Submitted:
01 April 2026
Posted:
02 April 2026
You are already at the latest version
Abstract
Keywords:
1. Introduction
2. Materials and Methods
2.1. Cell Lines and Reagents
2.2. Molecular Modeling
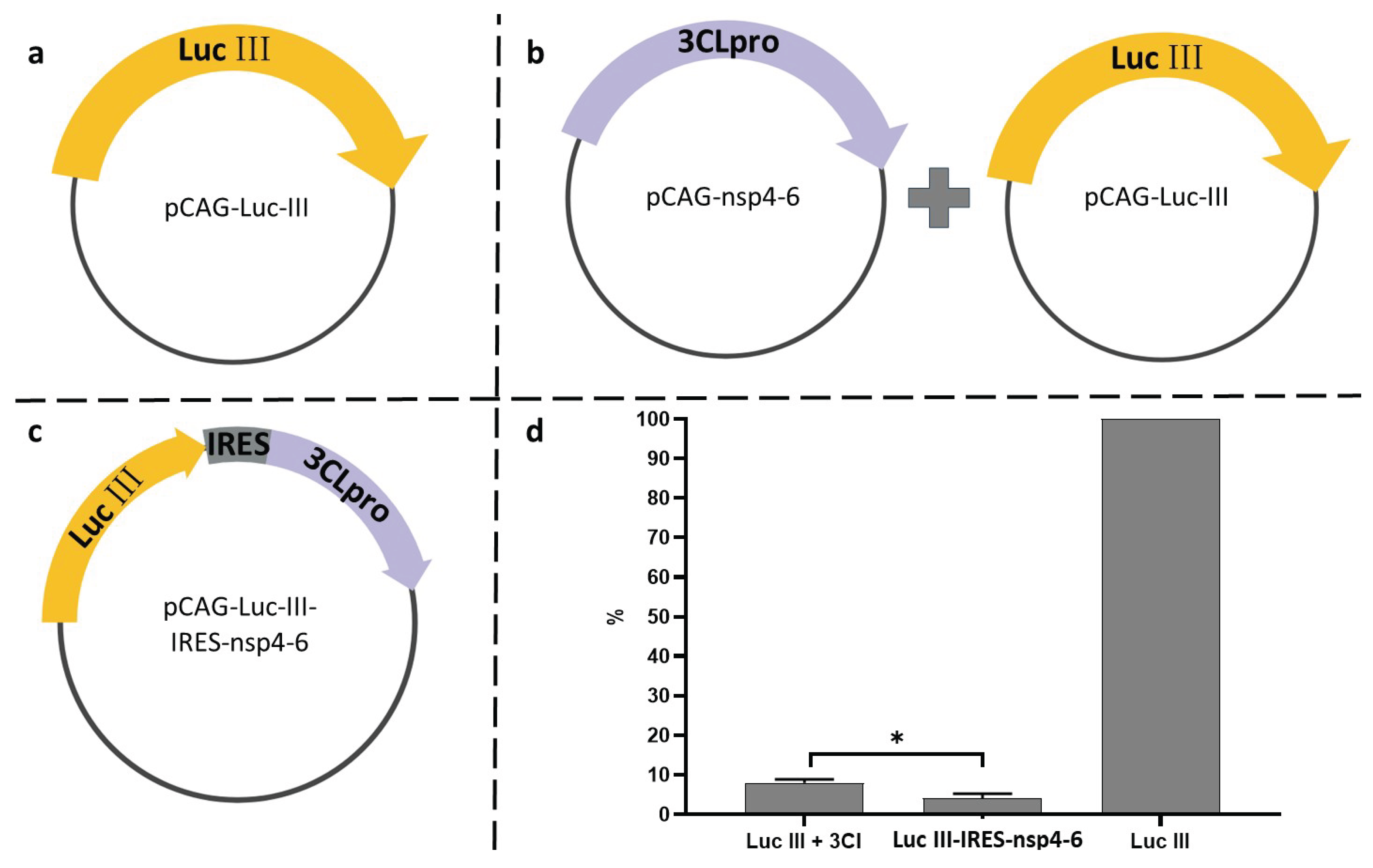
2.3. Plasmid Vector Construction
2.4. Transfections and Luciferase Assays
2.5. Evaluation of Inhibitor Activity in the Developed Cell-Based System
2.6. Evaluation of the Antiviral Activities against SARS-CoV-2 Viruses
2.7. Assessment of Inhibitor Activity on a Fluorescent Substrate
2.8. Statistical Analysis
3. Results
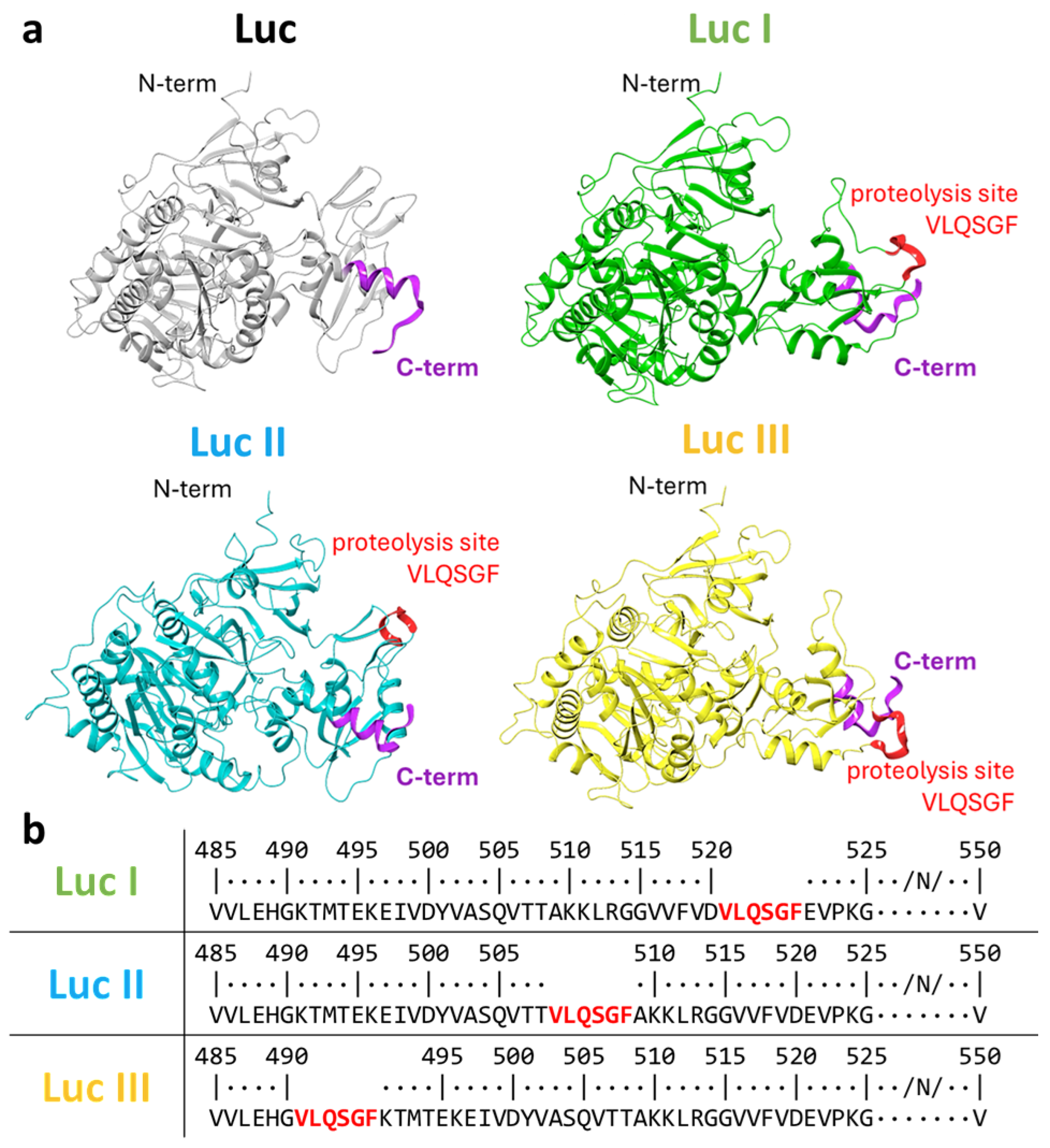
3.1. Selection of the Insertion Site in Luc for the SARS-CoV-2 3CLpro Cleavage Site
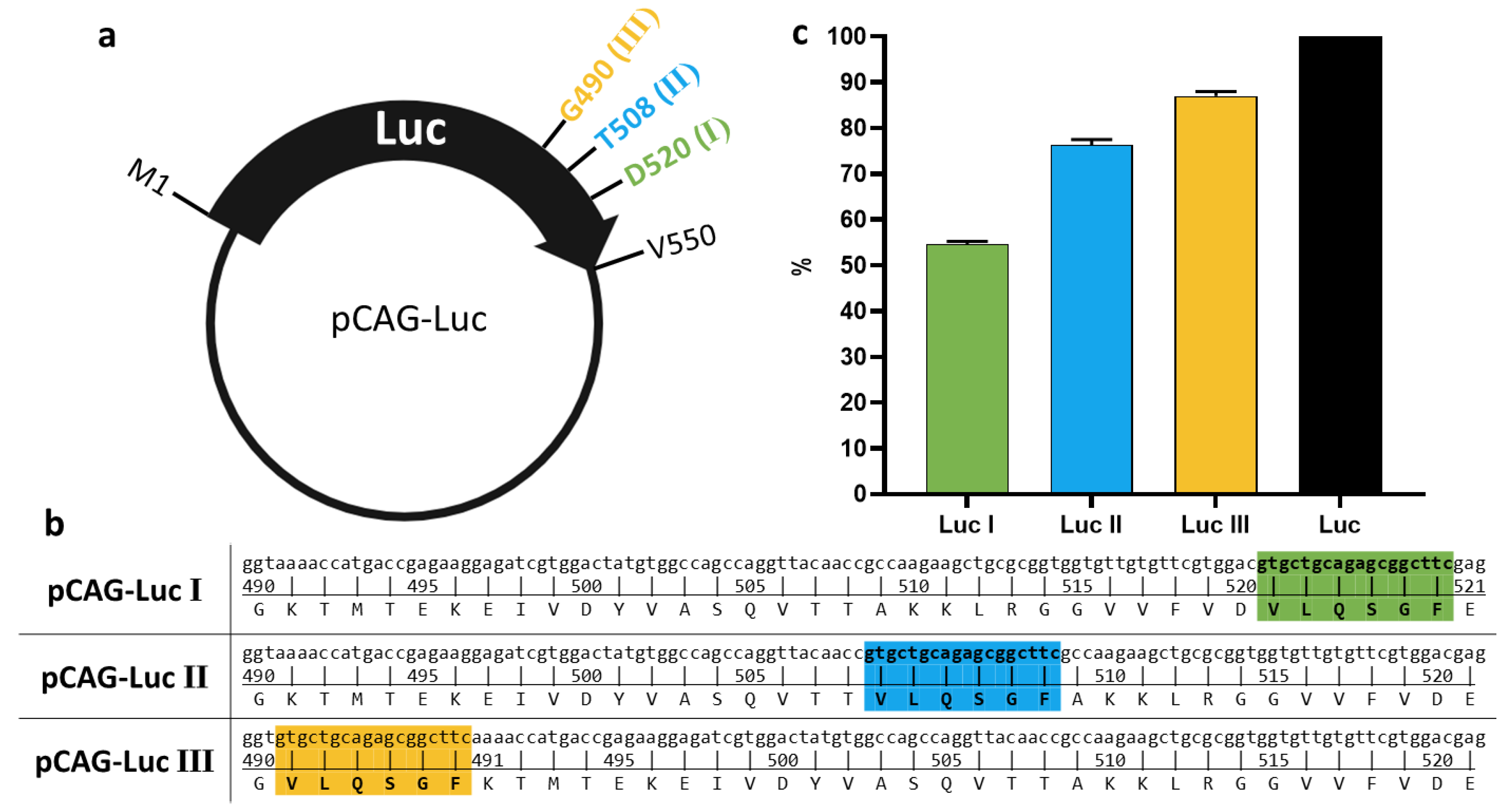
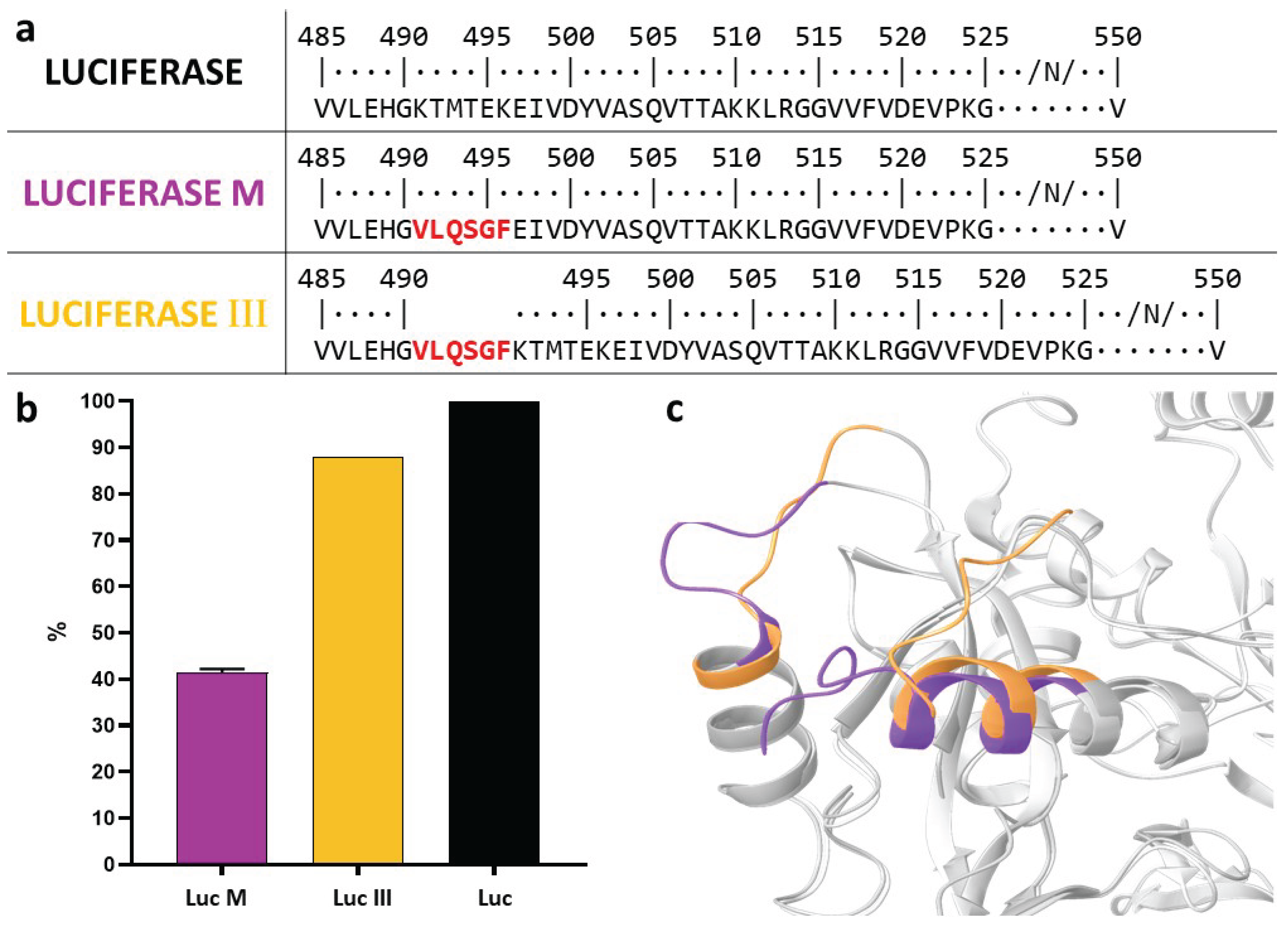
3.2. Assessment of Luc Variant Activity Containing the Cleavage Site
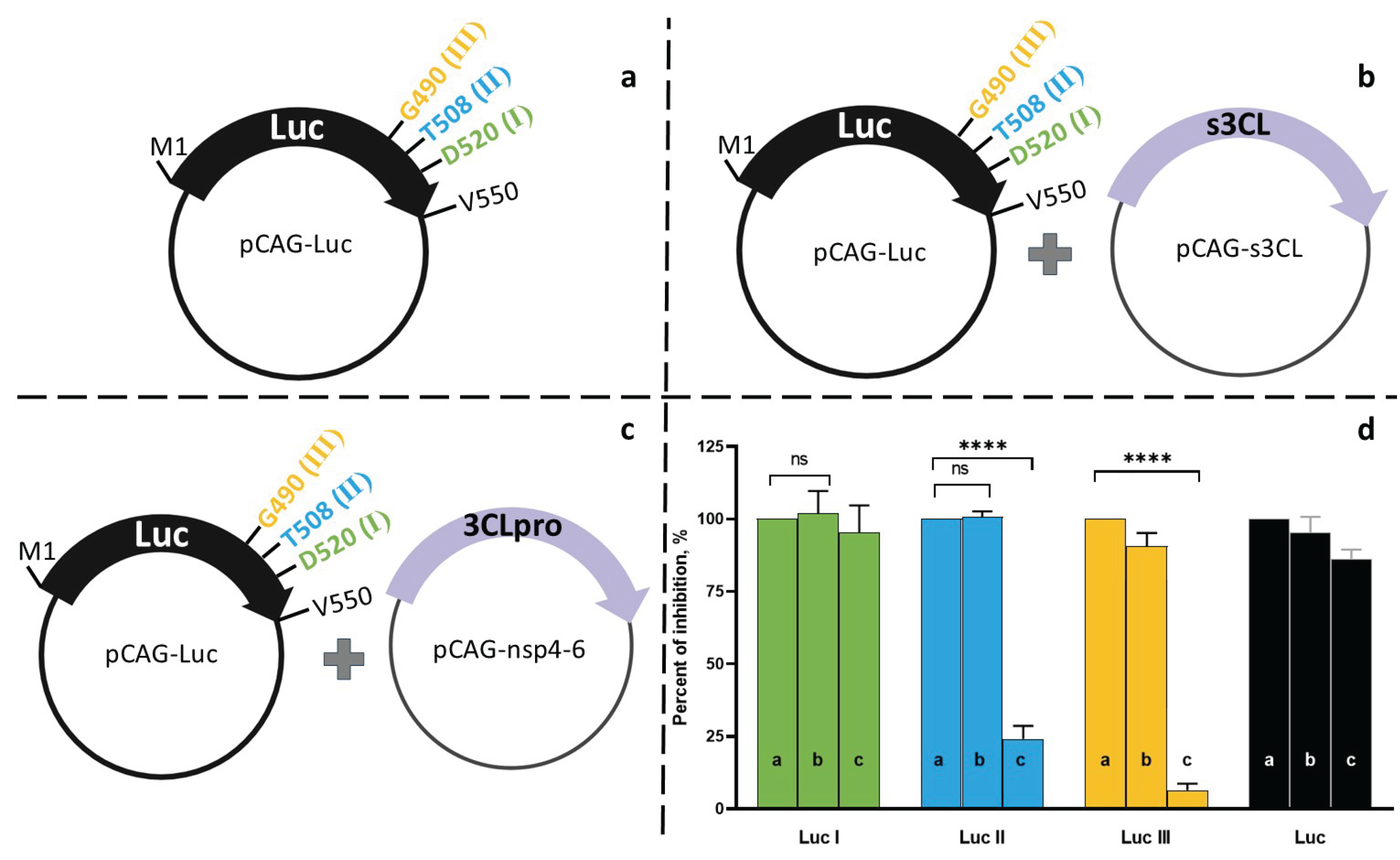
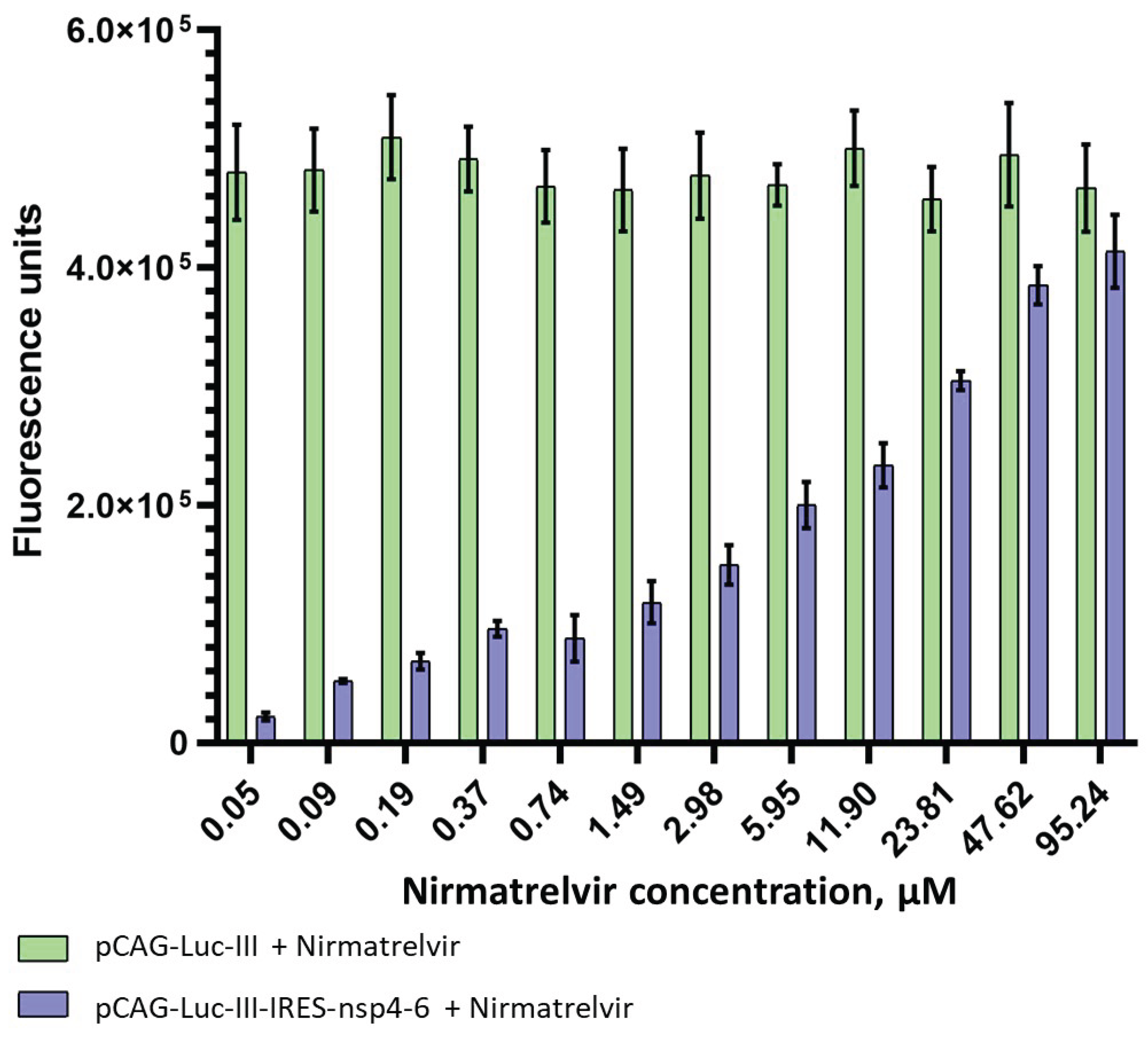
3.3. Validation of the System using 3CLpro Inhibitors
4. Discussion
5. Conclusions
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Conflicts of Interest
References
- Tong, L. Viral Proteases. Chem. Rev. 2002, 102, 4609–4626. [Google Scholar] [CrossRef]
- Swanstrom, R.; Anderson, J.; Schiffer, C.; Lee, S.K. Viral Protease Inhibitors. Handb. Exp. Pharmacol. 2009, 189, 85–110. [Google Scholar] [CrossRef]
- Babé, L.M.; Craik, C.S. Viral Proteases: Evolution of Diverse Structural Motifs to Optimize Function. Cell 1997, 91, 427–430. [Google Scholar] [CrossRef]
- Sittampalam, G.S.; Kahl, S.D.; Janzen, W.P. High-Throughput Screening: Advances in Assay Technologies. Curr. Opin. Chem. Biol. 1997, 1, 384–391. [Google Scholar] [CrossRef]
- Blay, V.; Tolani, B.; Ho, S.P.; Arkin, M.R. High-Throughput Screening: Today’s Biochemical and Cell-Based Approaches. Drug Discov. Today 2020, 25, 1807–1821. [Google Scholar] [CrossRef] [PubMed]
- An, W.F.; Tolliday, N. Cell-Based Assays for High-Throughput Screening. Mol. Biotechnol. 2010, 45, 180–186. [Google Scholar] [CrossRef]
- Lai, C.; Jiang, X.; Li, X. Development of Luciferase Reporter-Based Cell Assays. Assay. Drug Dev. Technol. 2006, 4, 307–315. [Google Scholar] [CrossRef]
- Jiang, T.; Xing, B.; Rao, J. Recent Developments of Biological Reporter Technology for Detecting Gene Expression. Biotechnol. Genet. Eng. Rev. 2008, 25, 41–76. [Google Scholar] [CrossRef] [PubMed]
- Cevenini, L.; Calabretta, M.M.; Calabria, D.; A. Roda, E.M. Luciferase Genes as Reporter Reactions: How to Use Them in Molecular Biology? Adv. Biochem. Eng. Biotechnol. 2015, 123, 127–141. [Google Scholar] [CrossRef]
- Close, D.M.; Ripp, S.; Sayler, G.S. Reporter Proteins in Whole-Cell Optical Bioreporter Detection Systems, Biosensor Integrations, and Biosensing Applications. Sensors 2009, 9, 9147–9174. [Google Scholar] [CrossRef]
- Gould, S.J.; Subramani, S. Firefly Luciferase as a Tool in Molecular and Cell Biology. Anal. Biochem. 1988, 175, 5–13. [Google Scholar] [CrossRef] [PubMed]
- Thorne, N.; Inglese, J.; Auld, D.S. Illuminating Insights into Firefly Luciferase and Other Bioluminescent Reporters Used in Chemical Biology. Chem. Biol. 2010, 17, 646–657. [Google Scholar] [CrossRef]
- Azad, T.; Janse Van Rensburg, H.J.; Morgan, J.; Rezaei, R.; Crupi, M.J.F.; Chen, R.; Ghahremani, M.; Jamalkhah, M.; Forbes, N.; Ilkow, C.; et al. Luciferase-Based Biosensors in the Era of the COVID-19 Pandemic. ACS Nanosci. Au 2021, 1, 15–37. [Google Scholar] [CrossRef]
- Wigdal, S.S.; Anderson, J.L.; Vidugiris, G.J.; Shultz, J.; Wood, K. V.; Fan, F. A Novel Bioluminescent Protease Assay Using Engineered Firefly Luciferase. Curr. Chem. Genom. 2008, 2, 16–28. [Google Scholar] [CrossRef]
- Zhou, J.; Wang, D.; Xi, Y.; Zhu, X.; Yang, Y.; Lv, M.; Luo, C.; Chen, J.; Ye, X.; Fang, L.; et al. Assessing Activity of Hepatitis A Virus 3C Protease Using a Cyclized Luciferase-Based Biosensor. Biochem. Biophys. Res. Commun. 2017, 488, 621–627. [Google Scholar] [CrossRef]
- Kilianski, A.; Mielech, A.M.; Deng, X.; Baker, S.C. Assessing Activity and Inhibition of Middle East Respiratory Syndrome Coronavirus Papain-Like and 3C-Like Proteases Using Luciferase-Based Biosensors. J. Virol. 2013, 87, 11955–11962. [Google Scholar] [CrossRef]
- Chen, K.Y.; Krischuns, T.; Varga, L.O.; Harigua-Souiai, E.; Paisant, S.; Zettor, A.; Chiaravalli, J.; Delpal, A.; Courtney, D.; O’Brien, A.; et al. A Highly Sensitive Cell-Based Luciferase Assay for High-Throughput Automated Screening of SARS-CoV-2 Nsp5/3CLpro Inhibitors. Antivir. Res. 2022, 201, 105272. [Google Scholar] [CrossRef]
- Rawson, J.M.O.; Duchon, A.; Nikolaitchik, O.A.; Pathak, V.K.; Hu, W.S. Development of a Cell-Based Luciferase Complementation Assay for Identification of Sars-Cov-2 3clpro Inhibitors. Viruses 2021, 13, 1–16. [Google Scholar] [CrossRef] [PubMed]
- Belenkaya, S. V.; Merkuleva, I.A.; Yarovaya, O.I.; Chirkova, V.Y.; Sharlaeva, E.A.; Shanshin, D. V.; Volosnikova, E.A.; Vatsadze, S.Z.; Khvostov, M. V.; Salakhutdinov, N.F.; et al. The Main Protease 3CLpro of the SARS-CoV-2 Virus: How to Turn an Enemy into a Helper. Front. Bioeng. Biotechnol. 2023, 11, 923–924. [Google Scholar] [CrossRef]
- Bradford, M.M. A Rapid and Sensitive Method for the Quantitation of Microgram Quantities of Protein Utilizing the Principle of Protein-Dye Binding; 1976; Vol. 72. [Google Scholar]
- Szecsi, P.B. The Aspartic Proteases. Scand. J. Clin. Lab. Invest. 1992, 52, 5–22. [Google Scholar] [CrossRef]
- Abramson, J.; Adler, J.; Dunger, J.; Evans, R.; Green, T.; Pritzel, A.; Ronneberger, O.; Willmore, L.; Ballard, A.J.; Bambrick, J.; et al. Accurate Structure Prediction of Biomolecular Interactions with AlphaFold 3. 2024, 630, 493–500. [Google Scholar] [CrossRef]
- Conti, E.; Franks, N.P.; Brick, P. Crystal Structure of Firefly Luciferase Throws Light on a Superfamily of Adenylate-Forming Enzymes. 1996, 4, 287–298. [Google Scholar] [CrossRef]
- Mariani, V.; Biasini, M.; Barbato, A.; Schwede, T. Structural Bioinformatics LDDT : A Local Superposition-Free Score for Comparing Protein Structures and Models Using Distance Difference Tests. 2013, 29, 2722–2728. [Google Scholar] [CrossRef] [PubMed]
- Frei, T.; Cella, F.; Tedeschi, F.; Gutiérrez, J.; Stan, G.B.; Khammash, M.; Siciliano, V. Characterization and Mitigation of Gene Expression Burden in Mammalian Cells. Nat. Commun. 2020, 11, 1–14. [Google Scholar] [CrossRef]
- Owen, D.R.; Allerton, C.M.N.; Anderson, A.S.; Aschenbrenner, L.; Avery, M.; Berritt, S.; Boras, B.; Cardin, R.D.; Carlo, A.; Coffman, K.J.; et al. An Oral SARS-CoV-2 Mpro Inhibitor Clinical Candidate for the Treatment of COVID-19. 2021, 3, 1586–1593. [Google Scholar] [CrossRef]
- Lockbaum, G.J.; Reyes, A.C.; Lee, J.M.; Tilvawala, R.; Nalivaika, E.A.; Ali, A.; Yilmaz, N.K.; Thompson, P.R.; Schiffer, C.A. Crystal Structure of Sars-Cov-2 Main Protease in Complex with the Non-Covalent Inhibitor Ml188. Viruses 2021, 13. [Google Scholar] [CrossRef]
- Sala-Newby, G.B.; Campbell, A.K. Stepwise Removal of the C-Terminal 12 Amino Acids of Firefly Luciferase Results in Graded Loss of Activity. Biochim. Biophys. Acta-Protein Struct. Mol. Enzymol. 1994, 1206, 155–160. [Google Scholar] [CrossRef]
- Pedersen, N.C.; Kim, Y.; Liu, H.; Galasiti Kankanamalage, A.C.; Eckstrand, C.; Groutas, W.C.; Bannasch, M.; Meadows, J.M.; Chang, K.O. Efficacy of a 3C-like Protease Inhibitor in Treating Various Forms of Acquired Feline Infectious Peritonitis. J. Feline Med. Surg. 2018, 20, 378–392. [Google Scholar] [CrossRef]
- Fu, L.; Ye, F.; Feng, Y.; Yu, F.; Wang, Q.; Wu, Y.; Zhao, C.; Sun, H.; Huang, B.; Niu, P.; et al. Both Boceprevir and GC376 Efficaciously Inhibit SARS-CoV-2 by Targeting Its Main Protease. Nat. Commun. 2020, 11, 1–8. [Google Scholar] [CrossRef]
- Jacobs, J.; Grum-Tokars, V.; Zhou, Y.; Turlington, M.; Saldanha, S.A.; Chase, P.; Eggler, A.; Dawson, E.S.; Baez-Santos, Y.M.; Tomar, S.; et al. Discovery, Synthesis, and Structure-Based Optimization of a Series of N -(Tert -Butyl)-2-(N -Arylamido)-2-(Pyridin-3-Yl) Acetamides (ML188) as Potent Noncovalent Small Molecule Inhibitors of the Severe Acute Respiratory Syndrome Coronavirus (SARS-CoV) 3CL. J. Med. Chem. 2013, 56, 534–546. [Google Scholar] [CrossRef] [PubMed]
- Lobo-Galo, N.; Terrazas-López, M.; Martínez-Martínez, A.; Díaz-Sánchez, Á.G. FDA-Approved Thiol-Reacting Drugs That Potentially Bind into the SARS-CoV-2 Main Protease, Essential for Viral Replication. J. Biomol. Struct. Dyn. 2021, 39, 3419–3427. [Google Scholar] [CrossRef] [PubMed]
- Gurard-Levin, Z.A.; Liu, C.; Jekle, A.; Jaisinghani, R.; Ren, S.; Vandyck, K.; Jochmans, D.; Leyssen, P.; Neyts, J.; Blatt, L.M.; et al. Evaluation of SARS-CoV-2 3C-like Protease Inhibitors Using Self-Assembled Monolayer Desorption Ionization Mass Spectrometry. Antivir. Res. 2020, 182, 104924. [Google Scholar] [CrossRef] [PubMed]
- Wang, X.; Cao, R.; Zhang, H.; Liu, J.; Xu, M.; Hu, H.; Li, Y.; Zhao, L.; Li, W.; Sun, X.; et al. The Anti-Influenza Virus Drug, Arbidol Is an Efficient Inhibitor of SARS-CoV-2 in Vitro. Cell Discov. 2020, 6, 28. [Google Scholar] [CrossRef] [PubMed]
| Compound | CC50, μM (HEK293T) |
IC50, μM (Luc System) |
IC50, μM (rec-3CLpro) |
IC50, μM (Virus) |
|---|---|---|---|---|
| Nirmatrelvir | >100 | 6.1 ± 0.8 | 0.105 ± 0.009 | 3.6 ± 0.5 |
| GC376 | >100 | 3.3 ± 1.1 | 0.023 ± 0.004 | 9.54 ± 2.03 |
| ML188 | 90±4 | NA | 1.56 ± 0.55 | 3.77 ± 0.87 |
| Disulfiram | 101±6 | - | 6.25 ± 1.97 | - |
| Remdesivir | >100 | - | - | 3.76 ± 0.94 |
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2026 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).




