Submitted:
26 March 2026
Posted:
31 March 2026
You are already at the latest version
Abstract
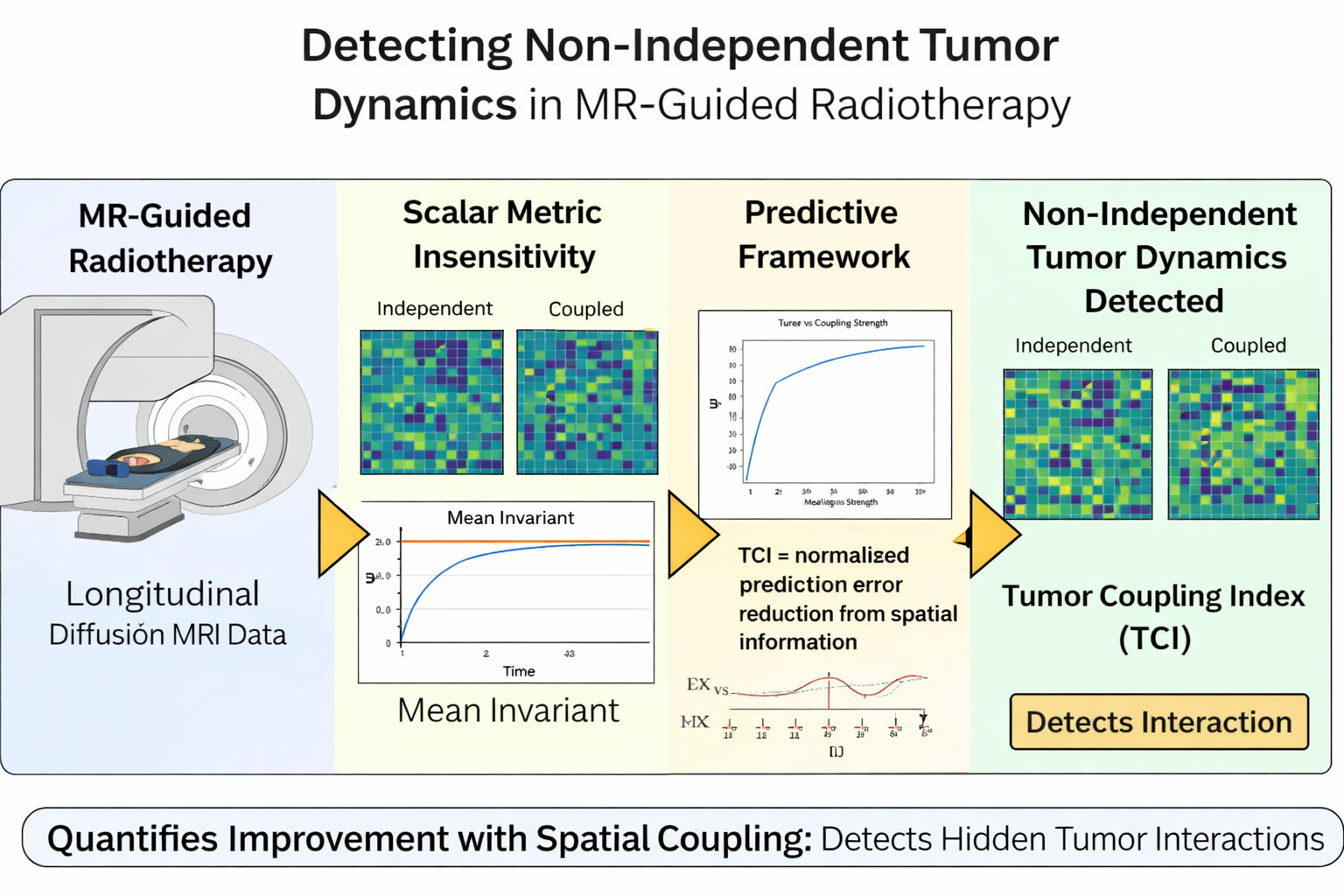
Keywords:
1. Introduction
1.1. Background and Motivation
1.2. Limitations of Existing Approaches
- mechanistic models, which explicitly incorporate interaction but are difficult to validate
- clinical metrics, which are robust but insensitive to spatial structure
1.3. A Predictive Perspective on Tumor Dynamics
- modeling what generates the data
- to measuring what can be predicted from the data
1.4. Contribution of This Work
-
Observable-based formulation:Spatial interaction is defined operationally through improvement in predictive performance rather than through explicit parameter estimation.
-
Quantitative metric:The Tumor Coupling Index provides a normalized measure of predictive gain attributable to spatial information.
-
Theoretical insight:We demonstrate that commonly used scalar metrics are inherently insensitive to spatial coupling, establishing a fundamental identifiability limitation.
-
Falsifiability:The framework is constructed such that the metric returns near-zero values under independent or randomized conditions, ensuring that detected structure is not spurious.
-
Simulation-based validation:Controlled experiments confirm that the proposed metric distinguishes independent and coupled systems and varies systematically with coupling strength.
1.5. Scope and Positioning
1.6. Structure of the Paper
2. Theoretical Framework
2.1. Tumor as a Spatially Distributed Dynamical System
2.2. Baseline Model: Independent Tumor Dynamics
- is the equilibrium value
- is the local response rate
- no interaction between voxels
- purely local evolution
2.3. Spatially Coupled Dynamics
- is the coupling strength
- denotes neighboring voxels
- local smoothing
- information propagation across the tumor
2.4. Identifiability Limitation of Scalar Metrics
- identical local dynamics
- different spatial coupling
2.5. From Identifiability to Observability
2.6. Predictive Observability Framework
2.7. Definition of Tumor Coupling Index (TCI)
- : independent dynamics
- : predictive gain → spatial interaction
2.8. Falsifiability
2.9. Conceptual Synthesis
- from estimating unobservable coupling parameters
- to measuring observable predictive consequences of interaction
3. Methods
3.1. Overview
3.2. Simulation of Tumor Dynamics
- controls decay (e.g., treatment effect)
- controls spatial coupling strength
- controls temporal dependence
- is a latent memory variable
- represents noise
- denotes the set of neighboring voxels
3.3. Simulation Scenarios
- Independent:
- Temporal-only:
- Spatial-only:
- Combined:
3.4. Data Preparation
- Independent model:
- Temporal model:
- Spatial model:
3.5. Predictive Models
- ensure interpretability
- avoid overfitting
- provide a conservative estimate of predictive gain
3.6. Tumor Coupling Index (TCI)
- : error of the independent model
- : error of the spatial model
3.7. Falsification Procedures
- Temporal shuffling: voxel time series randomly permuted
- Spatial shuffling: voxel locations randomized at each time point
- TCI ≈ 0 under shuffled conditions
3.8. Statistical Evaluation
- mean TCI
- standard deviation across runs
3.9. Parameter Settings
- (spatial regimes)
- (temporal regimes)
3.10. Implementation
4. Results
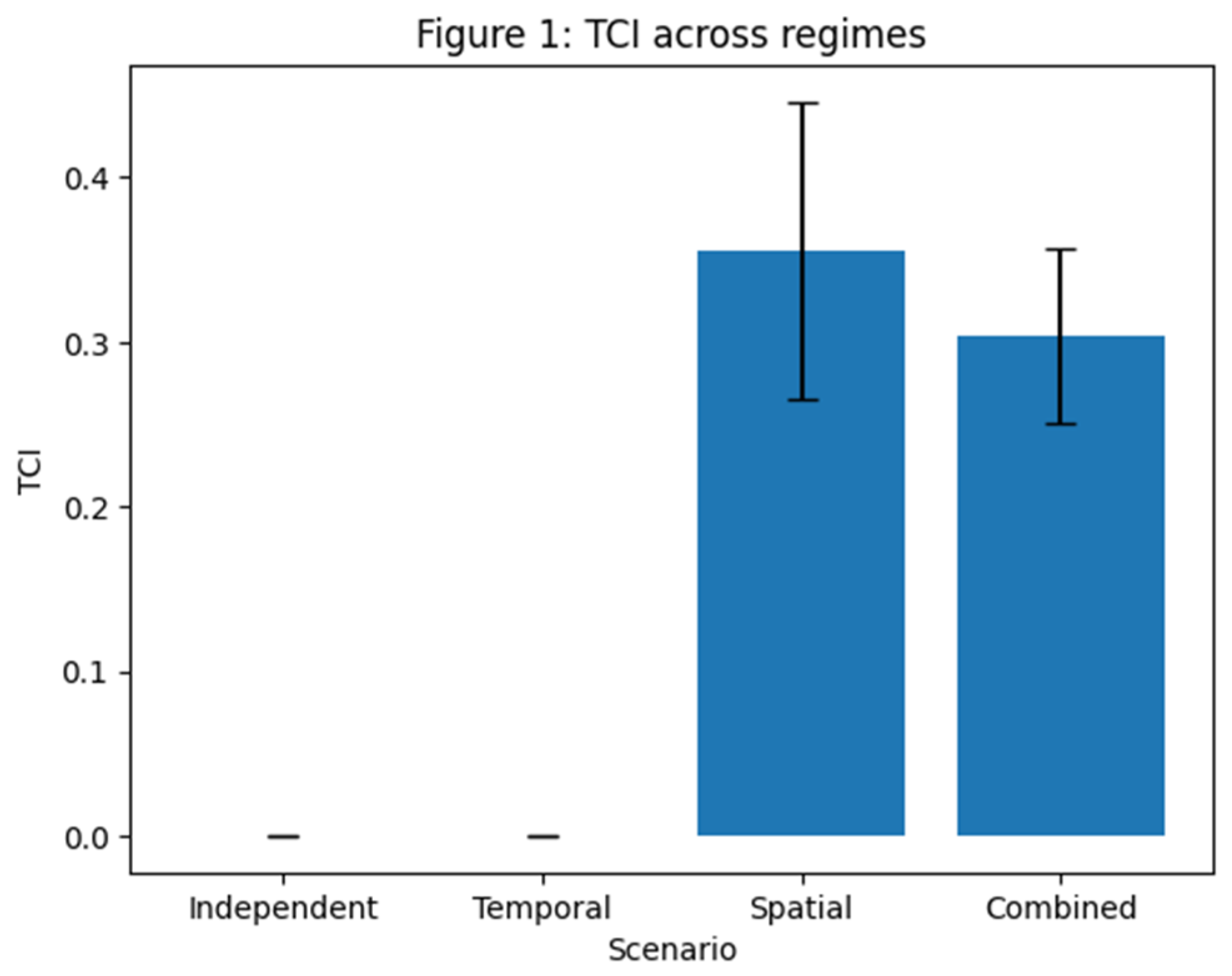
4.1. Detection of Non-Independent Dynamics Across Simulation Regimes
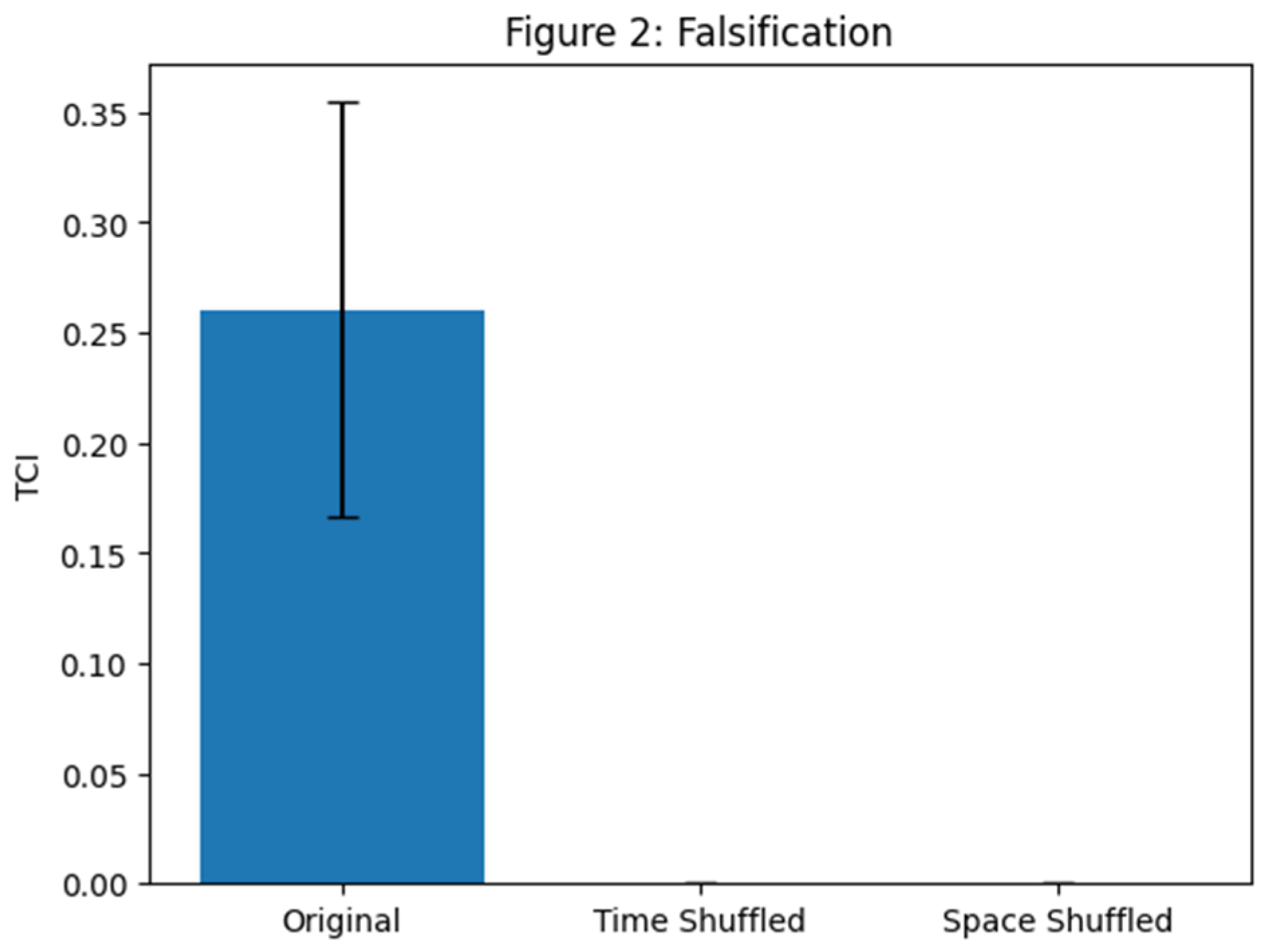
4.2. Falsifiability and Specificity of the Metric
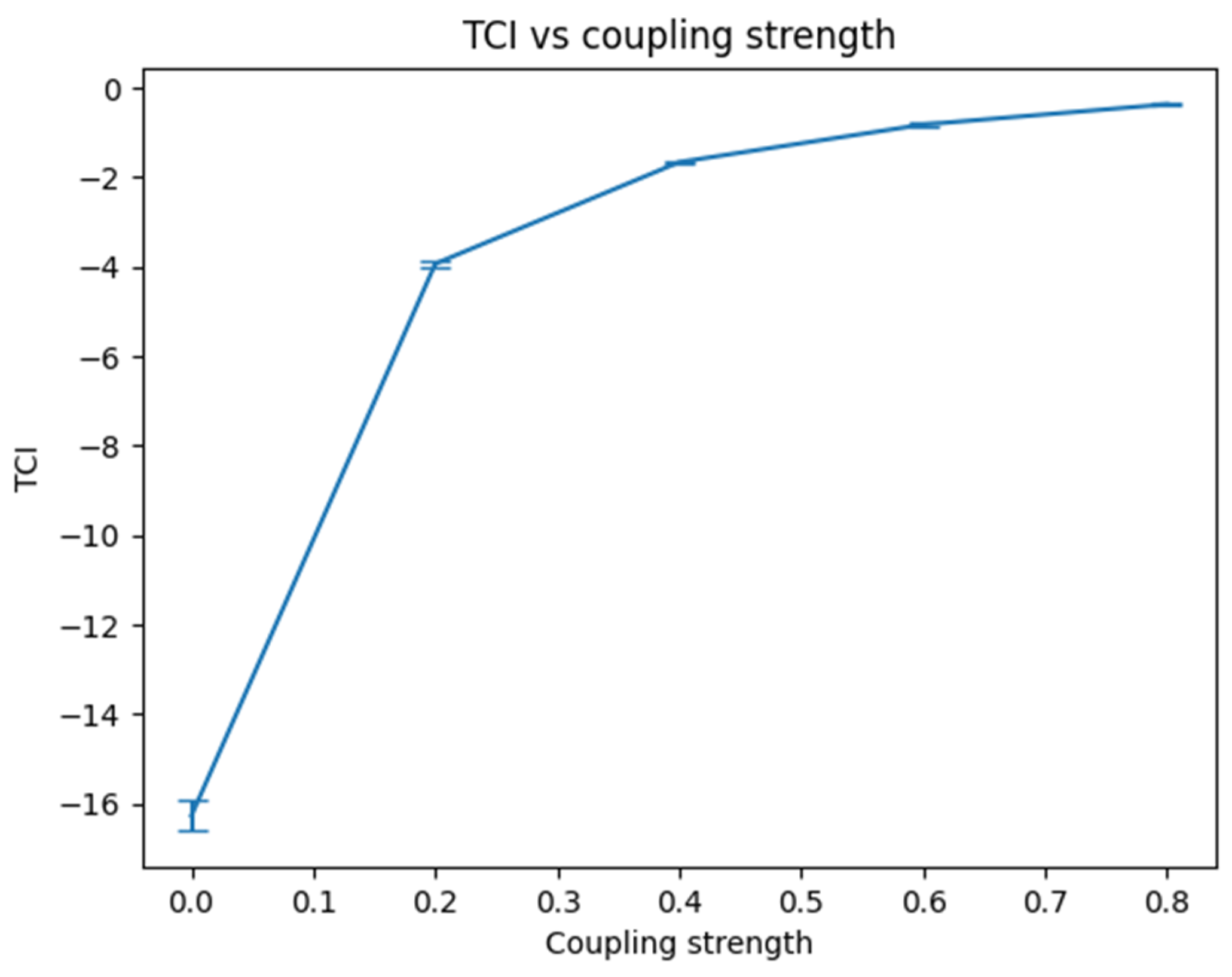
4.3. Dependence of TCI on Spatial Interaction Strength
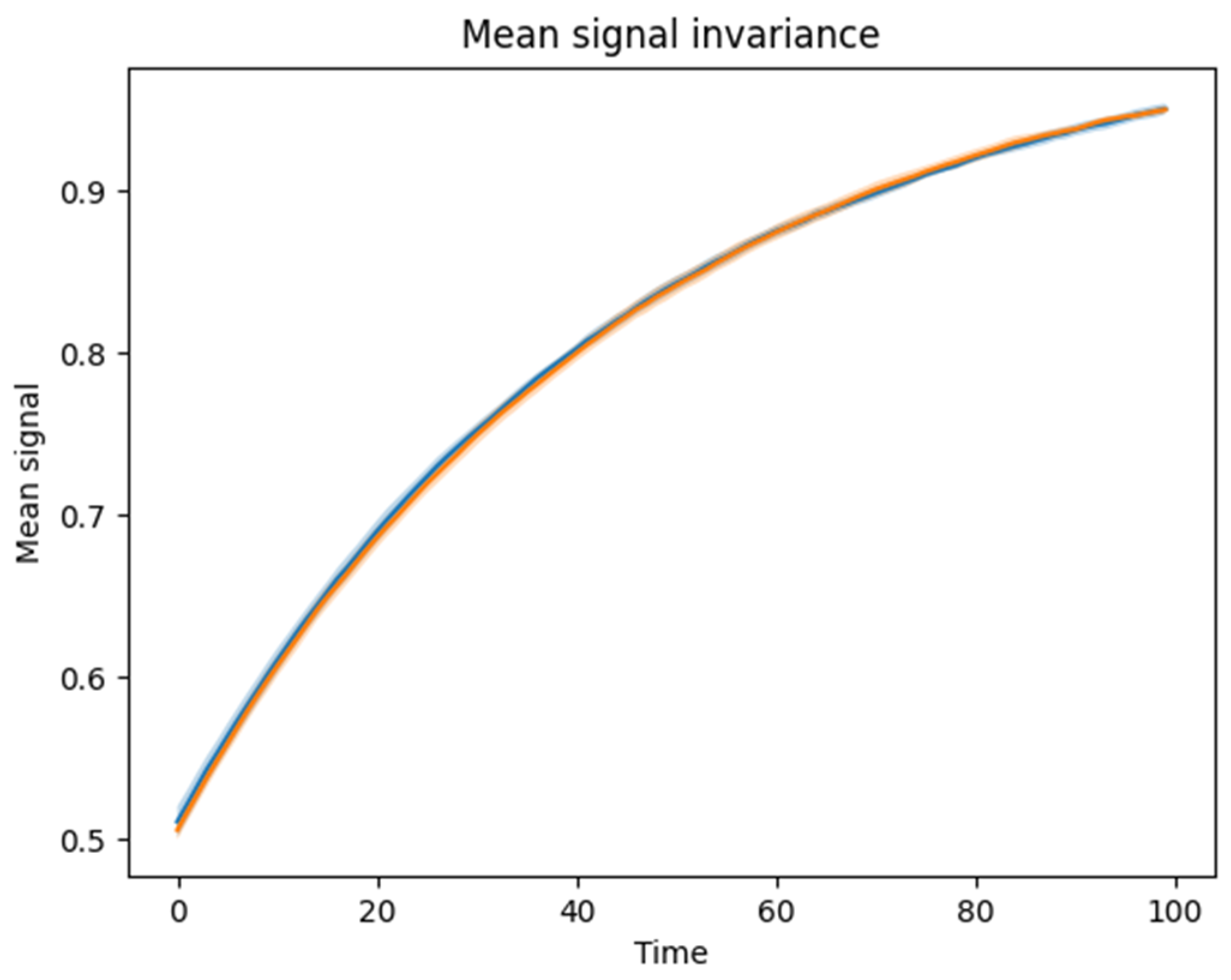
4.4. Invariance of Mean-Based Metrics to Spatial Coupling
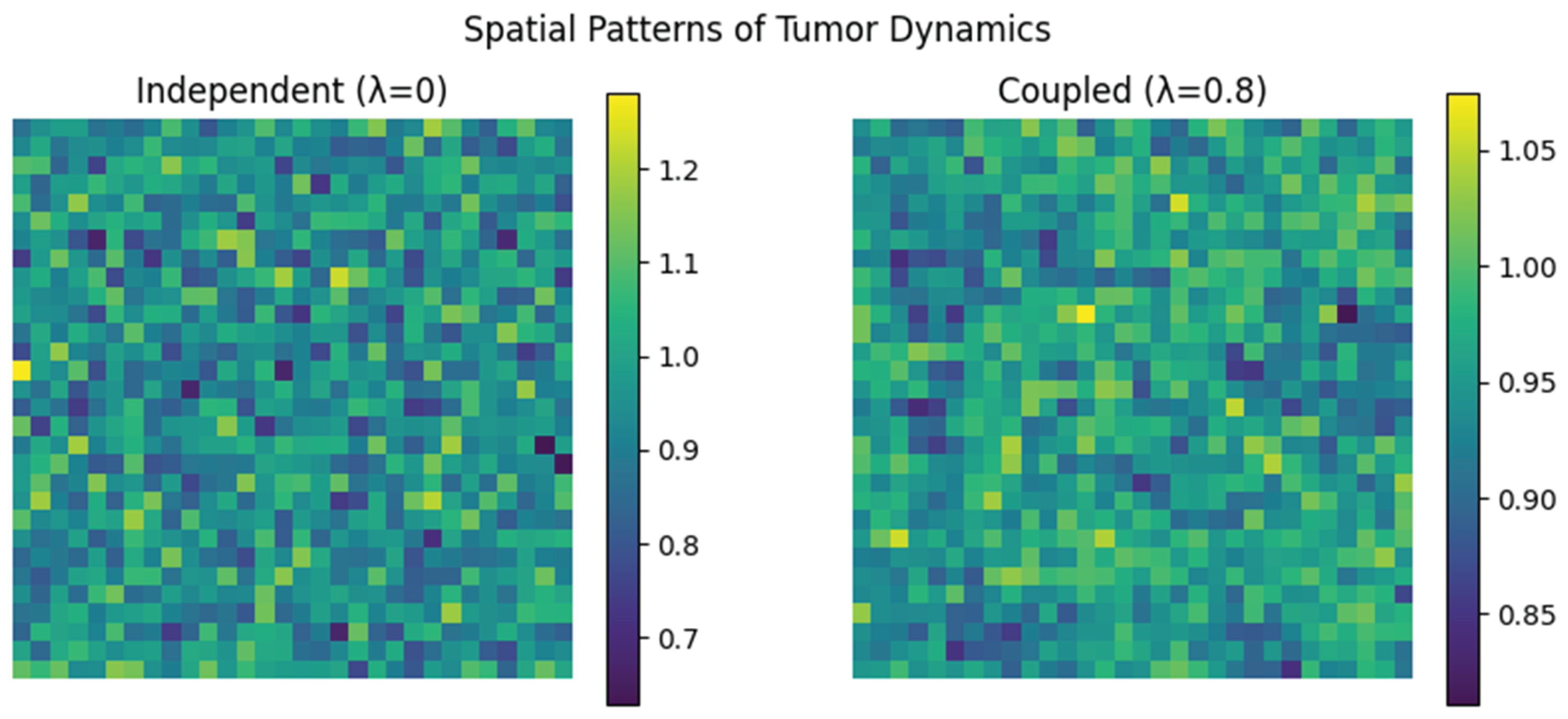
4.5. Spatial Patterns Reveal Qualitative Differences in Tumor Dynamics
5. Discussion
5.1. Summary of Findings
5.2. Relation to Existing Theory
5.3. Clinical Implications
5.3.1. From Static Metrics to Dynamic Monitoring
5.3.2. TCI as a Monitoring Signal During Radiotherapy
- Low TCI → dynamics consistent with independent or unstructured response
- Elevated TCI → presence of coordinated or structured behavior
- spatially coordinated resistance
- heterogeneous response patterns
- redistribution of tumor activity under treatment
5.3.3. Potential Role in Adaptive Radiotherapy
-
Stable, low TCI:Dynamics consistent with approximately independent voxel behavior, suggesting that standard independence-based response models remain applicable.
-
Increasing TCI during treatment:Emergence of structured or coordinated dynamics, indicating a potential deviation from independence assumptions and warranting closer monitoring or reassessment of model-based interpretation.
-
Reduction in TCI following intervention:Decrease in coordinated dynamics, which may reflect disruption of structured behavior and could be consistent with effective treatment response, although this interpretation requires further validation.
-
Persistently elevated TCI despite apparent response:Continued presence of coordinated dynamics even when scalar metrics (e.g., mean ADC) suggest favorable response. This may indicate that independence-based models are overestimating treatment effectiveness, and that model predictions should be interpreted with reduced confidence.
-
Divergence between scalar metrics and TCI trends:Situations in which conventional metrics indicate stable or improving response, while TCI increases over time. Such divergence may signal a breakdown of underlying model assumptions, suggesting the need for incorporating spatially informed or alternative modeling approaches.
5.3.4. Practical Considerations for Clinical Implementation
- using simple predictive models
- producing an output (TCI)
- avoiding reliance on unobservable parameters
5.4. Limitations
5.5. Future Work
6. Conclusion
Data Availability Statement
Conflicts of Interest
References
- Enderling, H; Chaplain, MAJ. Mathematical modeling of tumor growth and treatment. Curr Pharm Des. 2014, 20(30), 4934–40. [Google Scholar] [CrossRef] [PubMed]
- Gatenby, RA; Silva, AS; Gillies, RJ; Frieden, BR. Adaptive therapy. Cancer Res. 2009, 69(11), 4894–903. [Google Scholar] [CrossRef] [PubMed]
- Granger, CWJ. Investigating causal relations by econometric models and cross-spectral methods. Econometrica 1969, 37(3), 424–38. [Google Scholar] [CrossRef]
- Greaves, M; Maley, CC. Clonal evolution in cancer. Nature 2012, 481(7381), 306–13. [Google Scholar] [CrossRef] [PubMed]
- Hormuth, DA, 2nd; Phillips, CM; Wu, C; Lima, EABF; Lorenzo, G; Kalpathy-Cramer, J; et al. Biologically-based mathematical modeling of tumor growth and treatment response. Br J Radiol. 2018, 91(1090), 20180447. [Google Scholar]
- Jain, RK. Transport of molecules in the tumor interstitium: a review. Cancer Res. 1987, 47(12), 3039–51. [Google Scholar] [PubMed]
- Koh, TS; Collins, DJ. Perfusion MRI: the journey of an imaging biomarker. Br J Radiol. 2007, 80 Spec No 1, S10–17. [Google Scholar]
- Lagendijk, JJ; Raaymakers, BW; van Vulpen, M. The magnetic resonance imaging-guided radiotherapy (MRIgRT) system: a short introduction. Front Oncol. 2014, 4, 210. [Google Scholar]
- Mitchell, J; et al. Tumors as adaptive, integrated dynamical systems: Implications for radiotherapy response and emergent resistance. J Theor Biol. 2024, 578, 111682. [Google Scholar]
- Mora, T; Bialek, W. Selective erasure of physiological memory in biological networks without topological alteration. Phys Rev E 2025, 111(2), 024401. [Google Scholar]
- Mutic, S; Dempsey, JF. The ViewRay system: magnetic resonance-guided and adaptive radiotherapy. Semin Radiat Oncol. 2014, 24(3), 196–9. [Google Scholar] [CrossRef] [PubMed]
- O’Connor, JP; Aboagye, EO; Adams, JE; Aerts, HJ; Barrington, SF; Beer, AJ; et al. Imaging biomarker roadmap for cancer studies. Nat Rev Clin Oncol. 2017, 14(3), 169–86. [Google Scholar] [CrossRef] [PubMed]
- Osswald, M; Jung, E; Sahm, F; Solecki, G; Venkataramani, V; et al. Brain tumour cells interconnect to a functional and resistant network. Nature 2015, 528(7580), 93–8. [Google Scholar] [CrossRef] [PubMed]
- Padhani, AR; Liu, G; Koh, DM; Chenevert, TL; Thoeny, HC; Takahara, T; et al. Diffusion-weighted magnetic resonance imaging as a cancer biomarker: consensus and recommendations. Neoplasia 2009, 11(2), 102–25. [Google Scholar] [CrossRef] [PubMed]
- Pigozzi, G; et al. Resistance and recurrence as dynamical phenomena: A framework for history-dependent tumor organization. Nat Commun. 2026, 17(1), 412. [Google Scholar]
- Raaymakers, BW; Jürgenliemk-Schulz, IM; Bol, GH; Glitzner, M; Kotte, ANTJ; van Asselen, B; et al. First patients treated with a 1.5 T MRI-linac: clinical proof of concept of a high-precision, real-time adaptive radiotherapy system. Phys Med Biol. 2017, 62(23), L41–50. [Google Scholar] [CrossRef] [PubMed]
- Rajan, DH; Lee, B; Albright, A; Tang, E; Ressler, K; Maravillas, A; et al. Stentor stipatus is a new unicellular species that demonstrates habituation and unique phototaxis. Sci Rep. 2026, 16, 4027, (Note: Initial findings/preprints circulating in late 2024/2025 under bioRxiv). [Google Scholar] [CrossRef] [PubMed]
- Rockne, R; Rockhill, JK; Mrugala, M; Spence, AM; Kalet, I; Hendrickson, K; et al. Predicting the efficacy of radiotherapy in individual glioblastoma patients in vivo: a mathematical modeling approach. Phys Med Biol. 2010, 55(12), 3271–85. [Google Scholar] [CrossRef] [PubMed]
- Schreiber, T. Measuring information transfer. Phys Rev Lett. 2000, 85(2), 461–4. [Google Scholar] [CrossRef] [PubMed]
- Swanson, KR; Rostomily, RC; Alvord, EC, Jr. Virtual and real brain tumors: using mathematical modeling to quantify glioma growth and invasion. J Neurol Sci. 2003, 216(1), 1–10. [Google Scholar] [CrossRef] [PubMed]
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2026 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).



