Submitted:
30 March 2026
Posted:
30 March 2026
You are already at the latest version
Abstract
Keywords:
1. Introduction
2. Materials and Methods
2.1. Sample Collection
2.2. Nucleic Acids Extraction
2.3. RT-qPCR for HEV detection
2.4. Whole-Genome Amplification by RT-PCR
2.5. Sanger Sequencing
2.6. Phylogenetic and Phylogeographic Analysis
2.7. Recombination Analysis
3. Results
3.1. HEV Detection
3.2. Phylogenetic Analysis
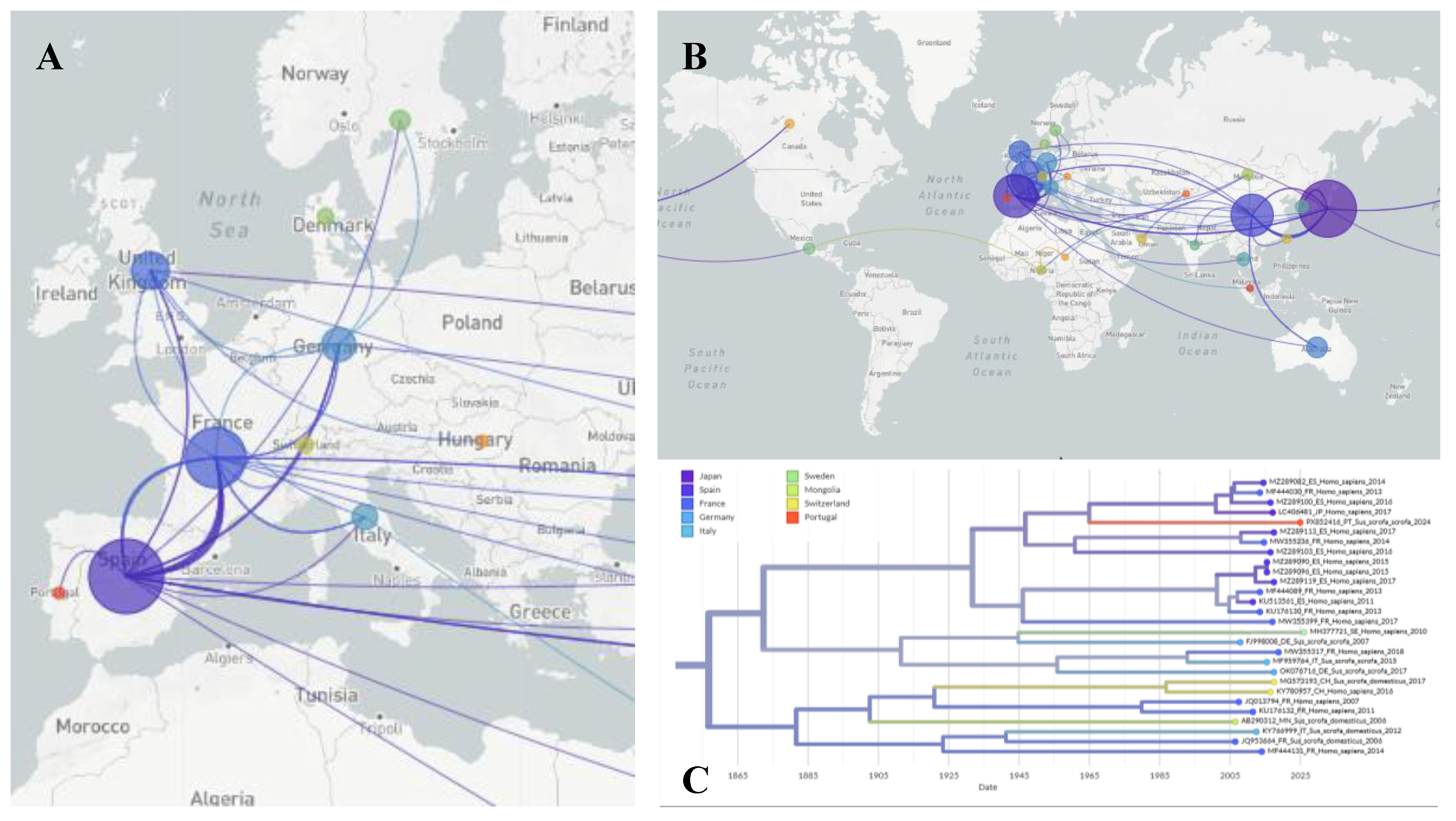
3.3. Phylogeographic Analysis
3.4. Recombination Analysis
4. Discussion and Conclusions
Supplementary Materials
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Acknowledgments
Conflicts of Interest
Abbreviations
| The following abbreviations are used in this manuscript | . |
References
- Massei, G.; Kindberg, J.; Licoppe, A.; Gačić, D.; Šprem, N.; Kamler, J.; Baubet, E.; Hohmann, U.; Monaco, A.; Ozoliņš, J.; et al. Wild Boar Populations up, Numbers of Hunters down? A Review of Trends and Implications for Europe. Pest Management Science 2015, 71, 492–500. [Google Scholar] [CrossRef]
- Raji, Y.E.; Toung, O.P.; Mohd Taib, N.; Sekawi, Z.B. A Systematic Review of the Epidemiology of Hepatitis E Virus Infection in South – Eastern Asia. Virulence 2020, 12, 114–129. [Google Scholar] [CrossRef] [PubMed]
- Sooryanarain, H.; Meng, X.-J. Hepatitis E Virus: Reasons for Emergence in Humans. Current Opinion in Virology 2019, 34, 10–17. [Google Scholar] [CrossRef]
- Pavio, N.; Doceul, V.; Bagdassarian, E.; Johne, R. Recent Knowledge on Hepatitis E Virus in Suidae Reservoirs and Transmission Routes to Human. Veterinary Research 2017, 48, 78. [Google Scholar] [CrossRef] [PubMed]
- Tene, S.D.; Diouara, A.A.M.; Sané, S.; Coundoul, S. Hepatitis E Virus (HEV) Infection in the Context of the One Health Approach: A Systematic Review. Pathogens 2025, 14, 704. [Google Scholar] [CrossRef]
- Wang, L.; Teng, J.L.L.; Lau, S.K.P.; Sridhar, S.; Fu, H.; Gong, W.; Li, M.; Xu, Q.; He, Y.; Zhuang, H.; et al. Transmission of a Novel Genotype of Hepatitis E Virus from Bactrian Camels to Cynomolgus Macaques. J Virol 2019, 93, e02014-18. [Google Scholar] [CrossRef]
- Schlosser, J.; Eiden, M.; Vina-Rodriguez, A.; Fast, C.; Dremsek, P.; Lange, E.; Ulrich, R.G.; Groschup, M.H. Natural and Experimental Hepatitis E Virus Genotype 3 - Infection in European Wild Boar Is Transmissible to Domestic Pigs. Veterinary Research 2014, 45, 121. [Google Scholar] [CrossRef]
- Spancerniene, U.; Grigas, J.; Buitkuviene, J.; Zymantiene, J.; Juozaitiene, V.; Stankeviciute, M.; Razukevicius, D.; Zienius, D.; Stankevicius, A. Prevalence and Phylogenetic Analysis of Hepatitis E Virus in Pigs, Wild Boars, Roe Deer, Red Deer and Moose in Lithuania. Acta Veterinaria Scandinavica 2018, 60, 13. [Google Scholar] [CrossRef] [PubMed]
- Nicot, F.; Dimeglio, C.; Migueres, M.; Jeanne, N.; Latour, J.; Abravanel, F.; Ranger, N.; Harter, A.; Dubois, M.; Lameiras, S.; et al. Classification of the Zoonotic Hepatitis E Virus Genotype 3 Into Distinct Subgenotypes. Front Microbiol 2021, 11, 634430. [Google Scholar] [CrossRef] [PubMed]
- Pavio, N.; Mansuy, J.-M. Hepatitis E in High-Income Countries. Curr Opin Infect Dis 2010, 23, 521–527. [Google Scholar] [CrossRef]
- Abrantes, A.C.; Santos-Silva, S.; Mesquita, J.; Vieira-Pinto, M. Hepatitis E Virus in the Wild Boar Population: What Is the Real Zoonotic Risk in Portugal? Tropical Medicine and Infectious Disease 2023, 8, 433. [Google Scholar] [CrossRef]
- Izopet, J.; Tremeaux, P.; Marion, O.; Migueres, M.; Capelli, N.; Chapuy-Regaud, S.; Mansuy, J.-M.; Abravanel, F.; Kamar, N.; Lhomme, S. Hepatitis E Virus Infections in Europe. J Clin Virol 2019, 120, 20–26. [Google Scholar] [CrossRef] [PubMed]
- Krain, L.J.; Nelson, K.E.; Labrique, A.B. Host Immune Status and Response to Hepatitis E Virus Infection. Clinical Microbiology Reviews 2014, 27, 139–165. [Google Scholar] [CrossRef]
- Aggarwal, A.; Perumpail, R.B.; Tummala, S.; Ahmed, A. Hepatitis E Virus Infection in the Liver Transplant Recipients: Clinical Presentation and Management. World J Hepatol 2016, 8, 117–122. [Google Scholar] [CrossRef]
- Damiris, K.; Meybodi, M.A.; Niazi, M.; Pyrsopoulos, N. Hepatitis E in Immunocompromised Individuals. World Journal of Hepatology 2022, 14, 482–494. [Google Scholar] [CrossRef] [PubMed]
- Hoofnagle, J.H.; Nelson, K.E.; Purcell, R.H. Hepatitis E. N Engl J Med 2012, 367, 1237–1244. [Google Scholar] [CrossRef]
- Barroso, P.; Risalde, M.A.; García-Bocanegra, I.; Acevedo, P.; Barasona, J.Á.; Caballero-Gómez, J.; Jiménez-Ruiz, S.; Rivero-Juárez, A.; Montoro, V.; Vicente, J. Long-Term Determinants of the Seroprevalence of the Hepatitis E Virus in Wild Boar (Sus Scrofa). Animals 2021, 11, 1805. [Google Scholar] [CrossRef]
- Rivero-Juarez, A.; Risalde, M.A.; Frias, M.; García-Bocanegra, I.; Lopez-Lopez, P.; Cano-Terriza, D.; Camacho, A.; Jimenez-Ruiz, S.; Gomez-Villamandos, J.C.; Rivero, A. Prevalence of Hepatitis E Virus Infection in Wild Boars from Spain: A Possible Seasonal Pattern? BMC Vet Res 2018, 14, 54. [Google Scholar] [CrossRef] [PubMed]
- Zhao, C.; Li, Z.; Yan, B.; Harrison, T.J.; Guo, X.; Zhang, F.; Yin, J.; Yan, Y.; Wang, Y. Comparison of Real-Time Fluorescent RT-PCR and Conventional RT-PCR for the Detection of Hepatitis E Virus Genotypes Prevalent in China. Journal of Medical Virology 2007, 79, 1966–1973. [Google Scholar] [CrossRef]
- Larsson, A. AliView: A Fast and Lightweight Alignment Viewer and Editor for Large Datasets. Bioinformatics 2014, 30, 3276–3278. [Google Scholar] [CrossRef]
- Castresana, J. Selection of Conserved Blocks from Multiple Alignments for Their Use in Phylogenetic Analysis. Mol Biol Evol 2000, 17, 540–552. [Google Scholar] [CrossRef]
- Talavera, G.; Castresana, J. Improvement of Phylogenies after Removing Divergent and Ambiguously Aligned Blocks from Protein Sequence Alignments. Syst Biol 2007, 56, 564–577. [Google Scholar] [CrossRef] [PubMed]
- Schmidt, H.A.; Strimmer, K.; Vingron, M.; von Haeseler, A. TREE-PUZZLE: Maximum Likelihood Phylogenetic Analysis Using Quartets and Parallel Computing. Bioinformatics 2002, 18, 502–504. [Google Scholar] [CrossRef]
- Strimmer, K.; von Haeseler, A. Likelihood-Mapping: A Simple Method to Visualize Phylogenetic Content of a Sequence Alignment. Proc Natl Acad Sci U S A 1997, 94, 6815–6819. [Google Scholar] [CrossRef]
- Hoang, D.T.; Chernomor, O.; von Haeseler, A.; Minh, B.Q.; Vinh, L.S. UFBoot2: Improving the Ultrafast Bootstrap Approximation. Mol Biol Evol 2018, 35, 518–522. [Google Scholar] [CrossRef]
- Kalyaanamoorthy, S.; Minh, B.Q.; Wong, T.K.F.; von Haeseler, A.; Jermiin, L.S. ModelFinder: Fast Model Selection for Accurate Phylogenetic Estimates. Nature Methods 2017, 14, 587–589. [Google Scholar] [CrossRef] [PubMed]
- Wong, T.K.F.; Ly-Trong, N.; Ren, H.; Baños, H.; Roger, A.J.; Susko, E.; Bielow, C.; Maio, N.D.; Goldman, N.; Hahn, M.W.; et al. IQ-TREE 3: Phylogenomic Inference Software Using Complex Evolutionary Models. 2025.
- Pas, S.D.; de Man, R.A.; Mulders, C.; Balk, A.H.M.M.; van Hal, P.T.W.; Weimar, W.; Koopmans, M.P.G.; Osterhaus, A.D.M.E.; van der Eijk, A.A. Hepatitis E Virus Infection among Solid Organ Transplant Recipients, the Netherlands. Emerg Infect Dis 2012, 18, 869–872. [Google Scholar] [CrossRef] [PubMed]
- Rambaut, A.; Lam, T.T.; Max Carvalho, L.; Pybus, O.G. Exploring the Temporal Structure of Heterochronous Sequences Using TempEst (Formerly Path-O-Gen). Virus Evol 2016, 2, vew007. [Google Scholar] [CrossRef]
- Borges, V.; Pinheiro, M.; Pechirra, P.; Guiomar, R.; Gomes, J.P. INSaFLU: An Automated Open Web-Based Bioinformatics Suite “from-Reads” for Influenza Whole-Genome-Sequencing-Based Surveillance. Genome Medicine 2018, 10, 46. [Google Scholar] [CrossRef]
- Santos, J.D.; Sobral, D.; Pinheiro, M.; Isidro, J.; Bogaardt, C.; Pinto, M.; Eusébio, R.; Santos, A.; Mamede, R.; Horton, D.L.; et al. INSaFLU-TELEVIR: An Open Web-Based Bioinformatics Suite for Viral Metagenomic Detection and Routine Genomic Surveillance. Genome Medicine 2024, 16, 61. [Google Scholar] [CrossRef]
- Hadfield, J.; Megill, C.; Bell, S.M.; Huddleston, J.; Potter, B.; Callender, C.; Sagulenko, P.; Bedford, T.; Neher, R.A. Nextstrain: Real-Time Tracking of Pathogen Evolution. Bioinformatics 2018, 34, 4121–4123. [Google Scholar] [CrossRef] [PubMed]
- Martin, D.P.; Murrell, B.; Golden, M.; Khoosal, A.; Muhire, B. RDP4: Detection and Analysis of Recombination Patterns in Virus Genomes. Virus Evol 2015, 1, vev003. [Google Scholar] [CrossRef]
- Martin, D.; Rybicki, E. RDP: Detection of Recombination amongst Aligned Sequences. Bioinformatics 2000, 16, 562–563. [Google Scholar] [CrossRef] [PubMed]
- Padidam, M.; Sawyer, S.; Fauquet, C.M. Possible Emergence of New Geminiviruses by Frequent Recombination. Virology 1999, 265, 218–225. [Google Scholar] [CrossRef]
- Smith, J.M. Analyzing the Mosaic Structure of Genes. J Mol Evol 1992, 34, 126–129. [Google Scholar] [CrossRef]
- Posada, D.; Crandall, K.A. Evaluation of Methods for Detecting Recombination from DNA Sequences: Computer Simulations. Proc. Natl. Acad. Sci. U.S.A 2001, 98, 13757–13762. [Google Scholar] [CrossRef] [PubMed]
- Gibbs, M.J.; Armstrong, J.S.; Gibbs, A.J. Sister-Scanning: A Monte Carlo Procedure for Assessing Signals in Recombinant Sequences. Bioinformatics 2000, 16, 573–582. [Google Scholar] [CrossRef]
- Lam, H.M.; Ratmann, O.; Boni, M.F. Improved Algorithmic Complexity for the 3SEQ Recombination Detection Algorithm. Mol Biol Evol 2018, 35, 247–251. [Google Scholar] [CrossRef]
- Weiller, G.F. Phylogenetic Profiles: A Graphical Method for Detecting Genetic Recombinations in Homologous Sequences. Mol Biol Evol 1998, 15, 326–335. [Google Scholar] [CrossRef]
- Lemey, P.; Lott, M.; Martin, D.P.; Moulton, V. Identifying Recombinants in Human and Primate Immunodeficiency Virus Sequence Alignments Using Quartet Scanning. BMC Bioinformatics 2009, 10, 126. [Google Scholar] [CrossRef]
- McGuire, G.; Wright, F. TOPAL 2.0: Improved Detection of Mosaic Sequences within Multiple Alignments. Bioinformatics 2000, 16, 130–134. [Google Scholar] [CrossRef] [PubMed]
- Holmes, E.C.; Worobey, M.; Rambaut, A. Phylogenetic Evidence for Recombination in Dengue Virus. Mol Biol Evol 1999, 16, 405–409. [Google Scholar] [CrossRef] [PubMed]
- Pires, H.; Cardoso, L.; Lopes, A.P.; Fontes, M. da C.; Santos-Silva, S.; Matos, M.; Pintado, C.; Figueira, L.; Matos, A.C.; Mesquita, J.R.; et al. Prevalence and Risk Factors for Hepatitis E Virus in Wild Boar and Red Deer in Portugal. Microorganisms 2023, 11, 2576. [Google Scholar] [CrossRef] [PubMed]
- Santos-Silva, S.; Moraes, D.F. da S.D.; López-López, P.; Palmeira, J.D.; Torres, R.T.; São José Nascimento, M.; Dashti, A.; Carmena, D.; Rivero-Juarez, A.; Mesquita, J.R. Survey of Zoonotic Diarrheagenic Protist and Hepatitis E Virus in Wild Boar (Sus Scrofa) of Portugal. Animals 2023, 13, 256. [Google Scholar] [CrossRef]
- Reuter, G.; Fodor, D.; Forgách, P.; Kátai, A.; Szűcs, G. Characterization and Zoonotic Potential of Endemic Hepatitis E Virus (HEV) Strains in Humans and Animals in Hungary. Journal of Clinical Virology 2009, 44, 277–281. [Google Scholar] [CrossRef]
- Roth, A.; Lin, J.; Magnius, L.; Karlsson, M.; Belák, S.; Widén, F.; Norder, H. Markers for Ongoing or Previous Hepatitis E Virus Infection Are as Common in Wild Ungulates as in Humans in Sweden. Viruses 2016, 8, 259. [Google Scholar] [CrossRef]
- Rutjes, S.A.; Lodder-Verschoor, F.; Lodder, W.J.; van der Giessen, J.; Reesink, H.; Bouwknegt, M.; de Roda Husman, A.M. Seroprevalence and Molecular Detection of Hepatitis E Virus in Wild Boar and Red Deer in The Netherlands. Journal of Virological Methods 2010, 168, 197–206. [Google Scholar] [CrossRef]
- Schielke, A.; Ibrahim, V.; Czogiel, I.; Faber, M.; Schrader, C.; Dremsek, P.; Ulrich, R.G.; Johne, R. Hepatitis E Virus Antibody Prevalence in Hunters from a District in Central Germany, 2013: A Cross-Sectional Study Providing Evidence for the Benefit of Protective Gloves during Disembowelling of Wild Boars. BMC Infectious Diseases 2015, 15, 440. [Google Scholar] [CrossRef]
- Fajardo-Alonso, T.; García-Bocanegra, I.; Risalde, M.A.; Rivero-Juárez, A.; Jiménez-Ruiz, S.; Cano-Terriza, D.; Casares-Jiménez, M.; Laguna, E.; Acevedo, P.; Frías, M.; et al. Longitudinal Survey of Hepatitis E Virus in Extensively Raised Pigs in Spain. Veterinary Microbiology 2024, 298, 110256. [Google Scholar] [CrossRef]
- Wang, H.; Castillo-Contreras, R.; Saguti, F.; López-Olvera, J.R.; Karlsson, M.; Mentaberre, G.; Lindh, M.; Serra-Cobo, J.; Norder, H. Genetically Similar Hepatitis E Virus Strains Infect Both Humans and Wild Boars in the Barcelona Area, Spain, and Sweden. Transboundary and Emerging Diseases 2018, 66, 978–985. [Google Scholar] [CrossRef]
- Ruiz-Ponsell, L.; Monastiri, A.; López-Roig, M.; Sauleda, S.; Bes, M.; Mentaberre, G.; Escobar-González, M.; Costafreda, M.I.; López-Olvera, J.R.; Serra-Cobo, J. Endemic Maintenance of Human-Related Hepatitis E Virus Strains in Synurbic Wild Boars, Barcelona Metropolitan Area, Spain. Science of The Total Environment 2024, 955, 176871. [Google Scholar] [CrossRef] [PubMed]
- Mesquita, J.R.; Oliveira, R.M.S.; Coelho, C.; Vieira-Pinto, M.; Nascimento, M.S.J. Hepatitis E Virus in Sylvatic and Captive Wild Boar from Portugal. Transboundary and Emerging Diseases 2014, 63, 574–578. [Google Scholar] [CrossRef] [PubMed]
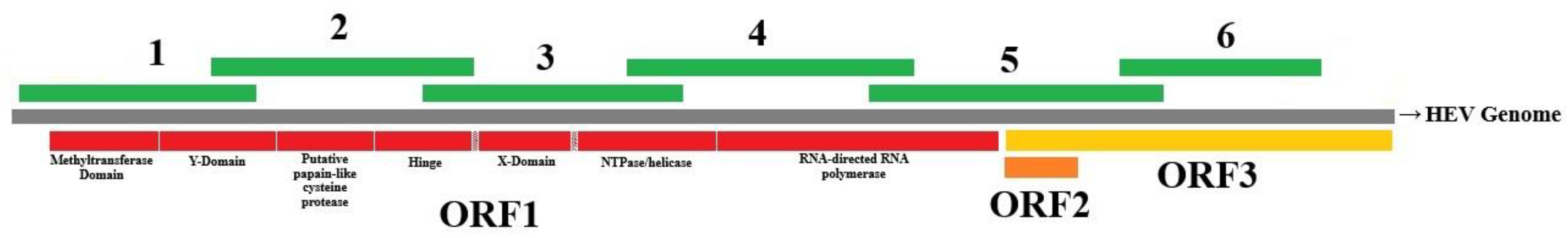
| Reaction(s) | Primer name | Sequence | Target Gene | Amplicon size |
| HEV RT-qPCR | HEV-AB-F | 5’ - CGGTGGTTTCTGGGGTGA - 3’ | Capsid / ORF3 | 75 bp |
| HEV-AB-R | 5’ - GCRAAGGGRTTGGTTGG - 3’ | |||
| HEV probe | 5’ - [FAM]TGATTCTCAGCCCTTCGC - 3’ | |||
| HEV RT-PCR 1 | Primer_HEV_Fw1 | 5’ - CTCCTGGCATTACTACTGCCA - 3’ | ORF1 | 1188 bp |
| Primer_HEV_Rv1 | 5’ - TTYTGGGCRTGCTCAACCTC - 3’ | |||
| HEV RT-PCR 2 | Primer_HEV_Fw2 | 5’ - CGGCTYATGACYTAYCTCCG - 3’ | ORF1 | 1358 bp |
| Primer_HEV_Rv2 | 5’ - RAGRAGVCGGCGRGTGCG - 3’ | |||
| HEV RT-PCR 3 | Primer_HEV_Fw3 | 5’ - GTCHACATCTGGYTTYTCTAG - 3’ | ORF1 | 1360 bp |
| Primer_HEV_Rv3 | 5’ - TRAAVGTGGCVCCCTGGGC - 3’ | |||
| HEV RT-PCR 4 | Primer_HEV_Fw4 | 5’ - GYGAGCTYATACGYGGGGC - 3’ | ORF1 | 1335 bp |
| Primer_HEV_Rv4 | 5’ - TAGCARTGTGCTATGATCGCC - 3’ | |||
| HEV RT-PCR 5 | Primer_HEV_Fw5 | 5’ - TCYCTYGGCCTTGAGTGTGT - 3’ | ORF1 / ORF2 / ORF3 | 1491 bp |
| Primer_HEV_Rv5 | 5’ - TCTCRACAGAGCGCCARCC - 3’ | |||
| HEV RT-PCR 6 | Primer_HEV_Fw6 | 5’ - TGGTGCCRAATGCYGTYGG - 3’ | ORF3 | 1019 bp |
| Primer_HEV_Rv6 | 5’ - ARAATGTYTTRGARTACTGCTG - 3’ |
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2026 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).




