Figure 1.
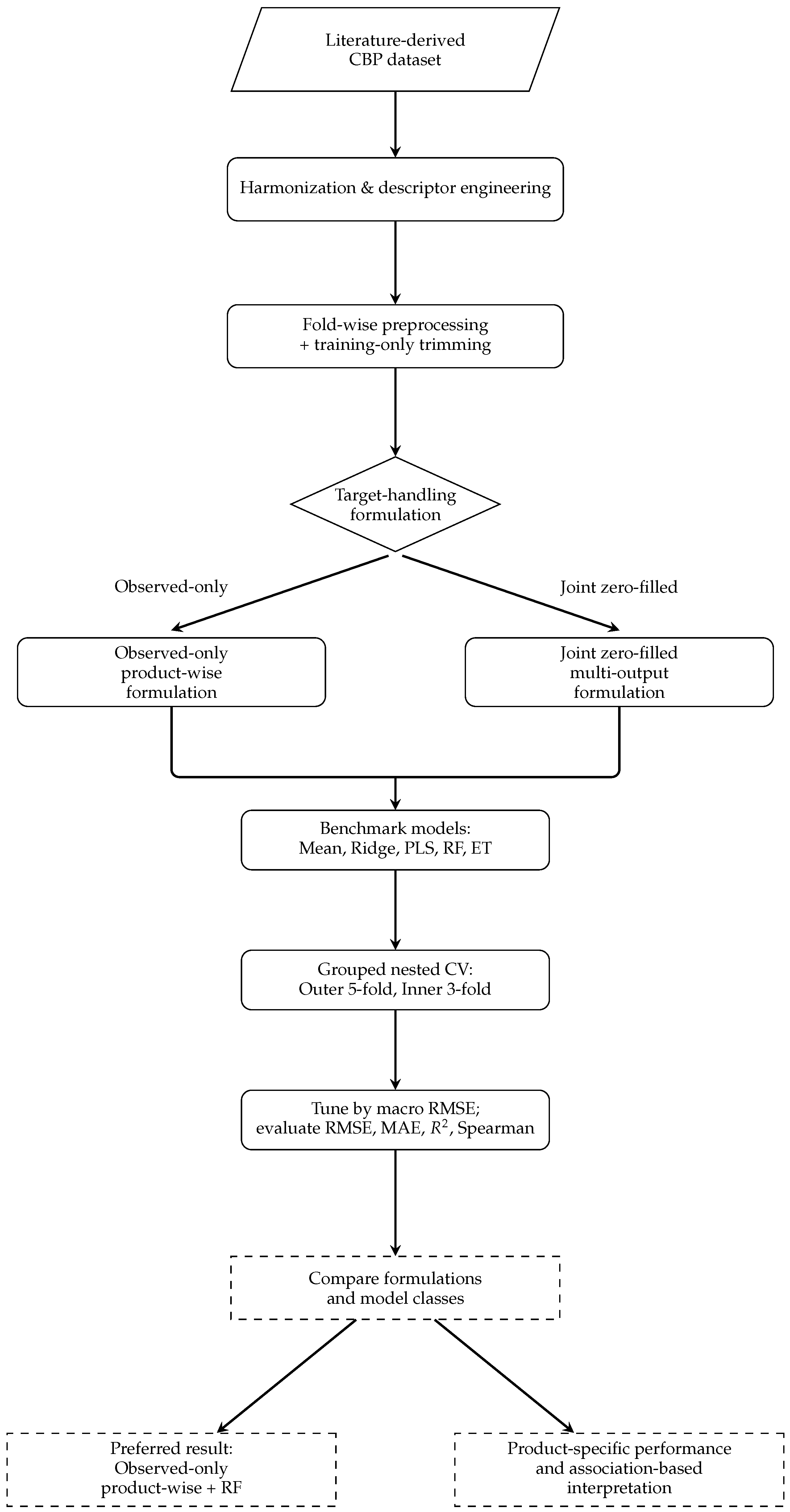
Simplified methodological workflow for literature-derived multi-product CBP modeling. The pipeline branches only at the target-handling step, contrasting the observed-only product-wise formulation with the joint zero-filled multi-output formulation, while preprocessing, benchmarking, grouped nested cross-validation, and evaluation remain shared across formulations.
Figure 1.
Simplified methodological workflow for literature-derived multi-product CBP modeling. The pipeline branches only at the target-handling step, contrasting the observed-only product-wise formulation with the joint zero-filled multi-output formulation, while preprocessing, benchmarking, grouped nested cross-validation, and evaluation remain shared across formulations.
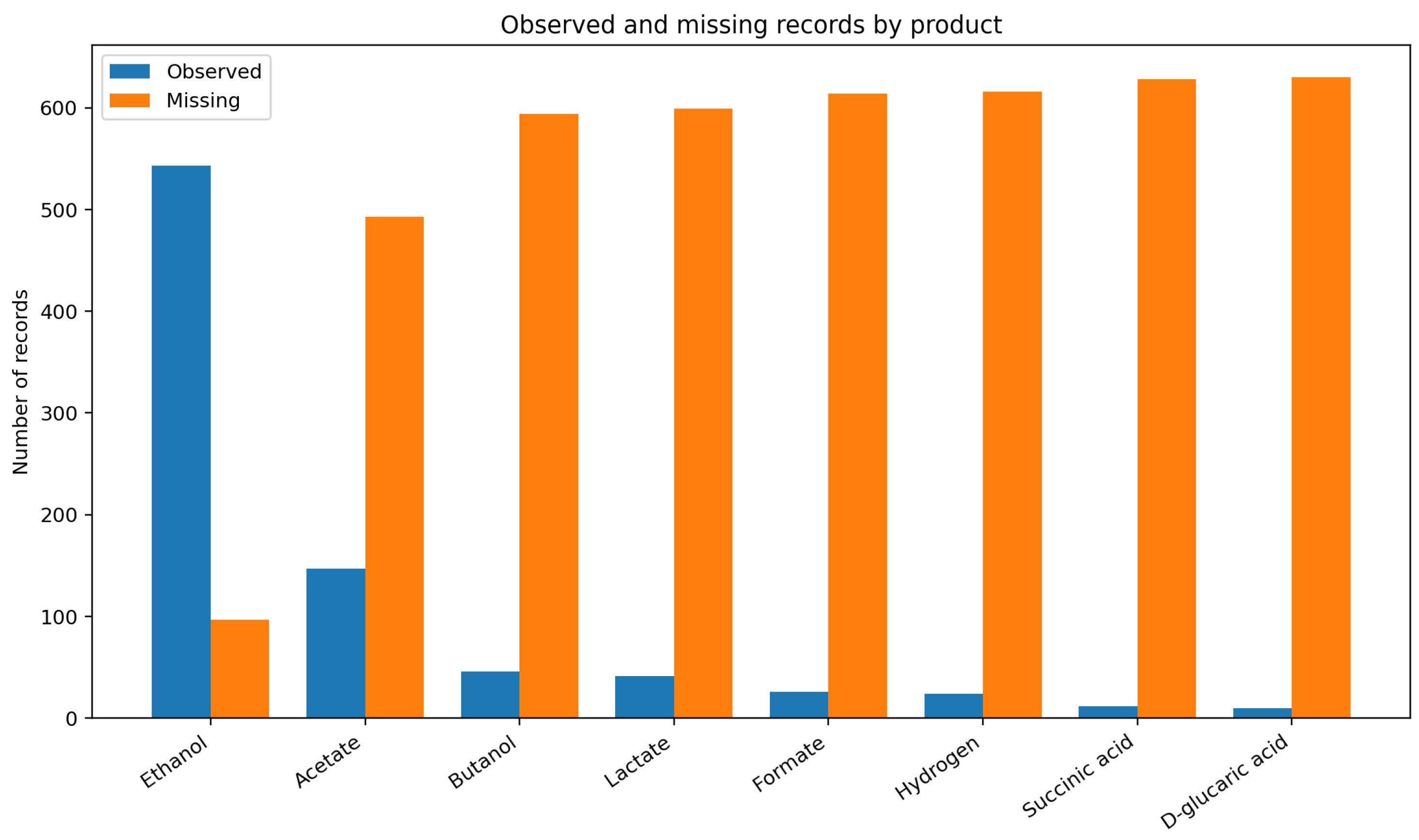
Figure 2.
Observed and unreported records by product in the literature-derived CBP dataset. Ethanol was the only well-supported response, whereas all co-products were much more sparsely reported. This imbalance is a defining feature of the dataset and constrains model training, validation, and interpretation.
Figure 2.
Observed and unreported records by product in the literature-derived CBP dataset. Ethanol was the only well-supported response, whereas all co-products were much more sparsely reported. This imbalance is a defining feature of the dataset and constrains model training, validation, and interpretation.
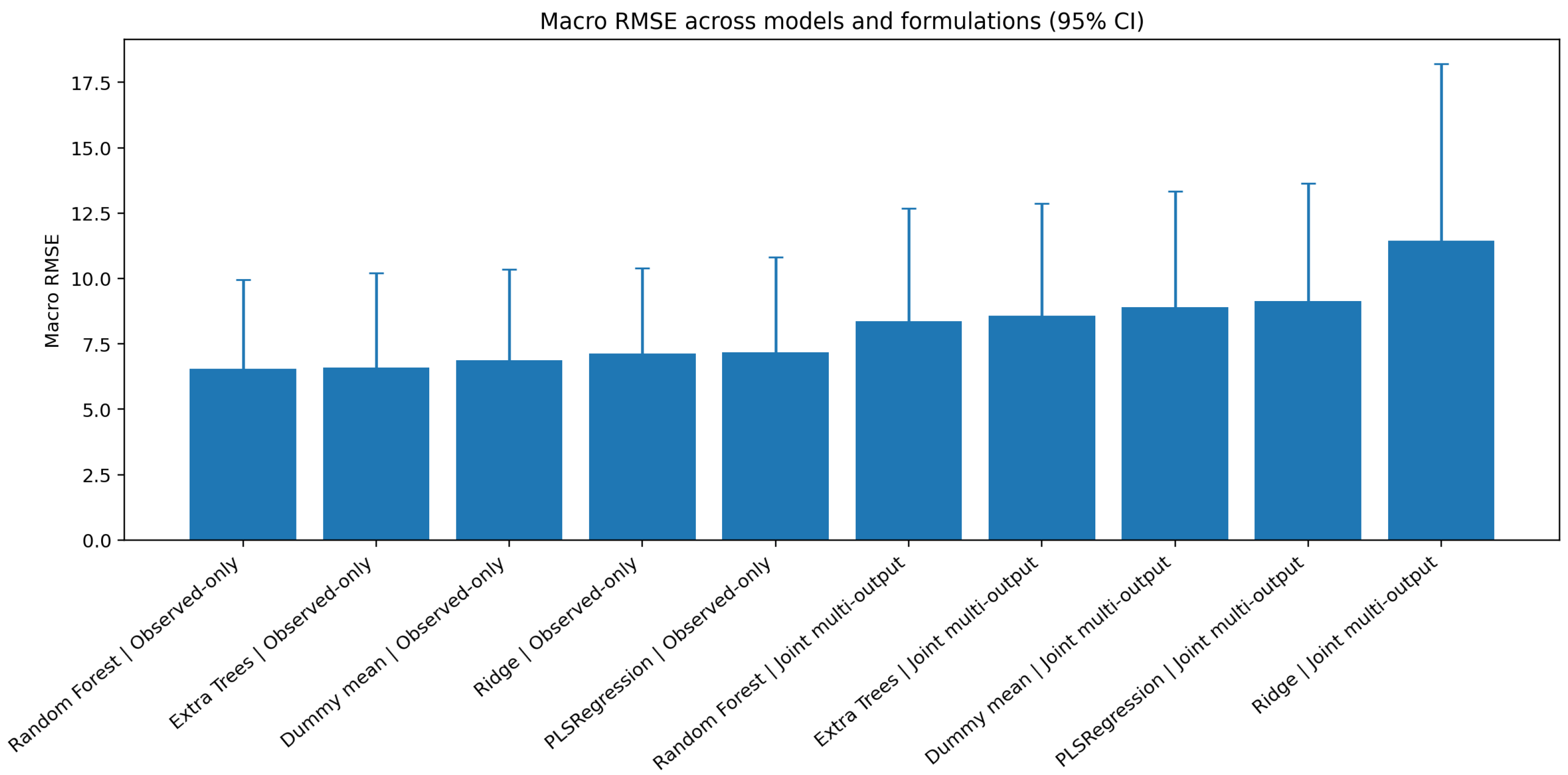
Figure 3.
Nested cross-validation macro RMSE across candidate models and target-handling formulations (95% confidence intervals). The observed-only product-wise formulation generally yielded lower error than the joint zero-filled multi-output formulation.
Figure 3.
Nested cross-validation macro RMSE across candidate models and target-handling formulations (95% confidence intervals). The observed-only product-wise formulation generally yielded lower error than the joint zero-filled multi-output formulation.
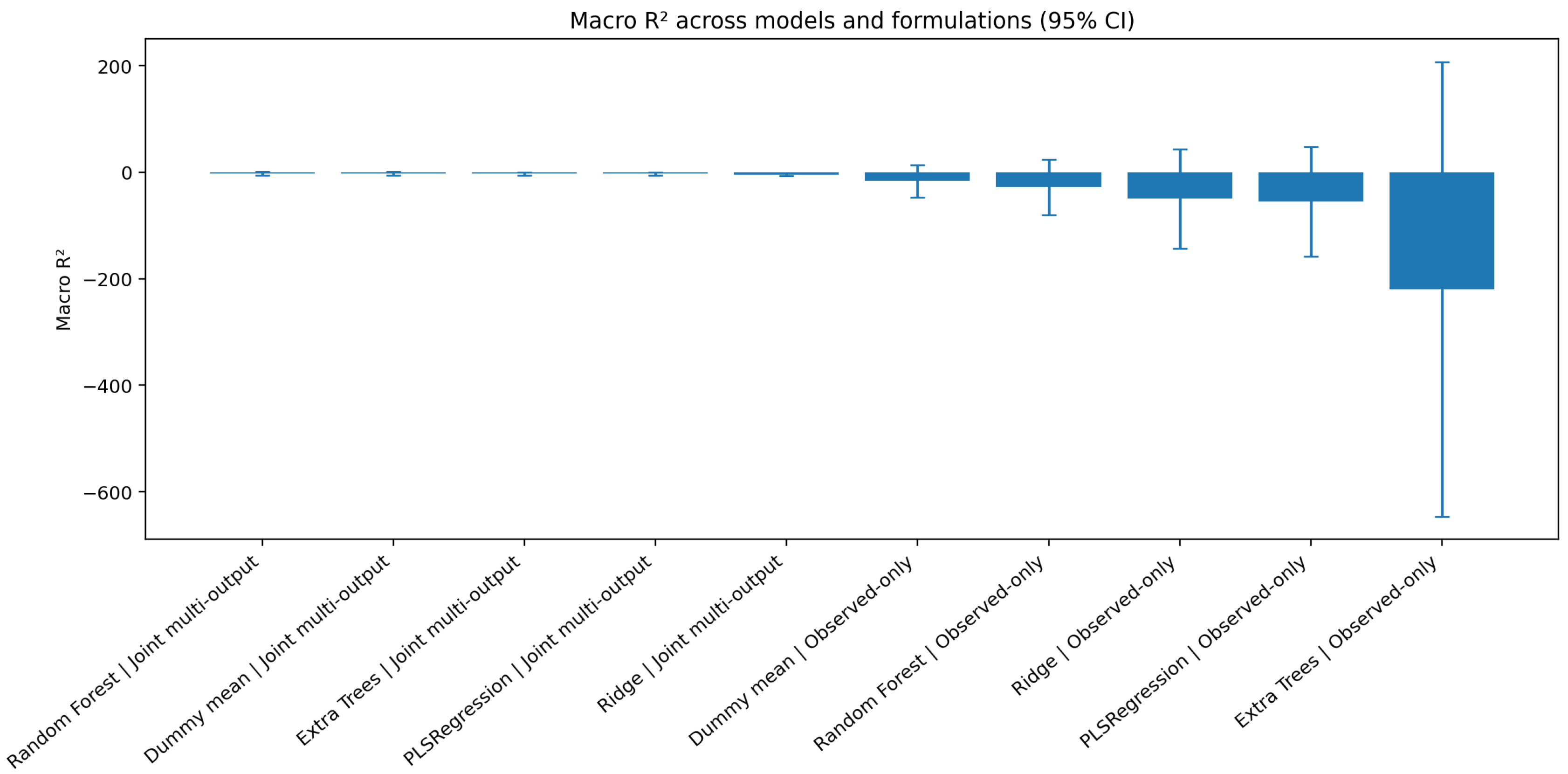
Figure 4.
Nested cross-validation macro across candidate models and target-handling formulations (95% confidence intervals). Although the observed-only product-wise formulation improved relative performance, remained weak or unstable in several model settings.
Figure 4.
Nested cross-validation macro across candidate models and target-handling formulations (95% confidence intervals). Although the observed-only product-wise formulation improved relative performance, remained weak or unstable in several model settings.
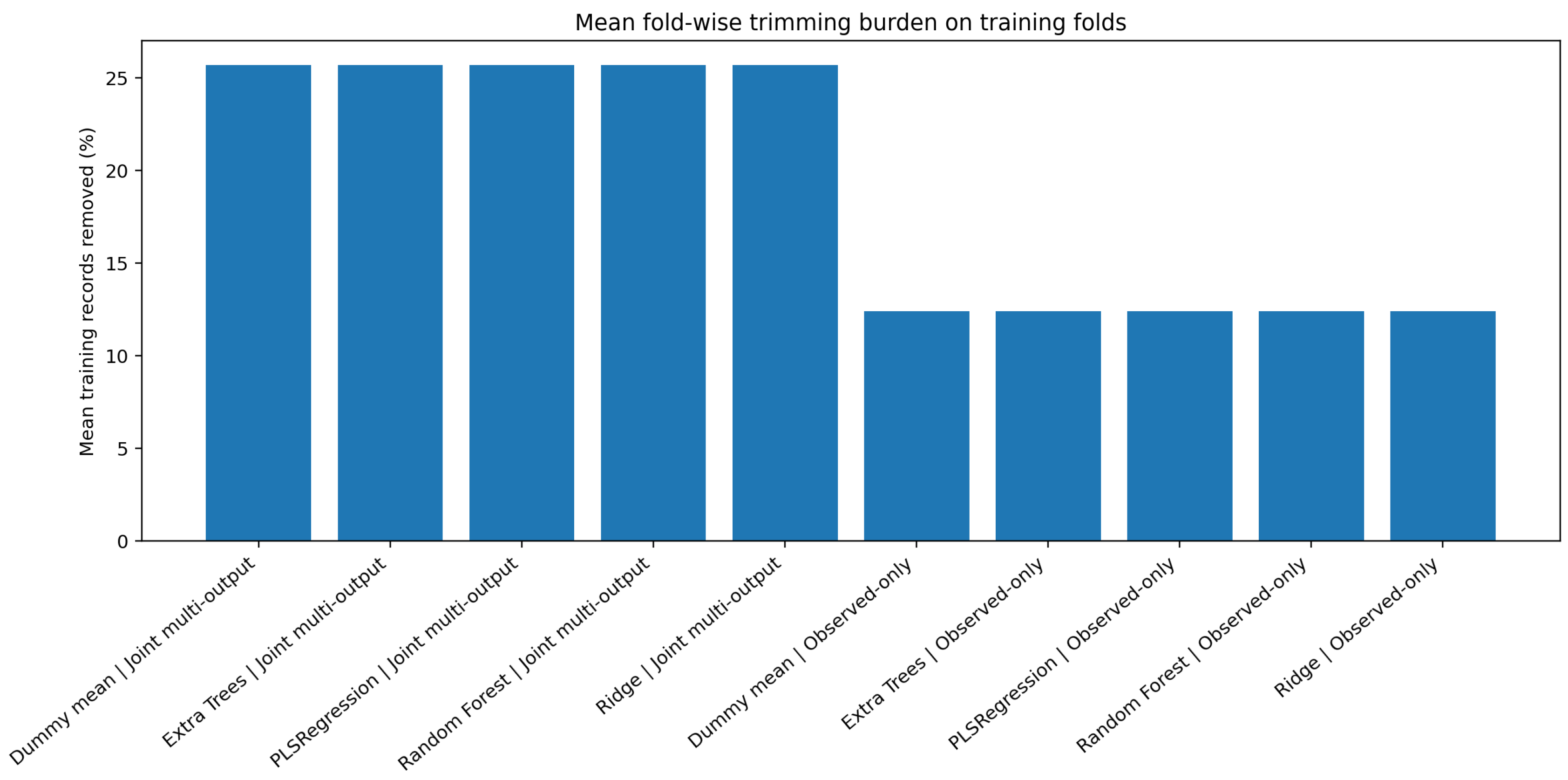
Figure 5.
Mean fold-wise trimming burden on training folds. The joint zero-filled multi-output formulation consistently required heavier filtering than the observed-only product-wise formulation.
Figure 5.
Mean fold-wise trimming burden on training folds. The joint zero-filled multi-output formulation consistently required heavier filtering than the observed-only product-wise formulation.
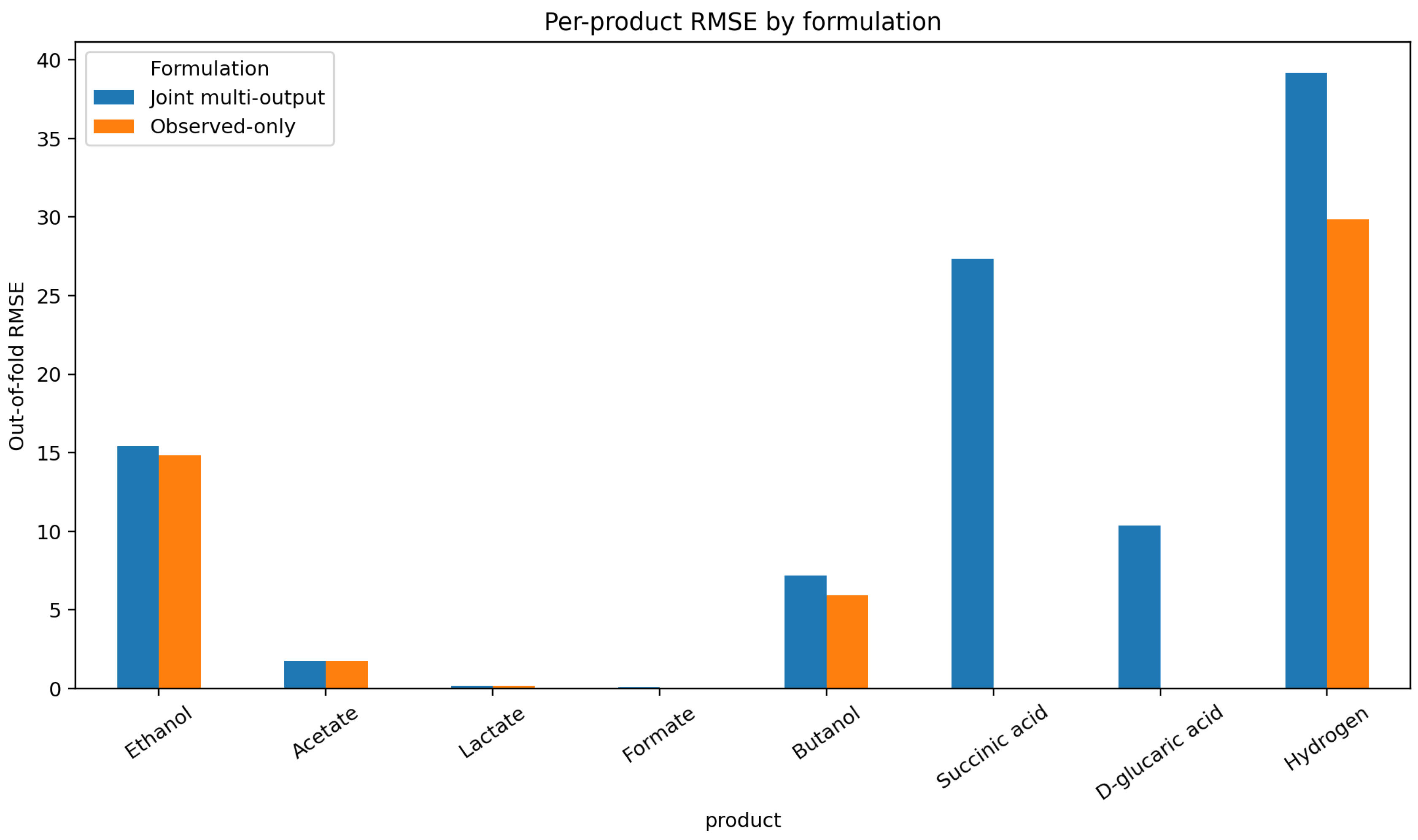
Figure 6.
Per-product RMSE under the two target-handling formulations. The observed-only product-wise formulation shows the clearest gains for butanol, hydrogen, and ethanol, whereas the sparsest products remain weak overall.
Figure 6.
Per-product RMSE under the two target-handling formulations. The observed-only product-wise formulation shows the clearest gains for butanol, hydrogen, and ethanol, whereas the sparsest products remain weak overall.
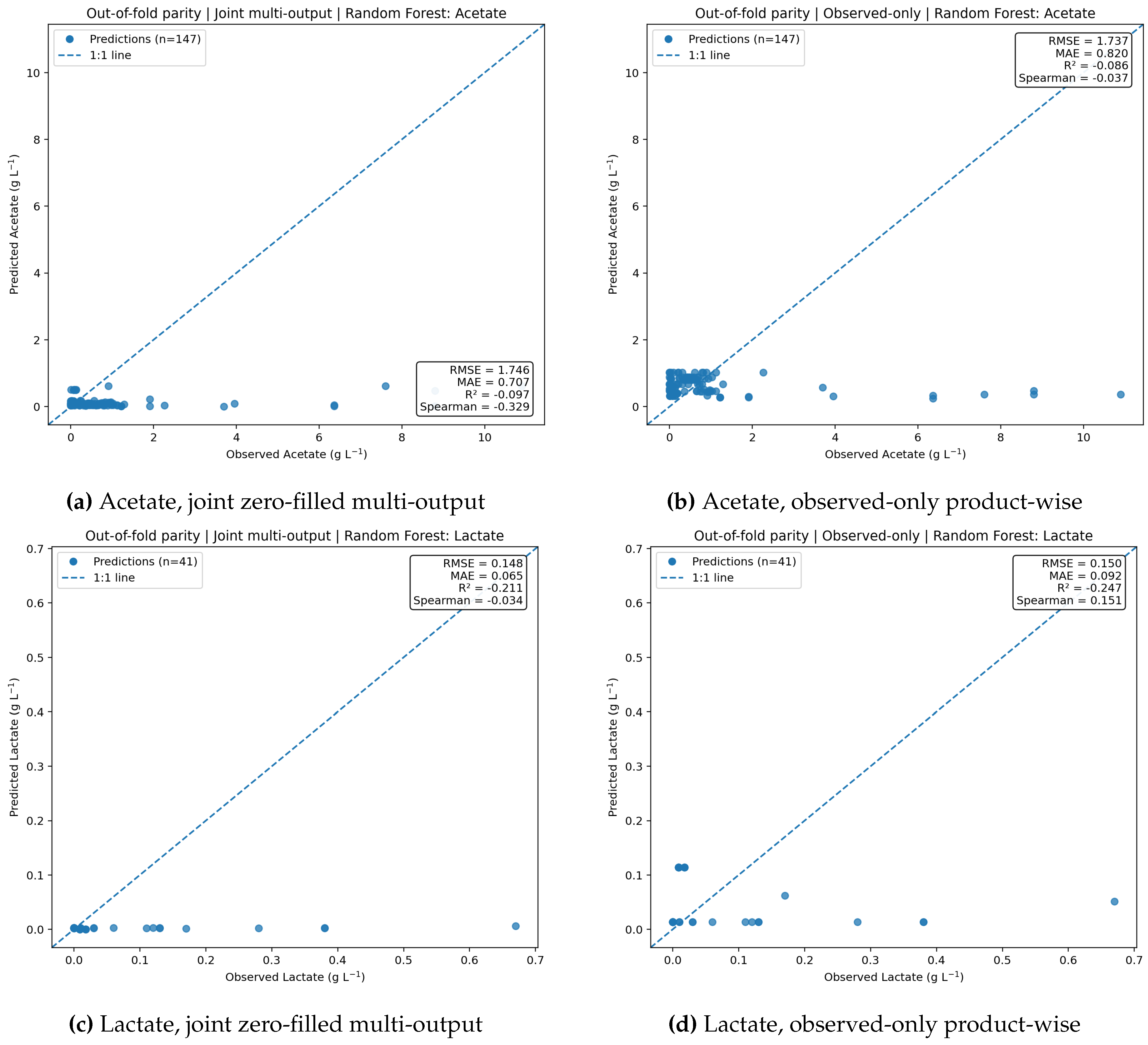
Figure 7.
Parity diagnostics for acetate and lactate under the two target-handling formulations. For both products, predictions remain compressed toward low values, and departures from the 1:1 line become more evident at higher observed responses. The observed-only product-wise formulation provides only limited improvement in calibration.
Figure 7.
Parity diagnostics for acetate and lactate under the two target-handling formulations. For both products, predictions remain compressed toward low values, and departures from the 1:1 line become more evident at higher observed responses. The observed-only product-wise formulation provides only limited improvement in calibration.
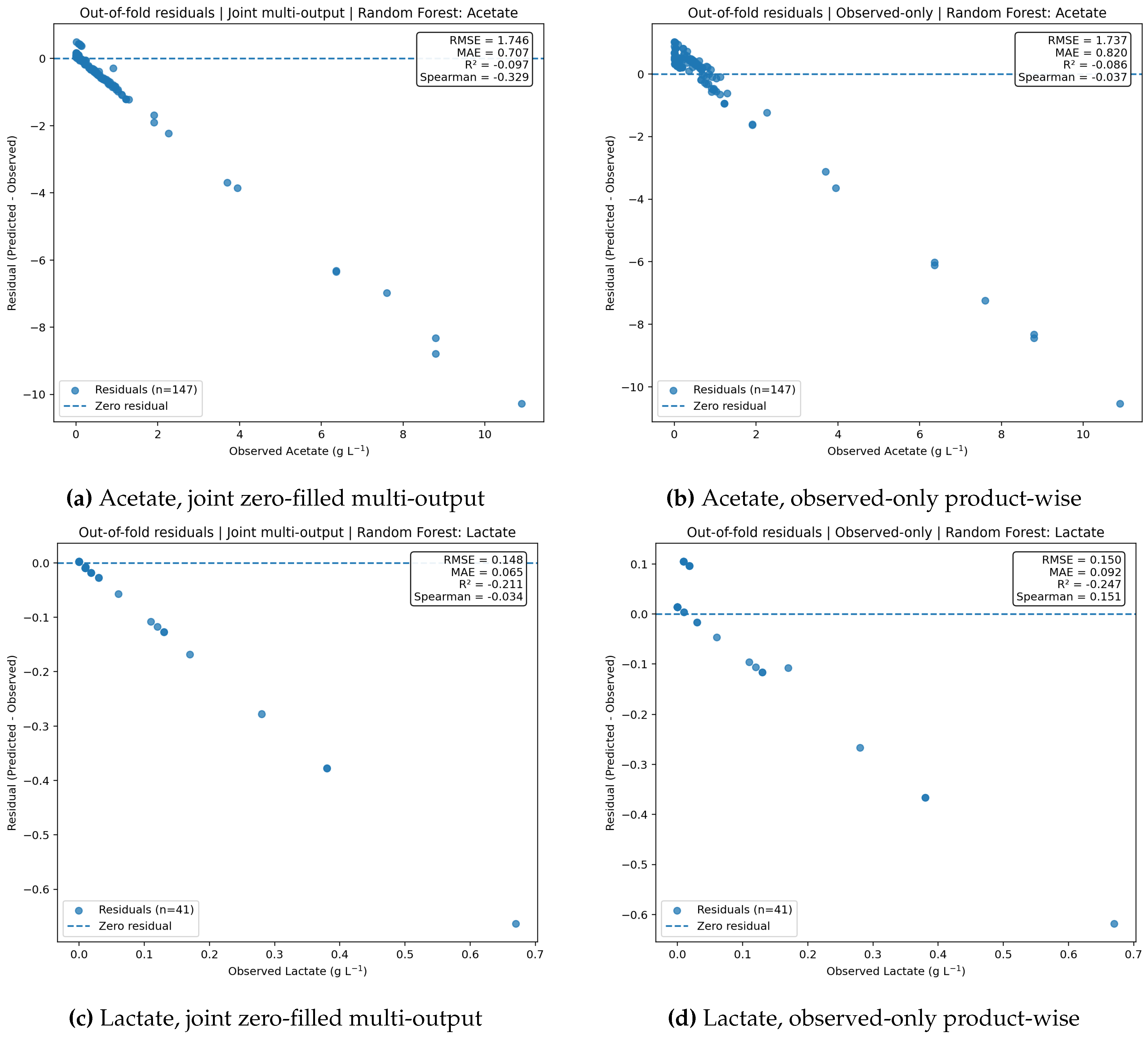
Figure 8.
Residual diagnostics for acetate and lactate under the two target-handling formulations. Both products show increasingly negative residuals at higher observed values, indicating persistent underprediction of upper-range responses. The observed-only product-wise formulation reduces this bias only modestly.
Figure 8.
Residual diagnostics for acetate and lactate under the two target-handling formulations. Both products show increasingly negative residuals at higher observed values, indicating persistent underprediction of upper-range responses. The observed-only product-wise formulation reduces this bias only modestly.
Table 1.
Short summary of the literature-derived CBP dataset used in this study.
Table 1.
Short summary of the literature-derived CBP dataset used in this study.
| Attribute |
Value |
| Records |
640 |
| Variables |
118 |
| Representation |
Experimental endpoint level |
| Supervised targets |
8 products |
| Input descriptor groups |
Biomass, pretreatment, microbial system, reactor/operation |
| Response standardization |
Liquid products in g L−1; hydrogen in mmol L−1
|
Table 2.
Selected Random Forest configuration and best inner-cross-validation performance for each formulation.
Table 2.
Selected Random Forest configuration and best inner-cross-validation performance for each formulation.
| Formulation |
Trees |
Max depth |
Min leaf |
Mean inner macro RMSE |
| Joint zero-filled multi-output |
300 |
8 |
3 |
10.688 |
| Observed-only product-wise |
300 |
8 |
3 |
7.266 |
Table 3.
Best-model out-of-fold macro metrics by target-handling formulation. In both cases, Random Forest was selected as the best overall model.
Table 3.
Best-model out-of-fold macro metrics by target-handling formulation. In both cases, Random Forest was selected as the best overall model.
| Formulation |
Macro RMSE |
Macro MAE |
Macro
|
Macro Spearman |
| Joint zero-filled multi-output |
12.68 |
9.40 |
-4.29 |
-0.003 |
| Observed-only product-wise |
10.49 |
6.16 |
-0.04 |
0.255 |
Table 4.
Mean outer-fold macro performance of candidate models under the preferred observed-only product-wise formulation.
Table 4.
Mean outer-fold macro performance of candidate models under the preferred observed-only product-wise formulation.
| Model |
Mean RMSE |
SD |
Mean MAE |
Spearman |
| Random Forest |
6.54 |
3.88 |
4.64 |
0.250 |
| Extra Trees |
6.59 |
4.11 |
4.75 |
0.250 |
| Mean baseline |
6.87 |
3.97 |
5.07 |
— |
| Ridge |
7.13 |
3.72 |
5.40 |
0.040 |
| PLS regression |
7.18 |
4.16 |
5.44 |
0.078 |
Table 5.
Outer-fold best-model frequency under each target-handling formulation.
Table 5.
Outer-fold best-model frequency under each target-handling formulation.
| Model |
Joint zero-filled multi-output |
Observed-only product-wise |
| Random Forest |
5/5 |
3/5 |
| Extra Trees |
0/5 |
2/5 |
| Ridge |
0/5 |
0/5 |
| PLS regression |
0/5 |
0/5 |
| Mean baseline |
0/5 |
0/5 |
Table 6.
Summary of trimming burden for the final selected formulations.
Table 6.
Summary of trimming burden for the final selected formulations.
| Formulation |
Mean fold-wise trimming burden (%) |
Final-fit trimmed fraction (%) |
| Joint zero-filled multi-output |
25.7 |
30.0 |
| Observed-only product-wise |
12.5 |
— |
Table 7.
Product-specific row support and trimming burden for the final observed-only product-wise Random Forest fit.
Table 7.
Product-specific row support and trimming burden for the final observed-only product-wise Random Forest fit.
| Product |
Final observed training rows |
Trimmed rows |
Trimmed fraction (%) |
| Ethanol |
543 |
162 |
29.8 |
| Acetate |
147 |
29 |
19.7 |
| Butanol |
46 |
1 |
2.2 |
| Lactate |
41 |
2 |
4.9 |
| Formate |
26 |
0 |
0.0 |
| Hydrogen |
24 |
7 |
29.2 |
| Succinic acid |
12 |
0 |
0.0 |
|
D-glucaric acid |
10 |
2 |
20.0 |
Table 8.
Observed support and best-model out-of-fold performance under the preferred observed-only product-wise formulation.
Table 8.
Observed support and best-model out-of-fold performance under the preferred observed-only product-wise formulation.
| Product |
Observed records |
RMSE |
Spearman |
| Ethanol |
543 |
14.82 |
0.476 |
| Acetate |
147 |
1.74 |
-0.037 |
| Butanol |
46 |
5.92 |
0.612 |
| Lactate |
41 |
0.150 |
0.151 |
| Hydrogen |
24 |
29.83 |
0.074 |