Submitted:
30 March 2026
Posted:
01 April 2026
You are already at the latest version
Abstract
Keywords:
1. Introduction
2. Materials and Methods
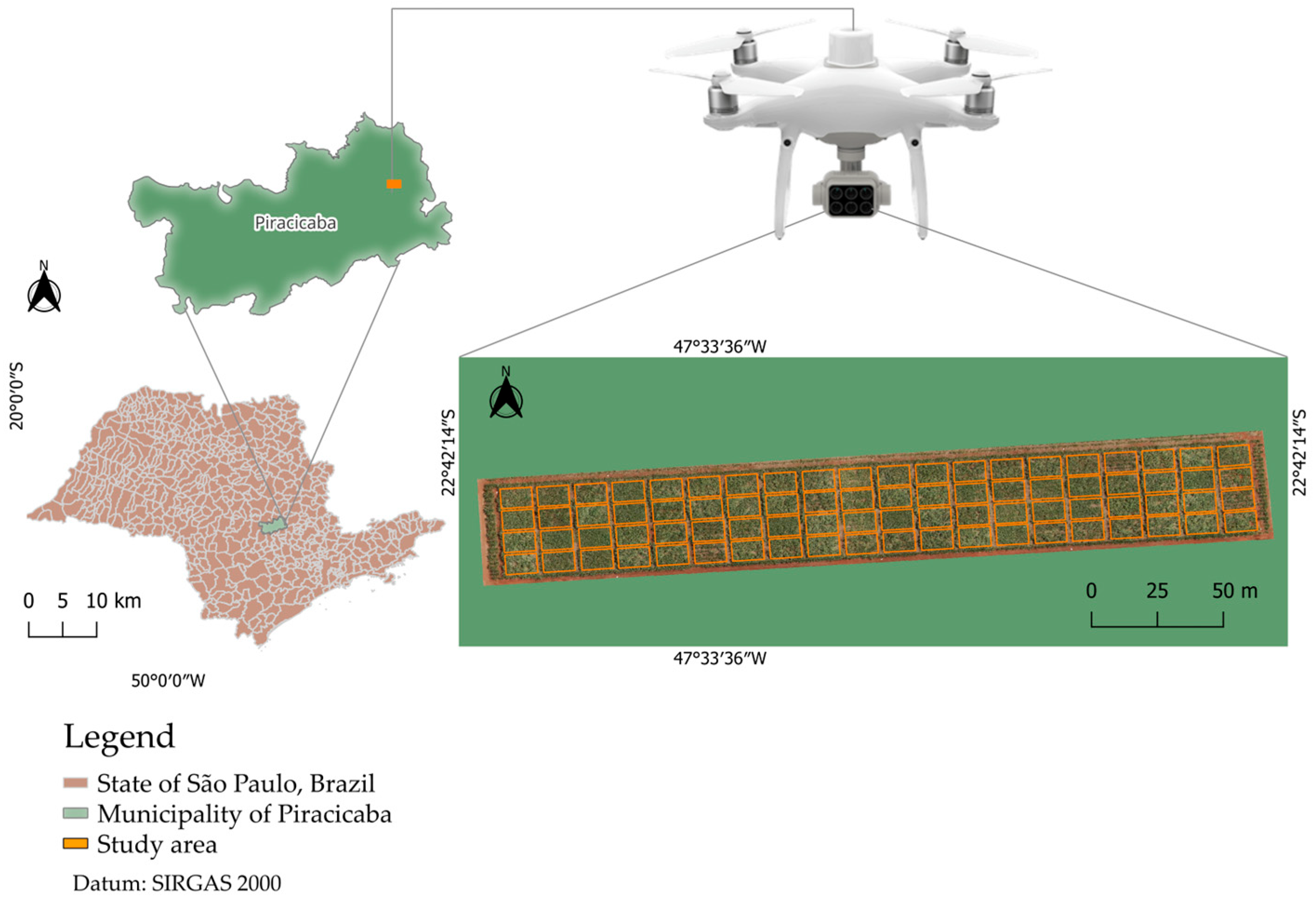
2.1. Study Area and Experimental Conditions
2.2. Field Experimental Design
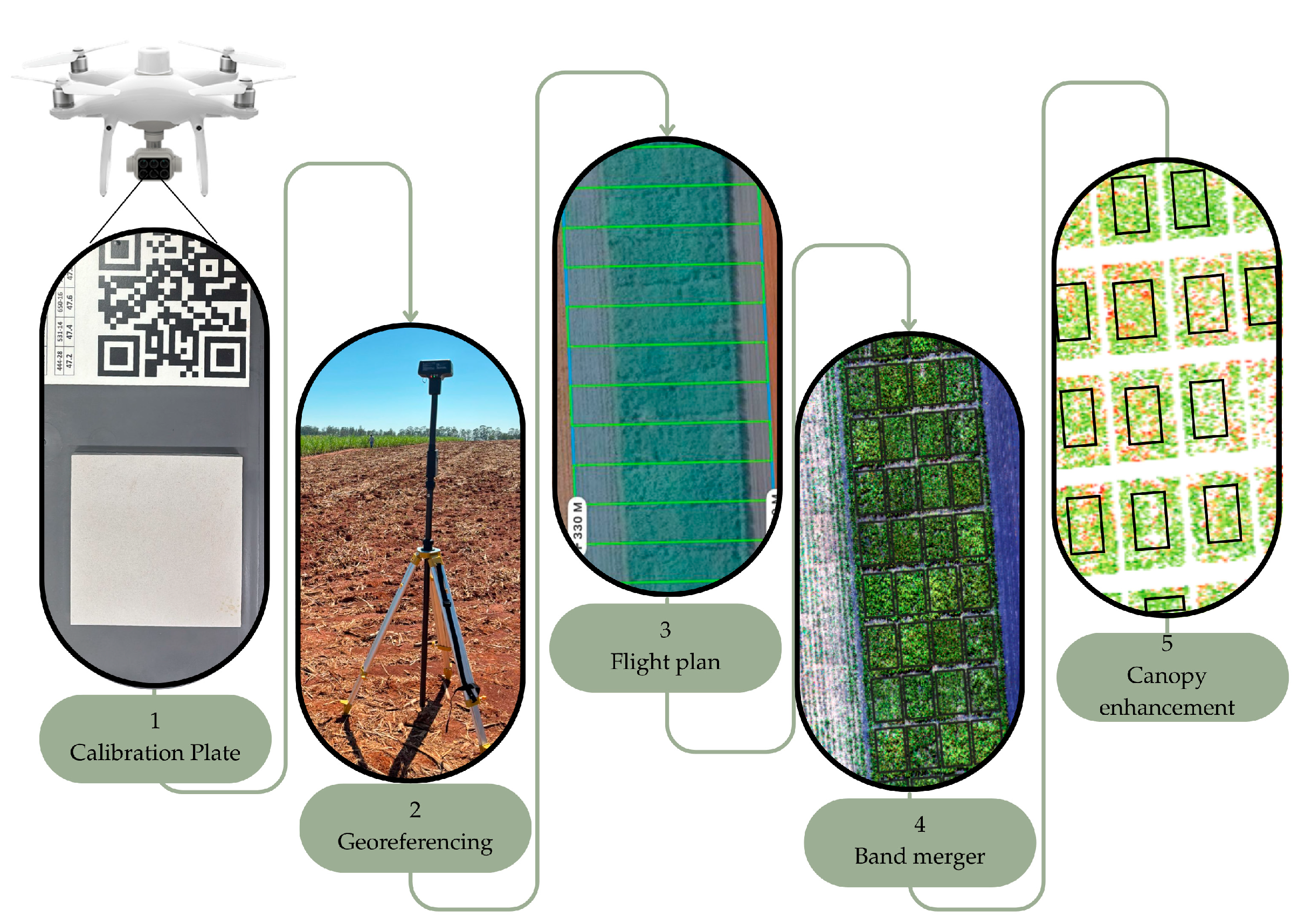
2.3. Acquisition of Multispectral Images Using a UAV
2.4. Acquisition of Leaf Samples
2.5. Acquisition of Laboratory Hyperspectral Data
2.6. Spectral Data Preprocessing
2.6.1. Calibration and Extraction of UAV Multispectral Data
2.6.2. Standardization of Hyperspectral Data
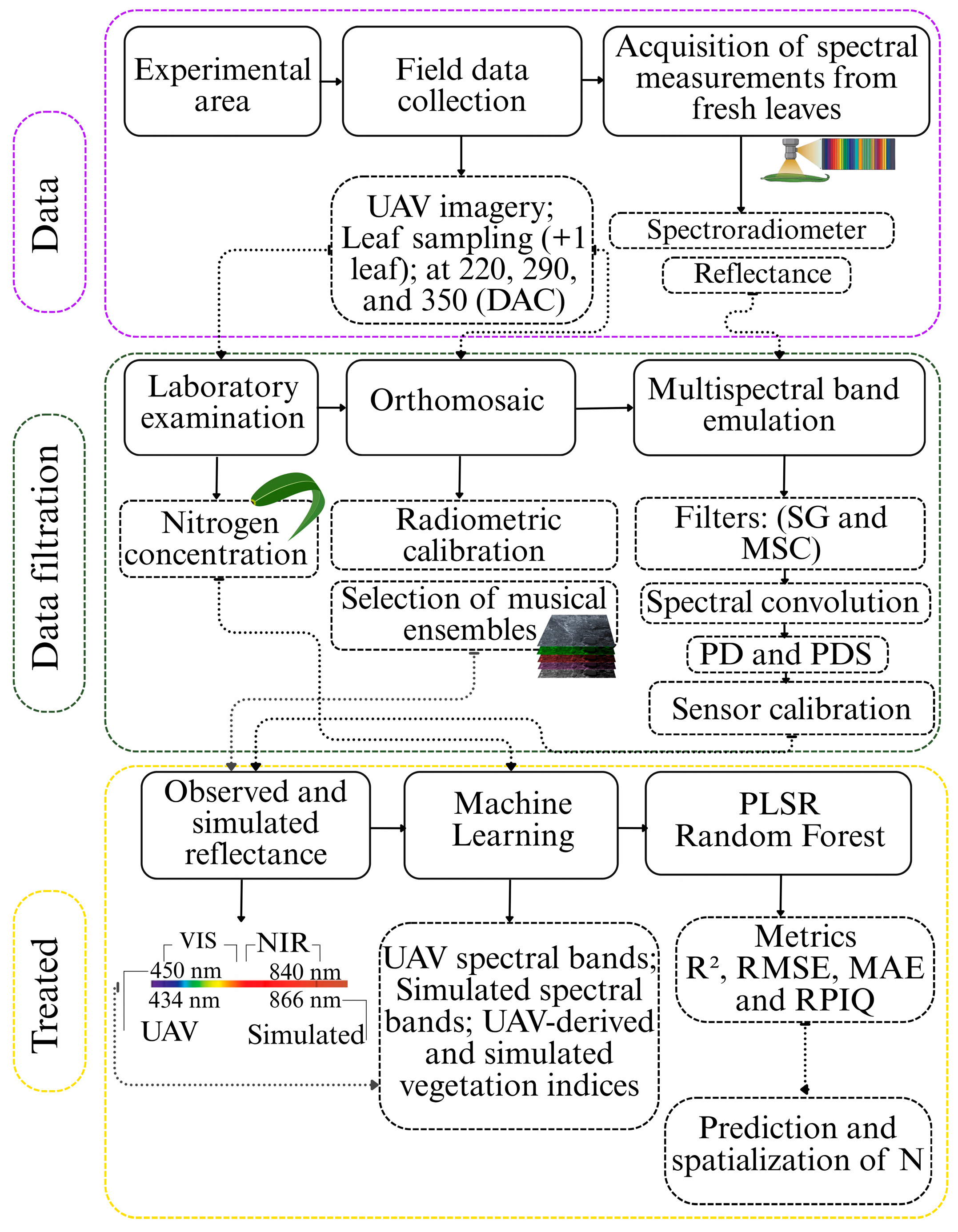
2.7. Methodological Sequence of the Simulation and Modeling Technique
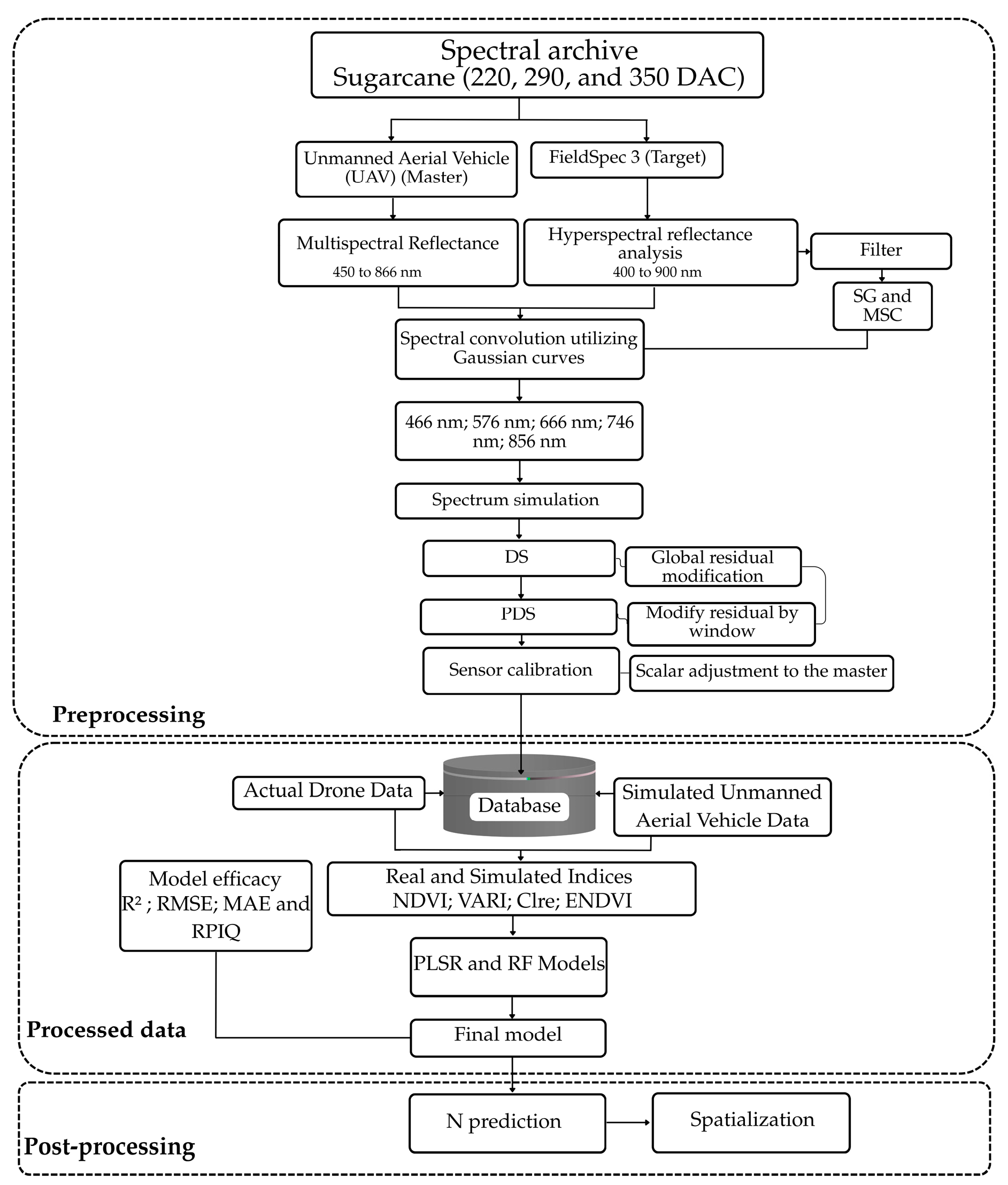
2.7.1. Spectral Convolution
2.7.2. Transfer Calibration Using (DS/PDS)
2.7.3. Sensor-to-Sensor Calibration
2.7.4. Nitrogen-Sensitive Vegetation Indices
2.8. Spearman Correlation
2.9. Modeling of Nitrogen Content Using Machine Learning
2.9.1. Linear Regression
2.9.2. Partial Least Squares Regression (PLSR)
2.9.3. Random Forest (RF)
2.9.4. Performance Indicators
3. Results
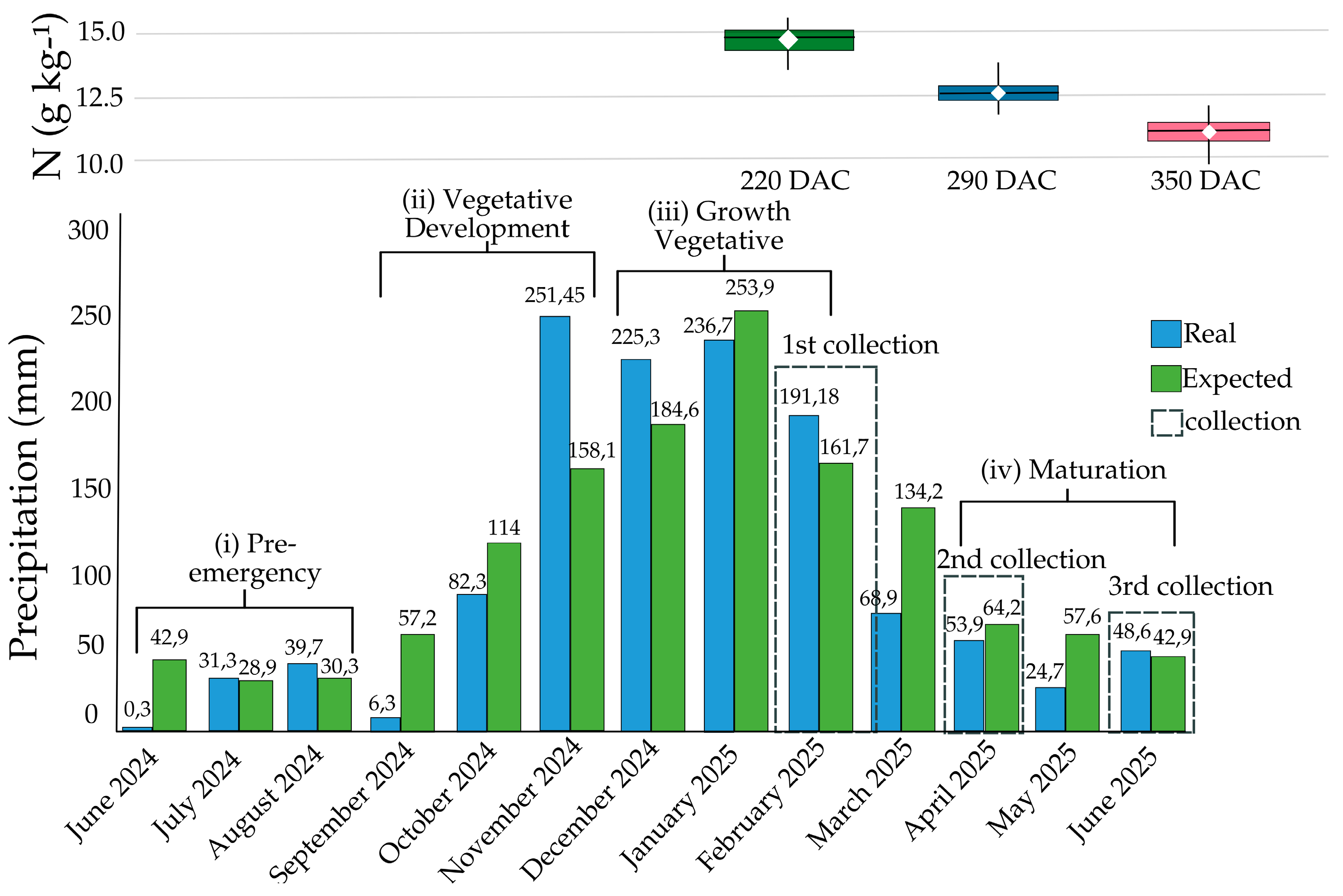
3.1. Leaf Nitrogen Content (LNC) and Precipitation Throughout the Season
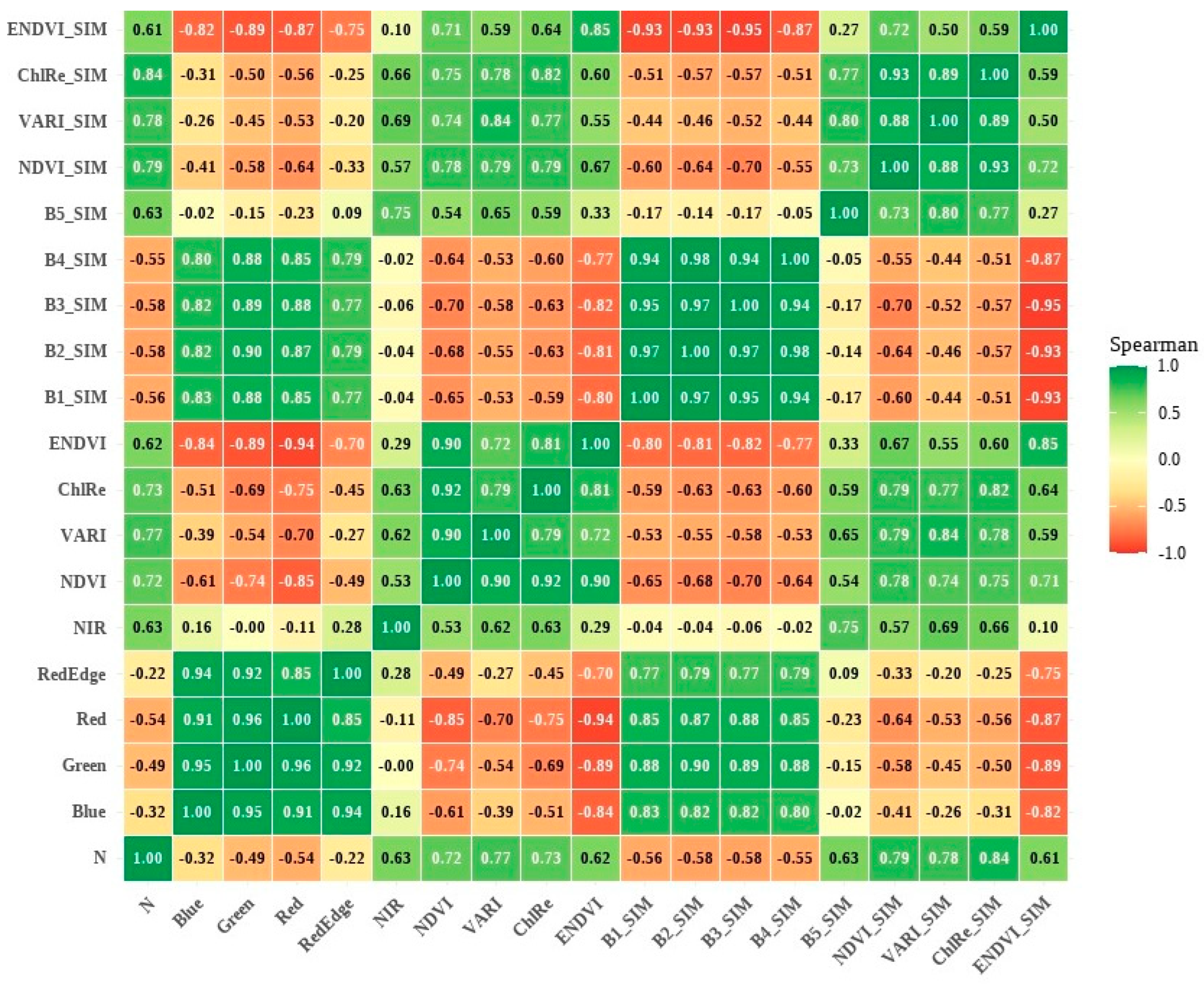
3.2. Spearman Correlation Between Original and Simulated Bands and Indices in Relation to LNC
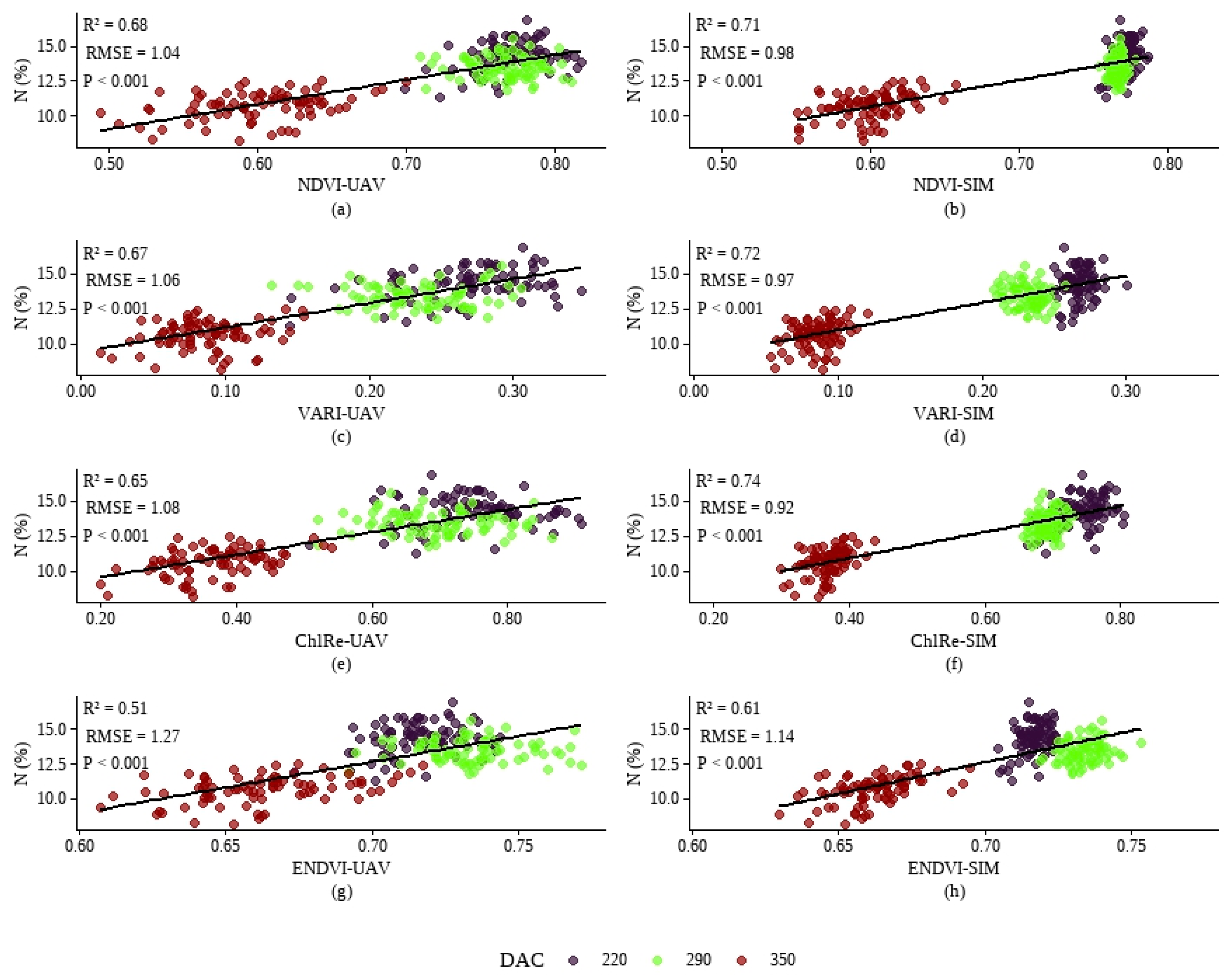
3.3. Exploratory Estimation of Nitrogen Concentration in Sugarcane Using Vegetation Indices from UAV Data and Derived from the Simulated Dataset
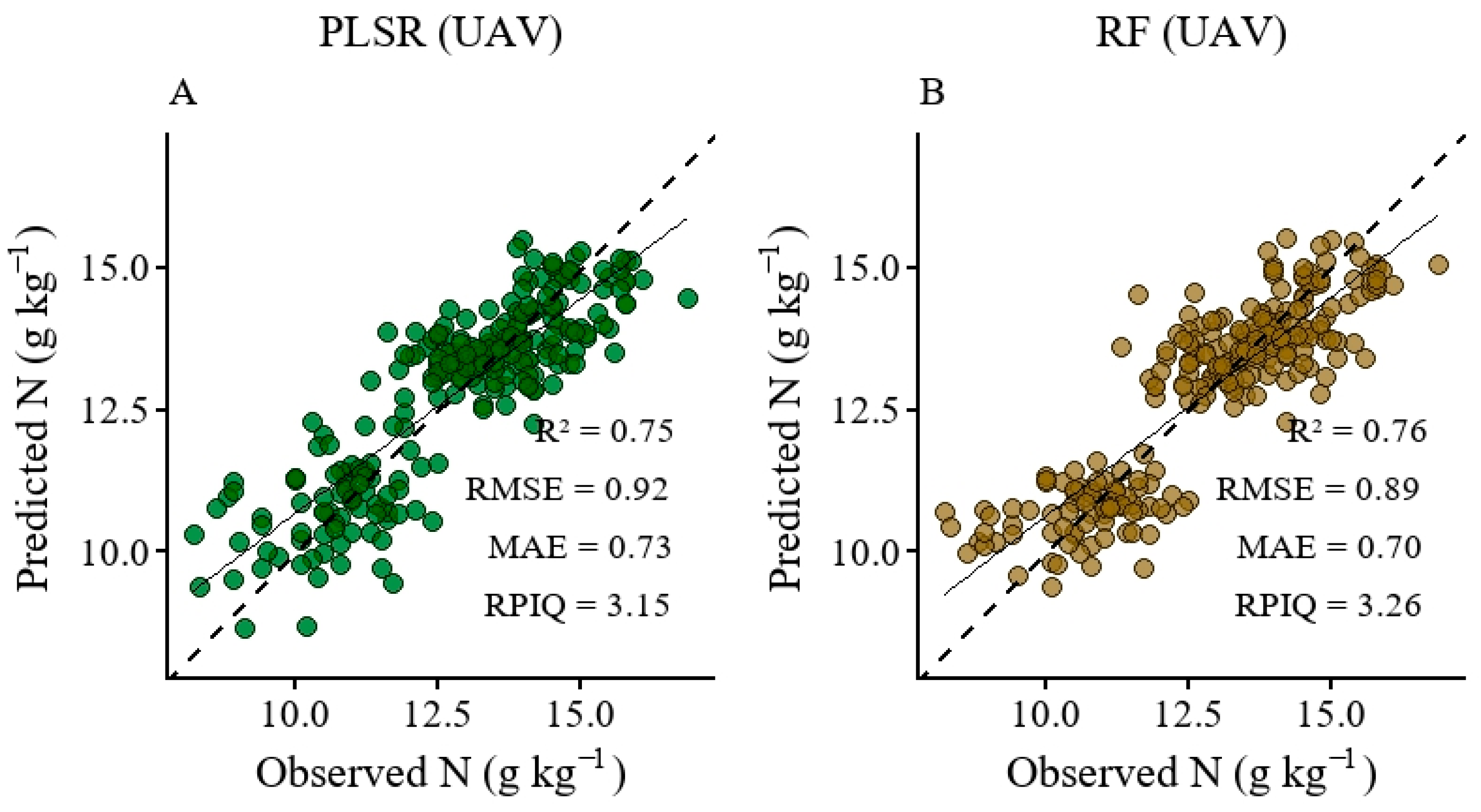
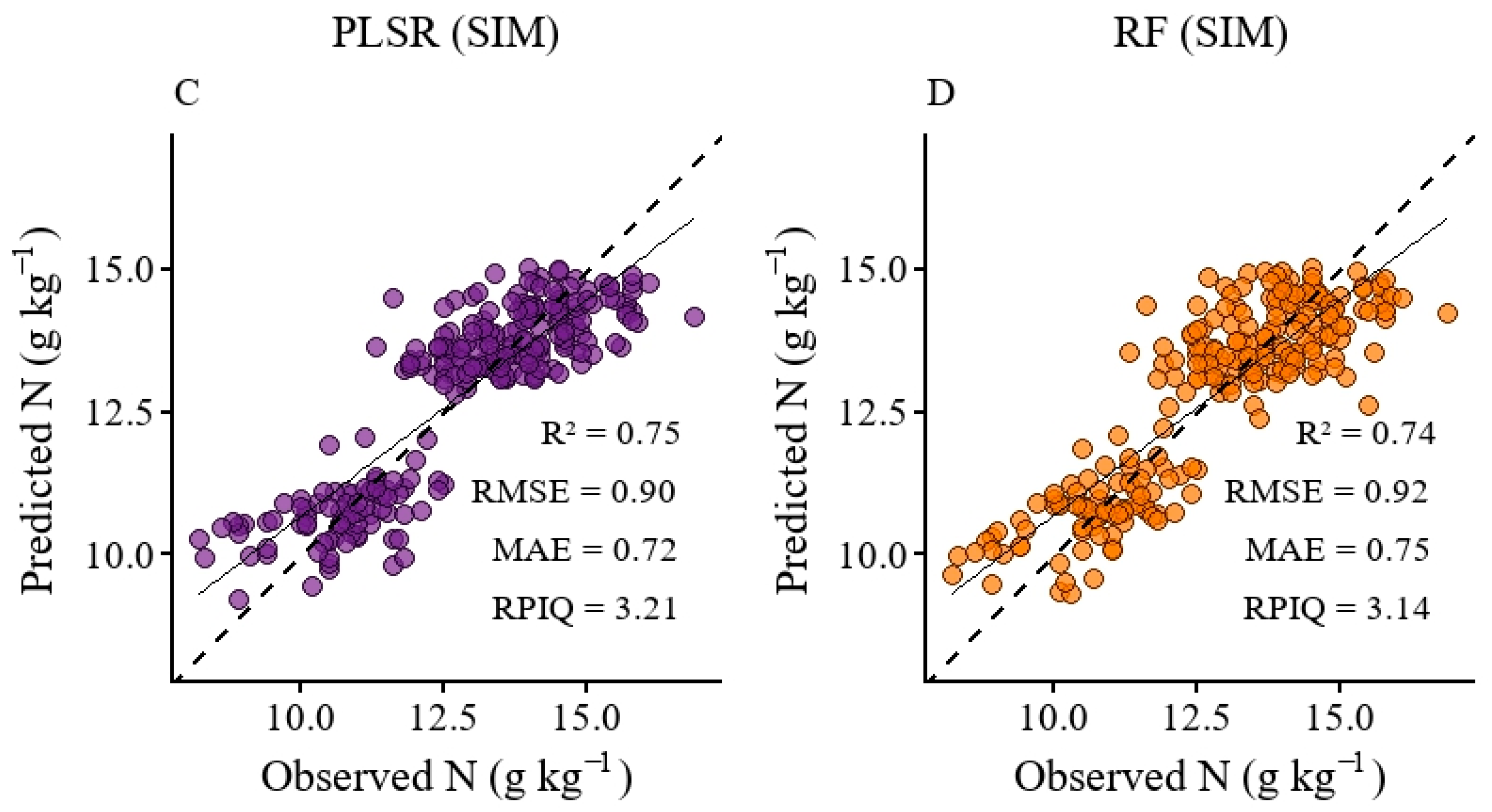
3.4. Modeling of Nitrogen Content Using a UAV-Derived Spectra and Simulated Data
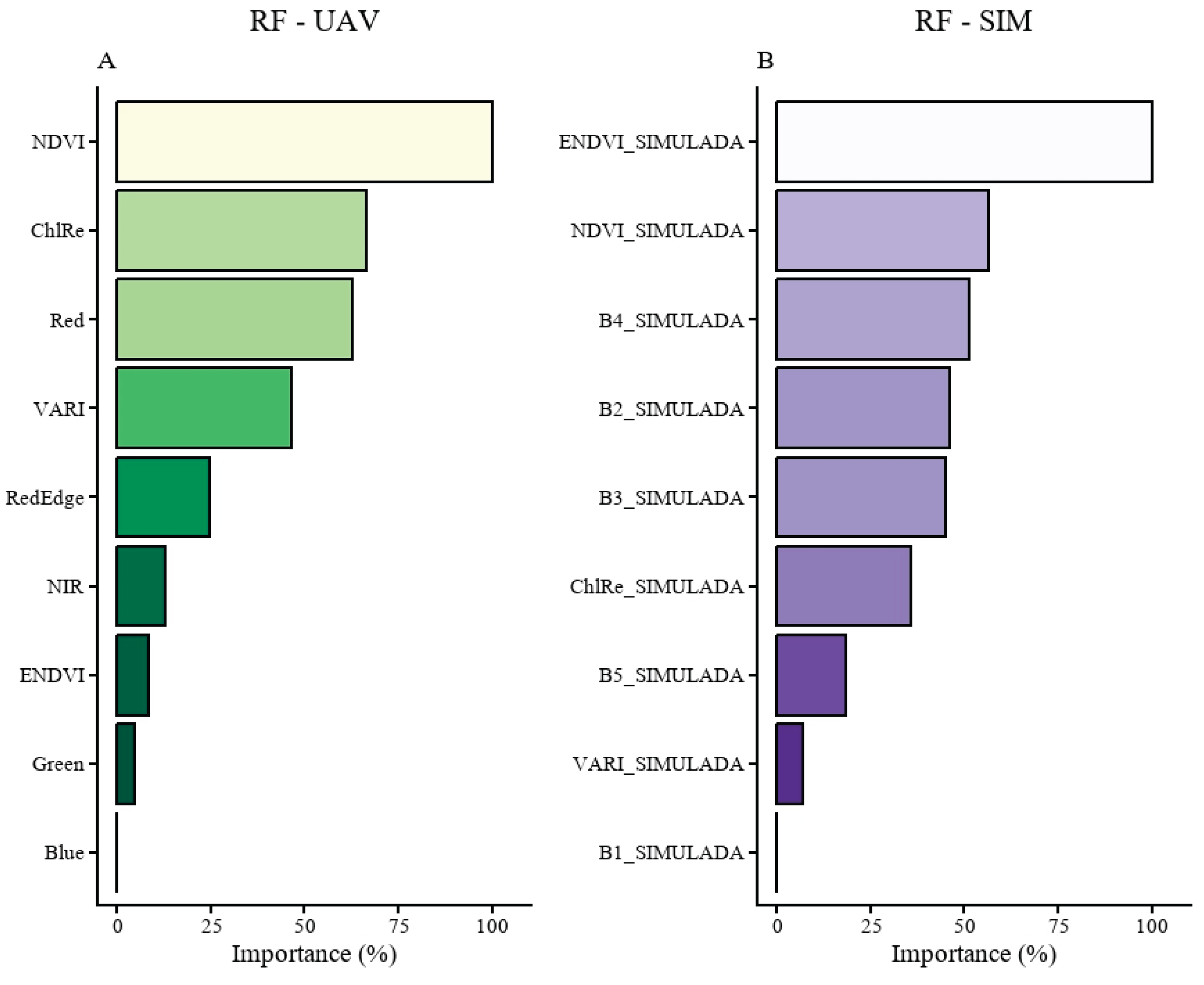
3.5. Variable Importance in Projection
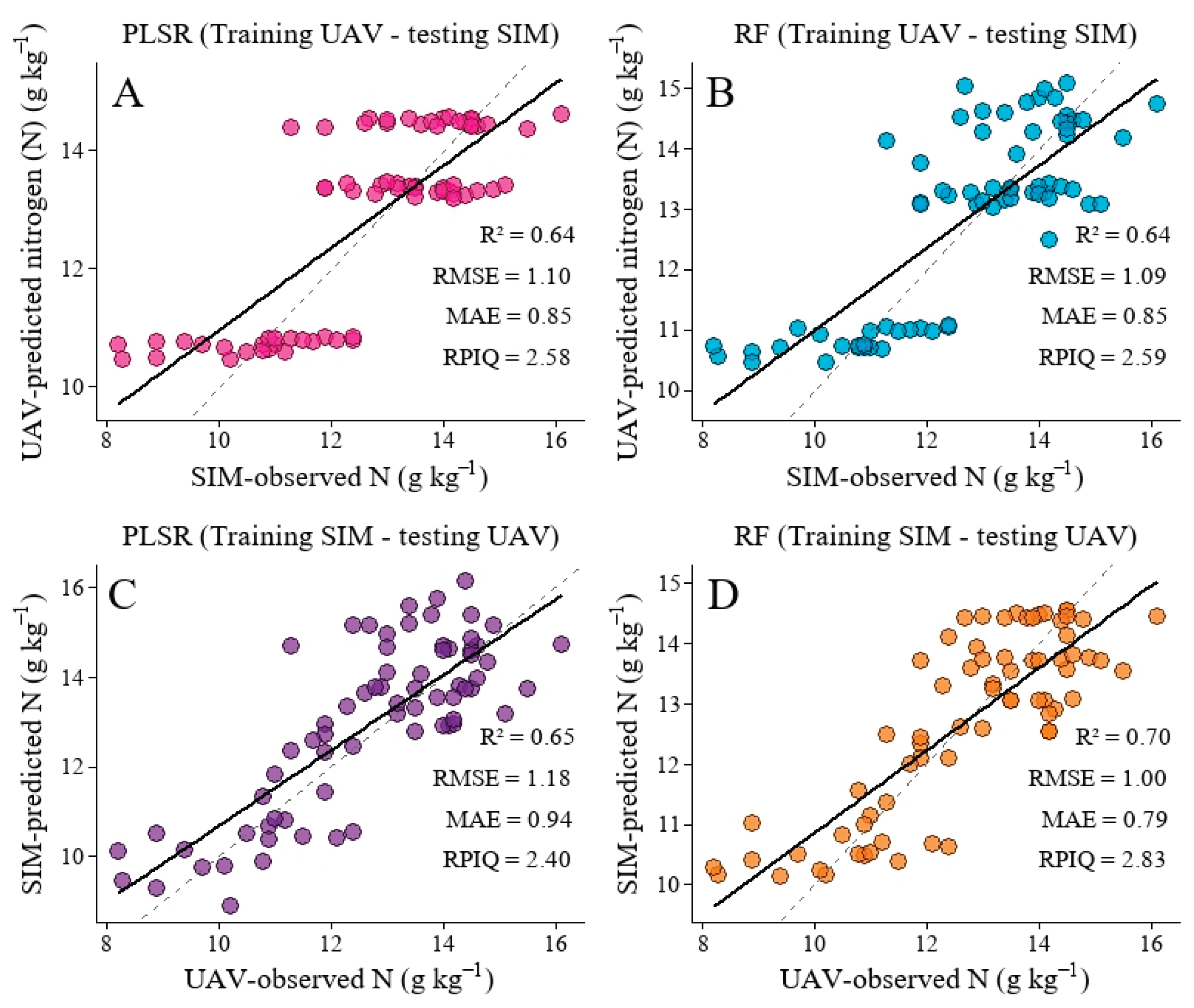
3.6. Independent Validation
3.7. Spatialization and Mapping of Leaf Nitrogen Content
4. Discussion
4.1. Analysis of the Correlations Between Original and Simulated Spectral Bands and Indices, and of the Linear Regression Applied to Spectral Indices for Predicting TFN
4.2. Model Performance in Nitrogen Prediction Using a UAV and Simulated Data (PLSR and Random Forest)
4.3. Validation
4.4. Spatial Distribution of Nitrogen Content in Sugarcane
5. Conclusions
Author Contributions
Funding
Data Availability Statement
Acknowledgments
Conflicts of Interest
References
- Barros, P.P.d.S.; Fiorio, P.R.; Demattê, J.A.d.M.; Martins, J.A.; Montezano, Z.F.; Dias, F.L.F. Estimation of leaf nitrogen levels in sugarcane using hyperspectral models. Cienc. Rural. 2022, 52. [Google Scholar] [CrossRef]
- Giordano, M.; Petropoulos, S.A.; Rouphael, Y. The Fate of Nitrogen from Soil to Plants: Influence of Agricultural Practices in Modern Agriculture. Agriculture 2021, 11, 944. [Google Scholar] [CrossRef]
- Li, X.; Ba, Y.; Zhang, M.; Nong, M.; Yang, C.; Zhang, S. Sugarcane Nitrogen Concentration and Irrigation Level Prediction Based on UAV Multispectral Imagery. Sensors 2022, 22, 2711. [Google Scholar] [CrossRef] [PubMed]
- Reyes-Trujillo, A.; Daza-Torres, M.C.; Galindez-Jamioy, C.A.; Rosero-García, E.E.; Muñoz-Arboleda, F.; Solarte-Rodriguez, E. Estimating canopy nitrogen concentration of sugarcane crop using in situ spectroscopy. Heliyon 2021, 7, e06566. [Google Scholar] [CrossRef]
- Barros, P.P.d.S.; Fiorio, P.R.; Demattê, J.A.d.M.; Martins, J.A.; Montezano, Z.F.; Dias, F.L.F. Estimation of leaf nitrogen levels in sugarcane using hyperspectral models. Cienc. Rural. 2022, 52. [Google Scholar] [CrossRef]
- Segl, K.; Richter, R.; Küster, T.; Kaufmann, H. End-to-end sensor simulation for spectral band selection and optimization with application to the Sentinel-2 mission. Appl. Opt. 2012, 51, 439–449. [Google Scholar] [CrossRef]
- Chen, X.; Miao, Y.; Kusnierek, K.; Li, F.; Wang, C.; Shi, B.; Wu, F.; Chang, Q.; Yu, K. Potential of Multi-Source Multispectral vs. Hyperspectral Remote Sensing for Winter Wheat Nitrogen Monitoring. Remote. Sens. 2025, 17, 2666. [Google Scholar] [CrossRef]
- Rehman, T.U.; Zhang, L.; Ma, D.; Wang, L.; Jin, J. Calibration transfer across multiple hyperspectral imaging-based plant phenotyping systems: I – Spectral space adjustment. Comput. Electron. Agric. 2020, 176, 105685–105685. [Google Scholar] [CrossRef]
- Matese, A.; Czarnecki, J.M.P.; Samiappan, S.; Moorhead, R. Are unmanned aerial vehicle-based hyperspectral imaging and machine learning advancing crop science? Trends Plant Sci. 2023, 29, 196–209. [Google Scholar] [CrossRef]
- Atherton, J.; Zhang, C.; Oivukkamäki, J.; Kulmala, L.; Xu, S.; Hakala, T.; Honkavaara, E.; MacArthur, A.; Porcar-Castell.
- Zhang, L.; Wang, A.; Zhang, H.; Zhu, Q.; Zhang, H.; Sun, W.; Niu, Y. Estimating Leaf Chlorophyll Content of Winter Wheat from UAV Multispectral Images Using Machine Learning Algorithms under Different Species, Growth Stages, and Nitrogen Stress Conditions. Agriculture 2024, 14, 1064. [Google Scholar] [CrossRef]
- Zou, M.; Liu, Y.; Fu, M.; Li, C.; Zhou, Z.; Meng, H.; Xing, E.; Ren, Y. Combining spectral and texture feature of UAV image with plant height to improve LAI estimation of winter wheat at jointing stage. Front. Plant Sci. 2024, 14, 1272049. [Google Scholar] [CrossRef] [PubMed]
- Yang, X.; Deng, X.; Li, Z.; Xu, J.; Zhou, Y. Monitoring sugarcane growth and nitrogen status using UAV-based multispectral imagery and machine learning. UAVs 2022, v. 6(n. 9), 230. [Google Scholar] [CrossRef]
- Li, X.; Ba, Y.; Zhang, M.; Nong, M.; Yang, C.; Zhang, S. Sugarcane Nitrogen Concentration and Irrigation Level Prediction Based on UAV Multispectral Imagery. Sensors 2022, 22, 2711. [Google Scholar] [CrossRef] [PubMed]
- Silva, C.A.A.C.; Rizzo, R.; Oliveira, A.K.d.S.; Castro, M.P.P.; Alexandre, M.L.d.S.; Lima, I.d.L.e.; Demattê, A.M.; Fiorio, P.R. Interspecies Prediction of Nitrogen Content in Processed Plant Samples Using Spectroscopic Modeling and Transfer Learning. Food Energy Secur. 2026, 15. [Google Scholar] [CrossRef]
- Martins, J.A.; Fiorio, P.R.; Silva, C.A.A.C.; Demattê, J.A.M.; Barros, P.P.d.S. Application of Vegetative Indices for Leaf Nitrogen Estimation in Sugarcane Using Hyperspectral Data. Sugar Tech 2023, 26, 160–170. [Google Scholar] [CrossRef]
- Fiorio, P.R.; Silva, C.A.A.C.; Rizzo, R.; Demattê, J.A.M.; Luciano, A.C.d.S.; da Silva, M.A. Prediction of leaf nitrogen in sugarcane (Saccharum spp.) by Vis-NIR-SWIR spectroradiometry. Heliyon 2024, 10, e26819. [Google Scholar] [CrossRef]
- Kumarasiri, D. S.; Wijetunga, C.; Gunathilaka, L. D. Use of UAV imagery to predict leaf nitrogen content of sugarcane cultivated under organic fertilizer application. Tropical Agricultural Research 2024, v. 35(n. 1), 67–79. Available online: https://tar.sljol.info/articles/8700.
- Alvares, C. A.; et al. Köppen’s climate classification map for Brazil. Meteorologische Zeitschrift 2013, v. 22(n. 6), 711–728. [Google Scholar] [CrossRef]
- Lee, M.A.; Huang, Y.; Yao, H.; Thomson, S.J.; Bruce, L.M. Determining the Effects of Storage on Cotton and Soybean Leaf Samples for Hyperspectral Analysis. IEEE J. Sel. Top. Appl. Earth Obs. Remote. Sens. 2014, 7, 2562–2570. [Google Scholar] [CrossRef]
- Tavares, T.R.; Fiorio, P.R.; Seixas, H.T.; Garcia, A.C.; Barros, P.P.d.S. Effects of storage on vis-NIR-SWIR reflectance spectra of Mombasa grass leaf samples. Cienc. Rural. 2020, 50. [Google Scholar] [CrossRef]
- ASD - Analytical Spectral Devices FieldSpec® 3 User Manual. 2010. Available online: Http://Www.Asdi.Com/.
- AGISOFT LLC. Agisoft Metashape Professional Edition, version 2.0.1; Agisoft LLC: St. Petersburg, Russia, 2022. [Google Scholar]
- Picado, E.F.; Romero, K.F.; Heenkenda, M.K. Mapping Spatial Variability of Sugarcane Foliar Nitrogen, Phosphorus, Potassium and Chlorophyll Concentrations Using Remote Sensing. Geomatics 2025, 5, 3. [Google Scholar] [CrossRef]
- QGIS DEVELOPMENT TEAM. QGIS Geographic Information System. Version 3.40. [S.l.]: QGIS Association, 2024. Disponível em: qgis.org.
- Li, W.; Zhu, X.; Yu, X.; Li, M.; Tang, X.; Zhang, J.; Xue, Y.; Zhang, C.; Jiang, Y. Inversion of Nitrogen Concentration in Apple Canopy Based on UAV Hyperspectral Images. Sensors 2022, 22, 3503. [Google Scholar] [CrossRef]
- Savitzky, A.; Golay, M.J.E. Smoothing and Differentiation of Data by Simplified Least Squares Procedures. Anal. Chem. 1964, 36, 1627–1639. [Google Scholar] [CrossRef]
- R Core Team. R: A Language and Environment for Statistical Computing_. R Foundation for Statistical Computing. 2024. Available online: https://www.r-project.org/.
- Burggraaff, O. Biases from incorrect reflectance convolution. Opt. Express 2020, 28, 13801–13816. [Google Scholar] [CrossRef]
- Wang, Y.; Kowalski, B.R. Calibration Transfer and Measurement Stability of Near-Infrared Spectrometers. Appl. Spectrosc. 1992, 46, 764–771. [Google Scholar] [CrossRef]
- Wang, Y.; Veltkamp, D.J.; Kowalski, B.R. Multivariate instrument standardization. Anal. Chem. 1991, 63, 2750–2756. [Google Scholar] [CrossRef]
- Mishra, P.; Nikzad-Langerodi, R.; Marini, F.; Roger, J.M.; Biancolillo, A.; Rutledge, D.N.; Lohumi, S. Are standard sample measurements still needed to transfer multivariate calibration models between near-infrared spectrometers? The answer is not always. TrAC Trends Anal. Chem. 2021, 143. [Google Scholar] [CrossRef]
- Ji, W.; Viscarra Rossel, R.A.; Shi, Z. Improved estimates of organic carbon using proximally sensed vis-NIR spectra corrected by piecewise direct standardization. Eur. J. Soil Sci. 2015, 66, 670–678. [Google Scholar] [CrossRef]
- Xue, J.; Su, B. Significant remote sensing vegetation indices: A review of developments and applications. J. Sens. 2017, 2017, 1353691. [Google Scholar] [CrossRef]
- Chen, X.; Miao, Y.; Kusnierek, K.; Li, F.; Wang, C.; Shi, B.; Wu, F.; Chang, Q.; Yu, K. Potential of Multi-Source Multispectral vs. Hyperspectral Remote Sensing for Winter Wheat Nitrogen Monitoring. Remote. Sens. 2025, 17, 2666. [Google Scholar] [CrossRef]
- Rouse, J.; Haas, R.; Schell, J.; Deering, D.; Harlan, J. Monitoring vegetation systems in the Great Plains with ERTS. In THIRD EARTH RESOURCES TECHNOLOGY SATELLITE-1 SYMPOSIUM; S.l.]: [s.n.], 1973; pp. 309–317. [Google Scholar]
- Gitelson, A.A.; Kaufman, Y.J.; Stark, R.; Rundquist, D. Novel algorithms for remote estimation of vegetation fraction. Remote Sens. Environ. 2002, 80, 76–87. [Google Scholar] [CrossRef]
- Gitelson, A.A.; Gritz, Y.; Merzlyak, M.N. Relationships between leaf chlorophyll content and spectral reflectance and algorithms for non-destructive chlorophyll assessment in higher plant leaves. J. Plant Physiol. 2003, 160, 271–282. [Google Scholar] [CrossRef]
- Maxmax. ENDVI. 2015. Available online: http://www.maxmax.com/endvi.htm.
- Schober, P.; Boer, C.; Schwarte, L.A. Correlation Coefficients: Appropriate Use and Interpretation. Anesth. Analg. 2018, 126, 1763–1768. [Google Scholar] [CrossRef]
- Wei, H.-E.; Grafton, M.; Bretherton, M.; Irwin, M.; Sandoval, E. Evaluation of Point Hyperspectral Reflectance and Multivariate Regression Models for Grapevine Water Status Estimation. Remote. Sens. 2021, 13, 3198. [Google Scholar] [CrossRef]
- Anku, K.E.; Percival, D.C.; Lada, R.; Heung, B.; Vankoughnett, M. Remote estimation of leaf nitrogen content, leaf area, and berry yield in wild blueberries. Front. Remote. Sens. 2024, 5, 1414540. [Google Scholar] [CrossRef]
- Yin, C.; Lv, X.; Zhang, L.; Ma, L.; Wang, H.; Zhang, L.; Zhang, Z. Hyperspectral UAV Images at Different Altitudes for Monitoring the Leaf Nitrogen Content in Cotton Crops. Remote. Sens. 2022, 14, 2576. [Google Scholar] [CrossRef]
- Chen, H.; Tan, C.; Lin, Z.; Wu, T. Classification and quantitation of milk powder by near-infrared spectroscopy and mutual information-based variable selection and partial least squares. Spectrochim. Acta Part A: Mol. Biomol. Spectrosc. 2018, 189, 183–189. [Google Scholar] [CrossRef]
- Breiman, L. Random forests. Mach. Learn. 2001, 45, 5–32. [Google Scholar] [CrossRef]
- Liu, Y.; Wang, Y.; Zhang, J. New Machine Learning Algorithm: Random Forest. In Information Computing and Applications; Springer: Berlin/Heidelberg, Germany, 2012; pp. 246–252. [Google Scholar]
- Tang, K.; Qin, M.; Han, B.; Shao, D.; Xu, Z.; Sun, H.; Wu, Y. Identifying the influencing factors of soil nitrous acid emissions using random forest model. Atmospheric Environ. 2024, 339. [Google Scholar] [CrossRef]
- Liu, M.; Liu, X.; Liu, D.; Ding, C.; Jiang, J. Multivariable integration method for estimating sea surface salinity in coastal waters from in situ data and remotely sensed data using random forest algorithm. Comput. Geosci. 2015, 75, 44–56. [Google Scholar] [CrossRef]
- Friedman, J.H.; Meulman, J.J. Multiple additive regression trees with application in epidemiology. Stat. Med. 2003, 22, 1365–1381. [Google Scholar] [CrossRef]
- Rodrigues, M.; Nanni, M.R.; Cezar, E.; dos Santos, G.L.A.A.; Reis, A.S.; de Oliveira, K.M.; de Oliveira, R.B. Vis–NIR spectroscopy: from leaf dry mass production estimate to the prediction of macro- and micronutrients in soybean crops. J. Appl. Remote. Sens. 2020, 14, 044505. [Google Scholar] [CrossRef]
- Leite, J.M.; Ciampitti, I.A.; Mariano, E.; Vieira-Megda, M.X.; Trivelin, P.C.O. Nutrient Partitioning and Stoichiometry in Unburnt Sugarcane Ratoon at Varying Yield Levels. Front. Plant Sci. 2016, 7, 466. [Google Scholar] [CrossRef]
- Hassan, M.A.; Yang, M.; Rasheed, A.; Yang, G.; Reynolds, M.; Xia, X.; Xiao, Y.; He, Z. A rapid monitoring of NDVI across the wheat growth cycle for grain yield prediction using a multi-spectral UAV platform. 4th IPPN International Plant Phenotyping Symposium; LOCATION OF CONFERENCE, COUNTRYDATE OF CONFERENCE; pp. 95–103.
- Zhang, C.; Yang, G.; Li, H.; Tang, F.; Liu, C.; Zhang, Y. Remote sensing inversion of leaf area index of winter wheat based on random forest algorithm. Scientia Agricultura Sin. 2018, 51(5), 855–867. [Google Scholar] [CrossRef]
- Osco, L.P.; Ramos, A.P.M.; Pereira, D.R.; Moriya, É.A.S.; Imai, N.N.; Matsubara, E.T.; Estrabis, N.; de Souza, M.; Junior, J.M.; Gonçalves, W.N.; et al. Predicting Canopy Nitrogen Content in Citrus-Trees Using Random Forest Algorithm Associated to Spectral Vegetation Indices from UAV-Imagery. Remote. Sens. 2019, 11, 2925. [Google Scholar] [CrossRef]
- Imran, H.A.; Gianelle, D.; Rocchini, D.; Dalponte, M.; Martín, M.P.; Sakowska, K.; Wohlfahrt, G.; Vescovo, L. VIS-NIR, Red-Edge and NIR-Shoulder Based Normalized Vegetation Indices Response to Co-Varying Leaf and Canopy Structural Traits in Heterogeneous Grasslands. Remote. Sens. 2020, 12, 2254. [Google Scholar] [CrossRef]
- Narmilan, A.; Gonzalez, F.; Salgadoe, A.S.A.; Kumarasiri, U.W.L.M.; Weerasinghe, H.A.S.; Kulasekara, B.R. Predicting Canopy Chlorophyll Content in Sugarcane Crops Using Machine Learning Algorithms and Spectral Vegetation Indices Derived from UAV Multispectral Imagery. Remote. Sens. 2022, 14, 1140. [Google Scholar] [CrossRef]
- Gutman, G.; Skakun, S.; Gitelson, A. Revisiting the use of red and near-infrared reflectances in vegetation studies and numerical climate models. Sci. Remote. Sens. 2021, 4. [Google Scholar] [CrossRef]
- Miphokasap, P.; Honda, K.; Vaiphasa, C.; Souris, M.; Nagai, M. Estimating Canopy Nitrogen Concentration in Sugarcane Using Field Imaging Spectroscopy. Remote. Sens. 2012, 4, 1651–1670. [Google Scholar] [CrossRef]
- Li, X.; Ba, Y.; Zhang, M.; Nong, M.; Yang, C.; Zhang, S. Sugarcane Nitrogen Concentration and Irrigation Level Prediction Based on UAV Multispectral Imagery. Sensors 2022, 22, 2711. [Google Scholar] [CrossRef]
- Martins, J.A.; Fiorio, P.R.; Silva, C.A.A.C.; Demattê, J.A.M.; Barros, P.P.d.S. Application of Vegetative Indices for Leaf Nitrogen Estimation in Sugarcane Using Hyperspectral Data. Sugar Tech 2023, 26, 160–170. [Google Scholar] [CrossRef]
- Hassani, K.; Gholizadeh, H.; Taghvaeian, S.; Natalie, V.; Carpenter, J.; Jacob, J. Assessing the impact of spatial resolution of UAS-based remote sensing and spectral resolution of proximal sensing on crop nitrogen retrieval accuracy. Int. J. Remote. Sens. 2023, 44, 4441–4464. [Google Scholar] [CrossRef]
- Lv, Z.; Zhao, W.; Kong, S.; Li, L.; Lin, S. Overview of molecular mechanisms of plant leaf development: a systematic review. Front. Plant Sci. 2023, 14, 1293424. [Google Scholar] [CrossRef]
- Yoneyama, T.; Suzuki, A. Light-Independent Nitrogen Assimilation in Plant Leaves: Nitrate Incorporation into Glutamine, Glutamate, Aspartate, and Asparagine Traced by 15N. Plants 2020, 9, 1303. [Google Scholar] [CrossRef]
- Miphokasap, P.; Wannasiri, W. Estimations of Nitrogen Concentration in Sugarcane Using Hyperspectral Imagery. Sustainability 2018, 10, 1266. [Google Scholar] [CrossRef]
- Biriukova, K.; Celesti, M.; Evdokimov, A.; Pacheco-Labrador, J.; Julitta, T.; Migliavacca, M.; Giardino, C.; Miglietta, F.; Colombo, R.; Panigada, C.; et al. Effects of varying solar-view geometry and canopy structure on solar-induced chlorophyll fluorescence and PRI. Int. J. Appl. Earth Obs. Geoinformation 2020, 89. [Google Scholar] [CrossRef]
- Lu, N.; Wang, W.; Zhang, Q.; Li, D.; Yao, X.; Tian, Y.; Zhu, Y.; Cao, W.; Baret, F.; Liu, S.; et al. Estimation of Nitrogen Nutrition Status in Winter Wheat From Unmanned Aerial Vehicle Based Multi-Angular Multispectral Imagery. Front. Plant Sci. 2019, 10, 1601. [Google Scholar] [CrossRef] [PubMed]
- Azizabadi, E.C.; El-Shetehy, M.; Cheng, X.; Youssef, A.; Badreldin, N. In-Season Potato Nitrogen Prediction Using Multispectral Drone Data and Machine Learning. Remote. Sens. 2025, 17, 1860. [Google Scholar] [CrossRef]
- Soltanikazemi, M.; Minaei, S.; Shafizadeh-Moghadam, H.; Mahdavian, A. Field-scale estimation of sugarcane leaf nitrogen content using vegetation indices and spectral bands of Sentinel-2: Application of random forest and support vector regression. Comput. Electron. Agric. 2022, 200. [Google Scholar] [CrossRef]
- Huang, Y.; Chen, W.; Tan, W.; Deng, Y.; Yang, C.; Zhu, X.; Shen, J.; Liu, N. Transfer learning for enhancing the generality of leaf spectroscopic models in estimating crop foliar nutrients across growth stages. Int. J. Appl. Earth Obs. Geoinformation 2025, 139. [Google Scholar] [CrossRef]
- Ji, F.; Li, F.; Hao, D.; Shiklomanov, A.N.; Yang, X.; Townsend, P.A.; Dashti, H.; Nakaji, T.; Kovach, K.R.; Liu, H.; et al. Unveiling the transferability of PLSR models for leaf trait estimation: lessons from a comprehensive analysis with a novel global dataset. New Phytol. 2024, 243, 111–131. [Google Scholar] [CrossRef] [PubMed]
- Helsen, K.; Bassi, L.; Feilhauer, H.; Kattenborn, T.; Matsushima, H.; Van Cleemput, E.; Somers, B.; Honnay, O. Evaluating different methods for retrieving intraspecific leaf trait variation from hyperspectral leaf reflectance. Ecol. Indic. 2021, 130. [Google Scholar] [CrossRef]
- Schiefer, F.; Schmidtlein, S.; Kattenborn, T. The retrieval of plant functional traits from canopy spectra through RTM-inversions and statistical models are both critically affected by plant phenology. Ecol. Indic. 2021, 121. [Google Scholar] [CrossRef]
- Chlus, A.; Townsend, P.A. Characterizing seasonal variation in foliar biochemistry with airborne imaging spectroscopy. Remote. Sens. Environ. 2022, 275. [Google Scholar] [CrossRef]
- Pan, Y.; Li, J.; Zhang, J.; He, J.; Zhang, Z.; Yao, X.; Cheng, T.; Zhu, Y.; Cao, W.; Tian, Y. Estimating Leaf Nitrogen Accumulation Considering Vertical Heterogeneity Using Multiangular Unmanned Aerial Vehicle Remote Sensing in Wheat. Plant Phenomics 2024, 6, 0276. [Google Scholar] [CrossRef]
- Silva, C.A.A.C.; Rizzo, R.; Oliveira, A.K.d.S.; Castro, M.P.P.; Alexandre, M.L.d.S.; Lima, I.d.L.e.; Demattê, A.M.; Fiorio, P.R. Interspecies Prediction of Nitrogen Content in Processed Plant Samples Using Spectroscopic Modeling and Transfer Learning. Food Energy Secur. 2026, 15. [Google Scholar] [CrossRef]
- Alharbi, K.; Haroun, S.A.; Kazamel, A.M.; Abbas, M.A.; Ahmaida, S.M.; AlKahtani, M.; AlHusnain, L.; Attia, K.A.; Abdelaal, K.; Gamel, R.M.E. Physiological Studies and Ultrastructure of Vigna sinensis L. and Helianthus annuus L. under Varying Levels of Nitrogen Supply. Plants 2022, 11, 1884. [Google Scholar] [CrossRef]
- Mu, X.; Chen, Y. The physiological response of photosynthesis to nitrogen deficiency. Plant Physiol. Biochem. 2021, 158, 76–82. [Google Scholar] [CrossRef]
- Alexandre, M.L.d.S.; Lima, I.d.L.e.; Nilsson, M.S.; Rizzo, R.; Silva, C.A.A.C.; Fiorio, P.R. Sugarcane (Saccharum officinarum) Productivity Estimation Using Multispectral Sensors in RPAs, Biometric Variables, and Vegetation Indices. Agronomy 2025, 15, 2149. [Google Scholar] [CrossRef]
- Yu, K.-Q.; Zhao, Y.-R.; Li, X.-L.; Shao, Y.-N.; Liu, F.; He, Y. Hyperspectral Imaging for Mapping of Total Nitrogen Spatial Distribution in Pepper Plant. PLOS ONE 2014, 9, e116205–e116205. [Google Scholar] [CrossRef]
- Pechanec, V.; Mráz, A.; Rozkošný, L.; Vyvlečka, P. Usage of Airborne Hyperspectral Imaging Data for Identifying Spatial Variability of Soil Nitrogen Content. ISPRS Int. J. Geo-Information 2021, 10, 355. [Google Scholar] [CrossRef]
- Verrelst, J.; Rivera-Caicedo, J.P.; Reyes-Muñoz, P.; Morata, M.; Amin, E.; Tagliabue, G.; Panigada, C.; Hank, T.; Berger, K. Mapping landscape canopy nitrogen content from space using PRISMA data. ISPRS J. Photogramm. Remote. Sens. 2021, 178, 382–395. [Google Scholar] [CrossRef]
- Bassi, D.; Menossi, M.; Mattiello, L. Nitrogen supply influences photosynthesis establishment along the sugarcane leaf. Sci. Rep. 2018, 8, 2327. [Google Scholar] [CrossRef] [PubMed]
- Do Amaral, L. R.; Molin, J. P. Sensor óptico no auxílio à recomendação de adubação nitrogenada em cana-de-açúcar. Pesquisa Agropecuária Brasileira 2011, v. 46(n. 12), 1633–1642. [Google Scholar] [CrossRef]
- Santana, A.C.d.A.; de Oliveira, E.C.A.; da Silva, V.S.G.; dos Santos, R.L.; da Silva, M.A.; Freire, F.J. Critical nitrogen dilution curves and productivity assessments for plant cane. Rev. Bras. De Eng. Agricola E Ambient. 2020, 24, 244–251. [Google Scholar] [CrossRef]
- Ollinger, S.V. Sources of variability in canopy reflectance and the convergent properties of plants. New Phytol. 2010, 189, 375–394. [Google Scholar] [CrossRef]


| Band (λ) | Central Wavelength (nm) | Bandwidth (nm) | |
|---|---|---|---|
| Blue (B) | 450 | ±16 | |
| Green (G) | 560 | ±16 | |
| Red (R) | 650 | ±16 | |
| Red-edge (RE) | 730 | ±16 | |
| Near-infrared (NIR) | 840 | ±26 | |
| RGB | 20 megapixels |
| ID | Vegetation Index | Formula | References |
|---|---|---|---|
| 1 | Normalized Difference Vegetation Index (NDVI) | (NIR - R) / (NIR + R) | [36] |
| 2 | Visible Atmospheric Resistance Index (VARI) | (G - R) / (G + R - B) | [37] |
| 3 | Chlorophyll Index - Red-Edge (ChlRe) | (NIR) / (RED)-1 | [38] |
| 4 | Improved Normalized Difference Vegetation Index (ENDVI) | (NIR - G) - (2 x B) / (NIR - G) + (2 x B) | [39] |
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2026 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).




