Submitted:
01 April 2026
Posted:
03 April 2026
You are already at the latest version
Abstract
Keywords:
Introduction
Methods
eQTL Data
pQTL Data
Mendelian Randomization
- -
- SNPs must be significantly associated with the transcript level (P<5e-8)
- -
- SNPs must be in linkage disequilibrium (r2<0.05)
- -
- SNPs must have pQTL summary association statistics
Genetic Correlation
Colocalization
Results
Mendelian Randomization
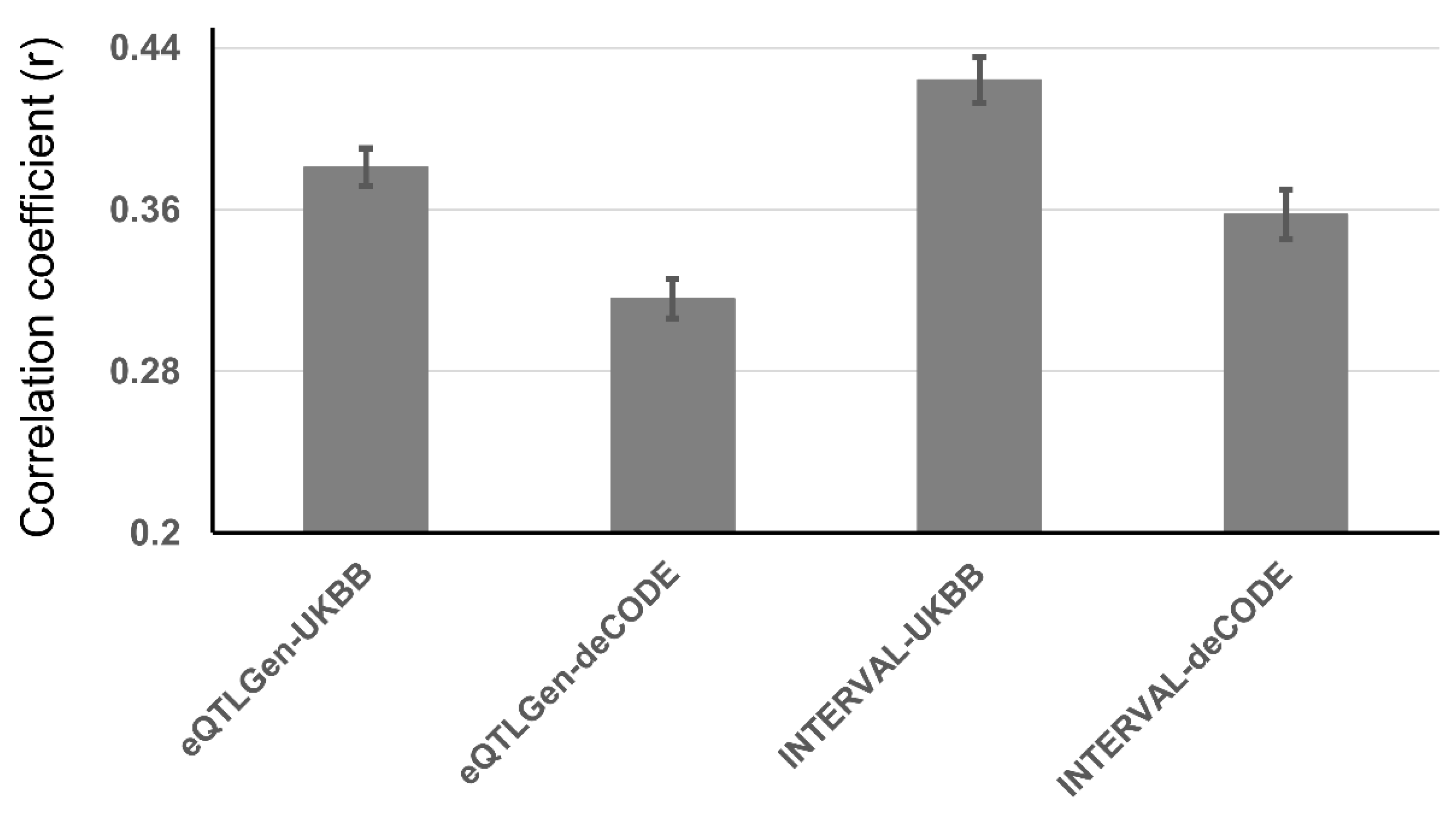
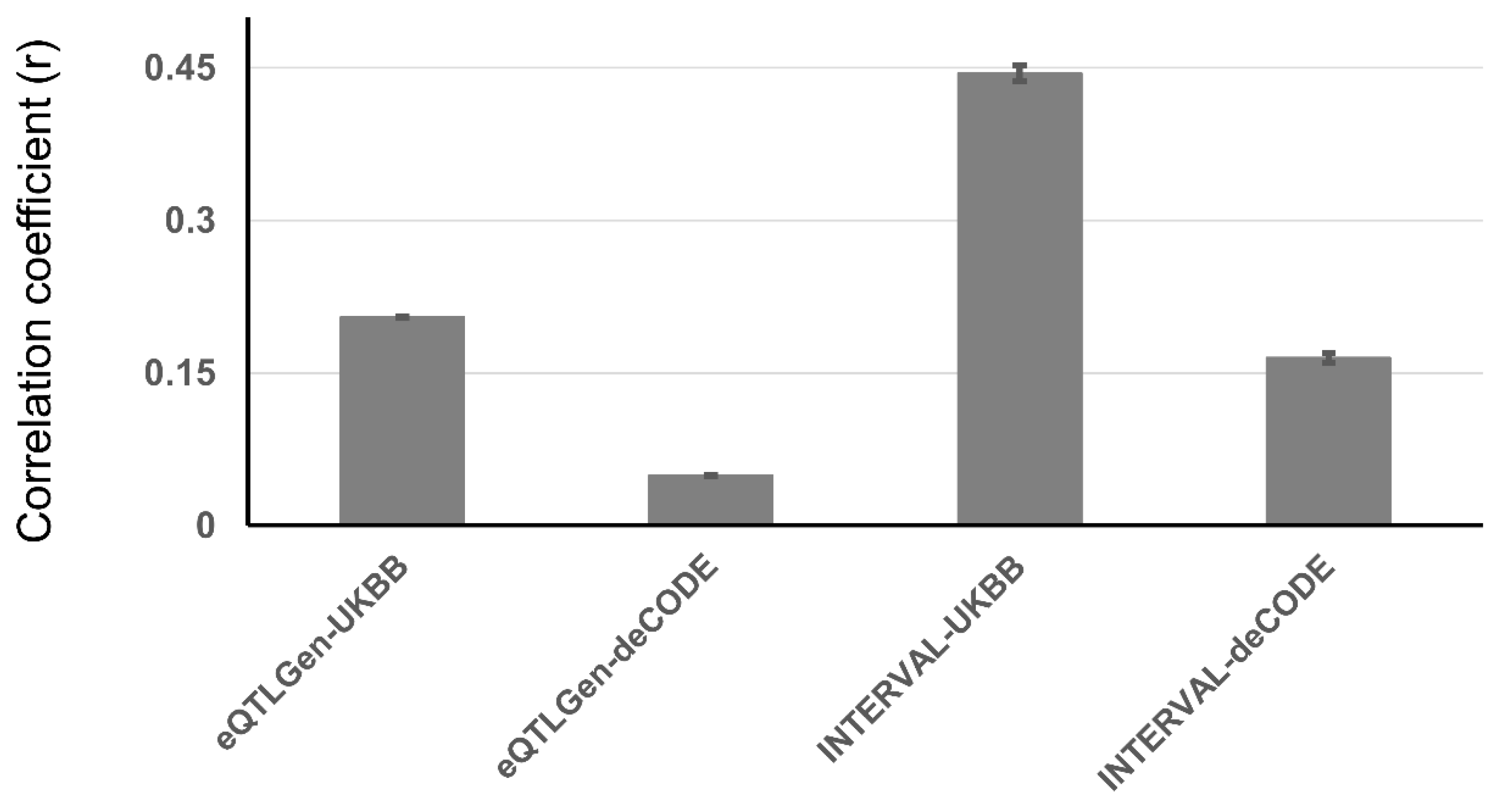
Genetic Correlation
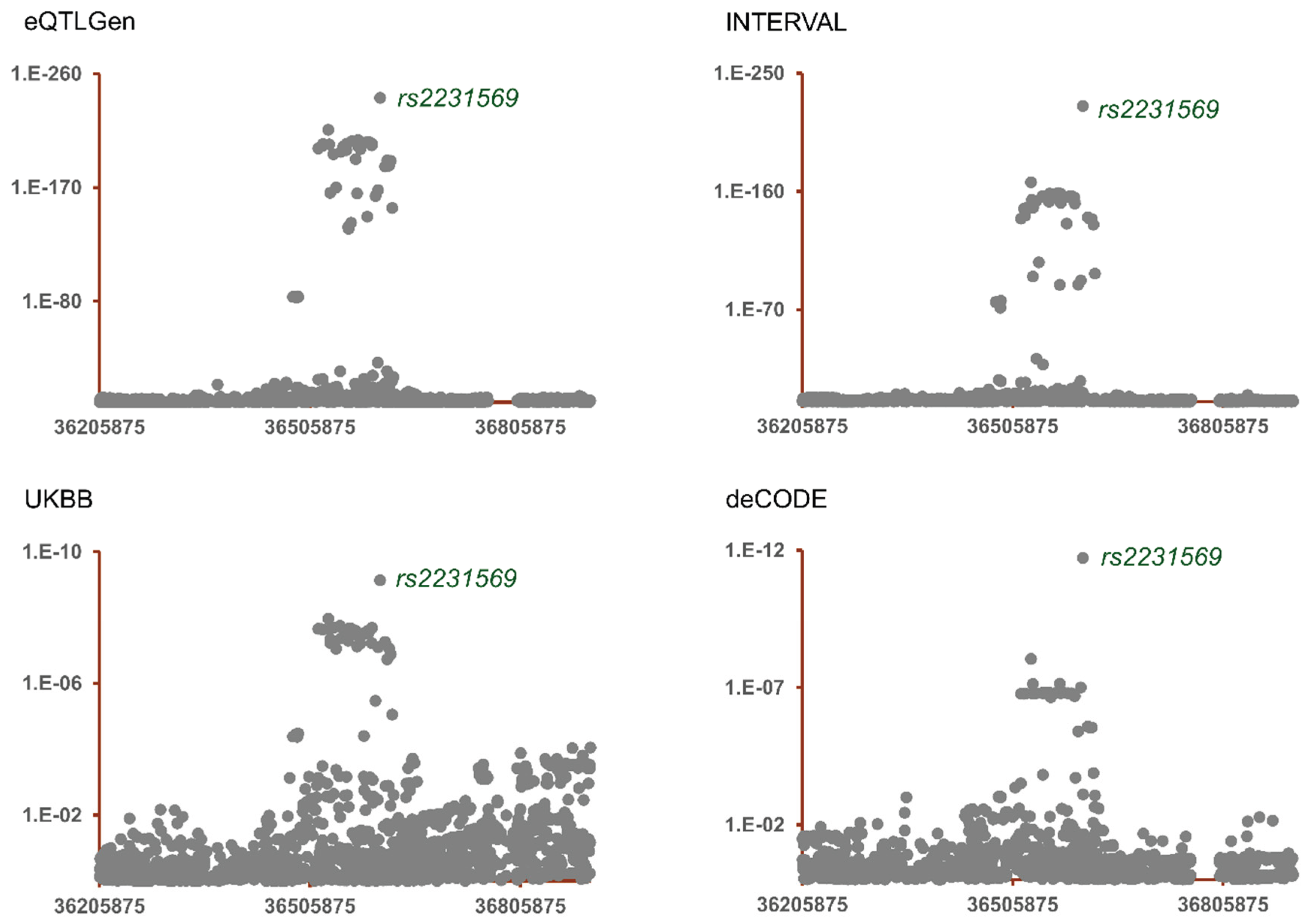
Colocalization
Discussion
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Acknowledgments
Competing interests
References
- Zhu, Z.; Zheng, Z.; Zhang, F.; Wu, Y.; Trzaskowski, M.; Maier, R.; Robinson, M.R.; McGrath, J.J.; Visscher, P.M.; Wray, N.R.; et al. Causal associations between risk factors and common diseases inferred from GWAS summary data. Nat. Commun. 2018, 9, 224. [Google Scholar] [CrossRef] [PubMed]
- Pasaniuc, B.; Price, A.L. Dissecting the genetics of complex traits using summary association statistics. Nat. Rev. Genet. 2017, 18, 117–127. [Google Scholar] [CrossRef] [PubMed]
- Boyle, E.A.; Li, Y.I.; Pritchard, J.K. An expanded view of complex traits: from polygenic to omnigenic. Cell 2017, 169, 1177–1186. [Google Scholar] [CrossRef] [PubMed]
- Price, A.L.; Helgason, A.; Thorleifsson, G.; McCarroll, S.A.; Kong, A.; Stefansson, K. Single-tissue and cross-tissue heritability of gene expression via identity-by-descent in related or unrelated individuals. PLoS Genet. 2011, 7, e1001317. [Google Scholar] [CrossRef] [PubMed]
- Jiang, D.; Cope, A.L.; Zhang, J.; Pennell, M. On the Decoupling of Evolutionary Changes in mRNA and Protein Levels. Mol. Biol. Evol. 2023, 40. [Google Scholar] [CrossRef] [PubMed]
- Liu, Y.; Beyer, A.; Aebersold, R. On the Dependency of Cellular Protein Levels on mRNA Abundance. Cell 2016, 165, 535–550. [Google Scholar] [CrossRef] [PubMed]
- Ponomarenko, E.A.; Krasnov, G.S.; Kiseleva, O.I.; Kryukova, P.A.; Arzumanian, V.A.; Dolgalev, G.V.; Ilgisonis, E.V.; Lisitsa, A.V.; Poverennaya, E.V. Workability of mRNA Sequencing for Predicting Protein Abundance. Genes 2023, 14, 2065. [Google Scholar] [CrossRef] [PubMed]
- Võsa, U.; Claringbould, A.; Westra, H.-J.; Bonder, M.J.; Deelen, P.; Zeng, B.; Kirsten, H.; Saha, A.; Kreuzhuber, R.; Yazar, S.; et al. Large-scale cis- and trans-eQTL analyses identify thousands of genetic loci and polygenic scores that regulate blood gene expression. Nat. Genet. 2021, 53, 1300–1310. [Google Scholar] [CrossRef] [PubMed]
- Xu, Y.; Ritchie, S.C.; Liang, Y.; Timmers, P.R.H.J.; Pietzner, M.; Lannelongue, L.; Lambert, S.A.; Tahir, U.A.; May-Wilson, S.; Foguet, C.; et al. An atlas of genetic scores to predict multi-omic traits. Nature 2023, 616, 123–131. [Google Scholar] [CrossRef] [PubMed]
- Sun, B.B.; Chiou, J.; Traylor, M.; Benner, C.; Hsu, Y.-H.; Richardson, T.G.; Surendran, P.; Mahajan, A.; Robins, C.; Vasquez-Grinnell, S.G.; et al. Plasma proteomic associations with genetics and health in the UK Biobank. Nature 2023, 622, 329–338. [Google Scholar] [CrossRef] [PubMed]
- Ferkingstad, E.; Sulem, P.; Atlason, B.A.; Sveinbjornsson, G.; Magnusson, M.I.; Styrmisdottir, E.L.; Gunnarsdottir, K.; Helgason, A.; Oddsson, A.; Halldorsson, B.V.; et al. Large-scale integration of the plasma proteome with genetics and disease. Nat Genet 2021, 53, 1712–1721. [Google Scholar] [CrossRef] [PubMed]
- Zhu, Z.; Zhang, F.; Hu, H.; Bakshi, A.; Robinson, M.R.; Powell, J.E.; Montgomery, G.W.; Goddard, M.E.; Wray, N.R.; Visscher, P.M.; et al. Integration of summary data from GWAS and eQTL studies predicts complex trait gene targets. Nat Genet 2016, 48, 481–487. [Google Scholar] [CrossRef] [PubMed]
- Sherman, B.T.; Hao, M.; Qiu, J.; Jiao, X.; Baseler, M.W.; Lane, H.C.; Imamichi, T.; Chang, W. DAVID: a web server for functional enrichment analysis and functional annotation of gene lists (2021 update). Nucleic Acids Res. 2022, 50, W216–W221. [Google Scholar] [CrossRef] [PubMed]
- Wang, Q.S.; Hasegawa, T.; Namkoong, H.; Saiki, R.; Edahiro, R.; Sonehara, K.; Tanaka, H.; Azekawa, S.; Chubachi, S.; Takahashi, Y.; et al. Statistically and functionally fine-mapped blood eQTLs and pQTLs from 1,405 humans reveal distinct regulation patterns and disease relevance. Nat. Genet. 2024, 56, 2054–2067. [Google Scholar] [CrossRef] [PubMed]
- Kirsher, D.Y.; Chand, S.; Phong, A.; Nguyen, B.; Szoke, B.G.; Ahadi, S. The Current Landscape of Plasma Proteomics: Technical Advances, Biological Insights, and Biomarker Discovery. 2025, 2025.02.14.638375. [Google Scholar] [CrossRef]
- Eldjarn, G.H.; Ferkingstad, E.; Lund, S.H.; Helgason, H.; Magnusson, O.Th.; Gunnarsdottir, K.; Olafsdottir, T.A.; Halldorsson, B.V.; Olason, P.I.; Zink, F.; et al. Large-scale plasma proteomics comparisons through genetics and disease associations. Nature 2023, 622, 348–358. [Google Scholar] [CrossRef] [PubMed]
- Haslam, D.E.; Li, J.; Dillon, S.T.; Gu, X.; Cao, Y.; Zeleznik, O.A.; Sasamoto, N.; Zhang, X.; Eliassen, A.H.; Liang, L.; et al. Stability and reproducibility of proteomic profiles in epidemiological studies: comparing the Olink and SOMAscan platforms. Proteomics 2022, 22, e2100170. [Google Scholar] [CrossRef] [PubMed]
| Study Name | Source (PMID) | Data type | Platform | Sample Size | Number of Biomarkers |
|---|---|---|---|---|---|
| eQTLGen | 34475573 | eQTL | Microarray | 31,684 | 19,960 |
| INTERVAL | 36991119 | eQTL | RNAseq | 4,732 | 15,298 |
| UKBB | 37794186 | pQTL | Olink | 34,557 | 2,923 |
| deCODE | 34857953 | pQTL | SomaScan | 35,559 | 4,719 |
| Term | Count in the network | Fold Enrichment | P-value | P-value* |
|---|---|---|---|---|
| Cell adhesion | 25 in 677 | 4.07 | 1.1e-8 | 1.6e-5 |
| Immune system process | 28 in 953 | 3.24 | 1.2e-7 | 1.8e-4 |
| Innate immune response | 22 in 621 | 3.91 | 2.1e-7 | 3.1e-4 |
| Negative regulation of activated T cell proliferation | 6 in 17 | 38.91 | 3.2e-7 | 4.7e-4 |
| Immune response | 20 in 572 | 3.85 | 1.1e-6 | 1.6e-3 |
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2026 by the author. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license.




