Submitted:
31 March 2026
Posted:
01 April 2026
You are already at the latest version
Abstract
Keywords:
1. Introduction
2. Materials and Methods
2.1. Participants
2.2. Molecular Karyotyping
2.3. Whole Exome Sequencing (WES)
2.4. Association of Genes with CHD and Its Dosage Sensitivity
3. Results
3.1. Patients’ Demographic and Clinical Data
3.2. Chromosomal Microarray Analysis Results
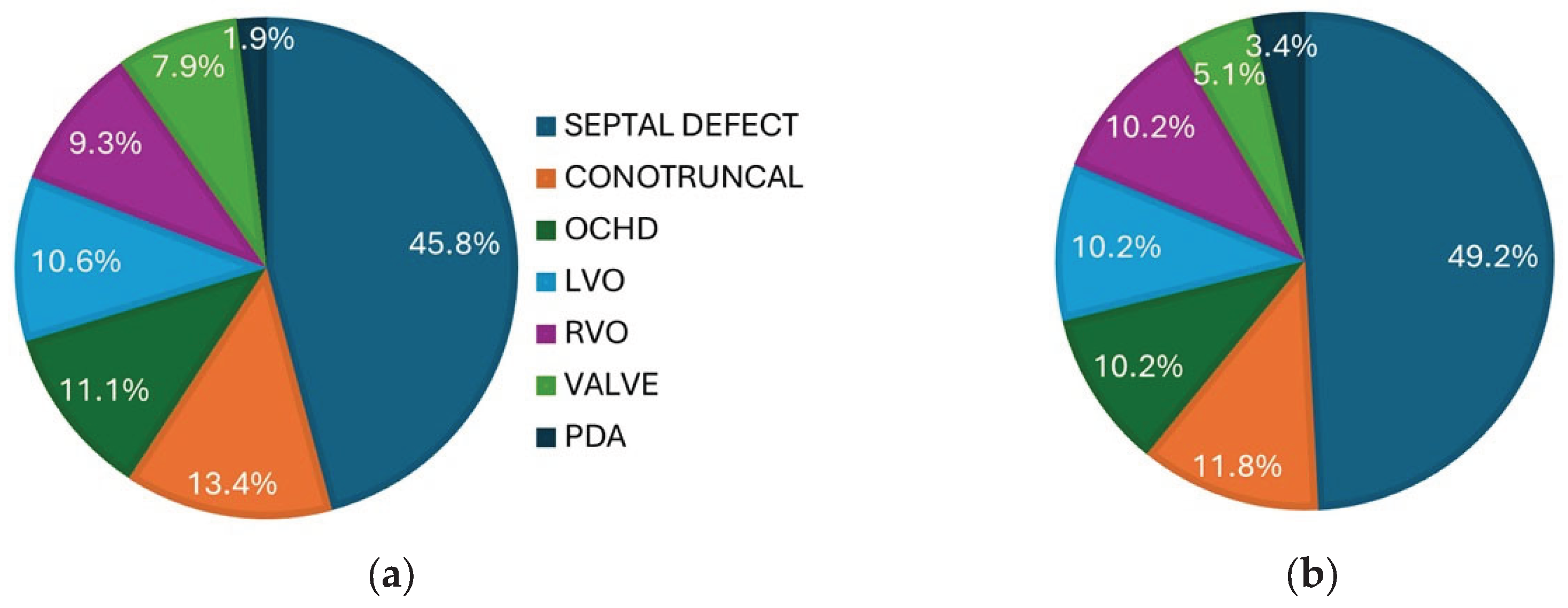
3.3. WES Analysis Results
3.4. Analysis of the Diagnostic Yield and Candidate Genes for CHD
4. Discussion
5. Conclusions
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Acknowledgments
Conflicts of Interest
Abbreviations
| CHD | congenital heart defect |
| CNV | copy number variation |
| SNV | single-nucleotide variants |
| DD/ID | developmental delay/intellectual disability |
| WES | whole-exome sequencing |
| INDELs | insertions/deletions |
| ASD | atrial septal defect |
| VSD | ventricular septal defect |
| ToF | tetralogy of Fallot |
| CMA | chromosomal microarray |
| ES | exome sequencing |
| AVSD | atrioventricular septal defect |
| TGA | dextro-transposition of the great arteries |
| DORV | double outlet right ventricle |
| TAC | truncus arteriosus communis |
| AS | aortic stenosis |
| PS | pulmonary stenosis |
| LVO | left ventricular outflow tract obstructions |
| RVO | right ventricular outflow tract obstructions |
| HLHS | hypoplastic left heart syndrome |
| CoA | coarctation of the aorta |
| PAVSD | pulmonary atresia with ventricular septal defect |
| OCHD | other complex congenital heart defects |
| PDA | patent ductus arteriosus |
| PAToA | pulmonary atresia with D-transposition of aorta and atrial septal defect |
| IUGR | intrauterine growth retardation |
| MVP | mitral valve prolapses |
| NEC | necrotizing enterocolitis |
| PFO | patent foramen ovale |
| TAPVR | total anomalous pulmonary venous return |
| VUR | vesicoureteral reflux |
| AAC | amino acid change |
| CCA | coronary cameral fistulas |
| IAA | interrupted aortic arch |
References
- Pierpont, M.E; Brueckner, M.; Chung, W.K.; Garg, V.; Lacro, R.V.; McGuire, A.L.; Mital, S.; Priest, J.R.; Pu, W.T.; Roberts, A.; et al. Genetic Basis for Congenital Heart Disease: Revisited: A Scientific Statement from the American Heart Association. Circulation 2018, 138, e653–e711. [Google Scholar] [CrossRef] [PubMed]
- Xu, J.; Li, Q.; Deng, L.; Xiong, J.; Cheng, Z.; Ye, C. Global, regional, and national epidemiology of congenital heart disease in children from 1990 to 2021. Front. Cardiovasc. Med. 2025, 12, 1522644. Available online: https://pubmed.ncbi.nlm.nih.gov/40454242/. [CrossRef]
- Cowan, JR; Ware, SM. Genetics and genetic testing in congenital heart disease. Clin Perinatol 2015, 42, 373–393, ix. [Google Scholar] [CrossRef] [PubMed]
- Manning, M.; Hudgins, L. Array-based technology and recommendations for utilization in medical genetics practice for detection of chromosomal abnormalities. Gen. Med. 2010, 12(11), 742–745. [Google Scholar] [CrossRef]
- Wilde, A.A.M.; Semsarian, C.; Márquez, M.F.; Shamloo, A.S.; Ackerman, M.J.; Ashley, E.A.; Sternick, E.B.; Barajas-Martinez, H.; Behr, E.R.; Bezzina, C.R.; et al. European Heart Rhythm Association (EHRA)/Heart Rhythm Society (HRS)/Asia Pacific Heart Rhythm Society (APHRS)/Latin American Heart Rhythm Society (LAHRS) Expert Consensus Statement on the state of genetic testing for cardiac diseases. Heart Rhythm 2022. [Google Scholar] [CrossRef]
- Nappi, F. In-Depth Genomic Analysis: The New Challenge in Congenital Heart Disease. Int. J. Mol. Sci. 2024, 25, 1734. [Google Scholar] [CrossRef]
- Christoffels, V.M.; Habets, P.E.; Franco, D.; Campione, M.; de Jong, F.; Lamers, W.H.; Bao, Z.Z.; Palmer, S.; Biben, C.; Harvey, R.P.; et al. Chamber formation and morphogenesis in the developing mammalian heart. Dev. Biol. 2000, 223, 266–278. [Google Scholar] [CrossRef]
- Astrof, S.; Arriagada, C.; Saijoh, Y.; Francou, A.; Kelly, R.G.; Moon, A. Aberrant differentiation of second heart field mesoderm prefigures cellular defects in the outflow tract in response to loss of FGF8. Dev. Biol. 2023, 499, 10–21. [Google Scholar] [CrossRef]
- Ivanovitch, K.; Soro-Barrio, P.; Chakravarty, P.; Jones, R.A.; Bell, D.M.; Mousavy Gharavy, S.N.; Stamataki, D.; Delile, J.; Smith, J.C.; Briscoe, J. Ventricular, atrial, and outflow tract heart progenitors arise from spatially and molecularly distinct regions of the primitive streak. PLoS Biol. 2021, 19, e3001200. [Google Scholar] [CrossRef]
- Simmons, M. A.; Brueckner, M. The Genetics of Congenital Heart diseaseUnderstanding and Improving Long Term Outcomes in Congenital Heart Disease: A Review for the General Cardiologist and Primary Care Physician. Curr. Opin. Pediatr. 2017, 29(5), 520–528. [Google Scholar] [CrossRef] [PubMed]
- Vey, C.; Melnik, N.; Dombrowsky, G.; Hitz, M.P. Current and future diagnostics of congenital heart disease. Medizinische Genetik 2025, 37(2), 95–102. [Google Scholar] [CrossRef]
- Miller, S.A.; Dykes, D.D.; Polesky, H.F. A simple salting out procedure for extracting DNA from human nucleated cells. Nucleic Acids Res. 1988, 11(16(3)), 1215. [Google Scholar] [CrossRef]
- Riggs, E.R.; Andersen, E.F.; Cherry, A.M.; Riggs, E.R.; Andersen, E.F.; Cherry, A.M.; Kantarci, S.; Kearney, H.; Patel, A.; Raca, G.; et al. Correction: Technical standards for the interpretation and reporting of constitutional copy-number variants: a joint consensus recommendation of the American College of Medical Genetics and Genomics (ACMG) and the Clinical Genome Resource (ClinGen). Genet. Med. 2020, 22(2), 245–257. [Google Scholar] [CrossRef]
- Bachman, K.K.; DeWard, S.J.; Chrysostomou, C.; Munoz, R.; Madan-Khetarpal, S. Array CGH as a first-tier test for neonates with congenital heart disease. Cardiology in the Young (2015) 25, 115–122. [CrossRef]
- Alankarage, D.; Ip, E.; Szot, J.O.; Munro, J.; Blue, G.M.; Harrison, K.; Cuny, H.; Enriquez, A.; Troup, M.; Humphreys, D.T.; et al. Identification of clinically actionable variants from genome sequencing of families with congenital heart disease. Genet. Med. 2019, 21, 1111–1120. [Google Scholar] [CrossRef] [PubMed]
- Shikany, A.R.; Landis, B.J.; Parrott, A.; Miller, E.M.; Coyan, A.; Walters, L.; Hinton, R.B.; Goldenberg, P.; Ware, S. M. A Comprehensive Clinical Genetics Approach to Critical Congenital Heart Disease in Infancy. J. Pediatr. 2020, 227, 231–238.e14. [Google Scholar] [CrossRef] [PubMed]
- Mone, F.; Eberhardt, R.Y.; Morris, R.K.; Hurles, M.E.; McMullan, D.J.; Maher, E.R.; Lord, J.; Chitty, L.S.; Giordano, J.L.; Wapner, R.J.; et al. Congenital heart disease and the Diagnostic yield with Exome sequencing (CODE) study: prospective cohort study and systematic review. Ultrasound Ob stet. Gynecol. 2021, 57, 43–51. [Google Scholar] [CrossRef] [PubMed]
- Shan, W.; Yuanqing, X.; Jing, Z.; Xi, W.; Huifeng, G.; Yi, W. Risk factor analysis for adverse prognosis of the fetal ventricular septal defect (VSD). BMC Pregnancy Childbirth 2023, 23, 683. [Google Scholar] [CrossRef]
- Slavotinek, A.M.; Thompson, M.L.; Martin, L. J.; Gelb, B.D. Diagnostic yield after next-generation sequencing in pediatric cardiovascular disease. Human Genetics and Genomics Advances 2024, 5, 100286. [Google Scholar] [CrossRef]
- Simrick, S.; Szumska, D.; Gardiner; Jones, J.R.; Sagar, K.; Morrow, B.; Bhattacharya, S.; Basson, M.A. Biallelic Expression of Tbx1 Protects the Embryo From Developmental Defects Caused by Increased Receptor Tyrosine Kinase. Signaling Developmental Dynamics 2012, 241, 1310–1324. [Google Scholar] [CrossRef]
- Cillo, F.; Coppola, E.; Habetswallner, F.; Cecere, F.; Pignata, L.; Toriello, E.; De Rosa, A.; Grilli, L.; Ammendola, A.; Salerno, P.; et al. Understanding the Variability of 22q11.2 Deletion Syndrome: The Role of Epigenetic Factors. Genes (Basel) 2024, 29(15(3)). [Google Scholar] [CrossRef]
- Mary, L.; Leclerc, D.; Gilot, D.; Belaud-Rotureau, M.A.; Jaillard, S. The TALE never ends: A comprehensive overview of the role of PBX1, a TALE transcription factor, in human developmental defects. Human Mutation 2022, 43(9), 1125–1148. [Google Scholar] [CrossRef]
- Huai, W.; Li, J.; Li, X.; Ding, Y.; Yu, T.; Zhang, H.; Wang, X.; Yao, R. Novel variants in. ZNF462 and phenotype update in patients with Weiss-Kruszka syndrome: a case series Transl Pediatr 2025, 14(8), 1991–2000. [Google Scholar] [CrossRef]
- Martin, R.; Splitt, M.; Genevieve, D.; Aten, E.; Collins, A.; de Bie, C. I.; Faivre, L.; Foulds, N.; Giltay, J.; Ibitoye, R. De novo variants in CNOT3 cause a variable neurodevelopmental disorder. Eur J Hum Genet 2019, 27(11), 1677–1682. [Google Scholar] [CrossRef]
- Jin, L.; Ming, W.J.; Ding, M.P. A novel pathogenic variant in CNOT3 causing neurodevelopmental delay and epilepsy. Seizure: European Journal of Epilepsy 2023, 107, 104–106. [Google Scholar] [CrossRef] [PubMed]
- Mustafa, H.J.; Najjariasl, P.; Faezeh, A.; Sambatur, E.V.; Rodenbarger, A.; Guseh, S.; Roberts, E.A.; Shamshirsaz, A.A. Yield of Whole Genome Sequencing for Pathogenic Single Nucleotide Variants in Congenital Heart Disease: A Systematic Review and Meta-Analysis. Prenatal Diagnosis 2025, 00, 1–39. [Google Scholar] [CrossRef]
- Li, Y.; Zhou, Z.; Xu, Y.; Wang, Z.R. Novel mutation c.2090_2091del in neurodevelopmental-craniofacial syndrome with variable renal and cardiac abnormalities in an 18.5-mo-old boy: A case report. World J Clin Cases 2023, 11(16), 3891–3898. [Google Scholar] [CrossRef] [PubMed]
- Yasuhara, J.; Garg, V. Genetics of congenital heart disease: a narrative review of recent advances and clinical implications. Transl. Pediatr. 2021, 10(9), 2366–2386. [Google Scholar] [CrossRef]
- Khasawneh, R.R.; Kist, R.; Queen, R.; Hussain, R.; Coxhead, J.; Schneider, J.E.; Mohun, T.J.; Zaffran, S.; Peters, H.; Phillips, H.M.; et al. Msx1 haploinsufficiency modifies the Pax9-deficient cardiovascular phenotype. BMC Dev Biol 2021, 21, 14. [Google Scholar] [CrossRef]
- Ferese, R.; Bonetti, M.; Consoli, F.; Guida, V.; Sarkozy, A.; Lepri, F.R.; Versacci, P.; Gambardella, S.; Calcagni, G.; Margiotti, K.; et al. Heterozygous missense mutations in NFATC1 are associated with atrioventricular septal defect. Human Mutation 2018, 39, 1428–1441. [Google Scholar] [CrossRef]
- Gutmann, J.L.; Spister, A.; Baticic, L. From Genetics to Phenotype: Understandingthe Diverse Manifestations of Cardiovascular Genetic Diseases in Pediatric Populations. Cardiogenetics 2025, 15, 29. [Google Scholar] [CrossRef]
- Jordan, V.K.; Zaveri, H.P.; Scott, D. A 1p36 deletion syndrome: an update. Appl Clin Genet. 2015, 27(8), 189–200. [Google Scholar] [CrossRef]
- Morris, C.A.; Mervis, C.B. Williams syndrome and related disorders. Annu Rev Genomics Hum Genet. 2000, 1, 461–84. [Google Scholar] [CrossRef]
- Muiño Mosquera, L.; Steijns, F.; Audenaert, T.; Meerschaut, I.; de Paepe, A.; Steyaert, W.; Symoens, S.; Coucke, P.; Callewaert, B.; Renard, M.; et al. Tailoring the American College of Medical Genetics and Genomics and the Association for Molecular Pathology Guidelines for the interpretation of sequenced variants in the FBN1 gene for Marfan syndrome: proposal for a disease- and gene-specific guideline. Circ Genom Precis Med. 2018, 11, e002039. [Google Scholar] [CrossRef] [PubMed]
- Davies, G.E.; Howard, C.M.; Farrer, M.J.; Coleman, M.M.; Bennett, L.B.; Cullen, L. M.; Wyse, R.K.H.; Burn, J.; Williamson, R.; Kessling, A.M. Genetic variation in the COL6A1 region is associated with congenital heart defects in trisomy 21 (Down’s syndrome). Ann. Hum. Genet. 1995, 59, 253–269. [Google Scholar] [CrossRef] [PubMed]
- Zhang, P.; Torres, K.; Liu, X.; Liu, CG.; Pollock, RE. An Overview of Chromatin-Regulating Proteins in Cells. Curr Protein Pept Sci 2016, 17(5), 401–410. [Google Scholar] [CrossRef] [PubMed]
- Wallace, R.; Raghu, S.; Hota, S.K. Chromatin remodelers in heart development and congenital heart defects The Nucleus 2026. [CrossRef]
- Diab, N.S.; Barish, S.; Dong, W.; Zhao, S.; Allington, G.; Yu, X.; Kahle, K.T.; Brueckner, M.; Jin, S.C. Molecular Genetics and Complex Inheritance of Congenital Heart Disease. Genes 2021, 12, 1020. [Google Scholar] [CrossRef]
- Petty, E.; Pillus, L. Balancing chromatin remodeling and histone modifications in transcription Trends Genet. 2013, 29(11), 621–629. [Google Scholar] [CrossRef]
- Faravelli, F. NSD1 mutations in Sotos syndrome. American Journal of Medical Genetics -C 2005, 137(1), 24–31. [Google Scholar] [CrossRef]
- Calcagni, G.; Ferrigno, F.; Sinibaldi, L.; Minotti, C.; Parlapiano, G.; Franceschini, A.; Dentici, M.L.; Micalizzi, A.; Alesi, V.; Capolino, R. Congenital Heart Defects in Patients with Molecularly Confirmed Sotos Syndrome Diagnostics. 2024, 14, 594. [Google Scholar] [CrossRef]
- Koolen, D.A.; Pfundt, R.; Linda, K.; Beunders, G.; Veenstra-Knol, H.E.; Conta, J.H.; Fortuna, A.M.; Gillessen-Kaesbach, G.; Dugan, S.; Halbach, S.; et al. The Koolen de Vries syndrome: a phenotypic comparison of patients with a 17q21.31 microdeletion versus a KANSL1 sequence variant. Eur J Hum Genet 2016, 24(5), 652–9. [Google Scholar] [CrossRef] [PubMed]
- Valer, J. A.; Sánchez-de-Diego, C.; Pimenta-Lopes, C.; Rosa, J.L.; Ventura, F. ACVR1 Function in Health and Disease. Cells 2019, 8(11), 1366. [Google Scholar] [CrossRef]
- Chang, L.; Ji, R.; Sa, R.; Huge, J.; An, C. Whole-exome sequencing of pathogenic genes in a family with congenital heart disease A case report. AMER1 gene. 2024, 10322. [Google Scholar] [CrossRef] [PubMed]
- Sumitaka, H.; Sato, T.; Akazawa, H.; Okada, H.; Maeno, A.; Ito, M.; Sugitani, Y.; Shibata, H.; Miyazaki, J.; Katsuki, M. Apoptosis in neural crest cells by functional loss of APC tumor suppressor gene. PNAS 2002, 99, 297–302. [Google Scholar] [CrossRef]
- Massey, H.; Tennant, S.; Dean, J. PACS2, PACS1, and VACTERL: A Clinical Overlap Mol. Syndromol. 2025, 16, 29–32. [Google Scholar] [CrossRef]
- Sánchez- Soler, M.J.; Serrano-Antón, A.T.; López-González, V.; Guillén-Navarro, E. New case with the recurrent c. Neurología 657–742. [CrossRef]
- Kury, S.; Besnard, T.; Ebstein, F.; Khan, T.N.; Gambin, T.; Douglas, J.; Bacino, C.A.; Craigen, W.J.; Sanders, S.J.; Lehmann, A.; et al. De Novo Disruption of the Proteasome Regulatory Subunit PSMD12 Causes a Syndromic Neurodevelopmental Disorder. Am J Hum Genet 2017, 100(4), 689. [Google Scholar] [CrossRef]
| Children with congenital heart defects | N (%) | |
|---|---|---|
| Gender | Boys | 108 (50.0) |
| Girls | 108 (50.0) | |
| Age | Mean (+SD) | 3.19 (5.18) |
| Median (min-max) | 0.9 (0.1-18) | |
| DD/ID | Unknown | 60 (27.8) |
| Yes | 108 (50.0) | |
| No | 48 (22.2) | |
| Facial dysmorphia | 150 (69.4) | |
| Head and CNS anomalies | 52 (24.1) | |
| Urogenital tract anomalies | 29 (13.4) | |
| Skeletal and joint system anomalies | 21 (9.8) | |
| Autism spectrum disorders | 9 (4.2) | |
| Epilepsy | 14 (6.5) | |
| Isolated CHD | 19 (8.8) | |
| Syndrome | Region | CNV Type | Size (Mb) | Gene (OMIM) | N (%) | CHD type (n) |
|---|---|---|---|---|---|---|
| DiGeorge | 22q11.2 | Deletion | 2.50 | TBX1 (602054) | 8 (13.6) | ToF (4), AVSD (2), VSD (2) PAToA (2), OCHD (1) |
| 22q11.2 duplication | 22q11.2 | Duplication | 2.80 | 3 (5.1) | ||
| Williams-Beuren | 7q11.23 | Deletion | 1.40 | ELN (130169) | 5 (8.5) | *PS (3), *AS (2) |
| Noonan | 2p22.1 | Duplication | 2.73 | SOS1 (182530) | 2 (3.4) | VSD (1) |
| 11p15.2 | Deletion | 4.20 | RRAS2 (600098) | PS (1) |
| CMA findings (GRCh37) | Size Mb | Syndromes (OMIM) | Gene(s) (OMIM) | Phenotype | |
|---|---|---|---|---|---|
| 1 | 1p31.3(61782914_62322790) x1 | 0.54 | NA | NA | OCHD, corpus callosum agenesia, congenital inguinal hernia |
| 2 | 1p36.13-p36.11(18900374_23966858)x1 | 5.1 | 1p36 microdeletion (#607872) |
CDC42 (16952) ECE1 (600423) |
ASD, facial dysmorphia, hypotonia, clubfoot, right sided hydronephrosis, |
| 3 | 1q21.1(145413388_145747269)x1 | 0.33 | Thrombocytopenia Absent Radius-TAR (#274000) |
RBM8A (605313) |
AVSD, TAR |
| 4 | 1q23.2-q23.3(160465291_165429037)x1 | 4.9 | NA |
PBX1 (176310) |
ASD, facial dysmorphia, ectopic kidney, glaucoma, cerebral palsy |
| 5 | 2q11.1-q11.2(96779631_98021592)x1 | 1.2 | NA | NA | PDA, preterm birth, microcephaly, craniosynostosis, metatarsus varus |
| 7 | 2q22.2-q22.3(143986161_146890297)x1 | 2.9 | Mowat-Wilson (#235730) |
ZEB2 (605802) |
CoA, aortic bicuspid valve, facial dysmorphia, microcephaly, corpus callosum agenesia |
| 8 | 2q23.3-q24.1(151373825_158622730)x1 | 7.3 | NA |
ACVR1 (102576) |
PDA, VSD, facial dysmorphia, microcephaly |
| 9 | 3q22.1-q29(133562250_197840339)x3 | 64.3 | Noonan type 11 (#618499) |
MRAS (608435) |
OCHD, facial dysmorphia, cleft soft palate, IUGR |
| 10 | 3q28-q29 (191861311_197840339)x3 5p15.33-p15.31 (151737_7144623)x1 |
6.0 7.0 |
3q29 microduplication (#611936) Cri du chat (#123450) |
PAK2 (605022) |
AV block I, PS, PFO, facial dysmorphia, hypospadias, hypopituitarism, hip dysplasia, scoliosis, myopia |
| 11 | 4p16.3-p16.1(71552_7760991)x1 17q25.3(76890486_81029941)x3 |
7.7 4.1 |
Wolf-Hirschhorn (#194190) |
MSX1 (142983) |
ASD, PS, facial dysmorphia, complete cleft palate, microcephaly, hypotonia |
| 12 | 4q21.21(81561965_82010110)x3 | 0.45 | NA | NA |
AVSD, facial dysmorphia, rhizomelia, hypotonia |
| 13 | 5q21.3-q23.1(105623245_116004172)x1 | 10.4 | Familial Adenomatous Polyposis (#175100) |
APC (175100) |
ToF, facial dysmorphia, imperforate anus |
| 14 | 7p15.3-p14.3(20993642_30739239)x1 | 9.7 | NA |
HOXA1 (142955) |
ASD, facial dysmorphia, IUGR, kidney hypoplasia, skeletal anomalies |
| 15 | 7q33-q36.3(133749674_158909738)x3 | 25.2 | NA |
KMT2C (606833) |
PDA, facial dysmorphia, cleft palate, brain anomalies |
| 16 | 8p23.3-p23.1(221611_9261350)x1 8q21.2-q24.3(86842195_146280020)x3 |
9.0 59.4 |
Cornelia de Lange Syndrome 4 (#614701) |
RAD21 (606462) |
AVSD, PDA, ptosis, Pierre Robin sequence |
| 17 | 8p23.3-p22 (524066_17541888)x3 9p24.3-p24.2 (271257_42776209x1 |
17.0 4.0 |
8p23.1 microduplication |
GATA4 (600576) |
OCHD, omphalocele, scoliosis, arachnodactyly |
| 18 | 8q23.3-q24.23(113589865_136427632)x4 | 22.8 | Cornelia de Lange Syndrome 4 (#614701) |
RAD21 (606462) |
VSD, facial dysmorphia, cryptorchidism |
| 19 | 8q22.2-q22.23(100973253_103335730)x1 | 2.4 | NA |
SPAG1 (603395) |
AVSD, facial dysmorphia, complete palate cleft |
| 20 | 9p23-p22.3(12772471_14680180)x1 | 1.9 | NA |
MPDZ (603785) |
AVSD, craniofacial dysmorphia, macrocephaly |
| 21 | 9q31.1-q31.3(106828041_112710753)x1 | 5.9 | 9q31.1-q31.3 microdeletion (#618619) |
ZNF462 (617371) |
VSD, facial dysmorphia, bilateral VUR |
| 22 | 12p13.3-p11(511504_34189943)x4 | 35 p arm |
Pallister- Killian (# 601803) |
NA | VSD, bicuspid aortic valve, facial dysmorphia, acromelia, brain anomalies, anal atresia, cryptorchidism |
| 23 | 14q32.3(105717621_106327993)x1 | 0.60 | NA |
PACS2 (610423) |
ASD, hypertelorism, micrognathia |
| 24 | 15q11.2-q13.1(23699701_28525460)x1 | 4.8 | Angelman type II (#105830) |
NA | mitral valve dysplasia, DD/ID, facial dysmorphia, strabismus |
| 25 | 15q13.2-q13.3(31014508_32510863)x1 | 1.5 | 15q13.3 microdeletion (#612001) |
KLF13 (605328) |
MVP, left ventricular hypertrophy facial dysmorphia |
| 26 | 15q21.1(48905243_49084691)x1 | 0.29 | Marfan (#154700) |
FBN1 (134797) |
MVP, voluminous left ventricle, Marfan-like phenotype |
| 27 | 15q26.2-q26.3(94447479_102383473)x1 | 7.9 | NA |
NR2F2 (107773) |
AVSD, facial dysmorphia, short stature, VUR bilateral |
| 28 | 16p11.2(29673954_30198600)x1 | 0.52 | 16p11.2 proximal microdeletion (#611913) |
TBX6 (602427) |
ToF, polycystic kidney disease |
| 29 | 16q13.11(15048751_16249607)x1 | 1.2 | NA |
MYH11 (160745) |
PDA, preterm birth, craniofacial dysmorphia, bilateral inguinal hernias, hyperbilirubinemia |
| 30 | 16q11.2-q12.1(46564557_49053314)x3 16q12.2-q22.2(55361181_71354431)x3 |
2.5 16.0 |
NA | NA | ASD, micrognathia, short neck, torticollis, umbilical hernia, pes varus |
| 31 | 16q11.2-q22.2(46564557_71127772)x3 | 24.6 | NA | NA | OCHD, facial dysmorphia, hypotonia |
| 32 | 16q24.2-q24.3(88653937_89429735)x3 | 0.78 | NA |
ANKRD11 (611192) |
PDA, PFO, mitral valve anomaly, hydrops fetalis, |
| 33 | 17q12(34817422_36168104)x1 |
1.4 | 17q12 microdeletion (#614527) |
NA | VSD, facial dysmorphia, polycystic kidney disease |
| 34 | 17p13.3-p13.2 (51885_3882130)x1 |
3.8 | Miller-Dieker (#247200) |
DPH1(60352) NXN (612895) |
VSD, IUGR, hypotrophy, brain anomalies, toe anomalies |
| 35 | 17q21.31(43717703_44159862)x1 | 0.44 | Koolen-De Vries (#610443) |
KANSL1 (612452) |
ASD, facial dysmorphia, kidney agenesis, unilateral cleft lip |
| 36 | 5q35.2-q35.3(176033642_177013961)x3 [0.412] 18p11.32-p11.21(142096_14748636)x1[0.412] 18q21.2-q23(49545872_77901872)x1[0.412] |
0.98 14.6 28.4 |
NA Chromosome 18 ring |
NSD1 (606681) NFACT1 (600488) |
OCHD, facial dysmorphia, microcephaly, cleft lip and palate |
| 37 | 18q21.33-q23(59653070_78621175)x1 | 17.2 | 18q microdeletion (#601808) |
NFACT1 (600488) |
mitral and tricuspid valve dysplasia, facial dysmorphia |
| 38 | 18q23(75814123_78014123)x1 | 2.2 | 18q microdeletion (#601808) |
NFACT1 (600488) |
ASD, facial dysmorphia, microphthalmia, hypertrichosis, microcephaly, brain atrophy |
| 39 | 21q22. (46411778_48067924)x1 | 1.7 | NA |
COL6A1 (120220) |
ASD, VSD, right aortic arch, pulmonary artery atresia, facial dysmorphia, NEC |
| 40 | 22q11.1-q11.21(17096855_18953065)x3 | 1.9 | Cat-eye (#115470) | NA | TAPVR, facial dysmorphia, congenital hypothyroidism |
| 41 | 22q11.23(23739437_24988455)x3 | 1.2 | NA |
SMARCB1 (601607) |
VSD, facial dysmorphia, epilepsy, dolichocephaly |
| Syndrome | Gene (OMIM) | Transcript | SNV/INDEL | AAC | Zyg | Class | CHD type | |||
|---|---|---|---|---|---|---|---|---|---|---|
|
1 2 |
Noonan type 1 | PTPN11 (176876) | NM_002834.5 NM_002834.5 |
c.767A>G c.228G>T |
p.Gln256Arg p.Glu76Asp | Het Het |
P P |
PS PS |
||
| 3 | Noonan type 7 | BRAF (164757) | NM_004333.6 | c.1785T>G | p.Phe595Leu | Het | P | PS | ||
| 4 | Coffin-Siris 1 | ARID1B (614556) | NM_001374828.1 | c.1520C>T | p.Pro507Leu | Het | VUS PP# | HA | ||
| 5 | Sotos | NSD1 (606681) | NM_022455.5 | c.6206_6209del TTTG | p.Val2069fs | Het | P | VSD | ||
| 6 | Alagille type 1 | JAG1 (601920) | NM_000214.3 | c.2113+1G>A | splice site variant |
Het | LP | PAVSD PDA | ||
| 7 | Kabuki type 1 | KMT2D (602113) | NM_003482.4 | c.12598C>T | p.Gln4200Ter | Het | P | ASD | ||
| 8 | Stankiewicz- Isidor |
PSMD12 (604450) | NM_002816.5 | c.47_56del | p.Met12Thrfs Ter16 |
Het | P | CCA | ||
| 9 | Aarskog-Scott | FGD1 (300546) | NM_004463.3 | c.2046G>T | p.Gln682His | Hem | VUS PP# | OCHD | ||
| 10 | MIM 300373 |
AMER1 (300647) |
NM_152424.4 | c.1275C>A | p.Tyr425Ter | Hem | P | VSD | ||
| 11 | MIM 618672 | CNOT3 (604910) | NM_014516.4 | c.1438dupG | p.Ala480fs | Het | LP | IAA | ||
| 12 | MIM 616977 MIM 619522 |
HIVEP2 (143054) ZMYM2 (602221) |
NM_006734.4 NM_197968.4 |
c.3566T>C c.2320C>T |
p.Leu1189Ter p.Gln774Ter |
het het |
P LP |
PDA | ||
| 13 | MIM 612621 | SYNGAP1 (603384) | NM_006772.3 | c.3361del | p.Ser1121Ala fs*9 |
Het | LP | ASD | ||
| Function | Genes | Type CHD (n) | Patients n (%) |
|---|---|---|---|
| Transcription Factors | TBX1, PBX1, CNOT3, ZEB2, HIVEP2, ZMYM2, NR2F2, MSX1, NFATC1, ZNF462 | ToF (4), PAToA (2), ASD (2), AVSD (3), VSD (3), CoA (1), IAA (1), PDA (1), OCHD (1), PS (1) | 19 (40.4) |
|
RAS signaling pathway |
RRAS2, PTN11, BRAF, SOS1, FGD1, CDC42, SYNGAP1 | PS (4), ASD (2), VSD (1), OCHD (1) | 8 (17.0) |
| Structural proteins | ELN, FBN1, MYH11, COL6A1 | PS* (3), AS* (2), MVP (1), PDA (1), OCHD (1) | 8 (17.0) |
| Chromatin regulating | ARID1B, SMARCB1 ANKRD11, NSD1, KMT2D, KANSL1 | HA (1), PDA (1), ASD (2), VSD (2) | 6 (12.8) |
| GFR and tumor suppressors | ACVR1, AMER1, APC | VSD (2), ToF (1) | 3 (6.4) |
| Ungrouped protein | PACS2, JAG1, PSMD12, | CCA (1), ASD (1), PAVSD (1) | 3 (6.4) |
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2026 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).




