Submitted:
31 March 2026
Posted:
01 April 2026
You are already at the latest version
Abstract
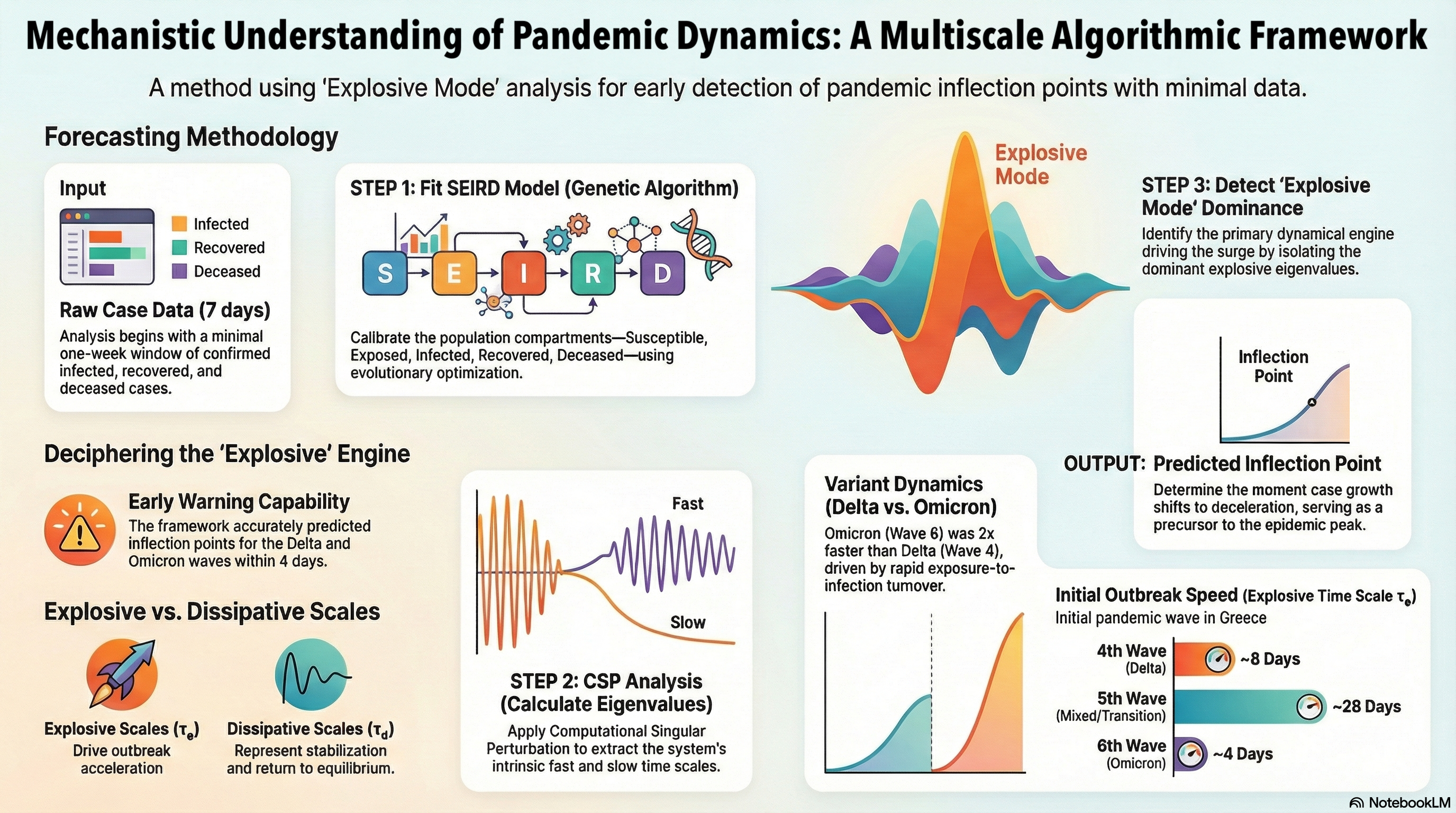
Keywords:
1. Introduction
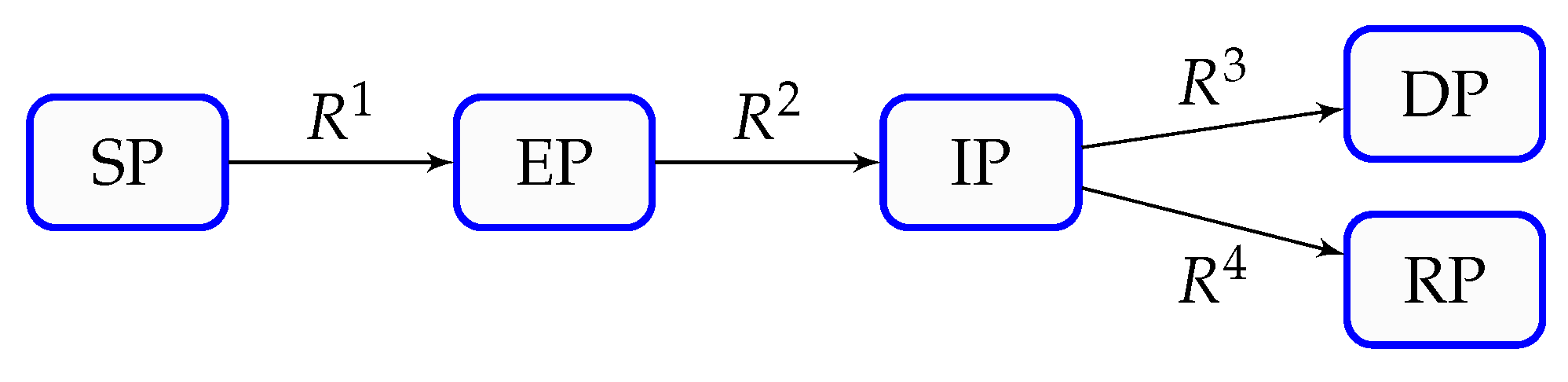
| Mathematical Term (CSP) | Biological / Epidemiological Translation | Practical Significance in This Study |
| Mode () | Dynamical component | A component of the model, the action of which is characterized by an explosive or dissipative time scale. The mode that is characterized by the fastest time scale drives the initial surge, while all other modes sustain or attenuate the epidemic wave before stabilization occurs. |
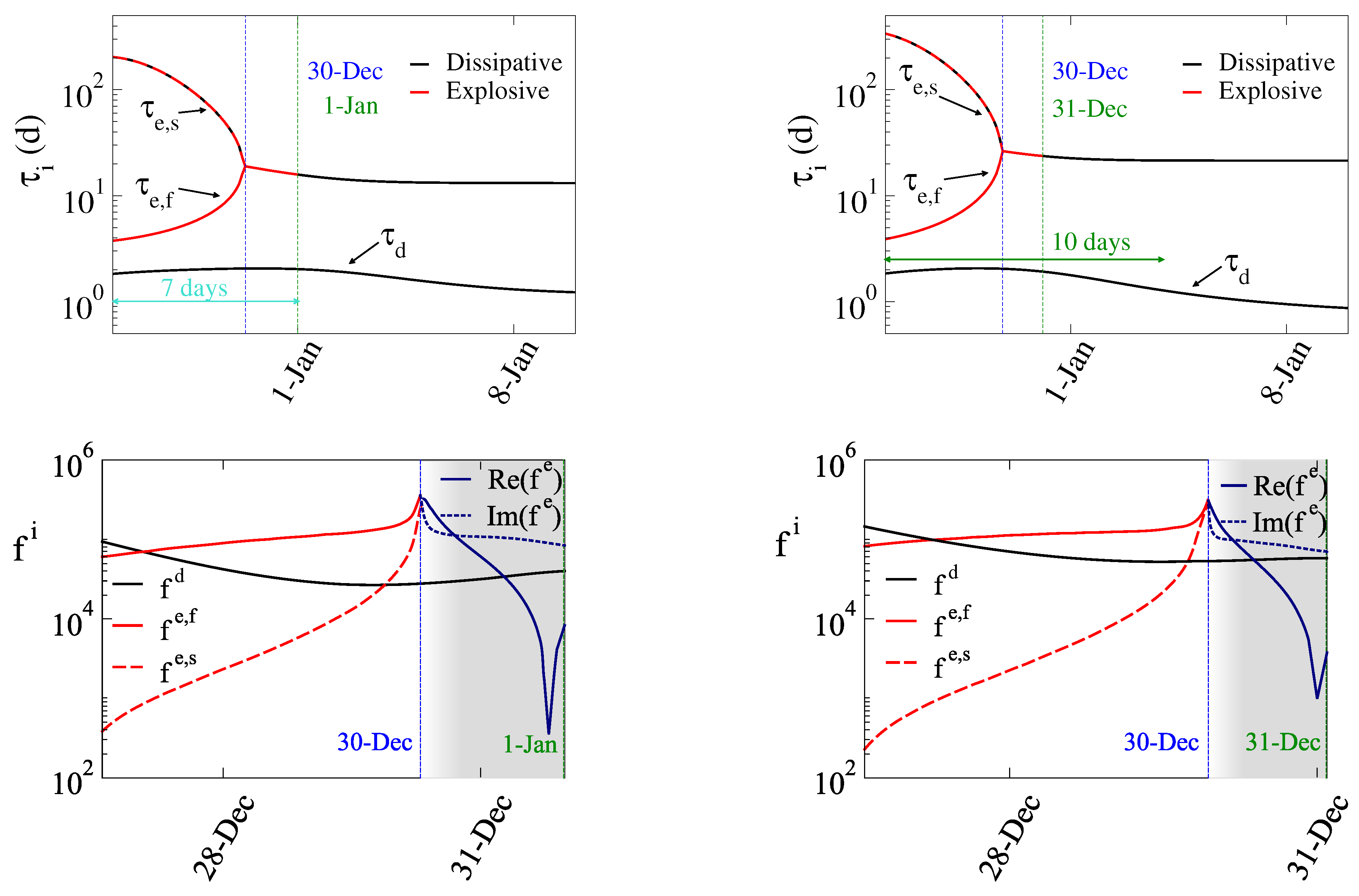
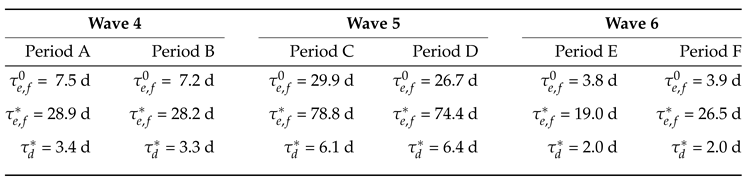
| Explosive Time Scale () | Characteristic expansion time | Represents the time frame of the action of a mode (explosive) that tends to drive the system away from equilibrium. A smaller value (e.g., ∼4 days during the 6th wave) indicates a more aggressive and rapidly spreading variant compared to larger values (e.g., ∼8 days during the 4th wave or ∼28 days during the 5th wave). |
| Dissipative Time Scale () | Stabilization or damping rate | Represents the time frame of the action of a mode (dissipative) that tends to drive the system towards equilibrium. When a dissipative mode dominates, the outbreak transitions into decay and active cases begin to decline. |
| Slow Invariant Manifold (SIM) | Epidemic trajectory | The reduced-dimensional path followed by the epidemic once fast transient processes subside. It represents the established dynamical regime during the outbreak phase. |
| Amplitude (f) | Driver intensity | Provides a measure of the impact of a mode in driving the epidemic wave. In the case of an explosive mode, a high amplitude indicates dominant transmission dynamics, while a low amplitude signals increasing control or immunity effects. |
| Time Scale Participation Index (TPI) | Mechanism identifier | Quantifies the contribution of individual biological transitions (e.g., transmission, incubation, recovery) to the time scale. In the case of an explosive time scale, it determines the degree to which the outbreak is promoted by high transmission () or by rapid progression from exposure to infection (), or it is obstructed by recovery (). |
| Pointer (Po) | Population influence index | Identifies which population compartment most strongly influenced by a specific mode. For example, a high Pointer value for the exposed population during the 6th wave indicates that rapid progression from exposure to infection drives the surge. |
| Inflection Point | Turning point in epidemic acceleration | The moment when outbreak growth shifts from acceleration to deceleration. This precedes the peak of infected population and serves as an early warning indicator of the impending peak. |
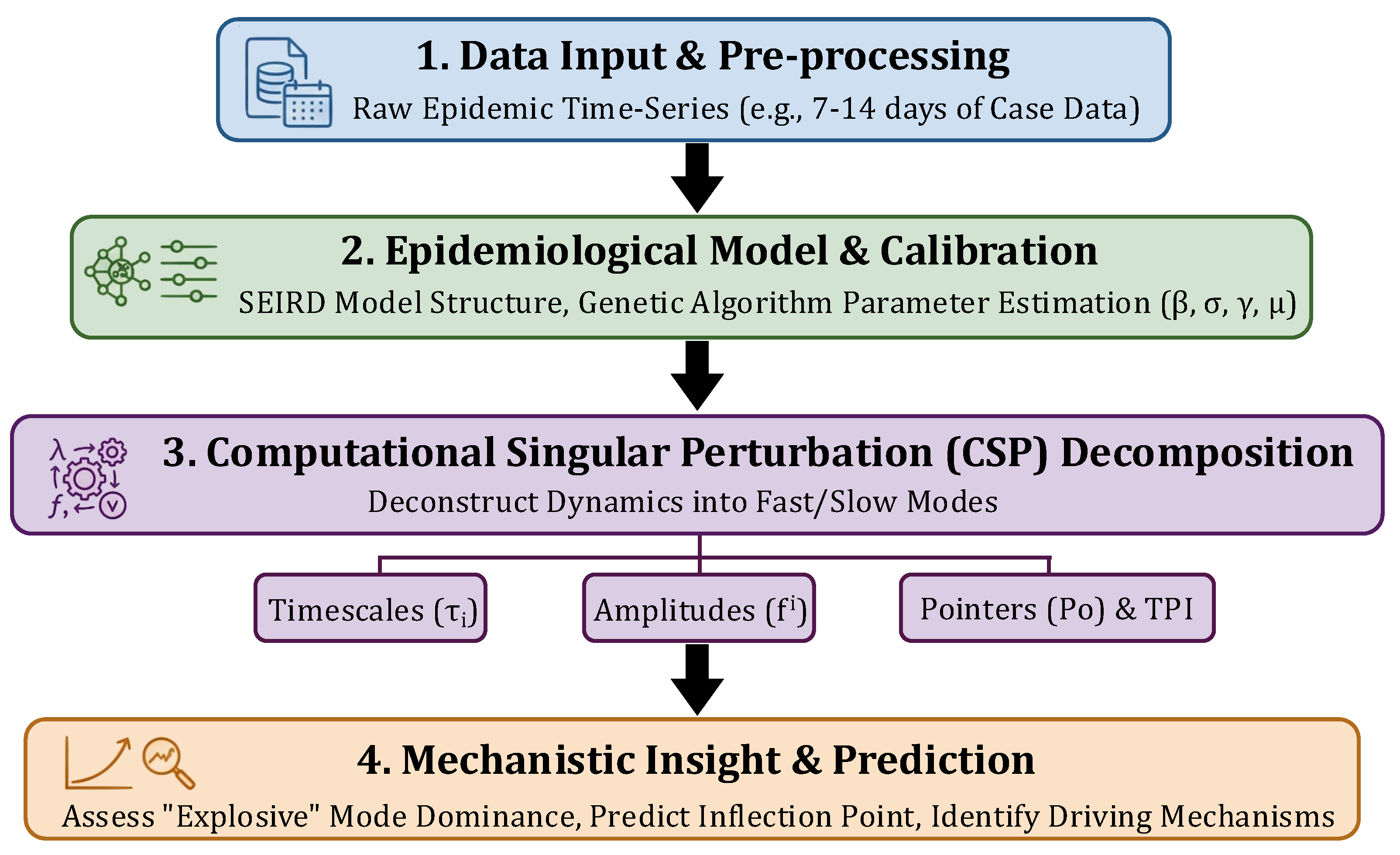
2. Materials and Methods
2.1. The Mathematical Model
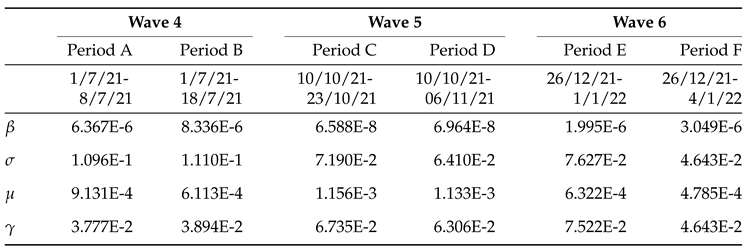
2.2. Model Calibration
2.3. Modeling Assumptions and Vaccination Effects
2.4. Time Scale Analysis and CSP Tools
3. Results
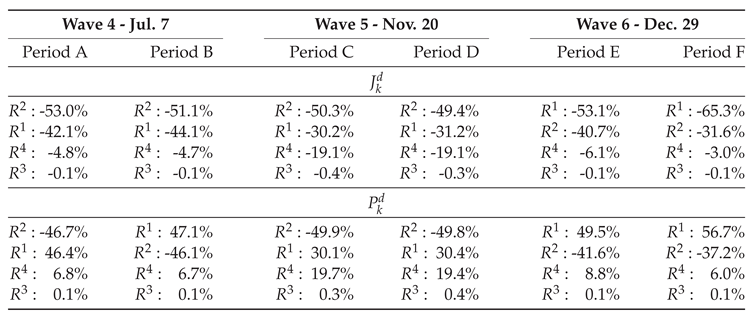
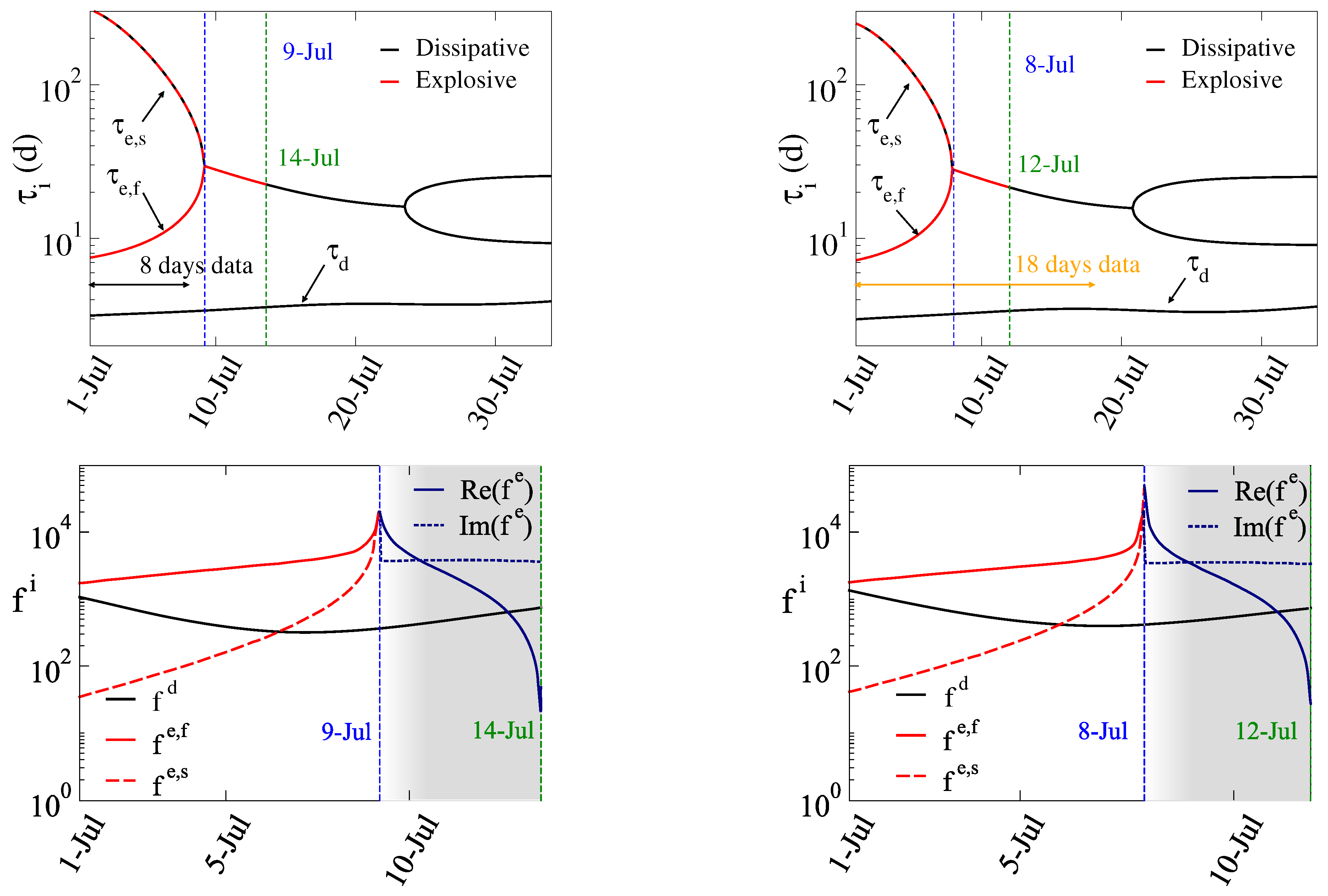
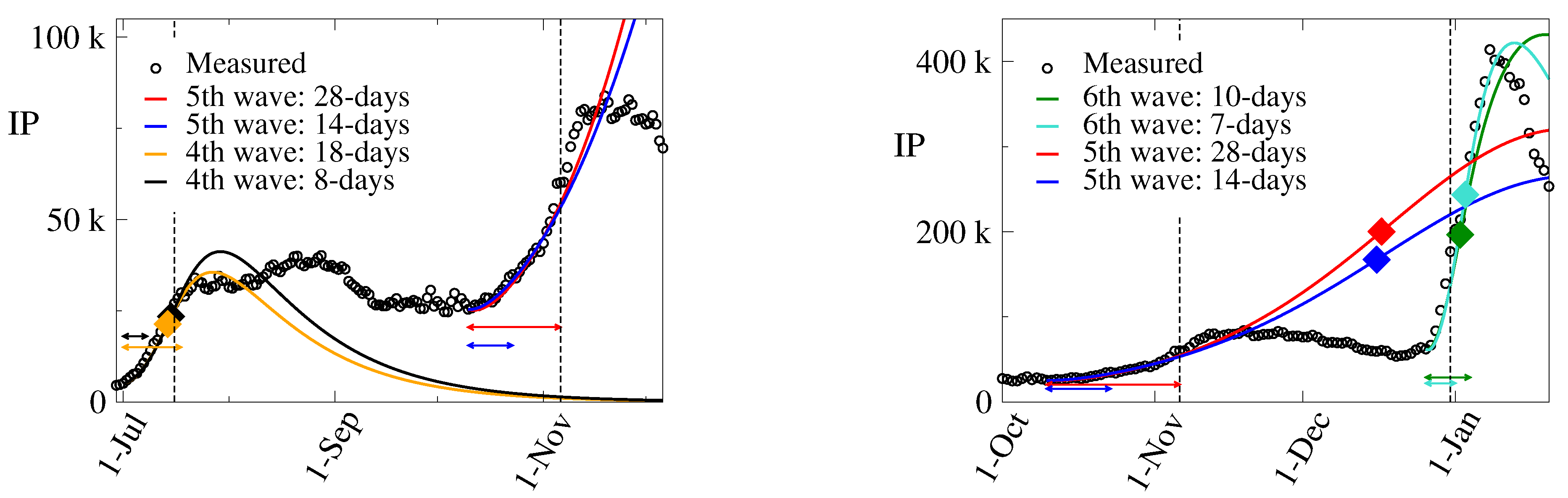
3.1. The 4th Wave
- i)
- Period A: July 1 – July 8 (8 days),
- ii)
- Period B: July 1 – July 18 (18 days),
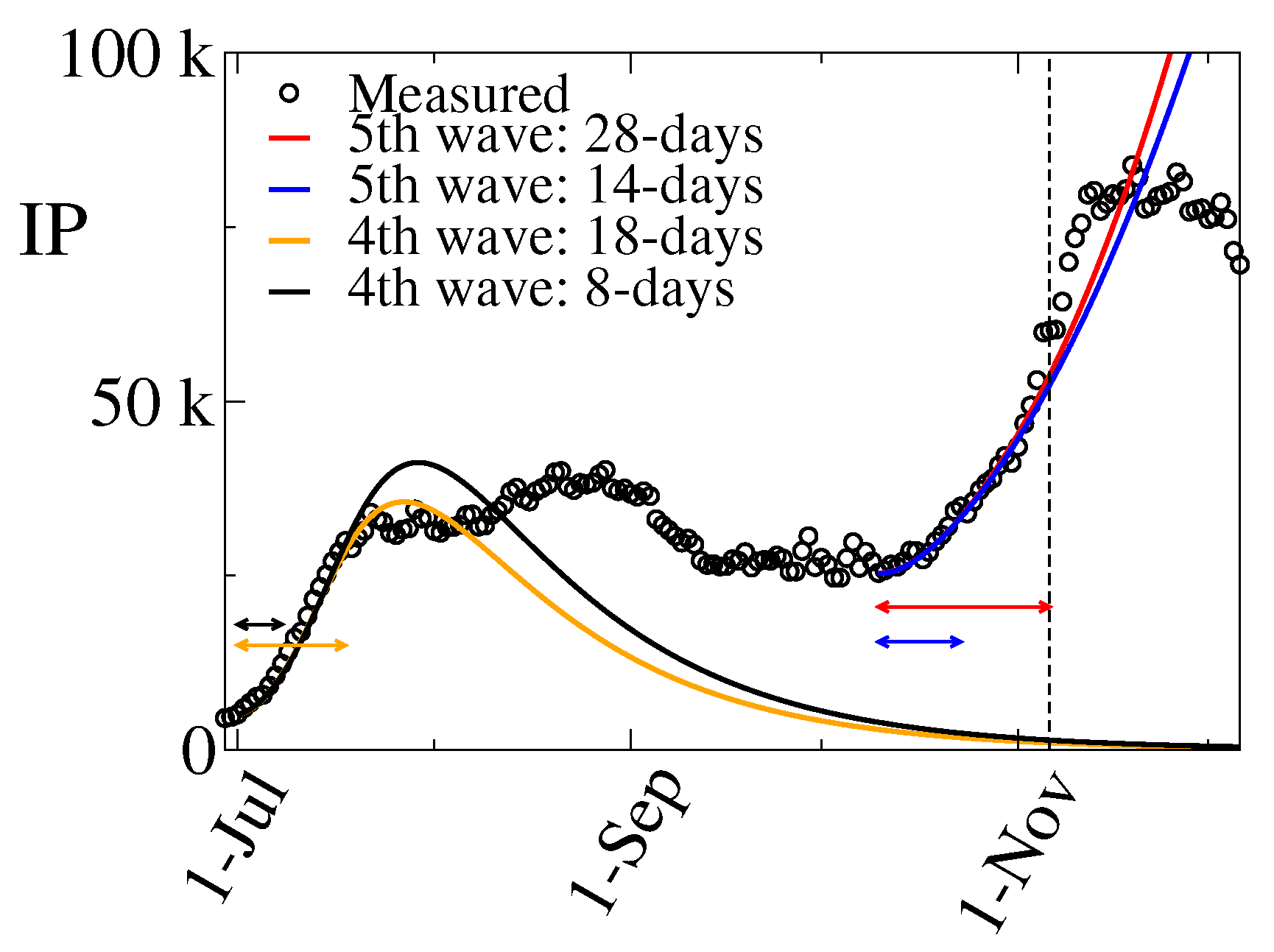
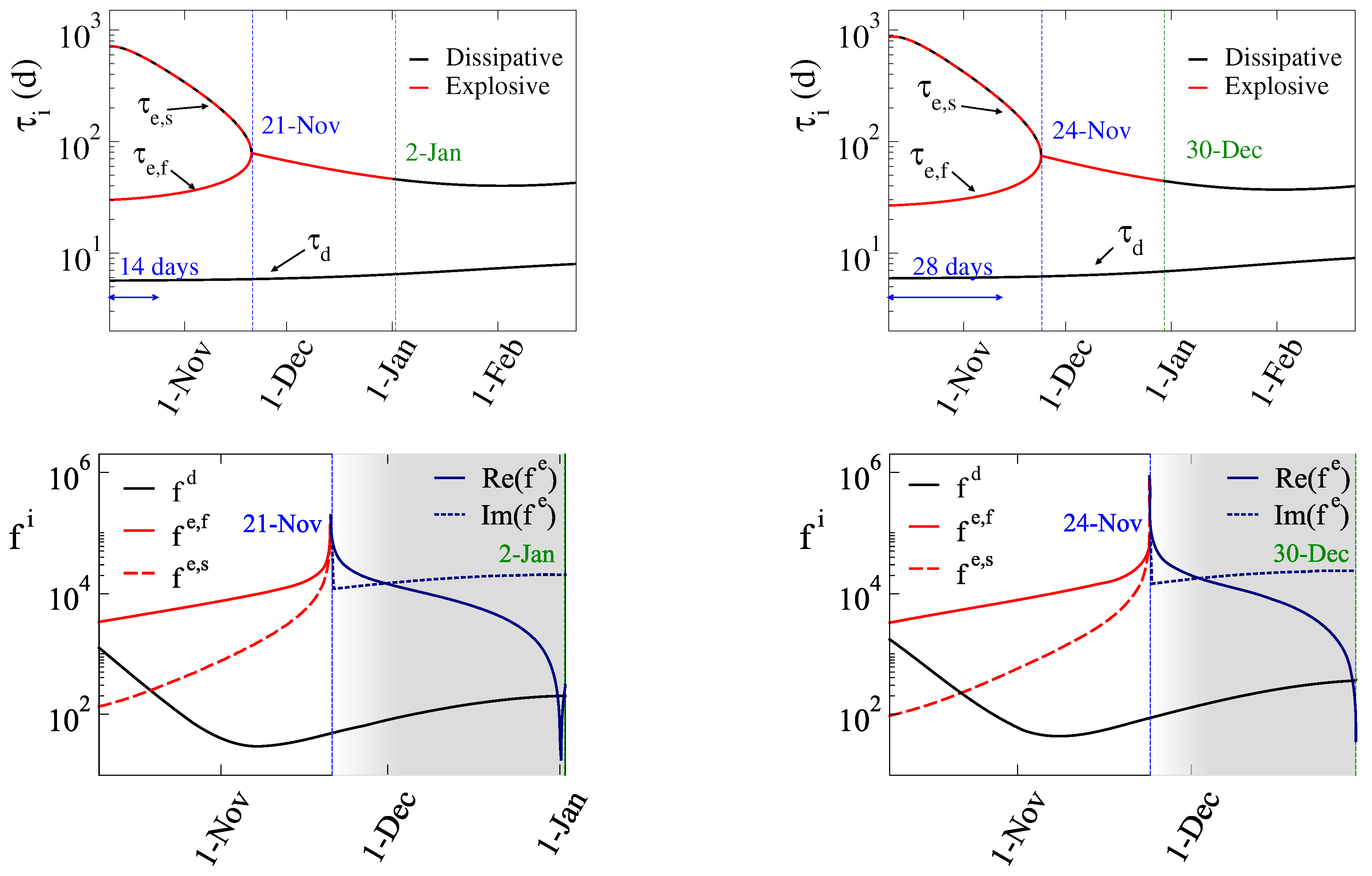
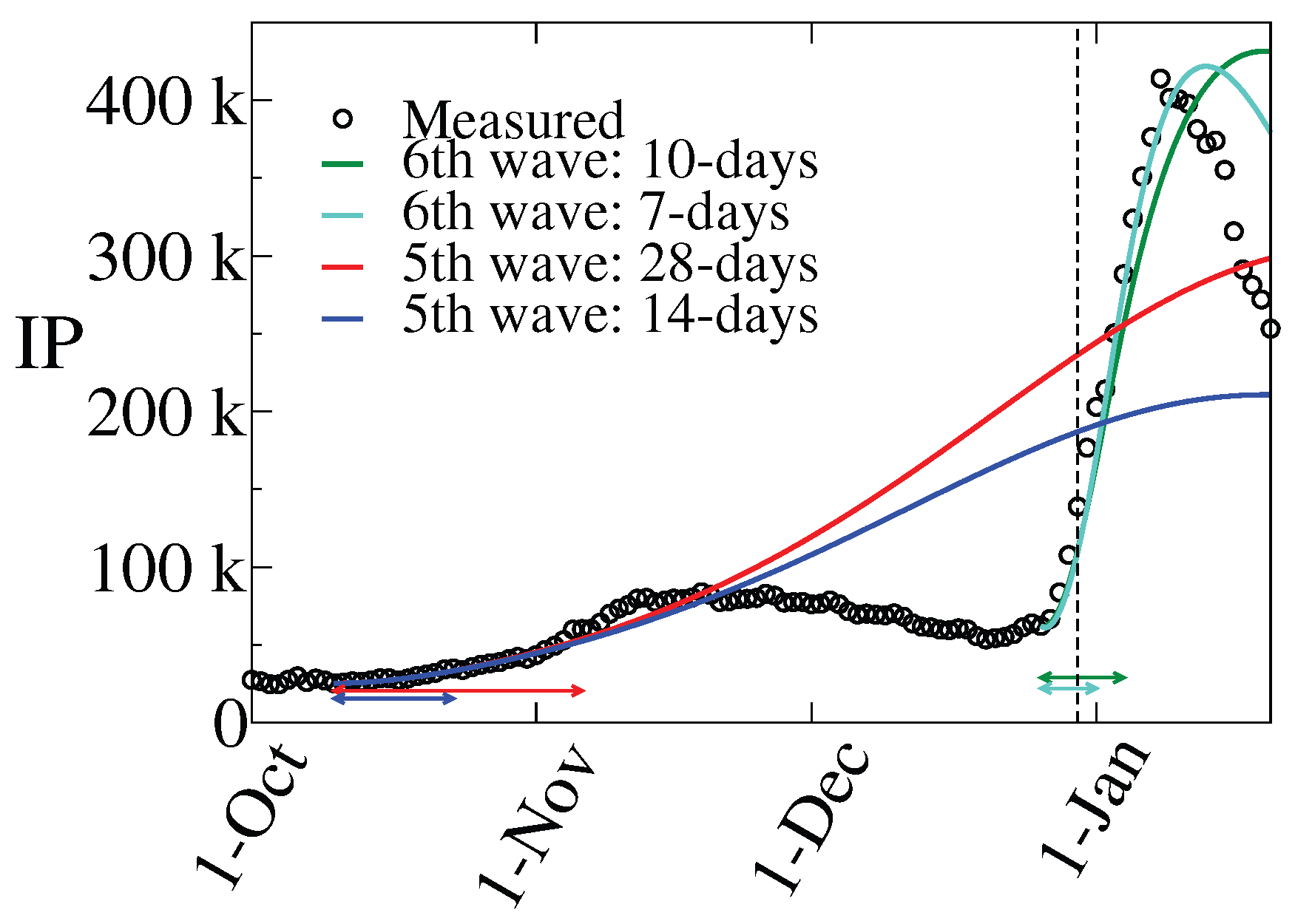
3.2. The 5th Wave
- i)
- Period C: October 10 – October 23 (14 days),
- ii)
- Period D: October 10 – November 6 (28 days),
3.3. The 6th Wave
- i)
- Period E: December 26 – January 1 (7 days),
- ii)
- Period F: December 26 – January 4 (10 days),
4. Physical Insights
5. Inflection Point for SP and Its Relation to the Explosive Dynamics
6. Conclusions
Author Contributions
Funding
Data Availability Statement
Acknowledgments
Conflicts of Interest
Abbreviations
| CSP | Computational Singular Perturbation method |
| TPI | Timescale Participation Index |
| Po | CSP Pointer |
| API | Amplitude Participation Index |
Appendix A. The Origin of the Fast Dissipative Mode
 |
References
- Röst, G.; Bartha, F.A.; Bogya, N.; Boldog, P.; Dénes, A.; Ferenci, T.; Horváth, K.J.; Juhász, A.; Nagy, C.; Tekeli, T.; et al. Early phase of the COVID-19 outbreak in Hungary and post-lockdown scenarios. Viruses 2020, 12, 708. [Google Scholar] [CrossRef]
- Nicolelis, M.A.; Raimundo, R.L.; Peixoto, P.S.; Andreazzi, C.S. The impact of super-spreader cities, highways, and intensive care availability in the early stages of the COVID-19 epidemic in Brazil. Sci. Rep. 2021, 11, 13001. [Google Scholar] [CrossRef]
- Velavan, T.P.; Meyer, C.G. The COVID-19 epidemic. Trop. Med. Int. Health 2020, 25, 278. [Google Scholar] [CrossRef]
- Flaxman, S.; Mishra, S.; Gandy, A.; Unwin, H.J.T.; Mellan, T.A.; Coupland, H.; Whittaker, C.; Zhu, H.; Berah, T.; Eaton, J.W.; et al. Estimating the effects of non-pharmaceutical interventions on COVID-19 in Europe. Nature 2020, 584, 257–261. [Google Scholar] [CrossRef]
- Barmparis, G.D.; Tsironis, G. Estimating the infection horizon of COVID-19 in eight countries with a data-driven approach. Chaos Solitons Fractals 2020, 135, 109842. [Google Scholar] [CrossRef]
- Peirlinck, M.; Linka, K.; Sahli Costabal, F.; Kuhl, E. Outbreak dynamics of COVID-19 in China and the United States. Biomech. Model. Mechanobiol. 2020, 19, 2179–2193. [Google Scholar] [CrossRef]
- Zheng, R.; Xu, Y.; Wang, W.; Ning, G.; Bi, Y. Spatial transmission of COVID-19 via public and private transportation in China. Travel Med. Infect. Dis. 2020, 34, 101626. [Google Scholar] [CrossRef] [PubMed]
- Odone, A.; Delmonte, D.; Scognamiglio, T.; Signorelli, C. COVID-19 deaths in Lombardy, Italy: data in context. Lancet Public Health 2020, 5, e310. [Google Scholar] [CrossRef] [PubMed]
- Prem, K.; Liu, Y.; Russell, T.W.; Kucharski, A.J.; Eggo, R.M.; Davies, N.; Flasche, S.; Clifford, S.; Pearson, C.A.; Munday, J.D.; et al. The effect of control strategies to reduce social mixing on outcomes of the COVID-19 epidemic in Wuhan, China: a modelling study. Lancet Public Health 2020. [Google Scholar] [CrossRef] [PubMed]
- Bayham, J.; Fenichel, E.P. Impact of school closures for COVID-19 on the US health-care workforce and net mortality: a modelling study. Lancet Public Health 2020, 5, e271–e278. [Google Scholar] [CrossRef]
- Colbourn, T. COVID-19: extending or relaxing distancing control measures. Lancet Public Health 2020, 5, e236–e237. [Google Scholar] [CrossRef]
- Martin, C.; Montesinos, I.; Dauby, N.; Gilles, C.; Dahma, H.; Van Den Wijngaert, S.; De Wit, S.; Delforge, M.; Clumeck, N.; Vandenberg, O. Dynamics of SARS-CoV-2 RT-PCR positivity and seroprevalence among high-risk healthcare workers and hospital staff. J. Hosp. Infect. 2020, 106, 102–106. [Google Scholar] [CrossRef]
- Abbas, M.; Robalo Nunes, T.; Martischang, R.; Zingg, W.; Iten, A.; Pittet, D.; Harbarth, S. Nosocomial transmission and outbreaks of coronavirus disease 2019: the need to protect both patients and healthcare workers. Antimicrob. Resist. Infect. Control 2021, 10, 1–13. [Google Scholar] [CrossRef]
- Lumley, S.F.; Wei, J.; O’Donnell, D.; Stoesser, N.E.; Matthews, P.C.; Howarth, A.; Hatch, S.B.; Marsden, B.D.; Cox, S.; James, T.; et al. The duration, dynamics, and determinants of severe acute respiratory syndrome coronavirus 2 (SARS-CoV-2) antibody responses in individual healthcare workers. Clin. Infect. Dis. 2021, 73, e699–e709. [Google Scholar] [CrossRef]
- Fokas, A.S.; Cuevas-Maraver, J.; Kevrekidis, P.G. Easing COVID-19 lockdown measures while protecting the older restricts the deaths to the level of the full lockdown. Sci. Rep. 2021, 11, 1–12. [Google Scholar] [CrossRef]
- Thompson, R.N.; Hollingsworth, T.D.; Isham, V.; Arribas-Bel, D.; Ashby, B.; Britton, T.; Challenor, P.; Chappell, L.H.; Clapham, H.; Cunniffe, N.J.; et al. Key questions for modelling COVID-19 exit strategies. Proc. R. Soc. B 2020, 287, 20201405. [Google Scholar] [CrossRef]
- Ruktanonchai, N.W.; Floyd, J.; Lai, S.; Ruktanonchai, C.W.; Sadilek, A.; Rente-Lourenco, P.; Ben, X.; Carioli, A.; Gwinn, J.; Steele, J.; et al. Assessing the impact of coordinated COVID-19 exit strategies across Europe. Science 2020, 369, 1465–1470. [Google Scholar] [CrossRef] [PubMed]
- Russo, L.; Anastassopoulou, C.; Tsakris, A.; Bifulco, G.N.; Campana, E.F.; Toraldo, G.; Siettos, C. Tracing day-zero and forecasting the COVID-19 outbreak in Lombardy, Italy: A compartmental modelling and numerical optimization approach. PLoS One 2020, 15, e0240649. [Google Scholar] [CrossRef] [PubMed]
- Kuniya, T. Prediction of the epidemic peak of coronavirus disease in Japan, 2020. J. Clin. Med. 2020, 9, 789. [Google Scholar] [CrossRef]
- Ma, J. Estimating epidemic exponential growth rate and basic reproduction number. Infect. Dis. Model. 2020, 5, 129–141. [Google Scholar] [CrossRef] [PubMed]
- Kucharski, A.J.; Russell, T.W.; Diamond, C.; Liu, Y.; Edmunds, J.; Funk, S.; Eggo, R.M.; Sun, F.; Jit, M.; Munday, J.D.; et al. Early dynamics of transmission and control of COVID-19: a mathematical modelling study. Lancet Infect. Dis. 2020. [Google Scholar] [CrossRef] [PubMed]
- Giordano, G.; Blanchini, F.; Bruno, R.; Colaneri, P.; Di Filippo, A.; Di Matteo, A.; Colaneri, M. Modelling the COVID-19 epidemic and implementation of population-wide interventions in Italy. Nat. Med. 2020, 1–6. [Google Scholar] [CrossRef] [PubMed]
- Tang, B.; Wang, X.; Li, Q.; Bragazzi, N.L.; Tang, S.; Xiao, Y.; Wu, J. Estimation of the transmission risk of the 2019-nCoV and its implication for public health interventions. J. Clin. Med. 2020, 9, 462. [Google Scholar] [CrossRef]
- Anastassopoulou, C.; Russo, L.; Tsakris, A.; Siettos, C. Data-based analysis, modelling and forecasting of the COVID-19 outbreak. PLoS One 2020, 15, e0230405. [Google Scholar] [CrossRef] [PubMed]
- Silva, P.C.; Batista, P.V.; Lima, H.S.; Alves, M.A.; Guimarães, F.G.; Silva, R.C. COVID-ABS: An agent-based model of COVID-19 epidemic to simulate health and economic effects of social distancing interventions. Chaos Solitons Fractals 2020, 139, 110088. [Google Scholar] [CrossRef]
- Kerr, C.C.; Stuart, R.M.; Mistry, D.; Abeysuriya, R.G.; Rosenfeld, K.; Hart, G.R.; Núñez, R.C.; Cohen, J.A.; Selvaraj, P.; Hagedorn, B.; et al. Covasim: an agent-based model of COVID-19 dynamics and interventions. PLoS Comput. Biol. 2021, 17, e1009149. [Google Scholar] [CrossRef]
- Adiga, A.; Dubhashi, D.; Lewis, B.; Marathe, M.; Venkatramanan, S.; Vullikanti, A. Mathematical models for covid-19 pandemic: a comparative analysis. J. Indian Inst. Sci. 2020, 100, 793–807. [Google Scholar] [CrossRef]
- Calvetti, D.; Hoover, A.P.; Rose, J.; Somersalo, E. Metapopulation network models for understanding, predicting, and managing the coronavirus disease COVID-19. Front. Phys. 2020, 8, 261. [Google Scholar] [CrossRef]
- Arenas, A.; Cota, W.; Gómez-Gardeñes, J.; Gómez, S.; Granell, C.; Matamalas, J.T.; Soriano-Paños, D.; Steinegger, B. Modeling the spatiotemporal epidemic spreading of COVID-19 and the impact of mobility and social distancing interventions. Phys. Rev. X 2020, 10, 041055. [Google Scholar] [CrossRef]
- Yang, Z.; Zeng, Z.; Wang, K.; Wong, S.S.; Liang, W.; Zanin, M.; Liu, P.; Cao, X.; Gao, Z.; Mai, Z.; et al. Modified SEIR and AI prediction of the epidemics trend of COVID-19 in China under public health interventions. J. Thorac. Dis. 2020, 12, 165. [Google Scholar] [CrossRef]
- Muhammad, L.; Algehyne, E.A.; Usman, S.S.; Ahmad, A.; Chakraborty, C.; Mohammed, I.A. Supervised machine learning models for prediction of COVID-19 infection using epidemiology dataset. SN Comput. Sci. 2021, 2, 1–13. [Google Scholar] [CrossRef]
- Fokas, A.; Dikaios, N.; Kastis, G. Mathematical models and deep learning for predicting the number of individuals reported to be infected with SARS-CoV-2. J. R. Soc. Interface 2020, 17, 20200494. [Google Scholar] [CrossRef]
- Fokas, A.S.; Dikaios, N.; Tsiodras, S.; Kastis, G.A. Simple formulae, deep learning and elaborate modelling for the COVID-19 pandemic. Encyclopedia 2022, 2, 679–689. [Google Scholar] [CrossRef]
- Yang, Z.; Bogdan, P.; Nazarian, S. An in silico deep learning approach to multi-epitope vaccine design: a SARS-CoV-2 case study. Sci. Rep. 2021, 11, 3238. [Google Scholar] [CrossRef] [PubMed]
- Zhang, J.; Litvinova, M.; Wang, W.; Wang, Y.; Deng, X.; Chen, X.; Li, M.; Zheng, W.; Yi, L.; Chen, X.; et al. Evolving epidemiology and transmission dynamics of coronavirus disease 2019 outside Hubei province, China: a descriptive and modelling study. Lancet Infect. Dis. 2020, 20, 793–802. [Google Scholar] [CrossRef]
- Chu, D.K.; Akl, E.A.; Duda, S.; Solo, K.; Yaacoub, S.; Schünemann, H.J.; El-Harakeh, A.; Bognanni, A.; Lotfi, T.; Loeb, M.; et al. Physical distancing, face masks, and eye protection to prevent person-to-person transmission of SARS-CoV-2 and COVID-19: a systematic review and meta-analysis. Lancet 2020, 395, 1973–1987. [Google Scholar] [CrossRef] [PubMed]
- Kontis, V.; Bennett, J.E.; Rashid, T.; Parks, R.M.; Pearson-Stuttard, J.; Guillot, M.; Asaria, P.; Zhou, B.; Battaglini, M.; Corsetti, G.; et al. Magnitude, demographics and dynamics of the effect of the first wave of the COVID-19 pandemic on all-cause mortality in 21 industrialized countries. Nat. Med. 2020, 26, 1919–1928. [Google Scholar] [CrossRef]
- Jewell, N.P.; Lewnard, J.A.; Jewell, B.L. Predictive mathematical models of the COVID-19 pandemic: Underlying principles and value of projections. JAMA 2020, 323, 1893–1894. [Google Scholar] [CrossRef] [PubMed]
- Poletto, C.; Scarpino, S.V.; Volz, E.M. Applications of predictive modelling early in the COVID-19 epidemic. Lancet Digit. Health 2020, 2, e498–e499. [Google Scholar] [CrossRef]
- Dehning, J.; Zierenberg, J.; Spitzner, F.P.; Wibral, M.; Neto, J.P.; Wilczek, M.; Priesemann, V. Inferring change points in the spread of COVID-19 reveals the effectiveness of interventions. Science 2020, 369, eabb9789. [Google Scholar] [CrossRef]
- Blackwood, J.C.; Childs, L.M. An introduction to compartmental modeling for the budding infectious disease modeler. Lett. Biomath. 2018, 5, 195–221. [Google Scholar] [CrossRef]
- Rella, S.A.; Kulikova, Y.A.; Dermitzakis, E.T.; Kondrashov, F.A. Rates of SARS-CoV-2 transmission and vaccination impact the fate of vaccine-resistant strains. Sci. Rep. 2021, 11, 15729. [Google Scholar] [CrossRef] [PubMed]
- Chowell, G.; Sattenspiel, L.; Bansal, S.; Viboud, C. Mathematical models to characterize early epidemic growth: A review. Phys. Life Rev. 2016, 18, 66–97. [Google Scholar] [CrossRef] [PubMed]
- Ivorra, B.; Ferrández, M.R.; Vela-Pérez, M.; Ramos, A.M. Mathematical modeling of the spread of the coronavirus disease 2019 (COVID-19) taking into account the undetected infections. The case of China. Commun.Nonlinear Sci. Numer. Simul. 2020, 88, 105303. [Google Scholar] [CrossRef]
- Arons, M.M.; Hatfield, K.M.; Reddy, S.C.; Kimball, A.; James, A.; Jacobs, J.R.; Taylor, J.; Spicer, K.; Bardossy, A.C.; Oakley, L.P.; et al. Presymptomatic SARS-CoV-2 infections and transmission in a skilled nursing facility. N. Engl. J. Med. 2020, 382, 2081–2090. [Google Scholar] [CrossRef]
- Demers, J.; Bewick, S.; Agusto, F.; Caillouët, K.A.; Fagan, W.F.; Robertson, S.L. Managing disease outbreaks: The importance of vector mobility and spatially heterogeneous control. PLoS Comput. Biol. 2020, 16, e1008136. [Google Scholar] [CrossRef]
- Lauer, S.A.; Grantz, K.H.; Bi, Q.; Jones, F.K.; Zheng, Q.; Meredith, H.R.; Azman, A.S.; Reich, N.G.; Lessler, J. The incubation period of coronavirus disease 2019 (COVID-19) from publicly reported confirmed cases: estimation and application. Ann. Intern. Med. 2020, 172, 577–582. [Google Scholar] [CrossRef] [PubMed]
- Li, Q.; Guan, X.; Wu, P.; Wang, X.; Zhou, L.; Tong, Y.; Ren, R.; Leung, K.S.; Lau, E.H.; Wong, J.Y.; et al. Early transmission dynamics in Wuhan, China, of novel coronavirus–infected pneumonia. N. Engl. J. Med. 2020. [Google Scholar] [CrossRef]
- Lin, Q.; Zhao, S.; Gao, D.; Lou, Y.; Yang, S.; Musa, S.S.; Wang, M.H.; Cai, Y.; Wang, W.; Yang, L.; et al. A conceptual model for the outbreak of Coronavirus disease 2019 (COVID-19) in Wuhan, China with individual reaction and governmental action. Int. J. Infect. Dis. 2020. [Google Scholar] [CrossRef]
- Maugeri, A.; Barchitta, M.; Battiato, S.; Agodi, A. Estimation of unreported novel coronavirus (SARS-CoV-2) infections from reported deaths: a susceptible–exposed–infectious–recovered–dead model. J. Clin. Med. 2020, 9, 1350. [Google Scholar] [CrossRef]
- Poonia, R.C.; Saudagar, A.K.J.; Altameem, A.; Alkhathami, M.; Khan, M.B.; Hasanat, M.H.A. An enhanced SEIR model for prediction of COVID-19 with vaccination effect. Life 2022, 12, 647. [Google Scholar] [CrossRef]
- Shams Eddin, M.; El Hajj, H.; Zayyat, R.; Lee, G. Systematic comparison of different compartmental models for predicting COVID-19 progression. Epidemiologia 2025, 6, 33. [Google Scholar] [CrossRef]
- Bastos, S.B.; Cajueiro, D.O. Modeling and forecasting the early evolution of the Covid-19 pandemic in Brazil. Sci. Rep. 2020, 10, 19457. [Google Scholar] [CrossRef]
- Tuncer, N.; Le, T.T. Structural and practical identifiability analysis of outbreak models. Math. Biosci. 2018, 299, 1–18. [Google Scholar] [CrossRef]
- Lintusaari, J.; Gutmann, M.U.; Kaski, S.; Corander, J. On the identifiability of transmission dynamic models for infectious diseases. Genetics 2016, 202, 911–918. [Google Scholar] [CrossRef]
- Raue, A.; Kreutz, C.; Maiwald, T.; Bachmann, J.; Schilling, M.; Klingmüller, U.; Timmer, J. Structural and practical identifiability analysis of partially observed dynamical models by exploiting the profile likelihood. Bioinformatics 2009, 25, 1923–1929. [Google Scholar] [CrossRef]
- Roda, W.C.; Varughese, M.B.; Han, D.; Li, M.Y. Why is it difficult to accurately predict the COVID-19 epidemic? Infect. Dis. Model. 2020, 5, 271–281. [Google Scholar] [CrossRef] [PubMed]
- Lee, C.; Li, Y.; Kim, J. The susceptible-unidentified infected-confirmed (SUC) epidemic model for estimating unidentified infected population for COVID-19. Chaos Solitons Fractals 2020, 139, 110090. [Google Scholar] [CrossRef]
- Fang, Y.; Nie, Y.; Penny, M. Transmission dynamics of the COVID-19 outbreak and effectiveness of government interventions: A data-driven analysis. J. Med. Virol. 2020, 92, 645–659. [Google Scholar] [CrossRef] [PubMed]
- Bertozzi, A.L.; Franco, E.; Mohler, G.; Short, M.B.; Sledge, D. The challenges of modeling and forecasting the spread of COVID-19. Proc. Natl. Acad. Sci. U.S.A. 2020, 117, 16732–16738. [Google Scholar] [CrossRef] [PubMed]
- Kheifetz, Y.; Kirsten, H.; Scholz, M. On the Parametrization of Epidemiologic Models—Lessons from Modelling COVID-19 Epidemic. Viruses 2022, 14, 1468. [Google Scholar] [CrossRef]
- Hu, W.H.; Sun, H.M.; Wei, Y.Y.; Hao, Y.T. Global infectious disease early warning models: An updated review and lessons from the COVID-19 pandemic. Infect. Dis. Model. 2024. [Google Scholar] [CrossRef] [PubMed]
- Looker, J.; Rock, K.S.; Dyson, L. Identifying COVID-19 peaks using early warning signals. PLoS Comput. Biol. 2025, 21, e1013524. [Google Scholar] [CrossRef]
- Chowell, G.; Echevarría-Zuno, S.; Viboud, C.; Simonsen, L.; Tamerius, J.; Miller, M.A.; Borja-Aburto, V.H. Characterizing the epidemiology of the 2009 influenza A/H1N1 pandemic in Mexico. PLoS Med 2011, 8, e1000436. [Google Scholar] [CrossRef] [PubMed]
- Park, S.W.; Champredon, D.; Weitz, J.S.; Dushoff, J. A practical generation-interval-based approach to inferring the strength of epidemics from their speed. Epidemics 2019, 27, 12–18. [Google Scholar] [CrossRef] [PubMed]
- Cao, Q.; Zheng, J.; Liu, Y.; Li, Z.; Xu, J.; Zhao, H.; Feng, L.; Jin, Y. Infectious disease prediction model based on optimized deep learning algorithm. Front. Public Health 2025, 13, 1703506. [Google Scholar] [CrossRef]
- Tretter, F.; Peters, E.M.; Sturmberg, J.; Bennett, J.; Voit, E.; Dietrich, J.W.; Smith, G.; Weckwerth, W.; Grossman, Z.; Wolkenhauer, O.; et al. Perspectives of (/memorandum for) systems thinking on COVID-19 pandemic and pathology. J. Eval. Clin. Pract. 2023, 29, 415–429. [Google Scholar] [CrossRef]
- Lam, S.H.; Goussis, D.A. Understanding complex chemical kinetics with computational singular perturbation. In Proceedings of the Proc. Combust. Inst. Elsevier; 1989; Vol. 22, pp. 931–941. [Google Scholar] [CrossRef]
- Hadjinicolaou, M.; Goussis, D.A. Asymptotic solution of stiff PDEs with the CSP method: the reaction diffusion equation. SIAM J. Sci. Comput. 1998, 20, 781–810. [Google Scholar] [CrossRef]
- Kuehn, C.; et al. Multiple time scale dynamics; Springer, 2015; Vol. 191. [Google Scholar]
- Patsatzis, D.G.; Goussis, D.A. A new Michaelis-Menten equation valid everywhere multi-scale dynamics prevails. Math. Biosci. 2019, 315, 108220. [Google Scholar] [CrossRef]
- Lam, S.H.; Goussis, D.A. The CSP method for simplifying kinetics. Int. J. Chem. Kinet. 1994, 26, 461–486. [Google Scholar] [CrossRef]
- Manias, D.M.; Tingas, E.A.; Frouzakis, C.E.; Boulouchos, K.; Goussis, D.A. The mechanism by which CH2O and H2O2 additives affect the autoignition of CH4/air mixtures. Combust. Flame 2016, 164, 111–125. [Google Scholar] [CrossRef]
- Patsatzis, D.G.; Wu, S.; Shah, D.K.; Goussis, D.A. Algorithmic multiscale analysis for the FcRn mediated regulation of antibody PK in human. Sci. Rep. 2022, 12, 6208. [Google Scholar] [CrossRef] [PubMed]
- Muhammed, I.; Manias, D.M.; Goussis, D.A.; Hatzikirou, H. Data-driven identification of biological systems using multi-scale analysis. PLoS Comput. Biol. 2025, 21, e1013193. [Google Scholar] [CrossRef] [PubMed]
- Patsatzis, D.G.; Maris, D.T.; Goussis, D.A. Asymptotic analysis of a target-mediated drug disposition model: algorithmic and traditional approaches. Bull. Math. Biol. 2016, 78, 1121–1161. [Google Scholar] [CrossRef] [PubMed]
- Michalaki, L.I.; Goussis, D.A. Asymptotic analysis of a TMDD model: when a reaction contributes to the destruction of its product. J. Math. Biol. 2018, 77, 821–855. [Google Scholar] [CrossRef]
- Patsatzis, D.G.; Tingas, E.A.; Goussis, D.A.; Sarathy, S.M. Computational singular perturbation analysis of brain lactate metabolism. PLoS One 2019, 14, e0226094. [Google Scholar] [CrossRef]
- Patsatzis, D.G.; Tingas, E.A.; Sarathy, S.M.; Goussis, D.A.; Jolivet, R.B. Elucidating reaction dynamics in a model of human brain energy metabolism. PLoS Comput. Biol. 2025, 21, e1013504. [Google Scholar] [CrossRef]
- Kourdis, P.D.; Steuer, R.; Goussis, D.A. Physical understanding of complex multiscale biochemical models via algorithmic simplification: Glycolysis in Saccharomyces cerevisiae. Physica D 2010, 239, 1798–1817. [Google Scholar] [CrossRef]
- Patsatzis, D.G. Algorithmic asymptotic analysis: Extending the arsenal of cancer immunology modeling. J. Theor. Biol. 2022, 534, 110975. [Google Scholar] [CrossRef]
- COVID-19 Data Repositoy by the Greek Government. Available online: https://covid19.gov.gr/covid19-live-analytics/ (accessed on 30 March 2026).
- Runarsson, T.P.; Yao, X. Stochastic ranking for constrained evolutionary optimization. IEEE Trans. Evol. Comput. 2000, 4, 284–294. [Google Scholar] [CrossRef]
- Hoops, S.; Sahle, S.; Gauges, R.; Lee, C.; Pahle, J.; Simus, N.; Singhal, M.; Xu, L.; Mendes, P.; Kummer, U. COPASI—a complex pathway simulator. Bioinformatics 2006, 22, 3067–3074. [Google Scholar] [CrossRef]
- Malli, F.; Lampropoulos, I.C.; Papagiannis, D.; Papathanasiou, I.V.; Daniil, Z.; Gourgoulianis, K.I. Association of SARS-CoV-2 Vaccinations with SARS-CoV-2 Infections, ICU Admissions and Deaths in Greece. Vaccines 2022, 10, 337. [Google Scholar] [CrossRef] [PubMed]
- Rabaan, A.; Al-Ahmed, S.; Albayat, H.; Alwarthan, S.; Alhajri, M.; Najim, M.; AlShehail, B.; Al-Adsani, W.; Alghadeer, A.; Abduljabbar, W.; et al. Variants of SARS-CoV-2: Influences on the Vaccines’ Effectiveness and Possible Strategies to Overcome Their Consequences. Medicina 2023, 59, 507. [Google Scholar] [CrossRef]
- Mohammed, H.; Pham-Tran, D.; Yeoh, Z.; Wang, B.; McMillan, M.; Andraweera, P.; Marshall, H. A Systematic Review and Meta-Analysis on the Real-World Effectiveness of COVID-19 Vaccines against Infection, Symptomatic and Severe COVID-19 Disease Caused by the Omicron Variant (B.1.1.529). Vaccines 2023, 11, 224. [Google Scholar] [CrossRef]
- Galani, A.; Markou, A.; Dimitrakopoulos, L.; Kontou, A.; Kostakis, M.; Kapes, V.; Diamantopoulos, M.A.; Adamopoulos, P.G.; Avgeris, M.; Lianidou, E.; et al. Delta SARS-CoV-2 variant is entirely substituted by the omicron variant during the fifth COVID-19 wave in Attica region. Sci. Total Environ. 2023, 856, 159062. [Google Scholar] [CrossRef] [PubMed]
- Diamantis, D.J.; Mastorakos, E.; Goussis, D.A. H2/air autoignition: the nature and interaction of the developing explosive modes. Combust. Theory Model. 2015, 19, 382–433. [Google Scholar] [CrossRef]
- Khalil, A.T.; Manias, D.M.; Kyritsis, D.C.; Goussis, D.A. NO Formation and Autoignition Dynamics during Combustion of H2O-Diluted NH3/H2O2 Mixtures with Air. Energies 2021, 14, 84. [Google Scholar] [CrossRef]
- Zagaris, A.; Kaper, H.G.; Kaper, T.J. Fast and slow dynamics for the computational singular perturbation method. Multiscale Model. Simul. 2004, 2, 613–638. [Google Scholar] [CrossRef]
- Kaper, H.G.; Kaper, T.J.; Zagaris, A. Geometry of the computational singular perturbation method. Math. Model. Nat. Phenom. 2015, 10, 16–30. [Google Scholar] [CrossRef]
- Goussis, D.; Lam, S. A study of homogeneous methanol oxidation kinetics using CSP. In Proceedings of the Proc. Combust. Inst. Elsevier; 1992; Vol. 24, pp. 113–120. [Google Scholar] [CrossRef]
- Manias, D.M.; Goldman, R.N.; Goussis, D.A. Physical insights from complex multiscale non-linear system dynamics: Identification of fast and slow variables. Commun.Nonlinear Sci. Numer. Simul. 2025, 148, 108858. [Google Scholar] [CrossRef]
- Goussis, D.A.; Najm, H.N. Model reduction and physical understanding of slowly oscillating processes: the circadian cycle. Multiscale Model. Simul. 2006, 5, 1297–1332. [Google Scholar] [CrossRef]
- Manias, D.M.; Diamantis, D.J.; Goussis, D.A. Algorithmic identification of the reactions related to the initial development of the time scale that characterizes CH 4/air autoignition. J. Energy Eng. 2015, 141, C4014015. [Google Scholar] [CrossRef]
- eKathimerini Greece 13-07-2021. Available online: https://www.kathimerini.gr/society/561431866/estiasi-kai-choroi-psychagogias-kalokairi-kathimenon-me-tsoychtera-prostima-kai-kleistoys-choroys-mono-gia-emvoliasmenoys (accessed on 30 March 2026).
- CNN Greece 06-11-2021. Available online: https://www.cnn.gr/ellada/story/288254/koronoios-nea-metra-gia-toys-anemvoliastoys-apo-simera-poioi-periorismoi-isxyoyn (accessed on 30 March 2026).
- eKathimerini Greece 29-12-2021. Available online: https://www.kathimerini.gr/society/561649636/nea-metra-live-oi-anakoinoseis-apo-ton-thano-pleyri (accessed on 30 March 2026).
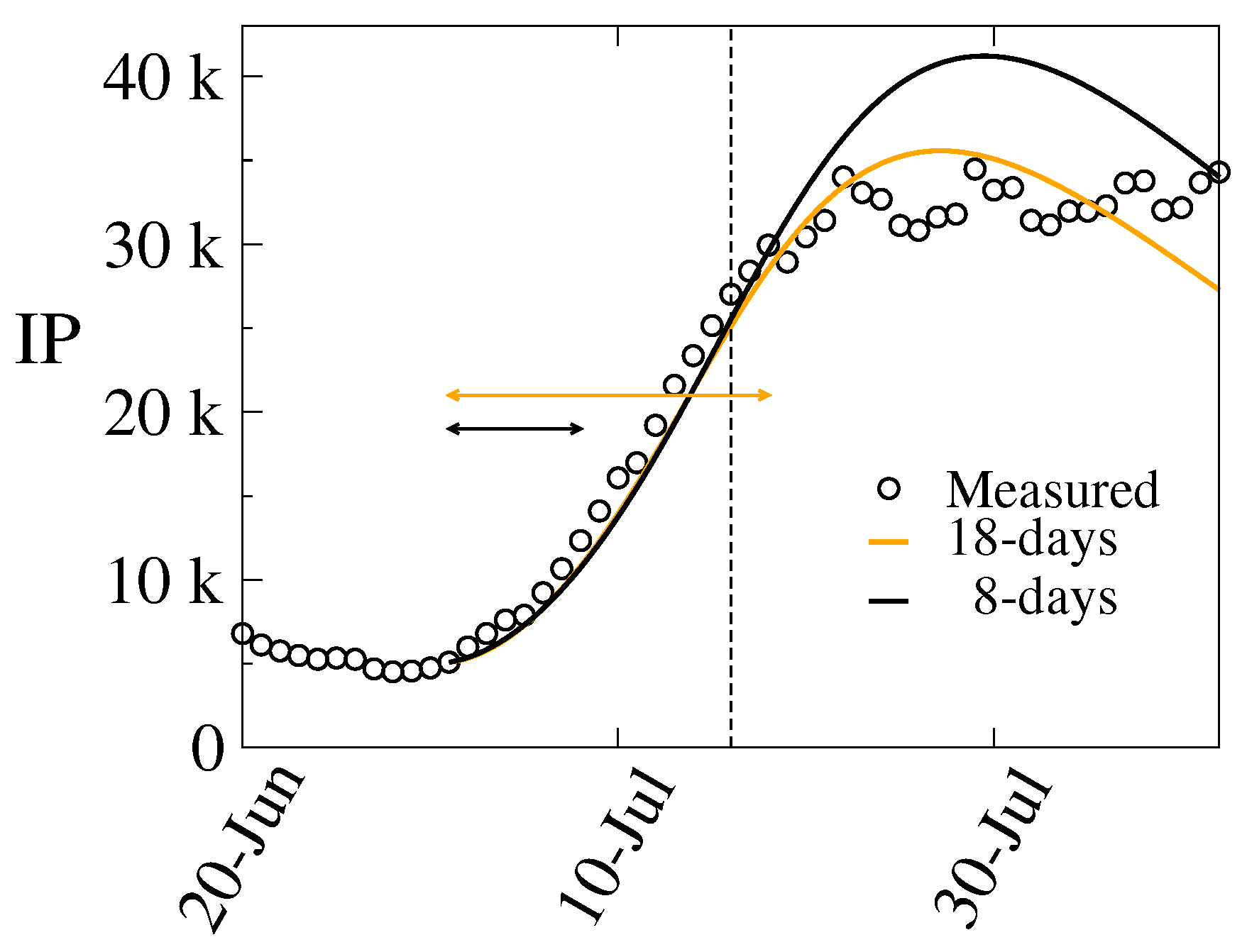
 |
| Period A | Period B | ||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|
| Jul-1 | Jul-7 | Jul-1 | Jul-7 | Jul-1 | Jul-7 | Jul-1 | Jul-7 | ||||
| 0.51 | 0.44 | 0.69 | 0.99 | 0.50 | 0.38 | 0.71 | 1.34 | ||||
| 0.35 | 0.31 | 0.49 | 0.62 | 0.36 | 0.31 | 0.51 | 0.80 | ||||
| -0.14 | -0.24 | -0.18 | -0.61 | -0.14 | -0.30 | -0.22 | -1.14 | ||||
| 0.00 | -0.01 | 0.00 | -0.01 | ||||||||
| Period C | Period D | ||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|
| Oct-10 | Nov-20 | Oct-10 | Nov-20 | Oct-10 | Nov-20 | Oct-10 | Nov-20 | ||||
| 0.49 | 0.43 | 0.55 | 1.90 | 0.49 | 0.43 | 0.54 | 0.92 | ||||
| -0.34 | -0.46 | 0.53 | 1.83 | -0.31 | -0.42 | 0.53 | 0.92 | ||||
| 0.16 | 0.10 | -0.08 | -2.73 | 0.19 | 0.14 | -0.07 | -0.84 | ||||
| -0.01 | -0.01 | -0.01 | -0.01 | ||||||||
| Period E | Period F | ||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|
| Dec-26 | Dec-29 | Dec-26 | Dec-29 | Dec-26 | Dec-29 | Dec-26 | Dec-29 | ||||
| 0.45 | 0.46 | 0.63 | 0.82 | 0.53 | 0.55 | 0.67 | 1.03 | ||||
| 0.41 | 0.32 | 0.63 | 0.82 | 0.37 | 0.22 | 0.68 | 1.03 | ||||
| -0.13 | -0.22 | -0.26 | -0.64 | -0.10 | -0.23 | -0.33 | -1.06 | ||||
| -0.01 | 0.00 | 0.00 | 0.00 | ||||||||
 |
| Wave 4 (Period A) | Wave 5 (Period C) | Wave 6 (Period E) |
|---|---|---|
| : 51.0 − 43.6% | : 49.0 − 42.6% | : 46.2 − 45.6% |
| : 34.8 − 32.3% | : -34.0 − -46.4 % | : 40.6 − 32.1% |
| : -13.9 −-23.5% | : 16.3 − 10.2% | : -13.1 − -22.0% |
| IP: 68.7 − 99.0% | IP: 54.9 − 190.1% | IP: 62.8 − 82.3% |
| EP: 48.6 − 62.4% | EP: 53.1 − 183.2% | EP: 62.8 − 82.1% |
| SP: -17.3 − -61.4% | SP: -8.0 − -273.8% | SP: -25.6 − -64.4% |
| Wave | Period | RM | SP inflection points | ||
|---|---|---|---|---|---|
| 4 | A | (Jul-1 to Jul-8) | Jul-16 | Jul-14 | Jul-12 |
| B | (Jul-1 to Jul-18) | Jul-16 | Jul-12 | Jul-10 | |
| 5 | C | (Oct-10 to Oct-23) | Nov-6 | Jan-2 | Jan-1 |
| D | (Oct-10 to Nov-6) | Nov-6 | Dec-30 | Dec-28 | |
| 6 | E | (Dec-26 to Jan-1) | Dec-30 | Jan-1 | Dec-31 |
| F | (Dec-26 to Jan-4) | Dec-30 | Dec-31 | Dec-30 | |
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2026 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).




