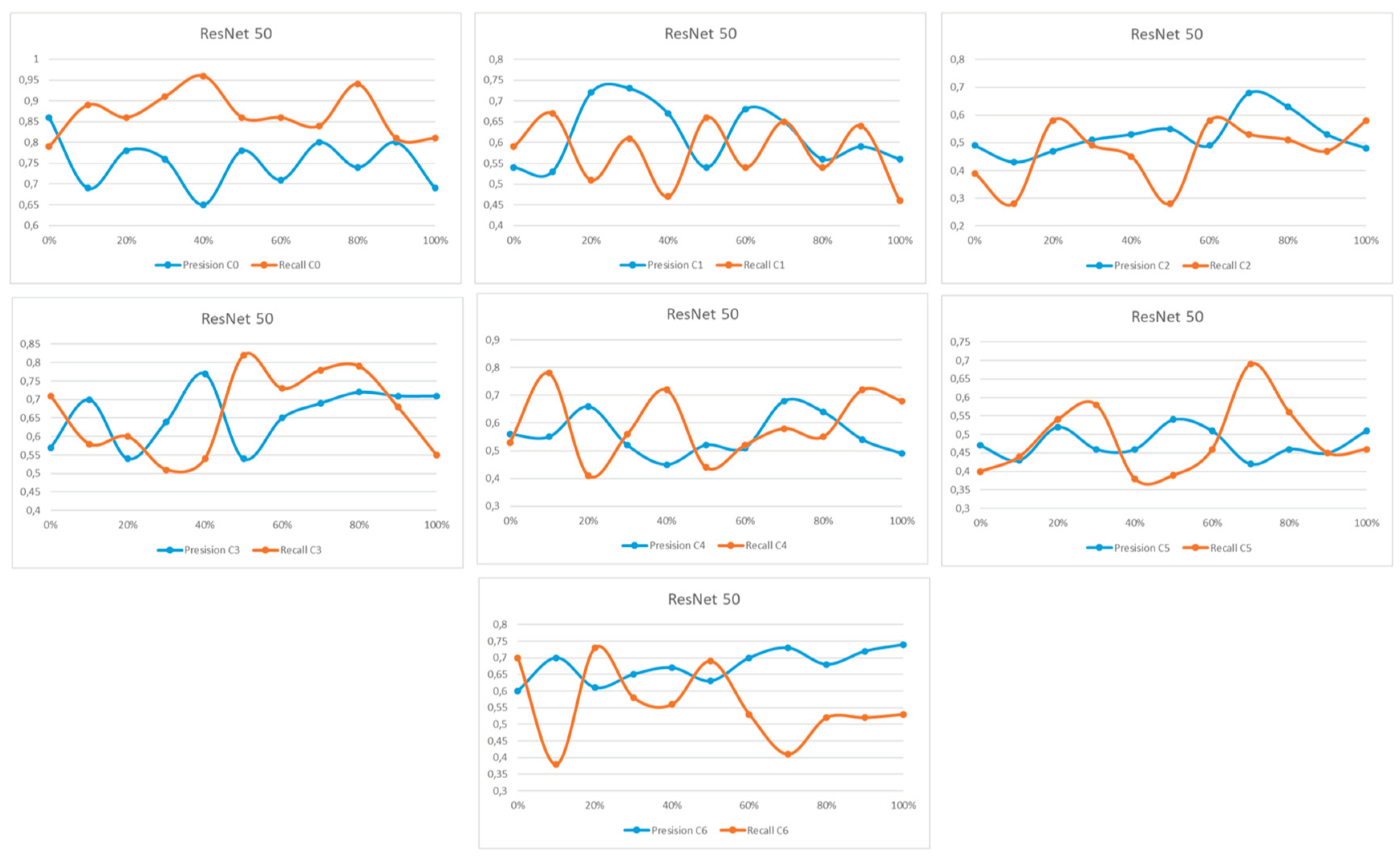
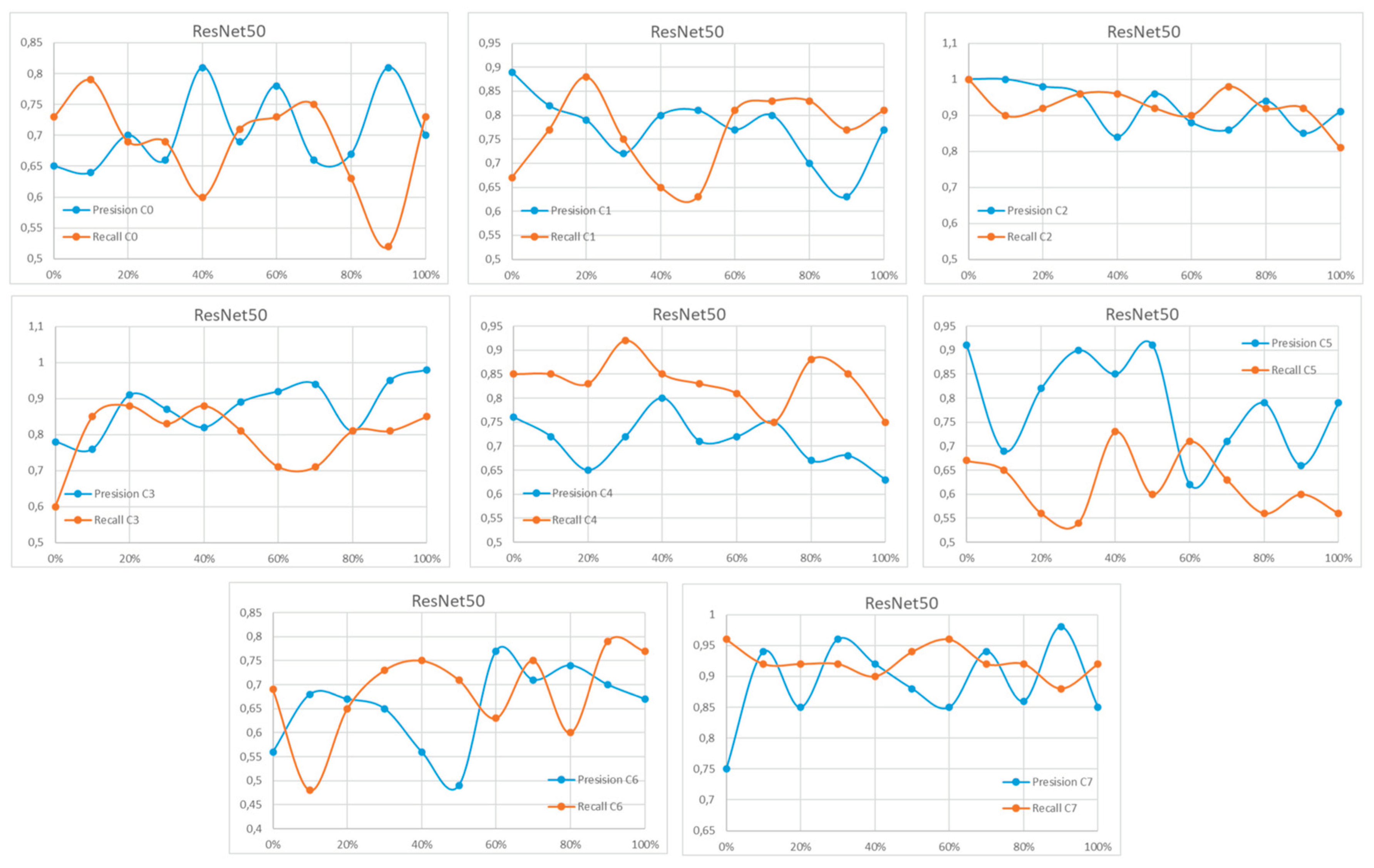
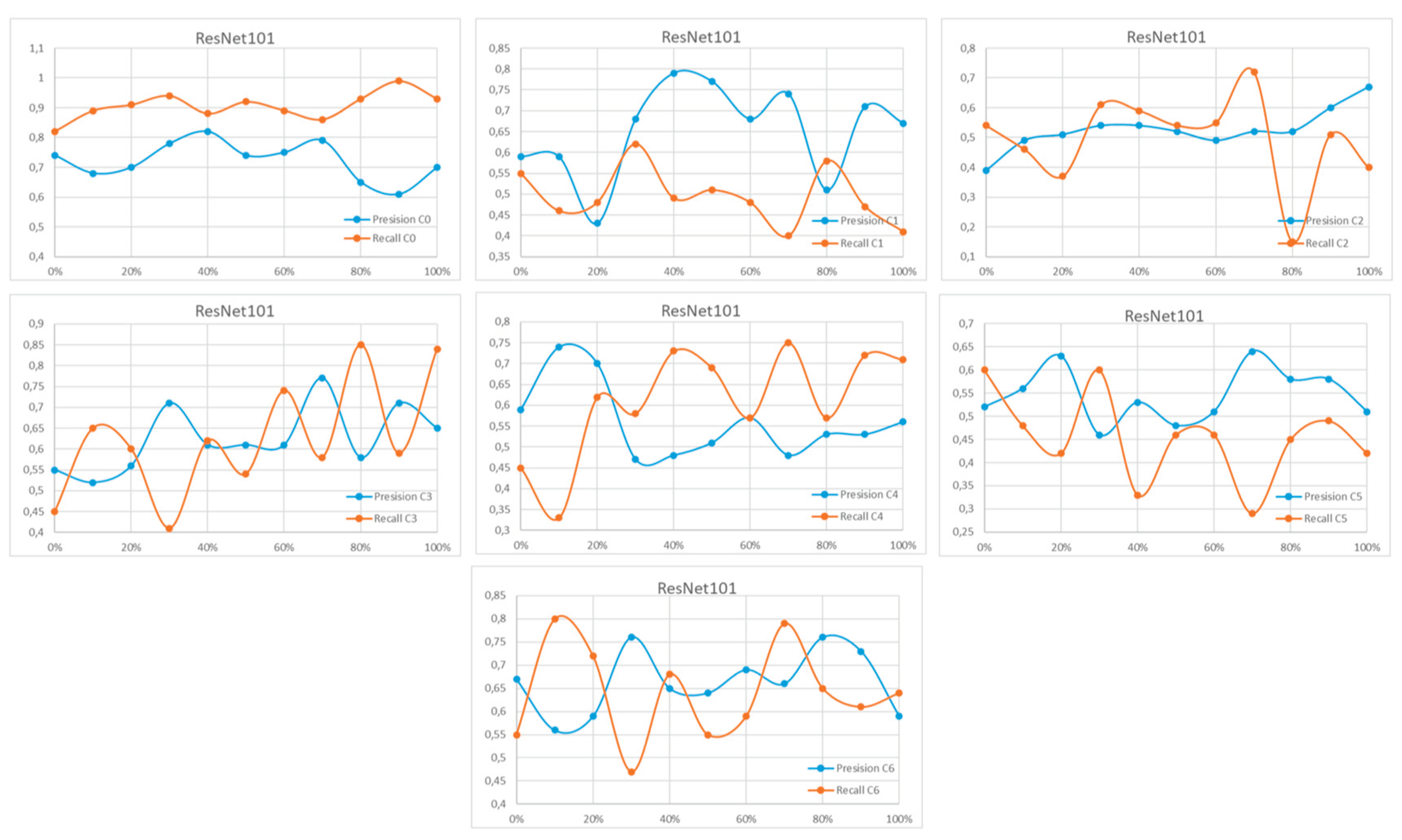
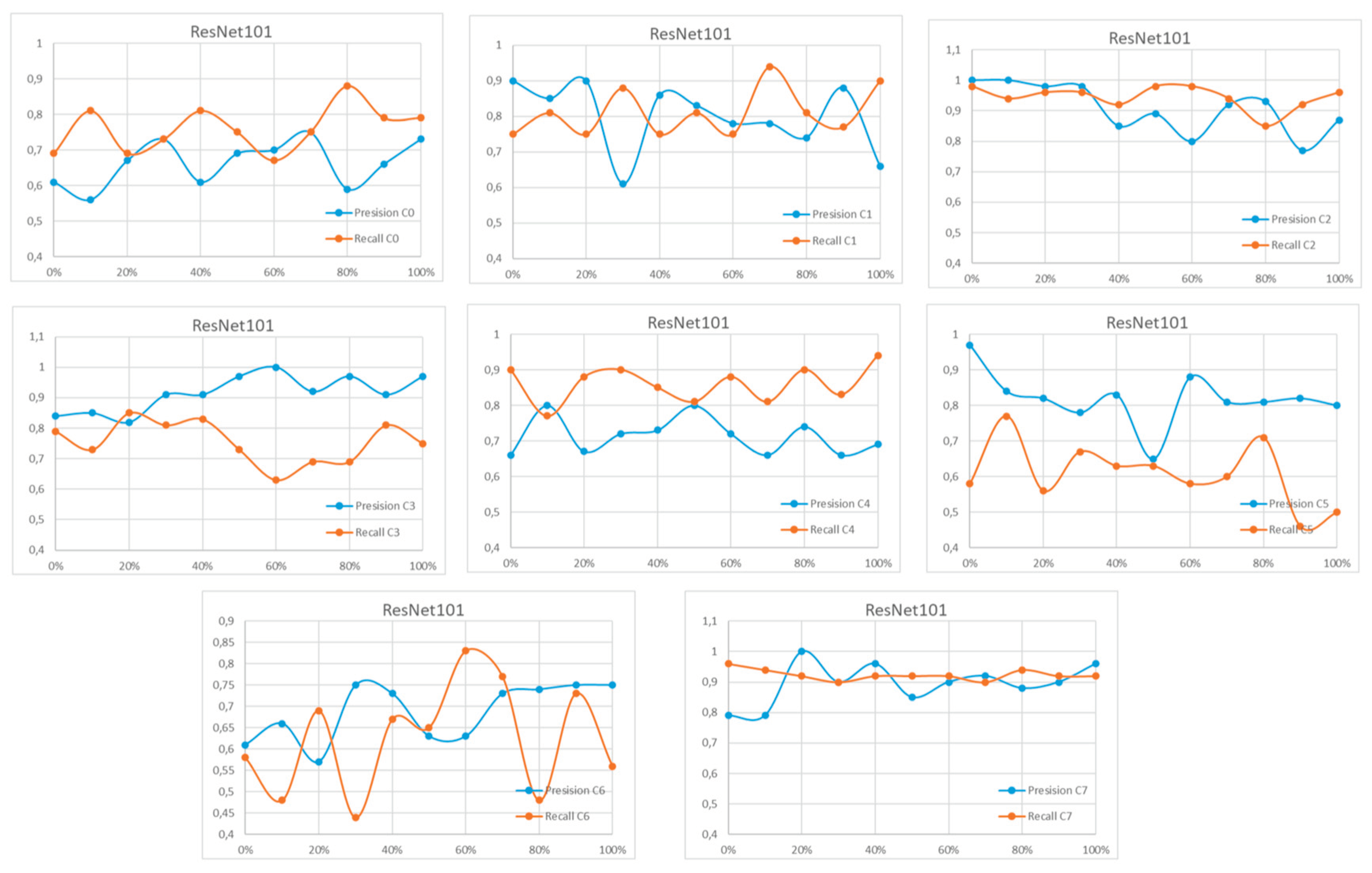
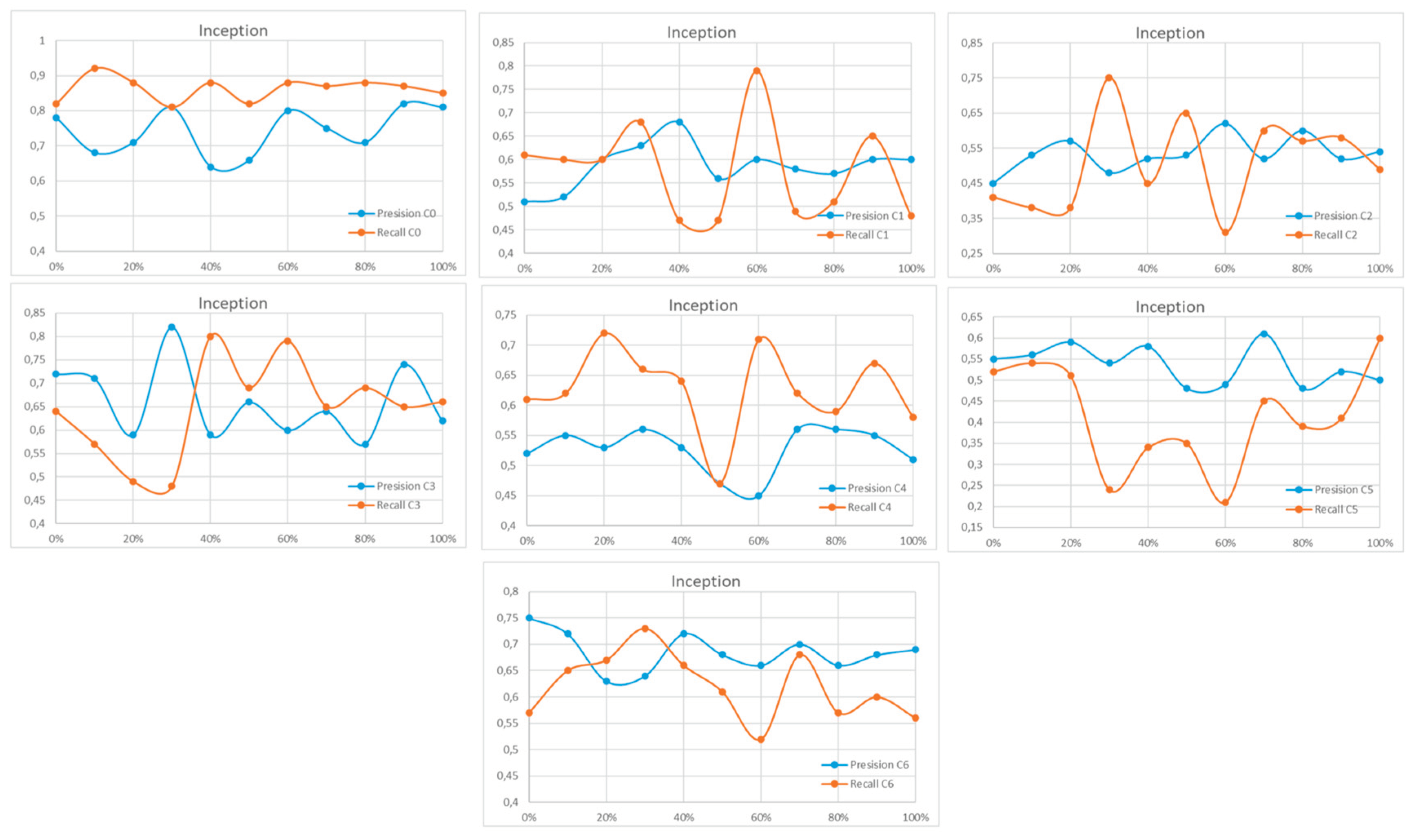
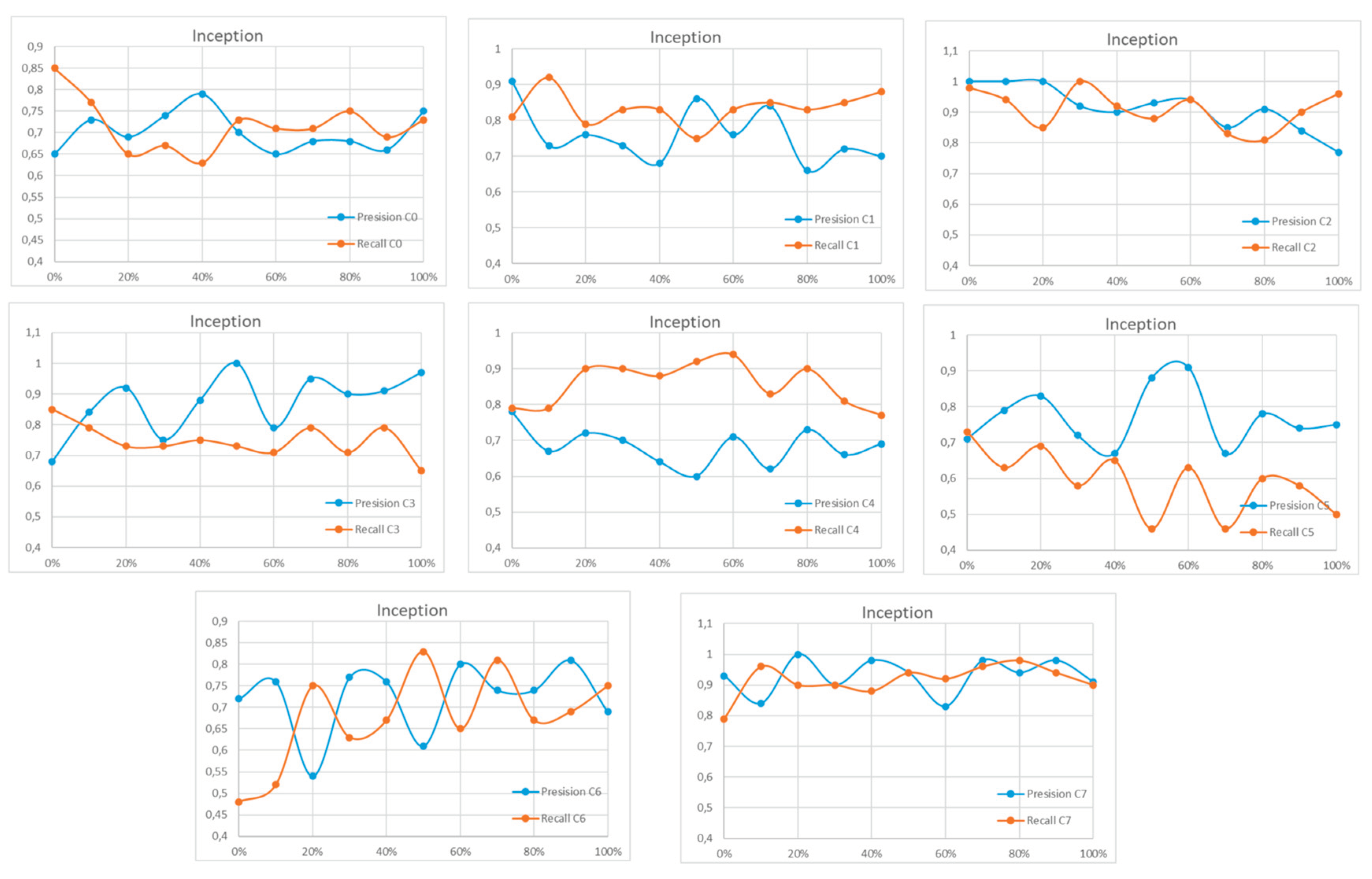
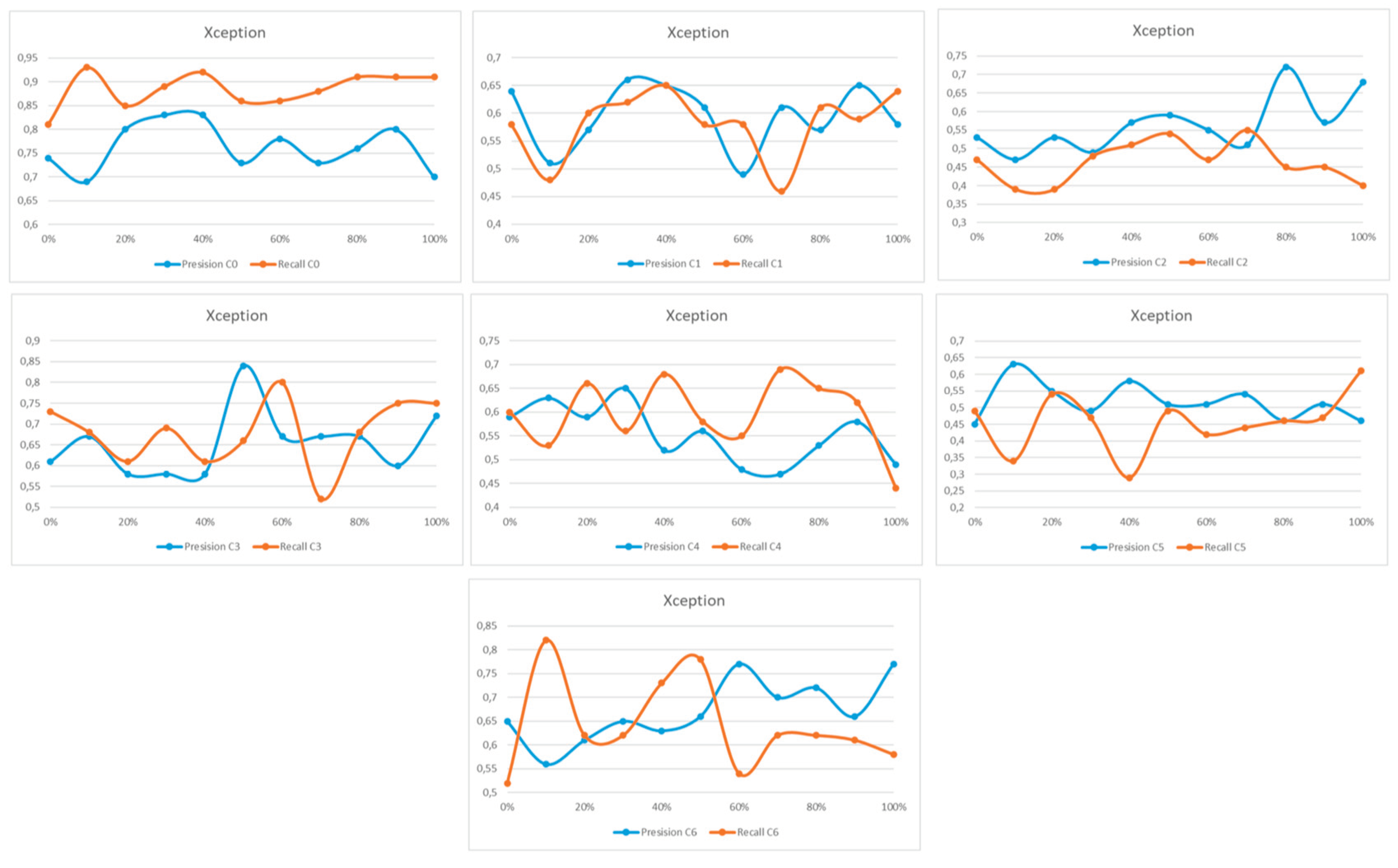
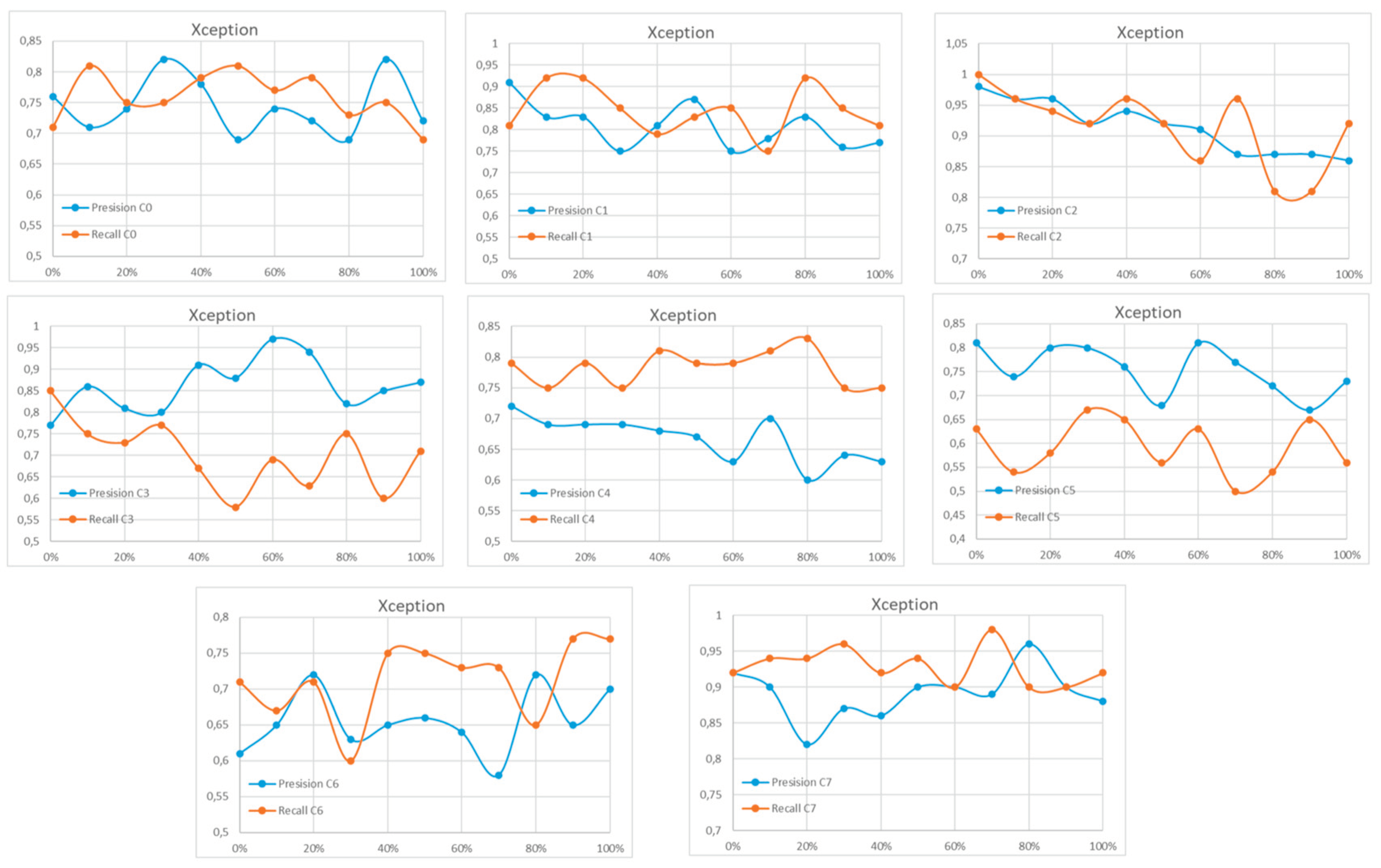
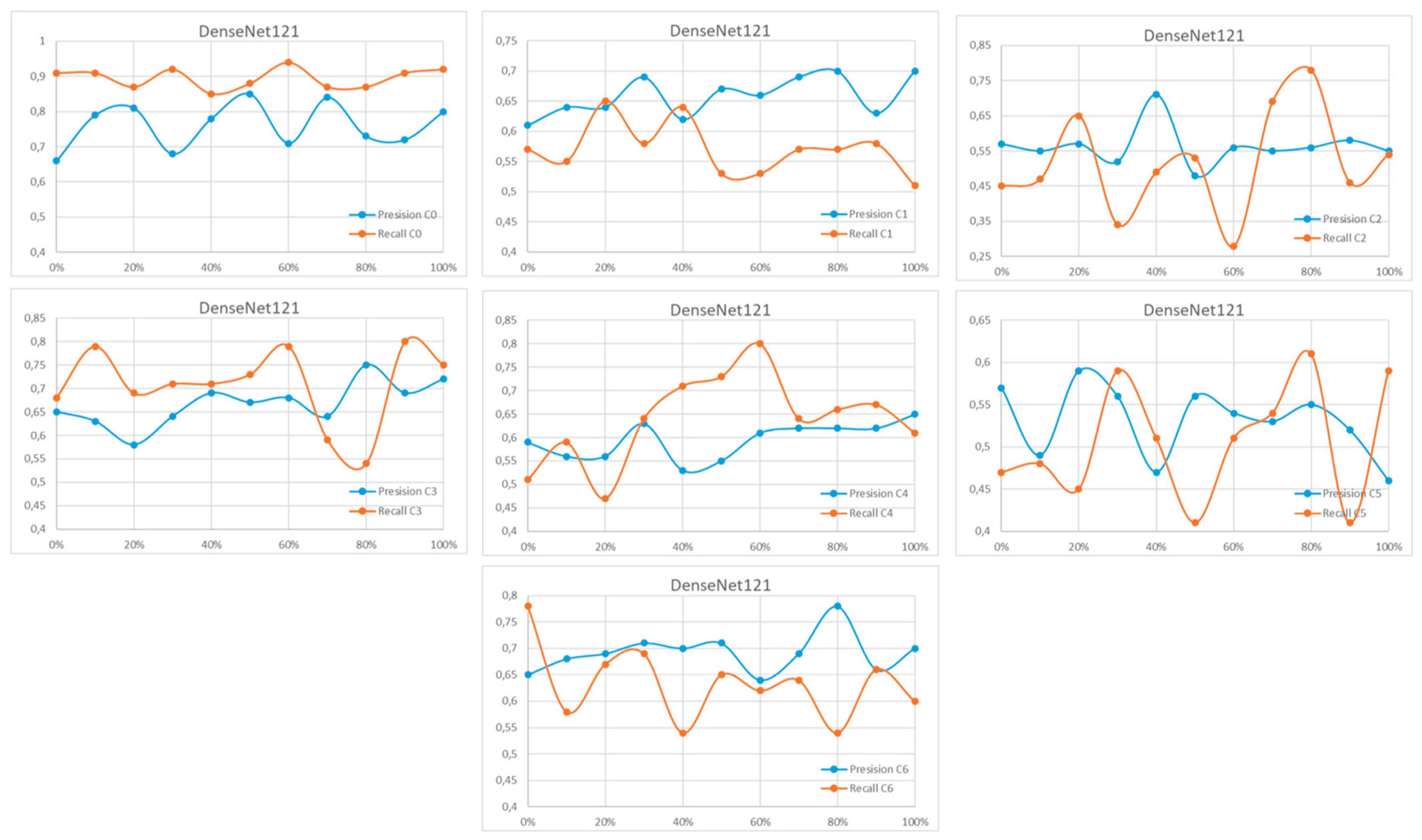
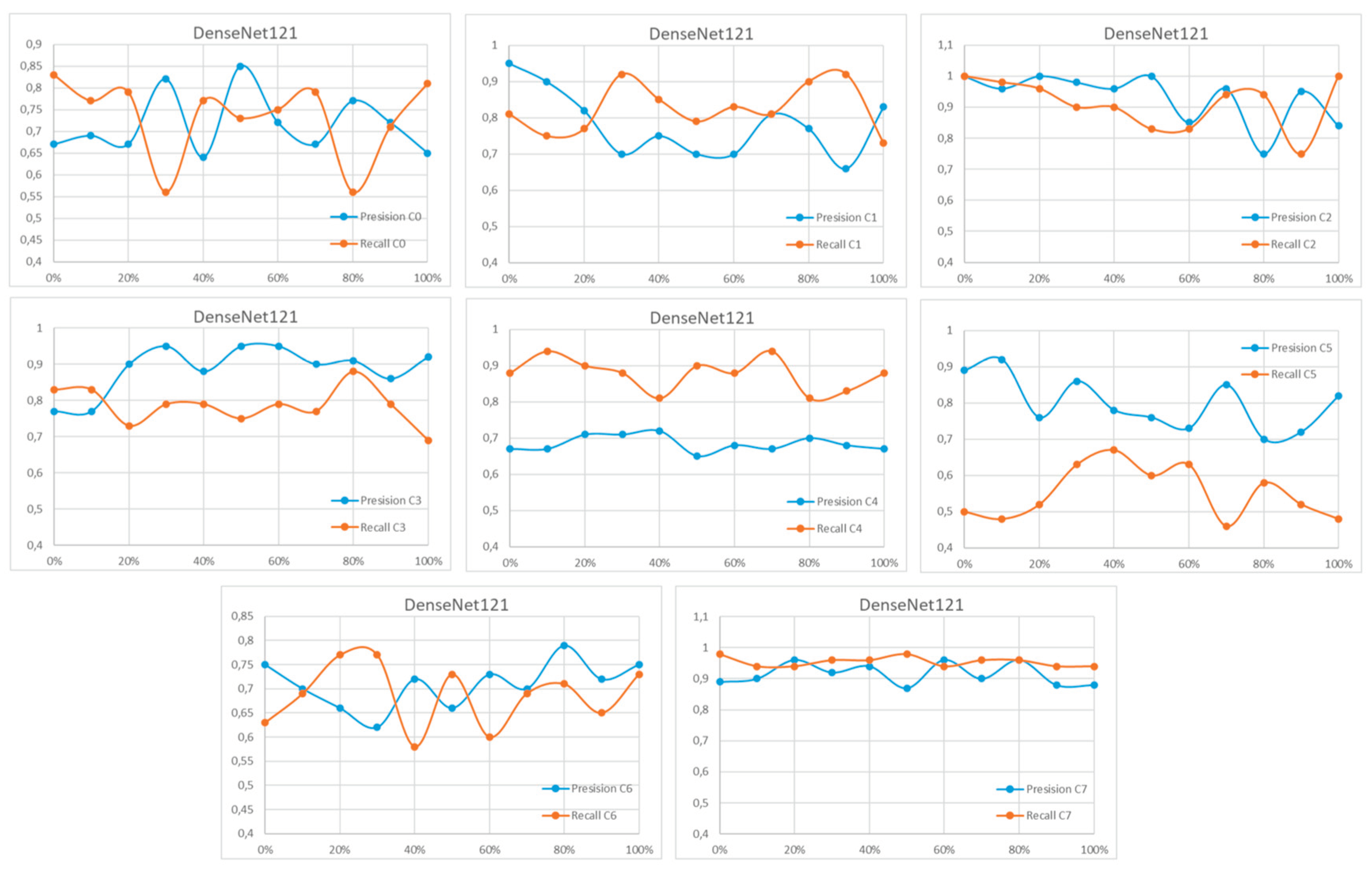
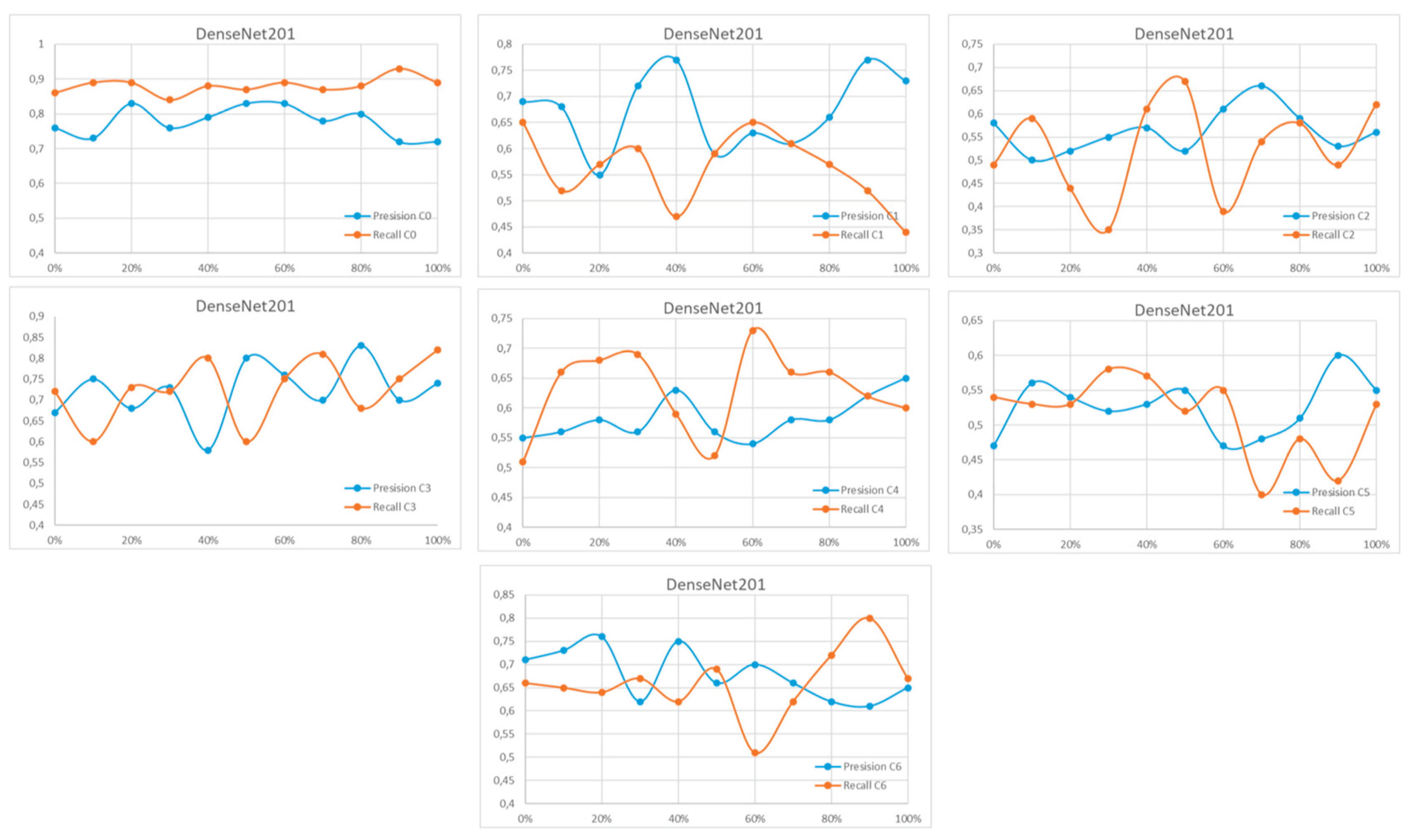
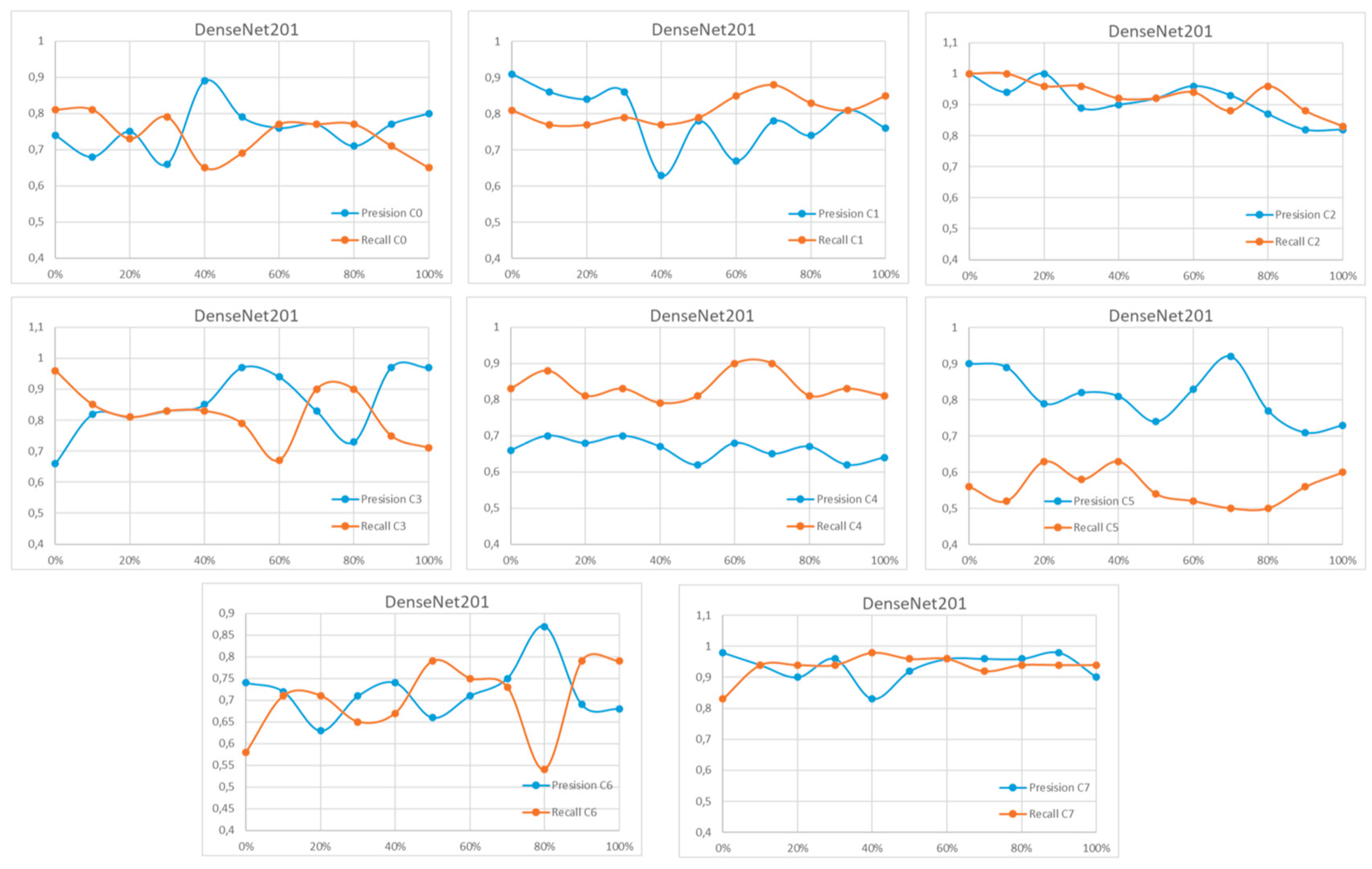
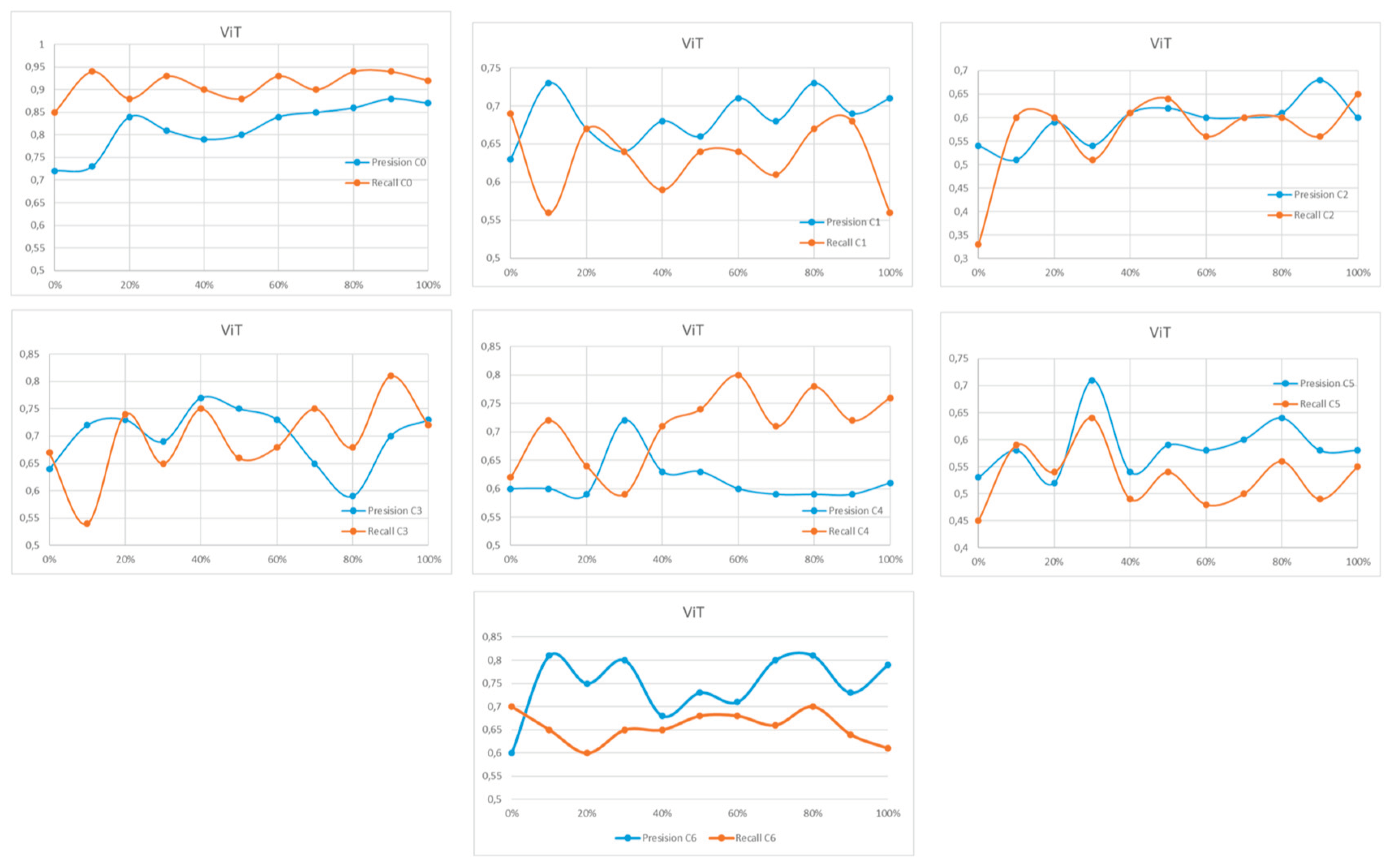
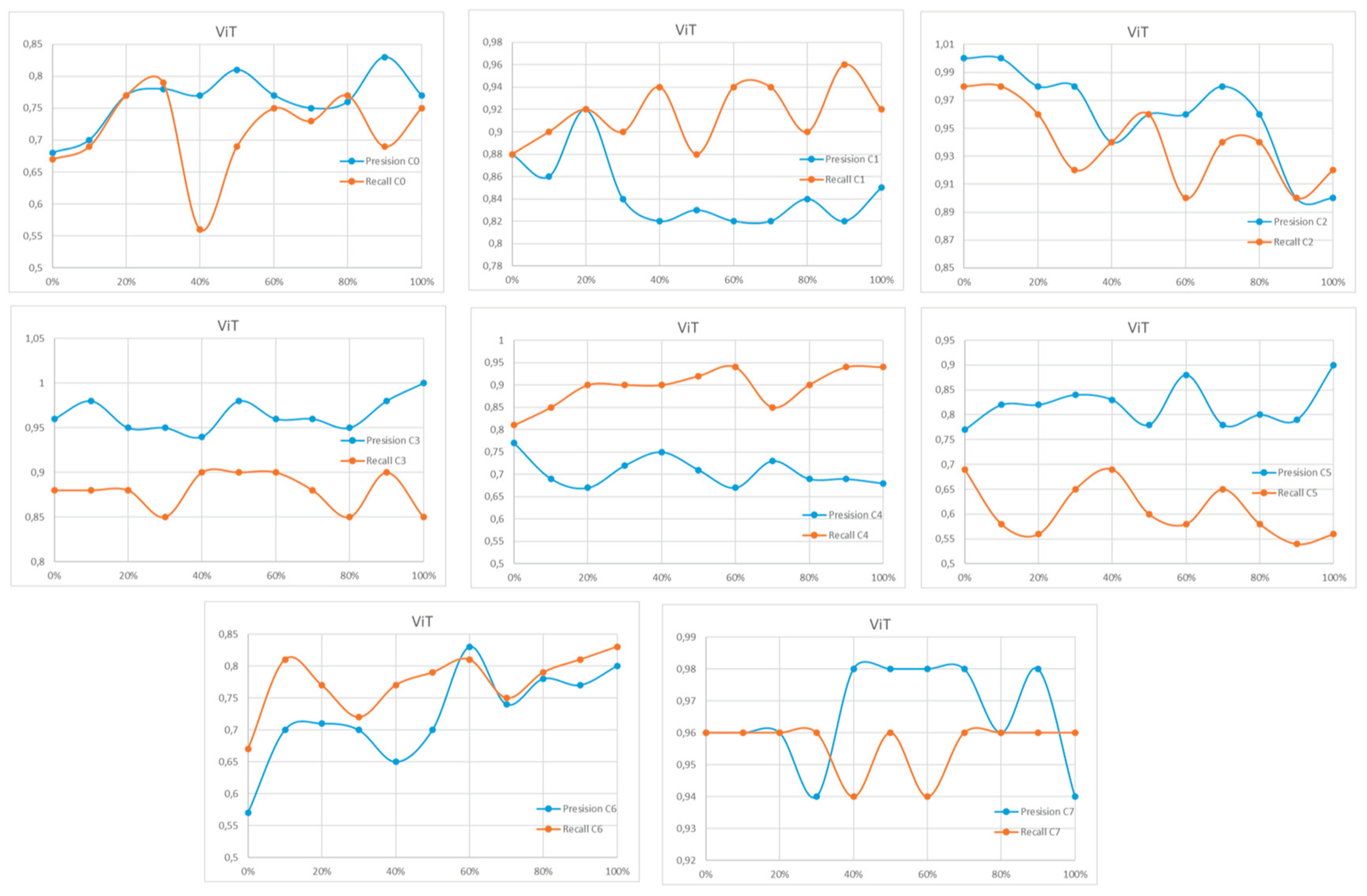
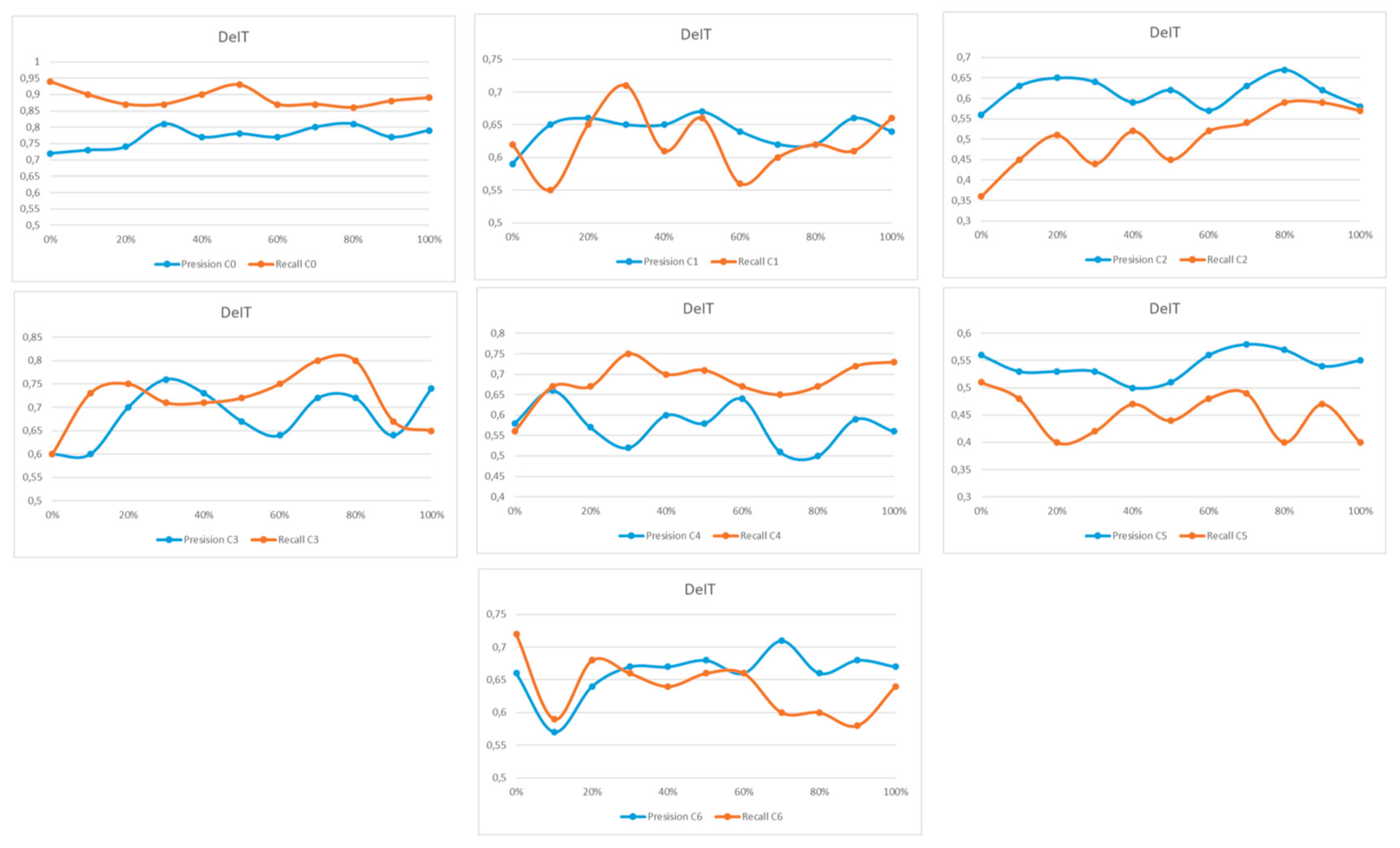
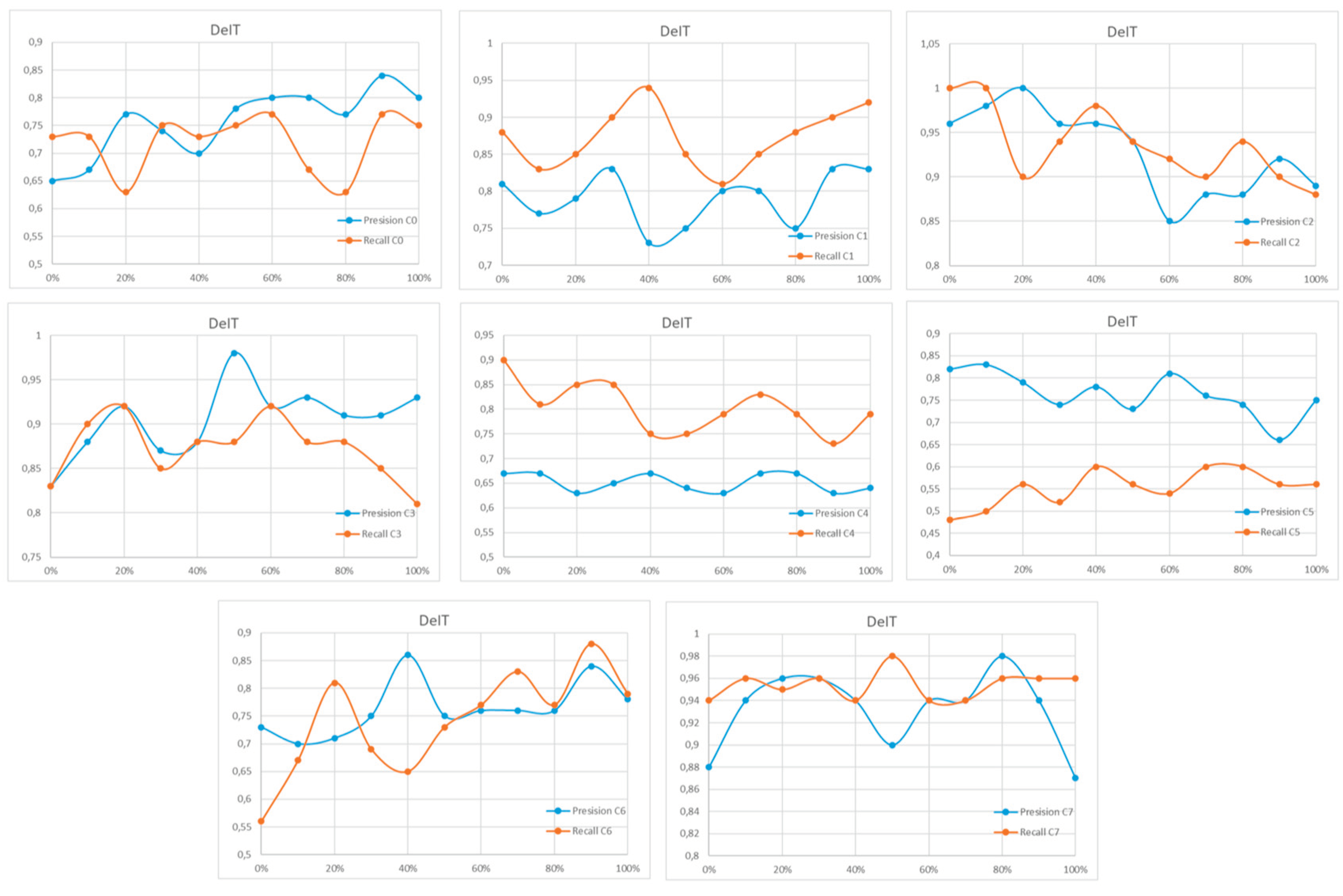
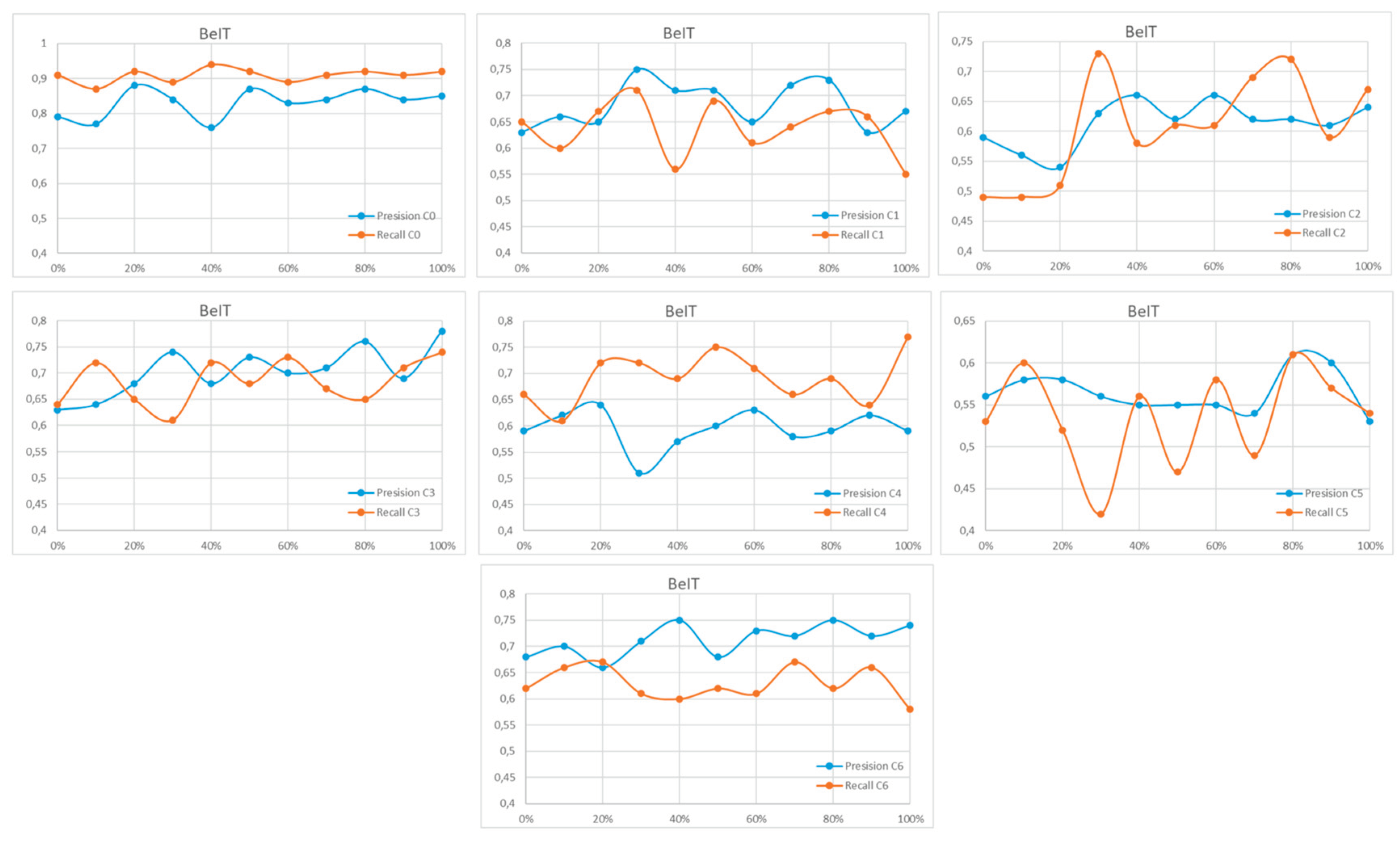
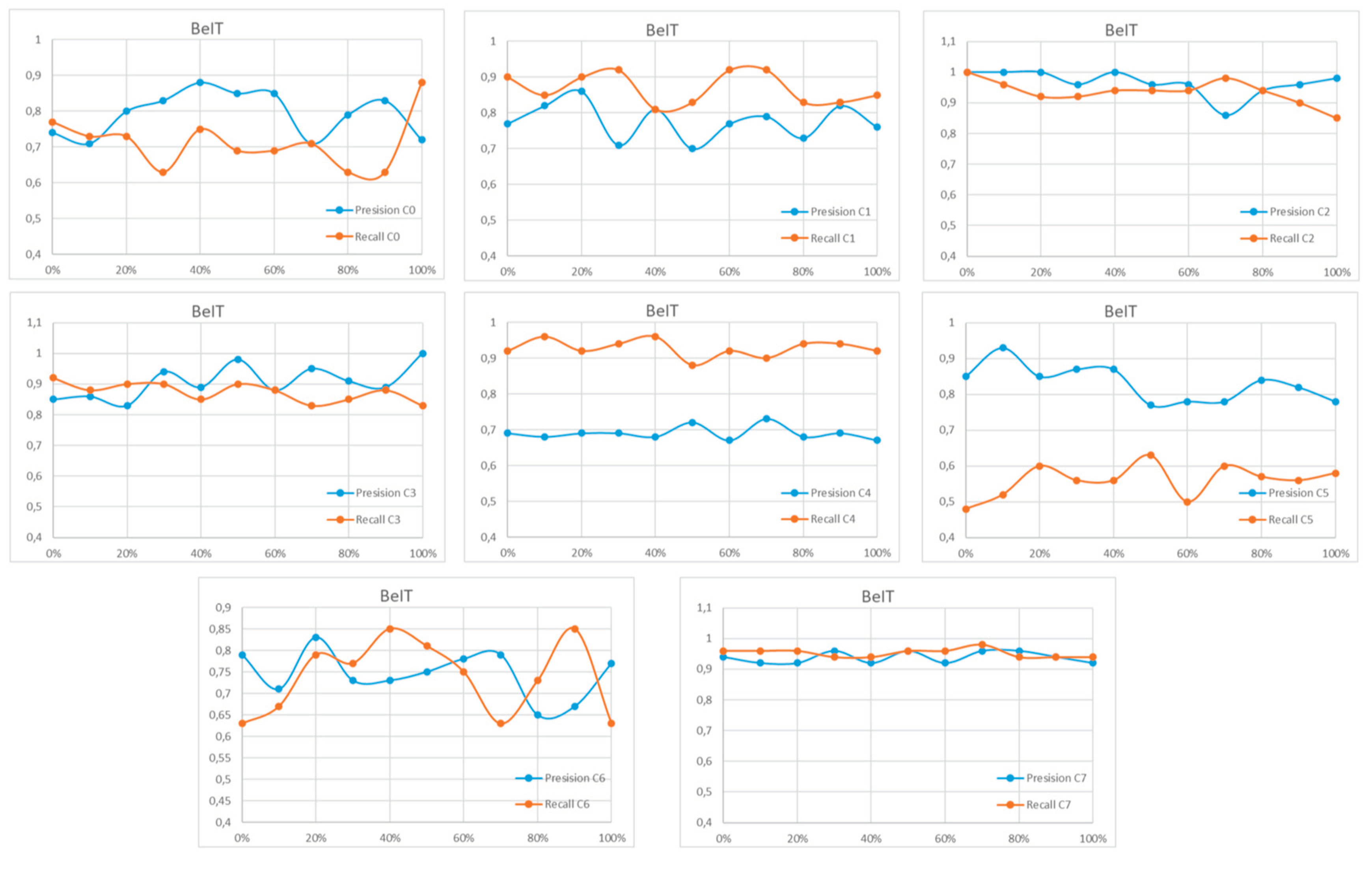
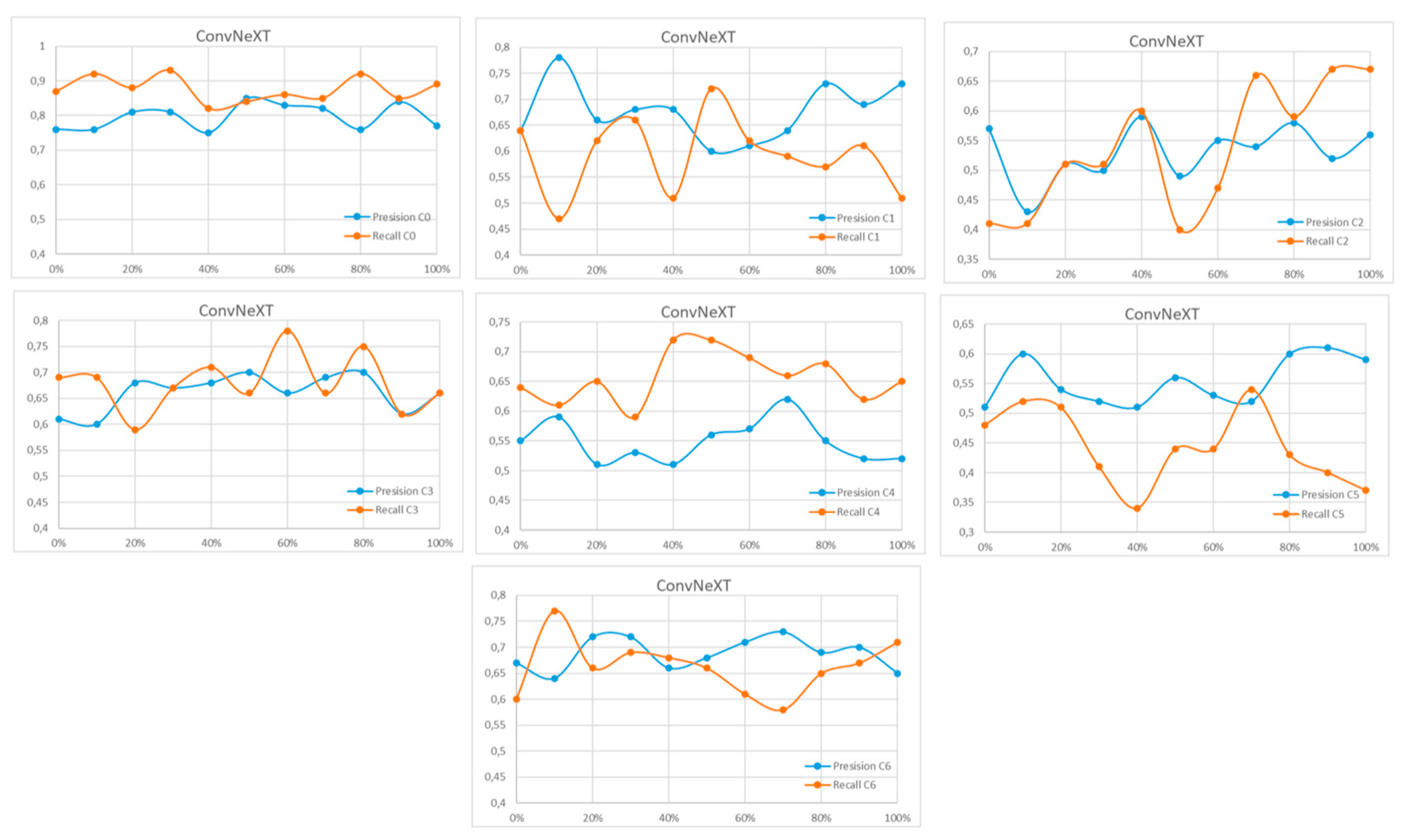
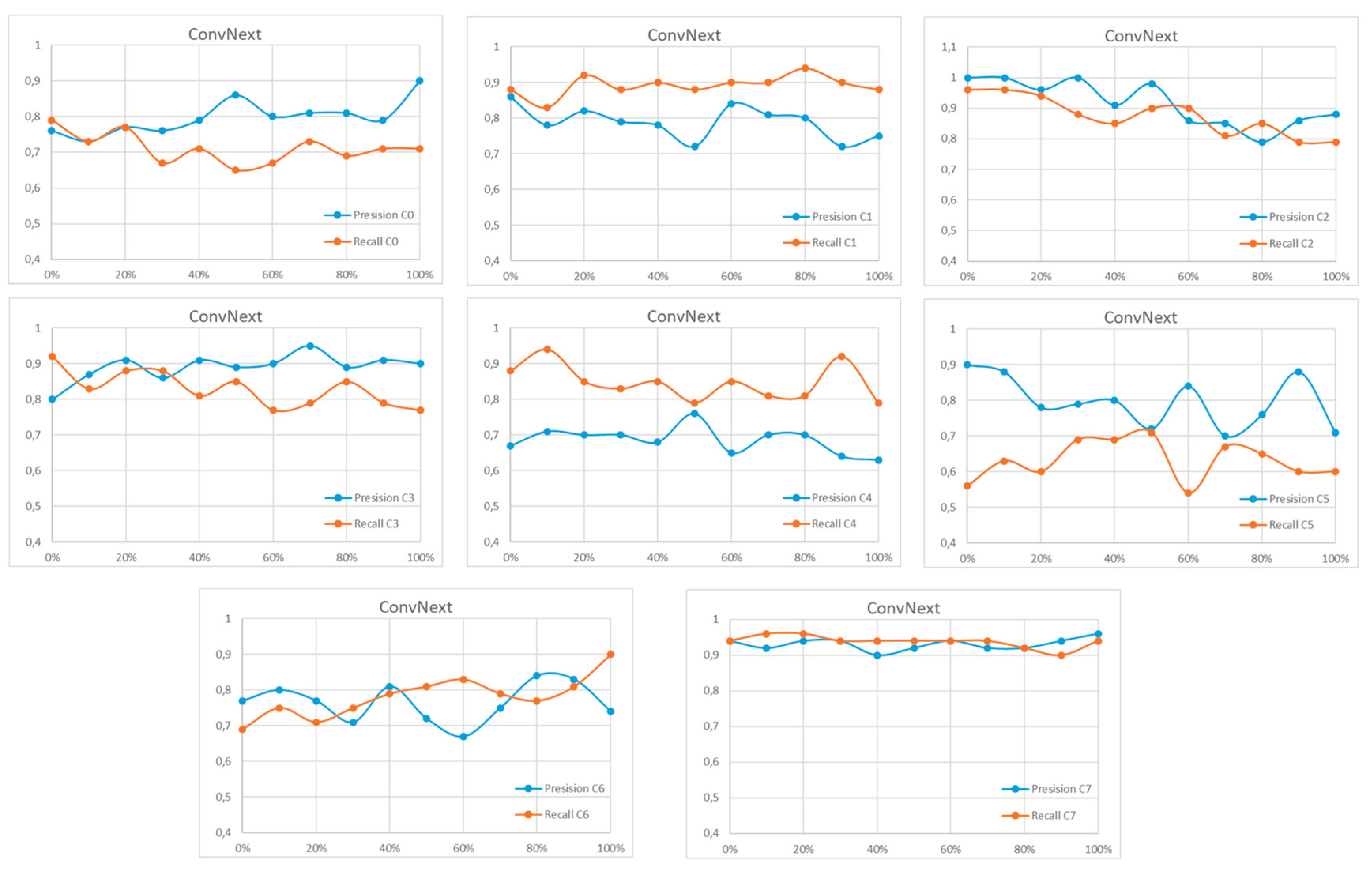
References
- Yang, S.; Xiao, W.; Zhang, M.; Guo, S.; Zhao, J.; Shen, F. Image Data Augmentation for Deep Learning: A Survey. arXiv 2022, arXiv:2204.08610. [Google Scholar]
- Alomar, K.; Aysel, H.I.; Cai, X. Data Augmentation in Classification and Segmentation: A Survey and New Strategies. J. Imaging 2023, 9, 46. [Google Scholar] [CrossRef] [PubMed]
- Shorten, C.; Khoshgoftaar, T.M. A survey on image data augmentation for deep learning. J. Big Data 2019, 6, 60. [Google Scholar] [CrossRef]
- Min, F.; Yu, T.; Zhang, C.; Xiao, Y.; Zhang, Z.; Chen, Y.; Xu, C.; Cai, J.; Chen, X.; Li, Z.; et al. A review of medical image data augmentation techniques for deep learning in healthcare. J. Med. Radiat. Sci. 2022, 69, 185–197. [Google Scholar]
- Antoniou, D.; Storkey, A.; Edwards, H. BAGAN: Data Augmentation with Balancing GAN. arXiv 2018, arXiv:1803.09655. [Google Scholar] [CrossRef]
- Wei, J.; Zou, C.; Coulon, J.A.; Klawonn, F.; Hammer, P.L. Text data augmentation for deep learning. J. Big Data 2021, 8, 134. [Google Scholar] [CrossRef] [PubMed]
- Alyasin, E.I.; Ata, O.; Mohammedqasim, H.; Mohammedqasem, R. Optimizing Prediction of Cardiac Conditions Using Hyperparameter Optimization and Ensemble Learning. Cogn. Comput. 2023, 15, 1–15. [Google Scholar]
- Barulina, M.; Sanbaev, A.; Okunkov, S. Deep Learning Approaches to Automatic Chronic Venous Disease Severity Assessment from Photographs. Mathematics 2022, 10, 3571. [Google Scholar] [CrossRef]
- Kaggle Skin Dataset. Available online: https://www.kaggle.com/datasets/ahmedxc4/skin-ds (accessed on 29 March 2026).
- He, K.; Zhang, X.; Ren, S.; Sun, J. Deep Residual Learning for Image Recognition. In Proceedings of the IEEE Conference on Computer Vision and Pattern Recognition (CVPR), Las Vegas, NV, USA, 27–30 June 2016. [Google Scholar]
- Dosovitskiy, A.; Beyer, L.; Kolesnikov, A.; Weissenborn, D.; Zhai, X.; Unterthiner, T.; Dehghani, M.; Minderer, M.; Heigold, G.; Gelly, S.; et al. An Image is Worth 16x16 Words: Transformers for Image Recognition at Scale. arXiv 2021, arXiv:2010.11929. [Google Scholar] [CrossRef]
- Huang, G.; Liu, Z.; van der Maaten, L.; Weinberger, K.Q. Densely Connected Convolutional Networks. In Proceedings of the IEEE Conference on Computer Vision and Pattern Recognition (CVPR), Honolulu, HI, USA, 21–26 July 2017. [Google Scholar]
- Szegedy, C.; Vanhoucke, V.; Ioffe, S.; Shlens, J.; Wojna, Z. Going Deeper With Convolutions. In Proceedings of the IEEE Conference on Computer Vision and Pattern Recognition (CVPR), Boston, MA, USA, 7–12 June 2015. [Google Scholar]
- Chollet, F. Xception: Deep Learning with Depthwise Separable Convolutions. In Proceedings of the IEEE Conference on Computer Vision and Pattern Recognition (CVPR), Honolulu, HI, USA, 21–26 July 2017. [Google Scholar]
- Liu, Z.; Mao, H.; Wu, C.-Y.; Feichtenhofer, C.; Darrell, T.; Xie, S. A ConvNet for the 2020s. In Proceedings of the IEEE/CVF Conference on Computer Vision and Pattern Recognition (CVPR), New Orleans, LA, USA, 18–24 June 2022. [Google Scholar]
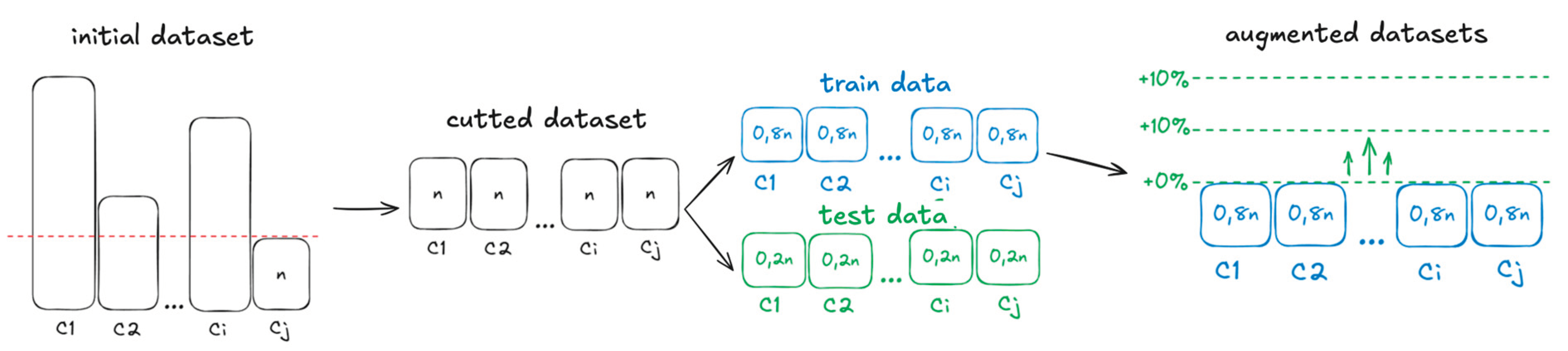
| Num of params | Image size | Batchsize | Num of hidden layers |
Optimizer | Learning rate | |
|---|---|---|---|---|---|---|
| ResNet34 | 21.7M | 224 | 64 | 34 | Adam | 1∙10-4 |
| ResNet50 | 25.5M | 224 | 64 | 50 | Adam | 1∙10-4 |
| ResNet101 | 44.5M | 224 | 64 | 101 | Adam | 1∙10-4 |
| VGG19 | 143.6M | 224 | 16 | 19 | Adam | 1∙10-4 |
| DenseNet161 | 28.7M | 224 | 8 | 161 | Adam | 1∙10-4 |
| DenseNet201 | 20M | 224 | 8 | 201 | Adam | 1∙10-4 |
| Inception_v3 | 27.2M | 224 | 16 | 48 | Adam | 1∙10-4 |
| Xception | 23M | 224 | 8 | 71 | Adam | 1∙10-4 |
| VIT | 86.4M | 224 | 32 | 12 | AdamW | 5∙10-5 |
| DeIT | 86.4M | 224 | 32 | 12 | AdamW | 5∙10-5 |
| BeIT | 86.9M | 384 | 2 | 12 | AdamW | 5∙10-5 |
| ConvNeXT | 28M | 224 | 8 | 36 | AdamW | 5∙10-5 |
| %, synthetic data | VGG19 | ResNet34 | ResNet50 | ResNet101 | Xception | Inception | DenseNet121 | DenseNet201 | VIT | DeIT |
BeIT |
ConvNeXT |
|---|---|---|---|---|---|---|---|---|---|---|---|---|
| 0 | 0,47 | 0,57 | 0,58 | 0,57 | 0,6 | 0,6 | 0,62 | 0,63 | 0,62 | 0,62 | 0,64 | 0,62 |
| 10 | 0,5 | 0,58 | 0,58 | 0,58 | 0,6 | 0,61 | 0,62 | 0,63 | 0,66 | 0,63 | 0,65 | 0,63 |
| 20 | 0,56 | 0,58 | 0,6 | 0,59 | 0,61 | 0,61 | 0,64 | 0,64 | 0,67 | 0,65 | 0,66 | 0,63 |
| 30 | 0,56 | 0,58 | 0,61 | 0,61 | 0,62 | 0,62 | 0,64 | 0,64 | 0,67 | 0,65 | 0,67 | 0,64 |
| 40 | 0,56 | 0,58 | 0,58 | 0,62 | 0,63 | 0,61 | 0,63 | 0,65 | 0,67 | 0,65 | 0,67 | 0,64 |
| 50 | 0,55 | 0,59 | 0,59 | 0,6 | 0,64 | 0,58 | 0,64 | 0,64 | 0,68 | 0,65 | 0,68 | 0,64 |
| 60 | 0,56 | 0,6 | 0,6 | 0,61 | 0,6 | 0,6 | 0,64 | 0,64 | 0,68 | 0,66 | 0,68 | 0,64 |
| 70 | 0,6 | 0,6 | 0,64 | 0,63 | 0,6 | 0,62 | 0,65 | 0,65 | 0,68 | 0,65 | 0,68 | 0,65 |
| 80 | 0,58 | 0,61 | 0,63 | 0,6 | 0,63 | 0,6 | 0,65 | 0,65 | 0,7 | 0,65 | 0,7 | 0,66 |
| 90 | 0,55 | 0,58 | 0,61 | 0,63 | 0,63 | 0,63 | 0,64 | 0,65 | 0,69 | 0,65 | 0,68 | 0,64 |
| 100 | 0,56 | 0,58 | 0,58 | 0,62 | 0,62 | 0,6 | 0,65 | 0,65 | 0,69 | 0,65 | 0,68 | 0,64 |
| %, synthetic data | VGG19 | ResNet34 | ResNet50 | ResNet101 | Xception | Inception | DenseNet121 | DenseNet201 | VIT | DeIT |
BeIT |
ConvNeXT |
| 0 | 0,71 | 0,76 | 0,77 | 0,78 | 0,8 | 0,79 | 0,8 | 0,8 | 0,82 | 0,79 | 0,82 | 0,82 |
| 10 | 0,72 | 0,78 | 0,78 | 0,78 | 0,8 | 0,79 | 0,81 | 0,81 | 0,83 | 0,8 | 0,82 | 0,82 |
| 20 | 0,73 | 0,79 | 0,79 | 0,79 | 0,79 | 0,78 | 0,8 | 0,8 | 0,84 | 0,81 | 0,84 | 0,83 |
| 30 | 0,73 | 0,79 | 0,79 | 0,79 | 0,78 | 0,78 | 0,8 | 0,8 | 0,84 | 0,81 | 0,83 | 0,82 |
| 40 | 0,73 | 0,79 | 0,79 | 0,8 | 0,79 | 0,77 | 0,79 | 0,78 | 0,84 | 0,81 | 0,83 | 0,82 |
| 50 | 0,71 | 0,78 | 0,77 | 0,78 | 0,77 | 0,78 | 0,79 | 0,79 | 0,84 | 0,81 | 0,83 | 0,82 |
| 60 | 0,7 | 0,78 | 0,78 | 0,78 | 0,78 | 0,79 | 0,78 | 0,79 | 0,84 | 0,81 | 0,82 | 0,81 |
| 70 | 0,7 | 0,78 | 0,79 | 0,8 | 0,77 | 0,78 | 0,8 | 0,81 | 0,84 | 0,81 | 0,82 | 0,81 |
| 80 | 0,7 | 0,77 | 0,77 | 0,78 | 0,77 | 0,78 | 0,79 | 0,78 | 0,84 | 0,81 | 0,82 | 0,81 |
| 90 | 0,69 | 0,76 | 0,77 | 0,78 | 0,76 | 0,78 | 0,76 | 0,78 | 0,84 | 0,82 | 0,82 | 0,8 |
| 100 | 0,72 | 0,77 | 0,78 | 0,79 | 0,77 | 0,77 | 0,78 | 0,77 | 0,84 | 0,81 | 0,81 | 0,8 |
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2026 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).




