Submitted:
31 March 2026
Posted:
01 April 2026
You are already at the latest version
Abstract
Keywords:
1. Introduction
2. Materials and Methods
2.1. Search Strategy and Data Sources
2.2. Inclusion Criteria
- Studies were included if they met all the following criteria:
- Focus on diagnostic US imaging;
- Use deep learning or machine learning methods for US image denoising or speckle reduction;
- Published in a peer-reviewed journal or a reputable medical imaging, computer vision, or machine learning conference;
2.3. Study Screening and Selection
2.4. Data Extraction
- A structured data charting process was employed to systematically extract and organize information from all included studies. The data extraction form was piloted on a subset of studies and calibrated by the review team prior to use to ensure consistency. The following variables were extracted from each study.
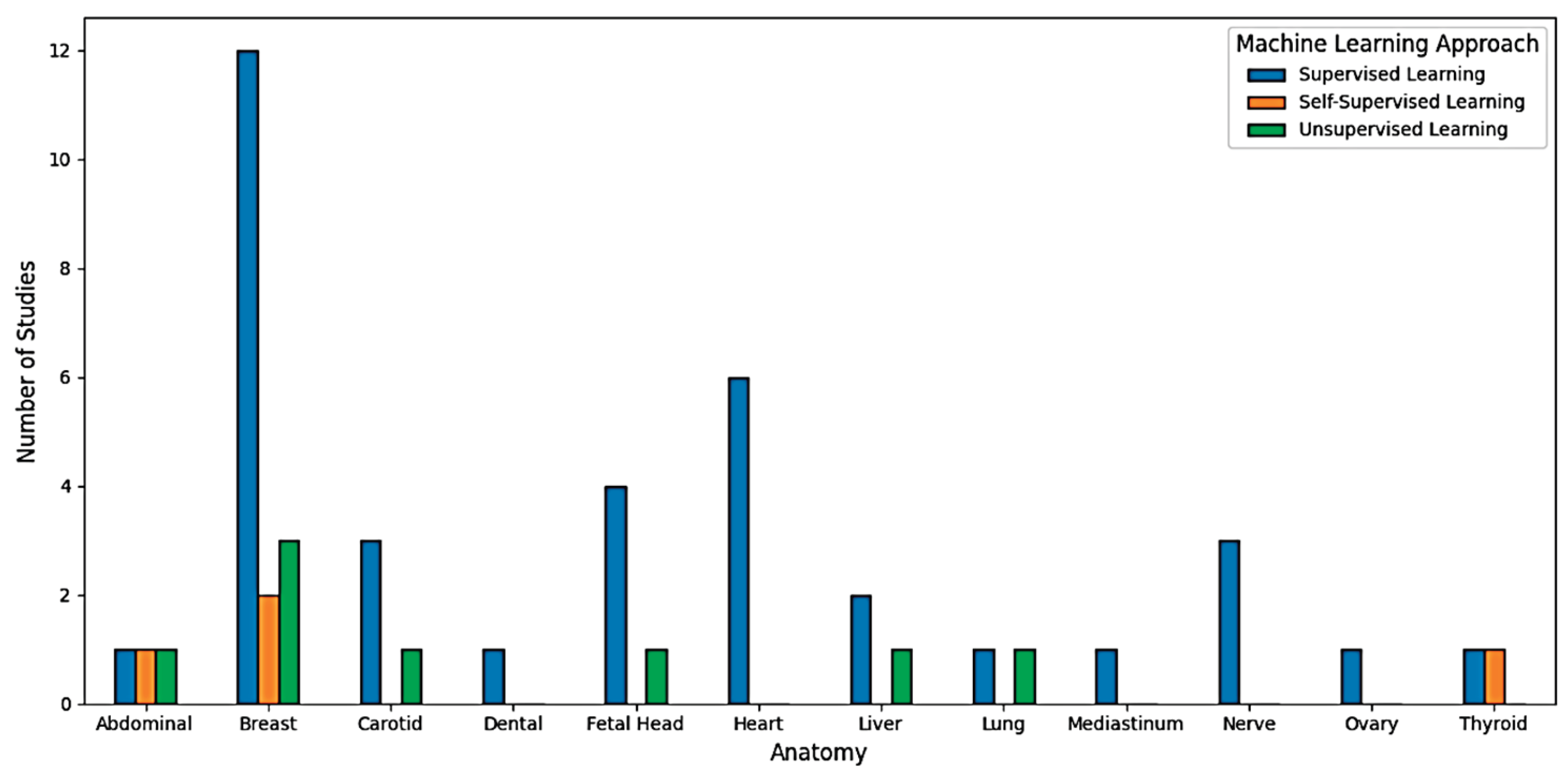
- Anatomy (Breast/ Fetal/ Cardiac/ Abdominal/Musculoskeletal/Others);
- Imaging dimensionality(2D/3D/Videos);
- Target noise type (Speckle, Gaussian noise);
- Learning paradigm (Supervised Learning/Self Supervised/Unsupervised);
- Deep learning architecture (CNN/Unet/GAN/Transformer);
- Summary of the proposed denoising methodology;
- Training data characteristics;
- Evaluation metrics used for quantitative assessment;
- Baseline methods used for comparison;
- Summary of performance results;
- Limitations of the study;
- Future research directions stated;
2.5. Data Handling and Summary
2.6. Limitations
3. Results and Discussion
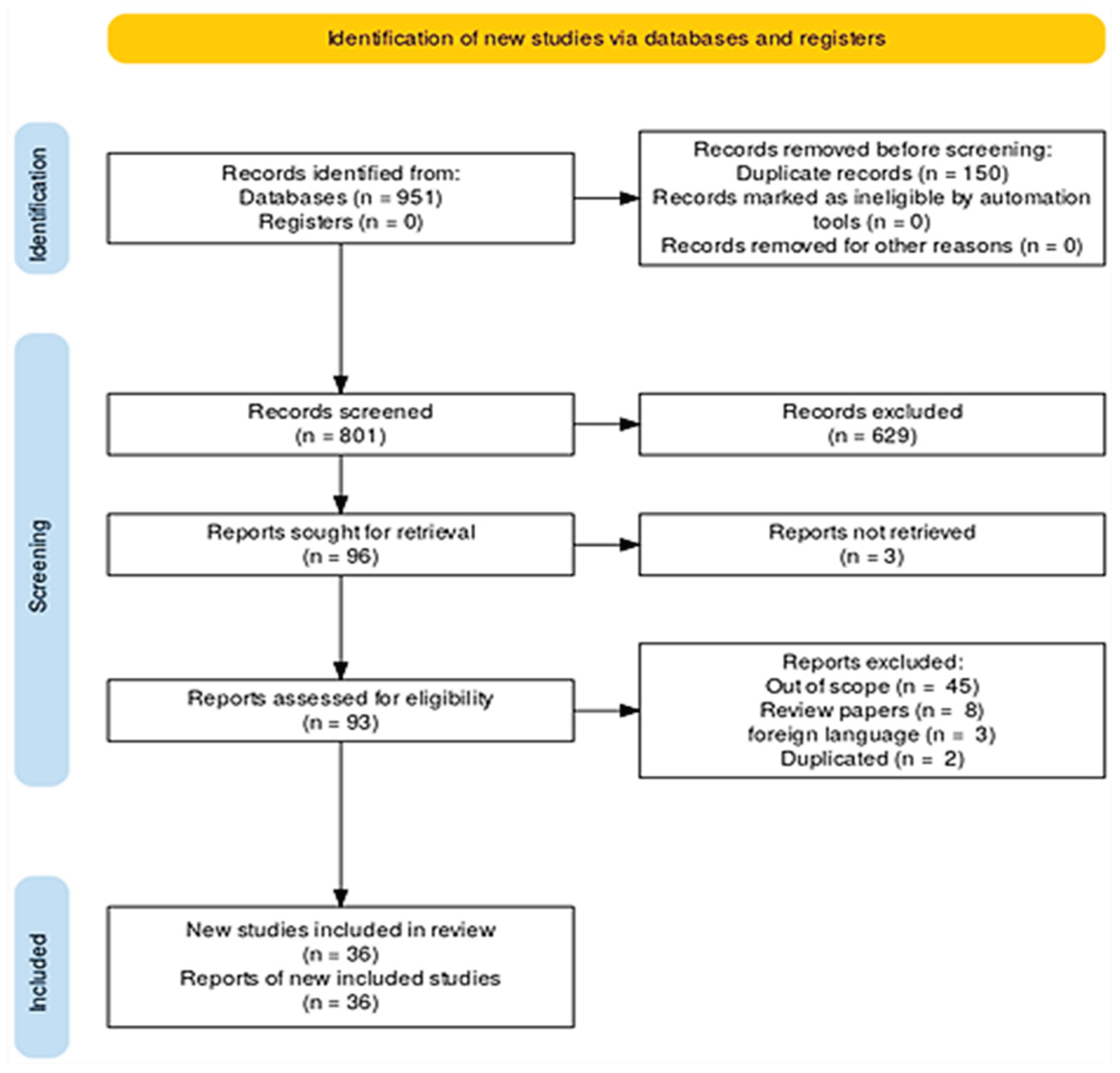
3.1. Study Selection
3.2. Characteristics of the Studies
3.3. Training Data and Noise Modelling Strategy
3.4. Evaluation Metrics
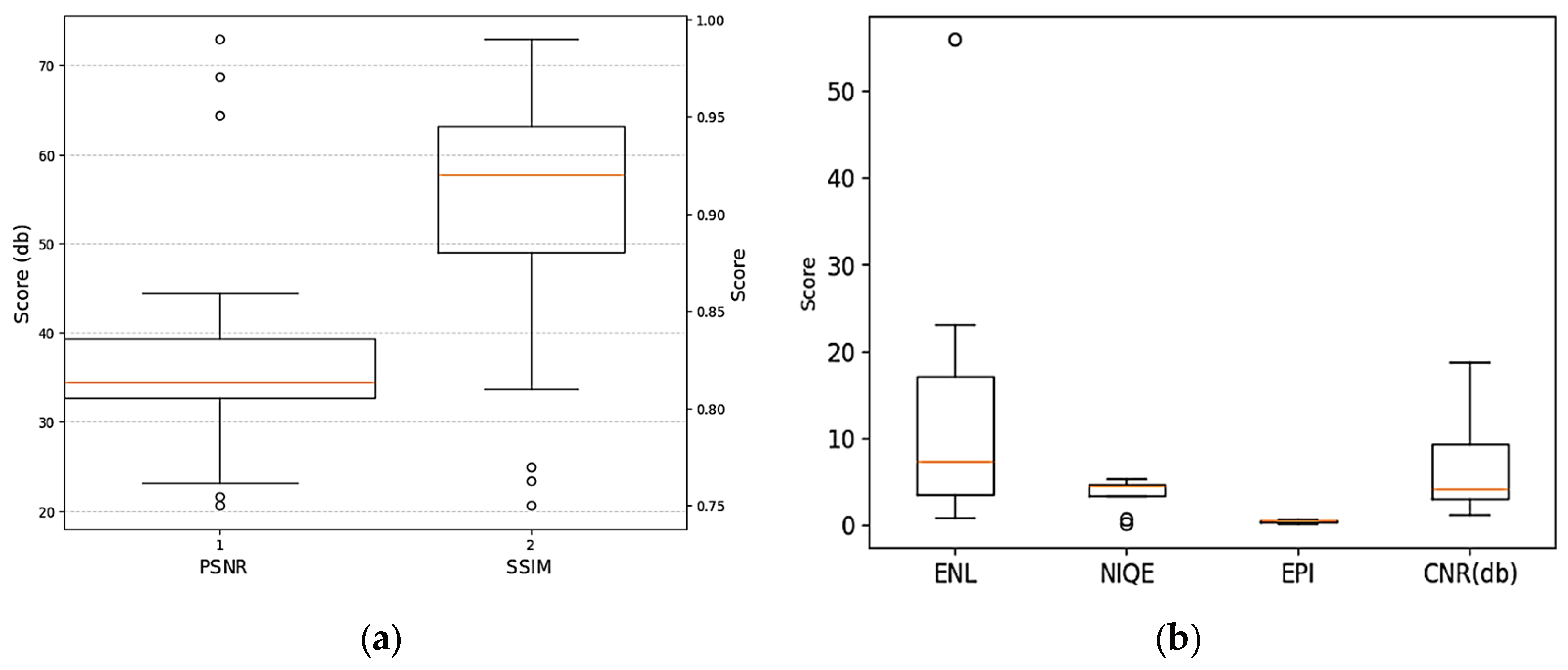
3.5. Meta Analysis of Quantitative Metrices
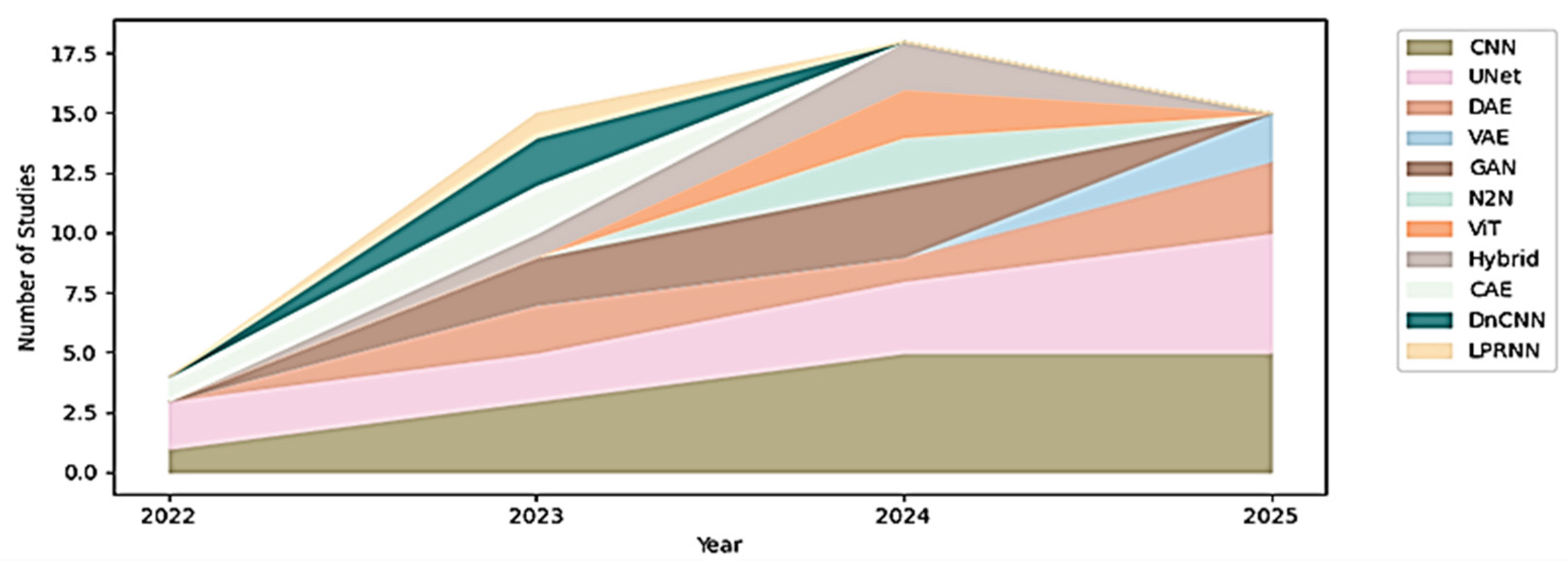
3.6. Methodological Trends
3.7. Identified Gap
4. Conclusions
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Conflicts of Interest
Abbreviations
| US | Ultrasound |
| PRISMA-DTA | Preferred Reporting Items for a Systematic Review and Meta-Analysis of Diagnostic Test Accuracy studies |
| CNN | Convolutional neural network |
| GAN | Generative adversarial neural network |
| VAE | Variational autoencoder |
| POCUS | Point of care ultrasound |
| CT | Computed tomography |
| MRI | Magnetic resonance imaging |
| SL | Supervised learning |
| SSL | Self-supervised learning |
| USL | Unsupervised learning |
| MSE | Mean squared error |
| RMSE | Root mean squared error |
| MSSIM | Mean structural similarity index |
| ENL | Equivalent number of looks |
| CNR | Contrast-to-noise ratio |
| SRN | Signal-to-noise ratio |
| FSIM | Feature similarity index measure |
| EPI | Edge preservation index |
| NIQE | Natural image quality evaluator |
| PIQE | Perception-based image quality evaluator |
| FOM | Figure of merit |
| ISNR | Improvement in signal-to-noise ratio |
| SI | Speckle index |
| SRE | Signal-to-reconstruction error |
| UIQ | Universal image quality |
| SSIM | Structural similarity index |
| PSNR | Peak signal to noise ratio |
References
- S. Wolstenhulme, “Peter Hoskins, Kevin Martin and Abigail Thrush (eds). Diagnostic Ultrasound: Physics and Equipment,” Ultrasound, vol. 28, no. 1, pp. 62-62, 2020. [CrossRef]
- T. Szabo, “Diagnostic Ultrasound Imaging—Inside Out,” 09/01 2004.
- M. Mahesh, “The Essential Physics of Medical Imaging, Third Edition,” Medical Physics, vol. 40, no. 7, p. 077301, 2013. [CrossRef]
- M. R. Torloni et al., “Safety of ultrasonography in pregnancy: WHO systematic review of the literature and meta-analysis,” Ultrasound in Obstetrics & Gynecology, vol. 33, no. 5, pp. 599-608, 2009. [CrossRef]
- C. M. I. Quarato et al., “A Review on Biological Effects of Ultrasounds: Key Messages for Clinicians,” (in eng), Diagnostics (Basel), vol. 13, no. 5, Feb 23 2023. [CrossRef]
- P. R. Atkinson et al., “Does Point-of-Care Ultrasonography Improve Clinical Outcomes in Emergency Department Patients With Undifferentiated Hypotension? An International Randomized Controlled Trial From the SHoC-ED Investigators,” (in eng), Ann Emerg Med, vol. 72, no. 4, pp. 478-489, Oct 2018. [CrossRef]
- A. P. Sarvazyan, M. W. Urban, and J. F. Greenleaf, “Acoustic waves in medical imaging and diagnostics,” (in eng), Ultrasound Med Biol, vol. 39, no. 7, pp. 1133-46, Jul 2013. [CrossRef]
- S. P. Grogan and C. A. Mount, “Ultrasound Physics and Instrumentation,” in StatPearls. Treasure Island (FL): StatPearls Publishing, Copyright © 2025, StatPearls Publishing LLC., 2025.
- G. F. Pinton, G. E. Trahey, and J. J. Dahl, “Sources of image degradation in fundamental and harmonic ultrasound imaging using nonlinear, full-wave simulations,” (in eng), IEEE Trans Ultrason Ferroelectr Freq Control, vol. 58, no. 4, pp. 754-65, Apr 2011. [CrossRef]
- M. M. Goodsitt, P. L. Carson, S. Witt, D. L. Hykes, and J. M. Kofler, Jr., “Real-time B-mode ultrasound quality control test procedures. Report of AAPM Ultrasound Task Group No. 1,” (in eng), Med Phys, vol. 25, no. 8, pp. 1385-406, Aug 1998. [CrossRef]
- C. L. Moore and J. A. Copel, “Point-of-care ultrasonography,” (in eng), N Engl J Med, vol. 364, no. 8, pp. 749-57, Feb 24 2011. [CrossRef]
- N. Yahya, N. S. Kamel, and A. S. Malik, “Subspace-based technique for speckle noise reduction in ultrasound images,” BioMedical Engineering OnLine, vol. 13, no. 1, p. 154, 2014/11/25 2014. [CrossRef]
- A. Sivaanpu et al., “Speckle Noise Reduction Techniques in Ultrasound Imaging: A comprehensive review of the last two decades (2005-2024),” (in eng), Comput Methods Programs Biomed, vol. 274, p. 109150, Nov 6 2025. [CrossRef]
- J. S. Lee, “Digital Image Enhancement and Noise Filtering by Use of Local Statistics,” IEEE Transactions on Pattern Analysis and Machine Intelligence, vol. PAMI-2, no. 2, pp. 165-168, 1980. [CrossRef]
- V. S. Frost, J. A. Stiles, K. S. Shanmugan, and J. C. Holtzman, “A model for radar images and its application to adaptive digital filtering of multiplicative noise,” (in eng), IEEE Trans Pattern Anal Mach Intell, vol. 4, no. 2, pp. 157-66, Feb 1982. [CrossRef]
- Y. Yongjian and S. T. Acton, “Speckle reducing anisotropic diffusion,” IEEE Transactions on Image Processing, vol. 11, no. 11, pp. 1260-1270, 2002. [CrossRef]
- A. Pizurica and W. Philips, “Estimating the probability of the presence of a signal of interest in multiresolution single- and multiband image denoising,” (in eng), IEEE Trans Image Process, vol. 15, no. 3, pp. 654-65, Mar 2006. [CrossRef]
- P. Coupé, P. Hellier, C. Kervrann, and C. Barillot, “Nonlocal means-based speckle filtering for ultrasound images,” (in eng), IEEE Trans Image Process, vol. 18, no. 10, pp. 2221-9, Oct 2009. [CrossRef]
- C. Duarte-Salazar, A. Castro-Ospina, M. Becerra, and E. Delgado-Trejos, “Speckle Noise Reduction in Ultrasound Images for Improving the Metrological Evaluation of Biomedical Applications: An Overview,” IEEE Access, vol. PP, pp. 1-1, 01/17 2020. [CrossRef]
- S. Wu, Q. Zhu, and Y. Xie, “Evaluation of various speckle reduction filters on medical ultrasound images,” (in eng), Annu Int Conf IEEE Eng Med Biol Soc, vol. 2013, pp. 1148-51, 2013. [CrossRef]
- J. Lehtinen et al., “Noise2Noise: Learning Image Restoration without Clean Data,” presented at the Proceedings of the 35th International Conference on Machine Learning, Proceedings of Machine Learning Research, 2018. [Online]. Available: https://proceedings.mlr.press/v80/lehtinen18a.html.
- A. Pizurica, A. m. Wink, E. Vansteenkiste, W. Philips, and J. Roerdink, “A Review of Wavelet Denoising in MRI and Ultrasound Brain Imaging,” Current Medical Imaging Reviews, vol. 2, pp. 247-260, 05/01 2006. [CrossRef]
- C. Tian, Y. xu, L. Fei, and K. Yan, Deep Learning for Image Denoising: A Survey. 2019, pp. 563-572.
- N. Gupta, A. P. Shukla, and S. Agarwal, “Despeckling of Medical Ultrasound Images: A Technical Review,” (in English), Int J Inf Eng Electron Bus. [CrossRef]
- A. Kaur and G. Dong, “A Complete Review on Image Denoising Techniques for Medical Images,” Neural Process. Lett., vol. 55, no. 6, pp. 7807–7850, 2023. [CrossRef]
- S. V. Mohd Sagheer and S. N. George, “A review on medical image denoising algorithms,” Biomedical Signal Processing and Control, vol. 61, p. 102036, 2020/08/01/ 2020. [CrossRef]
- W. Cui, Z. Pan, X. Li, Y. Tang, and S. Sun, “Physical imaging model-guided deep variational despeckling framework for ultrasound images,” Knowledge-Based Systems, vol. 329, p. 114409, 2025/11/04/ 2025. [CrossRef]
- A. Soy and V. V. Prakash, “Medical Image Denoising using Deep Convolutional Autoencoders for Ultrasound,” in 2025 International Conference on Automation and Computation (AUTOCOM), 4-6 March 2025 2025, pp. 262-267. [CrossRef]
- J. Chi, J. Miao, J. H. Chen, H. Wang, X. Yu, and Y. Huang, “DSTAN: A Deformable Spatial-temporal Attention Network with Bidirectional Sequence Feature Refinement for Speckle Noise Removal in Thyroid Ultrasound Video,” (in eng), J Imaging Inform Med, vol. 37, no. 6, pp. 3264-3281, Dec 2024. [CrossRef]
- A. Kavand and M. Bekrani, “Speckle noise removal in medical ultrasonic image using spatial filters and DnCNN,” Multimedia Tools and Applications, vol. 83, no. 15, pp. 45903-45920, 2024/05/01 2024. [CrossRef]
- M. Jha, R. Gupta, and R. Saxena, “Noise cancellation of polycystic ovarian syndrome ultrasound images using robust two-dimensional fractional fourier transform filter and VGG-16 model,” International Journal of Information Technology, vol. 16, pp. 2497 - 2504, 2024.
- N. A. El-Hag, H. M. El-Hoseny, and F. Harby, “DNN-driven hybrid denoising: advancements in speckle noise reduction,” Journal of Optics, vol. 54, no. 5, pp. 3126-3135, 2025/11/01 2025. [CrossRef]
- N. Reddy, C. Chitteti, S. Yesupadam, V. Desanamukula, S. S. Vellela, and N. Bommagani, “Enhanced Speckle Noise Reduction in Breast Cancer Ultrasound Imagery Using a Hybrid Deep Learning Model,” Ingénierie des systèmes d information, vol. 24, pp. 1063-1071, 08/31 2023. [CrossRef]
- M. Khalifa, H. M. Hamza, and K. M. Hosny, “De-speckling of medical ultrasound image using metric-optimized knowledge distillation,” Scientific Reports, vol. 15, no. 1, p. 23703, 2025/07/03 2025. [CrossRef]
- P. N. Devi et al., “Denoising of Medical Ultrasound Images Using Deep Learning With Channel And Spatial Attention Based Modified U-Net,” 2024 15th International Conference on Computing Communication and Networking Technologies (ICCCNT), pp. 1-5, 2024.
- W. T. Hsu, O. Agbodike, and J. Chen, “Attentive U-Net with Physics-Informed Loss for Noise Suppression in Medical Ultrasound Images,” in 2024 10th International Conference on Applied System Innovation (ICASI), 17-21 April 2024 2024, pp. 409-411. [CrossRef]
- S. Satish, N. Herald Anantha Rufus, M. Antony Freeda Rani, and R. Senthil Rama, “U-Net-Based Denoising Autoencoder Network for De-Speckling in Fetal Ultrasound Images,” Singapore, 2023: Springer Nature Singapore, in Fourth International Conference on Image Processing and Capsule Networks, pp. 323-338.
- P. Monkam et al., “US-Net: A lightweight network for simultaneous speckle suppression and texture enhancement in ultrasound images,” Computers in Biology and Medicine, vol. 152, p. 106385, 2023/01/01/ 2023. [CrossRef]
- R. S. S, S. S, R. S. S. K, B. V, S. Saranya, and B. Babu, “Ultrasound Image Denoising Using Cascaded Median Filter and Autoencoder,” in 2023 4th International Conference on Smart Electronics and Communication (ICOSEC), 20-22 Sept. 2023 2023, pp. 296-302. [CrossRef]
- T. Slimi, R. Ferjaoui, and A. B. Khalifa, “Ultrasound Imaging Enhancement Using Denoising AutoEncoders,” in 2025 IEEE 22nd International Multi-Conference on Systems, Signals & Devices (SSD), 17-20 Feb. 2025 2025, pp. 209-214. [CrossRef]
- S. Bhute, S. Mandal, and D. Guha, “Speckle Noise Reduction in Ultrasound Images using Denoising Auto-encoder with Skip connection,” in 2024 IEEE South Asian Ultrasonics Symposium (SAUS), 27-29 March 2024 2024, pp. 1-4. [CrossRef]
- Y. Jiménez-Gaona, M. J. Rodríguez-Alvarez, L. Escudero, C. Sandoval, and V. Lakshminarayanan, “Ultrasound breast images denoising using generative adversarial networks (GANs),” Intelligent Data Analysis, vol. 28, no. 6, pp. 1661-1678, 2024. [CrossRef]
- A. Sivaanpu et al., “A Lightweight Ultrasound Image Denoiser Using Parallel Attention Modules and Capsule Generative Adversarial Network,” Informatics in Medicine Unlocked, vol. 50, p. 101569, 2024/01/01/ 2024. [CrossRef]
- J. Liu et al., “Speckle noise reduction for medical ultrasound images based on cycle-consistent generative adversarial network,” Biomedical Signal Processing and Control, vol. 86, p. 105150, 2023/09/01/ 2023. [CrossRef]
- J. Gan, L. Wang, Z. Liu, and J. Wang, “Multi-scale ultrasound image denoising algorithm based on deep learning model for super-resolution reconstruction,” presented at the Proceedings of the 2023 4th International Conference on Control, Robotics and Intelligent System, Guangzhou, China, 2023. [Online]. Available: . [CrossRef]
- Y. Chen and Z. Guo, “TranSpeckle: An edge-protected transformer for medical ultrasound image despeckling,” IET Image Processing, vol. 17, no. 14, pp. 4014-4027, 2023. [CrossRef]
- D. Oliveira-Saraiva et al., “Make It Less Complex: Autoencoder for Speckle Noise Removal—Application to Breast and Lung Ultrasound,” Journal of Imaging, vol. 9, no. 10, p. 217, 2023. [Online]. Available: https://www.mdpi.com/2313-433X/9/10/217.
- Y. Li, X. Zeng, Q. Dong, and X. Wang, “RED-MAM: A residual encoder-decoder network based on multi-attention fusion for ultrasound image denoising,” Biomedical Signal Processing and Control, vol. 79, p. 104062, 2023/01/01/ 2023. [CrossRef]
- M. Jiang et al., “Controllable Deep Learning Denoising Model for Ultrasound Images Using Synthetic Noisy Image,” Cham, 2024: Springer Nature Switzerland, in Advances in Computer Graphics, pp. 297-308.
- O. Mahmoudi Mehr, M. R. Mohammadi, and M. Soryani, “Deep Learning-Based Ultrasound Image Despeckling by Noise Model Estimation,” (in eng), IRANIAN JOURNAL OF ELECTRICAL AND ELECTRONIC ENGINEERING, Research Paper vol. 19, no. 3, pp. 1-13, 2023. [CrossRef]
- Y. Chen, Z. Guo, J. Yuan, X. Li, and H. Yu, “Dual-TranSpeckle: Dual-pathway transformer based encoder-decoder network for medical ultrasound image despeckling,” Computers in Biology and Medicine, vol. 173, p. 108313, 2024/05/01/ 2024. [CrossRef]
- A. Sivaanpu et al., “Speckle Noise Reduction for Medical Ultrasound Images Using Hybrid CNN-Transformer Network,” IEEE Access, vol. 12, pp. 168607-168625, 2024. [CrossRef]
- Z. Bu, G. Zhou, and Y. Chen, A Complementary Global and Local Knowledge Network for Ultrasound Denoising with Fine-grained Refinement. 2024, pp. 1-5.
- B. B. Vimala et al., “Image Noise Removal in Ultrasound Breast Images Based on Hybrid Deep Learning Technique,” Sensors, vol. 23, no. 3, p. 1167, 2023. [Online]. Available: https://www.mdpi.com/1424-8220/23/3/1167.
- T. Slimi, A. Djeha, and A. B. Khalifa, “Medical Ultrasound Image Improvement Based on Denoising Convolutional Autoencoder,” in 2025 IEEE 22nd International Multi-Conference on Systems, Signals & Devices (SSD), 17-20 Feb. 2025 2025, pp. 715-720. [CrossRef]
- C. Yu, F. Ren, S. Bao, Y. Yang, and X. Xu, “Self-supervised ultrasound image denoising based on weighted joint loss,” Digital Signal Processing, vol. 162, p. 105151, 2025/07/01/ 2025. [CrossRef]
- C. Sun, J. Chi, H. Yu, B. Wu, Z. Li, and Y. Huang, “Self-Supervised Denoising of Thyroid Ultrasound Images Using SE-Module Enhanced U-Net with FPN,” in 2025 37th Chinese Control and Decision Conference (CCDC), 16-19 May 2025 2025, pp. 4212-4217. [CrossRef]
- T.-T. Zhang, H. Shu, K.-Y. Lam, C.-Y. Chow, and A. Li, “Feature decomposition and enhancement for unsupervised medical ultrasound image denoising and instance segmentation,” Applied Intelligence, vol. 53, no. 8, pp. 9548-9561, 2023/04/01 2023. [CrossRef]
- S. Goudarzi and H. Rivaz, Deep ultrasound denoising without clean data (SPIE Medical Imaging). SPIE, 2023.
- N. Chen, Y. Zhang, C. Fan, W. Zhao, C. Wang, and H. Wang, “DiffusionClusNet: Deep Clustering-Driven Diffusion Models for Ultrasound Image Enhancement,” IEEE Transactions on Consumer Electronics, vol. 71, no. 1, pp. 1495-1503, 2025. [CrossRef]
- P. Wei, L. Wang, J. Gan, X. Shi, and M. Shang, “Incorporation of Structural Similarity Index and Regularization Term into Neighbor2Neighbor Unsupervised Learning Model for Efficient Ultrasound Image Data Denoising,” Applied Sciences, vol. 14, no. 17, p. 7988, 2024. [Online]. Available: https://www.mdpi.com/2076-3417/14/17/7988.
- M. Basile et al., “Unsupervised Learning of Speckle Removal from Real Ultrasound Acquisitions without Clean Data,” in 2024 IEEE International Symposium on Medical Measurements and Applications (MeMeA), 26-28 June 2024 2024, pp. 1-6. [CrossRef]
- W. Al-Dhabyani, M. Gomaa, H. Khaled, and A. Fahmy, “Dataset of breast ultrasound images,” Data in Brief, vol. 28, p. 104863, 2020/02/01/ 2020. [CrossRef]
- Ultrasound cases [Online] Available: https://www.ultrasoundcases.info/.
- V. Pedraza, Narvaez, Duran, Munoz, Romero. DDTI Dataset: An open access database of thyroid ultrasound images. [Online]. Available: https://www.kaggle.com/datasets/dasmehdixtr/ddti-thyroid-ultrasound-images/data.
- P. S. Rodrigues. Breast Ultrasound Image. [Online]. Available: https://data.mendeley.com/datasets/wmy84gzngw/1.
- M. H. Yap et al., “Automated Breast Ultrasound Lesions Detection Using Convolutional Neural Networks,” IEEE Journal of Biomedical and Health Informatics, vol. 22, no. 4, pp. 1218-1226, 2018. [CrossRef]
- D. S. Anna Montoya, Hasnin, kaggle446, shirzad, Will Cukierski, and yffud. Ultrasound Nerve Segmentation. [Online]. Available: https://www.kaggle.com/c/ultrasound-nerve-segmentation.
- S. Leclerc et al., “Deep Learning for Segmentation Using an Open Large-Scale Dataset in 2D Echocardiography,” IEEE Transactions on Medical Imaging, vol. 38, no. 9, pp. 2198-2210, 2019. [CrossRef]
- N. L. o. Medicine. MedPix. [Online]. Available: https://lhncbc.nlm.nih.gov/medpix.html.
- I. A. PCOS Dataset. [Online]. Available: https://figshare.com/articles/dataset/PCOS_Dataset/27682557?file=50407062.
- R. Sawyer-Lee, Gimenez, F., Hoogi, A., & Rubin, D. urated Breast Imaging Subset of Digital Database for Screening Mammography (CBIS-DDSM). [CrossRef]
- C. Moreira, I. Amaral, I. Domingues, A. Cardoso, M. J. Cardoso, and J. S. Cardoso, “INbreast: toward a full-field digital mammographic database,” (in eng), Acad Radiol, vol. 19, no. 2, pp. 236-48, Feb 2012. [CrossRef]
- D. d. B. Thomas L. A. van den Heuvel, Chris L. de Korte and Bram van Ginneken. Automated measurement of fetal head circumference using 2D ultrasound images. [Online]. Available: http://doi.org/10.5281/zenodo.1322001.
- A. Momot. Common Carotid Artery Ultrasound Images. [CrossRef]
- W. Gómez-Flores, M. J. Gregorio-Calas, and W. Coelho de Albuquerque Pereira, “BUS-BRA: A breast ultrasound dataset for assessing computer-aided diagnosis systems,” (in eng), Med Phys, vol. 51, no. 4, pp. 3110-3123, Apr 2024. [CrossRef]
- Y. a. Z. Chen, Chunhui and Liu, Li and Feng, Cheng and Dong, Changfeng and Luo, Yongfang and Wan, Xiang. Pretraining deep ultrasound image diagnosis model through video contrastive representation learning. [Online]. Available: https://opendatalab.com/OpenDataLab/US-4.
| Studies | Machine earning paradigm | Deep Learning architecture | Dataset domain (Anatomy) | Metrics |
|---|---|---|---|---|
| Cui et al. [27],Soy et al. [28], Chi et al. [29], Kavand et al. [30], Jha et al. [31], El-Hag at al. [32], Reddy et al. [33] | SL | CNN | Breast, Thyroid, Ovary (PCOS), Carotid Artery, General US | PSNR, SSIM, MSE, RMSE, NIQE, PIQE, ENL, AGM, SSI, EI |
| Khalifa et al. [34], Devi et al. [35], Hsu et al. [36], Satish et al.[37], Monkam et al.[38] | SL | Unet | Breast, Liver, Lung, Fetal (Cardiac/Head), Carotid Artery, General US | PSNR, SSIM, MSE, EPI, ENL, CNR, SNR, AGM |
| Saranya et al.[39], Slimi et al. [40], Bhute et al. [41] | SL | DAE | Breast, General US | PSNR, SSIM, MSE |
| Jiménez-Gaona et al.[42], Sivaanpu et al. [43], Liu et al. [44], Gan et al. [45] | SL | GAN | Breast, Fetal Head, General US | PSNR, SSIM, MSSIM, MSE, RMSE, FOM, FSIM |
| Chen et al. [46],Oliveira et al. [47],Li et al.[48] | SL | CAE | Breast, Lung, Nerve, Cardiac, Fetal Head | PSNR, SSIM, RMSE |
| Jiang et al. [49], Mahmoudi et al. [50] | SL | DnCNN | General US, Carotid Artery | PSNR, SSIM |
| Chen et al. [51],Sivaanpu et al.[52], Bu et al. [53] | SL | Hybrid CNN + Transformer | Fetal Head, Breast, Dental, Cardiac Phantom | PSNR, SSIM, RMSE, MSE, NIQE, ENL, SNR, CNR, ISNR, SI |
| Vimala et al. [54] | SL | LPRNN (CNN+RNN) | ||
| Slimi et al. [55], Yu et al. [56], Sun et al. [57] | SSL | DAE / Unet | Breast, Thyroid, Abdominal, General US | PSNR, SSIM |
| Zhang et al. [58], Goudarzi et al. [59] | USL | CNN / Unet | Nerve | PSNR, SSIM, FSIM, EPI, CNR, SRE, UIQ, MSR |
| Chen et al. [60], Wei et al. [61], Basile et al. [62] | USL | N2N / VAE | Liver, Breast, Abdominal, Heart, Mediastinum | PSNR, SSIM, MSSIM, ENL, MSE, CNR, SNR |
| Study | Dataset | PSNR (dB) | SSIM | Other Metrics score |
|---|---|---|---|---|
| Slimi at al. [55] | BUS-BRA | 33.82 | 0.7625 | - |
| Saranya et al. [39] | PICMUS | 44.48 | 0.935 | - |
| Khalifa et al. [34] | Breast US | 40.72 | 0.940 | - |
| Cui et al.[27] | BUID | - | - | ENL=5.71, AGM=38.57, NIQE=4.25, PIQE=31.83 |
| BUSI | - | - | ENL=2.71, AGM=33.24, NIQE=4.74, PIQE=50.61 | |
| CCA | - | - | ENL=0.76, AGM=40.27, NIQE=4.36, PIQE=64.39 | |
| US-case | - | - | ENL=3.50, AGM=65.18, NIQE=5.38, PIQE=50.57 | |
| Chen et al. [60] | US-CASE | 35.19 | 0.90 | - |
| Slimi at al. [40] | BUS-BRA | 20.60 | 0.81 | - |
| Chi at al. [29] | DDTI | 36.82 | 0.93 | - |
| Jiménez-Gaona et al. [42] | BUSI | 39.79 | 0.96 | - |
| Wei at al. [61] | BUSI | 40.03 | - | SSI=0.80 |
| Chen at al. [51] | UNS | 32.82 | 0.9358 | SSI=0.79 |
| CAMUS | 35.29 | 0.9317 | SSI=0.78 | |
| Kavand et al. [30] | BUI + MedPix | 30.50 | 0.97 | UIQ=0.54 |
| Jha et al.[31] | PCOS | 72.96 | 0.99 | UIQ=0.23 |
| Sivaanpu et al. [52] | HC18 | - | 0.965 | ENL=7.26, NIQE=4.61, MSE=13.905, SRE=32.61, UIQ=0.04, |
| Sivaanpu at al. [43] | HC18 | 33.86 | 0.91 | ISNR=23.57dB |
| BUSI | 34.16 | 0.90 | ISNR=18.52dB | |
| El-Hag at al.[32] | BUSI | 28.72 | 0.77 | NIQE=4.50, MSE=157.3, SNR=40.95dB |
| Bhute et al. [41] | BUSI | 23.64 | 0.92 | MSE=0.0048 |
| Bu et al. [53] | HC18 | 40.62 | 0.98 | RMSE=2.33 |
| Hsu et al. [36] | BUSI + US-4 | 42.27 | 0.99 | - |
| Reddy et al. [33] | INBreast + CBIS-DDSM | 64.44 | - | NIQE=0.08, MSE=0.22 |
| Vimala et al. [54] | INBreast + CBIS-DDSM | 68.70 | - | SRE=63.8 |
| Monkam et al. [38] | HC18 | - | - | ENL=15.71, CNR=1.10, SNR=39.32dB, SRE=27.46 |
| BUSI | - | - | ENL=17.04, CNR=4.20, SNR=34.54dB, SRE=17.04 | |
| CAMUS | 32.77 | 0.87 | RMSE=6.05 | |
| Scheme | Dataset | PSNR (dB) | SSIM | Other Metrics score |
|---|---|---|---|---|
| Saranya et al. [39] | Private Fetus | 44.48 | 0.935 | - |
| Cehn et al. [60] | Private (abdominal) | 32.22 | 0.89 | - |
| Sun et al. [57] | Private Thyroid | 32.89 | 0.88 | - |
| Soy et al. [28] | Private Synthetic US | 34.38 | 0.93 | MSE=0.0021 |
| Devi et al. [35] | Clinical US private | 32.22 | 0.88 | MSE=0.0008, UIQ=0.65 |
| Sivaanpu et al. [52] | Private Heart Phantom | - | - | CNR=18.78dB, MSR=3.85 |
| Basile et al. [62] | Abdominal private | - | - | ENL=55.89, MSE=0.004, SSI=0.33, CNR=4.21dB, SNR=8.57dB |
| Jiang et al. [49] | Breast private | 23.13 | 0.81 | - |
| Liu et al. [44] | Private breast, heart, lymph node | 38.13 | - | RMSE=3.25, UIQ=0.98 |
| Vimala et al. [54] | Private CTS nerve | 41.27 | 0.97 | RMSE=0.85, CNR=11.05, EPI=0.18, SRE=51.7, UIQ=0.86, MSR=1.69, |
| Private CTS nerve | 51.78 | 0.86 | RMSE=1.69 | |
| Satish et al. [37] | Private Fetal cardiac | 29.07 | 0.86 | - |
| Goudarzi et al. [59] | Private Heart | 37.27 | 0.90 | MSE=0.006 |
| Private Chicken breast | 37.11 | 0.91 | MSE=0.008 | |
| Private Bovine liver | 31.28 | 0.88 | MSE=0.017 | |
| Li et al. [48] | Private Fetal Heart | 34.31 | 0.88 | RMSE=5.10 |
| Gan et al. [45] | Private Liver | - | - | NIQE=0.58, PIQE=0.79, RMSE=0.39 |
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2026 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).




