Submitted:
27 March 2026
Posted:
27 March 2026
You are already at the latest version
Abstract
Keywords:
1. Introduction
| LIGAND | BIOISOSTERIC REPLACEMENT | SMILES | CODE NAME |
|---|---|---|---|
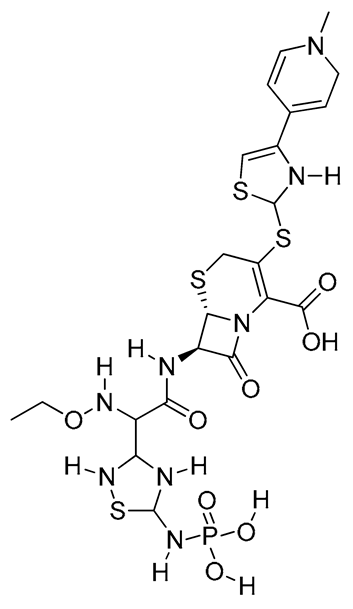 |
O=C(N2[C@H]1SCC(SC4SC=C(C5=CCN(C)C=C5)N4[H])=C2C(O)=O)[C@H]1N(C(C(C(N([H])C(S3)N([H])P(O[H])(O[H])=O)N3[H])N([H])OCC)=O)[H] | ||
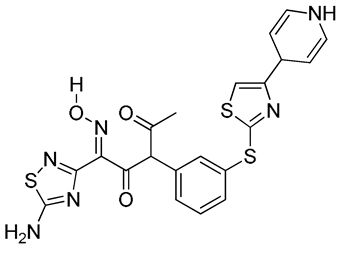 |
 |
O=C(/C(C4=NSC(N)=N4)=N/O[H])C(C1=CC=CC(SC2=NC(C3C=CNC=C3)=CS2)=C1)C(C)=O | 1 |
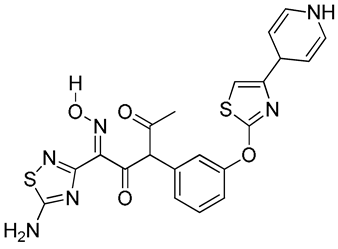 |
 |
O=C(/C(C4=NSC(N)=N4)=N/O[H])C(C1=CC=CC(OC2=NC(C3C=CNC=C3)=CS2)=C1)C(C)=O | 2 |
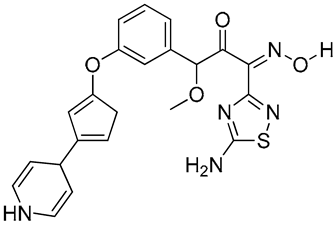 |
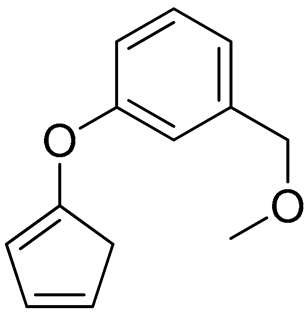 |
O=C(/C(C4=NSC(N)=N4)=N/O[H])C(OC)C1=CC=CC(OC2=CC(C3C=CNC=C3)=CC2)=C1 | 3 |
2. Materials and Methods
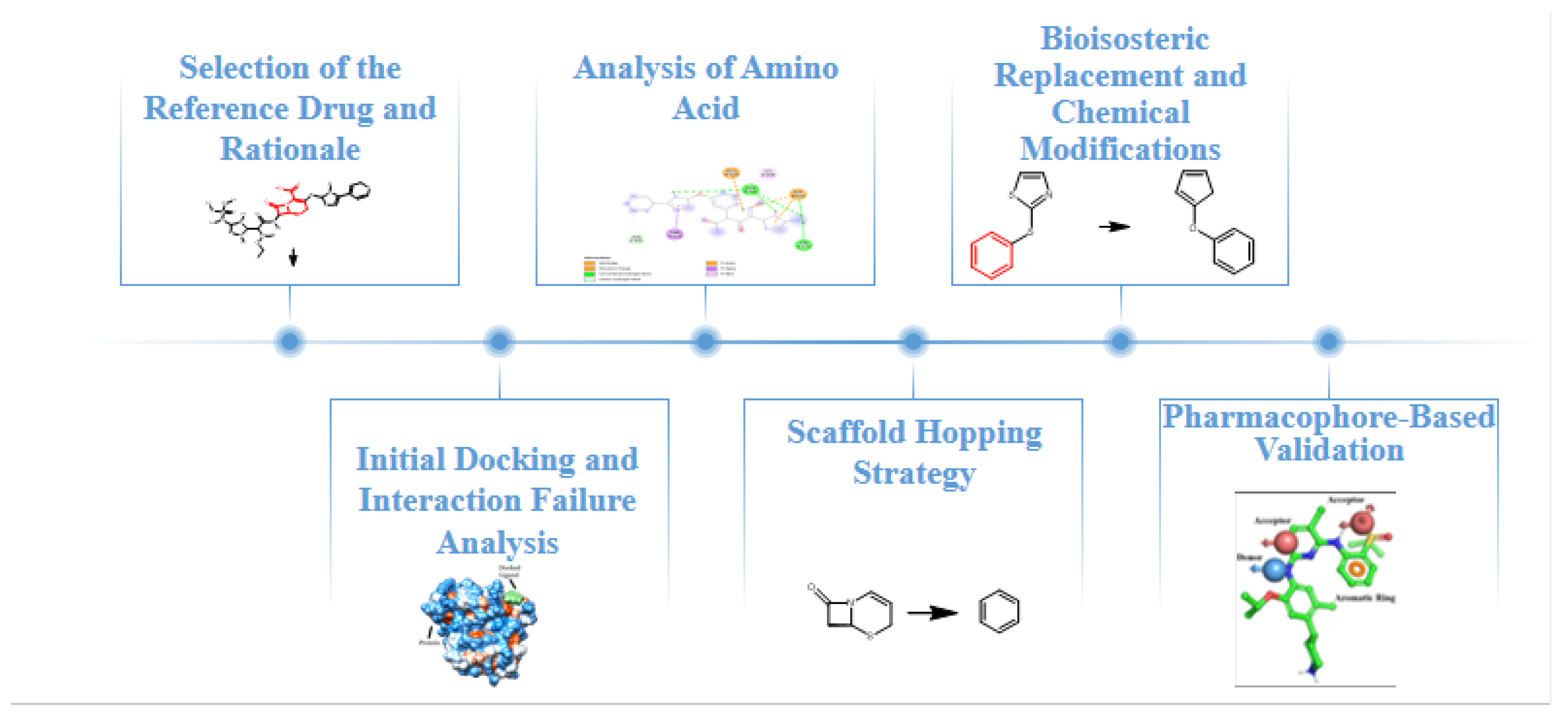
2.1. Selection of the Reference Drug and Rationale

2.2. Initial Docking and Interaction Failure Analysis
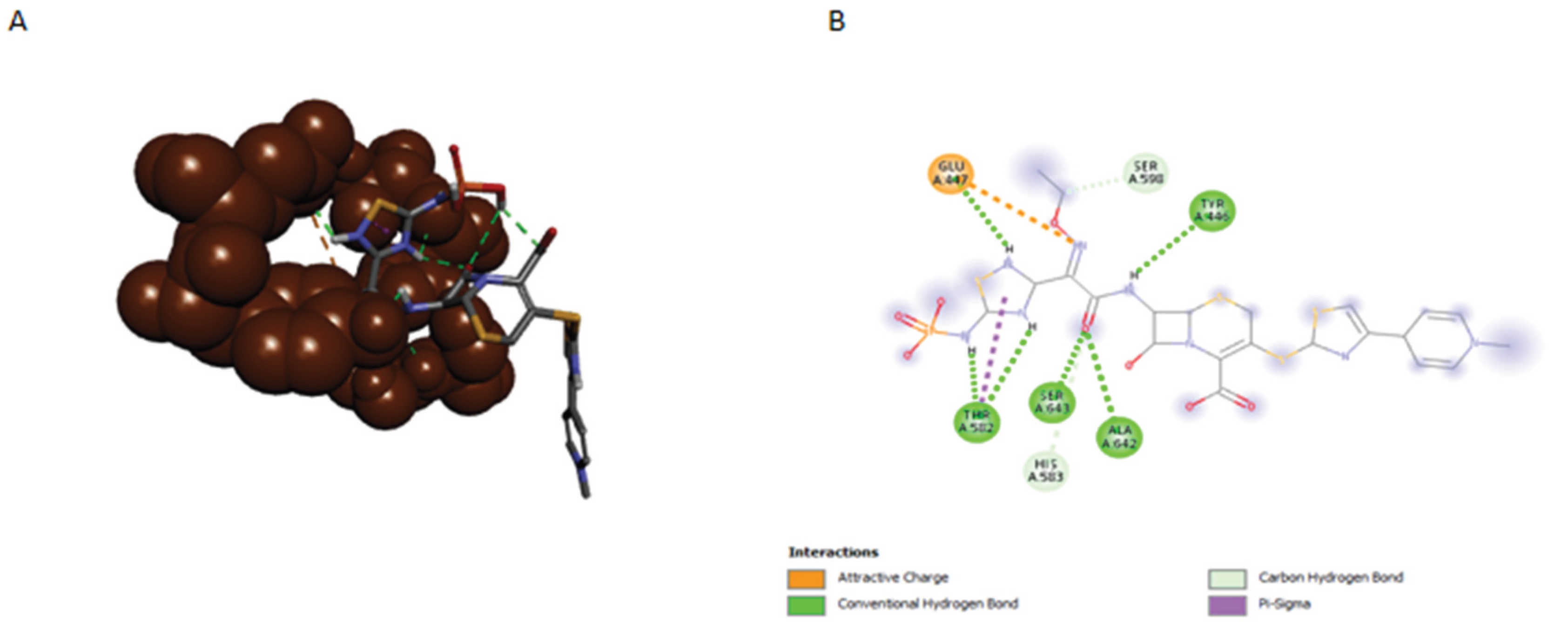
2.3. Analysis of Amino Acid Environment and Targetable Changes
2.4. Scaffold Hopping Strategy, Bioisosteric Replacement and Chemical Modifications
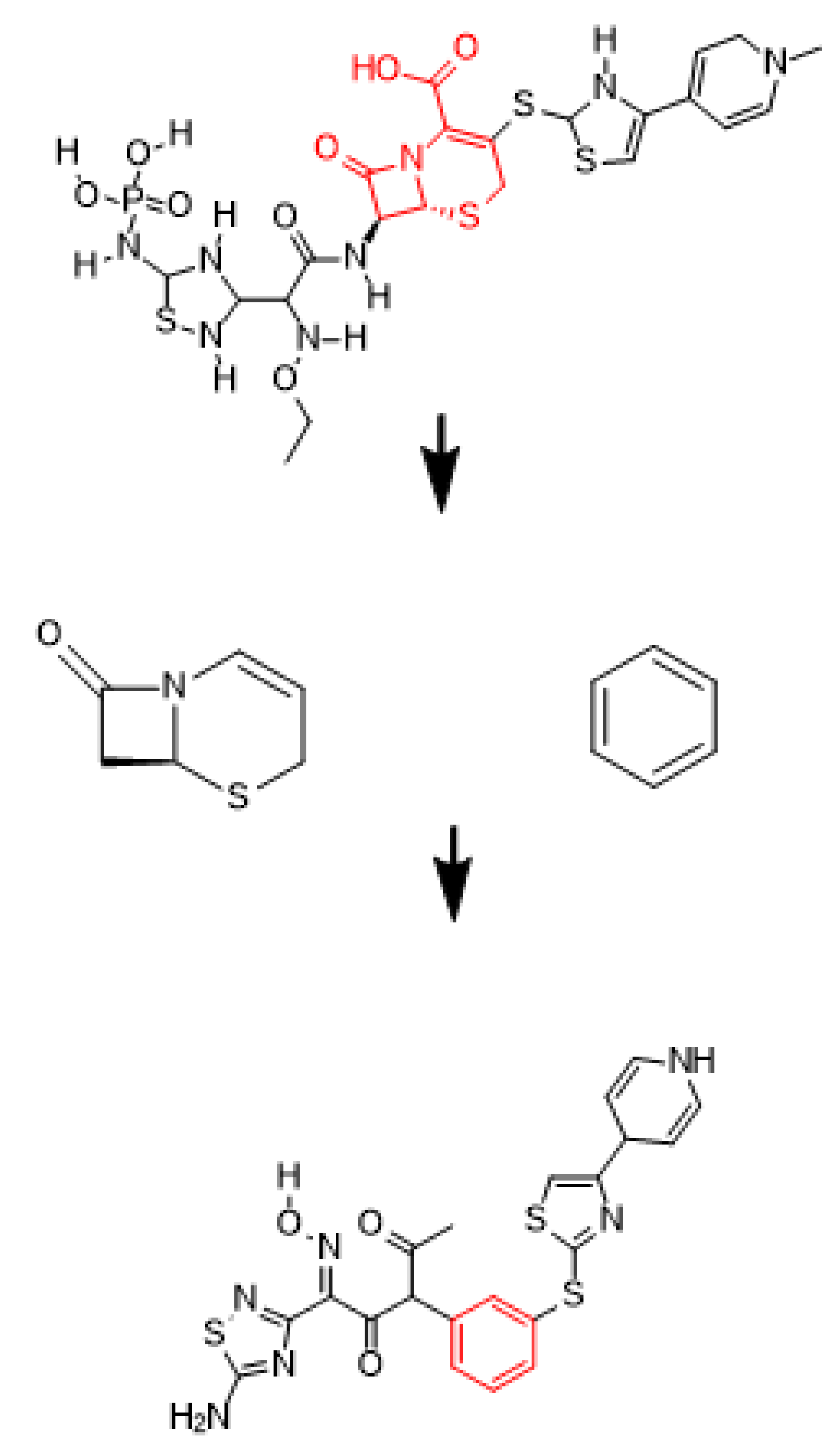
2.5. Bioisosteric Replacement and Chemical Modifications
| Ligand | Binding Affinity | rmsd/ub | rmsd/lb |
| CEFTAROLINE_Docking pose 1 | -7.9 | 0 | 0 |
| Folo drug analouge 1 _ pose 1 | -8.2 | 0 | 0 |
| Folo drug analouge 2 _ pose 1 | -8.4 | 0 | 0 |
| Folo drug analouge 3 _ pose 1 | -6.9 | 0 | 0 |
| Interaction Type | Analogue 1 | Analogue 2 | Analogue 3 |
| π–π stacking | 1 present | 1 present | 1 present |
| Hydrogen bonding | 1 present | 3 present | 2 present |
| Salt bridge | Absent | 1 present | 1 present |
| Electrostatic attraction | Minor clash | 1 present | 1 present |
| Peripheral residue contact | Limited | Extensive | Extensive |
2.6. Pharmacophore-Based Validation
3. Results
3.1. Docking Performance of Designed Ligands
| Ligand | Binding Affinity | rmsd/ub | rmsd/lb |
| CEFTAROLINE_Docking pose 1 | -7.9 | 0 | 0 |
| analouge 1 _ pose 1 | -8.2 | 0 | 0 |
| analouge 2 _ pose 1 | -8.4 | 0 | 0 |
| analouge 3 _ pose 1 | -6.9 | 0 | 0 |
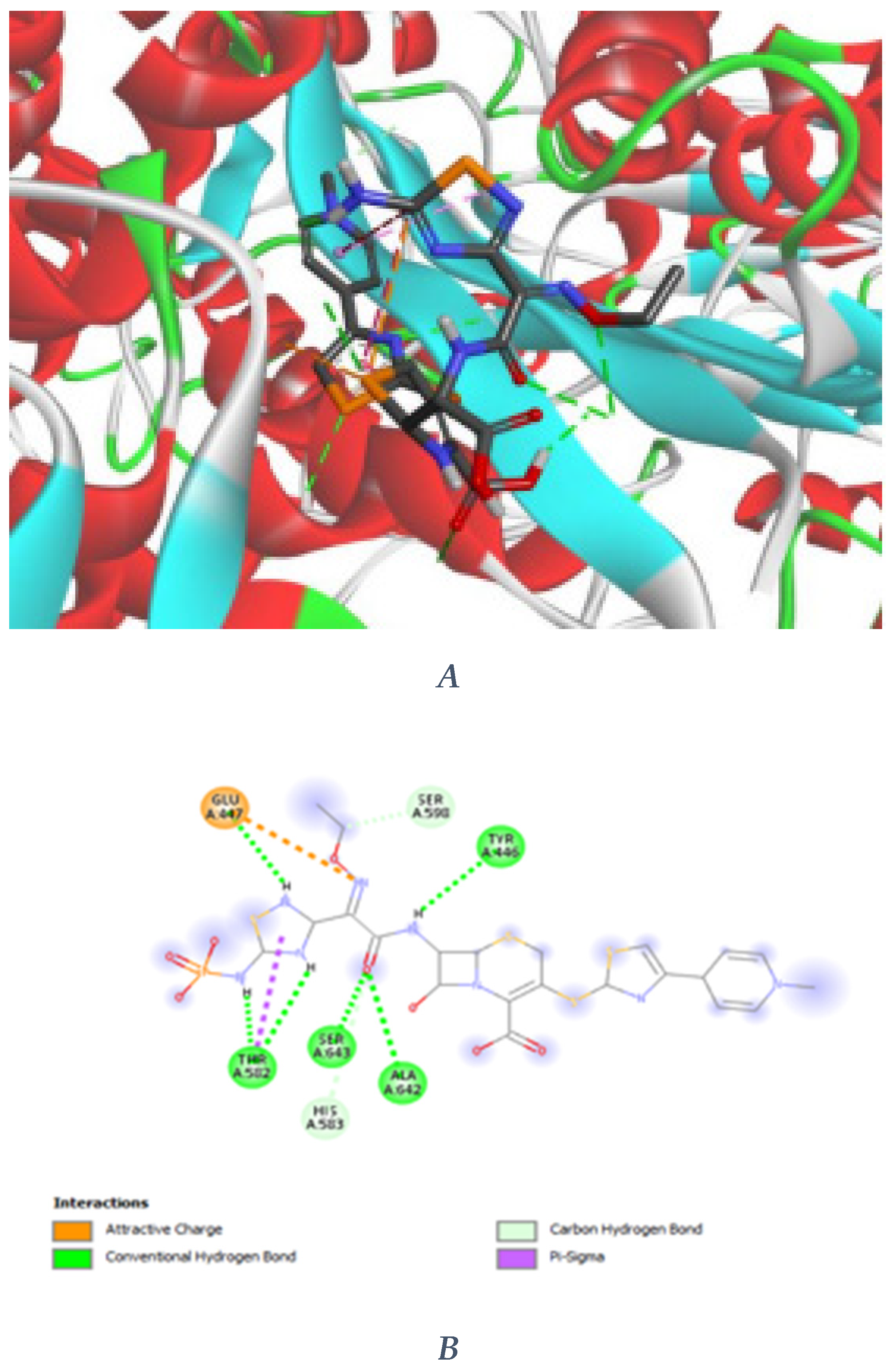
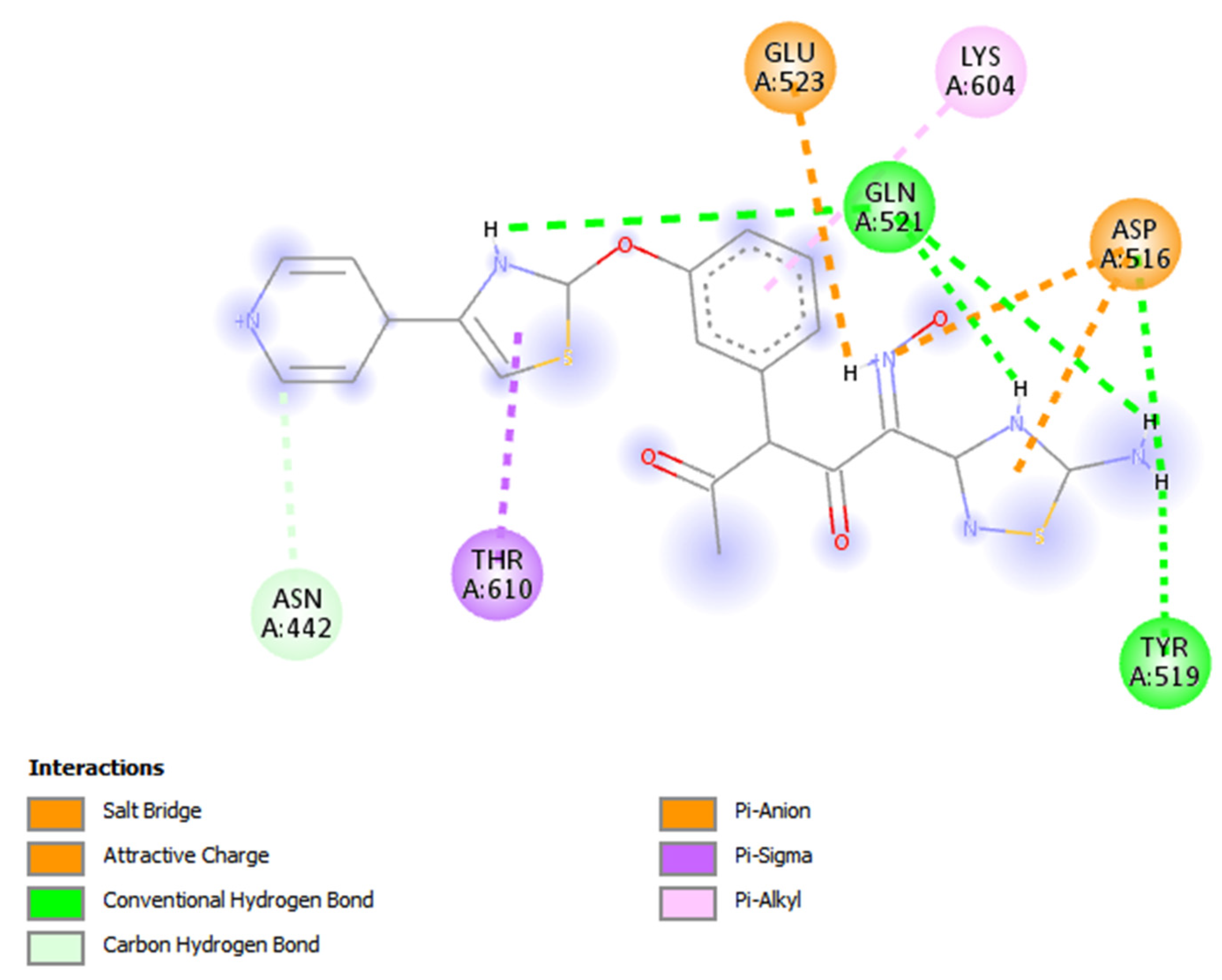
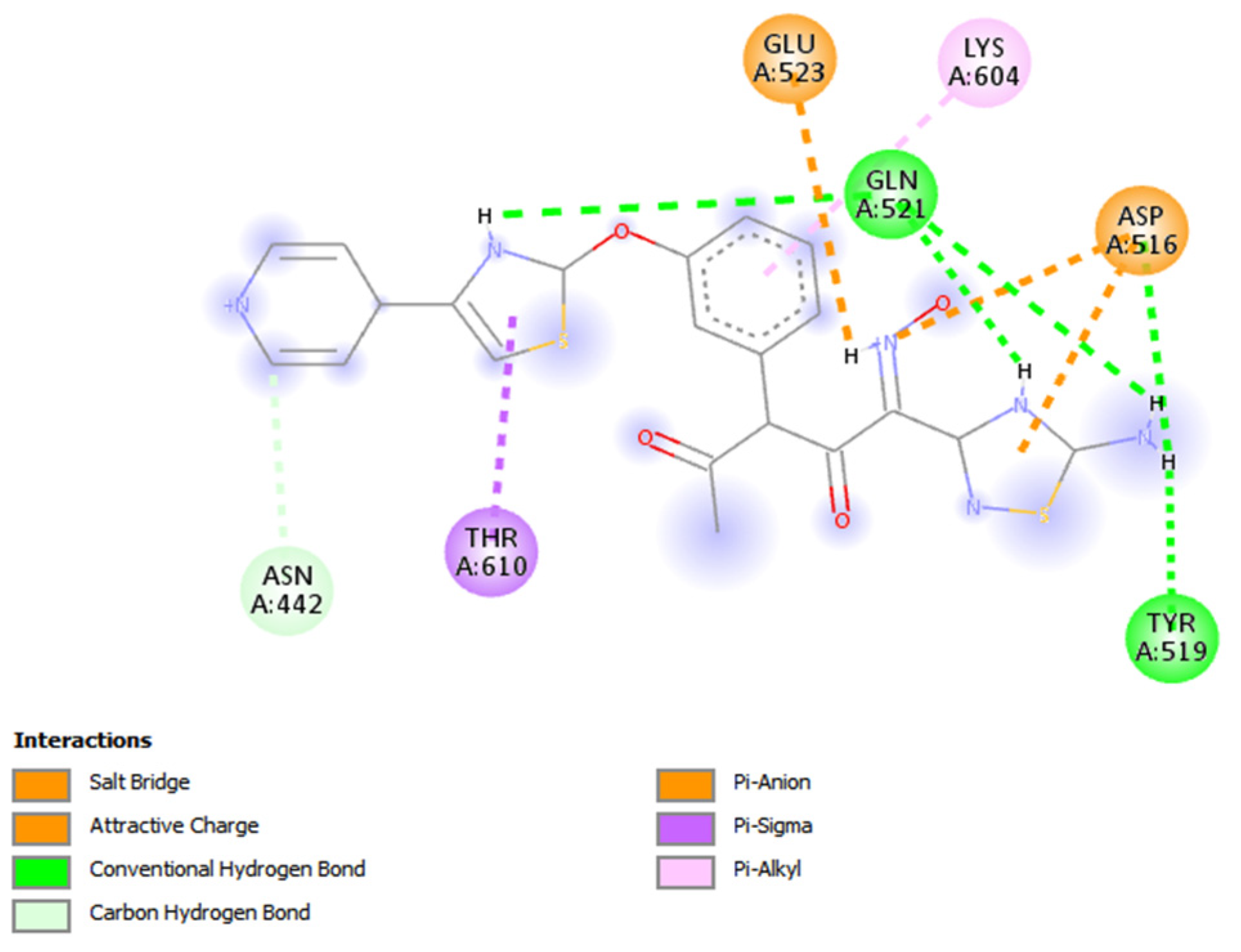
3.2. Novel Interaction Profile of Analogue 2
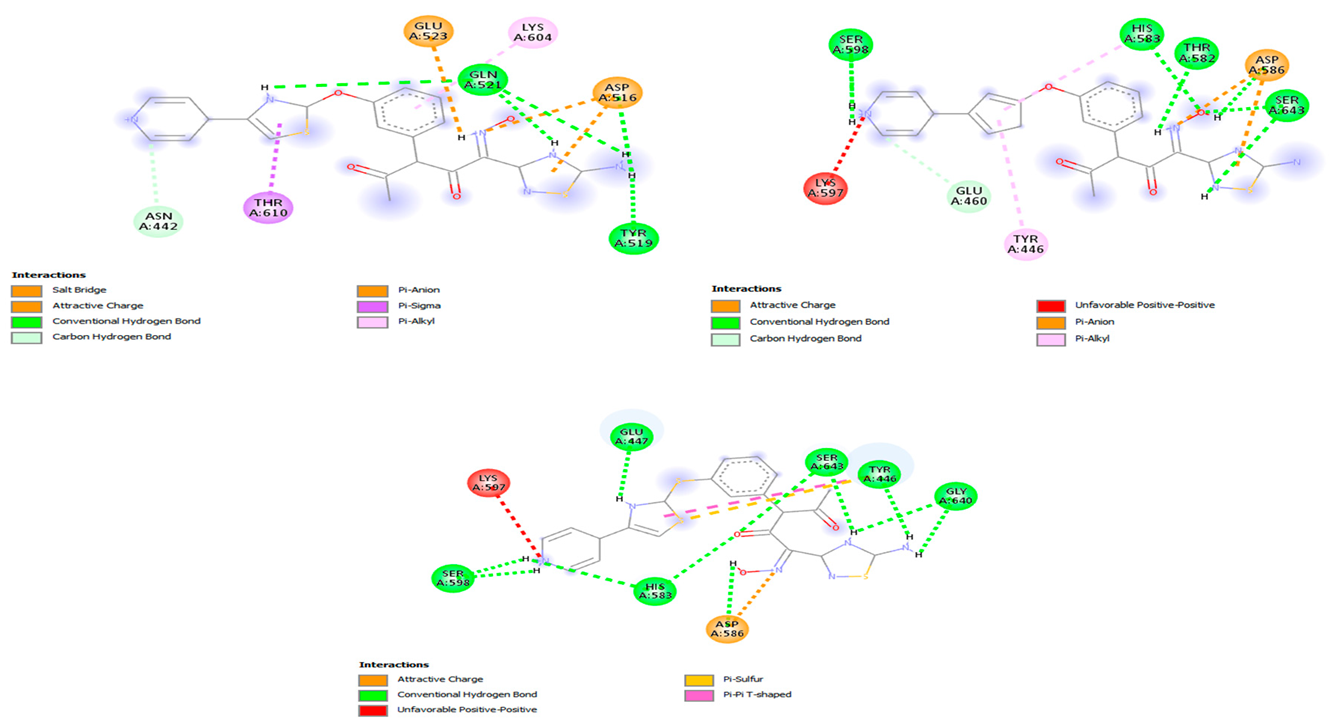
3.3. Comparative Performance of Novel Compounds
4. Discussion
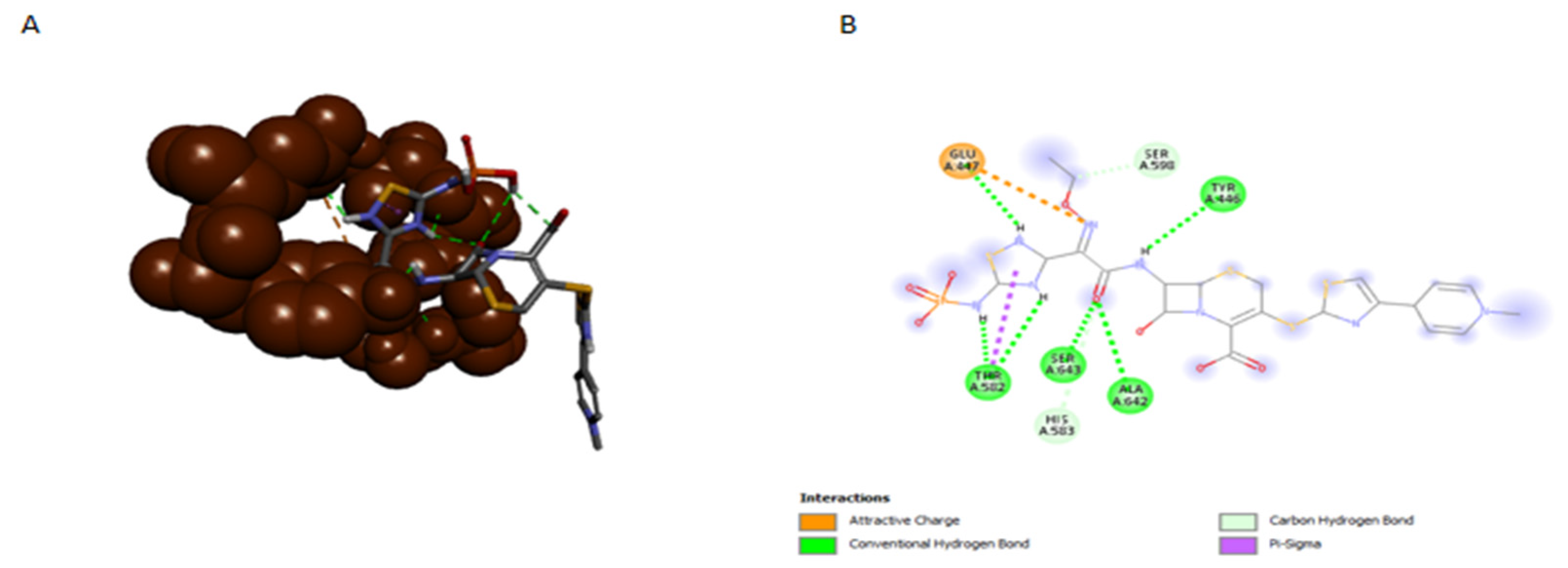
4.1. Confirmation of Allosteric Control Models of PBP2a
4.2. Extension Beyond Existing Non-β-Lactam Studies
4.3. Agreement and Constructive Tension with the Literature
4.4. Methodological Significance and Reproducibility
4.5. Broader Implications and the African Context
4.6. Concluding Perspective
Author Contributions
Funding
Conflicts of Interest
Ethics
References
- Abebe, A.A.; Birhanu, A.G. Methicillin Resistant Staphylococcus aureus: Molecular Mechanisms Underlying Drug Resistance Development and Novel Strategies to Combat. Infect. Drug Resist. 2023, 16, 7641–7662. [Google Scholar] [CrossRef] [PubMed]
- Antimicrobial resistance emerges as a bigger killer in Africa than malaria, HIV or TB [WWW Document], n.d. URL https://www.gavi.org/vaccineswork/antimicrobial-resistance-emerges-bigger-killer-africa-malaria-hiv-or-tb (accessed 3.12.26).
- Chapter 7. Structure-based drug design (SBDD) | Request PDF, 2026; ResearchGate. [CrossRef]
- Fishovitz, J.; Hermoso, J.A.; Chang, M.; Mobashery, S. Penicillin-Binding Protein 2a of Methicillin-Resistant Staphylococcus aureus. IUBMB Life 2014, 66, 572–577. [Google Scholar] [CrossRef] [PubMed]
- Jiao, F.; Bao, Y.; Li, M.; Zhang, Y.; Zhang, F.; Wang, P.; Tao, J.; Tong, H.H.Y.; Guo, J. Unraveling the mechanism of ceftaroline-induced allosteric regulation in penicillin-binding protein 2a: insights for novel antibiotic development against methicillin-resistant Staphylococcus aureus. Antimicrob. Agents Chemother. 2023, 67, e0089523. [Google Scholar] [CrossRef] [PubMed]
- Lin, Y. Bioisosteres and Scaffold Hopping: Strategic Tools in Drug Design 12. 2025. [Google Scholar]
- Loukas, C.; Varytimidis, C.; Rapantzikos, K.; Kanakis, M.A. Keyframe extraction from laparoscopic videos based on visual saliency detection. Comput. Methods Programs Biomed. 2018, 165, 13–23. [Google Scholar] [CrossRef] [PubMed]
- Mahasenan, K.V.; Molina, R.; Bouley, R.; Batuecas, M.T.; Fisher, J.F.; Hermoso, J.A.; Chang, M.; Mobashery, S. Conformational dynamics in penicillin-binding protein 2a of methicillin-resistant Staphylococcus aureus, allosteric communication network and enablement of catalysis. J. Am. Chem. Soc. 2017, 139, 2102–2110. [Google Scholar] [CrossRef] [PubMed]
- (PDF) Computational Screening of Approved Drugs for Inhibition of the Antibiotic Resistance Gene mecA in Methicillin-Resistant Staphylococcus aureus (MRSA) Strains [WWW Document], n.d. URL https://www.researchgate.net/publication/369727549_Computational_Screening_of_Approved_Drugs_for_Inhibition_of_the_Antibiotic_Resistance_Gene_mecA_in_Methicillin-Resistant_Staphylococcus_aureus_MRSA_Strains (accessed 3.12.26).
- Rosado, P.C.; Marques, M.M.; Justino, G.C. Targeting MRSA penicillin-binding protein 2a: structural insights, allosteric mechanisms, and the potential of adjuvant inhibitors. Biochem. Pharmacol. 2025, 239, 117048. [Google Scholar] [CrossRef] [PubMed]
- Zhang, C.; Shang, G.; Gui, X.; Zhang, X.; Bai, X.-C.; Chen, Z.J. Structural basis of STING binding with and phosphorylation by TBK1. Nature 2019, 567, 394–398. [Google Scholar] [CrossRef] [PubMed]
- Zhang, Y.; Luo, M.; Wu, P.; Wu, S.; Lee, T.-Y.; Bai, C. Application of Computational Biology and Artificial Intelligence in Drug Design. Int. J. Mol. Sci. 2022, 23, 13568. [Google Scholar] [CrossRef] [PubMed]
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2026 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).




