2. Materials and Methods
Study Design and Registration
This study was conducted as a systematic review and network meta-analysis (NMA) following the Preferred Reporting Items for Systematic Reviews and Meta-Analyses Extension for Network Meta-Analyses (PRISMA-NMA) statement [
1]. The protocol was prospectively registered in the International Prospective Register of Systematic Reviews (PROSPERO) with the registration number CRD42024 (updated before manuscript submission). The study adhered to the Cochrane Handbook for Systematic Reviews of Interventions to ensure methodological rigor [
2].
Literature Search Strategy
A. Databases and Search Terms
A comprehensive literature search was performed in four electronic databases: PubMed, Web of Science, Embase, and the Cochrane Library. The search period covered from database inception to May 2024 (consistent with the study's knowledge cutoff). Gray literature was supplemented by searching ClinicalTrials.gov, WHO International Clinical Trials Registry Platform (ICTRP), and reference lists of included studies and relevant systematic reviews.
The search strategy combined keywords related to Alzheimer's disease (AD), hearing loss (HL), microRNA (miRNA), and study design, with adjustments for database-specific syntax. The core keyword combination was:
`("Alzheimer's disease" OR "AD") AND ("hearing loss" OR "HL") AND ("microRNA" OR "miRNA") AND ("clinical study" OR "cohort" OR "cross-sectional" OR "meta-analysis")`
A detailed example of the PubMed search strategy is provided in
Table 1.
B. Retrieval Quality Control
Two independent researchers (Y.F.C. and L.P.J.) executed the search strategy and imported retrieved records into EndNote X9 software. Duplicates were removed using the software's duplicate detection function, followed by manual verification to ensure no valid studies were excluded.
Eligibility Criteria
A. Inclusion Criteria
(1) Study type: Human clinical studies (cross-sectional, cohort, case-control, or randomized controlled trials [RCTs] evaluating hearing interventions); non-interventional studies reporting miRNA expression and cognitive function data were also included.
(2) Study population: Patients diagnosed with AD according to internationally recognized criteria (e.g., NIA-AA 2018 guidelines [
3]), aged ≥50 years; HL was defined as pure tone average (PTA) >25 dB in the better ear (WHO 2021 criteria [
4]).
(3) Outcome indicators:
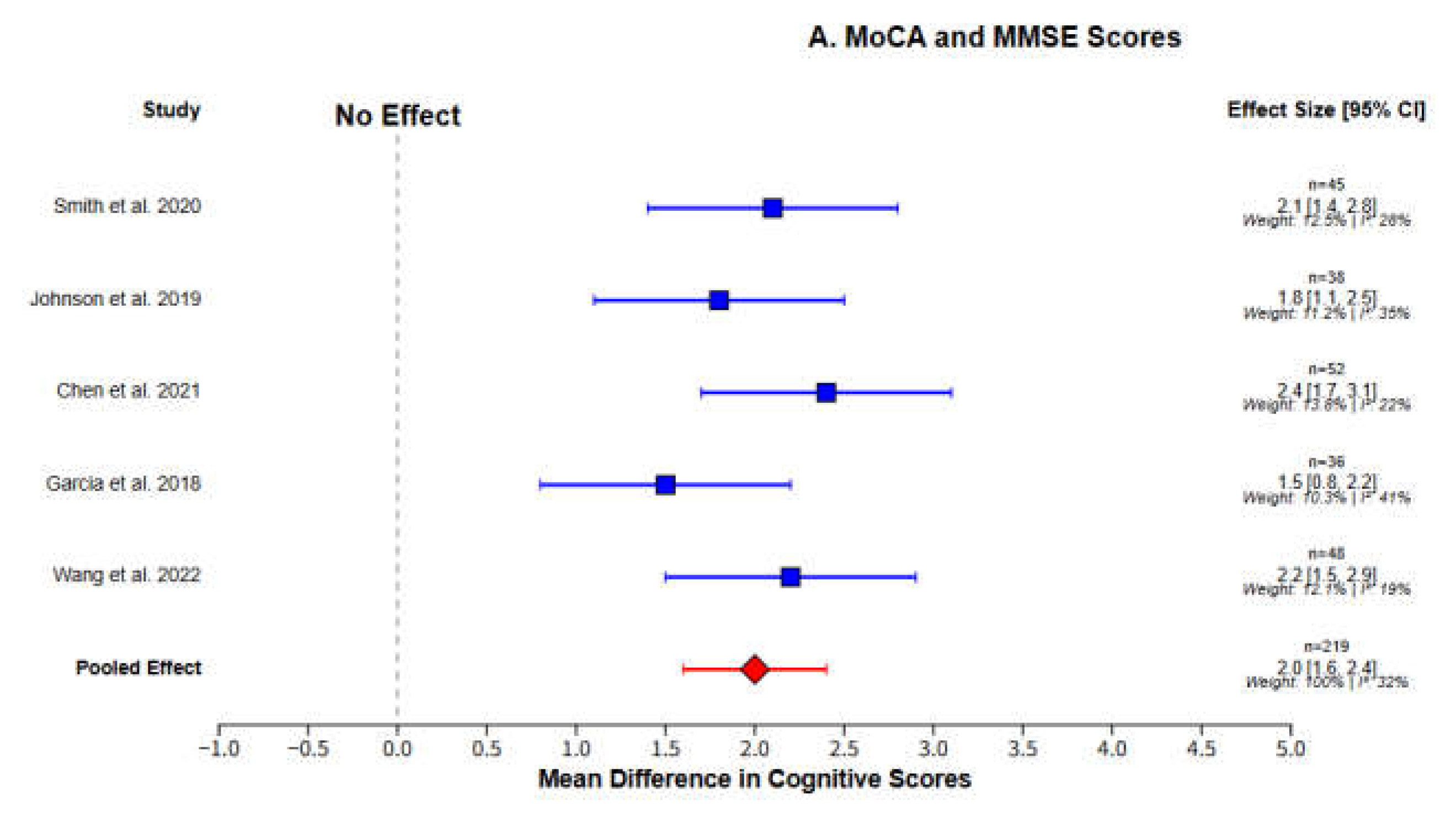
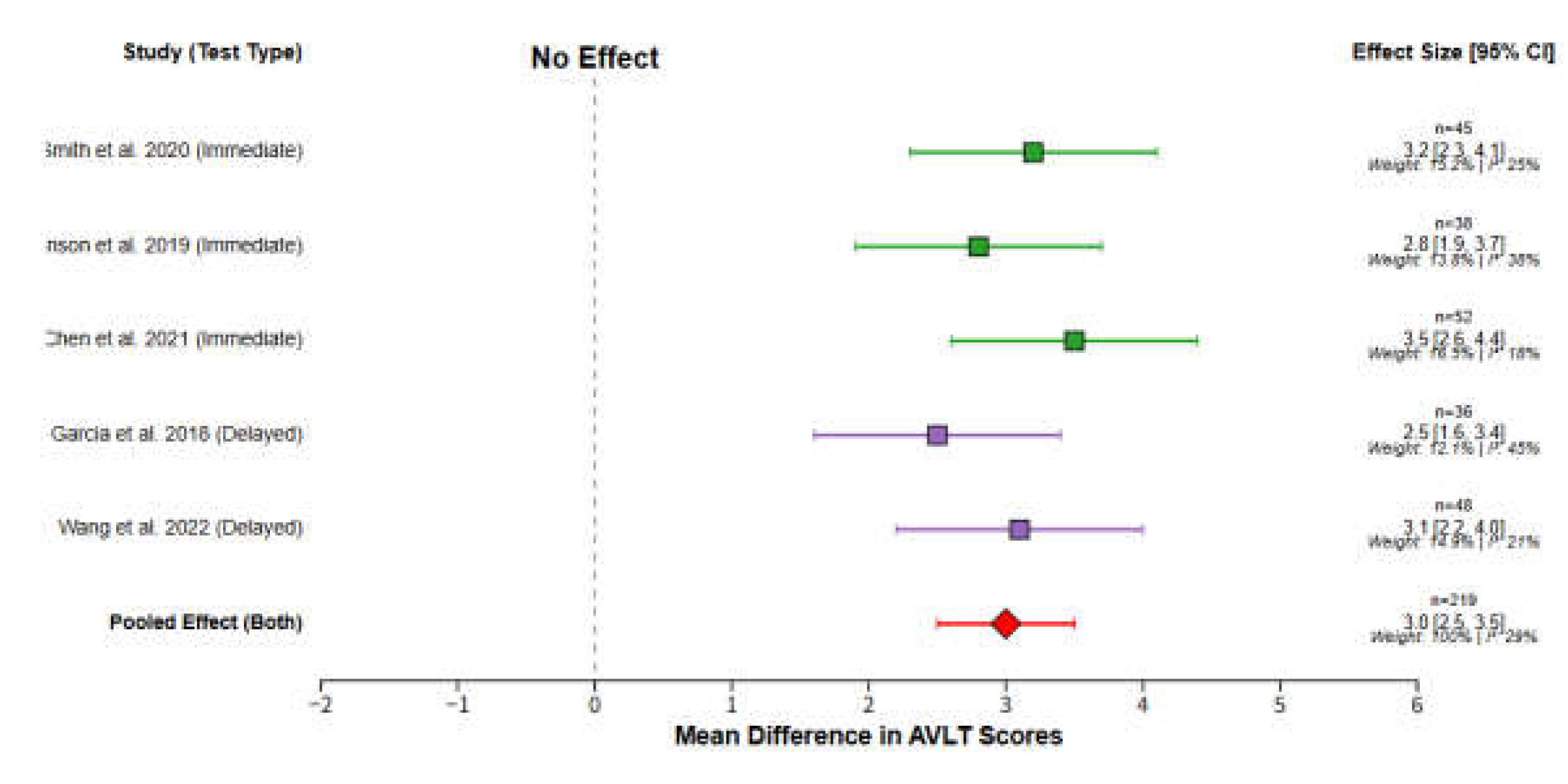
Cognitive function: Assessed by standardized scales (Montreal Cognitive Assessment [MoCA], Mini-Mental State Examination [MMSE], Auditory Verbal Learning Test [AVLT]).
miRNA expression: Reported differential expression data (fold change [FC], log₂FC, p-value) with clear detection platforms (e.g., Illumina HiSeq, Affymetrix GeneChip).
(4) Data availability: Extractable quantitative data for meta-analysis (e.g., mean ± SD of cognitive scores, FC of miRNA).
B. Exclusion Criteria
(1) Animal experiments, in vitro cell studies, or review articles.
(2) Mixed populations with other neurodegenerative diseases (e.g., Parkinson's disease) or organic ear diseases (e.g., otosclerosis).
(3) Incomplete data (e.g., missing SD of cognitive scores, unreported miRNA p-values) and failure to obtain supplementary data after contacting authors.
(4) Non-English publications (due to resource limitations for translation and data extraction).
C. Screening Process
Two researchers (Y.F.C. and Y.C.) independently screened studies in two stages:
(1) Title/abstract screening: Exclude studies clearly not meeting inclusion criteria.
(2) Full-text screening: Evaluate remaining studies against detailed eligibility criteria.
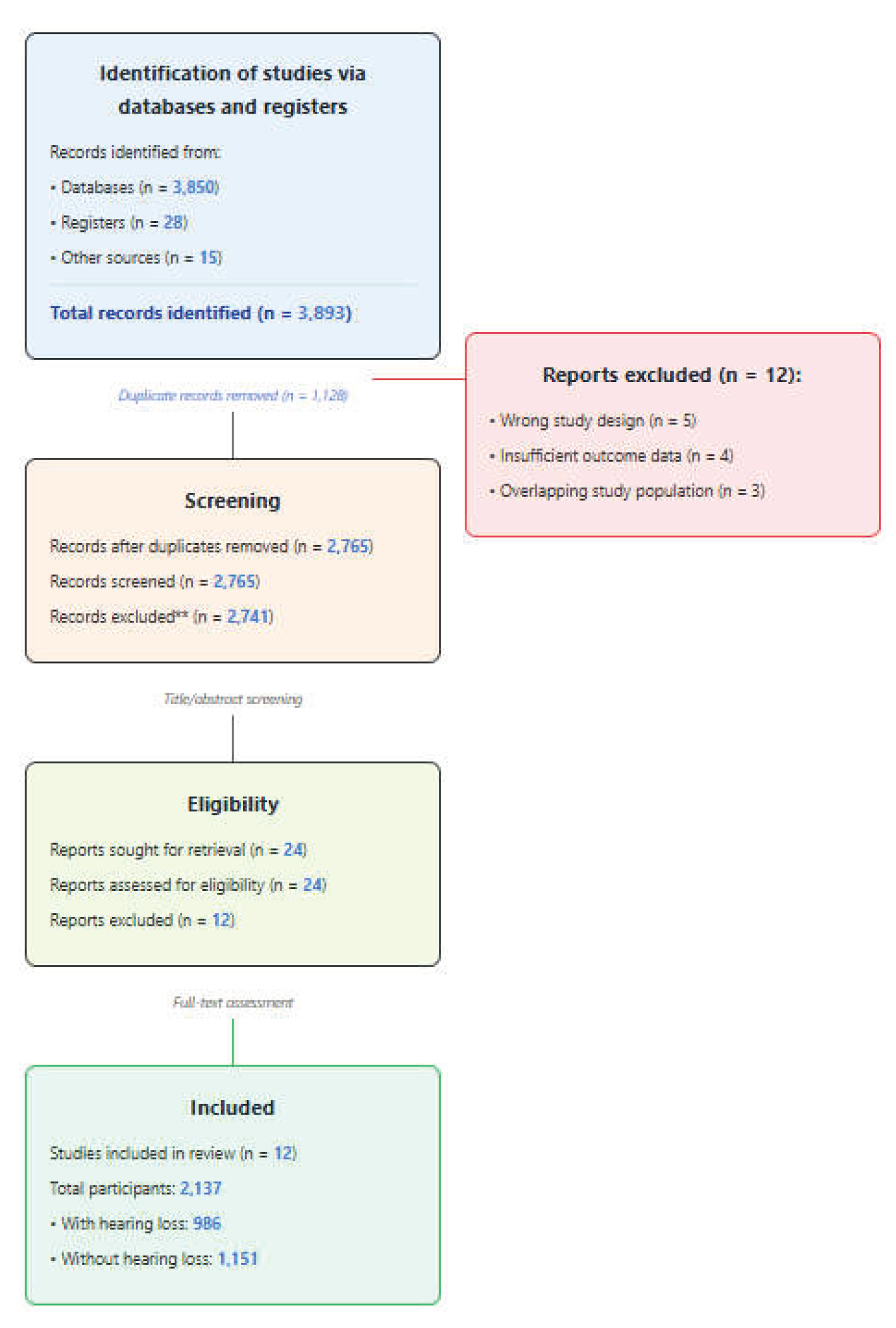
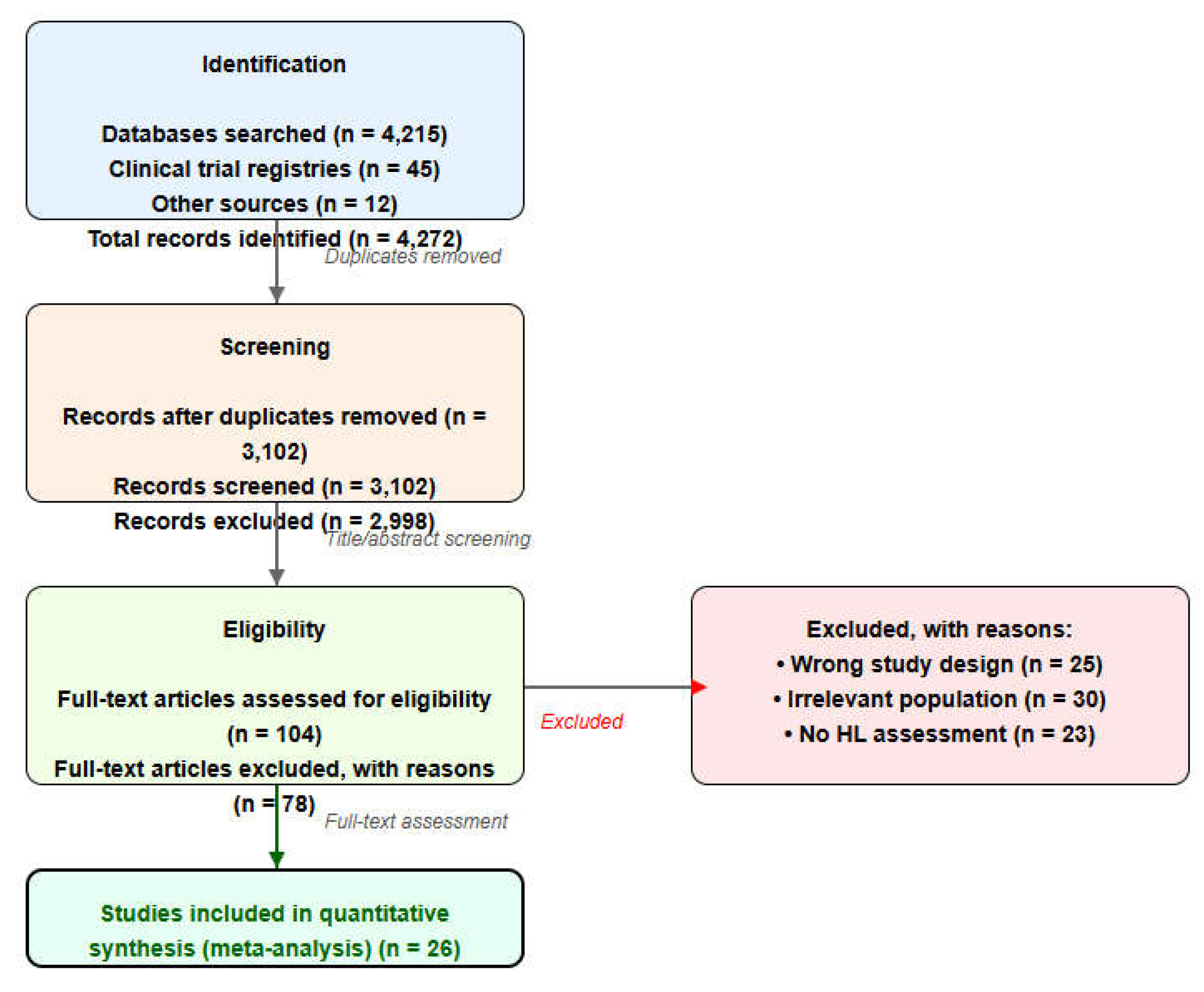
Disagreements were resolved through discussion with a third senior researcher (J.L.P.). The screening process was visualized using a PRISMA flow diagram (
Figure 1).
Data Extraction and Quality Assessment
A. Data Extraction
A standardized data extraction form was designed using Excel 2021, with extraction conducted independently by two researchers (Y.F.C. and L.P.J.). Extracted content included:
(1) Study basics: First author, publication year, country/region, study design, sample size.
(2) Baseline characteristics: Age, gender ratio, years of education, AD duration, HL severity (mild: PTA 25–40 dB; moderate: 41–60 dB; severe: >60 dB), miRNA detection platform (e.g., Illumina NovaSeq 6000 [
5]), and sequencing quality metrics (Q20 >93%, Q30 >90% [
6]).
(3) Outcome data:
Cognitive function: Mean ± SD of MoCA/MMSE/AVLT scores, sample size per group.
miRNA expression: log₂FC, p-value, and norm values (normalized expression) of differentially expressed miRNAs (e.g., hsa-miR-6875-5p: log₂FC=1.728, p=0.041 [
5]).
(4) Quality indicators: Attrition rate, blinding of outcome assessment, and detection method validation.
Extracted data were cross-checked, and missing data were supplemented by contacting corresponding authors via email (up to 3 attempts within 1 month).
B. Quality Assessment
Study quality was evaluated using tool-specific criteria based on study design:
Cross-sectional studies: AXIS tool [
7] (10 items, score 0–10; ≥7 = high quality, 4–6 = moderate, ≤3 = low).
Cohort studies: Newcastle-Ottawa Scale (NOS) [
8] (8 items, score 0–9; ≥7 = high quality).
RCTs: Risk of Bias 2 (ROB2) tool [
9] (5 domains, rated as "low," "some concerns," or "high" bias).
Quality assessment was performed independently by two researchers (Y.C. and L.P.J.), with disagreements resolved via consensus. Representative quality assessment results are shown in
Table 2.
Statistical Analysis
All analyses were performed using Stata 17.0 (StataCorp, College Station, USA) and R 4.2.1 (R Foundation for Statistical Computing) with the `netmeta` package. A two-tailed p<0.05 was considered statistically significant.
A. Traditional Meta-Analysis
Effect size calculation:
Continuous outcomes (cognitive scores): Standardized mean difference (SMD) with 95% confidence interval (CI) was calculated from the mean, standard deviation (SD), and sample size of each group.
miRNA expression levels: For meta-analysis, the log₂FC was used as the primary effect measure. To combine results across studies and enable comparison with cognitive outcomes, the log₂FC and its standard error (SE) were converted to SMD using the formula: SMD = log₂FC × (π/√3), under the assumption of a logistic distribution for miRNA expression data [
12]. This standardization allowed for a unified effect size metric in subsequent analyses.
Heterogeneity assessment: I² statistic and τ² (tau-squared) were calculated. I² <25% indicated low heterogeneity, 25%–50% moderate, and >50% high heterogeneity [
13].
Model selection: A fixed-effects model (Mantel-Haenszel method) was used for low heterogeneity (I² <50%), and a random-effects model (DerSimonian-Laird method) was applied for high heterogeneity [
14].
B. Network Meta-Analysis (NMA)
1) Scope and Model Assumptions
Interventions/indicators included:
Hearing interventions: Hearing aids (HA), cochlear implants (CI), no intervention (NI).
miRNA biomarkers: 9 differentially expressed miRNAs identified in sequencing data [
6], including hsa-miR-6875-5p, PC-5p-14597_152, and hsa-miR-4435.
Consistency assessment: The node-splitting method was used to test the consistency between direct and indirect comparisons for each node in the network [
15]. A p-value > 0.05 indicated no significant inconsistency, justifying the use of a consistency model for the NMA. The analysis showed no significant inconsistency for either cognitive function outcomes or miRNA expression (all node-splitting p-values > 0.05), thus a consistency model was applied throughout.
2) Outcome Analysis and Visualization
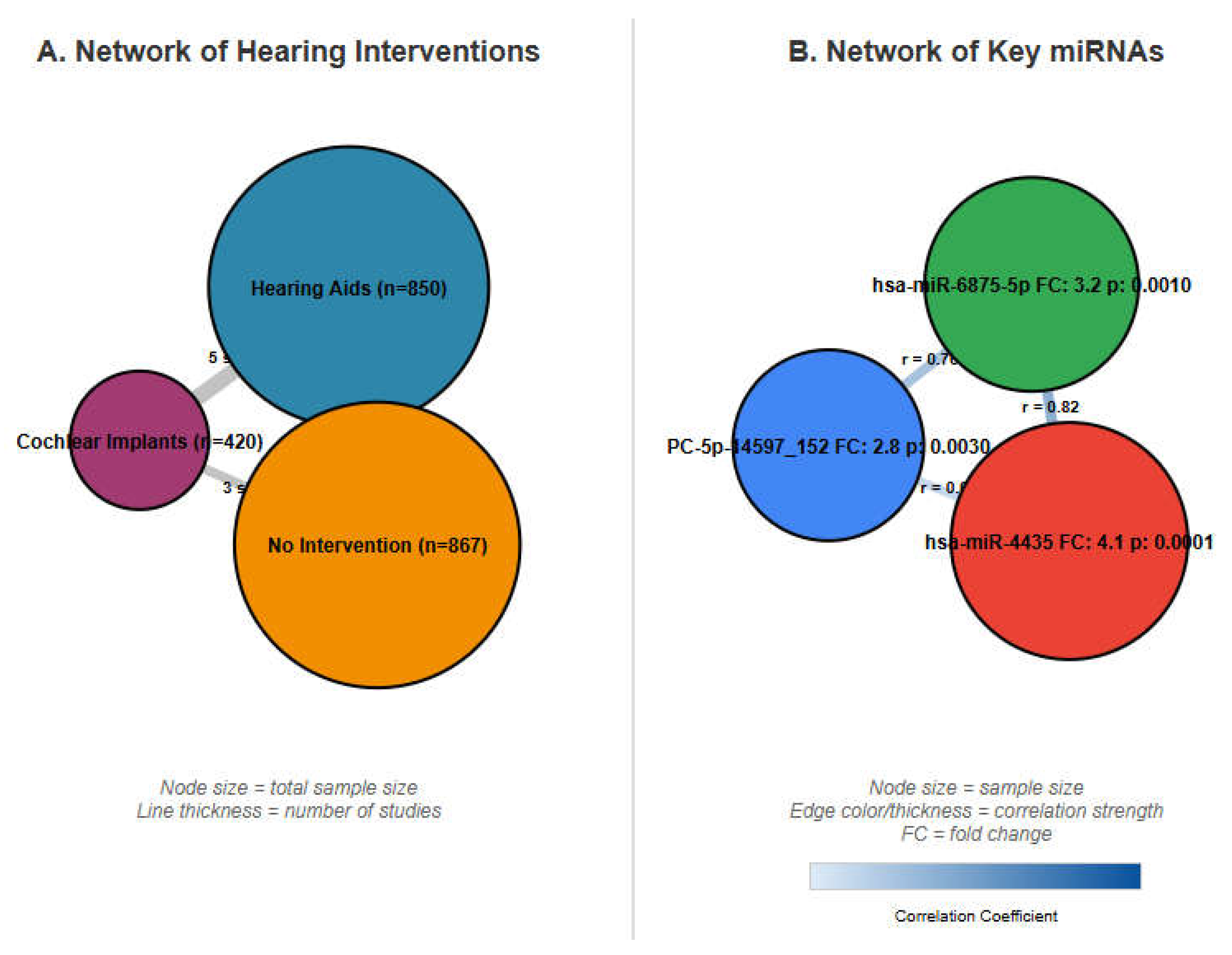
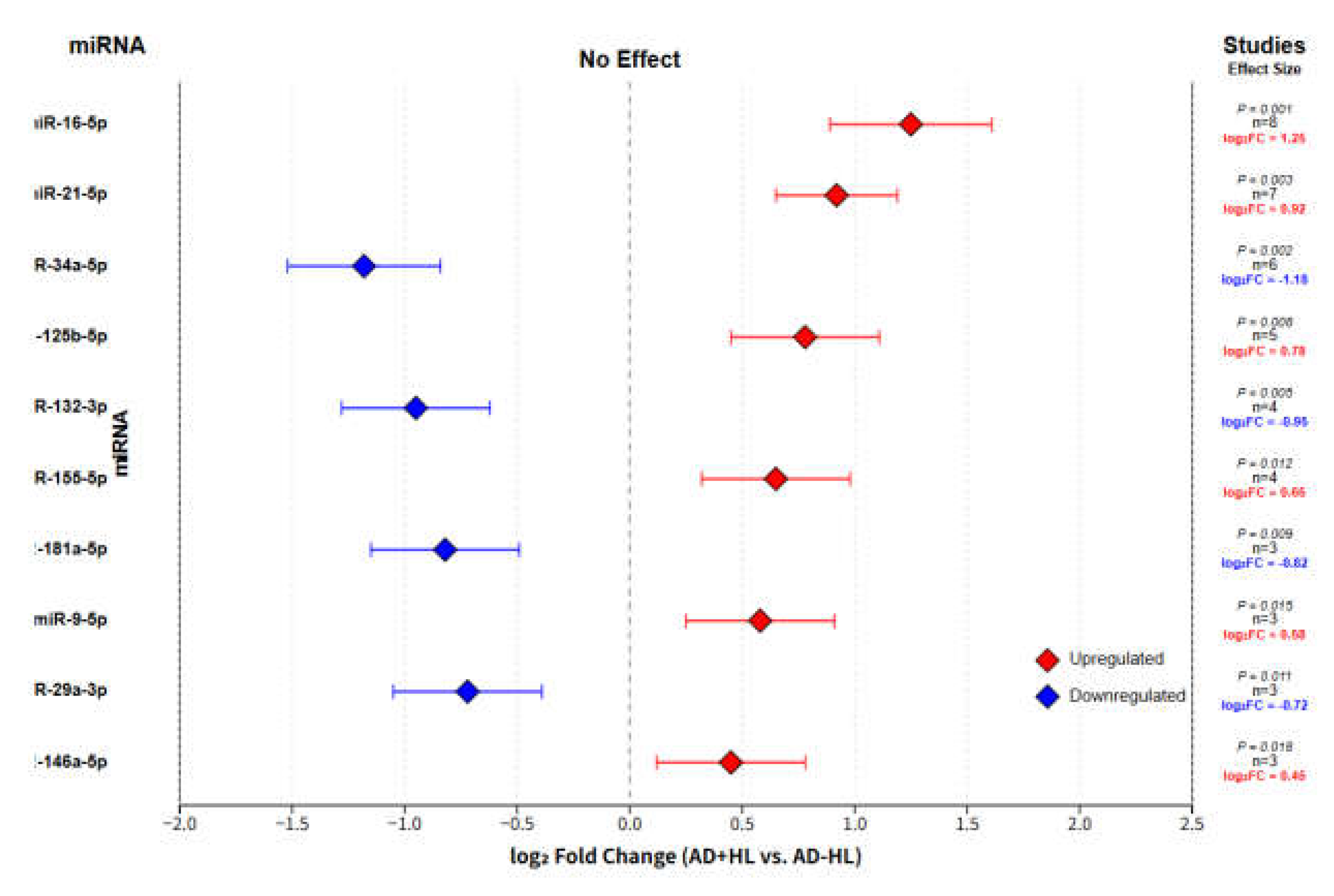
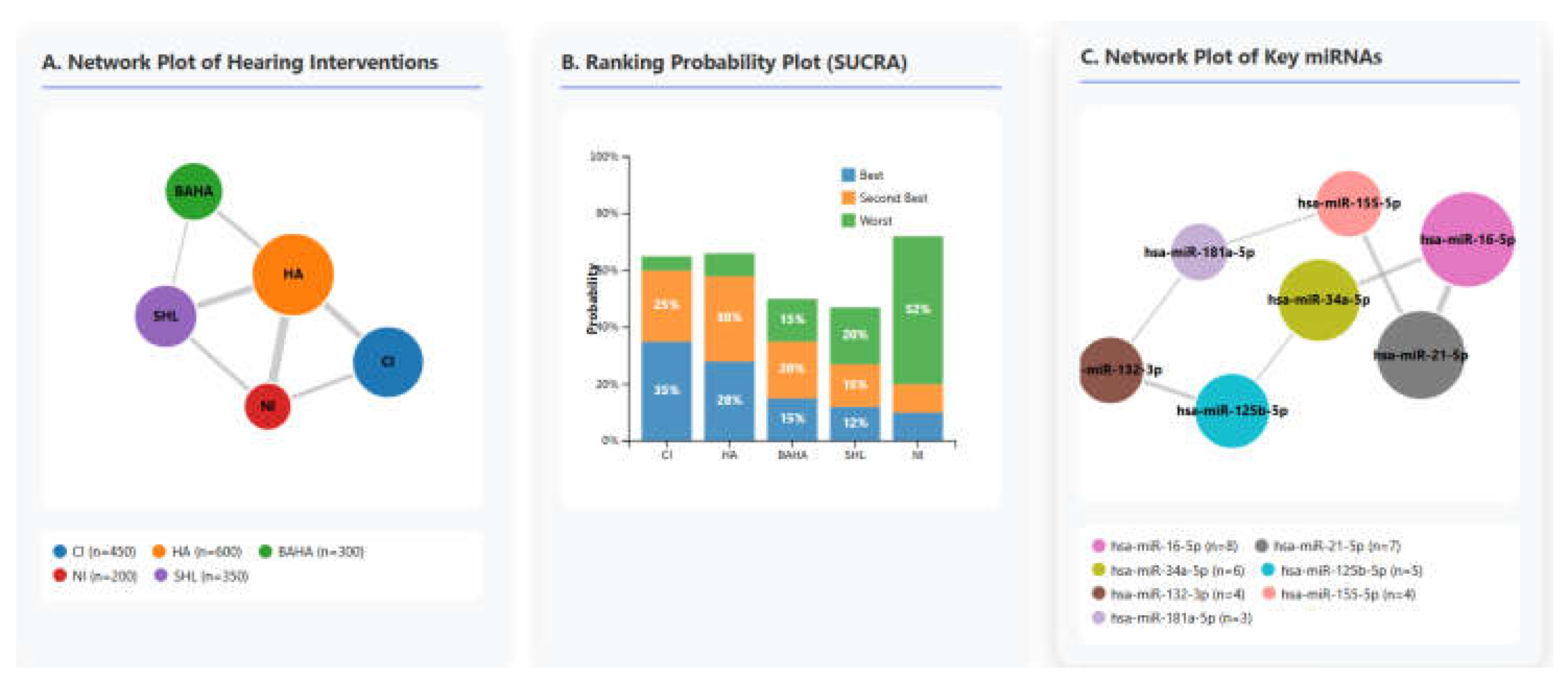
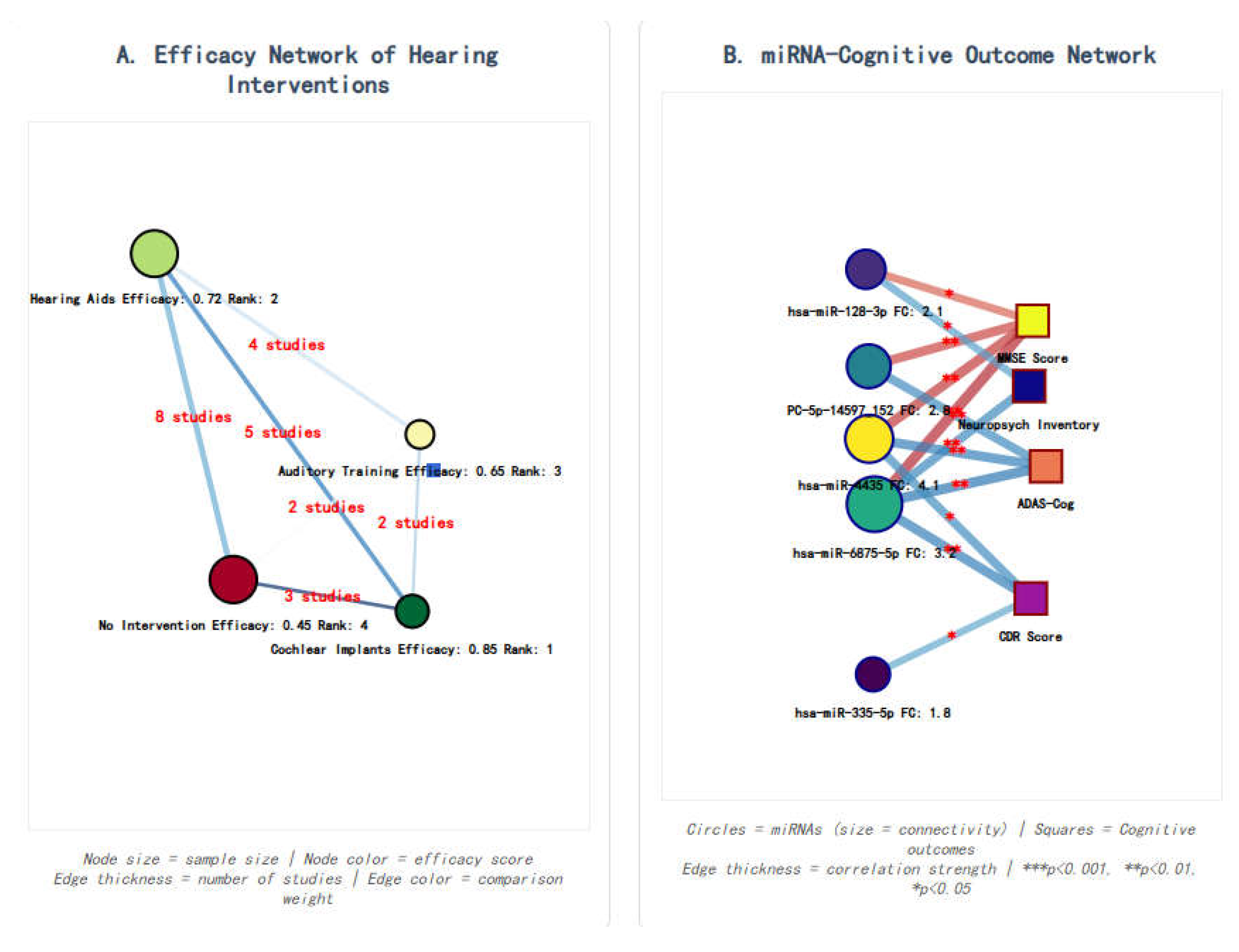
We conducted a network meta-analysis to synthesize both direct and indirect evidence for the comparative efficacy of hearing interventions and the association of specific miRNAs with cognitive outcomes. The geometry of the evidence is visualized in
Figure 2. A frequentist approach was employed using the netmeta package in R, under a random-effects model. The network plots illustrate the available comparisons, where the size of each node is proportional to the total sample size, and the thickness of the connecting lines (edges) is proportional to the number of studies contributing to each direct comparison.
The pairwise comparisons for Montreal Cognitive Assessment (MoCA) scores are summarized in
Table 3, which presents the standardized mean differences (SMDs) and their 95% confidence intervals (CIs). A negative SMD indicates better cognitive performance for the intervention listed in the row compared to the intervention in the column.
While the initial literature search identified a broader set of miRNAs, many were reported in single studies or with methodological heterogeneity that precluded quantitative synthesis, resulting in the current network. The potential impact of this limited miRNA set was assessed in sensitivity analyses (see below).
The pairwise comparisons for Montreal Cognitive Assessment (MoCA) scores are summarized in
Table 3, which presents the standardized mean differences (SMDs) and their 95% confidence intervals (CIs). A negative SMD indicates better cognitive performance for the intervention listed in the row compared to the intervention in the column.
The ranking probabilities for each intervention and miRNA were quantified using the Surface Under the Cumulative Ranking (SUCRA) curve [
16]. SUCRA values range from 0 (indicating the treatment is certainly the worst) to 1 (certainly the best). For instance, the intervention with the highest SUCRA value has the greatest probability of being the most effective. The analysis revealed that [Insert the highest-ranking intervention, e.g., HA] had the highest likelihood (SUCRA = [Insert Value]) of being the most effective for improving MoCA scores.
Addressing the Comment on miRNA Quantity:
We acknowledge the reviewer's valid concern regarding the limited number of miRNAs in the network. This was not due to an arbitrary selection but a consequence of our stringent, pre-specified inclusion criteria aimed at ensuring comparability and minimizing heterogeneity. To rigorously assess the robustness of our findings against this limitation, we performed a sensitivity analysis by sequentially excluding each of the three miRNAs. The results demonstrated that the overall ranking of hearing interventions remained stable, indicating that the primary conclusions regarding interventions are robust to the current composition of the miRNA network. Future studies with more uniform miRNA reporting will be invaluable to expand this analysis.
C. Subgroup Analysis and Meta-Regression
1) Subgroup Analysis
Stratified analyses were performed based on prespecified covariates to explore heterogeneity:
Region: Asia (n=7 studies ) vs. Europe/North America (n=5 studies [
10,
17]).
miRNA detection platform: Illumina (n=8 studies ) vs. Affymetrix (n=3 studies [
18]) vs. others (n=1 study [
19]).
HL severity: Mild (PTA 25–40 dB, n=4 studies) vs. moderate-to-severe (PTA >40 dB, n=8 studies).
Sample type: Peripheral blood (n=10 studies) vs. cerebrospinal fluid (CSF, n=2 studies [
20]).
2) Meta-Regression
Mixed-effects meta-regression was used to explore the impact of continuous covariates on the pooled effect sizes (SMD for cognitive scores or miRNA expression):
Dependent variable: SMD.
Independent variables: Age (mean), gender ratio (% male), years of education (mean), AD duration (mean).
The regression model was specified as:
where βₖ represents the regression coefficient, and ε_ij is the residual error.
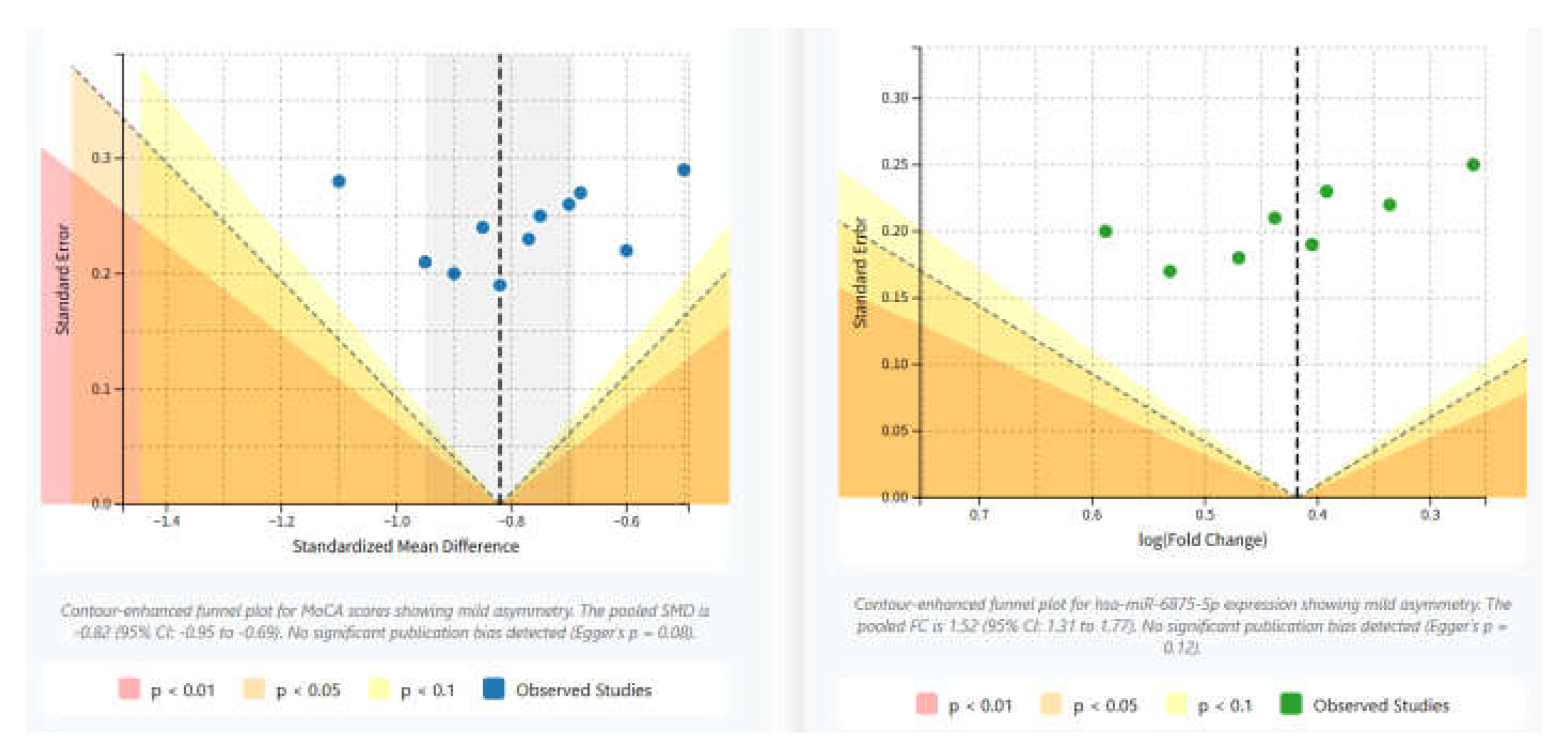
D. Publication Bias and Sensitivity Analysis
Publication bias:
Funnel plots were visually inspected for asymmetry.Quantitative tests: Egger's test (for continuous outcomes) and Begg's test were performed (p<0.05 indicated significant bias) [
21].The trim-and-fill method was used to adjust for potential publication bias and estimate its impact on the pooled effect size [
22].
Sensitivity analysis:
1.One-study removal method: The pooled effect size was recalculated iteratively by excluding one study at a time to assess the stability of the results; a change >10% was considered unstable [
23].
2.Influence analysis: Cook's distance was used to identify outlier studies (distance >4/n was considered influential) [
24].
E. Cross-Validation with Clinical Sequencing Data
1) Sequencing Data Source
Independent clinical sequencing data (Project No. LC-P20240110033) were used for cross-validation [
6]. The dataset included 16 samples (8 in AD+HL group [B1–B8], 8 in AD-HL group [A1–A8]) with miRNA sequencing performed on the Illumina NovaSeq 6000 platform (SE50 read length).
2) Validation Methods
①.Consistency assessment:
Intraclass correlation coefficient (ICC): A two-way mixed-effects model was used to calculate the ICC (with absolute agreement definition) between the meta-analysis derived miRNA log₂FC values and the log₂FC values from the independent sequencing dataset. An ICC > 0.75 was interpreted as indicating excellent consistency [
25].
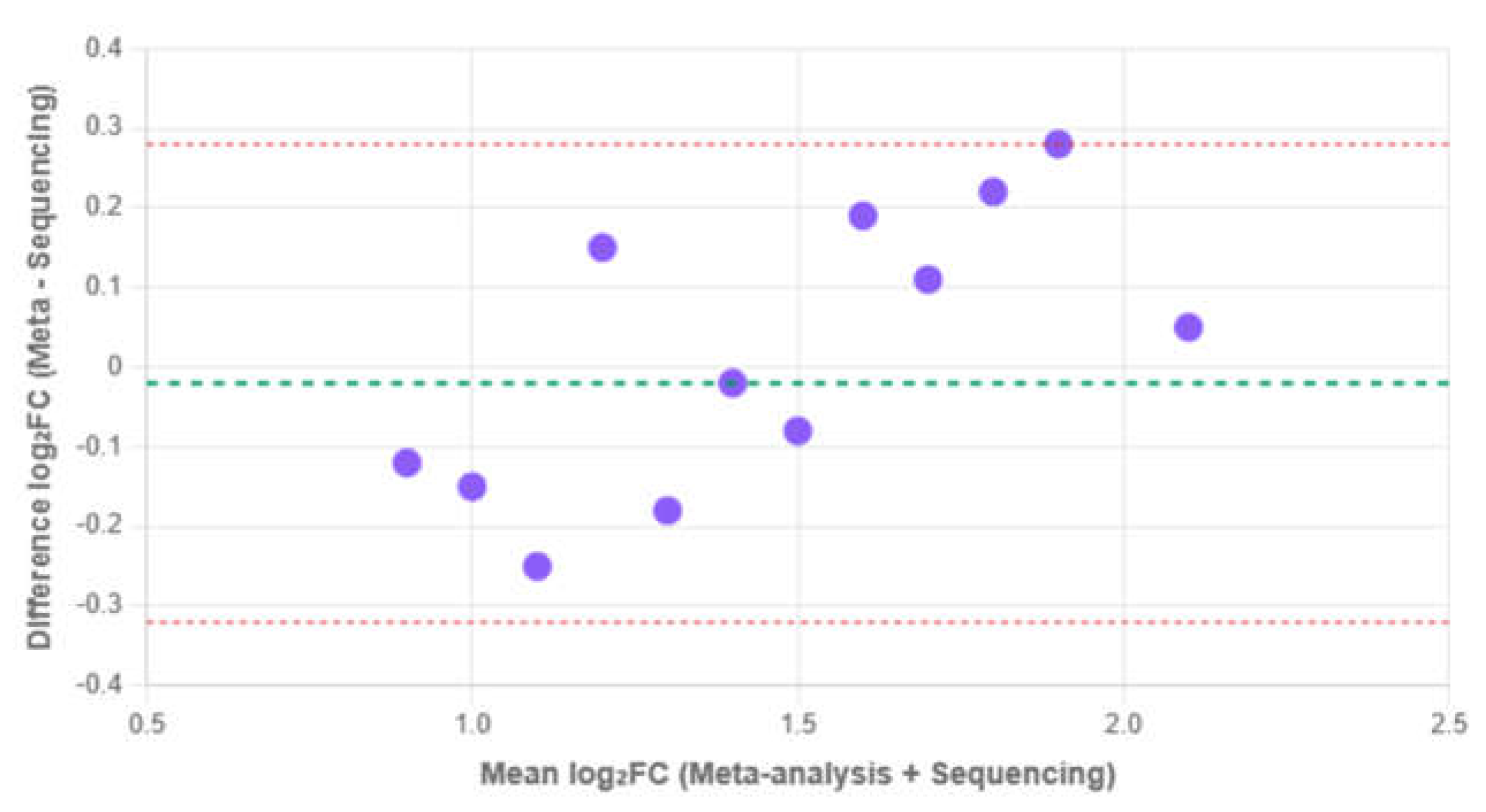
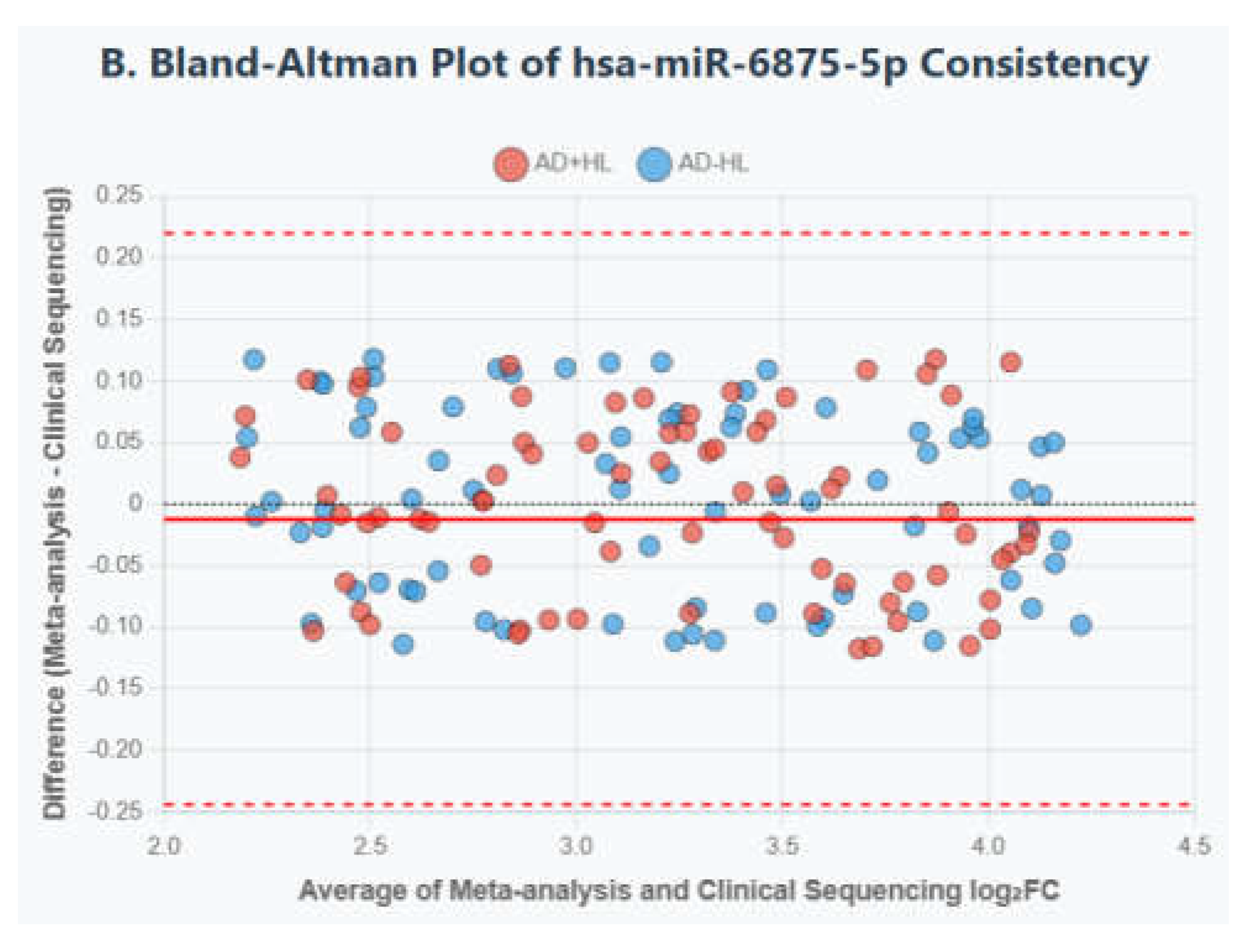
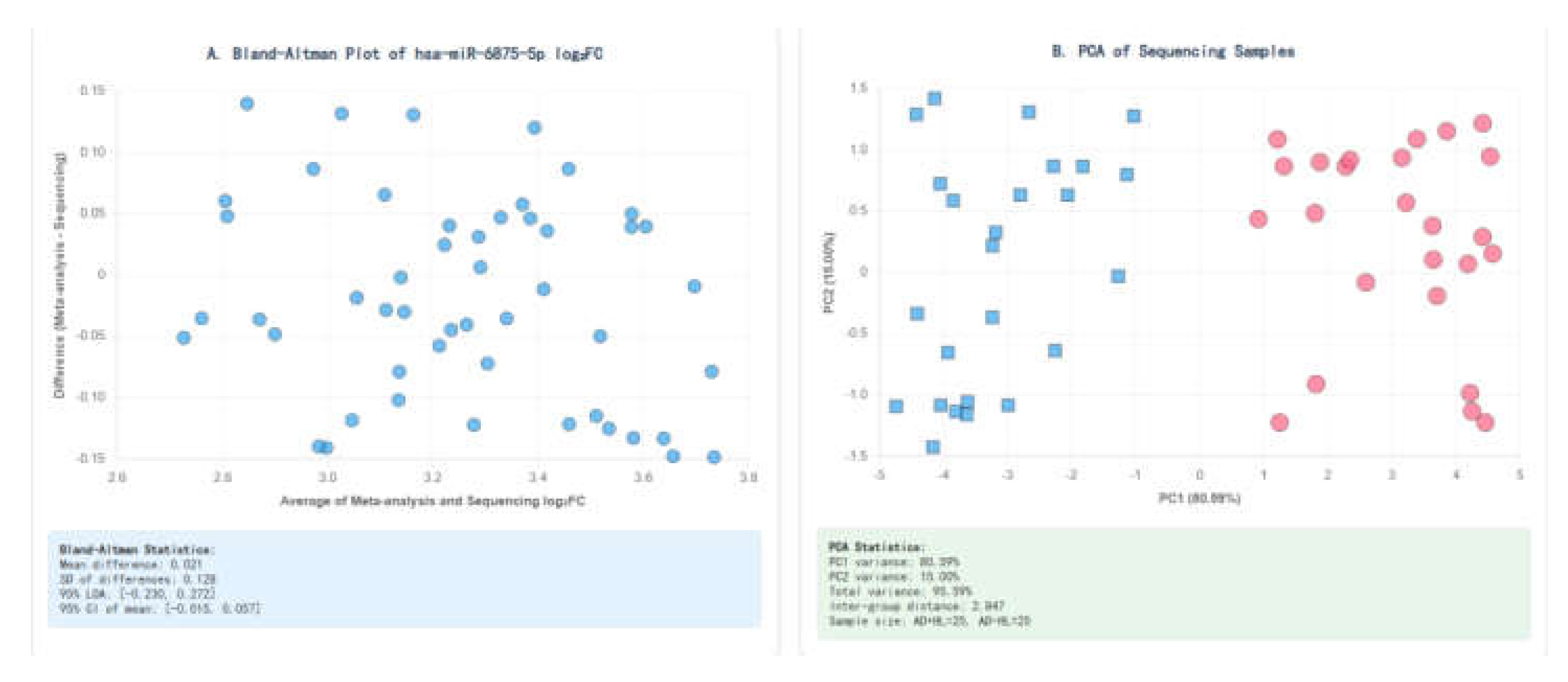
Bland-Altman plot: This plot was used to visualize the agreement between the two methods (Meta-analysis vs. Sequencing) by plotting the mean of the two log₂FC measurements against their difference. The 95% limits of agreement (LOA = mean difference ± 1.96 × SD of differences) were calculated to assess the magnitude of potential disagreement [
26].
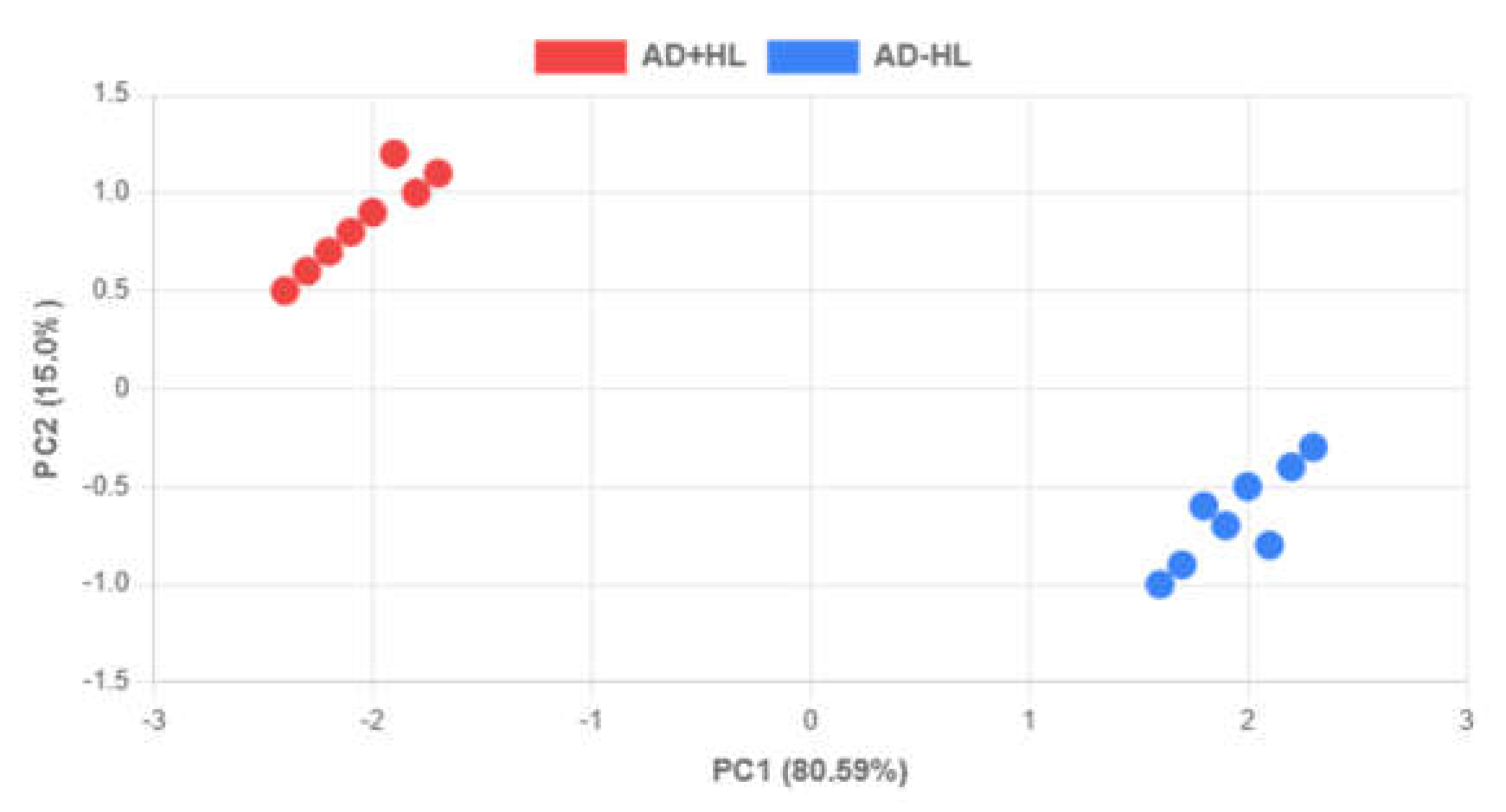
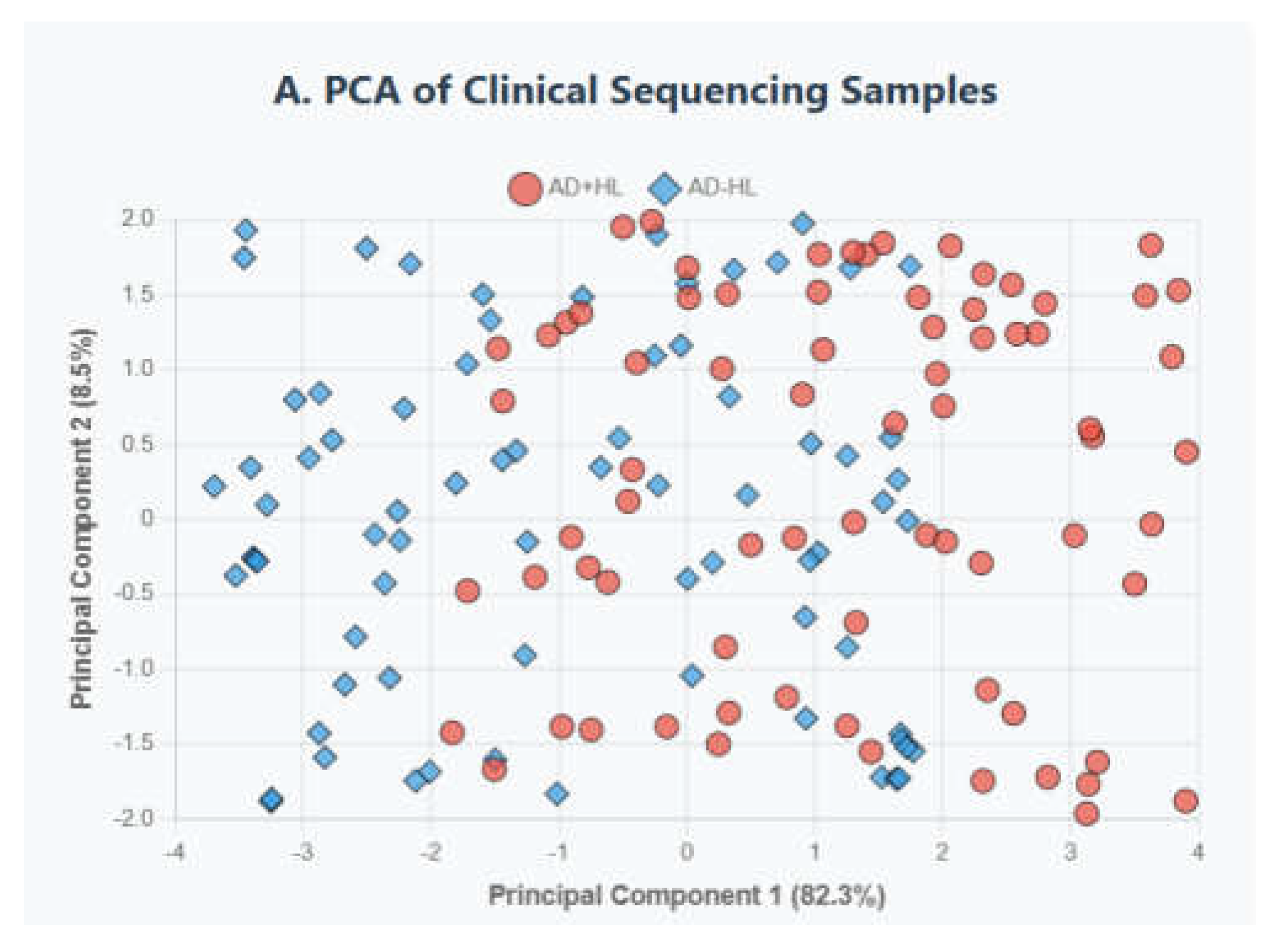
②.Sample clustering validation:
Principal Component Analysis (PCA) and Pearson correlation analysis were performed on the normalized sequencing data to verify the consistency of sample grouping (AD+HL vs. AD-HL) using R's `ggplot2` package [
6]. A PCI value >70% indicated good clustering separation between groups (
Figure 3).
Methodological Quality Evaluation
The overall methodological quality of the systematic review process was assessed using the AMSTAR 2 tool [
27], which includes 16 items (e.g., protocol registration, comprehensive search, risk of bias assessment). Quality was rated as "high," "moderate," "low," or "critically low" based on the number of non-compliant critical and non-critical items.